Introduction
Fetal hemoglobin (α2γ2; also
termed HbF) to adult Hb (α2β2; also termed
HbA) switching is usually completed within one year after birth and
results in a normal Hb profile in adulthood, comprising HbF
(<1%), HbA2 (α2δ2; 2-3%) and
HbA (≥95%) (1). This obscure scheme
is controlled by many zinc finger transcription factors, such as
Krüppel-like factor 1 (KLF1) (2),
B-cell lymphoma/leukemia 11A (BCL11A) (3) and leukemia/lymphoma-related factor
(LRF) (4). KLF1 is known as an
erythroid-specific transcription factor that orchestrates the
expression of hundreds of erythroid essential genes, including
fetal and adult globin genes (2,5). KLF1
regulates fetal to adult Hb switching by direct activation of
β-globin (HBB) gene expression and indirect repression of
the γ-globin (HBG) gene by stimulating the expression of
BCL11A and LRF, two major HBG gene repressors (6,7).
Furthermore, the docking of KLF1 on the CACCC core box within
HBB, BCL11A and LRF promoters was mentioned as
an initial step of gene expression (6,7).
Downregulation of KLF1 reduces BCL11A expression and further
increases HBG gene and HbF production (2,7). In
addition, KLF1 occupies the LRF promoter in human erythroid
cells and activates LRF gene expression (6). KLF1 is categorized as a member of the
KLF family. To date, 18 KLF genes have been described in
human chromosomes (8,9). KLF members are characterized by three
Cys2His2 zinc finger DNA binding domains at
the C-terminus, and variable transactivating domains at the
N-terminus. The binding domains of KLFs recognize specific DNA
motifs containing GC-rich or CACCC boxes, whereas the
transactivating domains harness transcriptional levels of the
target genes upon which co-activators or co-repressors are
recruited (10). Notably, the CACCC
motif has been identified as a common regulatory element in
erythroid gene promoters, particularly in globin gene promoters
(6,11,12).
Furthermore, KLF1 amino acid sequences have the
highest homology with KLF2 and KLF4(13). KLF2 (or lung KLF) regulates
embryonic globin gene expression and is essential for erythroid
maturation within the yolk sac in transgenic mice (14,15).
KLF4 (or gut KLF) binds to the CACCC motif within the HBG
gene promoter and activates HBG gene expression in early
erythroid progenitor cells from adults (16). Additionally, KLF4 is able to dock on
the α-globin (HBA) gene promoter and stimulate the gene
expression in the human chronic myeloid leukemia (K562) cell line
(17). In zebrafish, klf4
acts as an embryonic globin gene suppressor; however, the
arrangement of globin genes in zebrafish differs from that of
humans (18).
The present study proposed that additional KLF
members may be crucial for human globin gene regulation in
erythroid cells. To ascertain a potential for KLF in exploiting
globin gene regulation, the mRNA expression of 18 KLFs in
erythroid progenitor cells from fetal liver and adult peripheral
blood samples were screened for their relative differences in terms
of the globin gene expression status at specific developmental
stages. KLF with the greatest decrease of mRNA expression in
the fetus was chosen for a functional study using RNA interference
in adult peripheral blood-derived erythroid progenitor cells.
Effects of KLF knockdown during in vitro
erythropoiesis were assessed. Reactivation of embryonic or fetal
globin gene expression after the KLF knockdown was
investigated. These changes in globin gene expression can be
exploited for potential therapeutic implications for severe
thalassemias in which a specific globin gene was insufficient.
Materials and methods
Samples
Existing cDNA samples from a previous microarray
study (19) were used as templates
for investigating KLF expression patterns in the present
study. The cDNA samples were prepared from the RNA of hematopoietic
progenitor cells from fetal liver samples (n=3) and adult
peripheral blood samples (n=3). Whole blood samples were drawn from
six adults who were aged ≥18 years, consisting of healthy
volunteers (50 ml; n=3) and patients with
β0-thalassemia/HbE (25 ml; n=3) at Faculty of Allied
Health Sciences, Thammasat University (Pathum Thani, Thailand) in
June 2020 after obtaining written informed consent. Anticoagulant
Citrate Dextrose Solution, Solution A (Terumo Bct, Inc.) was used
as a preservative. Thalassemia and other hemoglobinopathies were
ruled out in the healthy subjects. The most common deletional
α-thalassemia mutations, including-α3.7,
-α4.2, --SEA and --THAI were
screened for exclusion, and HBB genes were genotyped in the
subjects with β0-thalassemia/HbE. At the time of
sampling, the patients had not received any blood transfusions for
≥1 month before venipuncture. All hematological parameters and Hb
typing were obtained from an automated blood cell analyzer (ADVIA
2120; Bayer AG) and the VARIANT II β-thalassemia Short Program
(Bio-Rad Laboratories, Inc.), respectively. The hematological data
of the subjects are presented in Table
SI.
