Introduction
Glioma is a primary tumour originating from glial
cells of the central nervous system, accounting for ~30% of all
central nervous system tumours and ~80% of all primary malignant
central nervous system tumours worldwide (1). Glioblastoma (GBM) is the most common
primary malignant tumour of the central nervous system in adults,
which originates from the neuroepithelium and accounts for 40-50%
of all intracranial tumours worldwide (2). Even with aggressive multimodal
therapy, GBM has poor prognosis, with a 5-year survival rate <5%
(3). Therefore, finding novel drug
targets and exploring the pathogenesis of the disease is of great
significance for the development of new drugs and the treatment of
GBM.
Kinesin family member 18A (KIF18A) is a member of
the kinesin-8 family, and is closely associated with cell division
(4). In recent years, the
expression and role of KIF18A in tumours have attracted increasing
attention in molecular oncology research (5). It has been shown that chromosomally
unstable tumour cells specifically require KIF18A for proliferation
(6). KIF18A mutant mice exhibit a
stable micronucleus envelope that does not promote the occurrence
of tumours (7). KIF18A gene
knockdown inhibits the proliferation, migration and invasion of
oesophageal cancer cells, while improves the sensitivity of cells
to radiotherapy (8). Stangeland
et al (9) showed that
KIF18A can be a potential therapeutic target for GBM. A relevant
clinical study demonstrated that KIF18A expression is higher in
human GBM tissues than normal tissues, and its expression is
closely associated with the recurrence of GBM in patients (10). However, to the best of our
knowledge, the specific mechanism of KIF18A in GBM has not been yet
elucidated.
Biological General Repository for Interaction
Datasets (BioGRID) and GeneMANIA databases were used to analyse the
genes interacting with KIF18A, and a binding relationship between
KIF18A and protein phosphatase 1 catalytic subunit α (PPP1CA) was
found. PPP1CA is an enzyme that is closely related to cell cycles
and it was shown that PPP1CA activates the ERK/MAPK signalling
pathway to promote the growth and metastasis of colorectal cancer
(11). Inhibition of PPP1CA can
significantly inhibit the proliferation and migration of breast
cancer cells (12). At the same
time, PPP1CA has been found to be a prognostic marker of GBM
through a clinical database analysis (13). However, to the best of our
knowledge, the regulatory mechanism of KIF18A and PPP1CA in GBM has
not been reported.
Therefore, the present study investigated the
expression of KIF18A and PPP1CA in GBM cells and further examined
the regulatory mechanism of these proteins in malignant GBM cells.
The present findings may provide a solid foundation for the
clinical treatment of GBM.
Materials and methods
Databases
The Chinese Glioma Genome Atlas (CGGA) database
(http://cgga.org.cn/analyse/RNA-data.jsp) was used to
analyse the expression levels of PPP1CA and KIF18A, survival
prognosis as well as their correlation. Pearson correlation
analysis was performed.
BioGRID (version 4.4; https://thebiogrid.org/) and GeneMANIA (https://genemania.org/) databases were used to analyse
the interaction between KIF18A and PPP1CA.
Cell culture
The normal microglia HMC3 (cat. no. CRL-3304),
glioblastoma A172 (cat. no. CRL-1620) and LN-18 (cat. no. CRL-2610)
and astrocytoma SW1783 (cat. no. HTB-13) cell lines were acquired
from the American Type Culture Collection and cultured in
Dulbecco's modified Eagle's medium (DMEM; Thermo Fisher Scientific,
Inc.) supplemented with 10% foetal bovine serum (FBS; Gibco; Thermo
Fisher Scientific, Inc.) and 1% penicillin-streptomycin (Gibco;
Thermo Fisher Scientific, Inc.) at 37˚C with 5% CO2.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from A172 cells with RNeasy
Mini Kit (Qiagen China Co., Ltd). Total RNA was reversely
transcribed into cDNA using M-MLV RTase and random primer kit
(GeneCopoeia, Inc.) according to the standard protocol. qPCR was
subsequently performed on an ABI PRISM 7900HT system (Applied
Biosystems; Thermo Fisher Scientific, Inc.) using SYBR Premix Ex
Taq™ (Takara Bio, Inc.) or TaqMan probes (cat. no. 450003; Applied
Biosystems; Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol. The thermocycling conditions were as
follows: Initial denaturation at 95˚C for 3 min; followed by 40
cycles of denaturation at 95˚C for 30 sec, annealing at 60˚C for 30
sec and extension at 72˚C for 30 sec. GAPDH was used as internal
reference and relative mRNA expression levels were calculated using
the 2-ΔΔCq method (14). The following primer pairs were used
for qPCR: KIF18A forward, 5'-GAGAGGCACATGAAGAGAAGT-3' and reverse,
5'-AAGTCCATGAACGACCACCC-3'; PPP1CA forward,
5'-CAGGGTCCTGACACCCCATT-3' and reverse, 5'-AGGTAAAAGAGACGCCACGG-3';
GAPDH forward, 5'-AATGGGCAGCCGTTAGGAAA-3' and reverse,
5'-GCGCCCAATACGACCAAATC-3'.
Western blotting
Total protein was extracted from A172 cells using
RIPA lysis buffer (Beyotime Institute of Biotechnology). Total
protein concentration was quantified with a BCA assay kit (Beyotime
Institute of Biotechnology) and equal amount of proteins (20 µg per
lane) was separated using 10% SDS-PAGE and subsequently transferred
onto PVDF membranes (MilliporeSigma) which were blocked with 5%
non-fat milk for 2 h at room temperature. The membranes were
incubated with primary antibodies targeting KIF18A (1:2,000; cat.
no. ab72417; Abcam), CDK1 (1:10,000; cat. no. ab133327; Abcam),
cyclin B (1:50,000; cat. no. ab32053; Abcam), MMP9 (1:1,000; cat.
no. ab76003; Abcam), MMP2 (1:1,000; cat. no. ab92536; Abcam),
PPP1CA (1:20,000; cat. no. ab52619; Abcam), or GAPDH (1:1,000; cat.
no. ab9485; Abcam) overnight at 4˚C. Following the rinse with PBS
for three times, membranes were incubated with HRP-conjugated
secondary antibodies (cat. no. ab6759; 1:5,000; Abcam) for 1.5 h.
