|
1
|
Barta JA, Powell CA and Wisnivesky JP:
Global epidemiology of lung cancer. Ann Glob Health.
85(8)2019.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Cao M and Chen W: Epidemiology of lung
cancer in China. Thorac Cancer. 10:3–7. 2019.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Taghvaee S, Sowlat MH, Hassanvand MS,
Yunesian M, Naddafi K and Sioutas C: Source-specific lung cancer
risk assessment of ambient PM2.5-bound polycyclic
aromatic hydrocarbons (PAHs) in central Tehran. Environ Int.
120:321–332. 2018.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Brasky TM, White E and Chen CL: Long-term,
supplemental, one-carbon metabolism-related vitamin B use in
relation to lung cancer risk in the vitamins and lifestyle (VITAL)
cohort. J Clin Oncol. 35:3440–3448. 2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Hicks BM, Filion KB, Yin H, Sakr L, Udell
JA and Azoulay L: Angiotensin converting enzyme inhibitors and risk
of lung cancer: Population based cohort study. BMJ.
363(k4209)2018.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Lin KF, Wu HF, Huang WC, Tang PL, Wu MT
and Wu FZ: Propensity score analysis of lung cancer risk in a
population with high prevalence of non-smoking related lung cancer.
BMC Pulm Med. 17(120)2017.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Fortunato O, Borzi C, Milione M, Centonze
G, Conte D, Boeri M, Verri C, Moro M, Facchinetti F, Andriani F, et
al: Circulating mir-320a promotes immunosuppressive macrophages M2
phenotype associated with lung cancer risk. Int J Cancer.
144:2746–2761. 2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
ENCODE Project Consortium. The ENCODE
(ENCyclopedia of DNA elements) project. Science. 306:636–340.
2004.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Huarte M: The emerging role of lncRNAs in
cancer. Nat Med. 21:1253–1261. 2015.PubMed/NCBI View
Article : Google Scholar
|
|
11
|
Wu Y, Shao A, Wang L, Hu K, Yu C, Pan C
and Zhang S: The role of lncRNAs in the distant metastasis of
breast cancer. Front Oncol. 9(407)2019.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Zhang Y and Tang L: The application of
lncRNAs in cancer treatment and diagnosis. Recent Pat Anticancer
Drug Discov. 13:292–301. 2018.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Lu T, Wang Y, Di Chen JL and Jiao W:
Potential clinical application of lncRNAs in non-small cell lung
cancer. Onco Targets Ther. 11:8045–8052. 2018.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Engle L, Simpson C and Landers J: Using
high-throughput SNP technologies to study cancer. Oncogene.
25:1594–1601. 2006.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Syvänen AC: Toward genome-wide SNP
genotyping. Nat Genet. 37 (Suppl):S5–S10. 2005.PubMed/NCBI View
Article : Google Scholar
|
|
16
|
Andrew AS, Gui J, Sanderson AC, Mason RA,
Morlock EV, Schned AR, Kelsey KT, Marsit CJ, Moore JH and Karagas
MR: Bladder cancer SNP panel predicts susceptibility and survival.
Hum Genet. 125:527–539. 2009.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Duboule D: The rise and fall of Hox gene
clusters. Development. 134:2549–2560. 2007.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Rinn JL, Kertesz M, Wang JK, Squazzo SL,
Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E and
Chang HY: Functional demarcation of active and silent chromatin
domains in human HOX loci by noncoding RNAs. Cell. 129:1311–1323.
2007.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Hajjari M and Salavaty A: HOTAIR: an
oncogenic long non-coding RNA in different cancers. Cancer Biol
Med. 12:1–9. 2015.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Loewen G, Jayawickramarajah J, Zhuo Y and
Shan B: Functions of lncRNA HOTAIR in lung cancer. J Hematol Oncol.