Erythroblast culture
Peripheral blood mononuclear cells (PBMCs) were
immediately harvested from the blood samples using Lymphoprep
(Axis-Shield). Hematopoietic stem cells or progenitor cells
(CD34+) were purified from the PBMCs using a CD34
MicroBead Kit (Miltenyi Biotec GmbH) with a magnetic activated cell
sorting column according to the manufacturer's instructions. The
isolated CD34+ cells were placed in the 3-phase
erythroblast culture system where the cells were able to grow,
expand and differentiate into erythroid cells. Briefly, Iscove's
modified Dulbecco's medium (Biochrom GmbH) supplemented with 20%
v/v fetal bovine serum (FBS; EMD Millipore), 300 µg/ml
holo-transferrin (ProSpec-Tany TechnoGene Ltd.) and 1% v/v
penicillin/streptomycin (Gibco; Thermo Fisher Scientific, Inc.) was
prepared as a basal medium. In phase 1 (days 0-4), the basal medium
was augmented with 10 ng/ml human interleukin-3 (IL-3; Miltenyi
Biotec GmbH), 50 ng/ml human stem cell factor (SCF; Miltenyi Biotec
GmbH) and 2 U/ml erythropoietin (EPO; Janssen-Cilag Ltd.). In phase
2 (days 4-8), the basal medium was fortified by 10 ng/ml SCF and 2
U/ml EPO. In phase 3 (days 8-14), the basal medium was only
supplemented with 4 U/ml EPO. As the number of isolated
CD34+ cells varied among individuals, cell numbers were
reserved at 1-2x106 cells/ml during the culture. The
cells were incubated at 37˚C in a 5% CO2 and 100%
humidity atmosphere.
KLF4 knockdown
KLF4sh1 (cat. no. TRCN0000231078) and KLF4sh2 (cat.
no. TRCN0000231079) short hairpin (sh)RNA sequences were obtained
from the MISSION® shRNA library (Sigma-Aldrich; Merck
KGaA; https://www.sigmaaldrich.com/catalog/genes/KLF4?lang=en®ion=GB#shRNA%20Products)
and cloned into the third-generation lentiviral vector,
pLL3.7-puro, containing the mouse U6 promoter for driving shRNA
expression and the puromycin resistance gene as a selectable
marker. pLL3.7-puro was in-house modified from pLL3.7 (cat. no.
11795; Addgene, Inc.) by replacing the EGFP gene with the puromycin
resistant gene. The 293T cells (cat. no. CRL-3216; American Type
Culture Collection) were cultured in Dulbecco's modified Eagle's
medium (Gibco; Thermo Fisher Scientific, Inc.) supplemented with
10% v/v FBS and 2 mM L-glutamine (Gibco; Thermo Fisher Scientific,
Inc.). When the cells reached ~80% confluency, lentiviruses
expressing different shRNAs targeting KLF4 mRNA were
produced by co-transfecting the expression constructs with
packaging plasmids, including 2.5 µg of pMD2.G (cat. no. 12259;
Addgene, Inc.), 3.75 µg of pMDLg/pRRE (cat. no. 12251; Addgene,
Inc.) and 3.75 µg of pRSV-Rev (cat. no. 12253; Addgene, Inc.) into
the 293T cells using the X-tremeGENE™ HP Transfection
Reagent (Roche Diagnostics GmbH). Supernatants were collected at 48
and 72 h after transfection and filtered through a 0.45-µm
membrane. The filtrates were concentrated using a Lenti-X
Concentrator (Clontech Laboratories, Inc.) and centrifuged at 4˚C,
1,500 x g for 1 h. Lentiviral titers were measured by transducing
the viruses into 293T cells in the presence of 4.0 µg/ml polybrene
(EMD Millipore) and were subsequently challenged by 2.0 µg/ml
puromycin (Invitrogen; Thermo Fisher Scientific, Inc.) at 48 h
post-transduction. A non-targeting control shRNA sequence (cat. no.
SHC016V; EMD Millipore) served as a negative control (shNTC).
Erythroid cells from day 4 of culture were
transduced overnight with the lentiviruses at a multiplicity of
infection of 20 in the phase 2 medium supplemented with 8.0 µg/ml
polybrene. Transduced cells were treated with 0.5 µg/ml puromycin
at 48 h post-transduction and were consequently placed in the phase
3 medium at 48 h after puromycin selection.
Erythroid differentiation and
morphology
Cultured cells (5x104 cells) were
harvested on day 12 and resuspended in Dulbecco's
phosphate-buffered saline (HyClone; Cytiva). Erythroid
differentiation was surveilled using antibodies against two
erythroid-specific surface markers, phycoerythrin-conjugated mouse
monoclonal anti-human CD71 (cat. no. CY1G4; BioLegend, Inc.) and
allophycocyanin-conjugated mouse monoclonal anti-human CD235a (cat.
no. GA-R2; BD Biosciences). Flow cytometry was achieved on BD
Accuri™ C6 Plus (BD Biosciences). Data were analyzed
using FlowJo version 10.3.0 (FlowJo LLC). To investigate erythroid
cell morphology, cultured cells (5x104 cells) were fixed
with absolute methanol and subsequently stained with modified
Giemsa stain (MilliporeSigma) according to the manufacturer's
instructions. The stained cells were examined under a light
microscope.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from cultured cells
(1x106 cells) on day 10 using the TRIzol®
Reagent (Ambion; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. The extracted RNA samples (1 µg each)
were treated with DNase I (Thermo Fisher Scientific, Inc.) to
remove DNA contamination and were reverse transcribed into cDNAs
using a RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher
Scientific, Inc.) according to the manufacturer's instructions.
