Contents
Introduction
Classic cytogenetics analysis
Molecular cytogenetics analysis
Fluorescence in situ hybridization (FISH)
Comparative genomic hybridization (CGH)
Microarray-based comparative genomic hybridization
(array CGH)
Postnatal diagnostic applications of array CGH
Conclusions and future perspectives
Introduction
Cytogenetics refers to studies of the cellular
aspects of heredity, especially the description of chromosome
structure and the identification of genomic aberrations that cause
disease (1). It has provided the
clues for solving fundamental biological questions, such as the
nature of inherited syndromes, the genomic changes that are
involved in tumorigenesis and the three-dimensional organization of
the human genome (2–5). Over the years, cytogenetic
techniques evolved and became part of routine laboratory testing,
providing valuable diagnostic and prognostic information in
congenital and developmental abnormalities.
Microarray-based CGH merges molecular diagnostics
with traditional chromosome analysis and is transforming the field
of clinical cytogenetics. Evaluation of copy number variation by
microarray analysis has significant advantages over standard
metaphase karyotyping and is quickly becoming the primary means of
postnatal genetic evaluation of patients with developmental delay
(DD), intellectual disability (ID), autism spectrum disorders (ASD)
and/or multiple congenital anomalies (MCA) (6,7).
The use of microarray technologies in these categories of disorders
is emerging and promises higher sensitivity for several highly
penetrant, clinically severe microdeletion and microduplication
syndromes. However, clinical application of microarrays for routine
postnatal diagnosis is still being investigated.
In this article, we present a brief overview of how
the field is moving away from traditional methods towards molecular
genetics approaches for the detection of pathogenic genomic
imbalances and highlight the potential utility of the microarray
technologies for clinical evaluation of patients with DD, ID, ASD
and/or MCA.
Classic cytogenetics analysis
To understand the importance of microarray
technologies to clinical practice, it is helpful to look at these
advances in the context of the evolution of clinical
cytogenetics.
Chromosome banding analysis was developed in the
1970s and is recognized as the gold standard for the diagnosis and
prognosis of congenital and acquired disorders. It has been used
for scanning the genome for aberrations that involve both gains and
losses of portions of the genome, as well as rearrangements within
and among chromosomes. Extensive experience gained in the past
decades has shown that karyotyping analysis have proven the causal
association between specific chromosomal abnormalities and clinical
syndromes observed in individuals with DD/ID, ASD or MCA.
However, investigation by standard metaphase
karyotyping is not always sufficient to identify the origin of
extra chromosome material or correctly assign other structural
chromosomal rearrangements (1).
In addition, the limited chromosome-specific banding resolution
makes the characterization and correct interpretation of complex
and cryptic chromosome aberrations difficult to ascertain and is
therefore often imprecise in nature.
Molecular cytogenetics analysis
Molecular cytogenetics, a new field of cytogenetics,
and the product of a combination of cytogenetics and molecular
biology, has increased the resolution and diagnostic utility of
cytogenetic analysis. It has been repeatedly proven effective in
genetic diagnostics and has been recognized as a valuable addition
or even alternative to chromosomal banding analysis (8–11).
The resolution and level of excellence of these techniques are
established against cytogenetic banding analysis, which remains the
golden standard in this instance.
The overwhelming majority of molecular cytogenetic
techniques are based on hybridization. In the last three decades,
molecular cytogenetic techniques based on hybridization
applications locating specific, fluorescence-labeled nucleic acid
sequences in interphase cells or metaphase chromosomes, have become
fast, sensitive, and important complementing tools in genetic
diagnostics (12).
The use of variable molecular cytogenetics
techniques enhances the thorough interpretation of numerical and
complex chromosome aberrations, bridging the gap between
conventional chromosome banding analysis and molecular genetics DNA
studies (13).
Fluorescence in situ hybridization
(FISH)
FISH is based on the use of chromosome
region-specific, fluorescent-labeled DNA probes. The technique
offers numerous possibilities for identifying genomic imbalances
with great accuracy and has become an important complementing
application in genetic diagnostics. The application of FISH permits
the determination of the number and location of specific DNA
sequences, both on metaphase chromosomes and in interphase nuclei,
thus significantly simplifying the preparation and evaluation of
samples (10). The use of
diverse, locus-specific FISH probes and multicolor assays enhances
the thorough characterization of numerical and complex chromosome
aberrations regardless of their complexity (14).
