Introduction
Globally, the most common primary malignant bone
tumor among teenagers and young adults is osteosarcoma (1). Due to surgery being combined with
chemotherapy, the 5-year survival rate for patients with
osteosarcoma has reached 50-80% (2). However, ~80% of osteosarcoma patients
will develop local recurrence or metastasis after surgical
treatment (3). Therefore, in order
to further improve the survival rate of patients with osteosarcoma,
there is an urgent requirement to develop new diagnostic biomarkers
and innovative treatment strategies. Prior studies have suggested
that there are a wide range of genetic and molecular alterations in
osteosarcoma (4,5); however, the highly complex molecular
mechanism of osteosarcoma has not been elucidated.
Kinesin superfamily proteins are responsible for the
movement of membrane-bound compartments and transport vesicles
(6). This family contains 45
genes, known as kinesins or kifs, which are divided into 14
different families (7). Kifs play
a vital role in mitosis, especially during spindle formation and
cytokinesis (8). Kinesin family
member 4A (KIF4A) is one member that appears to play a role in the
regulation of gene expression and heterochromatin formation
(9). A number of studies have
shown that abnormal expression of KIF4A can lead to tumorigenesis,
including that of breast, lung, colorectal and liver cancer
(10-13).
However, the role of KIF4A in osteosarcoma is unclear. In the
present study, the aim was to explore the role of KIF4A in human
osteosarcoma and confirm whether it could be considered as a tumor
induction gene, which may be used an indicator of prognosis.
Materials and methods
Bioinformatics analyses
KIF4A expression level and Kaplan-Meier survival
analysis were obtained using the Gene Expression Profiling
Interactive Analysis (GEPIA) website (http://gepia.cancer-pku.cn/index.html) using the term
‘KIF4A’ in the ‘Gene’ field and ‘SARC’ in the ‘Datasets Selection’
field. Differential expression of KIF4A was defined with the
following significance cutoff levels: |Log2FC|>1 and
P<0.01. For the Kaplan-Meier survival analysis, the same terms
were used and the group cutoff for KIF4A was the median expression
value. The Gene Expression Omnibus (GEO) database (www.ncbi.nlm.gov/geo) was used to download the
GSE28424 dataset (14). After
using 'osteosarcoma' as the search term, the GSE28424 dataset was
selected, as the study gives a comprehensive analysis of
osteosarcoma mRNA and provides new insight into the complex genetic
mechanisms of tumor development and the progression of
osteosarcoma. The common differentially expressed genes were
obtained using the GEO2R online tool (http://www.ncbi.nlm.nih.gov/geo/geo2r/). P<0.05 and
|Log2FC|>1 were set as cut-off criteria and KIF4A was
selected.
Patients
A total of 72 paired primary osteosarcoma and
adjacent noncancerous tissues were retrospectively obtained from
patients who received palliative surgery or radical resection at
Lianyungang No. 1 People's Hospital Affiliated to Xuzhou Medical
University (Lianyungang, China) between January 2005 and December
2018 (age range, 2-14 years; average age, 9.47±3.30 years). None of
the enrolled patients received radiotherapy or chemotherapy before
surgery. All patients underwent chemotherapy after surgery.
Inclusion criteria: Osteosarcoma patients without radiotherapy or
chemotherapy before surgery. Exclusion criteria: i) Osteosarcoma
patients received radiotherapy or chemotherapy before surgery; ii)
patients refused to sign the informed consent form. The
histological diagnosis of osteosarcoma was confirmed by two
professional pathologists and conformed to the World Health
Organization's and the American Joint Committee on Cancer
histological criteria (15). The
collected specimens were snap frozen in liquid nitrogen and stored
at -80˚C promptly until RNA extraction. All enrolled patient
guardians provided written informed consent. This research
conformed to the principles of the Declaration of Helsinki and was
approved by the Ethics Committee of The Lianyungang No. 1 People's
Hospital Affiliated to Xuzhou Medical University.
Cell lines
The human normal osteoblastic hFOB cell line and the
human osteosarcoma MG63, U2OS, HOS and Saos2 cell lines were
purchased from the American Type Culture Collection and cultured in
DMEM (Invitrogen; Thermo Fisher Scientific, Inc.) supplemented with
10% fetal bovine serum (Gibco; Thermo Fisher Scientific, Inc.) in a
humidified incubator containing 5% CO2 at 37˚C.
RNA isolation and reverse
transcription-quantitative PCR (RT-qPCR)
TRIzol® reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) was used to extract RNA from cell lines
(hFOB, MG63, U2OS, HOS and Saos2) and tissues (primary osteosarcoma
and adjacent noncancerous tissues). Following RNA extraction, the
EasyScript One Step gDNA Removal and cDNA Synthesis SuperMix
(TransGen Biotech Co., Ltd.) was used to acquire the first strand
cDNA according to the manufacturer's instructions. Next, RT-qPCR
was performed using SYBR-Green Master Mix kit (Roche Diagnostics)
with an Applied Biosystems 7900 Real-Time PCR System (Thermo Fisher
Scientific, Inc.). The reaction conditions were as follows:
Pre-denaturation at 95˚C for 15 min; followed by 35 cycles of
denaturation at 95˚C for 15 sec, annealing at 58˚C for 30 sec and
extension at 72˚C for 5 min; and final extension at 72˚C for 15
min. Expression levels of RNA were calculated based on the
comparative 2-ΔΔCq method. RT-qPCR was performed as in
previous publications (16,17).
The level of β-actin was used to normalize the relative LIF4A
expression level. Experiments were performed with the following
primers: KIF4A forward, 5'-TGGTGTGGAAACAAGCAGTGTG-3' and reverse,
5'-CAC CCTGTTGTTGCTGTAGCCAAA-3'; and β-actin forward,
5'-TCACCCACACTGTGCCCATCTACGA-3' and reverse,
5'-CAGCGGAACCGCTCATTGCCAATGG-3'. All experiments were performed in
triplicate.
siRNA transfection
siRNA oligos for KIF4A, commercially constructed by
Shanghai GenePharma Co., Ltd., were used to perform knockdown
experiments. KIF4A-siRNA oligo sequences were as follows: Sense,
5'-GGAAUGAGGUUGUGA UCUUTT-3' and anti-sense, 5'-AAGAUCACAACCUCA
UUCCTT-3'. Non-silencing siRNA (sense, 5'-UUCUCCGAAC GUGUCACGUTT-3'
and antisense, 5'-ACGUGACACGUUC GGAGAATT-3') was used as a negative
control. Before transfecting siRNA (20 nM) with
Lipofectamine® 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.), the cells were cultured in growth media in a
6-well plate until 70% confluence was reached according to the
manufacturer's instructions. Transfection was performed at 37˚C for
6 h before changing the transfection medium with full culture
medium. Then cells were harvested 48 h after transfection, and then
western blotting or RT-qPCR analyses were performed.