The protein bands were visualized using ECL detection reagent
(MilliporeSigma) and analysed with ImageJ software 1.8.0 (National
Institutes of Health). GAPDH was used as internal reference.
Cell transfection
The two GV248-green fluorescent protein (GFP)-short
harpin (sh)RNA-KIF18A lentiviral vectors, namely
GV248-GFP-sh-KIF18A#1 and GV248-GFP-sh-KIF18A#2, the negative
control (NC) GV248-GFP-sh-NC lentiviral vector, PPP1CA-specific
pcDNA overexpression vector (Oe-PPP1CA) and pcDNA3.1 empty vector
were constructed by Shanghai GeneChem Co., Ltd. A total of 100 nM
plasmids were transfected into A172 cells at 37˚C for 24 h using
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol. After
24 h, the harvested A172 cells were adopted for follow-up
experiments. The shRNA targeting sequences were as follows:
sh-KIF18A#1, 5'-TTGTTTCAGACTCACATATAA-3'; sh-KIF18A#2,
5'-AGTCCTGAGAGGAAGTCTTAA-3'; and sh-NC, 5'-TTCTCCGAACGTGTCACGT-3'.
Cells were then divided into the following groups: Control
(incubated in DMEM); sh-NC (transfected with GV248-GFP lentiviral
vector); sh-KIF18A (transfected with GV248-GFP-shRNA-KIF18A#2
lentiviral vector); sh-KIF18A + Oe-NC (transfected with
GV248-GFP-shRNA-KIF18A#2 lentiviral vector and Oe-NC empty vector);
and sh-KIF18A + Oe-PPP1CA (transfected with
GV248-GFP-shRNA-KIF18A#2 lentiviral vector and Oe-PPP1CA
vector).
Cell Counting Kit-8 (CCK-8) assay
A172 cells were plated in 96-well plates at the
density of 2x103 cells/well and incubated for 24, 48 and
72 h. Subsequently, the cells were incubated with CCK-8 solution
(MilliporeSigma) at 37˚C for 3 h. The absorbance of each well at
450 nm was measured with a multimode microplate reader (Synergy H1;
BioTek Instruments, Inc.).
Cell cycle analysis
Flow cytometry was used for cell cycle analysis. The
transfected A172 cells at a density of 4x105 cells per
well were fixed with 75% ethanol at 4˚C for 24 h, followed by the
staining with PI solution [200 µg/ml RNase A, 50 µg/ml PI and 0.1%
(v/v) Triton X-100 in PBS] for 30 min at room temperature. Finally,
the cell cycle was analysed using CellQuest software v.2.0 (BD
Biosciences) and ModFit LT v.2.0 (Verity Software House, Inc.).
Transwell assay
Transwell assays were performed using Transwell
inserts (8-µm pore size; Costar; Corning, Inc.) precoated with 50
µl Matrigel (BD Biosciences). The inserts were put into 24-well
plates. A172 cells that injected into the serum-free medium at a
density of 2x104 cells were seeded into the upper
chambers while the lower chambers were filled with DMEM
supplemented with 10% FBS. Following incubation at 37˚C for 24 h,
the residual cells on upper chambers were removed, and the cells
under the surface were stained with 0.5% crystal violet for 10 min
at room temperature and then fixed with 70% ethanol at 4˚C. The
number of cells in five randomly selected fields were counted with
a light microscope (Olympus Corporation) and were analyzed using
ImageJ 1.8.0 software (National Institutes of Health).
Wound healing assay
A172 cells were seeded in 6-well plates at the
density of 5x105 cells/well. After 95% cell confluence
was achieved, artificial wounds were gently made using a 200-µl
sterile pipette tip and confluent monolayers for wounding were
yielded. The serum-starved culture medium was replaced with DMEM
supplemented with 10 mg/ml mitomycin C (Sigma-Aldrich; Merck KGaA).
Cells in the scratched area were imaged at 0 and 24 h using a light
microscope (Thermo Fisher Scientific, Inc.). The relative migration
rate (%)=(wound width at 0 h-wound width at 24 h)/wound width at 0
h x100.
Immunoprecipitation
A172 cell lysates were prepared in RIPA buffer. The
supernatant was collected after centrifugation at 13,000 x g for 10
min at 4˚C. Subsequently, 0.2 mg protein A agarose beads (cat. no.
20366; Thermo Fisher Scientific, Inc.) washed with 100 µl PBS
buffer were added to 500 µg lysis buffer and incubated with 2 µg
IgG antibody (1:1,000; cat. no. ab172730; Abcam) or PPP1CA antibody
(1:500; cat. no. GTX105255; GeneTex, Inc.) or KIF18A antibody
(1:500; cat. no. ab72417; Abcam) overnight at 4˚C with slow
shaking. Following the IP reaction and centrifugation at 1,000 x g
at 4˚C for 2 min, the agarose beads were washed with lysis buffer
and boiled for 5 min at 100˚C by adding 15 µl 2X SDS loading dye
before being subjected to western blot analysis as
aforementioned.
Statistical analysis
The data from three independent experiments are
presented as the mean ± standard deviation and were evaluated using
SPSS 19 software (IBM Corp.). One-way ANOVA followed by Tukey's
post hoc test was used for statistical analysis. P<0.05 was
considered to indicate a statistically significant difference.
