1. Introduction
Osteoarthritis (OA) is a highly prevalent
degenerative joint disease characterized by the progressive
destruction of articular cartilage, synovial hyperplasia and
sclerosis of the subchondral bone (1–4).
In the United States, knee and/or hip OA are the most common causes
of walking-related disability among older adults (5). With the increasing incidence of
obesity and an aging population, the public health consequences of
OA and OA-related disability are have become a pressing issue, and
novel therapeutic strategies are urgently required (6).
Cartilage is classified histologically into hyaline,
elastic, and fibrocartilaginous, depending on the molecular
composition (7). Articular
cartilage is a form of hyaline cartilage and is divided into 4
zones: superficial, transitional, radial and calcified cartilage
(8). These zones are
characterized by a distinct organization of the collagen network,
and by differences in the amounts and types of proteoglycans. In
healthy articular cartilage, the principal molecular component is
type II collagen, although collagens III, VI, IX, X, XI, XII and
XIV all exist in the mature matrix (9–11).
Additionally, aggrecan, a large chondroitin sulfate proteoglycan,
is the main proteoglycan present in cartilage, which also includes
syndecans, glypican, decorin, biglycan, fibromodulin, lumican,
epiphycan and perlecan (12).
Apart from the extracellular matrix (ECM), chondrocytes are the
only cells found in healthy cartilage. Chondrocytes are arranged in
a zonal stratification and embedded in the arcade-like network of
collagen fibrils intermingled with proteoglycans (11). Chondrocytes respond to their
microenvironment to regulate articular cartilage homeostasis by
balancing the synthesis and degradation of ECM components. The ECM
is degraded by proteolytic enzymes, including matrix
metalloproteinases (MMPs) and the ‘A disintegrinand
metalloproteinase domain with thrombospondin-like motifs’ family
(ADAMTS) (13–15). In OA, cartilage degradation occurs
in 2 phases: the degradative and biosynthetic phase. In the
degradative phase, ADAMTS and MMPs digest the ECM in the presence
of inflammatory cytokines. Consequently, in the biosynthetic phase,
the chondrocytes attempt to repair the damaged ECM. However, matrix
synthesis is likewise inhibited by inflammatory factors through the
downregulation of ECM genes, thus accelerating cartilage erosion
(14).
microRNAs (miRNAs or miRs) are a class of non-coding
single-stranded RNAs of 18–22 nucleotides (nt). They regulate gene
expression at the post-transcriptional, level and therefore affect
cell fate during cell proliferation and differentiation (16). This regulatory mechanism is often
modulated by the promotion of mRNA degradation and/or the
repression of translation through sequence-specific interactions
with the 3′-untranslated regions (3′-UTRs) of specific mRNA
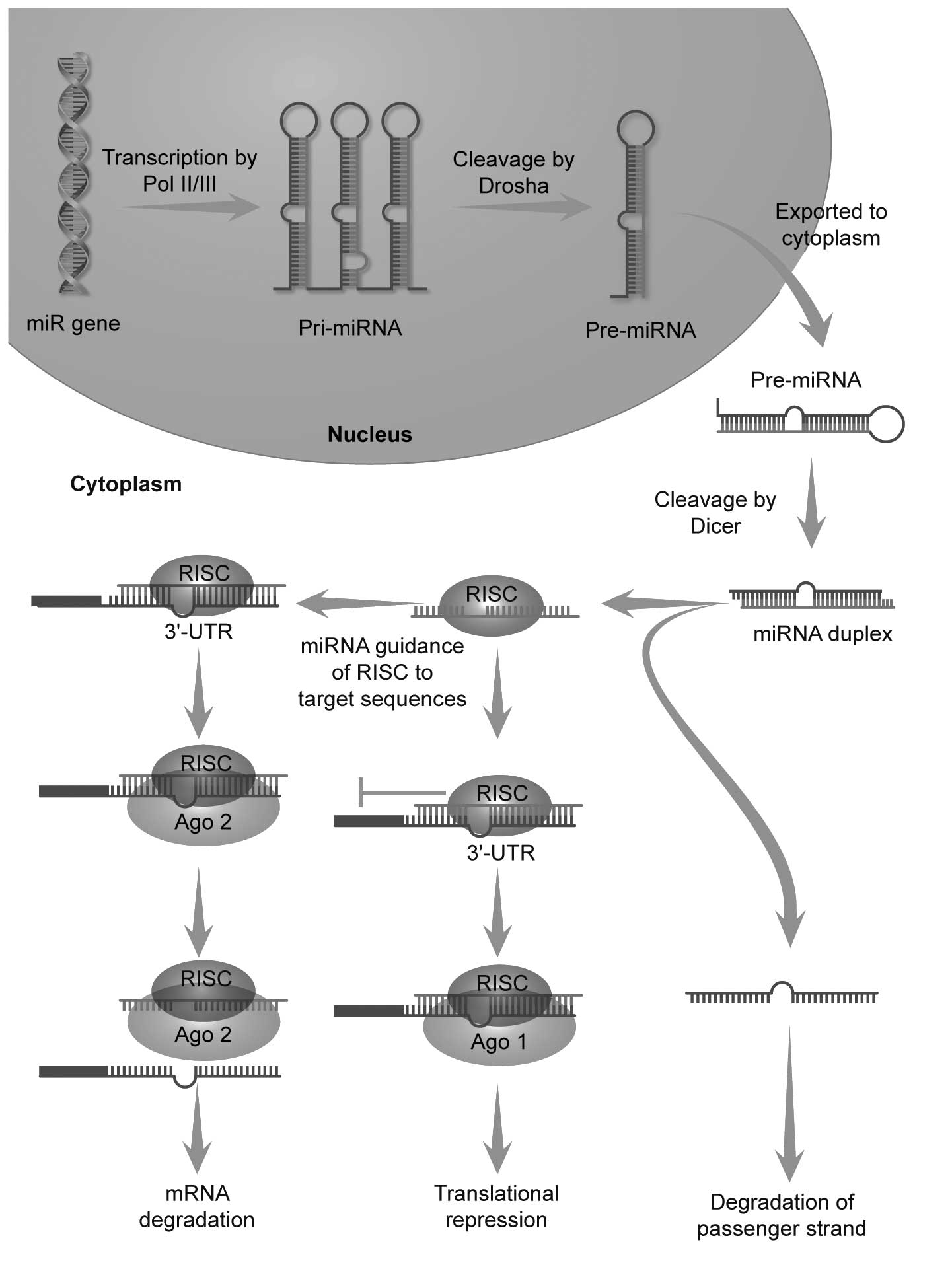
targets. Genes encoding miRNAs are transcribed by RNA polymerase II
or III to generate Pri-miRNAs, which are processed to Pre-miRNAs by
Drosha in the nucleus. Pre-miRNAs are exported to the cytoplasm
where they are further processed by the RNase III ribonuclease,
Dicer, to generate ~22-nt single-stranded mature miRNAs (17–21), which then combine with the
RNA-induced silencing complex (RISC) by the core unit Argonaute
(Ago) (16,22,23). The miRNA-RISC complex binds target
mRNAs and mediates mRNA translational repression or degradation
(24–27) (Fig.