Cell proliferation assay
At 48 h post-siRNA transfection, transfected cells
at a density of 1,000 cells per well were seeded in 96-well plates,
with 5 replicate wells. A total of 10 µl of 5 mg/ml MTT was added
into each well at 0, 24, 48 and 72 h after transfection, and then
the cells were cultured for 2 h in an incubator at 37˚C. Then, 150
µl DMSO was added per well. Finally, a microplate reader was used
to measure the absorbance at 490 nm. All viability experiments were
performed in triplicate.
Colony formation assay
At 6 h post-transfection, MG63 and U2OS cells were
resuspended as a single-cell suspension using trypsin, and then
seeded into 6-well plates with 2,000 cells/well, with 2 ml complete
medium added to each well. Next, the 6-well plates were placed in
an incubator at 37˚C. After 14 days, 4% paraformaldehyde was used
to fix the cells for 20 min at room temperature, 0.1% crystal
violet was applied to stain the cells for 30 min at room
temperature, and then tap water was used to rinse the cells, which
were photographed for manual counting. Colonies consisted of >50
cells.
Western blot analysis
After using pre-iced PBS to wash the cells, the
cells were lysed with RIPA buffer for 30 min, and then centrifuged
to collect the supernatant at 14,000 x g for 15 min at 4˚C.
Bradford protein assay (Bio-Rad Laboratories, Inc.) was used to
conduct protein quantitation; then per lane was loaded 30 µg of
protein. Electrophoresis was performed with 10% SDS-PAGE, and the
separated proteins were transferred onto nitrocellulose membranes.
After transferring the protein to membranes, 5% skimmed milk was
used to block the membranes for 1 h at room temperature, followed
by incubation with the following primary antibodies at 4˚C
overnight: Rabbit anti-KIF4A (1:1,000 dilution; catalog no.
ab124903), anti-GAPDH (1:2,500 dilution; catalog no. ab9485),
anti-ERK (1:5,000 dilution; ab265600), anti-MEK (1:20,000 dilution;
catalog no. ab178876); anti-cAMP responsive element binding protein
(CREB; 1:1,000 dilution; catalog no. ab31387) (all Abcam).
Subsequently, HRP-conjugated secondary antibodies (1:5,000
dilution; catalog no. S0001; Affinity Biosciences) were added for 1
h at room temperature. Pierce™ ECL Western Blotting Substrate (cat.
