Introduction
Prostate cancer (PC) is becoming one of the leading
male malignancies in a number of countries (1), which also has the highest incidence of
bone metastases among all urological malignancies (2). The high probability of bone metastasis
associated with prostate cancer places the patient at risk of
pathological skeletal events, such as fractures (3). An osteolytic response is characterized
by the destruction of normal bone due to osteoblast inactivation
coupled with osteoclast recruitment and activation in the
tumor-bone microenvironment (4).
Osteolytic lesions are characterized by soft sections of damaged
bone resulting from an osteolytic response that can cause bone pain
and fractures (5).
Prostate cancer cells can produce substantial
amounts of exosomes, which can contain important physiological and
pathological information and can circulate around the human body in
the blood or urine (6,7). However, the specific role of
PC-derived-exosomes in the bone microenvironment remain poorly
understood. Exosomes are extracellular vesicles with lipid bilayer
membranes that are 30-150 nm in diameter and are secreted by cells.
They have been reported to be involved in cell-cell communication,
due to their ability to hold contain DNAs, mRNAs, microRNAs
(miRNAs) and proteins (8,9). In particular, miRNAs are molecules
that can encode information that can be transferred among cells
(10,11). These findings provide evidence that
exosomal miRNAs can be used as biomarkers for diagnosing various
diseases, including cancer (12).
Numerous tumor types, including prostate, lung and breast cancer,
are particularly prone to bone metastasis and maintain substantial
crosstalk with bone cells in the bone microenvironment (13). Colorectal cancer-secreted miR-25-3p
can be transferred to HUVECs via exosomes (14). The promoting effect of exosomes on
transendothelial migration in cancer was previously found to be
positively correlated with miR-23a levels (15). However, it remains unknown which
molecules contained within exosomes secreted by prostate cancer
cells are involved in regulating bone homeostasis.
MicroRNAs (miRNAs) are small noncoding regulatory
RNA molecules that are ~22 nucleotides long and are encoded by
plants, animals and certain types of viruses (16). MiRNAs are considered to be important
regulators of bone metabolism and have differential expression
profiles between patients with cancer and healthy individuals
(17-19).
According to a previous study, miRNAs remain relatively stable in
clinical samples of plasma and serum (17). Circulating miRNAs identified in
human primary osteoporosis, have also been implicated in supporting
the progression of bone metastases in different tumors, such as PC,
breast and non-small-cell lung cancer, esophageal squamous cell
carcinoma and multiple myeloma, suggesting that these miRNAs can
trigger the homing of cancer cells to the bone (19). In particular, miR-148a-3p was
demonstrated to be significantly upregulated in osteoporotic human
samples compared with non-osteoporotic human samples, where the
increased miR-148a levels observed in the serum of patients with
osteoporosis appears to be associated with the aforementioned miRNA
upregulation observed in bone tissue of patients with osteoporosis
(20). The functional role of
miR-148a-3p in the bone suggests that this miRNA may serve as a
viable biomarker for postmenopausal osteoporosis (21,22).
Furthermore, exosomal miR-148a has been reported to serve a
positive regulatory role in osteoclast differentiation, where V-maf
musculoaponeurotic fibrosarcoma oncogene homolog B (MafB) was found
to be the target of miR-148a in osteoclast differentiation
(23). Therefore, MafB was
considered to be the marker of osteoclastic differentiation in the
present study.
The PI3K/AKT/mTOR signaling pathway has been
documented to regulate the migration and invasion of prostate
cancer, which is also mentioned frequently as the signaling pathway
that can be explored in other malignant tumors, such as breast
cancer and B-cell lymphoproliferative disorders (24,25).
Although previous findings found targeting this pathway to be an
option the prevention and treatment of bone metastasis caused by
prostate cancer (26-28),
the potential connection between PC-derived exosomes and the
PI3K/AKT/mTOR signaling pathway remain unclear.
In the present study, the possible function of
exosomes derived from prostate cancer on the bone microenvironment
and MafB expression in osteoclasts was examined. Exosomes isolated
from the prostate cancer cell line PC-3 were cocultured with
pre-osteoclast cells from volunteers. The present study
investigated the inhibitory effect of PC-3-derived exosomes on
osteoblast differentiation, which was detected by western blotting
and reverse transcription-quantitative (RT-q)PCR.
Materials and methods
PC-3 cell culture
The human prostate cancer epithelial cell line PC-3
(cat. no. CRL-1435; American Type Culture Collection) was used in
this study. PC-3 cells were cultured in RPMI-1640 (Gibco; Thermo
Fisher Scientific, Inc.) supplemented with 10% FBS (Biological
Industries), 100 U/ml penicillin and 100 µg/ml streptomycin (Gibco;
Thermo Fisher Scientific, Inc.) and incubated at 37˚C in 5%
CO2. The cell culture medium was changed every 2-3 days.
At 80% confluence, the cell culture medium was replaced by
RPMI-1640 medium with exosome-depleted serum (Thermo Fisher
Scientific, Inc.) for 48 h at 37˚C before the cell culture medium
was collected for exosome isolation.
Exosome isolation
The current ‘gold standard’ for the purification of
a subset of exosomes is differential centrifugation, which
typically consists of low-speed centrifugation to remove cells and
large vesicles and high-speed ultracentrifugation to pellet
exosomes (29). To isolate
exosomes, the cell culture medium (CCM) were collected. Briefly,
any detached and dead cells were removed from the CCM by serial
centrifugation at 300 x g for 10 min at 4˚C and 2,000 x g for 20
min at 4˚C. A further centrifugation procedure was performed at
10,000 x g at 4˚C for 30 min to remove any cell debris.
Supernatants were then collected and filtered through 0.22 mm
filters (Merck KGaA) to remove any contaminating apoptotic bodies,
microvesicles and cell debris. The clarified CCM was then
centrifuged in a Beckman Coulter Optima™ L-80XP Ultracentrifuge at
100,000 x g at 4˚C for 70 min with a Type 32.8 Ti rotor to pellet
the exosomes. The supernatant was carefully removed and the
resulting exosome pellet was resuspended in 100 µl PBS. Protein
concentration in the exosomes was determined by using a
bicinchoninic protein assay.
Exosome characterization
According to the manufacturer's guidelines, we
determined the number and size distribution of the exosomes with a
NanoSight LM10 (Malvern Panalytical). In total, 1 ml sample was
injected into the sample chamber using a sterile syringe. After
that, the morphology of exosomes was observed by transmission
electron microscopy (TEM). Sample (1:1,000) was dropped onto a
carbon-coated 200-mesh copper grid for a 1-min incubation in 2.5%
glutaraldehyde fixing solution at room temperature. Any extra
liquid was absorbed gently using a filter paper around the border
of the grids. The sample was stained with 2% aqueous solution of
phosphotungstic acid for 30 sec at room temperature. Extra liquid
was absorbed again by filter paper. The grids were examined using
an H-7650 TEM (Hitachi, Ltd.) at 80 kV. After the sample was heated
for 1 min at room temperature and embedded in epoxy resin at 37˚C
for 12 h, particle morphology was observed. Additionally,
nanoparticle tracking analysis was performed to assess the size
distribution of PC-3 exosomes using the NanoSight LM10 (Malvern
Panalytical).