1).
miRNAs are evolutionarily conserved and have been
found in various organisms (28).
In the human musculoskeletal system, the conditional knockout of
Dicer in the limb mesenchyme during the early stages of mice
embryonic development leads to the formation of smaller limbs
(29), suggesting that miRNAs
play a prominent role in skeletal development. Furthermore,
Dicer-null growth plates display a lack of chondrocyte
proliferation but an enhanced hypertrophic chondrocyte
differentiation, indicating that Dicer and miRNAs exhibit distinct
functional effects at different stages of chondrocyte development
(29). Given the importance of
miRNAs in chondrocyte development, we reviewed recent research on
miRNA regulation in cartilage biology, with the aim to further
understand chondrogenesis and OA development, and the development
of potential gene therapeutic strategies for OA prevention and
treatment.
2. Literature search and literature
selection
Literature search
A systematic literature review was conducted using
the PubMed and Embase databases on April 1, 2013. Several
combinations of terms and expressions were evaluated, including
both MeSH and free text terms. By analyzing the articles retrieved
from each combination, we selected the following final search
expression: (cartilage OR chondrocyte, chondrogenesis OR
chondrocytogenesis OR osteoarthritis) and (microRNA OR non-coding
RNA). Additionally, we manually searched the reference lists of the
identified aritcles. We used the Preferred Reporting Items of
Systematic Reviews and Meta-Analyses (PRISMA) guidelines in the
planning and execution of this study.
Literature selection
We selected articles, written in the English
language, in which scientific detail and reporting were sufficient
to enable our understanding of the material, and which presented
novel findings. In cases of multiple or serial reports, we selected
either the first or the most detailed report for inclusion. We
preferentially selected articles in which mechanistic data
supported observational findings.
Our literature search retrieved 457 studies. We
selected 59 studies which were of sufficient reporting rigor or
novelty, and provided mechanistic data of the biology of
chondrogenesis and OA.
3. miRNAs in chondrocytogenesis
Chondrocytes are derived from the chondrogenic
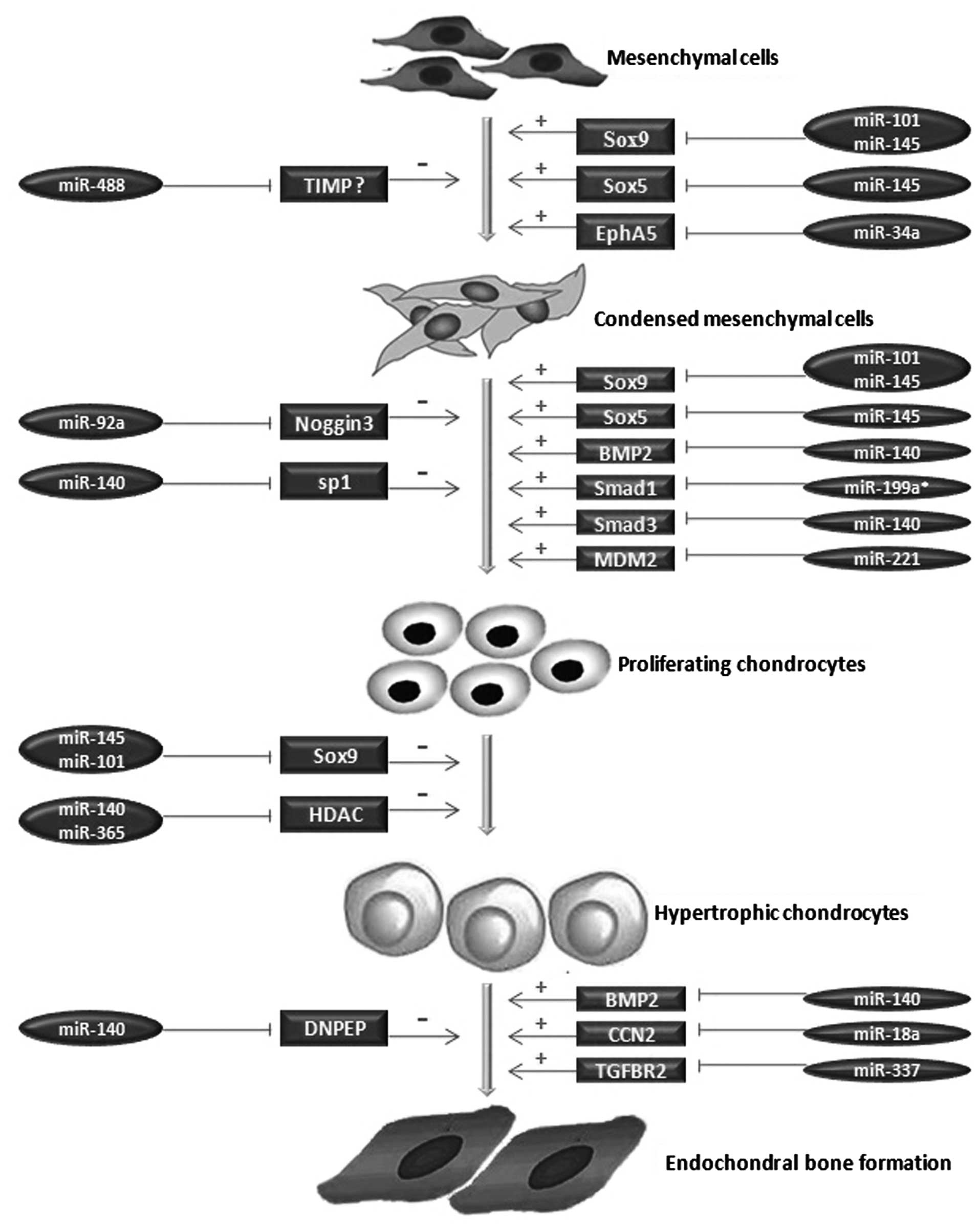
differentiation of mesenchymal stem cells (MSCs) (30). The process of chondrogenic
differentiation mainly includes 6 phases: mesenchymal cells
(chondroprogenitors), condensed mesenchymal cells, chondrocytes,
proliferating chondrocytes, pre-hypertrophic chondrocytes and
hypertrophic chondrocytes (31).