no. 32109; Thermo Fisher Scientific, Inc.) was used to visualize
the bound antibodies. The levels of protein were evaluated using
chemiluminescence detection system (GE Healthcare Biosciences), and
image analysis was performed using ImageJ software (version 1.42;
National Institutes of Health). Western blotting was performed as
in a previous publication (17).
Statistical analysis
Data are presented as the mean ± standard deviation.
Paired or unpaired Student's t-test or the Mann-Whitney U test was
used to compare the continuous variables in two groups. One-way
ANOVA was adopted to compare the continuous variables among
multiple groups, followed by Dunnett's post hoc test. The
differences in categorical variables were analyzed with Pearson's
χ2 and Fisher's exact tests and data were presented as n
(%). The prognostic value of patient survival was estimated by
Kaplan-Meier method and log-rank tests. Receiver operating curve
(ROC) analysis was used to assess predictive value of biomarkers,
together with calculation of the area under the ROC curve (AUC),
and ‘sensitivity + specificity -1’ was used as the best cut-off
value. Pearson correlation was used to evaluate the strength of
linear correlation between two continuous variables. GraphPad Prism
8.0 (GraphPad Software, Inc.) was used to analyze all statistical
data. P<0.05 was used to indicate a statistically significant
difference, and all P-values were two-sided.
Results
KIF4A upregulation in
osteosarcoma
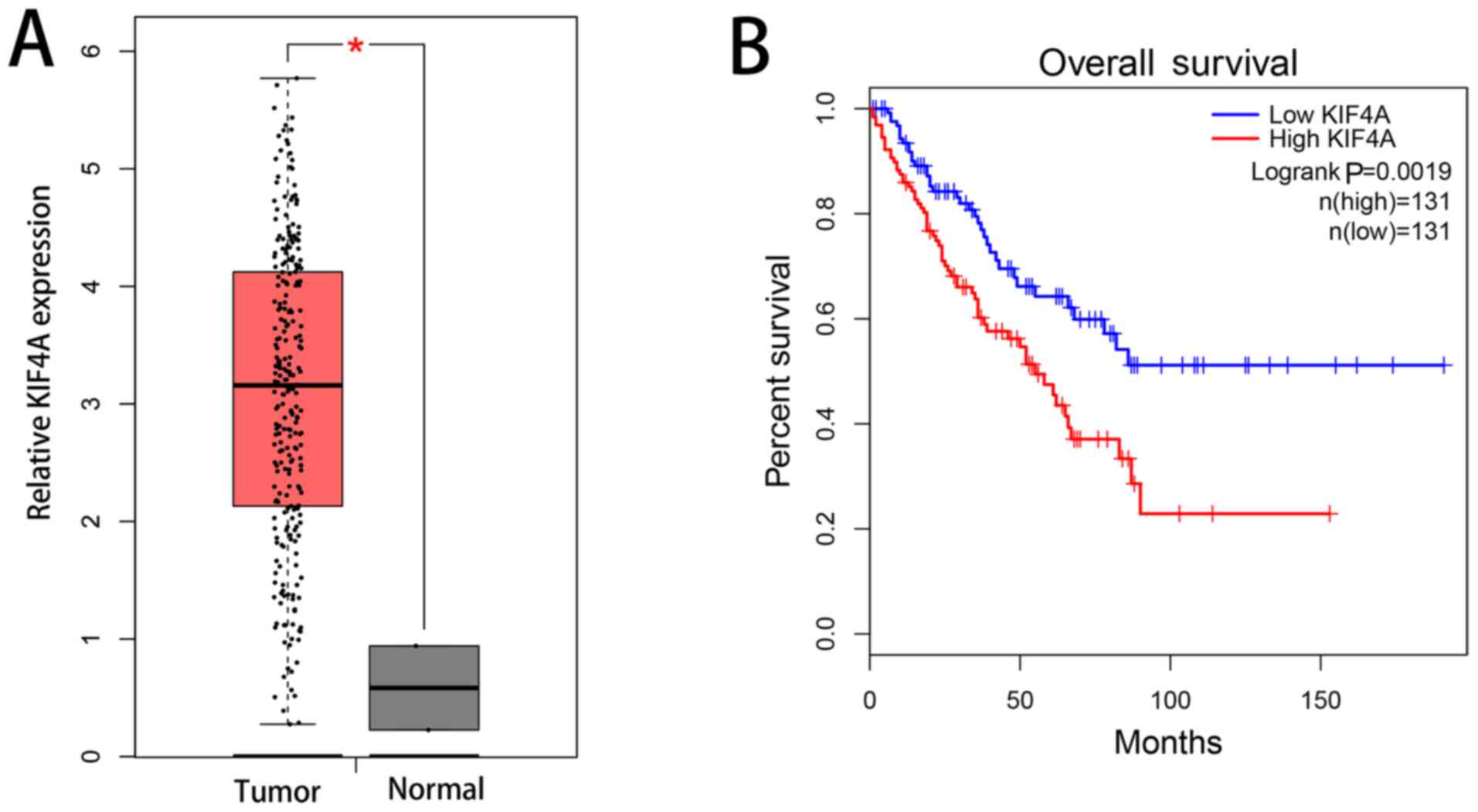
Using analysis of a clinical patient database, by
GEPIA, the expression level of KIF4A in sarcoma tissues (n=262) was
found to be upregulated in comparison with that non-sarcoma tissues
(n=2) at the mRNA level (Fig. 1A).
Upregulated expression of KIF4A was also associated with poor
overall survival (Fig. 1B).
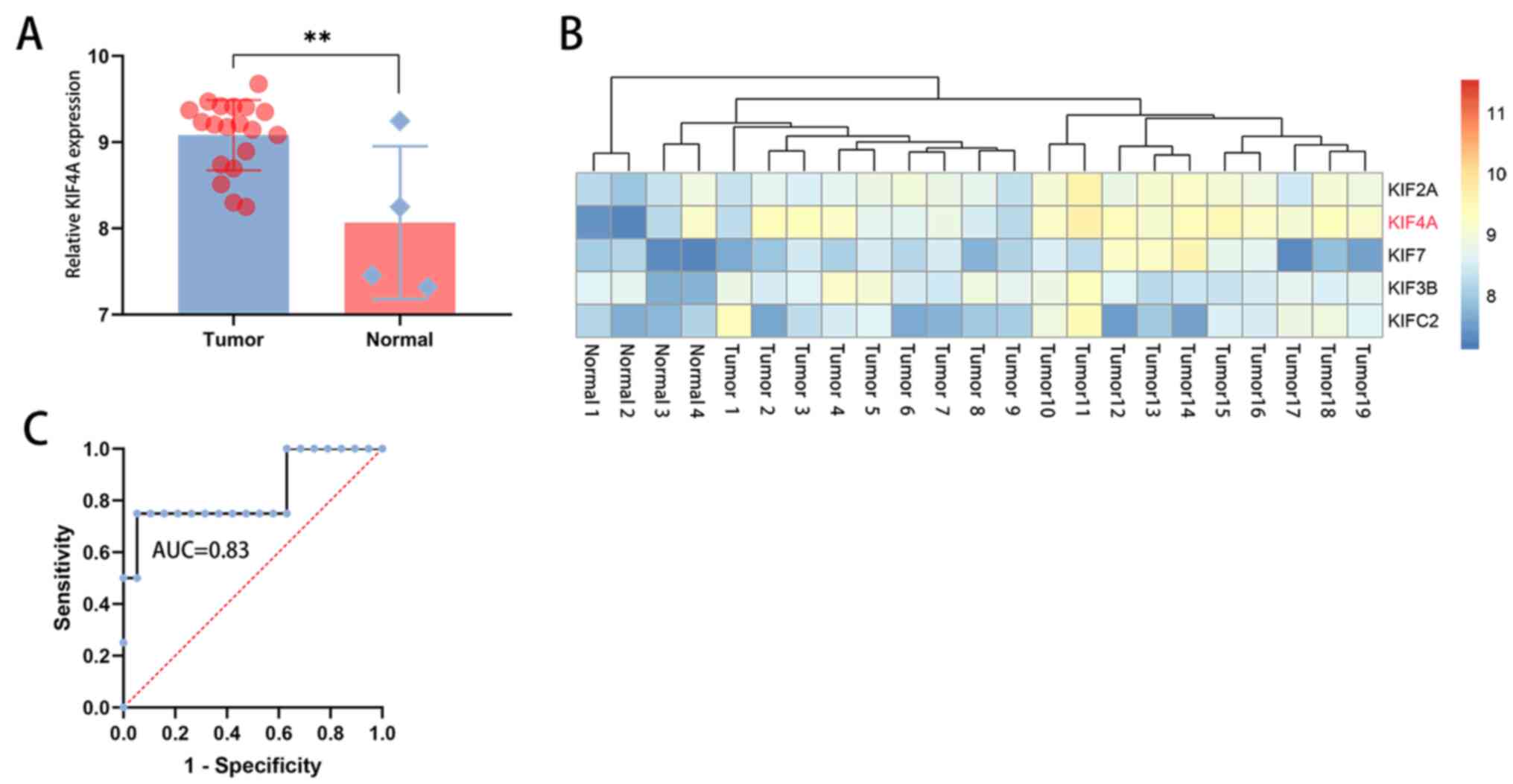
Analysis of the GEO dataset GSE28424 indicated that
KIF4A expression was upregulated in osteosarcoma (n=19) compared
with that in non-osteosarcoma (n=4) (Fig. 2A and B). The sensitivity and specificity of
KIF4A was evaluated by ROC-AUC, and the result showed that the AUC
was 0.83, suggesting that KIF4A should be considered as a potential
biomarker of osteosarcoma cases (Fig.
