Introduction
Lung cancer is a leading cause of cancer-associated
death worldwide (1). Non-small cell
lung cancer (NSCLC) accounts for 80-85% of all lung cancer cases
(2). As the overall survival of
patients with NSCLC remains poor, it is essential to identify novel
treatment targets for NSCLC.
MicroRNAs (miRNAs/miRs) are a class of non-coding
RNAs that lack the ability to code proteins but are capable of
regulating gene expression by binding with the 3'-untranslated
region (3'-UTR) of target mRNA (3).
miRNAs have been reported to serve crucial roles in regulating
numerous cellular processes, including proliferation, metastasis
and differentiation, and they may have dual roles in carcinogenesis
(4-6).
To date, the roles of miR-374a-5p in cancer
progression have remained controversial (7-9).
A previous study determined that the levels of miR-374a-5p, along
with those of miR-195-5p, miR-199a-3p and miR-320a were
significantly increased in patients with osteosarcoma (7). Another study revealed that miR-374a-5p
was also elevated in patients with triple-negative breast cancer
(8). Furthermore, in vitro
and in vivo studies demonstrated that miR-374a-5p was able
to regulate breast cancer cell proliferation and migration by
directly targeting arrestin β1 (ARRB1), indicating an oncogenic
role of miR-374-5p (8). By
contrast, the expression of miR-374a-5p was reduced in esophageal
squamous cell carcinoma and negatively regulated by
LINC00473(9). As the significant
roles of miR-374a-5p in various cancer types, including NSCLC, have
remained elusive, the biological roles of miR-374-5p in NSCLC were
herein investigated.
In the present study, the expression of miR-374a-5p
was determined in NSCLC cell lines and a normal cell line. The
effects of miR-374a-5p on NSCLC cell proliferation and migration
were investigated in vitro. A bioinformatics target
prediction and a luciferase reporter assay were utilized to
identify and confirm the direct binding of miR-374a-5p with NCK
adaptor protein 1 (NCK1). The present results may provide novel
mechanisms underlying the carcinogenesis of NSCLC and a basis for
targeted therapy.
Materials and methods
Cell culture
The NSCLC cell lines A549 and H1299 and the human
bronchial epithelial cell line 16-HBE were purchased from the Cell
Bank of the Chinese Academy of Sciences. These cells were incubated
in RPMI-1640 medium (Invitrogen; Thermo Fisher Scientific, Inc.)
containing 10% fetal bovine serum (Invitrogen; Thermo Fisher
Scientific, Inc.) and 1% Penicillin-Streptomycin at 37˚C in a
humidified incubator containing 5% CO2.
Cell transfection
NSCLC cells were transfected with 50 nM miR-374a-5p
mimics (cat. no. miR10000727-1-5), negative control miRNA (miR-con;
cat. no. miR1N0000001-1-5), 4 µg pcDNA3.1 containing the open
reading frame of NCK1 (pNCK1) or negative control (pcDNA3.1)
purchased from RiboBio with Lipofectamine® 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) at 37˚C. After 48 h
transfection, cells were collected for analysis.
Reverse transcription-quantitative PCR
(RT-qPCR)
RT-qPCR was performed to determine the expression
levels of miR-374a-5p or NCK1 in NSCLC cells and the normal cell
line. In brief, total RNA from cultured cells was isolated with
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.). To detect the expression levels of miR-374a-5p, cDNA was
synthesized from RNA using the TaqMan MicroRNA Reverse
Transcription kit (Applied Biosystems; Thermo Fisher Scientific,
Inc.). For the analysis of NCK1 expression levels, the PrimeScript
kit (Takara Biotechnology, Co., Ltd.) was used to synthesize cDNA
from RNA. RT-qPCR was performed using SYBR Green Mix (Takara
Biotechnology, Co., Ltd.) in an ABI 7500 system (Applied
Biosystems; Thermo Fisher Scientific, Inc.) with the following
thermocycling conditions: 95˚C for 10 min, followed by 40 cycles of
94˚C for 45 sec, 55˚C for 30 sec and 72˚C for 1 min. The primers
used were as follows: miR-374a-5p forward,
5'-GCGCGCTTATAATACAACCTGA-3' and reverse, 5'-GTGCAGGGTCCGAGGT-3';
U6 small nuclear (sn)RNA forward, 5'-CTCGCTTCGGCAGCACA-3' and
reverse, 5'-ACGCTTCACGAATTTGCGT-3'; NCK1 forward,
5'-GAACCATCACCTCCACAGTG-3' and reverse,
5'-AATCCCCTTCATGTCCTCTTTC-3'; GAPDH forward,
5'-GAAGGTGAAGGTCGGAGTC-3' and reverse, 5'-GAAGATGGTGATGGGATTTC-3'.