A number of transcription factors and cytokines influence discrete
steps in the chondrocyte differentiation pathway. These include
members of the Sox family [Sox9, Sox5 and Sox6 (32,33)], bone morphogenetic proteins (BMPs)
(34), connective tissue growth
factor/cysteine-rich 61/nephroblastoma overexpressed (CCN) family
protein 2 (CCN2) (35), as well
as others. Additionally, several miRNAs participate in the
regulation of chondrogenesis by targeting these transcription
factors and growth cytokines (Table
I and Fig. 2).
 | Table ISummary of microRNAs in
chondrocytes. |
Table I
Summary of microRNAs in
chondrocytes.
| microRNAs | Targets | Regulatory
role |
Authors/(Refs.) | Study models |
|---|
| miR-101 | Sox9 | Chondrocyte
differentiation (−) | Dai et al
(46) | Rat
chondrocytes |
| miR-145 | Sox9 | Chondrocyte
differentiation (−) | Yang et al
(32) | Murine MSCs |
|
miR-199a* | Smad1 | Early chondrocyte
differentiation (−) | Lin et al
(52) | Pluripotent
C3H10T1/2 stem cells |
| miR-92a | Noggin3 | Proliferation and
differentiation of chondrogenic progenitors (+) | Ning et al
(34) | Fish embryos |
| miR-488 | Not detected
(TIMP?) | Condensed
mesenchymal cells (+) | Song et al
(59) | Mesenchymal cells
derived from the distal tips of Hamburger-Hamilton (HH) stage 22/23
embryo leg buds of fertilized white Leghorn chicken eggs |
| miR-34a | EphA5 | Condensed
mesenchymal cells (−) | Kim et al
(60) | Chick limb
mesenchymal cells; human chondrocytes |
| miR-221 | MDM2 | Proliferation of
chondroprogenitors (−) | Kim et al
(60) | Mesenchymal cells
derived from the distal tips of HH stage 22/23 embryo leg buds of
fertilized White Leghorn chicken eggs |
| miR-194 | Sox5 | Differentiated
chondrocyte and proliferating chondrocyte (−) | Xu et al
(47) | The human ASCs; 2
weeks old Wistar male rats; articular chondrocytes isolated from
the knee joints of juvenile rats |
| miR-140 | HDAC4 | Hyperdrophic
chondrocyte (+) | Tuddenham et
al (71) | Mouse cells |
| CXCL12 | Hyperdrophic
chondrocyte (−) | Nicolas et
al (72) | Mouse cells |
| SP1 | Chondrocyte
proliferation (+) | Yang et al
(67) | Limb bud
mesenchymal cells |
| DNPEP | Endochondral bone
formation (+) | Nakamura et
al (58) | miR-140-null mice;
primary chondrocyte |
| BMP2 | Endochondral bone
formation (−) | Nicolas et
al (56) | Chicken primary
chondrocytes; primary human chondrocytes |
| Smad3 | Chondrocyte
differentiation (−) | Pais et al
(57) | Mouse cells |
| miR-1 | Not detected | Hyperdrophic
chondrocyte (−) | Sumiyoshi et
al (68) | Human chondrocytic
HCS-2/8 cells; primary chicken chondrocytes |
| miR-365 | HDAC4 | Hyperdrophic
chondrocyte (+) | Guan et al
(69) | Chicken primary
proliferative chondrocytes |
| miR-18a | CCN2/CTGF | Endochondral bone
formation (−) | Ohgawara et
al (35) | Human
chondrosarcoma-derived chondrocytic cell line, HCS-2/8; human
cervical cancer-derived cell line, HeLa; chicken sternum
chondrocytes |
| miR-337 | TGFBR2 | Endochondral bone
formation (−) | Zhong et al
(73) | Femoral head
cartilage tissues were obtained from Sprague-Dawley (SD) or Dark
Agouti (DA) rats |
| miR-23b | Protein kinase A
(PKA) | Chondrogenic
differentiation (+) | Ham et al
(74) | Human mesenchymal
stem cells (hMSCs); normal cell line Chinese hamsterovary (CHO)
cells |
miRNAs and the Sox family in
chondrocytogenesis: miR-101, miR-145 and miR-194
Sox proteins belong to the high-mobility group (HMG)
superfamily (36). Sox genes are
indispensable to multiple aspects of cartilage development by
activating the expression of cartilage-specific ECM components.