Furthermore, the expression of exosome markers was
measured by flow cytometry using an Accuri C6 flow cytometer (BD
Biosciences).
Exosome labeling
PKH fluorescent dyes, which label cell membranes by
the insertion of their aliphatic chains into the lipid bilayer,
have been widely used to label exosomes based on their intense
signal and long half-life (30,31).
PC-3-derived exosomes were labeled with PKH26 according to the
manufacturer's protocol (Umibio Science and Technology Group; cat.
no. UR52302). Briefly, the fluorescent stain solution contained 9
µl Diluent C and 1 µl PKH26. The exosome suspension made as
aforementioned was mixed with the stain solution and incubated for
10 min at 37˚C protected from light. After coculturing exosome (50
ng/1x103cells) with preosteoclasts (1x105) at
37˚C for 12 days, the culture media were removed before the cells
were fixed with 4% paraformaldehyde at room temperature for 5 min
and washed with PBS three times. The cells were then blocked with
5% BSA (Beyotime Institute of Biotechnology) for 10 min at room
temperature and the nuclei were stained with 5 µg/ml DAPI for 5 min
at room temperature. Images of the cells were acquired using a
fluorescence microscope (x100; magnification; Leica-DMi8; Leica
Microsystems GmbH).
Human osteoclast induction
To induce osteoclasts, human peripheral blood
mononuclear cells (PBMCs) were obtained from the peripheral blood
of six healthy male donors (age, 25-29 years; mean age, 26.7 years)
who have signed informed consent forms on August 10, 2019, in
Zhujiang Hospital Outpatient Blood Sampling Center (Guangzhou,
China), where the samples were collected into centrifuge tubes with
1,000 U/ml heparin under a protocol approved by the Committee of
Clinical Ethics of Zhujiang Hospital (Guangzhou, China). The human
peripheral blood samples were diluted with an equal volume of PBS
and centrifuged in a centrifuge tube containing the same volume of
Histopaque-1077 (Sigma-Aldrich; Merck KGaA) at 500 x g for 30 min
at 25˚C with slow deceleration. After centrifugation, PBMCs located
at the interface between the plasma and Histopaque-1077 were
collected and washed with PBS twice, followed by centrifugation at
250 x g for 10 min at 25˚C. The isolated PBMCs were suspended and
incubated in complete RPMI-1640 medium containing 50 ng/ml
macrophage colony-stimulating factor (M-CSF; Sino Biological, Inc.)
in six-well plates at a density of 1x106 cells/ml per
well in 5% CO2 at 37˚C for 3 days. Any non-adherent
cells were then washed away with PBS to obtain adherent mononuclear
cells. To stimulate osteoclast differentiation, 100 ng/ml receptor
activator of nuclear factor κB ligand (RANKL; Sino Biological,
Inc.) was added to complete exosome-free serum RPMI-1640 medium
after removing non-adherent cells in the absence (control) or
presence of PC-3 exosomes at three different concentrations (10, 30
and 50 ng per 1x103 cells seeded) in 5% CO2
at 37˚C for 12 days. The culture medium was replaced every 3 days
with fresh media supplemented with the aforementioned reagents
[RPMI-1640 containing 50 ng/ml macrophage colony-stimulating factor
and 100 ng/ml receptor activator of nuclear factor-κB ligand and
different concentrations of PC-3 exosomes (10, 30 and 50 ng per
1x103 cells seeded)]. Tartrate-resistant acid
phosphatase (TRAP) staining was used for osteoclast staining
whereas western blotting and reverse transcription-quantitative PCR
were used to demonstrate how osteoclast differentiation changed
among these concentrations of PC-3 exosomes.
TRAP staining
TRAP staining was used to confirm the shape of
osteoclasts according to the manufacturer's protocol (Sigma
Aldrich; Merck KGaA). CCM was first removed before the osteoclasts
(1x105) in six-well plates were washed twice with PBS.
Osteoclasts were fixed with Fixative Solution (a combination of 25
ml citrate solution, 65 ml acetone and 8 ml 37% formaldehyde) for
30 sec at room temperature. The samples were then rinsed with
deionized water three times. At the same time, 0.5 ml Fast Garnet
GBC base solution and 0.5 ml sodium nitrite solution were mixed for
30 sec and added into a 10-ml beaker containing 0.5 ml naphthol
AS-BI phosphate solution, 2 ml acetate solution and 1 ml tartrate
solution. Next, the osteoclasts were immersed in this mixture,
incubated at 37˚C for 1 h and rinsed with distilled water three
times. TRAP-positive cells appeared red or purple, where each of
them had ≥ three nuclei.
miRNA mimic/inhibitor
transfection
miR-148a mimics, miR-148a inhibitor and
corresponding negative controls (NC) were purchased from Shanghai
GenePharma Co., Ltd. Their sequences were as follows: miR-148a
mimic sense, 5'-UCAGUGCACUACAGAACUUUGU-3' and anti-sense,
5'AAAGUUCUGUAGUGCACUGAUU-3'; miR-148a inhibitor,
5'-ACAAAGUUCUGUAGUGCACUGA-3'; inhibitor NC,
5'-CAGUACUUUUGUGUAGUACAA-3' and mimic NC,
5'-UUGUACUACACAAAAGUACUG-3'. All NCs are non-targeting sequences.
Cells transfected with the mimic NC and inhibitor NC are designated
as control groups. PBMCs were transfected with 5 µl miR-148a mimic,
inhibitor, mimic NC or inhibitor NC by using the RFect siRNA
transfection reagent (cat. no. BIOG-11014; Changzhou EMI
Biotechnology Co., Ltd.) in six-well plates when cells reached 80%
confluence. After transfection for 6 h, the PBMCs were induced to
undergo osteoclast differentiation by M-CSF + RANKL treatment in 5%
CO2 at 37˚C for 12 days with the medium composition as
aforementioned.
Total RNA Extraction and RT-qPCR
RNA was isolated by TRIzol® extraction
(Thermo Fisher Scientific, Inc.) and reverse-transcribed using an
Evo M-MLV RT kit with gDNA Clean for qPCR at 42˚C for 2 min and
stored at 4˚C (Accurate Biotechnology Co., Ltd.; cat. no. AG11705).