However, FISH requires clinical suspicion that a
specific locus in the genome has undergone copy-number change and
therefore is not applicable for whole genome screening approaches
required in a diagnostic setting. Furthermore, FISH analysis on
metaphase chromosomes detects only microdeletions, and duplications
involving segments smaller than 3–5 Mb may be routinely missed even
by FISH of interphase nuclei (11).
Comparative genomic hybridization (CGH)
CGH represents a variation on FISH technology with
the clear advantage of revealing imbalances across the whole
genome. In CGH, test and reference genomic DNAs are differentially
labeled with fluorochromes and then co-hybridized onto normal
metaphase chromosomes. Following hybridization, the chromosomes are
scanned to measure the fluorescence intensities along the length of
the normal chromosomes to detect intensity ratio differences that
subsequently pinpoint to genomic imbalances (15). Differences between the
fluorescence intensities along the length of any given chromosome
will reveal gains or losses of genomic segments. In this way, a
global overview of chromosomal gains and losses throughout the
whole genome is available and many studies have proved that CGH is
a useful and reliable technique in the research and diagnostics of
both cancer and human genetic disorders.
Although the conventional CGH analysis allows the
entire genome to be scanned for aberrations, owing to the limited
resolution (5–10 Mb) of metaphase chromosomes, aberrations such as