Sox9 is a transcription factor belonging to the Sox protein family,
which is essential for chondrogenesis, and has been designated a
‘master regulator’ of the chondrocyte phenotype (37,38). Sox9 is expressed in all
chondroprogenitor cells, predominantly in mesenchymal condensations
and cartilage (39). Sox9 can
bind to and activate chondrocyte-specific enhancer elements in
Col2a1, Col9a1, Col11a2 and aggrecan to induce cartilage-specific
gene expression (40–43). Additionally, Sox9 prevents
chondrocyte hypertrophy and exerts a re-differentiation effect on
dedifferentiated osteoarthritic chondrocytes (44,45). Of note, miR-101 participates in
the interleukin (IL(-1β-induced downregulation of collagen type II
and aggrecan, possibly functioning through its target gene,
Sox9 (46). Therefore,
miR-101 inhibition is effective in preventing IL-1β-induced
chondrocyte ECM degradation (46). Additionally, miR-145 expression is
gradually reduced during transforming growth factor-β
(TGF-β)-induced chondrogenic differentiation. Through the
3′-UTR-reporter assay and gain- or loss-of-function experiments,
Sox9 has also identified as an miR-145 target (32). This was further supported by the
fact that Sox9 was identified as a direct target of miR-145
in human chondrocytes (33).
Thus, the attenuation of miR-145 expression can positively regulate
Sox9 expression, resulting in the promotion of chondrogenic
differentiation.
In addition to Sox9, Sox5 and
Sox6 (which share a high degree of sequence identity) belong
to a different subgroup of Sox proteins and present no sequence
homology with Sox9, apart from the HMG-box. Sox5 and Sox6 are
co-expressed with Sox9 during chondrogenic differentiation. Both
genes are also expressed in several non-chondrogenic tissues. Owing
to the presence of a highly conserved coiled-coil domain, Sox5 and
Sox6 form homodimers and heterodimers, which bind much more
efficiently to pairs of HMG-box-binding sites than to single
binding sites. Unlike Sox9, Sox5 and Sox6 do not contain a
transcriptional activation domain. Sox5 and Sox6 cooperate with
Sox9 to activate the Col2a1 enhancer and expression of Col2a1 and
aggrecan genes (39). Of note,
Sox5 was identified as a target of miR-194 based on luciferase
assay analysis (47). The
expression of miR-194 was gradually downregulated during the
chondrogenic differentiation of human adipose-derived stem cells
(hASCs) (47). However, miR-194
was upregulated under pathological conditions (e.g., OA). In gain-
or loss-of-function experiments, the downregulation of miR-194
increased the expression of its direct target gene, Sox5, resulting
in enhanced chondrogenic differentiation (47). Taken together, these findings
suggest that miR-101, miR-145 and miR-194 are important for
chondrogenesis by targeting the Sox family.
miRNAs and the BMP family in
chondrocytogenesis: miR-199a*, miR-140 and miR-92a
BMPs are members of the TGF-β family that bind to
type II and I serine-threonine kinase receptors, and transduce
signals through Smad and non-Smad signaling pathways (48). BMPs, particularly BMP2 and BMP4,
are very powerful growth factors that induce cartilage formation by
stimulating chondrocyte differentiation (36). Smad signaling is important for BMP
signaling. Smad1 is a downstream signaling molecule of BMP2
(49–51). Of note, Smad1 was identified as
the direct target of miR-199a*, which is specifically
expressed in the skeletal system (52). miR-199a* significantly
inhibited early chondrogenesis, as revealed by the reduced
expression of early chondrogenesis marker genes, such as cartilage
oligomeric matrix protein (COMP), type II collagen and Sox9,
whereas anti-miR-199a* increased the expression of these
genes. Therefore, miR-199a* acts as a negative regulator
of early chondrogenic differentiation through the suppression of
Smad1 in the BMP signaling pathway.
In addition to miR-199a*, miR-140
participates in the regulation of BMP signaling during cartilage
development (54). miR-140 is
highly expressed in chondrocytes, and miR-140 expression increases
during chondrogenic differentiation from MSCs (55). Standard luciferase assays have
confirmed that BMP2 is a direct target of miR-140 (56). In addition to BMP2, Smad3 was
identified as a direct target of miR-140, and the suppression of
Smad3 by miR-140 can subsequently inhibit the TGF-β pathway in
chondrocytes (57). Furthermore,
miR-140-null mice exhibit skeletal defects, possibly due to a
reduction in basal BMP signaling in miR-140-null chondrocytes
(58).
miR-92a has also been identified in the regulation
of the BMP signaling pathway. miR-92a is highly enriched in
chondrogenic progenitors. The inactivation of miR-92a has been
shown to result in poor proliferation, impaired differentiation and
the unsustainable survival of chondrogenic progenitors (34). The BMP antagonist gene, Noggin3
(Nog3), has been reported to be a direct target of miR-92a. Nog3
represses BMP activity to prevent apoptosis. Therefore, miR-92a
inactivation stabilizes Nog3 mRNA and represses BMP signaling,
resulting in abnormal behaviors of chondrogenic progenitors. By
contrast, the ectopic expression of the miR-92a duplex reduces Nog3
mRNA levels, and thereby suppresses BMP signaling and promotes cell
apoptosis. Therefore, miR-92a maintains BMP activity during
cartilage formation by targeting Nog3 (34). Collectively, these data
demonstrate that miR-199a*, miR-140 and miR-92a are
critical for chondrogenesis by regulating the BMP signaling
pathway.
miRNAs in the process of chondrogenic
differentiation
miRNAs in condensed mesenchymal cells:
miR-488 and miR-34a
In addition to their involvement in the
aforementioned 2 key families, miRNAs regulate chondrocyte
differentiation at different stages. Pre-cartilage mesenchymal
condensation is characterized by the production of sulfated
proteoglycans and the switch from type I to II collagen synthesis.
Thereafter, differentiated chondrocytes proliferate and secrete
increasing amounts of ECM macromolecules until each single cell is
completely surrounded by a matrix. In condensed MSCs, miR-488
expression is upregulated at the pre-condensation stage and then
downregulated at the post-condensation stage. The blocking of
miR-488 via antisense oligonucleotides reduces integrin b1 and
phosphorylated focal adhesion kinase (FAK) levels, and suppresses
cell motility and migration (59). In parallel with these
observations, treatment with anti-miR-488 oligonucleotides has been
shown to upregulate MMP-2 activity and inhibit cellular
condensation (59). The direct
target of miR-488 in the modulation of focal adhesion activity
remains unidentified, although a possible direct target is tissue
inhibitor of metalloproteinases (TIMP)-1. In summary, miR-488 is a
regulator in cell-ECM interaction through the modulation of focal
adhesion activity by MMP-2 during chondrogenesis of limb
mesenchymal cells (59).
