Introduction
Psoriasis is an immune-mediated, polygenic
dermatological condition characterized by the formation of scaly,
erythematous plaques or lesions. In some cases, patients may also
experience psoriatic arthritis as a comorbid condition (1). Additionally, intrinsic factors such
as hypertension, diabetes mellitus, obesity and mental stress,
along with external environmental factors including sun exposure,
trauma and infections, can exacerbate psoriasis. This condition
markedly impacts the quality of life of patients, a number of whom
experience feelings of shame due to the visible effects of the
disease. Consequently, >5% of patients may suffer from
depression and suicidal ideations (2,3).
Currently, the primary therapeutic strategies for psoriasis include
immunosuppressive therapy (such as methotrexate and cyclosporine)
(4) and biological agent therapy
(such as secukinumab and ustekinumab), which only offer transient
symptomatic relief (5). However,
these treatments are often associated with relapse upon
discontinuation, and prolonged use can lead to adverse effects such
as infection and hepatotoxicity. Therefore, developing innovative,
effective and safe anti-psoriasis treatments is a pressing
need.
Wogonin, a naturally occurring flavonoid derived
from the root of Scutellaria baicalensis Georgi, has gained
attention due to its broad pharmacological properties, including
anti-inflammatory, antioxidant and antiproliferative effects.
Chemically, wogonin (5,7-dihydroxy-8-methoxyflavone) has a
molecular formula of C16H12O5 and
a molecular weight of 284.26 g/mol. Although wogonin exhibits low
aqueous solubility, it exhibits good solubility in both DMSO and
ethanol, consistent with its lipophilic nature (predicted logP
value of ~2.7) (6).
Pharmacokinetically, wogonin exhibits moderate oral bioavailability
and has been shown to effectively penetrate skin tissue, supporting
its topical application in dermatological disorders (7). Used in Traditional Chinese Medicine
to treat inflammatory and infectious diseases, wogonin has
demonstrated therapeutic potential in various dermatological
conditions, such as atopic dermatitis, ultraviolet-induced skin
damage and psoriasis, by modulating inflammatory signaling pathways
and suppressing keratinocyte hyperproliferation (8-10).
Psoriasis is characterized by aberrant keratinocyte
proliferation, sustained activation of pro-inflammatory cytokines
(including TNF-α, IL-17 and IL-23) and dysregulation of innate
immune responses (11,12). In vitro research using human
keratinocyte (HaCaT) models has revealed that wogonin effectively
attenuates psoriasis-like inflammation by inhibiting the PI3K/AKT
pathway, thereby reducing the expression of inflammatory mediators,
such as TNF-α, IL-1β and IL-6(10). Furthermore, wogonin suppresses NLR
family pyrin domain containing 3 inflammasome activation, caspase-1
cleavage and gasdermin-D-mediated pyroptosis in keratinocytes,
outlining its dual role in mitigating both hyperproliferation and
inflammatory cascades in psoriatic pathogenesis (13). These findings suggest that wogonin
is a promising candidate for psoriasis therapy. However, despite
notable evidence from cellular models, the therapeutic efficacy and
mechanistic actions of wogonin in vivo remain unexplored.
Animal studies are therefore key in the validation of the
anti-psoriasis effects of wogonin, assessing its systemic safety
and elucidating tissue-specific interactions that cannot be fully
replicated in vitro. Addressing this gap is key for
translating preclinical findings into clinical applications.
Network pharmacology is an effective methodology for
investigating and elucidating pharmacological mechanisms,
encompassing chemical informatics, bioinformatics, network biology
and pharmacology (14). In the
present study, a network pharmacology approach was employed to
predict effective molecular targets and potential mechanisms of
wogonin underlying the treatment of psoriasis. Additionally, the
effects of wogonin on imiquimod (IMQ)-induced psoriatic lesions of
mice were examined and the targets of wogonin were validated. The
present study not only bridges the knowledge gap between cellular
and animal models but also provides a foundation for developing
wogonin-based therapies for psoriasis.
Materials and methods
Network pharmacological and molecular
docking analysis. Wogonin target screening
SwissTargetPrediction (15) (http://www.swisstargetprediction.ch/), PharmMapper
(16) (version 2017; http://www.lilab-ecust.cn/pharmmapper/),
Similarity Ensemble Approach (17)
(https://sea.bkslab.org/) and SuperPred databases
(18) (https://prediction.charite.de/index.php) were used to
predict potential targets of wogonin. The UniProt database
(https://www.uniprot.org/) was used to verify the
target information. Duplicate entries were removed to obtain the
final list of wogonin targets.
Identifying psoriasis differentially expressed
genes (DEGs) and obtaining the wogonin-psoriasis intersection
targets. GSE121212(19) and
GSE13355(20) datasets were
obtained from the Gene Expression Omnibus (GEO) database
(https://www.ncbi.nlm.nih.gov/geo/).
Each dataset contained >40 samples, with each group consisting
of >20 control and 20 psoriasis samples. GSE121212 included gene
expression profiles from 28 patients with psoriasis and 38 healthy
controls, while GSE13355 included gene expression profiles from 58
patients with psoriasis and 64 healthy skin samples. GEO2R
(https://www.ncbi.nlm.nih.gov/geo/geo2r/) was used to
analyze the DEGs of each dataset online and volcano plots of each
dataset were generated using the online platform SRplot (https://www.bioinformatics.com.cn) (21) for data analysis and visualization.
Significantly DEGs were identified using the thresholds of an
adjusted P-value <0.05 and |log2 fold change|>1.
Intersection target acquisition and action
network construction. Wogonin targets and psoriasis DEGs were
imported into Venny 2.1.0 software (https://bioinfogp.cnb.csic.es/tools/venny/index.html)
for common target identification. Venn diagrams of the intersecting
targets of wogonin in psoriasis were generated. These selected
targets were then imported into Cytoscape 3.10.1(22) for visualization and a
drug-target-disease action network diagram was constructed.
Construction of protein-protein interaction (PPI)
networks. A total of 30 potential targets were imported into
the Search Tool for the Retrieval of Interacting Genes/Proteins
(STRING) database, with the species set to human and the confidence
level set to 0.4, and the options set to ‘Hide disconnected nodes
in the network’. This allowed for the retrieval of protein
interaction network data after the removal of isolated proteins.
The data were then imported into Cytoscape 3.10.1 to construct the
PPI network. The degree value (the number of direct interactions
for a given protein node) of each target was analyzed using the
‘CentiScaPe2.2’ plug-in and the targets were ranked in descending
order of degree value to obtain relevant information regarding the
core targets. The ‘cytoHubba’ plug-in was employed to further
analyze the PPI network and determine the core targets, with the
top 10 targets selected based on their Matthews Correlation
Coefficient (MCC) values.
Gene Ontology (GO) and Kyoto Encyclopedia of
Genes and Genomes (KEGG) enrichment analysis. The Database for
Annotation, Visualization and Integrated Discovery (DAVID;
https://davidbioinformatics.nih.gov/)
(23) was employed with the
following parameters: i) ‘OFFICIAL_GENE_SYMBOL’ for the identifier;
ii) ‘Homo sapiens’ for the species; and iii) ‘gene list’ for the
submission list. The GO and KEGG enrichment analysis data of the
core targets were collected in an output table. The top 10 most
significant results for each category, including biological process
(BP), cellular component (CC) and molecular function (MF), were
selected for inclusion in the GO analysis. A total of 20 potential
signaling pathways, were obtained in descending order of count
value (P<0.05) and used for the analysis of KEGG pathways. The
resulting data were imported into the SRplot platform to generate a
visual representation of the GO function in conjunction with the
KEGG pathway enrichment analysis data forming the corresponding
enrichment bubble plot.
Molecular docking. Molecular docking was
performed using AutoDockTools 1.5.6(24). The X-ray crystal structures of the
key targets were obtained from the Protein Data Bank (PDB)
(www.rcsb.org). The 2D structures of the active
compounds were retrieved from PubChem (https://pubchem.ncbi.nlm.nih.gov/). The structures
were processed using AutoDockTools 1.5.6, including the removal of
water molecules, the addition of charges and non-polar hydrogen
atoms. Both the target and compound structures were saved in PDBQT
format. Molecular docking was performed using AutoDock Vina
software (25) and good affinity
conformations (affinity <-7 kcal/mol) were analyzed and
visualized using AutoDockTools 1.5.6 and PyMOL (26).
Experimental verification. Animal
modeling and group intervention
A total of 15 male BALB/c mice, aged 6-8 weeks and
weighing 20-22 g, were obtained from SPF (Beijing) Biotechnology
Co., Ltd. [animal license no. SCXK (Beijing) 2019-0010]. The mice
were maintained under the following conditions: Temperature was
controlled at 22±2˚C, relative humidity was 50±10%, and a 12-h
light/12-h dark cycle was maintained. All mice had free access to
standard rodent chow and autoclaved drinking water. Male mice were
selected for the present study to avoid potential fluctuations in
immune response and disease severity associated with the female
estrous cycle, thereby minimizing experimental variables, an
approach consistent with numerous previous studies utilizing the
IMQ-induced psoriasis model (27-29).
The mice were randomly assigned to three groups, a
control group, an IMQ group and an IMQ + wogonin group. The dorsal
fur of all mice was removed to create an exposed area measuring
~2x3 cm. Subsequently, 42 mg IMQ cream (Sichuan Mingxin
Pharmaceutical Co., Ltd.) was applied daily as a modeling inducer
for a total of 7 consecutive days (30), while saline was applied to the
control group. From days 1-7, the IMQ and IMQ + wogonin groups
received 42 mg of IMQ cream daily on their shaved backs, while the
mice in the IMQ + wogonin group additionally received an
intragastric gavage administration of 30 mg/kg/day wogonin (cat.
no. HY-N0400; MedChemExpress) (dissolved in sodium carboxymethyl
cellulose solvent). On day 8, all mice were euthanized. Prior to
euthanasia, mice were anesthetized through an intraperitoneal
injection of 1.5% pentobarbital sodium (50 mg/kg). The depth of
anesthesia was ensured by the absence of pedal and corneal
reflexes. Euthanasia was then performed by cervical dislocation
under deep anesthesia. Mortality was verified by ensuring the
cessation of breathing and heartbeat.