Relative expression levels were calculated using the
2-ΔΔCq method using U6 snRNA or GAPDH as internal
controls (10).
Cell proliferation assay
The cell proliferation rate was analyzed with a Cell
Counting Kit-8 (CCK-8; Takara Biotechnology, Co., Ltd.). Cells were
seeded in 96-well plates at 2x103 cells per well and
incubated at 37˚C for 0, 24, 48 or 72 h. A total of, 10 µl CCK-8
reagent (Beyotime Institute of Biotechnology) was added to each
well, followed by further incubation at 37˚C for 4 h. The
absorbance was measured at 450 nm using a microplate reader.
Wound healing assay
Serum starved cells were seeded in 6-well plates at
a density of 2x105 cells/well and incubated until ~100%
confluence was reached. A sterile pipette tip was used to create a
wound at the cell surface. The cells were then washed with PBS to
remove the cell debris. The migration distance was measured at 0
and 24 h under a light microscope at a magnification of x200.
Bioinformatics analysis
Targets for miR-374a-5p were predicted using
TargetScan (https://www.targetscan.org) and PicTar (http://pictar.mdc-berlin.de/).
Construction of luciferase activity
vectors
According to the results of the bioinformatics
prediction, NCK1 was selected as a putative target for miR-374a-5p.
The wild-type (wt) 3'-UTR containing the binding site for
miR-374a-5p was synthesized by GenScript, cloned into pGL3 vector
(Promega Corporation) and named as NCK1-wt. Mutant (mt) 3'-UTR of
NCK1 was constructed from NCK1-wt using a site-direct mutagenesis
kit (Takara Biotechnology, Co., Ltd.) and named as NCK1-mt.
Dual-luciferase reporter assay
For the luciferase activity assay, NSCLC cells were
co-transfected with NCK1-wt or NCK1-mt and miR-374a-5p mimics or
miR-con using Lipofectamine 2000 with the pGL3 vector described
above. After 48 h of transfection, the cells were lysed to analyze
the relative luciferase activities using a dual-luciferase
detection kit (Promega Corporation) following the manufacturer's
protocol and normalized to Renilla luciferase.
Statistical analysis
GraphPad Prism 5 software (GraphPad Software, Inc.)
was used for data analysis. Values are expressed as the mean ±
standard deviation. Differences between groups were analyzed using
Student's t-test or one-way analysis of variance with Tukey's
post-hoc test. P<0.05 was considered to indicate statistical
significance.