Results
KIF18A is upregulated in GBM
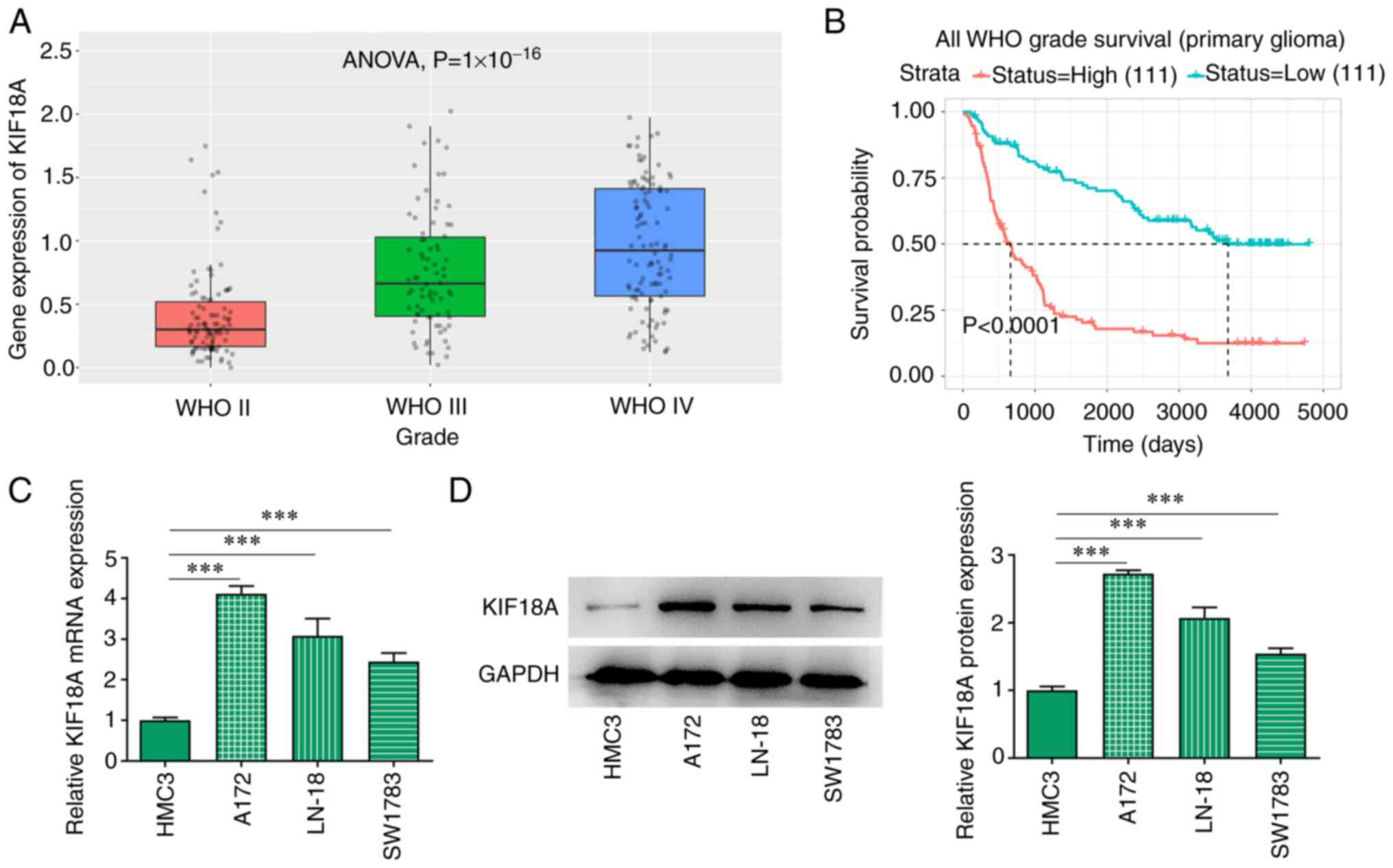
The CGGA database analysis showed that the
expression of KIF18A was markedly increased in WHO IV group
compared with the WHO II group (Fig.
1A). KIF18A expression was closely associated with the
prognosis of patients with GBM and the higher the KIF18A
expression, the lower the survival probability of patients with GBM
(Fig. 1B). RT-qPCR and western
blotting results showed that the mRNA and protein expression of
KIF18A in GBM cell lines was significantly increased compared with
the HMC3 group (Fig. 1C and
D). The expression level of KIF18A
in A172 cells was higher than that in LN-18 and SW1783 cells, and
consequently A172 cells were selected for subsequent
experiments.
Silencing KIF18A reduces cell
proliferation and induces G2/M cycle arrest in A172
cells
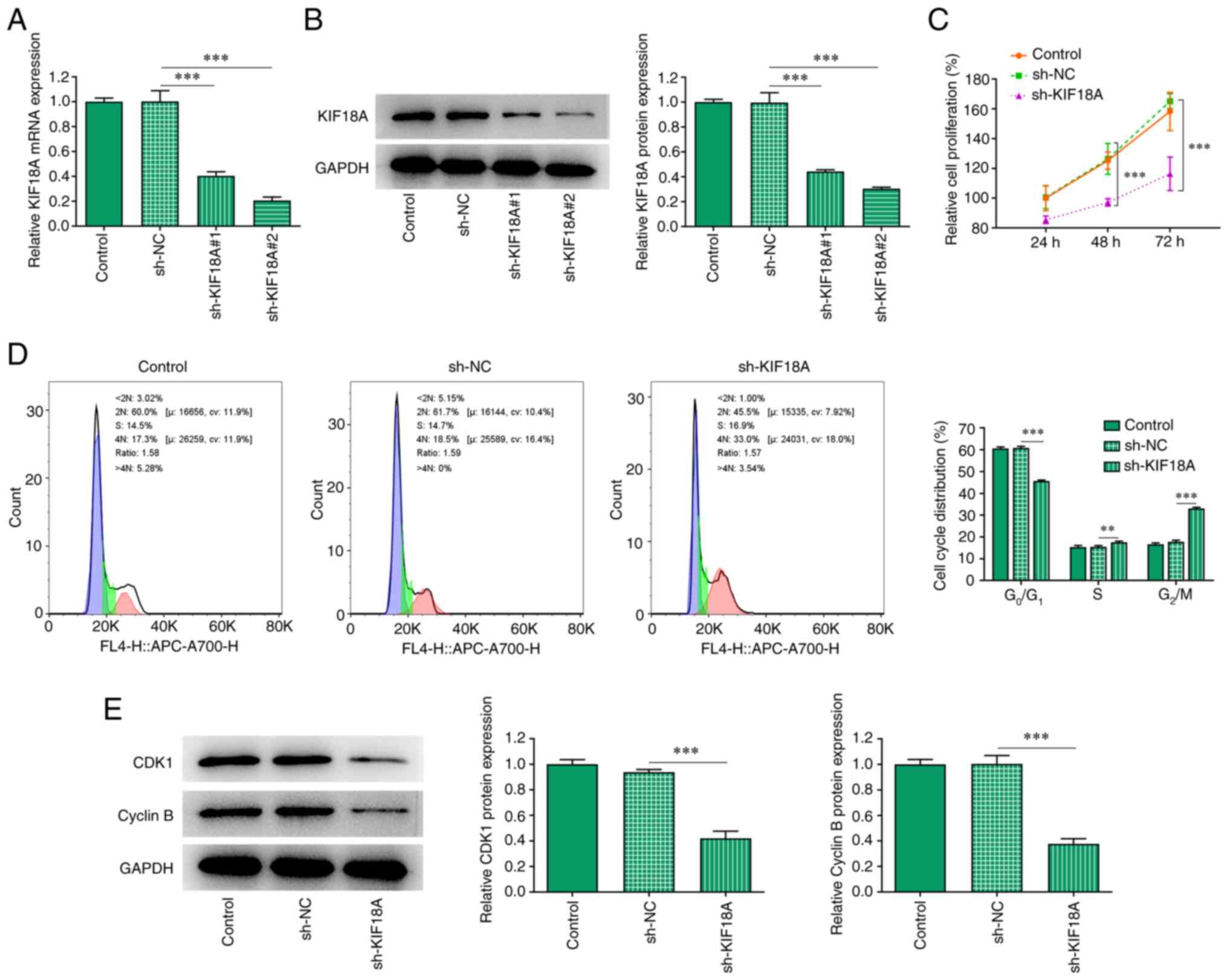
After silencing KIF18A expression, RT-qPCR and
western blotting were used to estimate the knockdown efficiency
(Fig. 2A and B). Compared with the sh-KIF18A#1 group,
KIF18A exhibited lower mRNA and protein expression in the
sh-KIF18A#2 group, likely indicating an improved transfection
efficacy for sh-KIF18A#2. Therefore, sh-KIF18A#2 was chosen for