Primers for qPCR were designed by Sangon Biotech Co., Ltd. as
follows: miR-148a forward, 5'-GCGCGTCAGTGCACTACAGAA-3' and reverse,
5'-AGTGCAGGGTCCGAGGTATT-3'; U6 forward,
5'-AGAGAAGATTAGCATGGCCCCTG-3' and reverse,
5'-AGTGCAGGGTCCGAGGTATT-3'; MafB forward,
5'-AGAAGCGGCGGACCCTGAAG-3' and reverse,
5'-GCTGCTCCACCTGCTGAATGAG-3'; MMP-9 forward,
5'-TCCTCTTATGCCTGCCTGTCTCC-3' and reverse,
5'-CTTGGTCCACCTGGTTCAACTCAC-3' and integrin β3 (Iβ3) forward,
5'-GTGACCTGAAGGAGAATCTGC-3' and reverse, 5'-CCGGAGTGCAATCCTCTGG-3'.
U6 and GAPDH (forward, 5'-CCGCATCTTCTTTTGCGTCG-3' and reverse,
5'-CCCAATACGACCAAATCCGTTG-3') were used as internal references.
qPCR was performed in triplicate using the SYBR Green Premix Pro
Taq HD qPCR kit (cat. no. AG11721; Accurate Biotechnology Co.,
Ltd.). PCRs were run and analyzed using the Applied Biosystems
StepOnePlus™ Real-Time PCR System (Thermo Fisher Scientific, Inc.).
The reaction conditions were pre-heating at 95˚C for 3 min,
followed by 40 cycles of denaturation at 95˚C for 30 sec and
annealing at 60˚C for 30 sec. Gene expression was represented as
ΔCq=Cqgene-Ctreference, and the
multiplication ratio was obtained using the 2-ΔΔCq
method (32).
Western Blotting
Osteoclasts were lysed in RIPA lysis buffer (CoWin
Biosciences) containing phosphatase (CoWin Biosciences) and
protease inhibitors (CoWin Biosciences). The cell lysates were
centrifuged at 12,000 x g at 4˚C for 15 min and the protein
concentration collected from the supernatant was determined using a
bicinchoninic acid Protein Assay kit. 60 µg of protein samples (80
µg) were separated by 8% SDS-PAGE and then transferred onto PVDF
membranes. The membranes were blocked in 5% nonfat dry milk diluted
in tris-buffered saline containing 0.1% Tween 20 (TBST) at room
temperature for 1 h. Afterwards, the membranes were incubated with
primary rabbit monoclonal antibodies against the following antigens
at 4˚C overnight: MafB (1:1,000; Zen-bio; cat. no. 862796), MMP-9
(1:1,000; Abcam; #EP1254), Iβ3 (1:1,000; Cell Signaling Technology,
Inc.), AKT, phosphorylated (p)-AKT, mTOR, p-mTOR and GAPDH (all
1:2,000; all Wuhan Boster Biological Technology, Ltd.). After
washing, the membranes were incubated with goat anti-rabbit IgG
secondary antibody (cat. no. BA1040, 1:10,000; Wuhan Boster
Biological Technology, Ltd.) at room temperature for 1 h and washed
with TBST three times. Proteins were visualized using a Tanon 4200
SF automated fluorescence chemiluminescence image analysis system
(Tanon Science and Technology Co., Ltd.) using an ECL kit (KeyGEN
Biotech; cat. no. KGP1121). The image lab software (v3.0; Bio-Rad
Laboratories, Inc.) was used to analyze the results of western
blotting.
Statistical Analysis
All data are presented as the mean ± standard
deviations. Comparisons were made using unpaired Student's t-test
and one-way ANOVA followed by the Tukey's post hoc test with
GraphPad Prism version 8.0 (GraphPad Software, Inc.). P<0.05 was
considered to indicate a statistically significant difference. All
experiments were repeated ≥ three times.
Results
Characteristics of PC-3 Exosomes
To investigate the specific effect of the exosomes
from PC-3 cells on osteoclast differentiation, ultracentrifugation
was used to collect exosomes from the PC-3 prostate cancer cells
that were incubated with exosome-free serum. The exosomes were
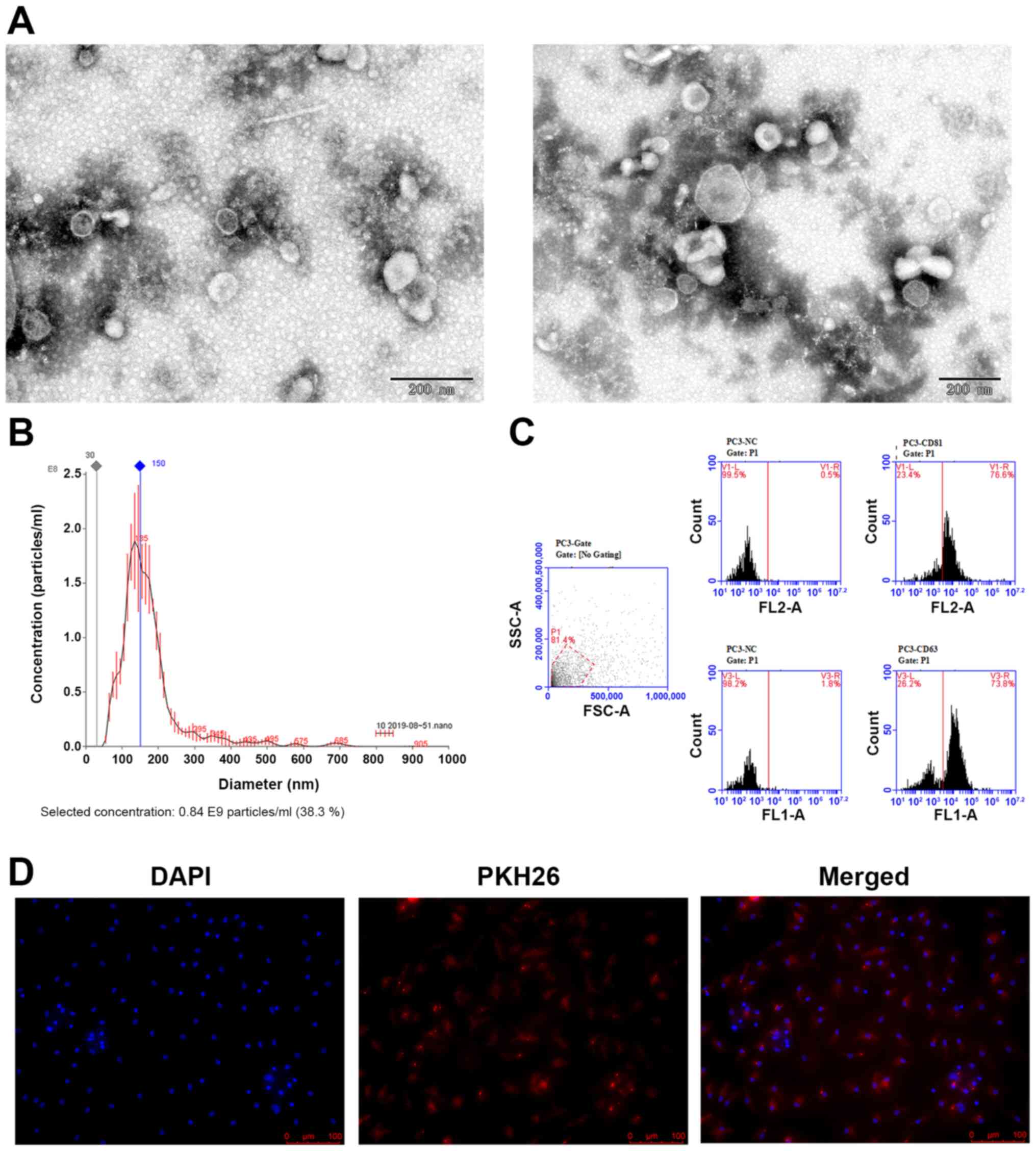
examined by TEM and nanoparticle tracking analysis. According to
the TEM results, the isolated exosomes appeared as round or oval
membrane vesicles (Fig. 1A) with
sizes within the characteristic diameter range of 45-150 nm. The
average size of the exosomes from PC-3 cells was 176.5 nm and the
peak diameter was 136 nm (Fig. 1B).