Animal health and behavior were monitored at least
twice daily throughout the present study. The specific humane
endpoints predefined for the present study, which would necessitate
immediate euthanasia, included: i) Severe lethargy or
unresponsiveness to gentle stimuli; ii) the inability to access
food or water; iii) weight loss >20% of baseline body weight; or
iv) signs of severe systemic illness, such as uncontrolled
bleeding, seizures or paralysis.
The present animal study protocol was approved by
the Animal Experiment Ethics Committee of Yunnan University of
Traditional Chinese Medicine (Kunming, China; approval no.
R-062021LH048).
Psoriasis Area and Severity Index (PASI) score of
mouse skin lesion tissues. Mouse dorsal lesions in each
treatment group were scored based on the area and severity of
psoriasis using the PASI score (31). The PASI score was assessed on a
five-point scale for erythema, infiltrates and scaling ranging from
0 to 4 (0=none, 1=mild, 2=moderate, 3=severe and 4=very severe),
and was calculated by combining the scores for erythema,
infiltration and scaling. The cumulative score served as a measure
of inflammation severity (scale 0-12).
H&E staining and observation of mouse skin
lesions. Following euthanasia on day 8, dorsal skin lesion
specimens were immediately collected from each mouse. Mouse skin
lesion tissues were fixed in 4% paraformaldehyde at 4˚C for 48 h,
with the fixative being changed once during this period. The
tissues were trimmed to a consistent size (~5x5 mm) and rinsed.
Subsequently, the tissue samples underwent a series of dehydration
steps using gradients of ethanol (100, 95, 85 and 75%), followed by
treatment with xylene and embedding in paraffin wax. The samples
were then sectioned to a thickness of 4 µm. Each section was dried,
dewaxed in xylene, and then hydrated through a descending ethanol
series (100, 95, 85 and 75%). Subsequently, the sectioned samples
were subjected to H&E staining, which included hematoxylin
staining (5 min), hydrochloric acid ethanol differentiation (1 min)
and eosin staining (1 min) at 25˚C. Finally, the samples were
dehydrated, rendered transparent and sealed. At the conclusion of
the experiment, histological alterations in the skin lesion tissues
were examined and documented under a light microscope (BX43;
Olympus Corporation). Finally, using ImageJ 1.8.0 software
(National Institutes of Health), three random fields per section
were selected to measure the epidermal thickness of the lesion.
Immunohistochemistry. Immunohistochemistry
was performed to assess the expression of matrix
metalloproteinase-9 (MMP9), cyclin-D1 (CCND1), cyclin-B1 (CCNB1),
cyclin-A2 (CCNA2), cyclin-dependent kinase 1 (CDK1) and androgen
receptor (AR) proteins in mouse skin lesion tissues. Mouse skin
lesion tissues were sectioned, baked, deparaffinized, hydrated (as
aforementioned for the HE staining process) and subjected to
antigen retrieval using 10 mM citrate buffer at 95˚C for 10 min.
Next, endogenous peroxidase activity was blocked by incubation with
3% H2O2 for 15 min at 25˚C. Following the
incubation of sections with 5% BSA (cat. no. A8012; Beijing
Solarbio Science & Technology Co., Ltd.) for 30 min at 25˚C.
Subsequently, the sections were incubated overnight at 4˚C with the
following primary antibodies diluted in 5% BSA: Anti-MMP9 (1:500;
cat. no. AF5228; Affinity Biosciences, Ltd.), anti-CCND1 (1:500;
cat. no. AF0931; Affinity Biosciences, Ltd.), anti-CCNA2 (1:500;
cat. no. AF0142; Affinity Biosciences, Ltd.), anti-CDK1 (1:500;
cat. no. DF6024; Affinity Biosciences, Ltd.), anti-CCNB1 (1:500;
cat. no. BF0062; Affinity Biosciences, Ltd.) and anti-AR (1:500;
cat. no. AF6137; Affinity Biosciences, Ltd.), followed by overnight
incubation with horseradish peroxidase-labeled goat anti-rabbit
secondary antibody (1:200; cat. no. S0001; Affinity Biosciences,
Ltd.) or horseradish peroxidase-labeled goat anti-mouse secondary
antibody (1:200, cat. no. S0002; Affinity Biosciences, Ltd.) at
4˚C. The sections were stained with 3,3'-diaminobenzidine,
counterstained with hematoxylin for 30-60 sec at 25˚C and sealed
after dehydration and clearing. Visualization was performed under
the BX43 light microscope (Olympus Corporation). Finally, using
ImageJ 1.8.0 software (National Institutes of Health), three random
fields per section were selected to measure the staining
intensity.
RNA sequencing (RNA-seq). Total RNA was
extracted from the three groups of skin samples using the RNeasy
Plus Mini kit (cat. no. 74134; Qiagen China Co., Ltd.) according to
the manufacturer's protocols. RNA concentration was determined
using a Qubit fluorometer (Invitrogen; Thermo Fisher Scientific,
Inc.) and RNA integrity was assessed using Agilent 4200 TapeStation
(Agilent Technologies, Inc.). The assay was performed strictly
following the manufacturer's instructions using an RNA ScreenTape
assay kit, and data were analyzed with the accompanying TapeStation
Analysis Software (version A.02.02). Samples with an RNA integrity
score of ≥8 were included in the present study. An indexed library
was constructed from 500 ng of total RNA according to the
manufacturer's instructions using the TruSeq Stranded mRNA sample
preparation kit (cat. no. 20020594; Illumina, Inc.). The library
concentration and fragment length distribution were assessed using
Qubit fluorometer and the Agilent 2100 Bioanalyzer (Agilent
Technologies, Inc.). The concentration was required to be >5
ng/µl and the fragment length was required to be between 300-400
bp. Paired-end sequencing (2x150 bp) was performed on an Illumina
NovaSeq 6000 platform using the NovaSeq 6000 S4 Reagent Kit (300
cycles; cat. no. 20028312; Illumina, Inc.). Bioinformatic analysis:
Raw sequencing reads (FASTQ files) were quality-controlled and
trimmed using fastp (version 0.23.2). Clean reads were then aligned
to the mouse reference genome (GRCm39) using STAR (version
2.7.10a). Gene-level counts were obtained using featureCounts (from
the Subread package; version 2.0.3). Differential expression
analysis was performed using the DESeq2 R package (version
1.38.3).
Statistical analysis
Experimental data were analyzed using SPSS 26.0 (IBM
Corp.) and data are presented as the median ± SD. One-way ANOVA
followed by Tukey's post hoc test was used to analyze the
statistical significance among multiple groups and unpaired
Student's t-test was used for comparisons between two groups (for
the analysis of DEGs from the two GEO datasets). Non-parametric
PASI score data are presented as the median ± interquartile range
and were analyzed with Kruskal-Wallis followed by Dunn's post hoc
test. P<0.05 was considered to indicate a statistically
significant difference.
Results
Acquisition of intersecting targets
between wogonin and psoriasis DEGs and construction of the action
network
A total of 100 targets of wogonin were obtained from
the SwissTargetPrediction database, 196 from the PharmMapper
database, 95 from the Similarity Ensemble Approach database and 80
from the SuperPred database. After eliminating duplicates, 423 drug
targets were identified (Table
SI).