subsequent experiments. CCK-8 was utilized to measure the cell
proliferation rate and the results showed that the proliferation of
the sh-KIF18A group was significantly decreased compared with sh-NC
(Fig. 2C). The cell cycle was
analysed using flow cytometry, and the results showed that compared
with sh-NC, the G0/G1 proportion in the
sh-KIF18A group was significantly decreased and the G2/M
proportion was significantly increased (Fig. 2D). Western blot analysis was used
to examine the expression of the cycle-related proteins
cyclin-dependent kinase 1 (CDK1) and cyclin B. The results showed
that KIF18A silencing could significantly inhibit the expression of
CDK1 and cyclin B (P<0.001; Fig.
2E).
Silencing of KIF18A inhibits the
migration and invasion of A172 cells
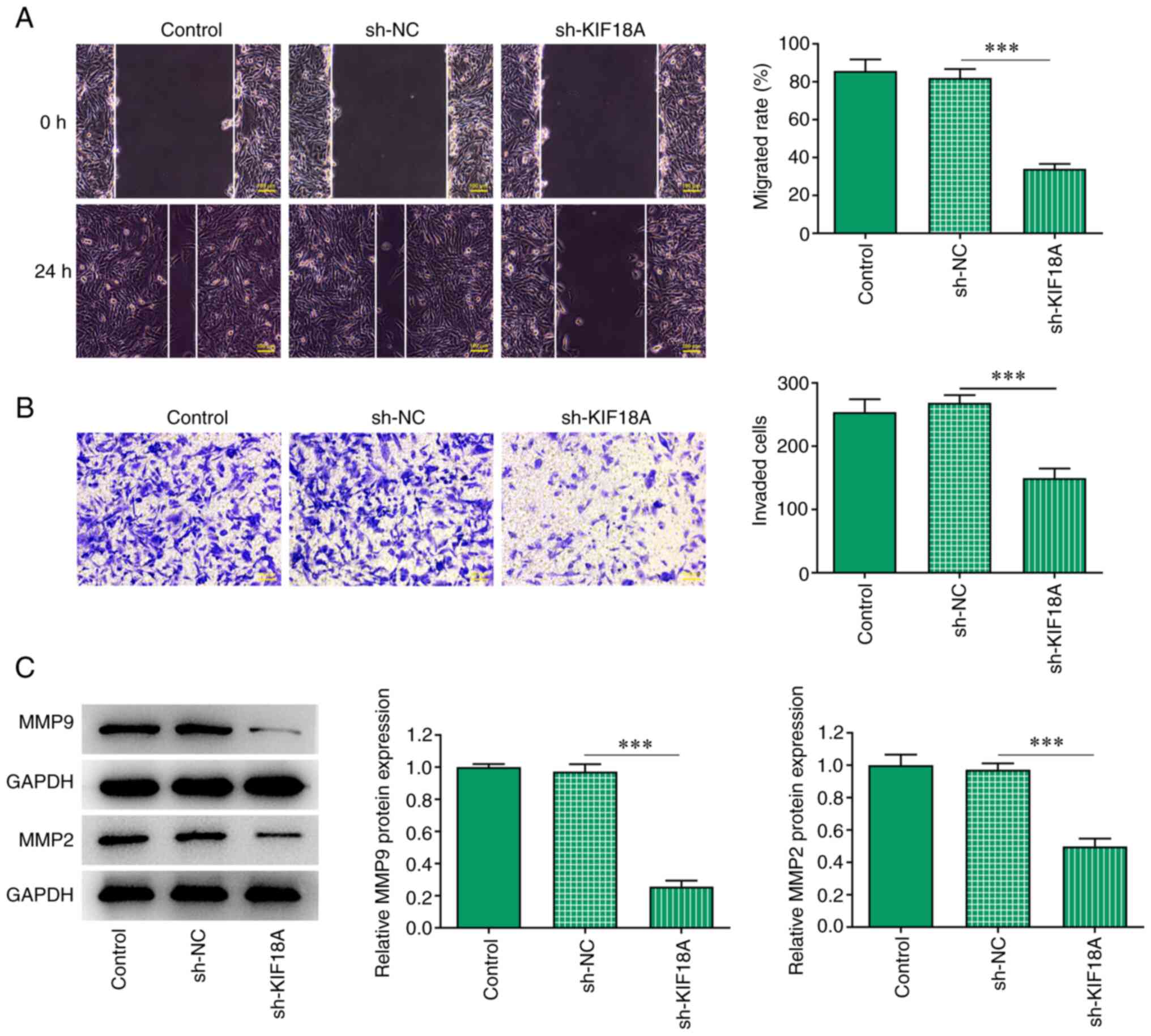
Transwell and wound healing assays were performed to
assess cell invasion and migration, and the results showed that the
migration and invasion of A172 cells were significantly decreased
after KIF18A knockdown (P<0.001 vs. sh-NC; Fig. 3A and B). Western blotting results showed that
the expression of the migration-related proteins MMP2 and MMP9 were
decreased significantly after KIF18A silencing (P<0.001 vs.
sh-NC; Fig. 3C).