Additionally, miR-34a expression is induced by c-Jun
N-terminal kinase (JNK) in the chondrogenic differentiation of
chick limb mesenchymal cells (60) and by IL-1β stimulation in rat
primary chondrocytes (61).
miR-34a is a negative modulator of chondrogenesis, particularly in
the migration of chondroblasts, by targeting EphA5, which inhibits
cellular condensation (60).
Furthermore, locked nucleotide analogue (LNA)-modified miR-34a
antisense can prevent the downregulation of Col2a1 and the
upregulation of iNOS in chondrocytes (61). Therefore, miR-488 and miR-34a
regulate condensed mesenchymal cells.
miRNAs in proliferating chondrocytes:
miR-221 and miR-140
Chondrocyte proliferation is the process for
acquiring a chondrocyte phenotype, leading to the formation of
unique tissues as cartilage (31). Of note, treatment with a JNK
pathway inhibitor has been shwon to upregulate miR-221 expression
in chondroprogenitor cells, leading to reduced proliferation and
pre-cartilage condensation (62).
Mouse double-minute 2 homolog (MDM2) is a relevant miR-221 target.
miR-221 has been found to be both necessary and sufficient for the
downregulation of MDM2 expression, which prevents slug protein
degradation, and thus negatively regulates chondroprogenitor
proliferation (62).
Additionally, in articular cartilage, miR-221 has been suggested to
be a potential regulator of the mechanotransduction pathway
(63). Sp1 has been previously
reported to be expressed in chondrocytes to activate ColII
expression (29,64) and is also a critical transcription
factor in cell cycle inhibition through the activation of
p15INK4b and p21Waf/Clip promoters in
vitro (65,66). Furthermore, the suppression of Sp1
activity by miR-140 has been implicated in the maintenance of
chondrocyte proliferation (67).
miRNAs in hypertrophic chondrocytes:
miR-1, miR-365 and miR-140
Following proliferation, chondrocytes in the center
region increase in size (hypertrophic chondrocytes) and secrete and
organize a different ECM (appearance of type X collagen). As the
miRNA most repressed following chondrocyte hypertrophic
differentiation, miR-1 was found to participate in chondrocytic
phenotype regulation during the late stage of the differentiation
process. Although the direct target remains unknown, miR-1
overexpression suppresses the expression of the major cartilaginous
proteoglycan gene, aggrecan, in both human chondrocytic HCS-2/8
cells and chicken normal chondrocytes, suggesting that miR-1
participates in maintaining the integrity of the cartilage tissue
(68).
By contrast, miR-365 expression is elevated in the
pre-hypertrophic zone to stimulate chondrocyte proliferation and
differentiation into hypertrophic chondrocytes (69). Of note, miR-365 is a
mechano-responsive microRNA that parallels the mechanical induction
of Indian hedgehog (Ihh) in primary chicken chondrocytes.
Similarly, miR-365 expression coincides with the Ihh expression
region in vivo. Indeed, miR-365 increases the expression of
Ihh and the hypertrophic marker, type X collagen, whereas
anti-miR-365 inhibits the expression of these genes (69). Further studies have demonstrated
that histone deacetylase 4 (HDAC4), an inhibitor of chondrocyte
hypertrophy (70), is an miR-365
target. miR-365 inhibits both endogenous HDAC4 protein levels and
the activity of a reporter gene bearing the 3′-UTR of HDAC4 mRNA.
Therefore, miR-365 inhibits HDAC4 and consequently induces the
expression of its downstream molecules, Ihh and Runx2, during
chondrocyte hypertrophy (69).
Additionally, studies have validated experimentally that miR-140
targets HDAC4 to regulate chondrocyte differentiation, and have
determined the potential role of miR-140 in long bone development
(71). Furthermore, Cxcl12 was
identified as a target of miR-140 by Northern blot analysis and a
luciferase reporter assay (72).
miRNAs in endochondral ossification:
miR-18a and miR-337
In endochondral ossification, the embryonic
cartilaginous model of most bones contributes to longitudinal
growth and is gradually replaced by bone. During endochondral
ossification, chondrocytes proliferate, undergo hypertrophy and
die. The cartilage ECM is then invaded by blood vessels,
osteoclasts, bone marrow cells and osteoblasts, the last of which
deposit bone on remnants of the cartilage matrix. miR-18a is the
most strongly downregulated miRNA in human chondrocytic HCS-2/8
cells compared with HeLa cells, suggesting a potential role of
miR-18a in chondrocytes (35).
Indeed, subsequent studies have confirmed that during the late
stage of chondrocyte differentiation, miR-18a targets the 3′-UTR of
the CCN family protein 2/connective tissue growth factor
(CCN2/CTGF), a molecule important for endochondral bone formation
(35). The introduction of
miR-18a effectively repressed CCN2 expression in chondrocytic cells
and thus significantly repressed the mature chondrocytic phenotype.
The overexpression of miR-18a also repressed aggrecan and Col2a1
expression (35). Furthermore,
many other targets, estrogen receptor, Smad2, insulin-like growth
factor (IGF)-1, and hypoxia inducible factor 1-α (HIF1-α), also
influence the final phenotypic changes.
The expression of miR-337 is significantly
downregulated and almost disappeares during the maturation phases
of endochondral ossification (73). miR-337 was found to directly
target the TGF-β type II receptor (TGFBR)2, which is well known for
its important roles in cartilage development (73). These data indicate that miR-18 and
miRNA-337 are associated with endochondral ossification of
chondrogenesis.
Other miRNAs involved in chondrogenesis:
miR-23b
The expression of miR-23b is induced by H-89, a
potential mediator of chondrogenic differentiation. When
upregulated by H-89, miR-23b can induce chondrogenic
differentiation by negatively inhibiting the protein kinase A (PKA)
signaling pathway, as evidenced by chondrogenic differentiation
observed following miR-23b overexpression in human mesenchymal stem
cells (hMSCs) (74).