mosaicism, balanced chromosomal translocations, inversions, and
whole genome ploidy changes cannot be detected using this approach.
In particular, apparently balanced translocations in patients with
abnormal phenotypes may hide deletions both at the breakpoint and
elsewhere in the genome. Preliminary data demonstrated that other
rearrangements, such as ring chromosomes, may be more complex than
anticipated (15). For the
detection of such abnormalities, a high-resolution technique is
required.
Overall, the resolution at which copy number changes
can be detected using these molecular cytogenetic techniques are
only slightly higher as compared to conventional karyotyping.
Furthermore, all experiments are labor-intensive and
time-consuming, especially when multiple genomic regions are
interrogated. Therefore, it is strongly advised to investigate
these rearrangements with higher resolution and excellent
throughput.
Microarray-based comparative genomic
hybridization (array CGH)
Microarray technology represents the technical
convergence of molecular genetics and cytogenetics and is rapidly
revolutionizing modern cytogenetics. The technology combines
fluorescence techniques with the microarray platform and allows the
comparison of DNA content in two differentially labeled genomes, a
test genome, and a reference genome. In this way the microarray
platform allows the use of thousands of individual DNA sequences
from throughout the genome, and provides precise information in a
single experiment about the locations of any identified
aberrations, thereby it holds a great potential for the analysis of
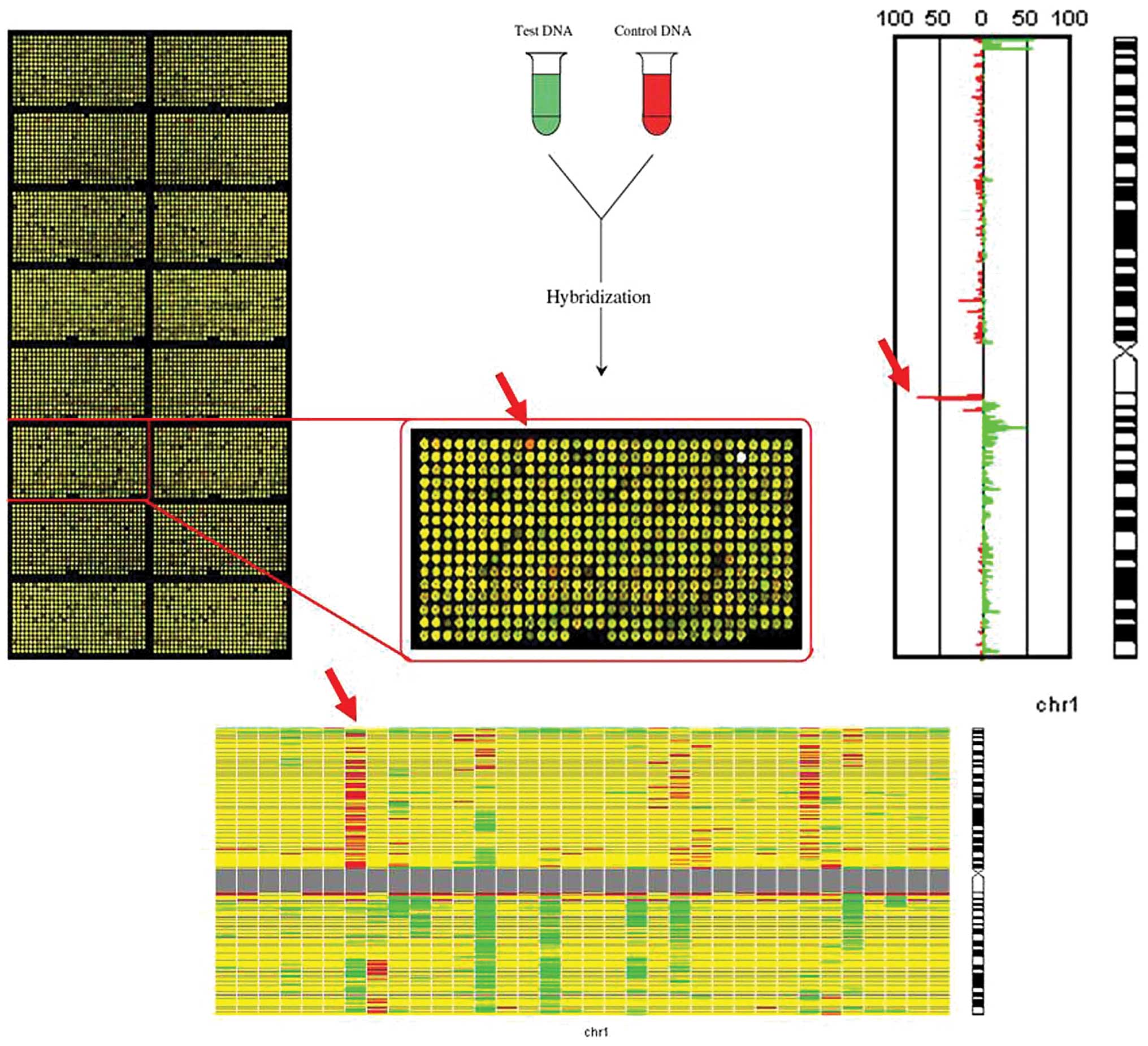
DNA copy number changes in clinical genetics (4). A schematic overview of the
microarray-based CGH technique is presented in Fig. 1.
The value of whole-genome array CGH highlights the
interpretation difficulties associated with copy number variations
of unclear significance. These changes include both numerical and
unbalanced structural abnormalities, such as deletions,
duplications, or amplifications at any locus as long as that region
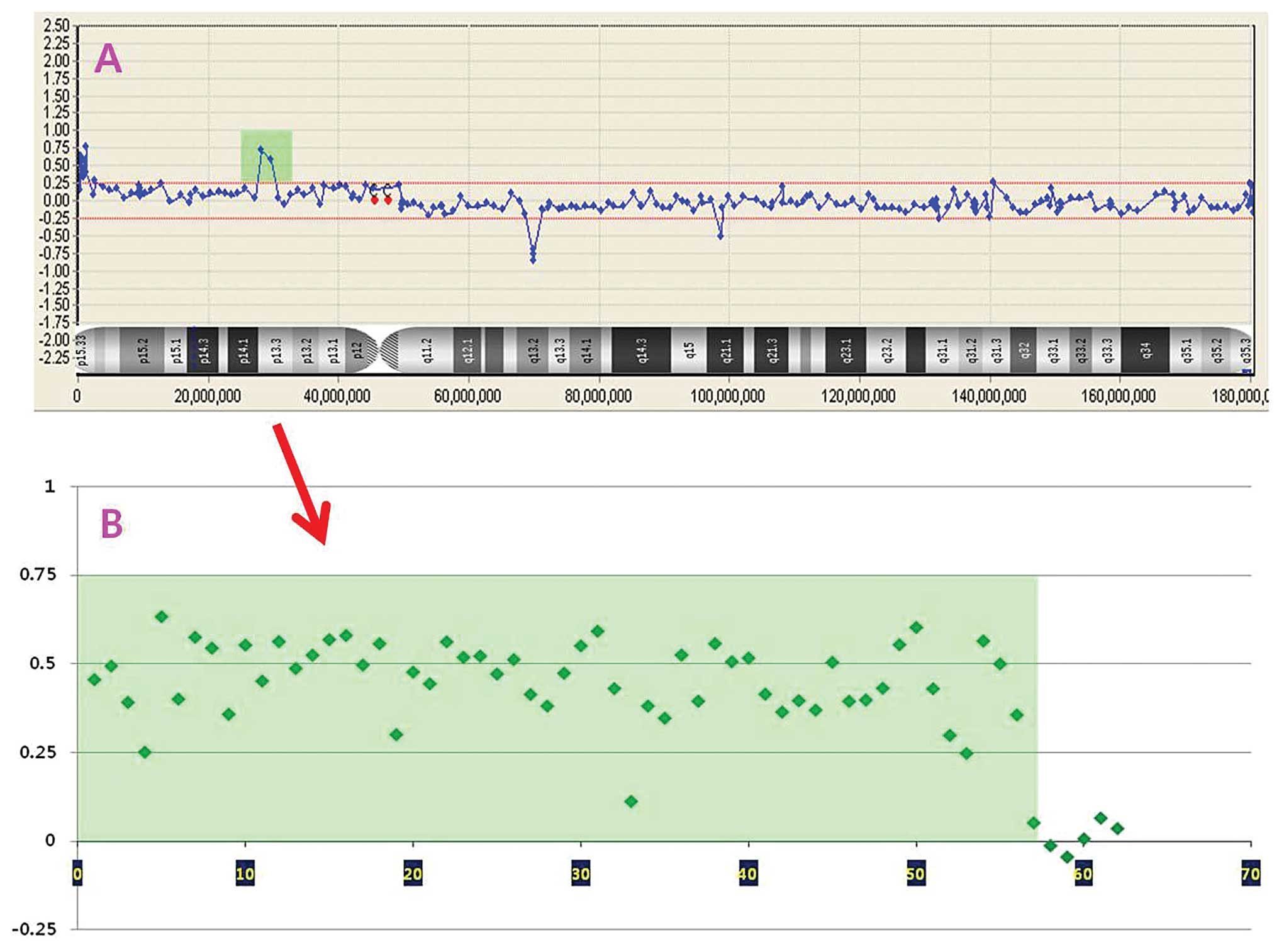
is represented on the array. An example of cryptic microduplication
detected by array CGH that could not be detected with the
conventional G-banded karyotyping is shown in Fig. 2.
Many studies aimed at assessing the diagnostic
capabilities of array CGH in screening for hidden chromosome
aberrations in patients with DD, ID, ASD or MCA who have an
apparently normal karyotype. It has been clearly established that
for these categories of disorders, the yield of array CGH is
significantly higher than karyotyping and any other molecular
approaches. The comparison of each technology, its uses and
limitations for understanding genomic imbalances is provided in
Table I.
 | Table IComparison of technical details
between conventional cytogenetic and major molecular
approaches. |
Table I
Comparison of technical details
between conventional cytogenetic and major molecular
approaches.