Results
miR-374a-5p is downregulated in
NSCLC
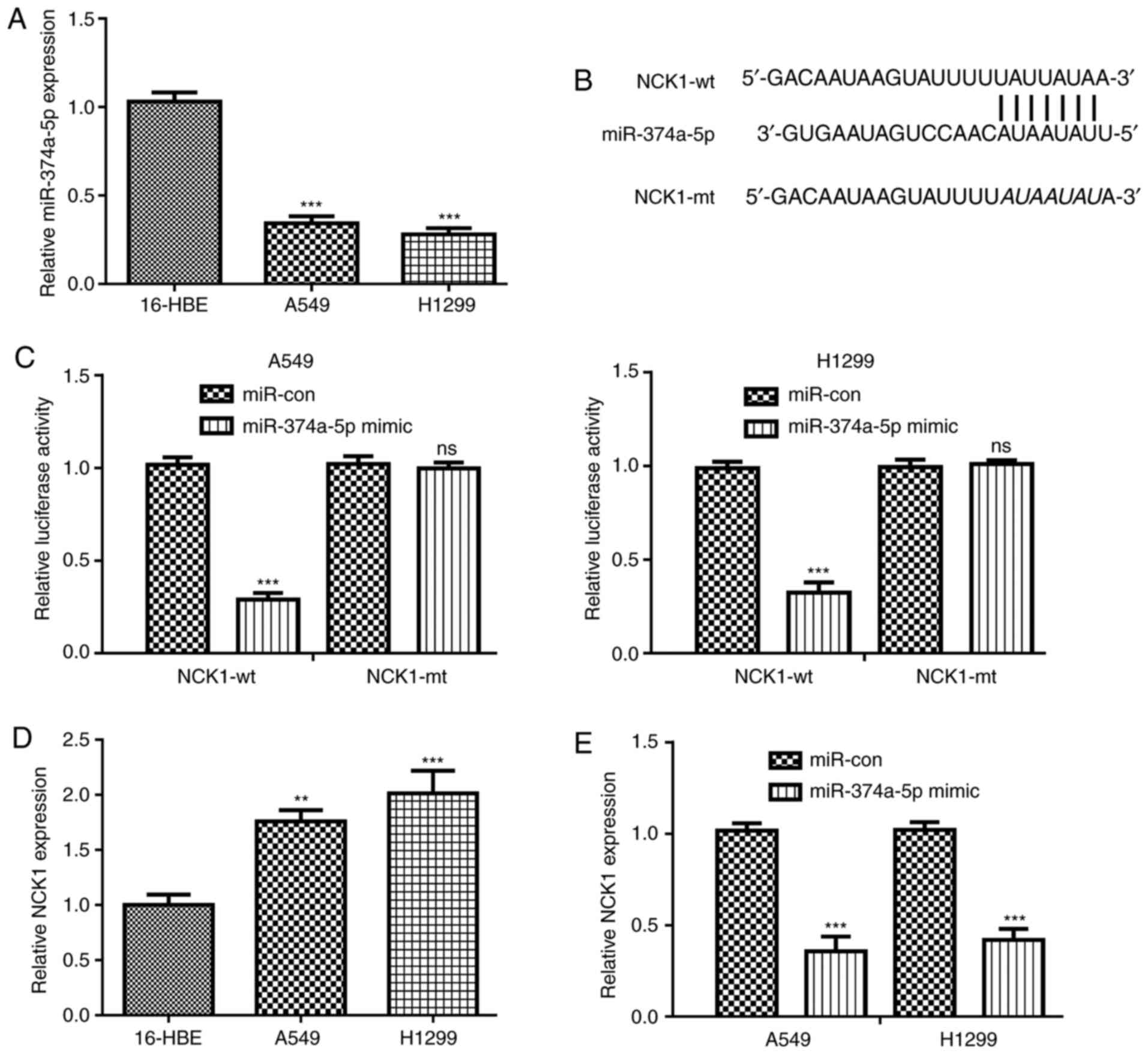
First, miR-374a-5p expression levels in NSCLC cells
were determined by RT-qPCR analysis. The results indicated that
miR-374a-5p expression levels were significantly decreased in NSCLC
cells in comparison with those in the normal cell line (Fig. 1A).
NCK1 is a direct target of miR-374a-5p
in NSCLC
The bioinformatics tools predicted that NCK1
contains a putative binding site for miR-374a-5p (Fig. 1B). To investigate the interaction of
miR-374a-5p and NCK1, luciferase reporter vectors were constructed.
The results indicated that miR-374a-5p overexpression decreased the
luciferase activity of cells transfected with NCK1-wt, but not of
those transfected with NCK1-mt (Fig.
1C). RT-qPCR analysis indicated that NCK1 expression was
increased in NSCLC cells compared with that in the normal cell line
(Fig. 1D). In addition, NCK1
expression was significantly decreased in NSCLC cells transfected
with miR-374a-5p mimics (Fig.
1E).
miR-374a-5p regulates NSCLC cell
proliferation and migration via regulating NCK1
The biological roles of miR-374a-5p and NCK1 in
NSCLC were investigated (Fig. 2).