These exosomes were verified further by detecting the
exosome-specific markers CD81 and CD63 by flow cytometry, which
were expressed at 76.6 and 73.8%, respectively (Fig. 1C).
Pre-osteoclasts uptake PC-3-derived
exosomes
The present study then investigated if the
PC-3-derived exosomes can be taken up by the pre-osteoclasts. The
exosomes were first labeled with the fluorescent dye PKH26 and then
added into the culture medium of preosteoclasts. After 12 h, the
preosteoclasts exhibited efficient uptake of the PC-3-derived
exosomes, as indicated by the presence of red fluorescence staining
in these cells (Fig. 1D).
PC-3-derived exosomes inhibit
osteoclast differentiation
According to the results of TRAP staining, marked
differences in osteoclast differentiation were found among the
groups of PBMCs exposed to three different concentrations of PC-3
exosomes after coculturing for 10 days with M-CSF and RANKL
(Fig. 2A). The differentiation of
osteoclasts decreased when the concentration of exosomes increased.
The most notable inhibition was observed in the group treated with
50 ng/1,000 cells (Fig. 2A). It was
also found that PC-3-derived exosomes reduced the expression level
of miR-148a and inhibited the differentiation of osteoclasts by
decreasing the protein and mRNA expression of osteoclastic markers,
including MMP-9 and Iβ-3 and increasing the protein and mRNA
expression of V-maf musculoaponeurotic fibrosarcoma oncogene
homolog B (MafB) (Fig. 2B and
C).
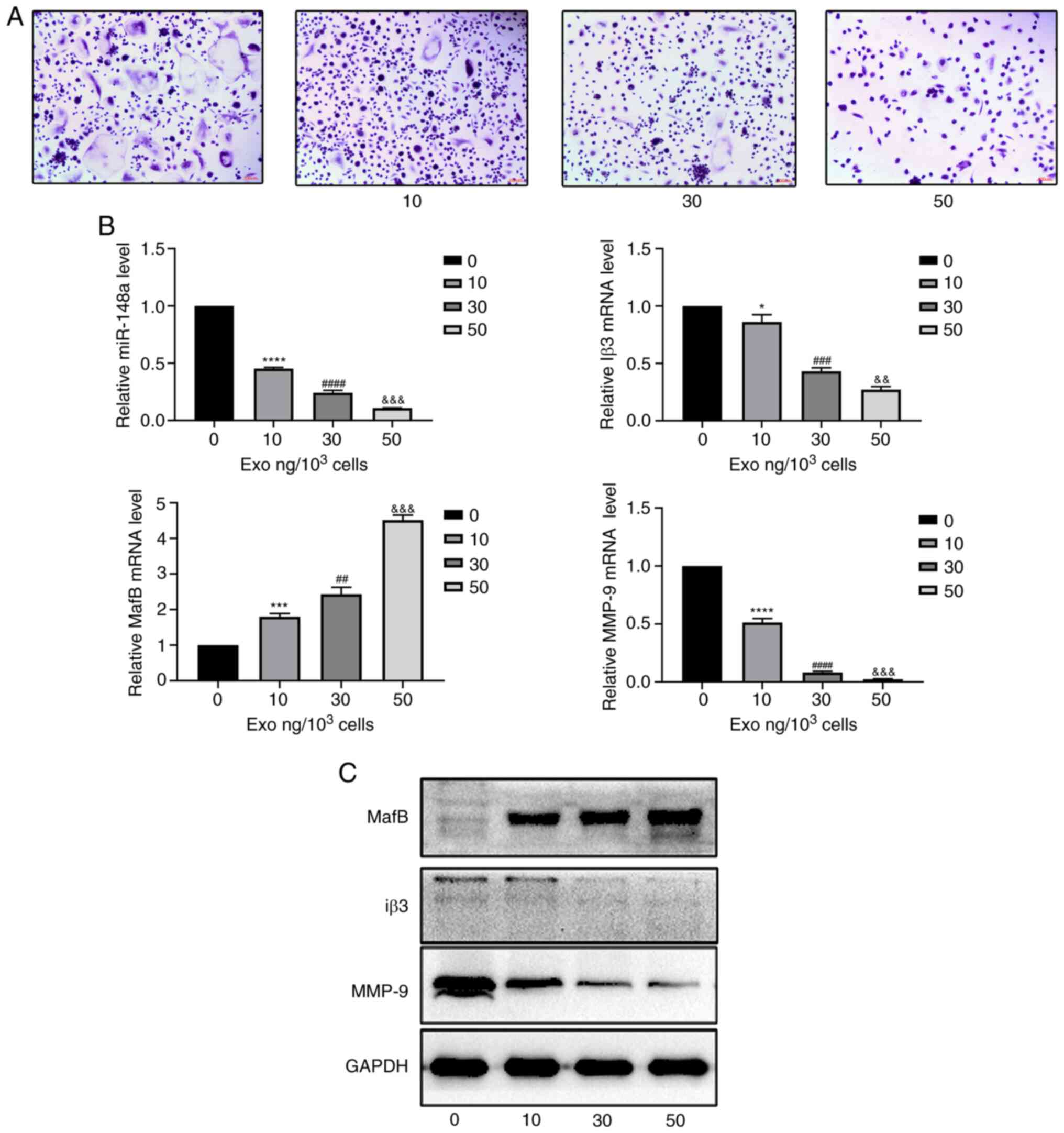 | Figure 2PC-3-derived exosomes inhibit
osteoclast differentiation. (A) Tartrate-resistant acid phosphatase
staining of osteoclasts treated with 0, 10, 30 or 50 ng
exosomes/1,000 cells. Scale bar, 100 µm. (B) Relative expression
levels of miR-148a, MafB, MMP-9 and Iβ3 in osteoclasts treated with
0, 10, 30, or 50 ng exosomes/1,000 cells. (C) Western blotting of
MafB, MMP-9 and Iβ3 in osteoclasts after treatment with 0, 10, 30
or 50 ng exosomes/1,000 cells. *P<0.05,
***P<0.001, ****P<0.0001 0 vs. 10;
##P<0.01, ###P<0.001,
####P<0.0001 10 vs. 30;
&&P<0.01,
&&&P<0.001 30 vs. 50 ng). MafB, V-maf
musculoaponeurotic fibrosarcoma oncogene homolog B; MMP, matrix
metalloproteinase; Iβ3, integrin β3; miR, microRNA; exo,
exosome. |
Downregulation of miR-148a inhibits
the differentiation of osteoclasts
To further investigate if miR-148a serves a role
during osteoclast differentiation, PBMCs were transfected with
miR-148a mimic, miR-148a inhibitor, mimic NC or inhibitor NC.