RT-qPCR was performed in duplicate using gene-specific primers, as
presented in Table SII and
FastStart™ Essential DNA Green Master (Roche Diagnostics
GmbH) according to the manufacturer's instructions. qPCR was
performed and analyzed using a CFX96 Real-Time System (Bio-Rad
Laboratories, Inc.) with pre-incubation at 95˚C for 5 min, followed
by 35 cycles of denaturation at 95˚C for 10 sec, annealing at 60˚C
for 10 sec and 72˚C for 10 sec. All target gene expression levels
were normalized to β-actin (ACTB) mRNA expression levels.
Relative gene expression data were calculated using the
2-ΔΔCq method (20).
Western blot analysis
Nuclear and cytoplasmic proteins were extracted from
a pellet of at least 5x106 cultured cells on day 10
using NE-PER™ Nuclear and Cytoplasmic Extraction
Reagents (Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. The protein concentration was
determined using the Quick Start™ Bradford Protein Assay
(Bio-Rad Laboratories, Inc.) according to the manufacturer's
instructions. A total of 10 µg protein was run on 10%
SDS-polyacrylamide gel, transferred to a polyvinylidene fluoride
membrane and subsequently blocked at room temperature for 1 h with
5% skimmed milk in PBS supplemented with 0.05% Tween-20.
Immunoblotting was performed by incubating the membranes with
specific antibodies against their target proteins (Table SIII) at room temperature for 1 h.
The membrane was washed three times for 10 min each with blocking
buffer. HRP-conjugated secondary antibodies were then added and
incubated at room temperature for 1 h before signal development.
Chemiluminescent detection was conducted with Amersham™
ECL™ Prime Western Blotting Detection Reagent (Cytiva)
according to the manufacturer's instructions.
Hb typing
Hemolysates were prepared from a minimum of
1x106 cultured cells on day 14 and subjected to
high-performance liquid chromatography for Hb type analysis using
the Bio-Rad VARIANT II Hemoglobin Testing System with β-Thalassemia
Short Program (Bio-Rad Laboratories, Inc.) according to the
manufacturer's protocols.
Statistical analysis
Statistical analysis was performed using SPSS
version 26.0.0.0 (IBM Corp.). Unpaired student's t-test and one-way
ANOVA with Tukey's post hoc test were performed to identify
significant differences. Data are presented as means ± SEM and
P<0.05 was considered to indicate a statistically significant
difference.
Results
KLF4 mRNA expression levels decrease
in erythroid progenitor cells from fetal liver samples
Our previous study conducted microarray analysis
using Affymetrix GeneChip® Human Gene 2.0 ST Arrays to
differentiate KLF expression patterns in erythroid
progenitor cells from fetal liver and adult peripheral blood
samples (19). According to the
microarray data, KLF4 and KLF9 emerged as the top
candidate genes due to their greater downregulation (0.79-fold;
P=0.06495) and upregulation (1.91-fold; P=0.00628), respectively,
in fetuses when compared with adults (Table SIV). The mRNA expression levels of
18 human KLFs, along with the key HbF regulators,
BCL11A and LRF, and eight globin genes were
subsequently validated by RT-qPCR using the existing cDNA samples
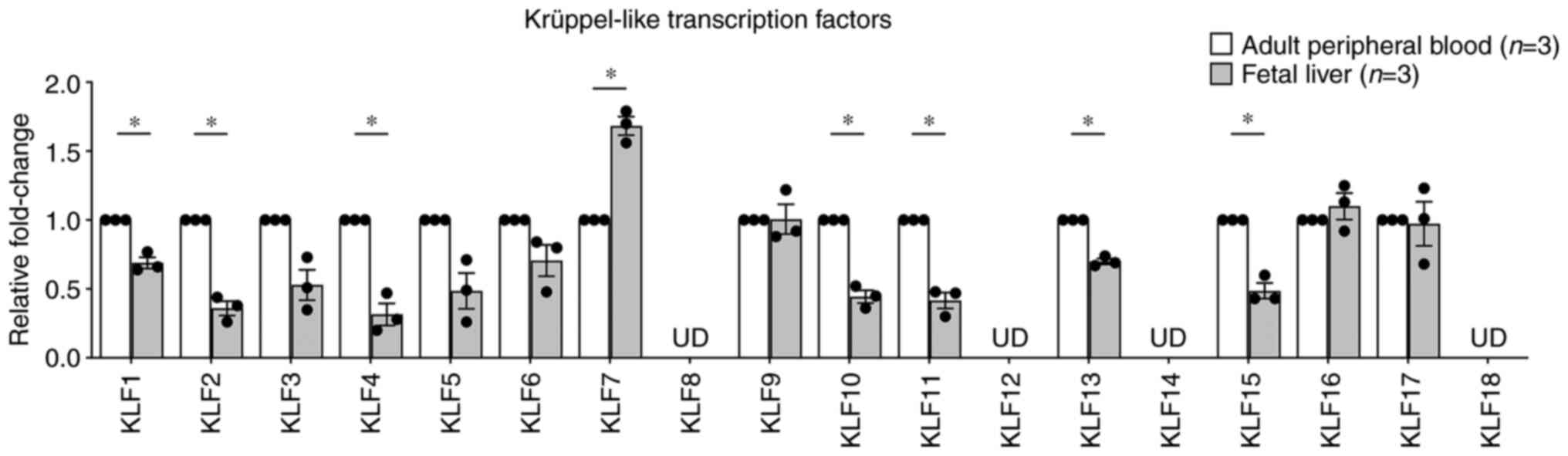
obtained from our previous study (19). The results demonstrated that
KLF4 mRNA expression was significantly downregulated
(0.32±0.08-fold; P=0.02413; Fig.