2C).
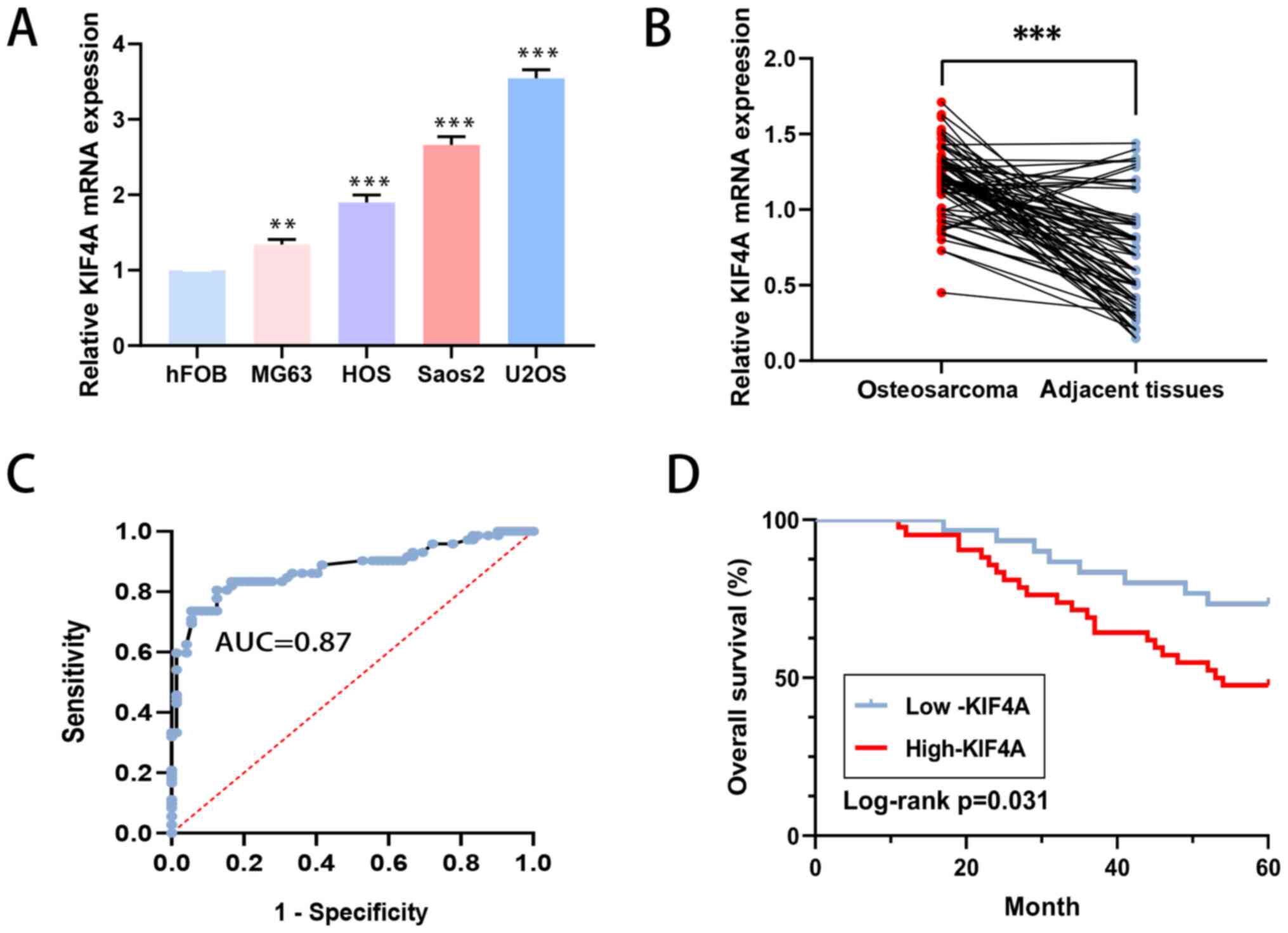
Compared with that in the normal osteoblastic hFOB
cell line, the level of KIF4A in the osteosarcoma MG63, U2OS, HOS
and Saos2 cell lines was higher according to RT-qPCR analysis
(Fig. 3A). In the 72 pairs of
human osteosarcoma and adjacent tissue collected, there were
similar results (Fig. 3B).
KIF4A expression level is associated
with a poor outcome in patients with osteosarcoma
Using the paired patient tissues, the AUC of KIF4A
was determined to be 0.87 (Fig.
3C). ROC analysis was used to acquire the best cut-off value
and then two groups were obtained, the low KIF4A expression group
(n=42) and the high KIF4A expression group (n=30) were created
(Table I). Clinical stage
(15), distant metastasis and
response to chemotherapy were found to be significantly associated
with high KIF4A expression (P=0.018, P=0.042 and P=0.020,
respectively). However, anatomical location, sex, age, lactate
dehydrogenase level, alkaline phosphatase level, and tumor size
were not associated with the expression of KIF4A. Moreover, it was
found that high expression level of KIF4A was associated with poor
overall survival in patients with osteosarcoma (P=0.031; Fig. 3D).
 | Table IAssociation between KIF4A level and
the clinicopathological features of osteosarcoma. |
Table I
Association between KIF4A level and
the clinicopathological features of osteosarcoma.