First, osteoclasts were subjected to TRAP staining (Fig. 3A). Transfection with the miR-148a
mimic increased the differentiation of osteoclasts and formed more
large particles of purplish red positive staining cells compared
with that the mimic NC group, whilst the miR-148a inhibitor reduced
osteoclast differentiation compared with that in the inhibitor NC
group. The expression level of miR-148a and osteoclastic markers
MafB, MMP-9, Iβ3 were also measured by RT-qPCR among the four
groups, where miR-148a, MMP-9 and Iβ3 expression in the mimic group
were significantly higher compared with that in the mimic NC group,
whilst expression in the inhibitor group were significantly lower
compared with that in the inhibitor NC group (Fig. 3B). MafB expression levels in the
mimic group were lower than in the mimic NC group, and higher in
the inhibitor group compared with inhibitor NC. From western
blotting, MafB expression was markedly upregulated, whereas MMP-9
and Iβ3 expression were markedly downregulated after transfection
with the miR-148a inhibitor compared with those in osteoclasts
transfected with inhibitor NC (Fig.
3C). By contrast, transfection with the miR-148a mimics
resulted in the opposite effects compared with the observations in
the miR-148a inhibitor groups (Fig.
3B and C). Therefore,
inhibiting the expression of miR-148a in PBMCs attenuated
differentiation into osteoclasts, whilst upregulating the
expression of miR-148a promoted differentiation into
osteoclasts.
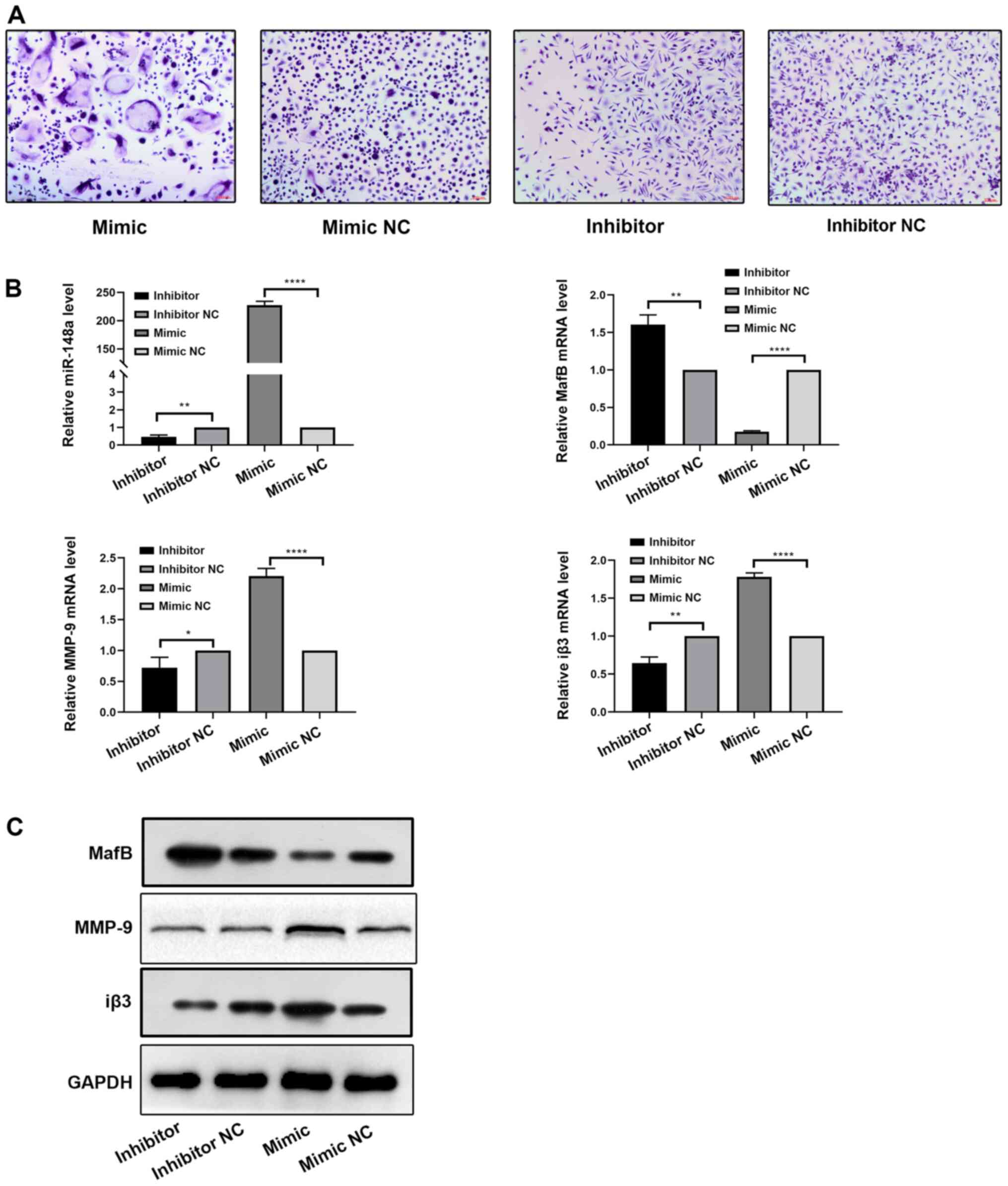 | Figure 3miR-148a downregulation inhibits
osteoclast differentiation. (A) Tartrate-resistant acid phosphatase
staining in osteoclasts expressing the miR-148a mimic, mimic NC,
inhibitor or inhibitor NC after induction with M-CSF and RANKL.