1), which was similar to that observed from the microarray data
(Table SIV), whereas the
upregulation of KLF9 mRNA expression was not detected by
RT-qPCR. Instead, the mRNA expression of KLF7 was
significantly upregulated (1.68±0.07-fold; P=0.03000; Fig. 1). However, certain KLF
expression patterns obtained from RT-qPCR, for example,
undetectable levels of KLF8, KLF12 and KLF14
(Fig. 1) were inconsistent with the
results from the microarray. The discrepancy of the association
between results of the microarray and RT-qPCR could be due to
different platform conditions. The mRNA expression levels of the
two well-characterized HbF regulators, BCL11A and LRF
were also analyzed for their function as essential HBG
repressors (3,4). Although the results demonstrated that
the two were relatively decreased in fetuses when compared with the
adults, only the mRNA expression level of BCL11A was
significantly downregulated in the fetuses (Fig. S1). These results indicated that
KLF4 mRNA expression might be developmentally regulated and
associated with changes in fetal globin expression, concomitant
with the BCL11A mRNA expression level.
KLF4 knockdown increases embryonic and
fetal globin gene expression
To determine the effects of KLF4 on globin
expression, the expression levels of eight human globin genes were
examined using RT-qPCR following KLF4 knockdown in human erythroid
progenitor cells obtained from healthy individuals and donors with
compound heterozygous β0-thalassemia/HbE, which exhibit
high baseline HbF levels. KLF4 expression was more pronounced upon
erythroid maturation in the primary erythroid culture system
(Fig. S2). KLF4sh1 and KLF4sh2
reduced the KLF4 expression levels by >0.5-fold when compared
with shNTC. Furthermore, downregulation of KLF4 did not
significantly affect the three known HbF suppressors, KLF1, BCL11A
and LRF (Fig. 2A and B). The results revealed that
downregulation of KLF4 in hematopoietic progenitor cells from
healthy individuals and donors with β0-thalassemia/HbE
evidently upregulated ζ-globin (HBZ), ε-globin (HBE)
and moderately increased HBG mRNA expression when compared
with shNTC in the two subject groups (Fig. 2C). This alteration was similar to
that of globin mRNA expression patterns in hematopoietic progenitor
cells from fetal liver samples (Fig.
S1). In addition, marginally decreased expression levels of
µ-globin (HBM), α-globin (HBA), θ-globin
(HBQ), δ-globin (HBD) and HBB genes were
observed when compared with the control (Fig. 2C). The increase of HbF (Δ%HbF)
executed by experimental KLF4sh1 knockdown in healthy and
β0-thalassemia/HbE erythroid cells was 2.0±0.8 and
2.7±1.4%, respectively, above the control. The Δ%HbF observed by
experimental KLF4sh2 knockdown in healthy and
β0-thalassemia/HbE erythroid cells was 1.5±1.0 and
5.0±1.3%, respectively, above the control (Fig. S3). These changes in HbF levels were
consistent with the marginally increased HBG mRNA expression
observed in Fig. 2C.