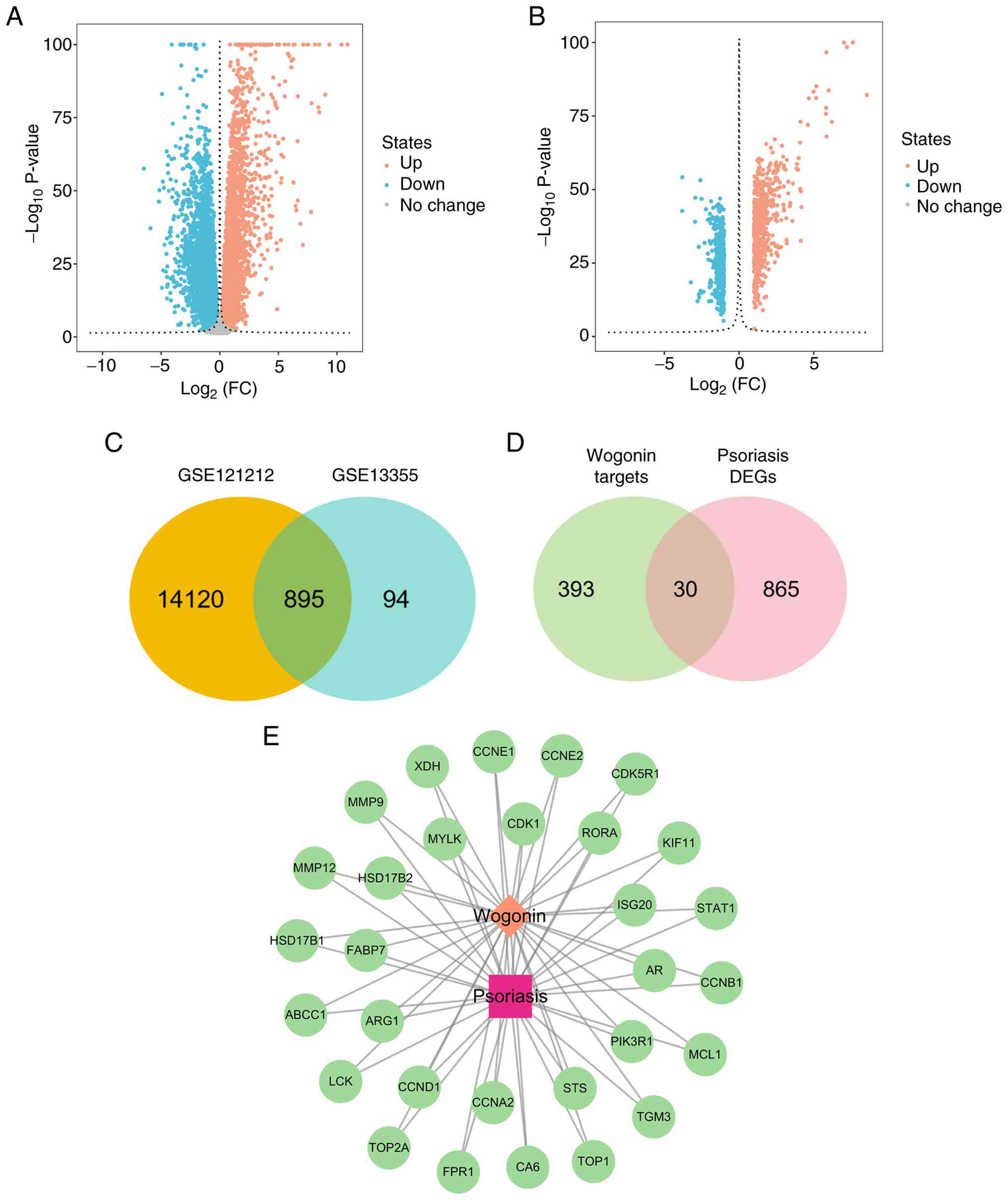
By comparing the differences between patients with
psoriasis and healthy controls, 15,015 and 989 DEGs were identified
in GSE121212 and GSE13355 datasets, respectively (Fig. 1A and B). The intersection of the two datasets
yielded 895 DEGs related to psoriasis (Fig. 1C). Subsequently, the overlap
between the DEGs of psoriasis and the targets of wogonin revealed
30 targets, which were defined as wogonin-psoriasis cross-targets
(Fig. 1D; Table I), suggesting that these targets
may serve a key role in mediating the therapeutic effects of
wogonin in psoriasis.
 | Table ITargets of wogonin for the treatment
of psoriasis. |
Table I
Targets of wogonin for the treatment
of psoriasis.
| Target name | Gene name | UniProt ID |
|---|
|
G2/mitotic-specific
cyclin-B1 | CCNB1 | P14635 |
| Signal transducer
and activator of transcription 1-α/β | STAT1 | P42224 |
| Cyclin-A2 | CCNA2 | P20248 |
| Cyclin-dependent
kinase 1 | CDK1 | P06493 |
| 17-β-hydroxysteroid
dehydrogenase type 1 | HSD17B1 | P14061 |
| DNA topoisomerase
2-α | TOP2A | P11388 |
| Protein-glutamine
γ-glutamyltransferase E | TGM3 | Q08188 |
| Multidrug
resistance-associated protein 1 | ABCC1 | P33527 |
|
Interferon-stimulated gene 20 kDa
protein | ISG20 | Q96AZ6 |
| Matrix
metalloproteinase-9 | MMP9 | P14780 |
| Induced myeloid
leukemia cell differentiation protein Mcl-1 | MCL1 | Q07820 |
|
G1/S-specific cyclin-E1 | CCNE1 | P24846 |
|
G1/S-specific cyclin-D1 | CCND1 | P24385 |
| Arginase-1 | ARG1 | Q61176 |
| Cyclin-dependent
kinase 5 activator 1 | CDK5R1 | Q15078 |
| Nuclear receptor
ROR-α | RORA | P35398 |
| Macrophage
metalloelastase | MMP12 | P39900 |
| Tyrosine-protein
kinase Lck | LCK | P06239 |
| Xanthine
dehydrogenase/oxidase | XDH | P47989 |
|
Phosphatidylinositol 3-kinase regulatory
subunit α | PIK3R1 | P27986 |
| Androgen
receptor | AR | P10275 |
| Kinesin-like
protein KIF11 | KIF11 | P52732 |
|
G1/S-specific cyclin-E2 | CCNE2 | O96020 |
| fMet-Leu-Phe
receptor | FPR1 | P21462 |
| Myosin light chain
kinase, smooth muscle | MYLK | Q15746 |
|
Steryl-sulfatase | STS | P08842 |
| Carbonic anhydrase
6 | CA6 | P23280 |
| Fatty acid-binding
protein, brain | FABP7 | O15540 |
| DNA topoisomerase
1 | TOP1 | P11387 |
| 17-β-Hydroxysteroid
dehydrogenase type 2 | HSD17B2 | P37059 |
Drug-target-disease network
construction and analysis
A drug-target-disease action network was constructed
to visualize the complex relationship between wogonin, its targets
and psoriasis (Fig. 1E). This
network illustrates the interaction between wogonin and the 30
cross-targets, highlighting the relationship between these targets
and psoriasis. The nodes in the network represent drugs (wogonin),
cross-targets and diseases (psoriasis), while the edges represent
their interactions. By mapping these interactions, the network
provides a comprehensive overview of the potential mechanisms
underlying the effects of wogonin on psoriasis.
PPI network analysis of
wogonin-psoriasis intersection targets and screening of core
genes
STRING was used to construct a PPI network of
wogonin-psoriasis cross-targets and the network was further
analyzed using Cytoscape 3.10.1. The PPI network consisted of 26
nodes and 71 edges, with 4 unconnected nodes hidden. Next, the
degree value of each target was obtained through the
‘CentiScaPe2.2’ plug-in. The top 10 targets with a higher degree