| | KIF4A expression | |
|---|
| Clinicopathological
features | Number of cases | High, n (%) | Low, n (%) | P-value |
|---|
| Age, years | | | | 0.417 |
|
<8 | 19 | 6 (31.58) | 13 (68.42) | |
|
≥8 | 53 | 24 (45.28) | 29 (54.72) | |
| Sex | | | | 0.809 |
|
Male | 41 | 18 (43.90) | 23 (56.10) | |
|
Female | 31 | 12 (38.71) | 19 (61.29) | |
| Tumor size, cm | | | | 0.634 |
|
<8 | 33 | 15 (45.45) | 18 (54.55) | |
|
≥8 | 39 | 15 (38.46) | 24 (61.54) | |
| Anatomical
location | | | | 0.063 |
|
Tibia/femur | 60 | 28 (46.67) | 32 (53.33) | |
|
Elsewhere | 12 | 2 (16.67) | 10 (83.33) | |
| Serum level of
lactate dehydrogenase | | | | 0.305 |
|
Elevated | 49 | 18 (36.73) | 31 (63.27) | |
|
Normal | 23 | 12 (52.17) | 11 (47.83) | |
| Serum level of
alkaline phosphatase | | | | 0.116 |
|
Elevated | 51 | 18 (35.29) | 33 (64.71) | |
|
Normal | 21 | 12 (57.14) | 9 (42.86) | |
| Clinical stage | | | | 0.018 |
|
I | 23 | 5 (21.74) | 18 (78.26) | |
|
II | 35 | 15 (42.86) | 20 (57.14) | |
|
III | 14 | 10 (71.43) | 4 (28.57) | |
| Distant
metastasis | | | | 0.042 |
|
Absent | 61 | 22 (36.07) | 39 (63.93) | |
|
Present | 11 | 8 (72.73) | 3 (27.27) | |
| Response to
chemotherapy | | | | 0.020 |
|
Good | 56 | 19 (33.93) | 37 (66.07) | |
|
Poor | 16 | 11 (68.75) | 5 (31.25) | |
Silencing of KIF4A inhibits the
proliferation of osteosarcoma cells
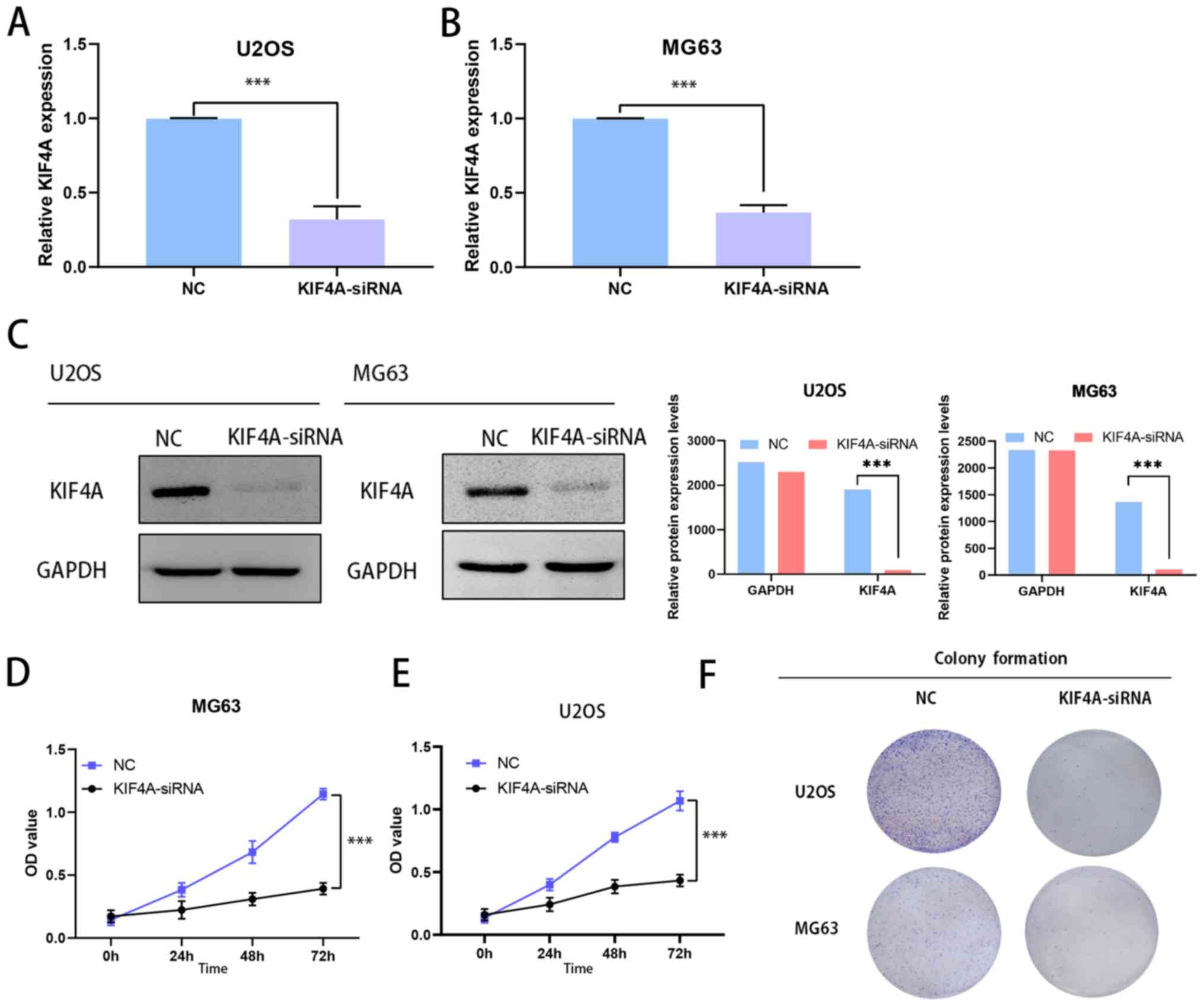
KIF4A siRNA plasmids were used to knock down KIF4A
expression in the osteosarcoma U2OS and MG63 cell lines, which were
selected as they are derived from children and teenagers. Next,
RT-qPCR was performed to show the efficiency of the KIF4A knockdown
(Fig. 4A and B). The transfection and expression
efficiency was also confirmed by western blotting (Fig. 4C). MTT assays were performed to
assess proliferation in KIF4A-siRNA-transfected osteosarcoma cells.
Results indicated that KIF4A knockdown inhibited U2OS and MG63 cell
proliferation (Fig. 4D and
E). Similarly, the colony
formation assays found that KIF4A knockdown inhibited colony
formation (Fig. 4F). Taken
together, both MTT and colony formation assay findings showed that
osteosarcoma cell proliferation was inhibited when KIF4A was
knocked down.