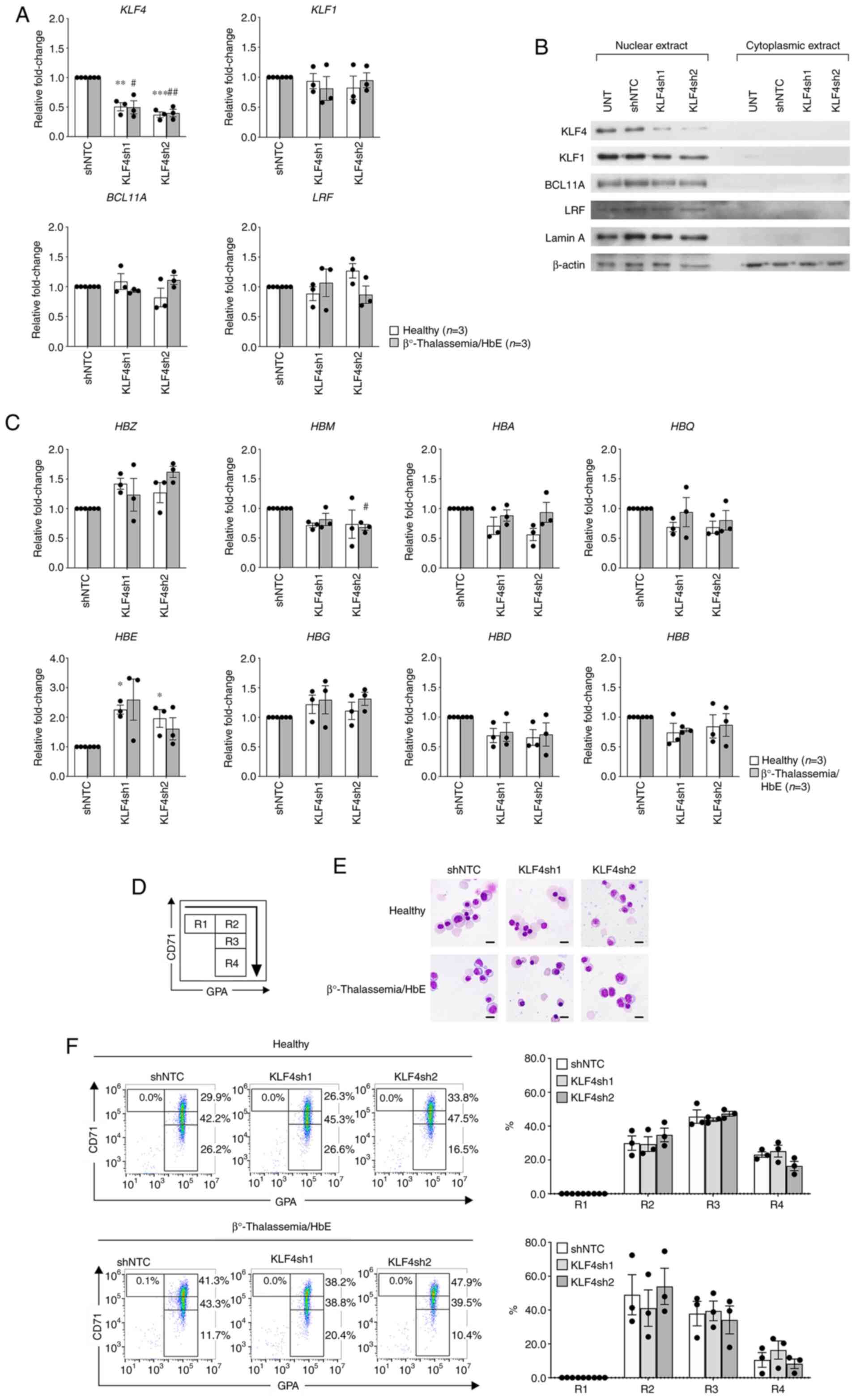 | Figure 2Effects of KLF4 knockdown in
erythroid progenitor cells from healthy individuals and patients
with β0-thalassemia/HbE. (A) RT-qPCR demonstrating KLF4
knockdown efficiency and effects on KLF1, BCL11A and
LRF mRNA expression levels. (B) Representative western blot
analysis showing KLF4 knockdown efficiency and effects on KLF1,
BCL11A and LRF protein expression levels. Lamin A and β-actin
served as loading controls for nuclear and cytoplasmic protein
origin, respectively. (C) RT-qPCR displaying relative fold-change
of α-like globin (upper panel) and β-like globin (lower panel) mRNA
expression levels. Relative fold-change represents the mRNA
expression levels normalized to β-actin of shRNAs targeting KLF4
(KLF4sh1 and KLF4sh2) vs. shNTC. Data are presented as the means ±
SEM of healthy individuals (n=3; *P<0.05,
**P<0.005, ***P<0.0005) and patients
with β0-thalassemia/HbE (n=3; #P<0.05,
##P<0.005). (D) Illustrative flow cytometric analysis
on the basis of transferrin receptor, CD71 and GPA expression
levels. Where R1=CD71high/GPAlow;
R2=CD71high/GPAhigh;
R3=CD71medium/GPAhigh; and
R4=CD71low/GPAhigh. (E) Representative
cytospin preparations of modified Giemsa-stained cells on day 12
visualized under a light microscope (x1,000 magnification; scale
bar, 10 µm). (F) Representative flow cytometric analysis of
experimental KLF4 knockdown. The bar graph represents proportions
of each cell population. KLF, Krüppel-like factor; HbE, hemoglobin
E; RT-qPCR, reverse transcription-quantitative PCR; BCL11A, B-cell
lymphoma/leukemia 11A; LRF, leukemia/lymphoma-related factor;
shRNA, short hairpin RNA; shNTC, non-targeting control shRNA; GPA,
glycophorin A; HBZ, ζ-globin; HBM, µ-globin;
HBA, α-globin; HBQ, θ-globin; HBE, ε-globin;
HBG, γ-globin; HBD, δ-globin; HBB,
β-globin. |
The effects of KLF4 knockdown on erythroid cell
differentiation and maturation were assessed by evaluating the
expression levels of two erythroid-specific surface markers, CD71
and glycophorin A (GPA), and by investigating erythroid morphology
under a light microscope. Erythroid differentiation and gating
proportions were analyzed based on the expression levels of CD71
and GPA, as follows: R1, CD71high/GPAlow; R2,
CD71high/GPAhigh; R3,
CD71medium/GPAhigh; and R4,
CD71low/GPAhigh (Fig. 2D). In the culture system,
β0-thalassemia/HbE-derived erythroblasts demonstrated
delayed erythroid differentiation when compared with the healthy
erythroblasts (Fig. 2E and F). Downregulation of KLF4 in healthy
erythroblasts using KLF4sh1 did not affect erythroid
differentiation or maturation. However, KLF4sh2 knockdown in the
healthy group insignificantly delayed erythroid differentiation as
demonstrated by the reduction in the R4 portion when compared with
the controls (Fig. 2F). Similar
findings of the fairly delayed differentiation were also observed
in the β0-thalassemia/HbE-derived erythroblasts with
KLF4sh2 knockdowns, in which the majority of the cell populations
retained in R2 (Fig. 2F).