7(90)2014.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Fatica A and Bozzoni I: Long non-coding
RNAs: New players in cell differentiation and development. Nat Rev
Genet. 15:7–21. 2014.PubMed/NCBI View
Article : Google Scholar
|
|
22
|
Somarowthu S, Legiewicz M, Chillón I,
Marcia M, Liu F and Pyle AM: HOTAIR forms an intricate and modular
secondary structure. Mol Cell. 58:353–361. 2015.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Bhan A and Mandal SS: LncRNA HOTAIR: A
master regulator of chromatin dynamics and cancer. Biochim Biophys
Acta. 1856:151–164. 2015.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Zhuang Y, Wang X, Nguyen HT, Zhuo Y, Cui
X, Fewell C, Flemington EK and Shan B: Induction of long intergenic
non-coding RNA HOTAIR in lung cancer cells by type I collagen. J
Hematol Oncol. 6(35)2013.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Gupta RA, Shah N, Wang KC, Kim J, Horlings
HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote cancer
metastasis. Nature. 464:1071–1076. 2010.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Wu Y, Zhang L, Wang Y, Li H, Ren X, Wei F,
Yu W, Wang X, Zhang L, Yu J and Hao X: Long noncoding RNA HOTAIR
involvement in cancer. Tumour Biol. 35:9531–9538. 2014.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Xu ZY, Yu QM, Du YA, Yang LT, Dong RZ,
Huang L, Yu PF and Cheng XD: Knockdown of long non-coding RNA
HOTAIR suppresses tumor invasion and reverses
epithelial-mesenchymal transition in gastric cancer. Int J Biol
Sci. 9:587–597. 2013.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Liu XH, Liu ZL, Sun M, Liu J, Wang ZX and
De W: The long non-coding RNA HOTAIR indicates a poor prognosis and
promotes metastasis in non-small cell lung cancer. BMC Cancer.
13(464)2013.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Zhao W, An Y, Liang Y and Xie XW: Role of
HOTAIR long noncoding RNA in metastatic progression of lung cancer.
Eur Rev Med Pharmacol Sci. 18:1930–1936. 2014.PubMed/NCBI
|
|
30
|
Nakagawa T, Endo H, Yokoyama M, Abe J,
Tamai K, Tanaka N, Sato I, Takahashi S, Kondo T and Satoh K: Large
noncoding RNA HOTAIR enhances aggressive biological behavior and is
associated with short disease-free survival in human non-small cell
lung cancer. Biochem Biophys Res Commun. 436:319–324.
2013.PubMed/NCBI View Article : Google Scholar
|
|
31
|
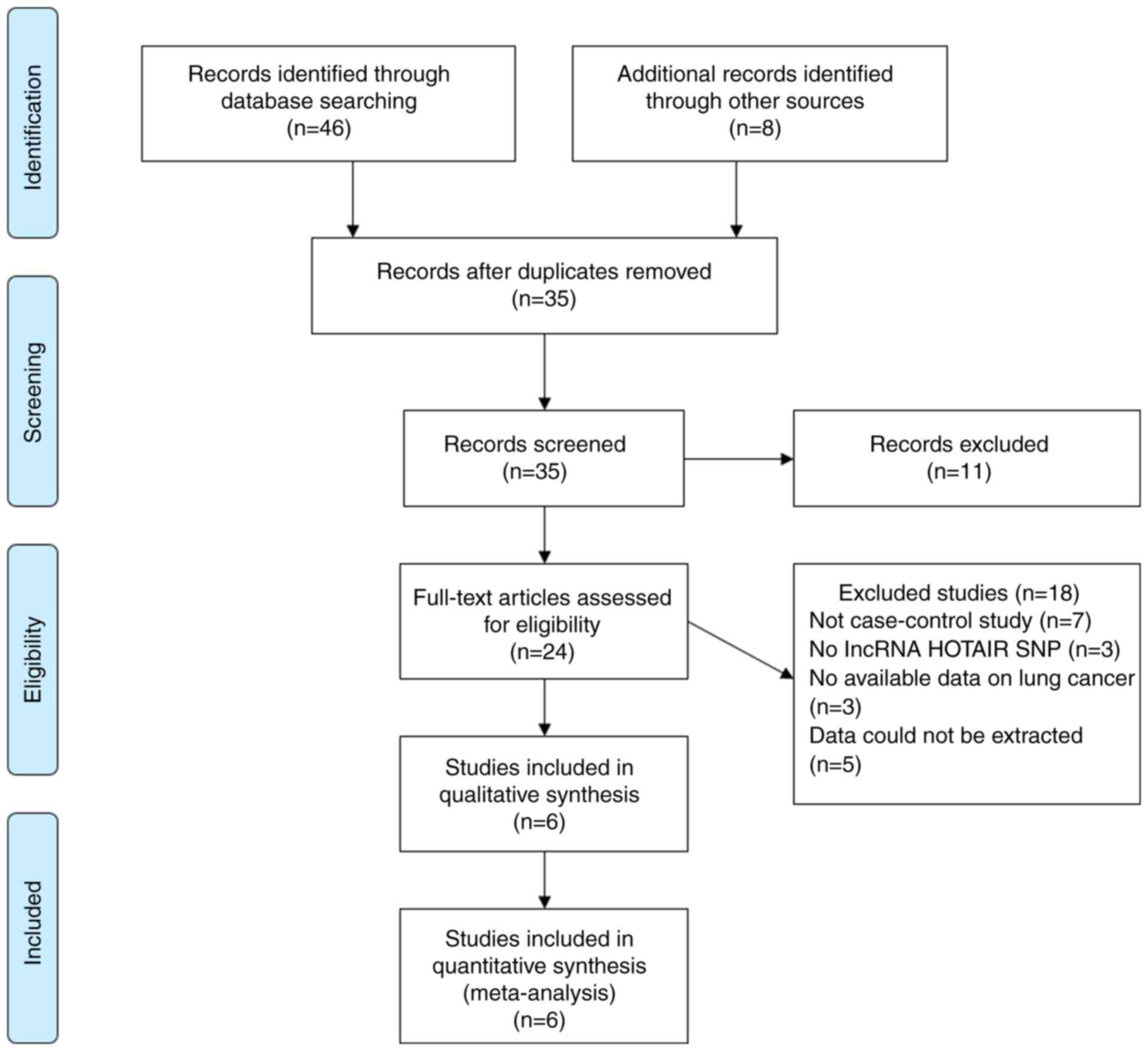
Moher D, Liberati A, Tetzlaff J and Altman
DG: PRISMA Group. Preferred reporting items for systematic reviews
and meta-analyses: The PRISMA statement. PLoS Med.
6(e1000097)2009.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Stang A: Critical evaluation of the
Newcastle-Ottawa scale for the assessment of the quality of