4. miRNAs in osteoarthritis
The regulatory effects of miRNAs on OA are evident
from studies comparing miRNA expression in both OA and normal
articular tissues. Iliopoulos et al (75) measured the expression of 365
miRNAs and identified 9 significantly upregulated miRNAs and 7
downregulated miRNAs in OA cartilage, compared with normal
controls, suggesting that miRNAs are involved in OA development. In
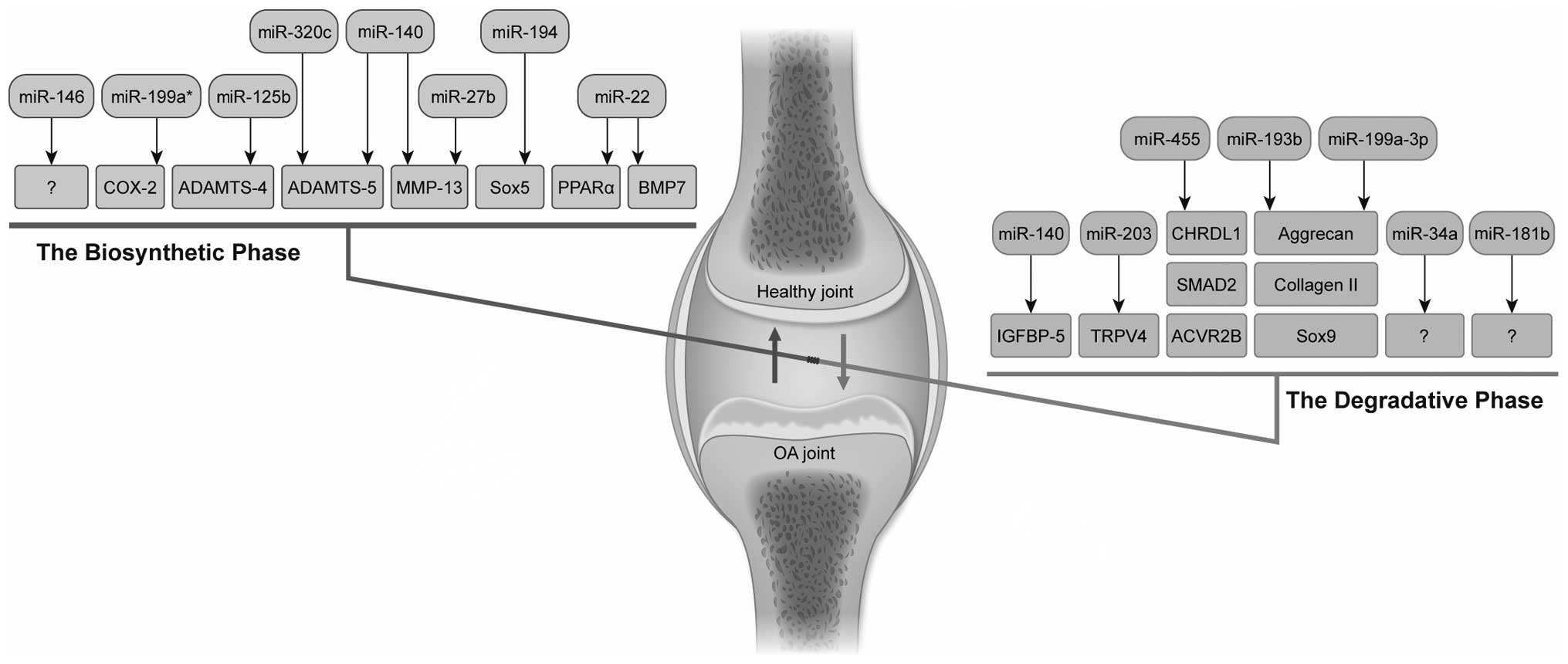
this review, we summarize the role of miRNAs in the development of
OA (Table II and Fig. 3).
 | Table IISummary of microRNAs in OA. |
Table II
Summary of microRNAs in OA.
| microRNAs | Targets | Regulatory
role |
Authors/(Refs.) | Study models |
|---|
| miR-140 | IGFBP-5 | OA pathogenesis
(+) | Tardif et al
(80) | Human cartilage
obtained from femoral condyles and tibial plateaus |
| ADAMTS-5 | OA pathogenesis
(−) | Miyaki et al
(78) | miR-140 null mice;
miR-140 transgenic (TG) mice; mouse OA model; mouse antigen-induced
arthritis (AIA) model |
| MMP-13 | OA pathogenesis
(−) | Liang et al
(53) | The human cartilage
cell line, C28/I2 |
| miR-194 | Sox5 | OA pathogenesis
(−) | Xu et al
(47) | The human ASCs; 2
weeks old Wistar male rats; articular chondrocytes isolated from
the knee joints of juvenile rats |
| miR-203 | TRPV4 | OA pathogenesis
(+) | Hu et al
(98) | Cartilage derived
from the temporomandibular joint (TMJ) of 1-week-old female SD
rats |
| miR-455 | ACVR2B, smad2,
CHRDL1 | Cartilage
homeostasis; OA pathogenesis (+) | Swingler et
al (95) | Human articular
cartilage obtained from the femoral heads of patients undergoing
total hip replacement surgery; SW-1353 cells; C3H10T1/2 cells; 3T3
cells; ATDC5 cells |
| miR-320c | ADAMTS-5 | OA pathogenesis
(−) | Ukai et al
(81) | Cartilage tissue
from patients with polydactylism, anterior cruciate ligament
injury, and osteoarthritis undergoing total knee arthroplasty |
| miR-125b | ADAMTS-4 | OA pathogenesis
(−) | Matsukawa et
al (82) | Cartilage tissues
obtained from OA patients undergoing total knee replacement |
| miR-27b | MMP-13 | OA pathogenesis
(−) | Akhtar et al
(87) | OA cartilage
samples obtained from patients who underwent total joint
arthroplasty; normal cartilage samples obtained from trauma
patients |
| miR-22 | PPARα, BMP7 | OA pathogenesis
(−) | Iliopoulos et
al (75) | Articular cartilage
obtained from femoral heads, femoral condyles and tibial plateaus
of patients with primary osteoarthritis undergoing hip or knee
replacement surgery |
|
miR-199a* | COX-2 | OA pathogenesis
(−) | Akhtar et al
(88) | Human
chondrocytes |
| miR-34a | Not detected | OA pathogenesis
(+) | Abouheif et
al (61) | Rat osteoarthritis
model |
| miR-181b | Not detected | OA pathogenesis
(+) | Song et al
(94) | Mesenchymal cells
derived from the distal tips of Hamburger-Hamilton (HH) stage 22/23
embryo leg buds of fertilized white Leghorn chicken eggs; human
chondrocytes |
| miR-146 | Not detected | OA pathogenesis
(−) | Jones et al
(91) | Cartilage and bone
obtained from patients with OA or not |
miR-193b
miR-199a-3p | Aggrecan, type 2
collagen, Sox9 | OA pathogenesis
(+) | Ukai et al
(81) | Cartilage tissue
from patients with polydactylism, anterior cruciate ligament
injury, and osteoarthritis undergoing total knee arthroplasty |
| miR-675 | Not detected | Cartilage
homeostasis (+) | Dudek et al
(96) | Human articular
cartilage |
miRNAs regulate proteolytic enzymes in
OA: miR-140, miR-320c, miR-125b and miR-27b
ADAMTS-5 is a critical proteolytic enzyme causing
aggrecan degradation in cartilage. ADAMTS-5-deficient animals are
resistant to cartilage degeneration in the surgical OA model and
inflammatory arthritis model (76,77). miR-140 is an miRNA that may
regulate ADAMTS-5 expression. miR-140 is specifically expressed in
the cartilage tissue of mouse embryos during long and flat bone
development, and is required for cartilage homeostasis and skeletal
development (71). miR-140
expression is decreased in OA chondrocytes following IL-1β
stimulation (55). Similarly,
mice lacking miR-140 are more prone to developing severe OA than