KIF18A interacts with PPP1CA in
GBM
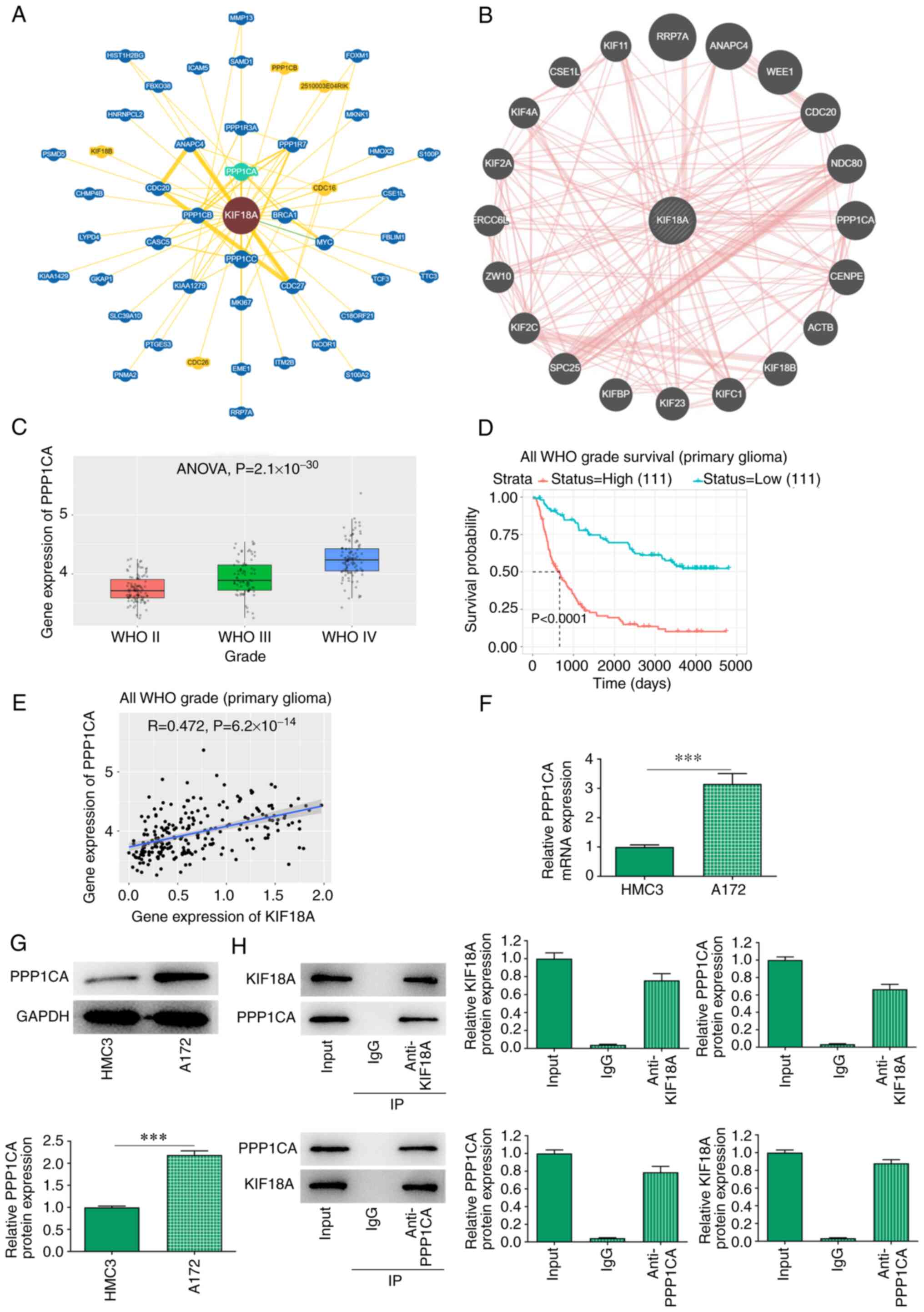
BioGRID and GeneMANIA database analysis revealed a
potential interaction between KIF18A and PPP1CA (Fig. 4A and B). Through the CGGA database, it was
found that the expression of PPP1CA was significantly increased in
WHO IV group compared with the WHO II group (Fig. 4C). The expression of PPP1CA was
closely related to the prognosis of GBM and the higher the KIF18A
expression, the lower the survival probability of patients with GBM
(Fig. 4D). In addition, CGGA
database also revealed a highly positive correlation between KIF18A
and PPP1CA in GBM (R=0.472, P=6.2x10-14; Fig. 4E). RT-qPCR and western blot
analysis showed that PPP1CA mRNA and protein expression were
significantly increased in A172 cells compared with that in HMC3
cells (P<0.001; Fig. 4F and
G). Co-immunoprecipitation assay
also verified the targeted binding relationship between KIF18A and
PPP1CA (Fig. 4H).