value were CCND1, MMP9, CCNB1, CCNA2, CDK1, AR, STAT1,
G1/S-specific cyclin-E1 (CCNE1), induced myeloid leukemia cell
differentiation protein Mcl-1 (MCL1) and DNA topoisomerase IIα
(TOP2A; Fig. 2A). The MCC value
for each target was obtained using the ‘cytoHubba’ plug-in to
identify the core targets. The top 10 targets with higher MCC
values were CDK1, CCNB1, CCND1, CCNA2, CCNE1, MMP9, AR, MCL1, TOP2A
and CCNE2 (Fig. 2B). By taking the
intersection of the top 10 targets ranked by degree values and the
top 10 targets ranked by MCC values, CCND1, MMP9, CCNB1, CCNA2,
CDK1 and AR were selected as core targets for subsequent molecular
docking and validation in animal experiments.
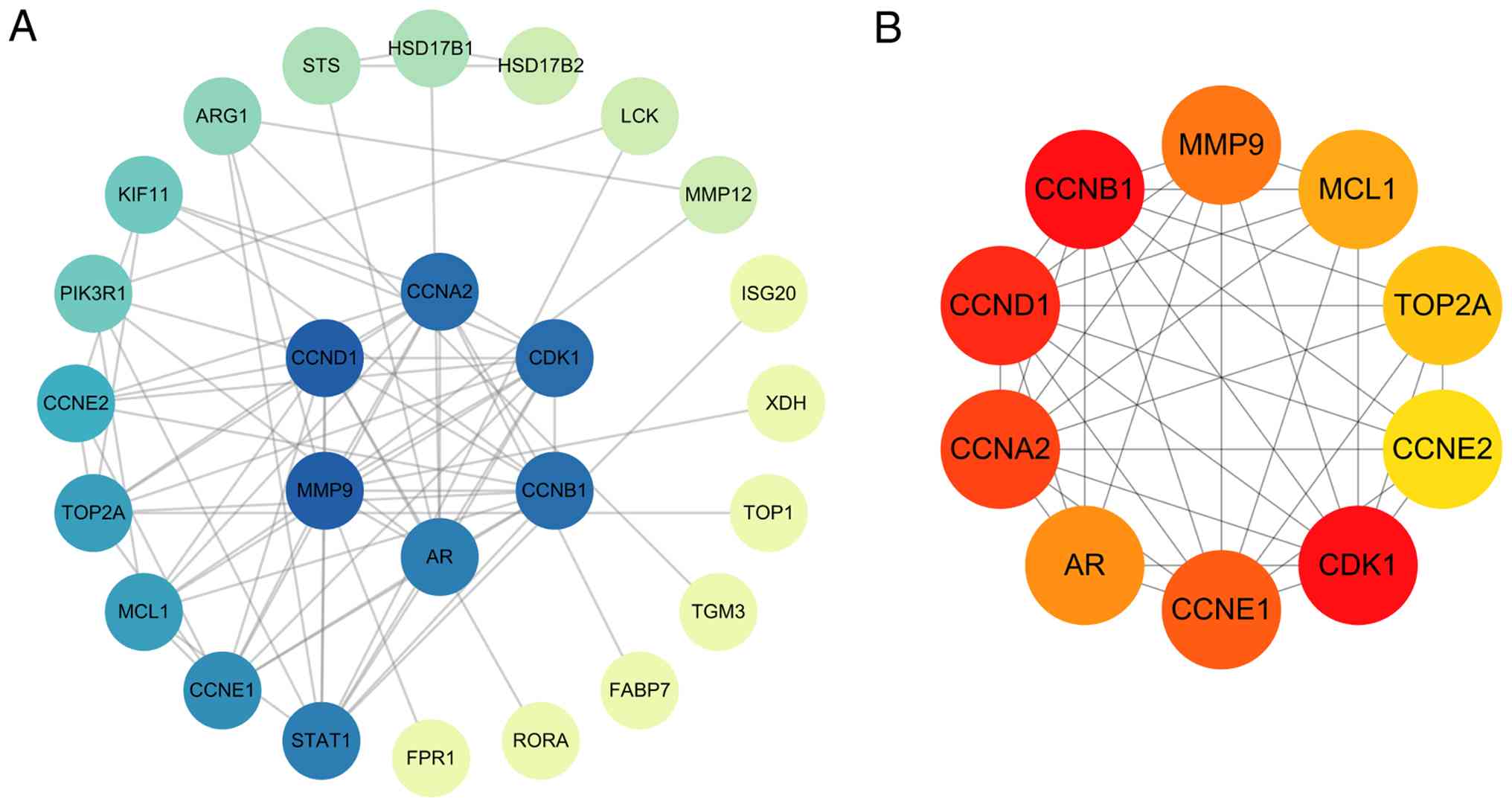 | Figure 2Construction of the PPI network and
identification of core target genes. (A) The PPI network of 30
intersection genes. The color depth indicates the degree value,
with darker colors representing higher degree values. (B) The top
10 genes based on MCC values. The color depth indicates the MCC
value, with darker colors representing higher MCC values. CCND1,
cyclin-D1; CCNA2, cyclin-A2; AR, androgen receptor; CCNE1,
cyclin-E1; CDK1, cyclin-dependent kinase 1; CCNE2, cyclin-E2;
TOP2A, DNA topoisomerase IIα; MCL1, induced myeloid leukemia cell
differentiation protein Mcl-1; MMP9, matrix metalloproteinase-9;
CCNB1, cyclin-B1. |
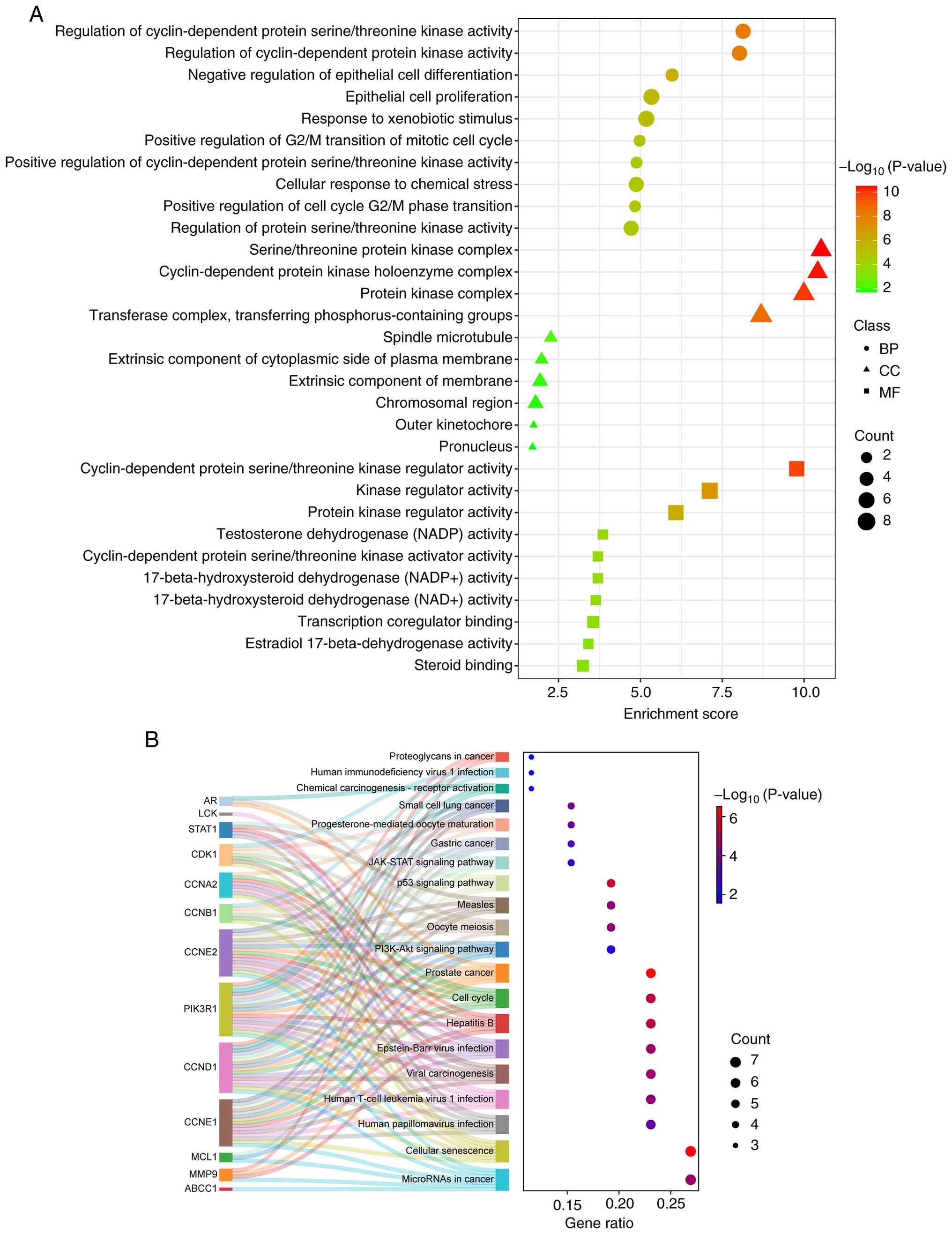
GO and KEGG pathway enrichment
analysis
A total of 30 intersecting targets were subjected to
gene enrichment analysis using DAVID. Among the 579 entries related
to BP, the majority of the targets were involved in the ‘regulation
of cyclin-dependent protein serine/threonine kinase activity’,
‘regulation of cyclin-dependent protein kinase activity’ and the
‘negative regulation of epithelial cell differentiation’. CC
yielded 29 entries, primarily ‘serine/threonine protein kinase
complex’, ‘cyclin-dependent protein kinase holoenzyme complex’ and
‘protein kinase complex’. MF yielded 93 entries, which
predominantly included ‘cyclin-dependent protein serine/threonine
kinase regulator activity’, ‘kinase regulator activity’ and
‘protein kinase regulator activity’. The results with the top 10
count values were selected for visualization through bar graphs,
demonstrating the BP, CC and MF components (Fig. 3A; Table SII).
KEGG pathway enrichment analysis was conducted using
DAVID, resulting in the identification of 51 signaling pathways.
These pathways were primarily associated with the ‘cell cycle’,
‘PI3K-Akt signaling pathway’, ‘p53 signaling pathway’ and ‘JAK-STAT
signaling pathway’ amongst others. Subsequently, the top 20
pathways were selected based on their count value and a KEGG
signaling pathway sankey diagram and dot plot was constructed
(Fig. 3B; Table SIII).
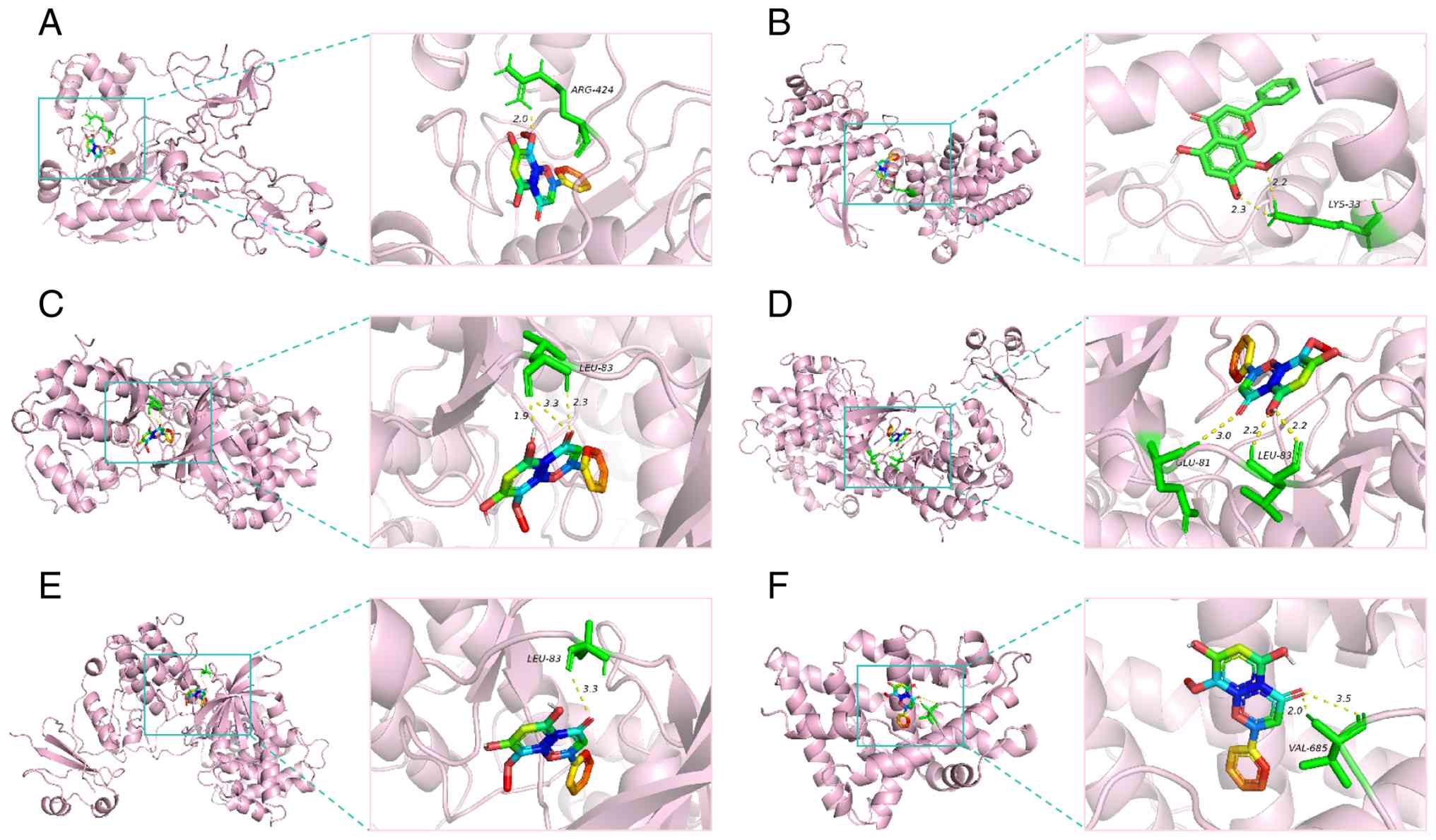
Molecular docking results
Molecular docking was used to analyze the binding
activities of six core targets (CCND1, MMP9, CCNB1, CCNA2, CDK1 and
AR) to wogonin. It is generally considered that a docking energy
value of <-7 kcal/mol indicates that the target protein binds
strongly to small molecules, while a docking energy value of <-5
kcal/mol indicates a moderate binding affinity (32). The molecular docking results
(Fig. 4; Table II) showed that the conformation of
the active compound exhibited a strong binding effect to the
protein targets, showing reliable interactions.