NSCLC cells were successfully transfected with miR-374a-5p mimics
(Fig. 2A). The results of the CCK-8
assay presented in Fig. 2B
illustrated that overexpression of miR-374a-5p decreased NSCLC cell
proliferation. The wound-healing assay suggested that miR-374a-5p
mimics significantly inhibited the migration ability of NSCLC cells
(Fig. 2C). To test whether
miR-374a-5p regulated NSCLC cell behavior via NCK1, rescue
experiments were performed. The overexpression of NCK1 by pNCK1 was
confirmed by RT-qPCR (Fig. 2D). The
CCK-8 and wound-healing assays indicated that overexpression of
NCK1 promoted NSCLC cell proliferation and migration (Fig. 2B and C, respectively). In addition,
co-transfection of pNCK1 partially abrogated the effects of
miR-374a-5p mimics on NSCLC cell behavior (Fig. 2B and C).
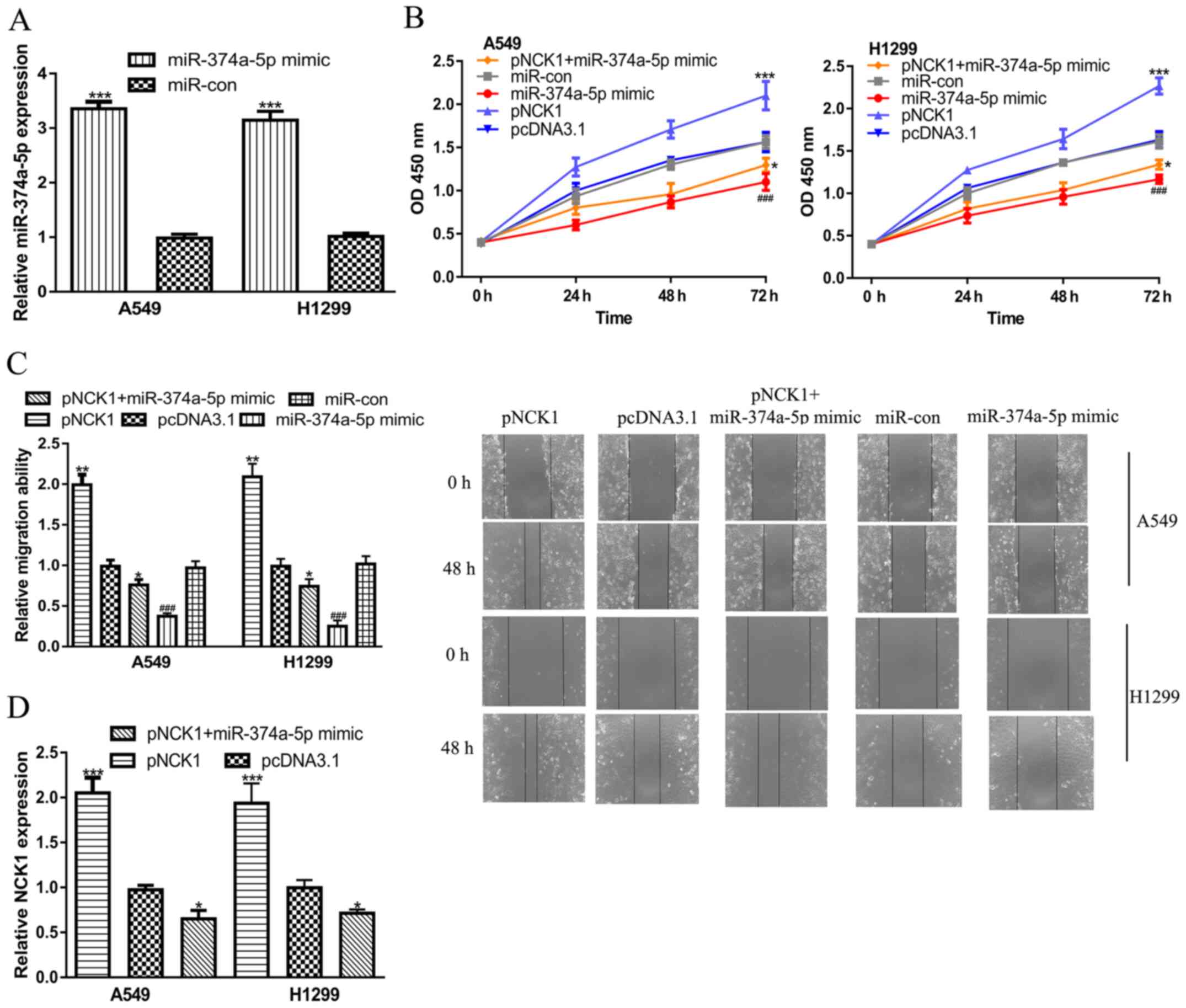 | Figure 2NCK1 reverses the effects of
miR-374a-5p on NSCLC cell behaviors. (A) miR-374a-5p expression in
NSCLC cells with miR-374a-5p mimics or miR-con transfection.