Scale bar, 100 µm. (B) Relative miR-148a, MafB, MMP-9 and Iβ3
expression in osteoclasts transfected with miR-148a mimic,
inhibitor, mimic NC or inhibitor NC after induction with M-CSF and
RANKL was detected by reverse transcription-quantitative PCR. (C)
Western blotting of MafB, MMP-9 and Iβ3 in osteoclasts transfected
with miR-148a mimic, inhibitor, mimic NC or inhibitor NC after
induction with M-CSF and RANKL. *P<0.05,
**P<0.01, ****P<0.0001. MafB, V-maf
musculoaponeurotic fibrosarcoma oncogene homolog B; MMP, matrix
metalloproteinase; Iβ3, integrin β3; miR, microRNA; M-CSF,
macrophage colony-stimulating factor; RANKL, receptor activator of
nuclear factor κB ligand; NC, negative control. |
PC-3-derived exosome downregulates the
AKT and mTOR pathways
To further investigate the underlying mechanism,
activities of the AKT and mTOR signaling pathways were focused
upon. The protein levels of AKT, p-AKT, p-mTOR and mTOR were first
measured in osteoclasts cultured with various concentrations of
PC-3-derived exosomes. The phosphorylation of AKT and mTOR were
significantly suppressed by PC-3-derived exosomes in a
concentration-dependent manner (Fig.
4A). However, the total AKT and mTOR levels remain unchanged.
Subsequently, the effect of miR-148a on the AKT and mTOR signaling
pathways were investigated. The levels of p-mTOR, mTOR, p-AKT, and
AKT were measured in the miR-148a mimic, inhibitor, mimic NC and
inhibitor NC groups. The levels of AKT and mTOR phosphorylation
were significantly increased in the miR-148a mimic group compared
with those in the mimic NC group (Fig.
4B). By contrast, their levels were significantly lower in the
miR-148a inhibitor group compared with those in the inhibitor NC
group (Fig. 4B). Furthermore,
miR-148a overexpression and inhibition exerted no effects on total
mTOR and AKT expression levels (Fig.
4B). These results suggest that PC-3-derived exosomes can block
the mTOR and AKT signaling pathways by miR-148a downregulation.
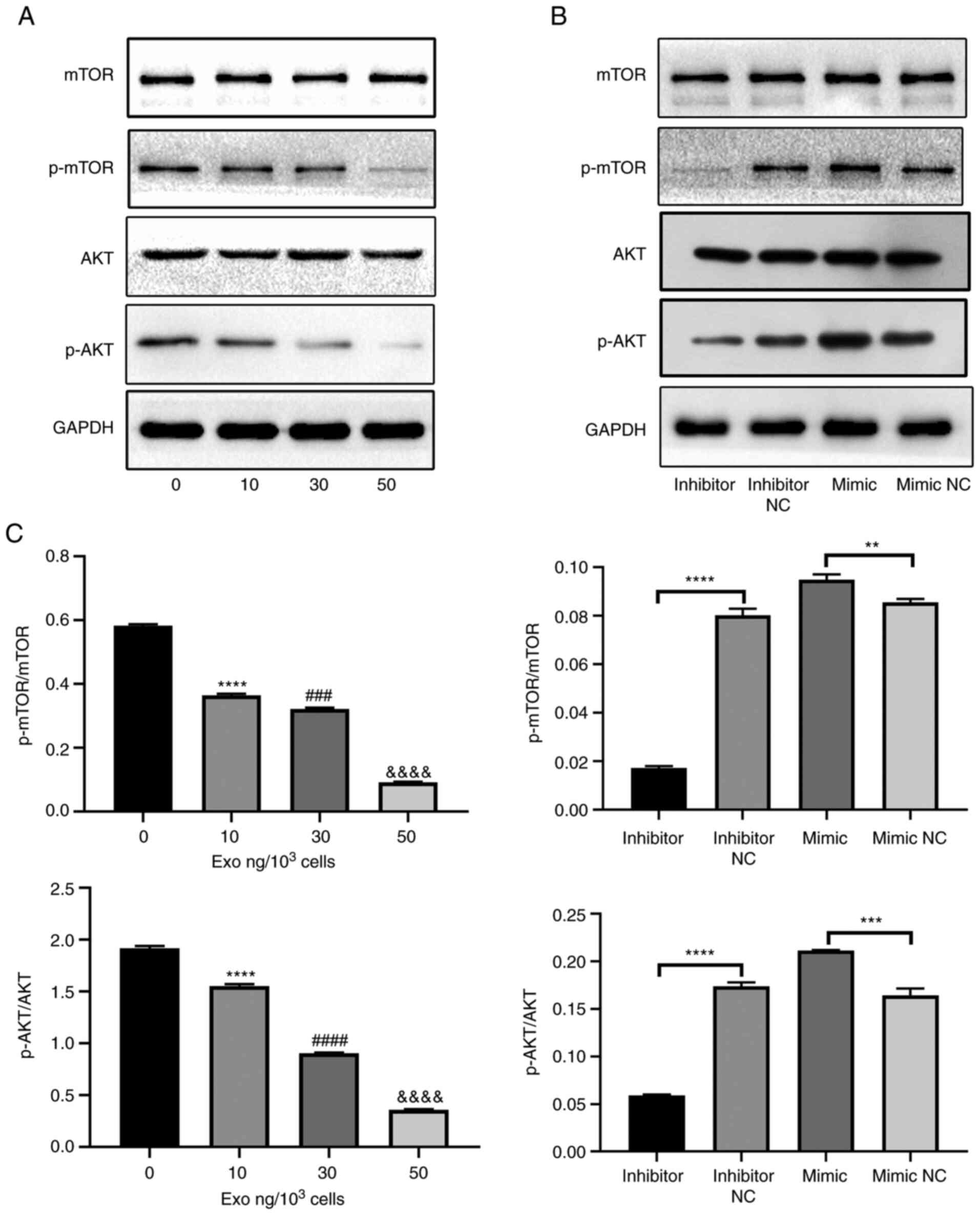 | Figure 4PC-3-derived exosomes block the AKT
and mTOR pathways by miR-148a downregulation. (A) Western blotting
of p-AKT, AKT, p-mTOR and mTOR in osteoclasts treated with 0, 10,
30 or 50 ng exosomes/1,000 cells. (B) Western blotting of p-AKT,
AKT, p-mTOR and mTOR in osteoclasts transfected with miR-148a
mimic, inhibitor, mimic NC or inhibitor NC after induction with
M-CSF and RANKL. (C) Semi-quantification of western blot analysis
of p-AKT/AKT and p-mTOR/mTOR. **P<0.01,
***P<0.001, ****P<0.0001 0 vs. 10;
###P<0.001, ####P<0.0001 10 vs. 30;
&&&&P<0.0001 30 vs. 50 ng. p-,
phosphorylation; miR, microRNA; NC, negative control; M-CSF,
macrophage colony-stimulating factor; RANKL, receptor activator of
nuclear factor κB ligand. |
Discussion
In the present study, the results suggest that
exosomes from the PC-3 prostate cancer cell line suppressed the
differentiation of osteoclasts. The osteoclastic maturation markers
Iβ3 and MMP-9 were found to be attenuated while MafB was
upregulated, where miR-148a expression was also downregulated. In
addition, the mTOR and AKT signaling pathways were suppressed by
exosomes from PC-3 cells. Combining these results and the previous
findings that osteoblastic bone metastases in prostate cancer are
caused by tumor-derived factors, which lead to osteoblast
proliferation, differentiation and bone formation (33), suggest that prostate cancer bone
metastases are mainly osteoblastic (34). The present study explored whether
miR-148a has a role in prostate cancer bone metastases. It was
shown that upregulating miR-148a expression promoted
differentiation of osteoclasts, whilst downregulation of miR-148a
suppressed osteoclast differentiation, thus this may be a novel
treatment method.