are wild-type mice. miR-140 overexpression in cartilage also
protects mice from cartilage degradation (78). Furthermore, miR-140 expression is
decreased in the knee synovial fluid in patients with OA, and is
negatively related to the severity of OA. This may be related to OA
occurrence and development, and may therefore represent a potential
molecular target for early diagnosis (79). It has also been revealed that
miR-140 suppresses ADAMTS-5 expression, thereby inhibiting OA-like
changes (78). Additionally,
miR-140 was identified as a negative feedback regulator of MMP-13,
a well-characterized key player in cartilage biology and OA
pathology owing to its capacity to degrade collagens and numerous
other matrix components (53).
Furthermore, IGFBP-5, a direct target of miR-140, is expressed in
human chondrocytes with its level being significantly lower in OA
(80).
Additionally, miR-320c was identified in the
regulation of ADAMTS-5 expression, although miR-320c expression is
apparently downregulated with age in human cartilage tissue. Of
note, ADAMTS-5 expression was downregulated in the miR-320c mimic
group and upregulated in the inhibitor group (81), suggesting that miR-320c is
involved in OA by regulating ADAMTS-5.
miR-125b expression is significantly lower in OA
chondrocytes than in normal chondrocytes. miR-125b seeds sequence
in 3′-UTR of human aggrecanase-1 (ADAMTS-4) mRNA, and suppresses
ADAMTS-4 mRNA expression by 72% and protein production by 62%
following IL-1β stimulation (82). Increased ADAMTS-4 expression is an
important factor in OA and other joint diseases, and therefore, it
is important for the control of miR-125b expression, which may
represent a novel approach for OA prevention and treatment.
Additionally, in other cell types (83,84), miR-125b suppresses MMP-13, an
important collagenase in articular cartilage (85), and vascular endothelialc adherin,
a regulator of angiogenesis, whose levels are increased in
OA-affected joints (86).
The human genome contains 2 miR-27 genes (miR-27a
and miR-27b) on chromosomes 19 and 9, respectively. Their mature
products differ by only 1 nucleotide in the 3′ region. Of note, no
change in miR-27a expression in OA has been reported. By contrast,
miR-27b was identified as a downregulated miRNA in IL-1β-stimulated
human OA chondrocytes. By using the 3′-UTR reporter assay and gain-
or loss-of-function experiments, MMP-13 was identified as a direct
target of miR-27b. MMP-13 exhibits broad substrate specificity and
can cleave types I, II, III, IV, X and XIV collagen; aggrecan; and
fibronectin, with the highest activity being directed toward type
II collagen in OA. Thus, upregulating miR-27b expression or
preventing its downregulation in vivo may present a novel
therapeutic and/or preventive approach in OA treatment (87). Thus, several miRNAs, including
miR-140, miR-320c, miR-125b and miR-27b, participate OA
pathogenesis by targeting the proteolytic enzymes.
miRNAs are involved in both
chondrogenesis and OA: miR-194, miR-199a* and
miR-34a
As discussed above, several miRNAs play important
roles in regulating chondrogenesis. In addition to their
physiological roles, these miRNAs appear to participate in OA
pathogenesis. For instance, the miR-194 level has been shown to be
reduced during chondrogenic differentiation by directly targeting
Sox5 expression. By contrast, miR-194 expression was upregulated
and Sox5 expression was downregulated in OA (47). Additionally, miR-199a*
targeted not only Sox9 in chondrocytes but also cyclooxygenase-2
(COX-2) in OA chondrocytes (88).
miR-199a* directly suppressed the luciferase activity of
a COX-2 3′-UTR reporter construct and inhibited IL-1β-induced
expression of COX-2 in OA chondrocytes. The modulation of
miR-199a* expression also significantly inhibited the
IL-1β-induced upregulation of mPGES1 and prostaglandin E2
production in OA chondrocytes. The activation of p38-MAPK
downregulated miR-199a* expression and induced COX-2
expression (88). Collectively,
these data suggest that miR-199a* participates in OA
development, and thus represents another potential target for OA
treatment.