***P<0.001 vs. the miR-con group. (B) Cell
proliferation was assessed. ***P<0.001 vs. the
pcDNA3.1 group, *P<0.05 vs. the pcDNA3.1 group and
###P<0.001 vs. the miR-con group. (C) cell migration
(magnification, x40) was determined. **P<0.01 vs. the
pcDNA3.1 group, *P<0.05 vs. the pcDNA3.1 group and
###P<0.001 vs. the miR-con group. (D) NCK1 expression
of NSCLC cells transfected with miR-374a-5p mimics, miR-con, pNCK1,
pcDNA3.1 or pNCK1 and miR-374a-5p mimics is presented.
***P<0.001 and *P<0.05 vs. the pcDNA3.1
group. NCK1, NCK adaptor protein 1; pNCK1, NCK1 overexpression
plasmid; pcDNA3.1, empty vector; miR-374a-5p, microRNA-374a-5p;
NSCLC, non-small cell lung cancer; miR-con, negative control miR;
OD, optical density. |
Discussion
miRNAs may function as either oncogenic or tumor
suppressor miRNAs in cancer (11).
To date, numerous miRNAs have been indicated to regulate NSCLC
progression. Among these, miR-199a-5p was indicated to be markedly
decreased in NSCLC tissues (12).
In addition, miR-199a-5p overexpression inhibited NSCLC cell
proliferation by causing cell cycle arrest at the G1 phase via
targeting mitogen-activated kinase kinase kinase 11(12). Furthermore, miR-221-3p expression
was determined to be elevated in NSCLC tissues and cell lines
(13). Overexpression of miR-221-3p
was also indicated to promote cell cycle progression of NSCLC cells
via targeting p27(13). Increased
expression of miR-421 was identified in NSCLC tissues and cells,
and its overexpression was able to promote cancer cell migration
and invasion through HOP homeobox via the Wnt/β-catenin signaling
pathway (14).
To the best of our knowledge, the present study was
the first to reveal that miR-374a-5p was significantly
downregulated in NSCLC cell lines compared with those in a normal
cell line. Increased miR-374a-5p expression was also indicated to
inhibit NSCLC cell proliferation and migration in vitro.
Collectively, the present results suggested a tumor suppressor role
of miR-374-5p in NSCLC, which is consistent with its role in
esophageal squamous cell carcinoma (9).
miR-374a-5p exerts its effects in cancers by
regulating the expression of tumor-specific genes (8,9). For
instance, ARRB1 and spindlin 1 were identified as direct targets
for miR-374a-5p in different cancer types (8,9).
However, the targets of miR-374a-5p in NSCLC remain to be verified.
By utilizing TargetScan and PicTar, NCK1 was identified as a
putative target for miR-374a-5p. NCK1 is a protein that has been
reported to be closely associated with cancer progression (15,16).
For instance, NCK1 was determined to be a downstream effector of
STAT3 and to promote colorectal cancer cell metastasis and
angiogenesis via activating the p21-activated serine-threonine
kinase (PAK1)/ERK pathway (15).
Furthermore, NCK1 was indicated to promote cervical squamous
carcinoma cell angiogenesis via the Rac1/PAK1/matrix
metalloproteinase 2 signaling pathway (16). In the present study, the molecular
mechanisms of the effects of miR-374a-5p on NSCLC cell behaviors
were investigated. Rescue experiments were performed, which
demonstrated that NCK1 overexpression partially reversed the
effects of miR-374a-5p. These results indicated that miR-374a-5p
exerted a tumor-suppressive role through NCK1 in NSCLC. However, as
a limitation of the present study, the signaling pathways involved
in the regulatory effect of miR-374a-5p on NSCLC cell behaviors
were not further investigated.