| Techniques | Resolution | Applications | Specific advantages
of the techniques | Limitations |
|---|
| G-banded
karyotyping | 10–15 Mb | Identification of
numerical and structural chromosomal anomalies | Provides global
information in a single assay; simple and robust procedures | Inability to detect
minor rearrangements (submicroscopic or cryptic
rearrangements) |
| FISH | 1–3 Mb | Identification of
the presence, number of copies per cell, and localization probe
DNA | Fast
characterization of marker chromosome origin, high sensitivity | Structural
anomalies cannot be detected in non dividing cells |
| Multicolor
karyotyping | 1–2 Mb | Detection of subtle
rearrangements, complex translocation, small marker, ring and
double minute chromosomes | Accurate origin
identification of all segments in complex rearrangements; fast
characterization of euchromatic marker chromosome content | Inability to detect
paracentromeric inversion, duplicatons, microdeletions and cryptic
translocation; difficulty to distinguish between artifacts and
aneuploidy/polyploidy |
| CGHb | 3–5 Mb | Interpretation of
highly complex karyotype with accurate chromosomal location of
imbalance | Whole genome
scanning technique without need of proliferating patient material,
locus-specific detection of gene amplification, band-specific
information on imbalance size | Balanced
rearrangements and imbalances present in low frequencies remain
undetected |
| Array CGH | 20–150 Mb | Identification of
cryptic rearrangements (aneuploidy, deletions, duplications, or
amplifications) of any locus represented on the array | Precise information
of whole genome in a single experiment; high-resolution
target-specific detection of gene amplification, submicroscopic
information on imbalances | Inability to detect
balanced translocation, inversion, mosaicism less than 50%, and
heterochromatic region |
Postnatal diagnostic applications of array
CGH
Array CGH has given the clinician a greater
appreciation of variability in the clinical presentation of many
well-described conditions and allowed for the discovery of
unsuspected imbalances in individuals with congenital and
developmental abnormalities. The use of array CGH circumvents the
problems associated with conventional cytogenetic testing and it
has become a first-tier test in the evaluation of atypical
deletions and microduplications in these population groups.
Based on the literature review, the overall rate of
detection of genomic abnormalities using a microarray approach in
patients with DD, ID or MCA with normal results from prior routine
cytogenetic testing is estimated to be between 8–20% (4,5,16,17). In the study by Dave and Sanger
(16) the additional application
of array CGH increased the identification of genomic imbalances by
∼8% in the population of DD with normal conventional banding
results. Similar to this result, an array analysis on 13,926
subjects with ID and/or MCA, whom had normal conventional
cytogenetic studies, reported an overall diagnostic rate of 10% for
pathogenic genomic imbalances (4). Another retrospective analysis of
36,325 patients with DD also demonstrated that a pathogenic
abnormality could be detected in ∼19% of unselected DD/ID patients
via genome-wide array approach (5). A review by the International
Standard Cytogenomic Array (ISCA) consortium (17) stated that the increased diagnostic
yield of array CGH in individuals with congenital anomalies and DD
was up to 15–20%, versus the 3% yield of conventional
karyotyping.
During the last few years, improved resolution of
array techniques has led to a significant increase in the detection
rate of chromosomal aberrations in patients with MR. Many studies
aimed to assess the diagnostic capabilities of array CGH to
identify hidden chromosome aberrations in patients with MR who had
an apparently normal karyotype (18–22). The study of de Vries et al
(18) suggested that the
diagnostic yield of array CGH in the general population of patients
with MR is at least twice as high as that of standard GTG-banded
karyotyping. Also, it has been demonstrated in several other groups
of hundreds of patients with idiopathic MR who had an apparently
normal karyotype that the diagnostic yield of rearrangements have
successfully increased up to 15–20% after exclusion of inherited
anomalies using array platforms (19–21). In addition, the identification of
submicroscopic subtelomeric alterations of idiopathic MR patients,
as well as the sporadic studies of submicroscopic interstitial
chromosomal rearrangements, suggests that a substantial portion of
idiopathic MR may be caused by smaller chromosomal rearrangements.
These observations make it clear that higher resolution screening
techniques for the detection of small deletions or duplications at
any chromosomal position will drastically increase the elucidation
of human genetic diseases (22).
Many recently published data concluded that array
technologies are being increasingly used in patients with DD, ID,
ASD or MCA, and microarrays should now be adopted as a first-tier
test in place of conventional karyotyping and/or FISH analysis in
these population groups (6,7,23).
Miller et al (23)
recommended a microarray approach as a first-tier test based on
studies of 21,698 patients with DD/ID, MCA and/or ASDs in whom the
diagnostic yield was 12.2% higher than that of a G-banded metaphase
karyotype. Shen et al (6)
also investigated the utility of microarray approach in the
evaluation of children with ASD and provided evidence that
microarray analysis should be the first-line genetic investigation
for children with autism.