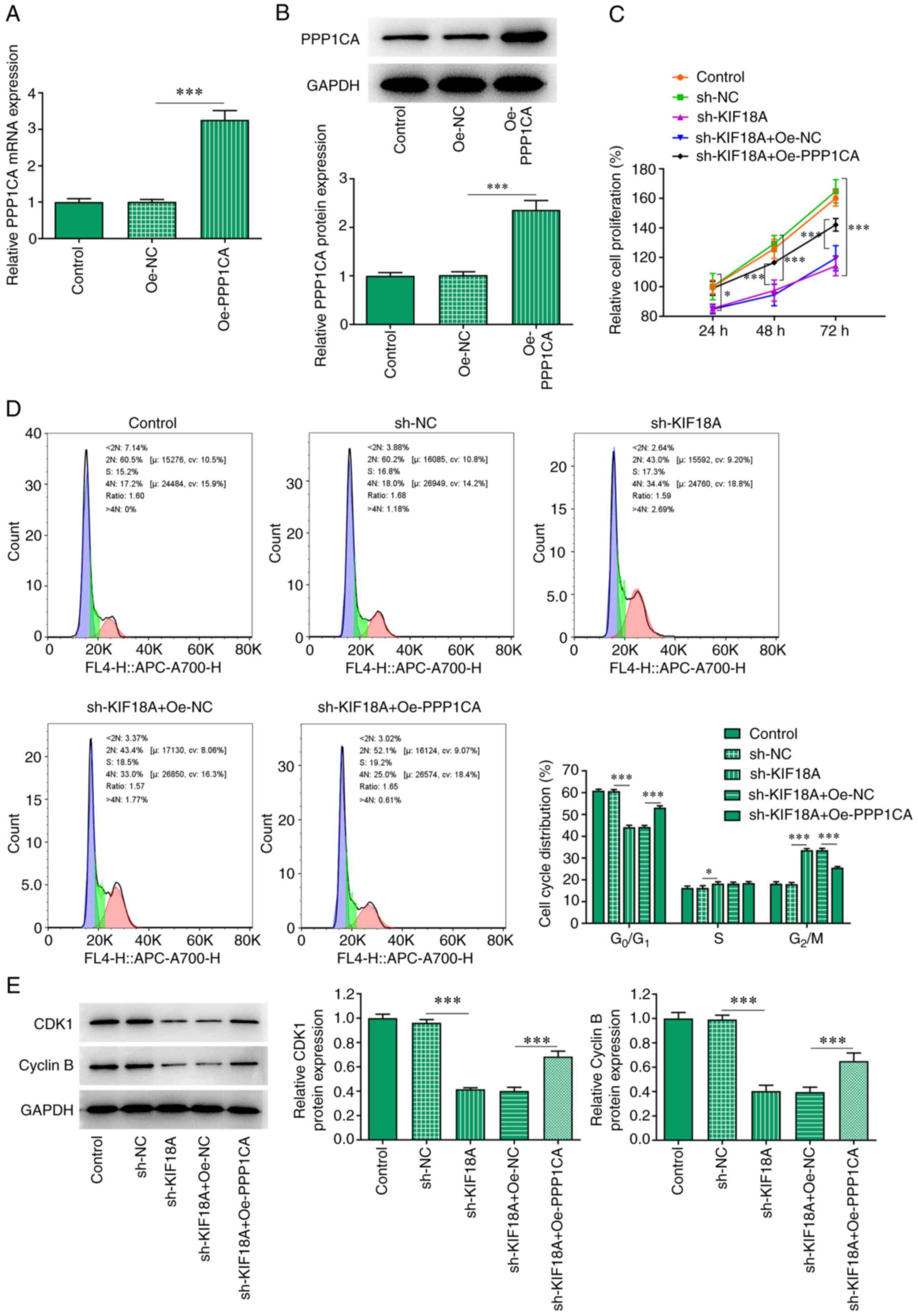
Overexpression of PPP1CA reduces the
effect of KIF18A knockdown on the proliferation and G2/M
cycle arrest of A172 cells
The Oe-PPP1CA vector was constructed and the
transfection efficiency was examined using RT-qPCR and western blot
analysis (Fig. 5A and B). Cells were divided into control,
sh-NC, sh-KIF18A, sh-KIF18A + Oe-NC and sh-KIF18A + Oe-PPP1CA
groups. CCK-8 results showed that KIF18A knockdown reduced the
proliferation of A172 cells compared with the sh-NC group.
Nevertheless, the reduced proliferation in KIF18A-depleted A172
cells was significantly increased by PPP1CA overexpression compared
with that of sh-KIF18A + Oe-NC group (Fig. 5C). As presented in Fig. 5D, KIF18A interference decreased the
number of cells at G0/G1 and S phase but
increased that of cells at G2/M phase relative with the
sh-NC, whereas PPP1CA overexpression exhibited the opposite impacts
on them, evidenced by an increased number of cells at
G0/G1 and S phase and decreased number of
cells at G2/M phase relative with the sh-KIF18A + Oe-NC
group (Fig. 5D). Moreover, results
obtained from western blot analysis demonstrated the declined
contents of CDK1 and cyclin B in A172 cells resulted from KIF18A
knockdown were enhanced after overexpressing PPP1CA expression in
contrast with those in sh-KIF18A + Oe-NC group (Fig. 5E).