In conclusion, the present study revealed the
reduced expression status of miR-374a-5p in NSCLC and validated
that miR-374a-5p functions as a tumor suppressor to regulate NSCLC
progression via targeting NCK1. The study provided evidence to
unravel a novel miR-374-5p and NCK1 axis in NSCLC, which may help
to develop targeted therapies for NSCLC.
Acknowledgements
Not applicable.
Funding
Funding: This project was supported by Pre-clinical
Transformation Research of panel, a Multigene Molecular Marker for
Early Diagnosis of Lung Adenocarcinoma (‘Office-School Consultation
Fund-Science and Technology Innovation’ Project of Hebei Science
and Technology Department-Hebei Medical University; grant. no.
2020TXZH04).
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
QG, HW, YX, MW and ZT were involved in designing the
study, performed the experiments and interpreted the data. QG and
ZT confirmed the authenticity of raw data, and drafted and revised
the manuscript. All authors read and approved the final version of
the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
3
|
He L and Hannon GJ: MicroRNAs: Small RNAs
with a big role in gene regulation. Nat Rev Genet. 5:522–531.
2004.PubMed/NCBI View
Article : Google Scholar
|
|
4
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297.
2004.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Lages E, Ipas H, Guttin A, Nesr H, Berger
F and Issartel JP: MicroRNAs: Molecular features and role in
cancer. Front Biosci (Landmark Ed). 17:2508–2540. 2012.PubMed/NCBI View
Article : Google Scholar
|
|
6
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006.PubMed/NCBI View
Article : Google Scholar
|
|
7
|
Lian F, Cui Y, Zhou C, Gao K and Wu L:
Identification of a plasma four-microRNA panel as potential
noninvasive biomarker for osteosarcoma. PLoS One.
10(e0121499)2015.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Son D, Kim Y, Lim S, Kang HG, Kim DH, Park
JW, Cheong W, Kong HK, Han W, Park WY, et al: miR-374a-5p promotes
tumor progression by targeting ARRB1 in triple negative breast
cancer. Cancer Lett. 454:224–233. 2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Chen W, Zhang Y, Wang H, Pan T, Zhang Y
and Li C: LINC00473/miR-374a-5p regulates esophageal squamous cell
carcinoma via targeting SPIN1 to weaken the effect of radiotherapy.
J Cell Biochem. 120:14562–14572. 2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Zhang B, Pan X, Cobb GP and Anderson TA:
MicroRNAs as oncogenes and tumor suppressors. Dev Biol. 302:1–12.
2007.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Li Y, Wang D, Li X, Shao Y, He Y, Yu H and
Ma Z: MiR-199a-5p suppresses non-small cell lung cancer via
targeting MAP3K11. J Cancer. 10:2472–2479. 2019.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Yin G, Zhang B and Li J: miR-221-3p
promotes the cell growth of non-small cell lung cancer by targeting
p27. Mol Med Rep. 20:604–612. 2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Liang H, Wang C, Gao K, Li J and Jia R:
ΜicroRNA-421 promotes the progression of non-small cell lung cancer
by targeting HOPX and regulating the Wnt/β-catenin signaling
pathway. Mol Med Rep. 20:151–161. 2019.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Zhang F, Lu YX, Chen Q, Zou HM, Zhang JM,
Hu YH, Li XM, Zhang WJ, Zhang W, Lin C and Li XN: Identification of
NCK1 as a novel downstream effector of STAT3 in colorectal cancer
metastasis and angiogenesis. Cell Signal. 36:67–78. 2017.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Xia P, Huang M, Zhang Y, Xiong X, Yan M,
Xiong X, Yu W and Song E: NCK1 promotes the angiogenesis of
cervical squamous carcinoma via Rac1/PAK1/MMP2 signal pathway.
Gynecol Oncol. 152:387–395. 2019.PubMed/NCBI View Article : Google Scholar
|