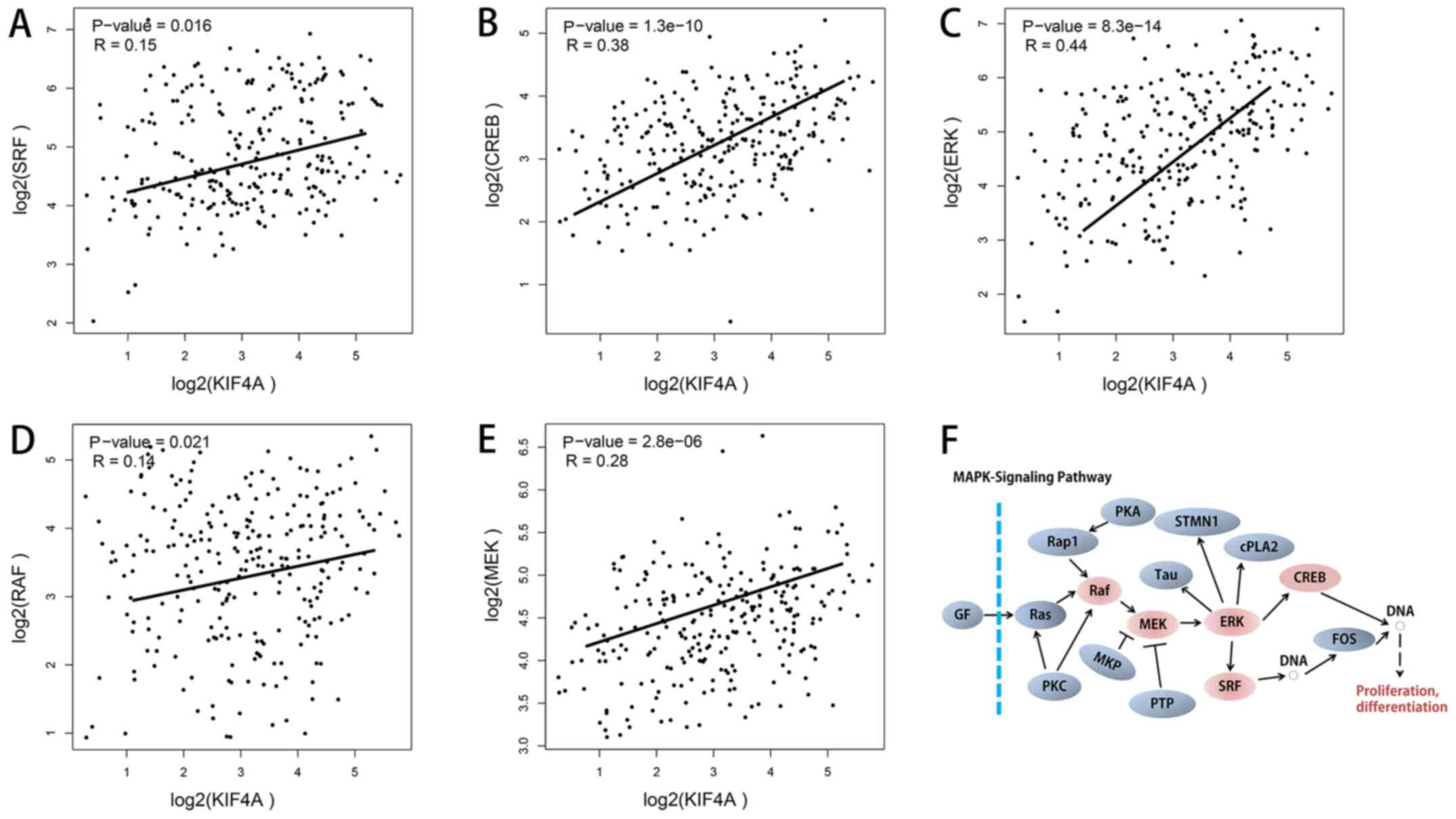
KIF4A knockdown may suppress MAPK
signaling pathway activity in osteosarcoma cells
The MAPK signaling pathway is an important
regulatory pathway that regulates the growth of cancer cells.
Positive correlations were found between KIF4A and the five key
genes in the MAPK signaling pathway, as determined in GEPIA using
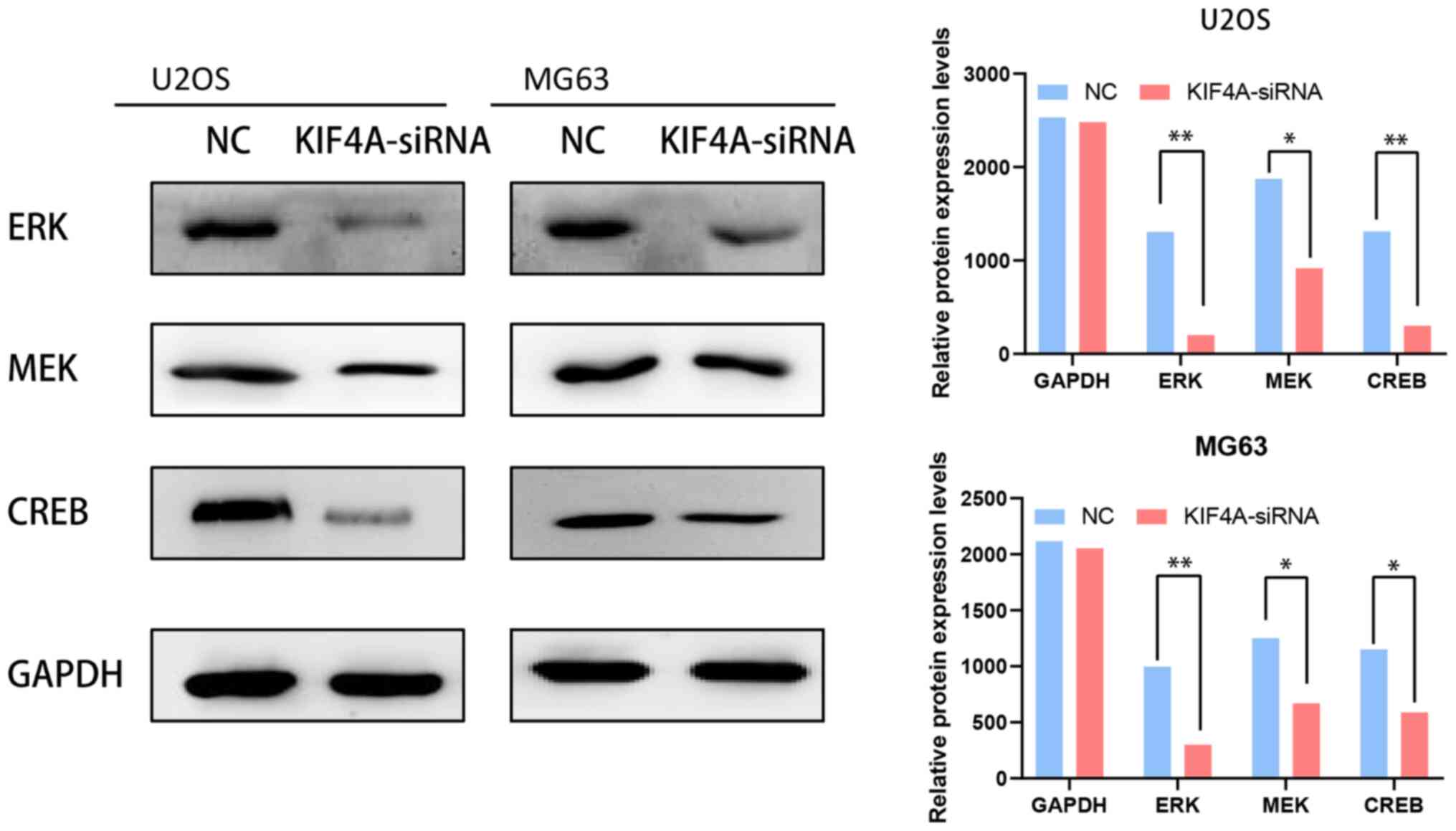
bioinformatics analysis of co-expression (Fig. 5A-F). To confirm our hypothesis of a
positive correlation between the KIF4A and MAPK signaling pathways,
the protein expression levels of ERK, MEK and CREB were obtained
after knockdown of KIF4A in osteosarcoma U2OS and MG63 cells. The
expression of the three proteins was found to be significantly
decreased when KIF4A was knocked down (Fig. 6). These findings show that KIF4A
may facilitate tumor proliferation by affecting the MAPK
pathway.
Discussion
In the pediatric age group, osteosarcoma is the most
frequent malignant tumor of the bone, with a poor prognosis
(4). Currently, the therapeutic
efficacy of osteosarcoma treatment strategy is not ideal, and it is
especially important to find a sensitive biomarker for the
diagnosis of early phase osteosarcoma and as a target for
treatment. Osteosarcoma is a disease resulting from a combination
of multiple genes (5). However,
the key genes in osteosarcoma have not been determined. Hence, it
is still necessary to explore novel therapeutic targets.
The results of the present study suggested that the
expression level of KIF4A was high in osteosarcoma, as observed in
other malignant tumors, as reported by previous studies (11,18-21).
In osteosarcoma, KIF4A upregulation has been confirmed by
bioinformatics analysis, and this has been validated by cell
experiments (22). However, to the
best of our knowledge, no clinical data exists. The present study
also found KIF4A upregulation in osteosarcoma. Furthermore, KIF4A
was found to promote the proliferation of osteosarcoma cells
through the MAPK signaling pathway. In the present study, in the
ROC curve analysis based on the patient tissue data, the AUC was
0.87, meaning that KIF4A was a highly sensitive diagnostic
predictor of osteosarcoma. Furthermore, in the patients with
osteosarcoma, a higher expression level of KIF4A was associated
with a shorter overall survival time, according to Kaplan-Meier
analysis (P=0.031). KIF4A may therefore be considered as a
therapeutic target in osteosarcoma and a factor that affects the
outcome of patients with this disease.
KIF4A is a member of the kif subfamily; various
studies have previously suggested that KIF4A has a critical role in
tumorigenesis and tumor progression (11,20).
KIF4A, in humans, is a motor protein for the action of microtubules
and plays a role in spindle formation, as well as in the regulation
of chromosome segregation (23).
In 2007, scientists first discovered that KIF4A was upregulated in