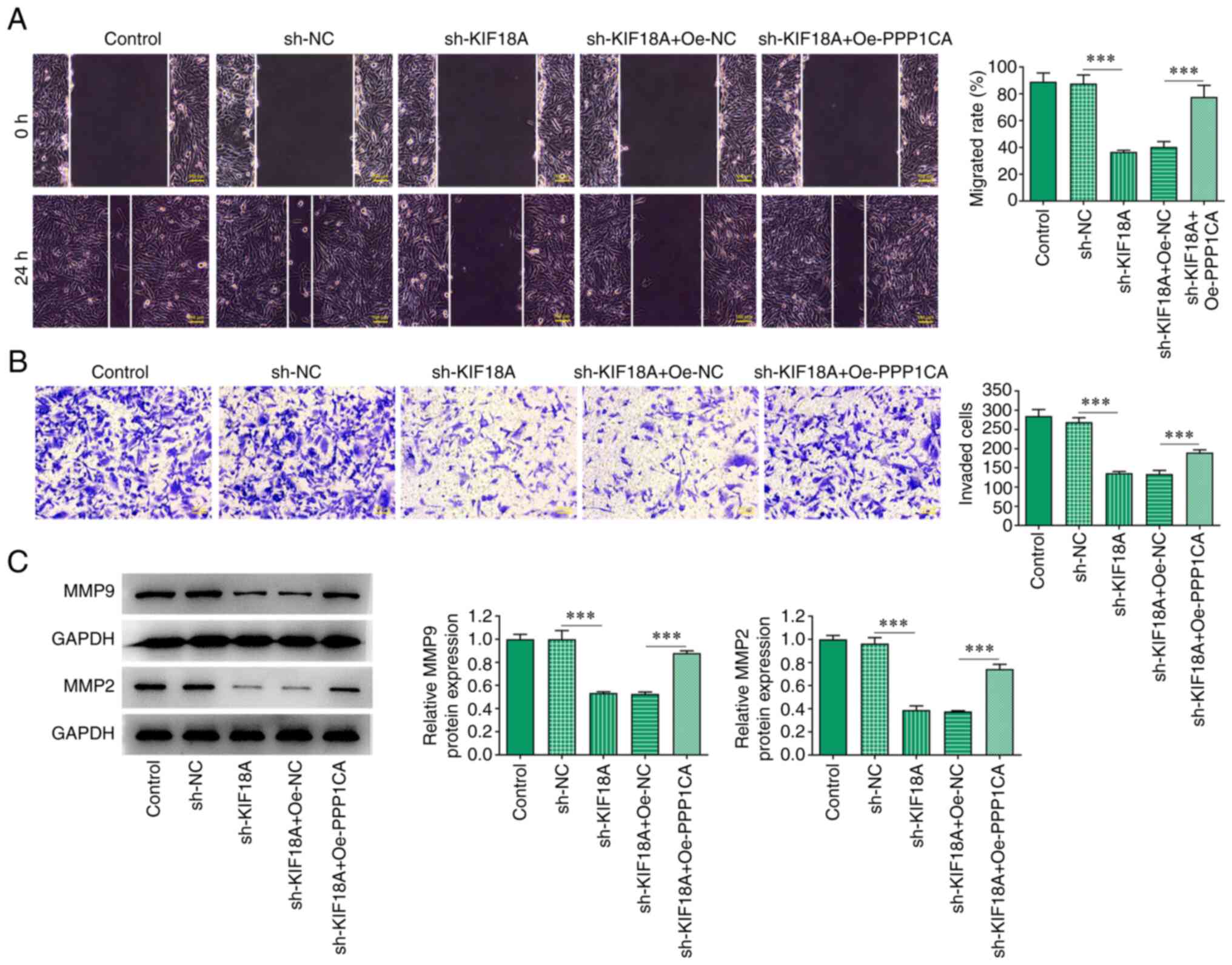
Overexpression of PPP1CA reduces the
effect of KIF18A knockdown on the migration and invasion of A172
cells
Wound healing and Transwell assays showed that
compared with the sh-KIF18A + Oe-Nc group, cell migration and
invasion were significantly increased in the sh-KIF18A + Oe-PPP1CA
group (Fig. 6A and B). Western blotting results showed that
MMP9 and MMP2 expression was significantly increased in the
sh-KIF18A + Oe-PPP1CA group compared with that in the sh-KIF18A +
Oe-NC group (P<0.001; Fig.
6C).
Discussion
The incidence of GBM has been increasing annually
lately, whereas the effective treatment methods for GBM were not
improved, with radiotherapy and chemotherapy still being the
standard treatment for patients with GBM after surgical resection
(15). Moreover, the use of
chemotherapy drugs only increased the median survival time of
patients with GBM by 2-3 months, while it did not significantly
prolong the survival time of patients (16). Therefore, it is of great urgency to
find more effective and sensitive therapeutic targets to improve
the treatment of GBM.
Through CGGA database analysis, the present study
found that the expression of KIF18A in patients with GBM is
significantly increased in the WHO IV group compared with the WHO
II group and is closely related to the prognosis of GBM. The
present study revealed that the expression of KIF18A is also
significantly increased in GBM cell lines when compared with the
normal microglia HMC3. Previously, it has been shown that KIF18A
can be used as a potential therapeutic target for GBM (9). Therefore, it is reasonable to
hypothesize that the abnormal expression of KIF18A is closely
related to the occurrence and development of GBM. A previous study
has shown that knockdown of KIF18A could suppress the
proliferation, migration and invasion of oesophageal cancer cells
(8). In hepatocellular carcinoma
cells, KIF18A may promote cell proliferation, invasion and
metastasis by promoting the cell cycle and Akt and MMP7/MMP9
related signalling pathways (17).
In prostate cancer, KIF18A silencing in PC-3 prostate cells has
been shown to significantly inhibit cell proliferation and
metastasis (18). These results
indicated that KIF18A may have an oncogenic role in multiple
tumours. The present results showed that KIF18A knockdown decreased
the proliferation, invasion and migration of GBM A172 cells but
induced the cell cycle. These results indicated that KIF18A may
have an oncogenic role in GBM.
The current study further explored the mechanism of
KIF18A in regulating GBM. Firstly, the interaction between KIF18A
and PPP1CA was investigated through BioGRID and GeneMANIA database
analysis. Moreover, CGGA database analysis showed a high
correlation between PPP1CA and KIF18A in GBM. Furthermore, the
interaction between PPP1CA and KIF18A was experimentally
investigated and demonstrated. A previous study has shown that the
transcriptional level of PPP1CA in pancreatic cancer is higher than
that in normal pancreas and the high transcriptional level of
PPP1CA is associated with the poor survival rate of patients with
pancreatic cancer (19). In the
current study, PPP1CA was also found to be associated with GBM
survival prognosis, which is consistent with the study by Shastry
et al (13). It has been
shown that PPP1CA deficiency may significantly inhibit the
proliferation and migration of breast cancer cells (12). PPP1CA is upregulated in clinical
specimens of maxillary sinus squamous cell carcinoma. Silencing
PPP1CA gene can significantly inhibit the proliferation and
invasion of cancer cells (20).
Therefore, PPP1CA shows oncogenic characteristics. However, it has
also been reported that PPP1CA downregulation could significantly
promote the proliferation, migration and invasion of gastric cancer
cells (21). Therefore, PPP1CA can
also act as a tumour suppressor gene. This raises questions about
the role of PPP1CA in GBM. In this context, the present study found
that PPP1CA expression is significantly elevated in A172 cells
compared with that in HMC3 cells. Overexpression of PPP1CA
countervailed the inhibitory effects of KIF18A knockdown on the
proliferation, cell cycle arrest, migration and invasion of GBM
cells, suggesting that PPP1CA may act as an oncogenic gene in
GBM.