 | Table IIMolecular docking results of wogonin
and corresponding core targets. |
Table II
Molecular docking results of wogonin
and corresponding core targets.
| Target name | PDB ID | Affinity,
kcal/mol | Binding site (amino
acid residues) |
|---|
| MMP9 | 1L6J | -8.080 | ARG-424 |
| CCND1 | 2W96 | -7.644 | LYS-33 |
| CCNA2 | 6GVA | -9.378 | LEU-83 |
| CDK1 | 4Y72 | -9.703 | LEU-83 and
GLU-81 |
| CCNB1 | 6GU2 | -8.178 | LEU-83 |
| AR | 1E3G | -7.730 | VAL-685 |
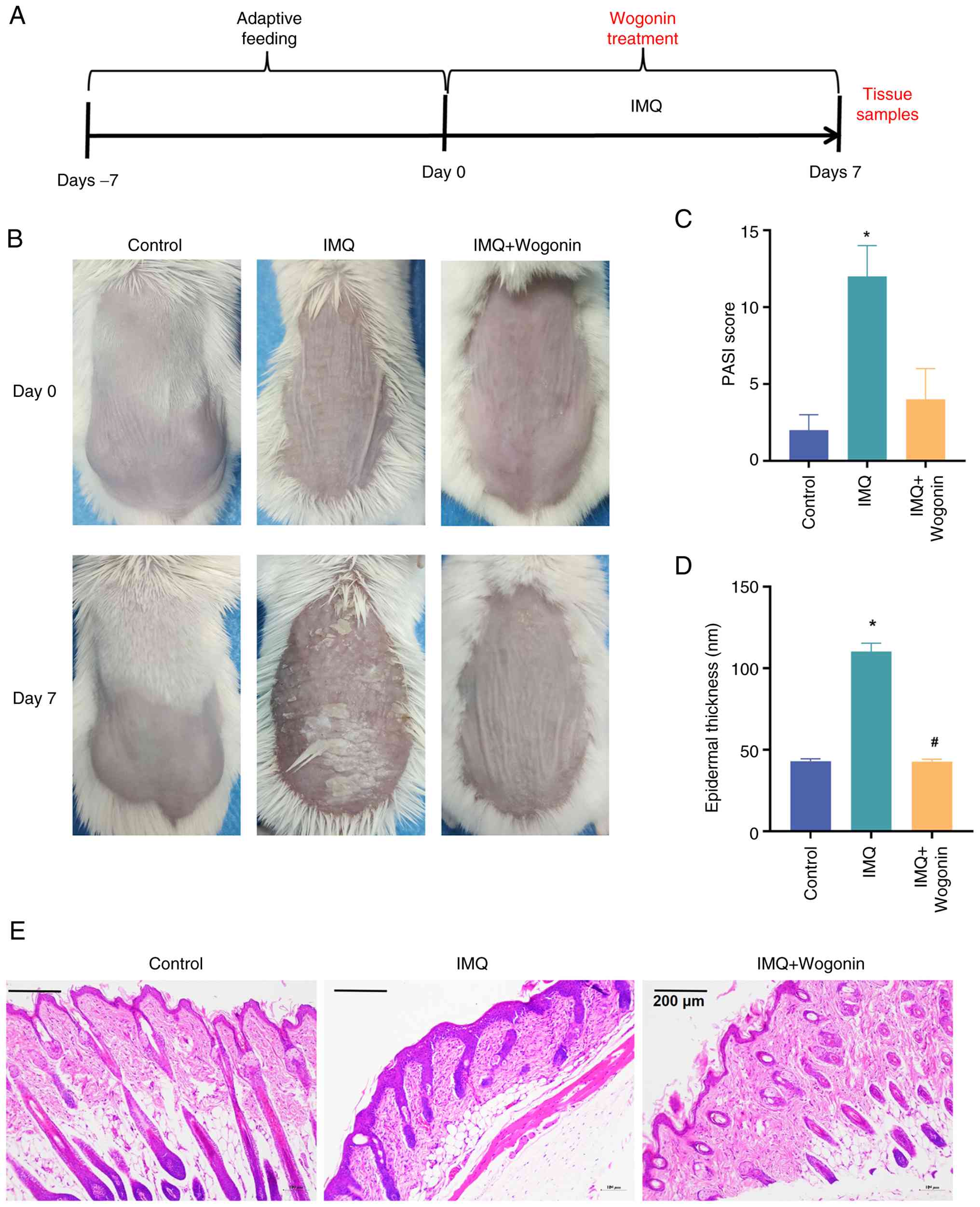
Wogonin alleviates psoriasis-like
lesions in the IMQ-induced mouse model
To evaluate the ameliorative effect of wogonin on
psoriasis-like lesions in IMQ-induced mice, wogonin was
administered to psoriasis mice (Fig.
5A). Following a 7-day intervention period, skin lesions were
scored using the PASI system and pathological changes in the skin
were observed using H&E staining. The PASI scores (Fig. 5B and C) of mice in the IMQ intervention group
were significantly higher than those in the control group
(P<0.05), while the PASI scores of mice in the IMQ + wogonin
group were lower than those in the IMQ group, albeit not
significantly different. The H&E staining results (Fig. 5D and E) demonstrated that the epidermis was
thickened and epidermal protrusions were elongated in the IMQ group
compared with those in the control (P<0.05), whereas the
thickness of the epidermis was significantly reduced after wogonin
treatment (P<0.05). These fundings suggest that wogonin may be
an effective intervention to improve skin damage in
psoriasis-induced mice.
Regulation of core target proteins by
wogonin
To further clarify the effect of wogonin on the
expression of CCND1, MMP9, CCNB1, CCNA2, CDK1 and AR proteins in
the skin of psoriasis-induced mice, protein expression was detected
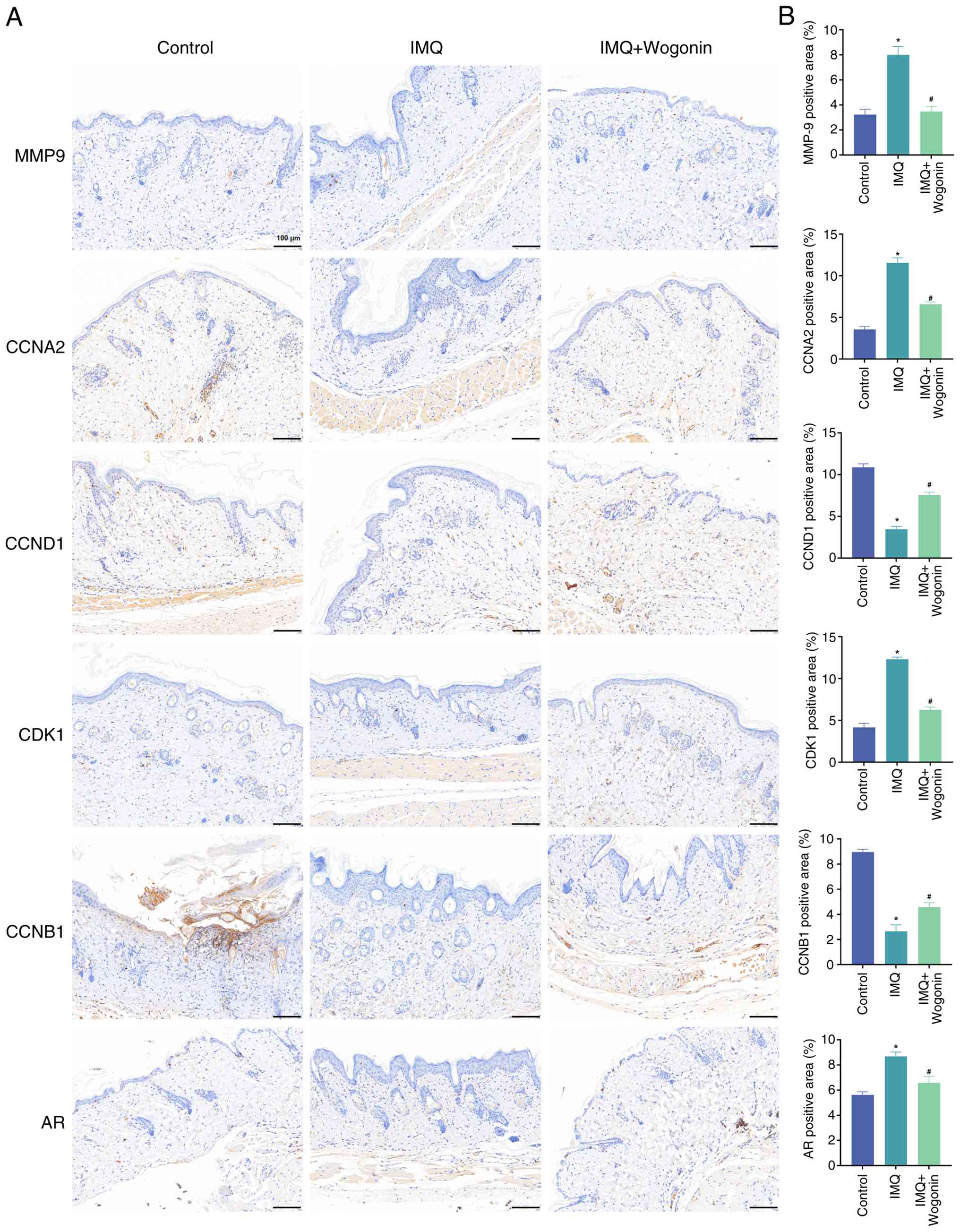
by immunohistochemistry. The results (Fig. 6) showed that, compared with that in
the control group, CCND1 and CCNB1 expression in the skin tissues
of mice in the IMQ group was significantly decreased (P<0.05),
while MMP9, CCNA2, CDK1 and AR protein expression was significantly
increased (P<0.05). Compared with that in the IMQ group, CCND1
and CCNB1 expression in the skin tissues of mice in the IMQ +
wogonin group was significantly increased (P<0.05), while MMP9,
CCNA2 and CDK1 protein expression was significantly decreased.