Torrealba et al previously suggested that the
PI3K/AKT/mTOR pathway is an important therapeutic target and a
viable option as a predictive biomarker for the onset, behavior and
progression of PC after prostatectomy (35). Therefore, the present study mainly
aimed to investigate activities of the mTOR and AKT pathways
following treatment with PC-3-derived exosomes in prostate cancer
bone metastases. Reduced expression of miR-148a inhibited the mTOR
and AKT pathways in the present study. It is therefore possible
that targeting miR-148a could be a new therapeutic strategy to
protect the bone from damage by prostate cancer metastasis.
Previously, miR-181b-5p upregulation was reported to suppress the
proliferation, migration and invasion of prostate cancer cells by
targeting oncostatin M to modulate RAW264.7 preosteoclast cell
osteoclast differentiation (36).
Another study previously suggested that hsa-miR-143 in exosomes and
trefoil factor 3 are associated with the development of prostate
cancer (37). According to the
findings aforementioned, prostate tumor cells interact closely with
osteoblasts and osteoclasts (38).
However, multiple myeloma cell-derived exosomes can directly induce
the expression of specific osteoclast markers and modulate the
secretion of proteases involved in bone resorption (39). In summary, these aforementioned
studies indicated that different kinds of cancer-derived exosomes
have different effects on bone; the present study suggested that
exosomes from the PC-3 prostate cancer cell line suppressed the
differentiation of osteoclasts.
Furthermore, this present study showed that
PC-3-derived exosomes significantly repressed the expression of
miR-148a in osteoclasts, which is consistent with the finding that
downregulating miR-148a attenuated osteoclast differentiation by
upregulating MafB (23). Exosomes
can deliver cytokines, including receptor activator of nuclear
factor-κB (RANK) and its ligand RANKL and microRNAs, including
miR-218 and miR-148a, to modulate osteoclast differentiation during
bone resorption (23). Therefore,
miR-148a mimics may reduce the progression of PC bone metastases.
However, additional studies are necessary to investigate the
detailed mechanism of miR-148a-mediated regulation of
osteoclastogenesis. Further flow cytometry analysis for the
characterization of cell surface markers is also required (40). In addition, changes in osteoblast
physiology and its surrounding microenvironment following prostate
metastasis to be bone would require further analysis.
Overall, the present study aimed to explore the
effect of PC-3-derived exosomes on the differentiation of
osteoclasts. However, it remains incomplete due to the limitations
of not identifying the specific molecules within the exosomes and
not determining the effect of osteoblasts on bone homeostasis
following PC-3-derived exosomes treatment. In addition, the present
study was only performed in vitro. Therefore, experiments
in vivo and on clinical specimens would also need to be
performed.
In the present study, PC-3-derived exosomes
downregulated the expression level of miR-148a and differentiation
of osteoclasts by blocking the PI3K/AKT/mTOR signaling pathway.
According to these findings, downregulating miR-148a in prostate
cancer may serve as a new therapeutic strategy to prevent PC bone
metastases. In future studies, the effect of prostate cancer on
osteoblast physiology and underlying mechanism would be assessed.
Regarding the limitations of the study, since only the PI3K pathway
was studied, the next step would be to explore the IKB pathway.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the National Natural
Science Foundation of China (grant no. 81672140) and the Natural
Science Foundation of Guangdong Province, China (grant nos.
2018A030313500 and 2017A030313111).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
SN and LZ conceptualized the study and performed the
experiments. GT and KH wrote and edited the manuscript and
collected and analyzed data. GT and KH confirmed the authenticity
of all the raw data. SQ, YX and YC analyzed data. All authors read
and approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Committee of
Clinical Ethics of Zhujiang Hospital (Guangzhou, China; approval
no. 5574813)). All participants provided written informed
consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare they have no competing
interests.
References
|
1
|
Taitt HE: Global trends and prostate
cancer: A review of incidence, detection, and mortality as
influenced by race, ethnicity, and geographic location. Am J Mens
Health. 12:1807–1823. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Parkin DM, Bray F, Ferlay J and Pisani P:
Global cancer statistics, 2002. CA Cancer J Clin. 55:74–108.
2005.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Gilbert SM and McKiernan JM: Epidemiology
of male osteoporosis and prostate cancer. Curr Opin Urol. 15:23–27.
2005.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Sturge J, Caley MP and Waxman J: Bone
metastasis in prostate cancer: Emerging therapeutic strategies. Nat
Rev Clin Oncol. 8:357–368. 2011.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Wong SK, Mohamad NV, Giaze TR, Chin KY,
Mohamed N and Ima-Nirwana S: Prostate cancer and bone metastases:
The underlying mechanisms. Int J Mol Sci. 20(2587)2019.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Boukouris S and Mathivanan S: Exosomes in
bodily fluids are a highly stable resource of disease biomarkers.
Proteomics Clin Appl. 9:358–367. 2015.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Mitchell PJ, Welton J, Staffurth J, Court
J, Mason MD, Tabi Z and Clayton A: Can urinary exosomes act as
treatment response markers in prostate cancer? J Transl Med.
7(4)2009.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Conde-Vancells J, Rodriguez-Suarez E,
Embade N, Gil D, Matthiesen R, Valle M, Elortza F, Lu SC, Mato JM
and Falcon-Perez JM: Characterization and comprehensive proteome
profiling of exosomes secreted by hepatocytes. J Proteome Res.
7:5157–5166. 2008.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Théry C, Zitvogel L and Amigorena S:
Exosomes: Composition, biogenesis and function. Nat Rev Immunol.
2:569–579. 2002.PubMed/NCBI View
Article : Google Scholar
|
|
10
|
Zhang J, Li S, Li L, Li M, Guo C, Yao J
and Mi S: Exosome and exosomal microRNA: Trafficking, sorting, and
function. Genomics Proteomics Bioinformatics. 13:17–24.
2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Png KJ, Halberg N, Yoshida M and Tavazoie
SF: A microRNA regulon that mediates endothelial recruitment and
metastasis by cancer cells. Nature. 481:190–194. 2011.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Lee J, Kwon MH, Kim JA and Rhee WJ:
Detection of exosome miRNAs using molecular beacons for diagnosing
prostate cancer. Artif Cells Nanomed Biotechnol. 46 (Sup3):S52–S63.
2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Xue M, Zhuo Y and Shan B: MicroRNAs, long
noncoding RNAs, and their functions in human disease. Methods Mol
Biol. 1617:1–25. 2017.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Zeng Z, Li Y, Pan Y, Lan X, Song F, Sun J,
Zhou K, Liu X, Ren X, Wang F, et al: Cancer-derived exosomal
miR-25-3p promotes pre-metastatic niche formation by inducing
vascular permeability and angiogenesis. Nat Commun.