Early-stage erythroid morphology was found in experimental KLF4sh2
knockdown in both healthy and β0-thalassemia/HbE-derived
erythroblasts when compared with the controls (Fig. 2E), confirming a slightly delayed
erythroid maturation. Notably, modest promotion of cell
differentiation was apparent in
β0-thalassemia/HbE-derived erythroblasts upon KLF4sh1
knockdown as evidenced by the increase of R4 when compared with the
controls (Fig. 2F). Following
KLF4sh1 knockdown, erythroid morphology accelerated erythroid
maturation in β0-thalassemia/HbE-derived erythroblasts
when compared with the controls (Fig.
2E). However, this acceleration was not particularly noticeable
in healthy subjects under the same circumstances.
Discussion
Hb switching is specific to human developmental
stages, resulting in the production of different Hb molecules
during ontogeny (1). However, the
mechanisms underlying the switch remain unclear. KLFs are
transcription factors classified by highly conserved three
Cys2His2 zinc finger DNA binding domains at
the C terminus (8-10).
KLFs bind to a specific CACCC DNA motif, which are commonly found
within human globin gene promoters (7,11,12).
KLF1 is a member of the KLF family, regulating fetal to adult Hb
switching by activating the expression of the major HBG
repressor, BCL11A. Furthermore, KLF1 can dock on the CACCC boxes
within the HBB gene promoter and can further activate
HBB gene expression. Notably, the amino acid sequences of
KLF1 are markedly similar to those of KLF2 and KLF4(13). This similarity may indicate
comparable functions between KLF1, KLF2 and KLF4 in relation to
globin gene regulation.
In the present study, the mRNA expression pattern of
18 human KLFs in fetal and adult erythroblasts were
investigated using RT-qPCR. The expression of KLF4 was
revealed to be the most significantly downregulated in erythroid
progenitor cells from the fetal liver when compared with those from
adults. The result was consistent with our previously reported
microarray data (19). This
indicates that KLF4 might be developmentally controlled.
Furthermore, the lower mRNA expression level of KLF4 was
concomitant with the greater mRNA expression levels of embryonic
and fetal globin, including HBZ, HBE and HBG.
This result revealed the association between the modest expression
of the KLF4 mRNA level and the pronounced mRNA expression
levels of embryonic and fetal globin in erythroid progenitor cells
from the fetus. However, it was found that the mRNA levels of two
major HBG gene repressors, KLF1 and BCL11A
were significantly downregulated in the fetus group when compared
with the adult group. Therefore, the elevation of fetal globin mRNA
expression in the present study might be due to the reduction of
these two transcription factors.
To elaborate on the function of KLF4 on globin
expression regulation at diverse HbF baseline levels, lentiviral
shRNA-mediated KLF4 knockdowns were performed in erythroid
progenitor cells from healthy adults and patients with
β0-thalassemia/HbE. The results demonstrated that
experimental KLF4 knockdown could induce embryonic
(HBZ and HBE) and fetal (HBG) globin mRNA
expression in healthy subjects and those with
β0-thalassemia/HbE. Moreover, a marginal surge in the
HbF level also corresponded with the modest elevated HBG
mRNA expression. However, these findings were inconsistent with
previous studies showing that KLF4 triggers HBG and
HBA gene expression in human erythroid cells and the K562
cell line, respectively (16,17).
This may be due to the disparate erythroid cell stage during
experimental KLF4 knockdown and the dissimilar erythroid culture
system. Notably, downregulation of KLF4 did not significantly
affect the expression levels of well-known fetal globin gene
repressors, KLF1, BCL11A and LRF, suggesting a possible independent
globin regulation mechanism. Although KLF4 knockdown revealed
insignificant effects on erythroid cell maturation, delayed
erythroid differentiation was observed in subjects with
β0-thalassemia/HbE when compared with the healthy
subjects. To date, the effort to induce embryonic HBZ
expression in patients with α-thalassemia major (ATM) has been
proposed as a novel therapy to improve the phenotype and survival
rate of patients with ATM (21).