These results suggest that CCND1, MMP9, CCNB1, CCNA2 and CDK1 may
be key targets of wogonin in the treatment of psoriasis.
Verification of core targets through
RNA-seq
To further validate the selected core targets,
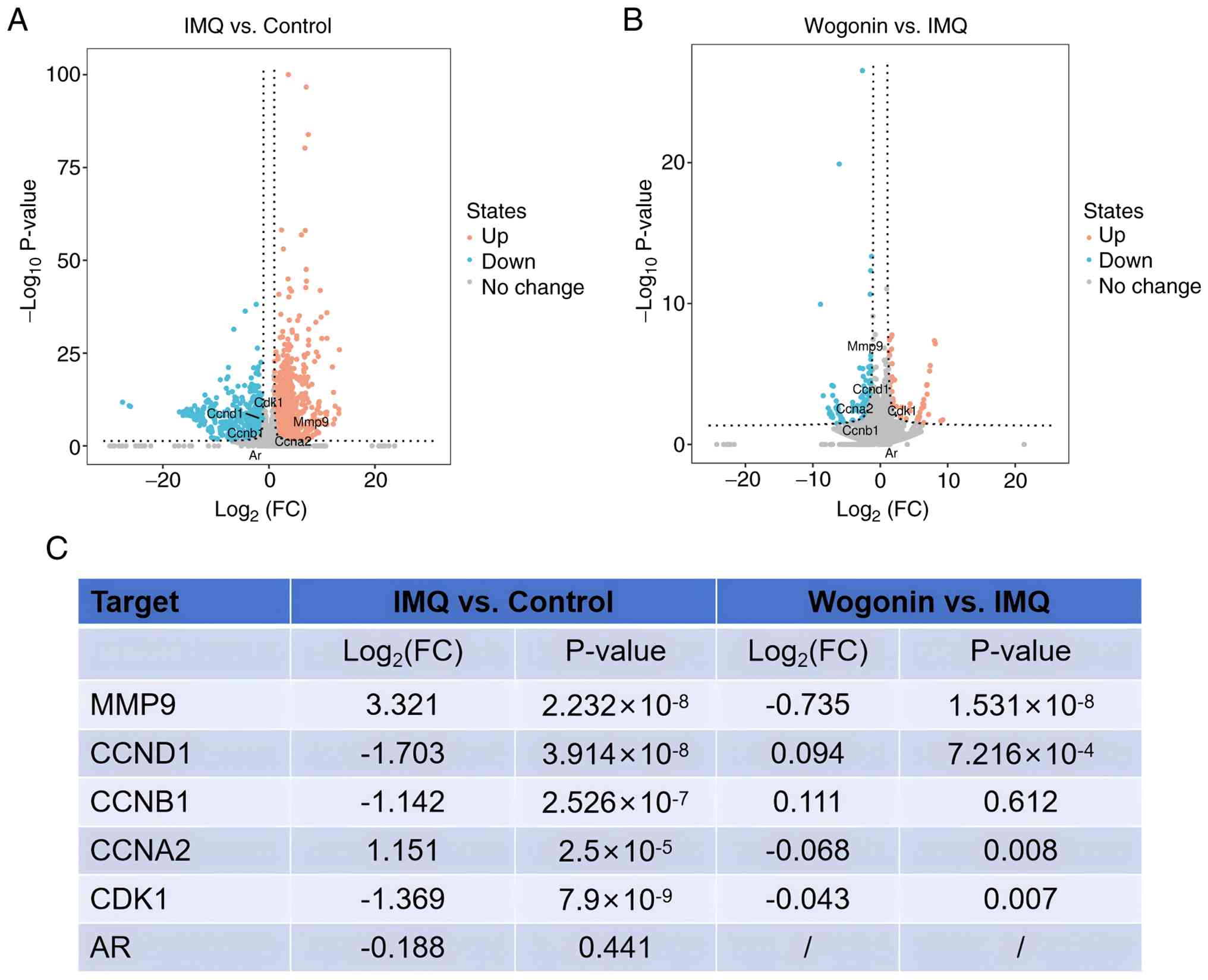
RNA-seq analysis was performed. The volcano plot illustrating the
DEGs between the IMQ and control groups is shown in Fig. 7A, while Fig. 7B displays the corresponding plot
for the wogonin group vs. the IMQ group. Fig. 7C summarizes the log2
(fold change) values and adjusted P-values of the core targets
across pairwise comparisons among the three groups. The results
showed that, compared with the control group, CCND1, CCNB1 and CDK1
were significantly downregulated in the IMQ group, while MMP9 and
CCNA2 were significantly upregulated. By contrast, compared with
the IMQ group, MMP9, CCNA2 and CDK1 were significantly
downregulated in the wogonin group, while CCND1 was significantly
upregulated. These changes in MMP9, CCNA2 and CCND1 expression were
consistent with the immunohistochemical findings.
Discussion
In the present study, the potential therapeutic
effects of wogonin in the treatment of psoriasis were investigated
using an integrative methodology that combined network
pharmacology, molecular docking and in vivo experiments. The
present findings suggested that wogonin may alleviate psoriasis
symptoms by targeting key proteins involved in the pathogenesis of
psoriasis, including CCND1, MMP9 and CCNA2 (33-35).
These results provide notable insights into the molecular
mechanisms underlying the therapeutic effects of wogonin and
emphasize its potential as a treatment option for psoriasis.
A key finding from the present network pharmacology
analysis was the prominence of cell cycle regulators (CCND1 and
CCNA2) among the core targets of wogonin. While MMP9 is a
well-established mediator of inflammation, the identification of
cell cycle components suggests a mechanism primarily focused on
controlling keratinocyte hyperproliferation. However, accumulating
evidence indicates that dysregulated cell cycle progression in
keratinocytes is closely associated with the initiation and
perpetuation of inflammation in psoriasis, creating a feedback loop
(36,37). The accelerated proliferation of
keratinocytes disrupts their normal differentiation, leading to the
release of damage-associated molecular patterns and a repertoire of
pro-inflammatory cytokines (such as IL-1β, IL-6 and TNF-α), further
amplifying the local immune response (38). Therefore, wogonin potentially
disrupts this key source of inflammation by targeting and
normalizing the cell cycle.
Additionally, the present network pharmacology
analysis and molecular docking results identified CCND1, MMP9,
CCNB1, CCNA2, CDK1 and AR as potential targets of wogonin.
Immunohistochemical and RNA-seq analyses in vivo revealed
that MMP9, CCNA2 and CCND1 may be key targets of wogonin in
psoriasis. It is important to note that the observed incomplete
consistency between RNA-seq (transcriptome level) and
immunohistochemistry (protein level) for some targets (e.g., AR and
CCNB1) is a recognized biological phenomenon. This discrepancy
primarily arises as mRNA expression levels and final protein
abundance are not always linearly correlated, being regulated by
complex mechanisms including post-transcriptional regulation,
translation efficiency and protein degradation. Consequently,
differences in the magnitude of change and statistical significance
between the two levels are not uncommon. The present study design
followed the standard paradigm in genomic research: Using
high-throughput RNA-seq for initial large-scale screening, followed
by independent validation (e.g., immunohistochemistry) of core
targets at the protein and tissue localization level, which is
closer to the pathophysiological reality. The general concordance
in the directional trends for the core targets (MMP9, CCNA2 and
CCND1) between the two techniques supports the conclusions, while
the specific variations highlight the value and necessity of
integrated multi-omics analysis. These proteins are involved in
various cellular processes, such as cell cycle regulation, matrix
remodeling and immune response modulation, all of which are key for
the development and progression of psoriasis (39-41).
CCND1 is a key regulator of the cell cycle,
facilitating the transition from the G1 phase to the S
phase (42,43). In psoriasis, CCND1 is often
upregulated, leading to keratinocytes hyperproliferation, a
hallmark of the disease (44).
Notably, CCND1 overexpression has been shown to directly influence
the expression of inflammatory genes in keratinocytes, thereby
associating uncontrolled proliferation to a pro-inflammatory
environment (45). Contrary to
these studies, the present study found that CCND1 was significantly
downregulated in the skin of mice induced by IMQ. However,
treatment with wogonin could effectively restore its expression.
This indicates that in the present model, the therapeutic effect of
wogonin involves correcting the dysregulation of CCND1, which may
help normalize the cell cycle process and inhibit the excessive
proliferation of keratinocytes.