carcinoma of the lungs and that silencing KIF4A using siRNA can
suppress the invasive activity of tumor cells (11). In a recent study of renal cancer,
Liu et al (24) reported
that KIF4A might be an independent factor that predicts
recurrence-free survival and overall survival in renal carcinoma.
These findings are consistent with the experimental results from
the present study.
The present study also found that the proliferation
of osteosarcoma cells was suppressed when KIF4A expression was
inhibited. In colorectal cancer, KIF4A can mediate p21 to arrest
the cell cycle, promoting the proliferation of tumor cells. The
protein level of ERK, MEK and AKT were decreased when silencing
KIF4A, indicating that the PI3K/AKT signaling pathway was
inactivated (12). However, in the
present study, when KIF4A activation was inhibited, the expression
of AKT was not changed. KIF4A downregulation resulted in changes in
the protein expression levels of MEK, ERK and CREB. Further
research is required for this different mechanism.
As reported in a number of studies, upregulation of
KIF4A is associated with a number of different cancer types
(12,19,22),
but the molecular mechanisms underlying KIF4A function in
osteosarcoma are not well understood. The present study revealed
the association between KIF4A and clinicopathological features in
osteosarcoma, and found that KIF4A indeed promoted the
proliferation of osteosarcoma via the MAPK signaling pathway.
Overall, KIF4A in osteosarcoma may be considered as
a potential target for tumor therapy. This finding should help to
normalize potential new targets for the diagnosis and treatment of
osteosarcoma. However, there were some limitations faced in the
present study. First, it was only a retrospective study; second,
the study was conducted in vitro; third, the clinical sample
size was not large at only 72 cases; fourth, the GEPIA data only
contained 2 normal samples, and the GEO data contained 4, making
any statistical analysis difficult to interpret. We acknowledge the
limited number of control samples as a limitation in this study.
Fifth, only the proliferation of osteosarcoma cells affected by
KIF4A was analyzed. Finally, only knockdown experiments, but no
overexpression experiments, were performed. Future studies will
continue to reveal the role of KIF4A in osteosarcoma.
In conclusion, in the present study, KIF4A was found
to promote tumor progression via activation of the MAPK signaling
pathway in osteosarcoma, and could be considered as a novel
biomarker and potential target for osteosarcoma treatment.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
DZ and TW conceived and directed the project. DZ
contributed to the writing of the manuscript. DZ, XX and MZ
analyzed the data. DZ, XX and MZ collected the clinical data. DZ,
XX, MZ and TW confirmed the authenticity of the raw data. All
authors have read and approved the final manuscript.