Therefore, manipulations of KLF4 expression using chemical
compounds or molecular approaches may also be considered as
potential therapeutic strategies for HBA substitution via
HBZ induction in those patients.
Thus, the present study revealed that the
KLF4 mRNA expression level is relatively downregulated in
hematopoietic progenitor cells from the fetal liver. Additionally,
experimental knockdown of KLF4 promoted embryonic and fetal globin
mRNA expression in human hematopoietic progenitor cells. Despite a
small sample size, this evidenced the inverse association between
KLF4 and embryonic and fetal globin mRNA expression in human
erythroid progenitor cells. Moreover, the development of drugs or
molecular methods that reduce KLF4 and further increase embryonic
globin gene expression for adult globin substitution may provide an
alternative treatment for severe α-thalassemia.
Supplementary Material
Relative mRNA expression profile in
erythroid progenitor cells from FL and AB samples as examined by
reverse transcription-quantitative PCR. (A) The relative mRNA
expression of two major fetal hemoglobin repressors, BCL11A
and LRF. The relative mRNA expression of (B) four α-like
globin genes, HBZ, HBM, HBA and HBQ and
(C) four β-like globin genes, HBE, HBG, HBD
and HBB. Gene expression levels were normalized to β-actin.
Data are presented as the means ± SEM of relative fold-change of FL
(n=3) relative to AB (n=3). *P<0.05, **P<0.005 and
****P<0.0001. FL, fetal liver; AB, adult peripheral blood;
BCL11A, B-cell lymphoma/leukemia 11A; LRF,
leukemia/lymphoma-related factor; HBZ, ζ-globin; HBM,
μ-globin; HBA, α-globin; HBQ, θ-globin; HBE,
ϵ-globin; HBG, γ-globin; HBD, δ-globin; HBB,
β-globin.
Time-course analysis of gene
expression during erythroid differentiation. (A) Reverse
transcription-quantitative PCR analysis for KLF4,
KLF1, BCL11A and LRF mRNA expression levels
upon time intervals of erythroid differentiation. Gene expression
levels were normalized to β-actin. Data are presented as the means
of relative fold-change of day 8 ± SEM of the healthy (n=3) and β
0-thalassemia/HbE (n=3) groups. (B) Representative nuclear protein
as evaluated by western blot analysis for KLF4, KLF1, BCL11A and
LRF during erythroid differentiation of a healthy subject. Lamin A
served as a loading control. KLF, Krüppel-like factor; HbE,
hemoglobin E; BCL11A, B-cell lymphoma/leukemia 11A; LRF,
leukemia/lymphoma-related factor; D, day.
Hemoglobin composition after KLF4
knockdown. Representative high-performance liquid chromatography
chromatograms from cultured cells on day 14. The bar graph
represents the increase in the percentage of HbF (Δ%HbF) in
KLF4-knockdown cells from the baseline level in shNTC-treated
cells. Data are presented as means ± SEM in the healthy (n=3) and
β0-thalassemia/HbE (n=3) groups. KLF, Krüppel-like factor; HbF,
fetal hemoglobin; KLF4sh, shRNAs targeting KLF4; shRNA, short
hairpin RNA; shNTC, non-targeting control.
Hematological data of subjects from
the present study.
Primers used for reverse
transcription-quantitative PCR in the present study.
Antibodies used for western blot
analysis in the present study.
KLF gene expression in hematopoietic
progenitor cells from FL (n=3) and AB (n=3) analyzed from previous
microarray data (18).
Acknowledgements
The authors would like to thank Miss Thongperm
Munkongdee, Miss Nattrika Buasuwan and Miss Nurmeeha Hinna,
Thalassemia Research Center, Institute of Molecular Biosciences,
Mahidol University (Nakhon Pathom, Thailand) for their assistance
with evaluating DNA for thalassemia and hemoglobin analyses. The
authors would also like to thank Associate Professor Alisa Tubsuwan
(Stem Cell Research Unit, Institute of Molecular Biosciences,
Mahidol University) who provided resources for lentivirus
production.
Funding
Funding: The present study was supported by a grant from the
Faculty of Allied Health Sciences, Thammasat University (grant no.
1/2562; Pathum Thani).
Availability of data and materials
The datasets used and/or analyzed during the
current study are available from the corresponding author on
reasonable request.
Authors' contributions
PK conceptualized the idea, designed the research,
performed experiments, analyzed data and wrote the manuscript. NJ
performed erythroid culture and supplied resources. AT and SH
provided microarray data, analyzed microarray data and supplied
cDNA samples. PK and AT confirm the authenticity of all the raw
data. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
The study procedures were approved in accordance
with compliance to the Declaration of Helsinki, The Belmont Report,
Council for International Organizations of Medical Sciences
guidelines and the International Conference on Harmonisation-Good
Clinical Practice by the Ethical Review Sub-Committee Board for
Human Research Involving Sciences (Thammasat University, Pathum
Thani, Thailand; approval nos. 198/2561 and 031/2563). In our
previous study (19), women who
underwent pregnancy termination due to medical complication
provided written informed consent to donate their fetus for
research purposes. The protocols for the previous study were
approved in accordance with the Declaration of Helsinki by the
Committee on Human Rights related to research involving human
subjects (Faculty of Medicine, Ramathibodi Hospital, Mahidol
University, Bangkok, Thailand; Institutional Review Board no.