MMP9 is involved in the degradation of extracellular
matrix components and is upregulated under inflammatory conditions,
including psoriasis (46,47). This regulation contributes to the
exacerbation of inflammation. Recent research has emphasized the
involvement of MMP9 in the pathogenesis of psoriasis, with elevated
levels of this enzyme being associated with greater disease
severity (48). In the present
study, wogonin was observed to downregulate MMP9, potentially
contributing to the attenuation of inflammatory responses in
psoriasis.
CCNA2 serves a key role in cell cycle regulation,
particularly during the transition from the G1 to the S
phase (49). Dysregulation of
CCNA2 has been observed in psoriasis, leading to abnormal
proliferation of epidermal cells (50). A previous study reported
upregulation of CCNA2 in psoriatic lesions, highlighting its
importance in the pathogenesis of psoriasis (51). The present findings suggested that
wogonin may downregulate CCNA2 to control keratinocyte
proliferation and mitigate skin damage in psoriasis.
The in vivo animal experiments conducted in
the present study corroborated the findings of network pharmacology
and molecular docking analyses. Wogonin treatment resulted in a
notable improvement in the skin lesions in a mouse model of
psoriasis, as demonstrated by reductions in epidermal
thickness.
Furthermore, immunohistochemical and RNA-seq
analyses indicated that wogonin modulated the expression of key
targets, including MMP9, CCNA2 and CCND1, in the skin tissue,
thereby elucidating its potential mechanism of action in psoriasis
treatment. Collectively, the present findings suggested that
wogonin exerts therapeutic effects against psoriasis by modulating
key targets involved in cell cycle regulation, matrix remodeling
and immune response modulation.
We hypothesize that the anti-proliferative effect
achieved through cell cycle modulation is associated with an
anti-inflammatory outcome. Rather than being independent,
normalizing the aberrant proliferation of keratinocytes directly
may contribute to mitigating inflammation in psoriatic lesions. By
targeting MMP9, CCNA2 and CCND1, wogonin may mitigate excessive
cell proliferation and inflammation, which are hallmark features of
psoriasis, thus providing a novel therapeutic strategy for managing
this challenging dermatological condition.
Furthermore, KEGG pathway enrichment analysis
revealed that the identified core targets are significantly
involved in key inflammatory and immune-related signaling pathways
implicated in psoriasis, such as the PI3K-Akt, JAK-STAT and p53
pathways (52-54).
This suggests that the therapeutic effect of wogonin may extend
beyond the direct modulation of individual targets such as MMP9,
and also involve the broader normalization of dysregulated
keratinocyte phenotype and immune response through these pivotal
pathways.
However, the present study did not further validate
these pathways, which constitutes a limitation. Although the
present study provided promising preclinical evidence regarding the
therapeutic effects of wogonin, further research is required to
fully elucidate the specific molecular mechanisms underlying its
action. In addition, clinical trials are essential to validate the
efficacy and safety of wogonin in patients with psoriasis. The
present study has also other limitations that should be considered
when interpreting the results. Firstly, the anti-psoriatic efficacy
of wogonin was evaluated at a single oral dose of 30 mg/kg/day.
This dosage was selected as it lies within the established
effective dose range (25-100 mg/kg) reported for wogonin in murine
inflammatory disease models (55)
and was supported by positive trends observed in preliminary
experiments of the present study. While this dose was effective in
alleviating psoriatic phenotypes, the lack of a full dose-response
analysis means that the optimal therapeutic dose for psoriasis
remains to be precisely determined. Secondly, a dedicated vehicle
control group (IMQ-induced mice receiving the 0.5% sodium
carboxymethyl cellulose solvent through gavage) was not included.
This omission, primarily due to logistical constraints, limits the
ability to completely rule out potential effects of the solvent or
the gavage procedure itself. Future studies should incorporate
multiple dosing regimens and a complete set of control groups to
build upon these promising initial findings.
In summary, the present findings suggested that
wogonin is a promising therapeutic agent for psoriasis, potentially
exerting its effects by modulating multiple molecular targets,
including MMP9, CCNA2 and CCND1. These results provide a foundation
for future investigations into the application of wogonin as a
potential treatment for psoriasis and other inflammatory skin
disorders.
Supplementary Material
Wogonin targets from various
databases.
Top 10 enriched terms of biological
process, cellular component and molecular function.
Top 20 enriched terms of Kyoto
Encyclopedia of Genes and Genomes pathways.
Acknowledgements
Not applicable.
Funding
Funding: The present research was funded by the Yunnan
Traditional Chinese Medicine Basic Research Joint Special Project
(grant nos. 202301AZ070001-090 and 202101AZ070001-124).
Availability of data and materials
The sequencing data generated in the present study
may be found in the Sequence Read Archive database of the National
Center for Biotechnology Information under accession number
PRJNA1344434 or at the following URL: https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA1344434.
The other data generated in the present study may be requested from
the corresponding author.
Authors' contributions
YL and XZ conceptualized the present study. DY, YG
and JG contributed towards the design and formulation of the
experimental plan. FZ was responsible for the statistical
analysis/bioinformatics. FZ, JW and YX contributed towards to
evaluation of the ultimate credibility of research results. Results
analysis was conducted by YX and DY. The experimental investigation
was conducted by YL, FZ and YL. DY contributed towards the
acquisition of funding. JW and YL were responsible for data
curation. YL was responsible for writing and preparing the original
draft. DY reviewed and edited the manuscript. XZ contributed
towards molecular docking visualization. DY supervised the study.
YL performed project administration. DY acquired the funding. All
authors read and approved the final version of the manuscript. YL
and DY confirm the authenticity of all the raw data.
Ethics approval and consent to
participate
The animal study protocol was approved by the Animal
Experiment Ethics Committee of Yunnan University of Traditional
Chinese Medicine (approval no. R-062021LH048).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Menter A: Psoriasis and psoriatic
arthritis treatment. Am J Manag Care. 22 (8 Suppl):S225–S237.
2016.PubMed/NCBI
|
|
2
|
Anutraungkool T, Chaowattanapanit S,
Choonhakarn C, Limpawattana P, Julanon N, Wongjirattikarn R,
Kruahong K, Mokkarat A and Sawanyawisuth K: Factors associated with
exacerbation in psoriasis compared to eczema. Dermatol Pract
Concept. 15(e20255630)2025.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Bardazzi F, Bonci C, Sacchelli L, DI
Altobrando A, Iommi M, Rucci P, Sacchelli P, Berardi D, Patrizi A
and Tengattini V: Suicide risk and depression in patients with
psoriasis. Ital J Dermatol Venerol. 157:497–501. 2022.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Kormeili T, Lowe NJ and Yamauchi PS:
Psoriasis: Immunopathogenesis and evolving immunomodulators and
systemic therapies; U.S. experiences. Br J Dermatol. 151:3–15.
2004.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Liu T, Li S, Ying S, Tang S, Ding Y, Li Y,
Qiao J and Fang H: The IL-23/IL-17 pathway in inflammatory skin
diseases: From bench to bedside. Front Immunol.
11(594735)2020.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Tuli HS, Rath P, Chauhan A, Parashar G,
Parashar NC, Joshi H, Rani I, Ramniwas S, Aggarwal D, Kumar M and
Rana R: Wogonin, as a potent anticancer compound: From chemistry to
cellular interactions. Exp Biol Med (Maywood). 248:820–828.
2023.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Chi YS, Lim H, Park H and Kim HP: Effects
of wogonin, a plant flavone from Scutellaria radix, on skin
inflammation: In vivo regulation of inflammation-associated gene
expression. Biochem Pharmacol. 66:1271–1278. 2003.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Liu C, Shu S, Xia Z, Chen G and Xu Y:
Exploring the latent mechanism of Huanglian Jiedu decoction formula
for anti-atopic dermatitis by systems pharmacology. Comb Chem High
Throughput Screen. 26:610–629. 2023.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Ibrahim M, Singh H, Fahim M, Khan S, Khan
J, Arun JK, Mishra AK, Virmani T, Sharma A, Kumar G, et al:
Nutraceutical interventions for mitigating skin ageing: Analysis of
mechanisms and efficacy. Curr Pharm Des. 31:2385–2401.
2025.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Zhang XH, Liu J, Ding X and Chen XL:
Chinese herbal medicines and its active ingredient wogonin can
improve immune inflammation in psoriatic arthritis. Int
Immunopharmacol. 147(113984)2025.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Zhang B, Roesner LM, Traidl S, Koeken V,
Xu CJ, Werfel T and Li Y: Single-cell profiles reveal distinctive
immune response in atopic dermatitis in contrast to psoriasis.
Allergy. 78:439–453. 2023.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Wu X, Liu C, Zhang C, Kuai L, Hu S, Jia N,
Song J, Jiang W, Chen Q and Li B: The role of lactate and
lactylation in the dysregulation of immune responses in psoriasis.
Clin Rev Allergy Immunol. 68(28)2025.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Ma J, Ji C, Sun Y, Liu D, Pan K and Wei Y:
Wogonin ameliorates the proliferation, inflammatory response, and
pyroptosis in keratinocytes via NOD-like receptor family pyrin
domain containing 3/Caspase-1/Gasdermin-D pathway. Immun Inflamm
Dis. 12(e1303)2024.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Zhang P, Zhang D, Zhou W, Wang L, Wang B,
Zhang T and Li S: Network pharmacology: Towards the artificial
intelligence-based precision traditional Chinese medicine. Brief
Bioinform. 25(bbad518)2023.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Daina A, Michielin O and Zoete V:
SwissTargetPrediction: Updated data and new features for efficient
prediction of protein targets of small molecules. Nucleic Acids
Res. 47(W1):W357–W364. 2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Wang X, Shen Y, Wang S, Li S, Zhang W, Liu
X, Lai L, Pei J and Li H: PharmMapper 2017 update: A web server for
potential drug target identification with a comprehensive target
pharmacophore database. Nucleic Acids Res. 45(W1):W356–W360.