From all these considerations, it is clear that a
high-resolution array platform covering the whole genome would
provide much more informative results than one containing only low
coverage limited to postnatal disease-associated regions. Truly
balanced rearrangements and low-level mosaicism are generally not
detectable by arrays, but these are relatively infrequent causes of
abnormal phenotypes in these categories of patients (<1%)
(23). Bui et al (24) also raised the question whether it
matters if the balanced alterations are not detected. For example,
carriers of balanced Robertsonian translocations are at risk for
uniparental disomy (UPD). Even though, array CGH cannot identify
UPD, single nucleotide polymorphism (SNP) arrays can detect regions
or chromosomes with copy-neutral absence of heterozygosity, which
may indicate an isodisomy (25).
Although, cytogeneticists will continuously require
additional training and laboratories must become appropriately
equipped, array CGH holds the promise of being the initial
diagnostic tool in the identification of visible and submicroscopic
chromosome abnormalities, the latter of which may be missed by
routine cytogenetics (11).
Further research is required before we can answer the question of
an absolute detection rate over conventional karyotyping in these
areas on a large cohort that has undergone both karyotyping and
analysis using a commercial reproducible array.
Conclusions and future perspectives
In this review, we present a critical appraisal of
the literature to date and provide a summary for clinicians for
patients with DD, ID, ASD or MCA. This systematic review provides
evidence of the relative advantage of using array testing in
postnatal diagnosis, even when the karyotype is normal (26).
Given the potential described in this article, we
anticipate array CGH to be the initial postnatal diagnostic
approach for the identification of chromosomal abnormalities and
envision that array CGH will play a major discovery role to reveal
the cryptic and/or complex nature of chromosome arrangements in the
very near future.
Acknowledgements
This study was financially supported
by the Korea Nazarene University Research Foundation in 2012.
References
|
1.
|
MR SpeicherNP CarterThe new cytogenetics:
blurring the boundaries with molecular biologyNat Rev
Genet6782792200510.1038/nrg169216145555
|
|
2.
|
JP MaciejewskiGJ MuftiWhole genome
scanning as a cytogenetic tool in hematologic
malignanciesBlood112965974200810.1182/blood-2008-02-13043518505780
|
|
3.
|
JU KangSH KooKC KwonJW ParkJM
KimIdentification of novel candidate target genes, including EPHB3,
MASP1 and SST at 3q26.2–q29 in squamous cell carcinoma of the
lungBMC Cancer9237200919607727
|
|
4.
|
GS SagooAS ButterworthS SandersonC
Shaw-SmithJP HigginsH BurtonArray CGH in patients with learning
disability (mental retardation) and congenital anomalies: updated
systematic review and meta-analysis of 19 studies and 13,926
subjectsGenet
Med11139146200910.1097/GIM.0b013e318194ee8f19367186
|
|
5.
|
R HochstenbachE van BinsbergenJ
EngelenArray analysis and karyotyping: workflow consequences based
on a retrospective study of 36,325 patients with idiopathic
developmental delay in the NetherlandsEur J Med
Genet52161169200910.1016/j.ejmg.2009.03.015
|
|
6.
|
Y ShenKA DiesIA HolmClinical genetic
testing for patients with autism spectrum
disordersPediatrics125727735201010.1542/peds.2009-168420231187
|
|
7.
|
L EdelmannK HirschhornClinical utility of
array CGH for the detection of chromosomal imbalances associated
with mental retardation and multiple congenital anomaliesAnn NY
Acad Sci1151157166200910.1111/j.1749-6632.2008.03610.x19154522
|
|
8.
|
SG VorsanovaYB YurovIY IourovHuman
interphase chromosomes: a review of available molecular cytogenetic
technologiesMol Cytogenet31201010.1186/1755-8166-3-120180947
|
|
9.
|
BA BejjaniLG ShafferApplication of
array-based comparative genomic hybridization to clinical
diagnosticsJ Mol
Diagn8528533200610.2353/jmoldx.2006.06002917065418
|
|
10.