The present study only detected the expression of
PPP1CA in GMB A172 cells and investigated its regulatory mechanism.
In the future, the mechanism of PPP1CA and KIF18A in different
stages of GBM should be further examined.
In conclusion, the present study results
demonstrated that KIF18A interacts with PPP1CA to promote the
proliferation, cell cycle arrest, migration and invasion of GBM
A172 cells.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
JY and XL conceived and designed the study. QZ, ZY,
JS, LZ, YY and XW performed the experiments. JY and XL analysed the
experimental data and wrote and revised the manuscript. JY and XL
confirm the authenticity of all the raw data. All authors have read
and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Davis ME: Epidemiology and overview of
gliomas. Semin Oncol Nurs. 34:420–429. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Preusser M, Lim M, Hafler DA, Reardon DA
and Sampson JH: Prospects of immune checkpoint modulators in the
treatment of glioblastoma. Nat Rev Neurol. 11:504–514.
2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Chen R, Smith-Cohn M, Cohen AL and Colman
H: Glioma subclassifications and their clinical significance.
Neurotherapeutics. 14:284–297. 2017.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Malaby HL, Lessard DV, Berger CL and
Stumpff J: KIF18A's neck linker permits navigation of
microtubule-bound obstacles within the mitotic spindle. Life Sci
Alliance. 2(e201800169)2019.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Tao BY, Liu YY, Liu HY, Zhang ZH, Guan YQ,
Wang H, Shi Y and Zhang J: Prognostic biomarker KIF18A and its
correlations with immune infiltrates and mitosis in glioma. Front
Genet. 13(852049)2022.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Marquis C, Fonseca CL, Queen KA, Wood L,
Vandal SE, Malaby HLH, Clayton JE and Stumpff J: Chromosomally
unstable tumour cells specifically require KIF18A for
proliferation. Nat Commun. 12(1213)2021.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Sepaniac LA, Martin W, Dionne LA, Stearns
TM, Reinholdt LG and Stumpff J: Micronuclei in Kif18a mutant mice
form stable micronuclear envelopes and do not promote
tumorigenesis. J Cell Biol. 220(e202101165)2021.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Qian LX, Cao X, Du MY, Ma CX, Zhu HM, Peng
Y, Hu XY, He X and Yin L: KIF18A knockdown reduces proliferation,
migration, invasion and enhances radiosensitivity of oesophageal
cancer. Biochem Biophys Res Commun. 557:192–198. 2021.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Stangeland B, Mughal AA, Grieg Z, Sandberg
CJ, Joel M, Nygard S, Meling T, Murrell W, Vik Mo EO and Langmoen
IA: Combined expressional analysis, bioinformatics and targeted
proteomics identify new potential therapeutic targets in
glioblastoma stem cells. Oncotarget. 6:26192–26215. 2015.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Wang LB, Zhang XB, Liu J and Liu QJ: The
proliferation of glioblastoma is contributed to kinesin family
member 18A and medical data analysis of GBM. Front Genet.
13(858882)2022.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Sun H, Ou B, Zhao S, Liu X, Song L, Liu X,
Wang R and Peng Z: USP11 promotes growth and metastasis of
colorectal cancer via PPP1CA-mediated activation of ERK/MAPK
signalling pathway. EBioMedicine. 48:236–247. 2019.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Xie W, Sun Y, Zeng Y, Hu L, Zhi J, Ling H,
Zheng X, Ruan X and Gao M: Comprehensive analysis of PPPCs family
reveals the clinical significance of PPP1CA and PPP4C in breast
cancer. Bioengineered. 13:190–205. 2022.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Shastry AH, Thota B, Srividya MR,
Arivazhagan A and Santosh V: Nuclear Protein Phosphatase 1 α (PP1A)
expression is associated with poor prognosis in p53 expressing
glioblastomas. Pathol Oncol Res. 22:287–292. 2016.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Hombach-Klonisch S, Mehrpour M, Shojaei S,
Harlos C, Pitz M, Hamai A, Siemianowicz K, Likus W, Wiechec E,
Toyota BD, et al: Glioblastoma and chemoresistance to alkylating
agents: Involvement of apoptosis, autophagy, and unfolded protein
response. Pharmacol Ther. 184:13–41. 2018.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Stupp R, Mason WP, van den Bent MJ, Weller
M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn
U, et al: Radiotherapy plus concomitant and adjuvant temozolomide
for glioblastoma. N Engl J Med. 352:987–996. 2005.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Luo W, Liao M, Liao Y, Chen X, Huang C,
Fan J and Liao W: The role of kinesin KIF18A in the invasion and
metastasis of hepatocellular carcinoma. World J Surg Oncol.
16(36)2018.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Zhang H, Shen T, Zhang Z, Li Y and Pan Z:
Expression of KIF18A is associated with increased tumor stage and
cell proliferation in prostate cancer. Med Sci Monit. 25:6418–6428.
2019.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Hang J, Lau SY, Yin R, Zhu L, Zhou S, Yuan
X and Wu L: The role of phosphoprotein phosphatases catalytic
subunit genes in pancreatic cancer. Biosci Rep.
41(BSR20203282)2021.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Nohata N, Hanazawa T, Kikkawa N, Sakurai
D, Fujimura L, Chiyomaru T, Kawakami K, Yoshino H, Enokida H,
Nakagawa M, et al: Tumour suppressive microRNA-874 regulates novel
cancer networks in maxillary sinus squamous cell carcinoma. Br J
Cancer. 105:833–841. 2011.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Verdugo-Sivianes EM, Navas L,
Molina-Pinelo S, Ferrer I, Quintanal-Villalonga A, Peinado J,
Garcia-Heredia JM, Felipe-Abrio B, Muñoz-Galvan S, Marin JJ, et al:
Coordinated downregulation of Spinophilin and the catalytic
subunits of PP1, PPP1CA/B/C, contributes to a worse prognosis in
lung cancer. Oncotarget. 8:105196–105210. 2017.PubMed/NCBI View Article : Google Scholar
|