2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Keiser MJ, Roth BL, Armbruster BN,
Ernsberger P, Irwin JJ and Shoichet BK: Relating protein
pharmacology by ligand chemistry. Nat Biotechnol. 25:197–206.
2007.PubMed/NCBI View
Article : Google Scholar
|
|
18
|
Gallo K, Goede A, Preissner R and Gohlke
BO: SuperPred 3.0: Drug classification and target prediction-a
machine learning approach. Nucleic Acids Res. 50(W1):W726–W731.
2022.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Novoszel P, Holcmann M, Stulnig G, De Sa
Fernandes C, Zyulina V, Borek I, Linder M, Bogusch A, Drobits B,
Bauer T, et al: Psoriatic skin inflammation is promoted by
c-Jun/AP-1-dependent CCL2 and IL-23 expression in dendritic cells.
EMBO Mol Med. 13(e12409)2021.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Swindell WR, Stuart PE, Sarkar MK,
Voorhees JJ, Elder JT, Johnston A and Gudjonsson JE: Cellular
dissection of psoriasis for transcriptome analyses and the
post-GWAS era. BMC Med Genomics. 7(27)2014.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Tang D, Chen M, Huang X, Zhang G, Zeng L,
Zhang G, Wu S and Wang Y: SRplot: A free online platform for data
visualization and graphing. PLoS One. 18(e0294236)2023.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Dennis G Jr, Sherman BT, Hosack DA, Yang
J, Gao W, Lane HC and Lempicki RA: DAVID: Database for annotation,
visualization, and integrated discovery. Genome Biol.
4(P3)2003.PubMed/NCBI
|
|
24
|
Zhang H, Xiong P, Zheng T, Hu Y, Guo P,
Shen T and Zhou X: Combination of berberine and evodiamine
alleviates obesity by promoting browning in 3T3-L1 cells and
high-fat diet-induced mice. Int J Mol Sci. 26(4170)2025.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Trott O and Olson AJ: AutoDock Vina:
Improving the speed and accuracy of docking with a new scoring
function, efficient optimization, and multithreading. J Comput
Chem. 31:455–461. 2010.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Seeliger D and de Groot BL: Ligand docking
and binding site analysis with PyMOL and Autodock/Vina. J Comput
Aided Mol Des. 24:417–422. 2010.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Cervantes-Durán C, Avalos-Viveros M,
Torner L, Sánchez-Ceja SG, Rodríguez-Orozco AR, Martínez-Flores HE
and García-Pérez ME: The 5-HT(1A) receptor agonist, 8-OH-DPAT,
attenuates long-lasting pain in imiquimod-induced psoriasis in
mice. Exp Dermatol. 31:600–607. 2022.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Chen X, Fu C, Zheng Y, Li X, Liao Y, Zheng
Y, Liang W, Zhao Y, Huang J, Huang T, et al: Intermittent fasting
alleviates IMQ-induced psoriasis-like dermatitis via reduced γδT17
and monocytes in mice. Arch Dermatol Res. 316(176)2024.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Yu Z, Wang Y, Guo Y, Zhu R, Fang Y, Yao Q,
Fu H, Zhou A, Ma L and Shou Q: Exploring the therapeutic and gut
microbiota-modulating effects of qingreliangxuefang on IMQ-induced
psoriasis. Drug Des Devel Ther. 19:3269–3291. 2025.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Lei MJ, Bai F, Zhang QY, Yang QQ and Tian
Z: Total glucosides of paeony regulate immune imbalance mediated by
dermal mesenchymal stem cells in psoriasis mice. Chin J Integr Med.
29:517–525. 2023.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Chen C, Hou G, Zeng C, Ren Y, Chen X and
Peng C: Metabolomic profiling reveals amino acid and carnitine
alterations as metabolic signatures in psoriasis. Theranostics.
11:754–767. 2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Gromiha MM and Harini K: Protein-nucleic
acid complexes: Docking and binding affinity. Curr Opin Struct
Biol. 90(102955)2025.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Gong T, Chen J, Xiao Z, Luo R, Tong Z, Ke
H, Liu Z, Xiao C, Xiang N and Ji C: Proteomic profiling and
clinical insights: The role of MMP9 in differentiating psoriasis
vulgaris from generalized pustular psoriasis. J Inflamm Res.
18:3795–3805. 2025.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Kılıç ŞB, Taheri S, Duman EM, Solak EO,
Şükranlı ZY, Rassoulzadegan M and Borlu M: Psoriatic skin
transcript phenotype: Androgen/estrogen and cortisone/cortisol
imbalance with increasing DNA damage response. Mol Biol Rep.
51(933)2024.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Choudhary S, Anand R, Pradhan D, Bastia B,
Kumar SN, Singh H, Puri P, Thomas G and Jain AK: Transcriptomic
landscaping of core genes and pathways of mild and severe psoriasis
vulgaris. Int J Mol Med. 47:219–231. 2021.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Albanesi C, De Pità O and Girolomoni G:
Resident skin cells in psoriasis: A special look at the
pathogenetic functions of keratinocytes. Clin Dermatol. 25:581–588.
2007.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Hawkes JE, Chan TC and Krueger JG:
Psoriasis pathogenesis and the development of novel targeted immune
therapies. J Allergy Clin Immunol. 140:645–653. 2017.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Benhadou F, Mintoff D and Del Marmol V:
Psoriasis: Keratinocytes or immune cells-which is the trigger?
Dermatology. 235:91–100. 2019.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Yao L, Sai HV, Shippy T and Li B: Cellular
and transcriptional response of human astrocytes to hybrid protein
materials. ACS Appl Bio Mater. 7:2887–2898. 2024.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Feng C, Yan W, Mei Z and Luo X: Exploring
the toxicological impact of bisphenol a exposure on psoriasis
through network toxicology, machine learning, and multi-dimensional
bioinformatics analysis. J Environ Manage.
385(125708)2025.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Buttars B, Baltazar D, Charest G, Flaherty
A, Hamann D, Whittemore D and Hamann C: Two cases of drug-induced
psoriasis from second-generation androgen receptor blockers. Clin
Exp Dermatol. 50:1035–1037. 2025.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Liu J, Lin J, Wang X, Zheng X, Gao X,
Huang Y, Chen G, Xiong J, Lan B, Chen C, et al: CCND1 Amplification
profiling identifies a subtype of melanoma associated with poor
survival and an immunosuppressive tumor microenvironment. Front
Immunol. 13(725679)2022.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Knudsen ES, Kumarasamy V, Nambiar R,
Pearson JD, Vail P, Rosenheck H, Wang J, Eng K, Bremner R, Schramek
D, et al: CDK/cyclin dependencies define extreme cancer cell-cycle
heterogeneity and collateral vulnerabilities. Cell Rep.
38(110448)2022.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Dorai S and Anand DA: Differentially
expressed cell cycle genes and STAT1/3-driven multiple cancer
entanglement in psoriasis, coupled with other comorbidities. Cells.
11(3867)2022.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Fernández-Hernández R, Rafel M, Fusté NP,
Aguayo RS, Casanova JM, Egea J, Ferrezuelo F and Garí E: Cyclin D1
localizes in the cytoplasm of keratinocytes during skin
differentiation and regulates cell-matrix adhesion. Cell Cycle.
12:2510–2517. 2013.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Dong C, Lin JM, Lu X, Zhu J, Lin L, Xu J
and Du J: Fibroblasts with high matrix metalloproteinase 2
expression regulate CD8+ T-cell residency and inflammation via
CD100 in psoriasis. Br J Dermatol. 191:405–418. 2024.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Liu L, Zhang H, Tang X, Zhang M, Wu Y,
Zhao Y, Lu C and Zhao R: Geniposide ameliorates psoriatic skin
inflammation by inhibiting the TLR4/MyD88/NF-κB p65 signaling
pathway and MMP9. Int Immunopharmacol. 133(112082)2024.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Zhou X, Zhou H, Luo X and Wu RF: Discovery
of biomarkers in the psoriasis through machine learning and dynamic
immune infiltration in three types of skin lesions. Front Immunol.
15(1388690)2024.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Ma Q: MiR-219-5p suppresses cell
proliferation and cell cycle progression in esophageal squamous
cell carcinoma by targeting CCNA2. Cell Mol Biol Lett.
24(4)2019.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Zhu X, Zhang E and Qin L: The high
expression of TOP2A and MELK induces the occurrence of psoriasis.
Aging (Albany NY). 16:3185–3199. 2024.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Kong Y, Wu R, Zhang S, Zhao M, Wu H, Lu Q,
Fu S and Su Y: Wilms' tumor 1-associating protein contributes to
psoriasis by promoting keratinocytes proliferation via regulating
cyclinA2 and CDK2. Int Immunopharmacol. 88(106918)2020.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Ren Y, Dong H, Jin R, Jiang J and Zhang X:
TRIM22 actives PI3K/Akt/mTOR pathway to promote psoriasis through
enhancing cell proliferation and inflammation and inhibiting
autophagy. Cutan Ocul Toxicol. 41:304–309. 2022.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Furtunescu AR, Georgescu SR, Tampa M and
Matei C: Inhibition of the JAK-STAT pathway in the treatment of
psoriasis: A review of the literature. Int J Mol Sci.
25(4681)2024.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Wang N, Xu X, Guan F, Lin Y, Ye Y, Zhou J,
Feng J, Li S, Ye J, Tang Z, et al: FGF12 Positively regulates
keratinocyte proliferation by stabilizing MDM2 and inhibiting p53
activity in psoriasis. Adv Sci (Weinh). 11(e2400107)2024.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Li HD, Chen X, Yang Y, Huang HM, Zhang L,
Zhang X, Zhang L, Huang C, Meng XM and Li J: Wogonin attenuates
inflammation by activating PPAR-γ in alcoholic liver disease. Int
Immunopharmacol. 50:95–106. 2017.PubMed/NCBI View Article : Google Scholar
|