9(5395)2018.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Hsu YL, Hung JY, Chang WA, Lin YS, Pan YC,
Tsai PH, Wu CY and Kuo PL: Hypoxic lung cancer-secreted exosomal
miR-23a increased angiogenesis and vascular permeability by
targeting prolyl hydroxylase and tight junction protein ZO-1.
Oncogene. 36:4929–4942. 2017.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297.
2004.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Mitchell PS, Parkin RK, Kroh EM, Fritz BR,
Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O'Briant
KC, Allen A, et al: Circulating microRNAs as stable blood-based
markers for cancer detection. Proc Natl Acad Sci USA.
105:10513–10518. 2008.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Landgraf P, Rusu M, Sheridan R, Sewer A,
Iovino N, Aravin A, Pfeffer S, Rice A, Kamphorst AO, Landthaler M,
et al: A mammalian microRNA expression atlas based on small RNA
library sequencing. Cell. 129:1401–1414. 2007.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Bellavia D, Salamanna F, Raimondi L, De
Luca A, Carina V, Costa V, Alessandro R, Fini M and Giavaresi G:
Deregulated miRNAs in osteoporosis: Effects in bone metastasis.
Cell Mol Life Sci. 76:3723–3744. 2019.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Seeliger C, Karpinski K, Haug AT, Vester
H, Schmitt A, Bauer JS and van Griensven M: Five freely circulating
miRNAs and bone tissue miRNAs Are associated with osteoporotic
fractures. J Bone Miner Res. 29:1718–1728. 2014.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Bedene A, Mencej Bedrač S, Ješe L, Marc J,
Vrtačnik P, Preželj J, Kocjan T, Kranjc T and Ostanek B: MiR-148a
the epigenetic regulator of bone homeostasis is increased in plasma
of osteoporotic postmenopausal women. Wien Klin Wochenschr. 128
(Suppl 7):S519–S526. 2016.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Kelch S, Balmayor ER, Seeliger C, Vester
H, Kirschke JS and van Griensven M: miRNAs in bone tissue correlate
to bone mineral density and circulating miRNAs are gender
independent in osteoporotic patients. Sci Rep.
7(15861)2017.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Cheng P, Chen C, He HB, Hu R, Zhou HD, Xie
H, Zhu W, Dai RC, Wu XP, Liao EY and Luo XH: miR-148a regulates
osteoclastogenesis by targeting V-maf musculoaponeurotic
fibrosarcoma oncogene homolog B. J Bone Miner Res. 28:1180–1190.
2013.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Chen H, Zhou L, Wu X, Li R, Wen J, Sha J
and Wen X: The PI3K/AKT pathway in the pathogenesis of prostate
cancer. Front Biosci (Landmark Ed). 21:1084–1091. 2016.PubMed/NCBI View
Article : Google Scholar
|
|
25
|
Mayer IA and Arteaga CL: The PI3K/AKT
pathway as a target for cancer treatment. Annu Rev Med. 67:11–28.
2016.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Edlind MP and Hsieh AC: PI3K-AKT-mTOR
signaling in prostate cancer progression and androgen deprivation
therapy resistance. Asian J Androl. 16:378–386. 2014.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Zhu W, Hu X, Xu J, Cheng Y, Shao Y and
Peng Y: Effect of PI3K/Akt signaling pathway on the process of
prostate cancer metastasis to bone. Cell Biochem Biophys.
72:171–177. 2015.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Tang LA, Dixon BN, Maples KT, Poppiti KM
and Peterson TJ: Current and investigational agents targeting the
phosphoinositide 3-kinase pathway. Pharmacotherapy. 38:1058–1067.
2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Théry C, Amigorena S, Raposo G and Clayton
A: Isolation and characterization of exosomes from cell culture
supernatants and biological fluids. Curr Protoc Cell Biol Chapter.
3(Unit 3.22)2006.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Skog J, Würdinger T, van Rijn S, Meijer
DH, Gainche L, Sena-Esteves M, Curry WT Jr, Carter BS, Krichevsky
AM and Breakefield XO: Glioblastoma microvesicles transport RNA and
proteins that promote tumour growth and provide diagnostic
biomarkers. Nat Cell Biol. 10:1470–1476. 2008.PubMed/NCBI View
Article : Google Scholar
|
|
31
|
Urabe F, Kosaka N, Kimura T, Egawa S and
Ochiya T: Extracellular vesicles: Toward a clinical application in
urological cancer treatment. Int J Urol. 25:533–543.
2018.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Schmittgen T and Livak K: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Janssen JC, Woythal N, Meißner S, Prasad
V, Brenner W, Diederichs G, Hamm B and Makowski MR:
[68Ga]PSMA-HBED-CC uptake in osteolytic, osteoblastic,
and bone marrow metastases of prostate cancer patients. Mol Imaging
Biol. 19:933–943. 2017.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Fang J and Xu Q: Differences of
osteoblastic bone metastases and osteolytic bone metastases in
clinical features and molecular characteristics. Clin Transl Oncol.
17:173–179. 2015.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Torrealba N, Rodriguez-Berriguete G,
Fraile B, Olmedilla G, Martínez-Onsurbe P, Sánchez-Chapado M,
Paniagua R and Royuela M: PI3K pathway and Bcl-2 family.
Clinicopathological features in prostate cancer. Aging Male.
21:211–222. 2018.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Shao B, Fu X, Yu Y and Yang D: Regulatory
effects of miRNA-181a on FasL expression in bone marrow mesenchymal
stem cells and its effect on CD4+T lymphocyte apoptosis in estrogen
deficiency-induced osteoporosis. Mol Med Rep. 18:920–930.
2018.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Che Y, Shi X, Shi Y, Jiang X, Ai Q, Shi Y,
Gong F and Jiang W: Exosomes derived from miR-143-overexpressing
MSCs inhibit cell migration and invasion in human prostate cancer
by downregulating TFF3. Mol Ther Nucleic Acids. 18:232–244.
2019.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Fornetti J, Welm AL and Stewart SA:
Understanding the bone in cancer metastasis. J Bone Miner Res.
33:2099–2113. 2018.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Raimondi L, De Luca A, Amodio N, Manno M,
Raccosta S, Taverna S, Bellavia D, Naselli F, Fontana S, Schillaci
O, et al: Involvement of multiple myeloma cell-derived exosomes in
osteoclast differentiation. Oncotarget. 6:13772–13789.
2015.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Fathi E, Valipour B, Sanaat Z, Nozad
Charoudeh H and Farahzadi R: Interleukin-6, -8, and TGF-β secreted
from mesenchymal stem cells show functional role in reduction of
telomerase activity of leukemia cell via Wnt5a/β-catenin and P53
pathways. Adv Pharm Bull. 10:307–314. 2020.PubMed/NCBI View Article : Google Scholar
|