Ethics approval and consent to
participate
All of the enrolled patient guardians provided
written informed consent. This research conformed to the principles
of the Declaration of Helsinki and was approved by the Ethics
Committee of The Lianyungang No. 1 People's Hospital Affiliated to
Xuzhou Medical University (December 2018; approval no. 20180104;
Lianyungang, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ritter J and Bielack SS: Osteosarcoma. Ann
Oncol. 21 (Suppl 7):vii320–vii325. 2010.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Mirabello L, Troisi RJ and Savage SA:
Osteosarcoma incidence and survival rates from 1973 to 2004: Data
from the Surveillance, Epidemiology, and End Results Program.
Cancer. 115:1531–1543. 2009.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Marina N, Gebhardt M, Teot L and Gorlick
R: Biology and therapeutic advances for pediatric osteosarcoma.
Oncologist. 9:422–441. 2004.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Joko R, Yamada D, Nakamura M, Yoshida A,
Takihira S, Takao T, Lu M, Sato K, Ito T, Kunisada T, et al: PRRX1
promotes malignant properties in human osteosarcoma. Transl Oncol.
14(100960)2021.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Tian W, Li Y, Zhang J, Li J and Gao J:
Combined analysis of DNA methylation and gene expression profiles
of osteosarcoma identified several prognosis signatures. Gene.
650:7–14. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Hirokawa N and Tanaka Y: Kinesin
superfamily proteins (KIFs): Various functions and their relevance
for important phenomena in life and diseases. Exp Cell Res.
334:16–25. 2015.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Camlin NJ, McLaughlin EA and Holt JE:
Motoring through: The role of kinesin superfamily proteins in
female meiosis. Hum Reprod Update. 23:409–420. 2017.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Samejima K, Samejima I, Vagnarelli P,
Ogawa H, Vargiu G, Kelly DA, de Lima Alves F, Kerr A, Green LC,
Hudson DF, et al: Mitotic chromosomes are compacted laterally by
KIF4 and condensin and axially by topoisomerase IIα. J Cell Biol.
199:755–770. 2012.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Mazumdar M, Sung MH and Misteli T:
Chromatin maintenance by a molecular motor protein. Nucleus.
2:591–600. 2011.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Xue D, Cheng P, Han M, Liu X, Xue L, Ye C,
Wang K and Huang J: An integrated bioinformatical analysis to
evaluate the role of KIF4A as a prognostic biomarker for breast
cancer. OncoTargets Ther. 11:4755–4768. 2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Taniwaki M, Takano A, Ishikawa N, Yasui W,
Inai K, Nishimura H, Tsuchiya E, Kohno N, Nakamura Y and Daigo Y:
Activation of KIF4A as a prognostic biomarker and therapeutic
target for lung cancer. Clin Cancer Res. 13:6624–6631.
2007.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Matsumoto Y, Saito M, Saito K, Kanke Y,
Watanabe Y, Onozawa H, Hayase S, Sakamoto W, Ishigame T, Momma T,
et al: Enhanced expression of KIF4A in colorectal cancer is
associated with lymph node metastasis. Oncol Lett. 15:2188–2194.
2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Hu G, Yan Z, Zhang C, Cheng M, Yan Y, Wang
Y, Deng L, Lu Q and Luo S: FOXM1 promotes hepatocellular carcinoma
progression by regulating KIF4A expression. J Exp Clin Cancer Res.
38(188)2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Namløs HM, Meza-Zepeda LA, Barøy T,
Østensen IH, Kresse SH, Kuijjer ML, Serra M, Bürger H,
Cleton-Jansen AM and Myklebost O: Modulation of the osteosarcoma
expression phenotype by microRNAs. PLoS One.
7(e48086)2012.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Cates JM: Simple staging system for
osteosarcoma performs equivalently to the AJCC and MSTS systems.
Journal of orthopaedic research : official publication of the
Orthopaedic Research Society. 36:2802–2808. 2018.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Mu J, Fan L, Liu D and Zhu D:
Overexpression of shugoshin1 predicts a poor prognosis for prostate
cancer and promotes metastasis by affecting epithelial-mesenchymal
transition. OncoTargets Ther. 12:1111–1118. 2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Narayan G, Bourdon V, Chaganti S,
Arias-Pulido H, Nandula SV, Rao PH, Gissmann L, Dürst M, Schneider
A, Pothuri B, et al: Gene dosage alterations revealed by cDNA
microarray analysis in cervical cancer: Identification of candidate
amplified and overexpressed genes. Genes Chromosomes Cancer.
46:373–384. 2007.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Gao J, Sai N, Wang C, Sheng X, Shao Q,
Zhou C, Shi Y, Sun S, Qu X and Zhu C: Overexpression of
chromokinesin KIF4 inhibits proliferation of human gastric
carcinoma cells both in vitro and in vivo. Tumour Biol. 32:53–61.
2011.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Minakawa Y, Kasamatsu A, Koike H, Higo M,
Nakashima D, Kouzu Y, Sakamoto Y, Ogawara K, Shiiba M, Tanzawa H,
et al: Kinesin family member 4A: A potential predictor for
progression of human oral cancer. PLoS One.
8(e85951)2013.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Zou JX, Duan Z, Wang J, Sokolov A, Xu J,
Chen CZ, Li JJ and Chen HW: Kinesin family deregulation coordinated
by bromodomain protein ANCCA and histone methyltransferase MLL for
breast cancer cell growth, survival, and tamoxifen resistance. Mol
Cancer Res. 12:539–549. 2014.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Pan J, Lei X and Mao X: Identification of
KIF4A as a pan-cancer diagnostic and prognostic biomarker via
bioinformatics analysis and validation in osteosarcoma cell lines.
PeerJ. 9(e11455)2021.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Mazumdar M, Sundareshan S and Misteli T:
Human chromokinesin KIF4A functions in chromosome condensation and
segregation. J Cell Biol. 166:613–620. 2004.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Liu G, Lu Y, Li L, Jiang T, Chu S, Hou P,
Bai J and Chen M: The kinesin motor protein KIF4A as a potential
therapeutic target in renal cell carcinoma. Invest New Drugs.
38:1730–1742. 2020.PubMed/NCBI View Article : Google Scholar
|