|
BA BejjaniLG ShafferClinical utility of
contemporary molecular cytogeneticsAnnu Rev Genomics Hum
Genet97186200810.1146/annurev.genom.9.081307.16420718949852
|
|
11.
|
LG ShafferBA BejjaniMedical applications
of array CGH and the transformation of clinical
cytogeneticsCytogenet Genome
Res115303309200610.1159/00009592817124414
|
|
12.
|
H TönniesModern molecular cytogenetic
techniques in genetic diagnosticsTrends Mol Med82462502002
|
|
13.
|
KD LichtenbeltNV KnoersGH
Schuring-BlomFrom karyotyping to array-CGH in prenatal
diagnosisCytogenet Genome
Res135241250201110.1159/00033406522086062
|
|
14.
|
BA BejjaniR SalekiBC BallifEA RoremK
SundinA TheisenCD KashorkLG ShafferUse of targeted array-based CGH
for the clinical diagnosis of chromosomal imbalance: is less
more?Am J Med Genet A134259267200510.1002/ajmg.a.3062115723295
|
|
15.
|
JR VermeeschH FieglerN de LeeuwGuidelines
for molecular karyotyping in constitutional genetic diagnosisEur J
Hum Genet1511051114200710.1038/sj.ejhg.520189617637806
|
|
16.
|
BJ DaveWG SangerRole of cytogenetics and
molecular cytogenetics in the diagnosis of genetic imbalancesSemin
Pediatr Neurol1426200710.1016/j.spen.2006.11.00317331878
|
|
17.
|
American College of Obstetricians and
GynecologistsACOG Committee Opinion No. 446: array comparative
genomic hybridization in prenatal diagnosisObstet
Gynecol11411611163200910.1097/AOG.0b013e3181c33cad20168129
|
|
18.
|
BB de VriesR PfundtM LeisinkDiagnostic
genome profiling in mental retardationAm J Hum
Genet77606616200516175506
|
|
19.
|
P StankiewiczAL BeaudetUse of array CGH in
the evaluation of dysmorphology, malformations, developmental
delay, and idiopathic mental retardationCurr Opin Genet
Dev17182192200710.1016/j.gde.2007.04.00917467974
|
|
20.
|
LE VissersBB de VriesK OsoegawaArray-based
comparative genomic hybridization for the genomewide detection of
submicroscopic chromosomal abnormalitiesAm J Hum
Genet7312611270200310.1086/37997714628292
|
|
21.
|
C Shaw-SmithR RedonL RickmanMicroarray
based comparative genomic hybridisation (array-CGH) detects
submicroscopic chromosomal deletions and duplications in patients
with learning disability/mental retardation and dysmorphic
featuresJ Med Genet412412424128200410.1136/jmg.2003.017731
|
|
22.
|
JR VermeeschC MelotteG FroyenMolecular
karyotyping: array CGH quality criteria for constitutional genetic
diagnosisJ Histochem
Cytochem53413422200510.1369/jhc.4A6436.200515750031
|
|
23.
|
DT MillerMP AdamS AradhyaConsensus
statement: chromosomal microarray is a first-tier clinical
diagnostic test for individuals with developmental disabilities or
congenital anomaliesAm J Hum
Genet86749764201010.1016/j.ajhg.2010.04.00620466091
|
|
24.
|
TH BuiA VetroO ZuffardiLG ShafferCurrent
controversies in prenatal diagnosis 3: is conventional chromosome
analysis necessary in the post-array CGH era?Prenat
Diagn31235243201110.1002/pd.272221374637
|
|
25.
|
BH FaasI van der BurgtAJ
KooperIdentification of clinically significant, submicroscopic
chromosome alterations and UPD in fetuses with ultrasound anomalies
using genome-wide 250k SNP array analysisJ Med
Genet47586594201010.1136/jmg.2009.075853
|
|
26.
|
SC HillmanS PretloveA CoomarasamyDJ
McMullanEV DavisonER MaherMD KilbyAdditional information from array
comparative genomic hybridization technology over conventional
karyotyping in prenatal diagnosis: a systematic review and
meta-analysisUltrasound Obstet Gynecol37614201110.1002/uog.7754
|