nonrandomized studies in meta-analyses. Eur J Epidemiol.
25:603–605. 2010.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Ren MM, Xu S, Wei YB, Yang JJ, Yang YN,
Sun SS, Li YJ, Wang PY and Xie SY: Roles of HOTAIR in lung cancer
susceptibility and prognosis. Mol Genet Genomic Med.
8(e1299)2020.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Minn AKK, Sato N, Mieno MN, Arai T and
Muramatsu M: Association study of long non-coding RNA HOTAIR
rs920778 polymorphism with the risk of cancer in an elderly
Japanese population. Gene. 729(144263)2020.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Wang C, Li Y, Li YW, Zhang HB, Gong H,
Yuan Y, Li WT, Liu HY and Chen J: HOTAIR lncRNA SNPs rs920778 and
rs1899663 are associated with smoking, male gender, and squamous
cell carcinoma in a Chinese lung cancer population. Acta Pharmacol
Sin. 39:1797–1803. 2018.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Li H, Yang Z, Li J, Lv X, Gao M, Bi Y,
Zhang Z, Wang S, Li S, Li N, et al: Genetic variants in lncRNA
HOTAIR are associated with lung cancer susceptibility in a Chinese
Han population in China: A case-control study. Cancer Manag Res.
10:5209–5218. 2018.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Dadaş E and Aydin M: Effect of HOTAIR
rs12826786 and rs1899663 polymorphisms on lung cancer
susceptibility and clinicopathological characteristics in a Turkish
population: A hospital-based case-control study. Cell Mol Biol
(Noisy-le-grand). 64:97–102. 2018.
|
|
38
|
Gong WJ, Yin JY, Li XP, Fang C, Xiao D,
Zhang W, Zhou HH, Li X and Liu ZQ: Association of
well-characterized lung cancer lncRNA polymorphisms with lung
cancer susceptibility and platinum-based chemotherapy response.
Tumour Biol. 37:8349–8358. 2016.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Dor Y and Cedar H: Principles of DNA
methylation and their implications for biology and medicine.
Lancet. 392:777–786. 2018.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Shimizu Y, Sato S, Noguchi Y, Koyamatsu J,
Yamanashi H, Higashi M, Nagayoshi M, Kadota K, Kawashiri SY, Nagata
Y, et al: Impact of single nucleotide polymorphism on short stature
and reduced tongue pressure among community-dwelling elderly
Japanese participants: A cross-sectional study. Environ Health Prev
Med. 22(62)2017.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Leaché AD and Oaks JR: The utility of
single nucleotide polymorphism (SNP) data in phylogenetics. Annu
Rev Ecol Evol Syst. 48:69–84. 2017.
|
|
42
|
Zhang X, Zhou L, Fu G, Sun F, Shi J, Wei
J, Lu C, Zhou C, Yuan Q and Yang M: The identification of an ESCC
susceptibility SNP rs920778 that regulates the expression of lncRNA
HOTAIR via a novel intronic enhancer. Carcinogenesis. 35:2062–2067.