MURA2013/363).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Weatherall DJ and Clegg JB: The
Thalassaemia Syndromes. 4th edition. Blackwell Science, Oxford,
2008.
|
|
2
|
Zhou D, Liu K, Sun CW, Pawlik KM and
Townes TM: KLF1 regulates BCL11A expression and gamma- to
beta-globin gene switching. Nat Genet. 42:742–744. 2010.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Sankaran VG, Menne TF, Xu J, Akie TE,
Lettre G, Van Handel B, Mikkola HK, Hirschhorn JN, Cantor AB and
Orkin SH: Human fetal hemoglobin expression is regulated by the
developmental stage-specific repressor BCL11A. Science.
322:1839–1842. 2008.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Masuda T, Wang X, Maeda M, Canver MC, Sher
F, Funnell AP, Fisher C, Suciu M, Martyn GE, Norton LJ, et al:
Transcription factors LRF and BCL11A independently repress
expression of fetal hemoglobin. Science. 351:285–289.
2016.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Magor GW, Tallack MR, Gillinder KR, Bell
CC, McCallum N, Williams B and Perkins AC: KLF1-null neonates
display hydrops fetalis and a deranged erythroid transcriptome.
Blood. 125:2405–2417. 2015.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Norton LJ, Funnell APW, Burdach J, Wienert
B, Kurita R, Nakamura Y, Philipsen S, Pearson RCM, Quinlan KGR and
Crossley M: KLF1 directly activates expression of the novel fetal
globin repressor ZBTB7A/LRF in erythroid cells. Blood Adv.
1:685–692. 2017.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Borg J, Papadopoulos P, Georgitsi M,
Gutiérrez L, Grech G, Fanis P, Phylactides M, Verkerk AJ, van der
Spek PJ, Scerri CA, et al: Haploinsufficiency for the erythroid
transcription factor KLF1 causes hereditary persistence of fetal
hemoglobin. Nat Genet. 42:801–805. 2010.PubMed/NCBI View
Article : Google Scholar
|
|
8
|
Pollak NM, Hoffman M, Goldberg IJ and
Drosatos K: Krüppel-like factors: Crippling and un-crippling
metabolic pathways. JACC Basic Transl Sci. 3:132–156.
2018.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Pei J and Grishin NV: A new family of
predicted Krüppel-like factor genes and pseudogenes in placental
mammals. PLoS One. 8(e81109)2013.PubMed/NCBI View Article : Google Scholar
|
|
10
|
McConnell BB and Yang VW: Mammalian
Krüppel-like factors in health and diseases. Physiol Rev.
90:1337–1381. 2010.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Cao A and Moi P: Regulation of the globin
genes. Pediatr Res. 51:415–421. 2002.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Raich N and Romeo PH: Erythroid regulatory
elements. Stem Cells. 11:95–104. 1993.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Chen X, Whitney EM, Gao SY and Yang VW:
Transcriptional profiling of Krüppel-like factor 4 reveals a
function in cell cycle regulation and epithelial differentiation. J
Mol Biol. 326:665–677. 2003.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Basu P, Morris PE, Haar JL, Wani MA,
Lingrel JB, Gaensler KM and Lloyd JA: KLF2 is essential for
primitive erythropoiesis and regulates the human and murine
embryonic beta-like globin genes in vivo. Blood. 106:2566–2571.
2005.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Vinjamur DS, Wade KJ, Mohamad SF, Haar JL,
Sawyer ST and Lloyd JA: Krüppel-like transcription factors KLF1 and
KLF2 have unique and coordinate roles in regulating embryonic
erythroid precursor maturation. Haematologica. 99:1565–1573.
2014.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Kalra IS, Alam MM, Choudhary PK and Pace
BS: Krüppel-like factor 4 activates HBG gene expression in primary
erythroid cells. Br J Haematol. 154:248–259. 2011.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Marini MG, Porcu L, Asunis I, Loi MG,
Ristaldi MS, Porcu S, Ikuta T, Cao A and Moi P: Regulation of the
human HBA genes by KLF4 in erythroid cell lines. Br J Haematol.
149:748–758. 2010.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Gardiner MR, Gongora MM, Grimmond SM and
Perkins AC: A global role for zebrafish klf4 in embryonic
erythropoiesis. Mech Dev. 124:762–774. 2007.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Tangprasittipap A, Kaewprommal P,
Sripichai O, Sathirapongsasuti N, Satirapod C, Shaw PJ,
Piriyapongsa J and Hongeng S: Comparison of gene expression
profiles between human erythroid cells derived from fetal liver and
adult peripheral blood. PeerJ. 6(e5527)2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Gregory GL, Wienert B, Schwab M, Lianoglou
BR, Hollis RP, Kohn DB, Conklin BR and MacKenzie TC: Investigating
zeta globin gene expression to develop a potential therapy for
alpha thalassemia major. Blood. 136 (Suppl 1):S3–S4. 2020.
|