2014.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Zhu H, Lv Z, An C, Shi M, Pan W, Zhou L,
Yang W and Yang M: Onco-lncRNA HOTAIR and its functional genetic
variants in papillary thyroid carcinoma. Sci Rep.
6(31969)2016.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Pan W, Liu L, Wei J, Ge Y, Zhang J, Chen
H, Zhou L, Yuan Q, Zhou C and Yang M: A functional lncRNA HOTAIR
genetic variant contributes to gastric cancer susceptibility. Mol
Carcinog. 55:90–96. 2016.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Liu Z, Sun M, Lu K, Liu J, Zhang M, Wu W,
De W, Wang Z and Wang R: The long noncoding RNA HOTAIR contributes
to cisplatin resistance of human lung adenocarcinoma cells via
downregualtion of p21(WAF1/CIP1) expression. PLoS One.
8(e77293)2013.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Fan L, Lei H, Lin Y, Zhou Z, Li J, Wu A,
Shu G, Ruger S and Yin G: Hotair promotes the migration and
proliferation in ovarian cancer by miR-222-3p/CDK19 axis. Cell Mol
Life Sci. 79(254)2022.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Chen J, Hou SF, Tang FJ, Liu DS, Chen ZZ,
Zhang HL and Wang SH: HOTAIR/Sp1/miR-199a critically regulates
cancer stemness and malignant progression of cutaneous squamous
cell carcinoma. Oncogene. 41:99–111. 2022.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Zhang W, Wu Q, Liu Y, Wang X, Ma C and Zhu
W: LncRNA HOTAIR promotes chemoresistance by facilitating
epithelial to mesenchymal transition through miR-29b/PTEN/PI3K
signaling in cervical cancer. Cells Tissues Organs. 211:16–29.
2022.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Zhang W, Liu J, Wu Q, Liu Y and Ma C:
HOTAIR contributes to stemness acquisition of cervical cancer
through regulating miR-203 interaction with ZEB1 on
epithelial-mesenchymal transition. J Oncol.
2021(4190764)2021.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Zhang M, Wu K, Zhang P, Qiu Y, Bai F and
Chen H: HOTAIR facilitates endocrine resistance in breast cancer
through ESR1/miR-130b-3p axis: Comprehensive analysis of
mRNA-miRNA-lncRNA network. Int J Gen Med. 14:4653–4663.
2021.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Deng S, Wang J, Zhang L, Li J and Jin Y:
LncRNA HOTAIR promotes cancer stem-like cells properties by
sponging miR-34a to activate the JAK2/STAT3 pathway in pancreatic
ductal adenocarcinoma. Onco Targets Ther. 14:1883–1893.
2021.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Sun N, Zhang W, Liu J, Yang X and Chu Q:
Propofol inhibits the progression of cervical cancer by regulating
HOTAIR/miR-129-5p/RPL14 axis. Onco Targets Ther. 14:551–564.
2021.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Wu D, Zhu J, Fu Y, Li C and Wu B: LncRNA
HOTAIR promotes breast cancer progression through regulating the
miR-129-5p/FZD7 axis. Cancer Biomark. 30:203–212. 2021.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Zhan Y, Abuduwaili K, Wang X, Shen Y,
Nuerlan S and Liu C: Knockdown of long non-coding RNA HOTAIR
suppresses cisplatin resistance, cell proliferation, migration and
invasion of DDP-resistant NSCLC cells by targeting
miR-149-5p/doublecortin-like kinase 1 axis. Cancer Manag Res.
12:7725–7737. 2020.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Liang H and Peng J: LncRNA HOTAIR promotes
proliferation, invasion and migration in NSCLC cells via the CCL22
signaling pathway. PLoS One. 17(e0263997)2022.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Tang Y, Song G, Liu H, Yang S, Yu X and
Shi L: Silencing of long non-coding RNA HOTAIR alleviates
epithelial-mesenchymal transition in pancreatic cancer via the
Wnt/β-catenin signaling pathway. Cancer Manag Res. 13:3247–3257.
2021.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Dai ZY, Jin SM, Luo HQ, Leng HL and Fang
JD: LncRNA HOTAIR regulates anoikis-resistance capacity and
spheroid formation of ovarian cancer cells by recruiting EZH2 and
influencing H3K27 methylation. Neoplasma. 68:509–518.
2021.PubMed/NCBI View Article : Google Scholar
|