Additionally, miR-34a is a negative modulator of
chondrogenesis, particularly in chondroblast migration, by
targeting EphA5, which inhibits cellular condensation.
Interestingly, miRNA-34a silencing can prevent chondrocyte
apoptosis, suggesting the involvement of miR-34a in cell apoptosis
(61,89,90). In agreement with these findings,
miR-34a expression was also upregulated in human OA cartilage
(91). Together, these data
suggest that miR-34a participates in promoting chondrocyte
apoptosis during OA development.
Other miRNAs in OA
Downregulated miRNAs in OA:
miR-146
miR-146 expression is high in low-grade OA cartilage
and inversely correlates with cartilage degeneration in late-stage
OA cartilage (91,92). Thus, the miR-146 expression
pattern appears to depend on OA severity. A target of miR-146 in
chondrocytes has not been detected yet; however, TNF-α production
is significantly reduced by miR-146 overexpression, indicating that
miR-146 participates in suppressing inflammation in OA (91). This was further evidenced by the
finding that miR-146a controls knee-joint homeostasis and
OA-associated algesia by balancing inflammatory responses in
cartilage and synovium (93).
Hence, miR-146a may target inflammation and thus be useful for the
treatment of both cartilage degeneration and pain symptoms in
OA.
Upregulated miRNAs in OA: miR-193b,
miR-199a-3p, miR-181b, miR-22, miR-455 and miR-675
miR-193b and miR-199a-3p expression are upregulated
with age in human cartilage tissue, and these may be involved in
chondrocyte aging by regulating aggrecan, type 2 collagen and Sox9
(81). Therefore, miR-199a-3p and
miR-193b are involved in the senescence of chondrocytes with
currently unknown targets.
miR-181b expression is significantly downregulated
during chondrogenic differentiation of TGF-β3-stimulated limb
mesenchymal cells, but is significantly upregulated in OA
chondrocytes (94). Although a
direct target has not identified, miR-181b attenuation reduced
MMP-13 expression, while inducing type II collagen expression.
Furthermore, anti-miR-181b overexpression significantly reduced
cartilage destruction in mice (94). These data suggest that miR-181b is
a negative regulator of cartilage development, and miR-181b
inhibition may be an effective therapeutic strategy for
cartilage-related diseases.
miR-22 is another miRNA upregulated in human OA
cartilage. miR-22 can directly downregulate BMP7 at the mRNA level
and PPARα at the protein level, which subsequently upregulates
IL-1β and MMP-13 expression in chondrocytes, which is detrimental
to the cartilage ECM. As previously demonstrated, miR-22 inhibition
in osteoarthritic chondrocytes upregulates PPARα (4.9-fold) and
BMP7 (5.8-fold) expression, blocks the inflammatory process through
the inhibition of IL1B (7.6-fold), inhibits catabolic changes, such
as MMP-13 expression (7.9-fold) and activates the cartilage repair
protein aggrecan (3.1-fold), suggesting the therapeutic potential
of miRNA inhibition in OA (75).
miR-455 resides within an intron of Col27a1 that
encodes a cartilage collagen. When human OA cartilage was compared
with cartilage obtained from patients with femoral neck fractures,
miR-140-5p expression increased in OA cartilage. In situ
hybridization revealed miR-455-3p expression in the developing
limbs of chicks and mice and in human OA cartilage (95). ACVR2B, Smad2 and CHRDL1 are direct
targets of miR-455-3p, and these may mediate its functional impact
on TGF signaling. miR-455 is expressed during chondrogenesis and in
adult articular cartilage, where it can regulate TGF signaling,
suppressing the Smad2/3 pathway. Diminished signaling through this
pathway during the aging process and in OA chondrocytes contributes
to cartilage destruction. The increased miR-455 expression in OA
may exacerbate this process and may contribute to the pathogenesis
of the disease (95).
miR-675 is encoded by lncRNA H19, which is highly
expressed as the most abundant cartilage matrix genes Col2a1 and
aggrecan (96). Both miR-675 and
lncRNA H19 are highly expressed in articular cartilage and are
upregulated in OA cartilage. Stress-induced regulation of H19
expression by hypoxic signaling and inflammation suggests that
lncRNA H19 acts as a metabolic correlate in cartilage (97). Though a direct target gene remains
to be identified, the H19-encoded miR-675 may modulate collagen
type II levels via an unrecognized molecule (97).
miR-203
Transient receptor potential vanilloid 4 (TRPV4) is
a member of TRP superfamily of Ca2+-permeable
non-selective cation channels. miR-203 can upregulate nitric oxide
(NO) expression in female rat mandibular condylar chondrocytes by
targeting TRPV4. NO is a critical mediator of the disrupted
processes implicated in OA pathophysiology (98).
5. Conclusion
Undoubtedly, miRNAs, which play a critical role in
one-third of human transcriptional regulation, represent a novel
breakthrough in the field of genetic engineering. Over the past few
years, remarkable progress has been made in the study of miRNAs, as
an increasing number of tissue-specific miRNAs have been identified
in chondrocytes. In conclusion, miRNAs appear to have significant
potential for OA diagnostics and therapeutics and cartilage tissue
engineering. However, their potential will only be realized through
further research to identify potential miRNA target sites and the
associated mechanisms.
Acknowledgements
This study was supported by a grant from the
National Natural Science Foundation of China (no. 81171729), a
scientific research grant from the National Natural Science
Foundation for the Youth of China (Grant No. 81201364), a
scientific research grant for the youth of Shanghai (Grant No.
ZZjdyx 2097), a scientific research grant from the 985 project -
Stem Cell and Regenerative Medicine Centre, an innovative research
grant from the Shanghai Municipal Education Commission (Grant No.
13YZ031) and a fund for Key Disciplines of Shanghai Municipal
Education Commission (J50206).
References
|
1
|
Iannone F and Lapadula G: The
pathophysiology of osteoarthritis. Aging Clin Exp Res. 15:364–372.
2003. View Article : Google Scholar
|
|
2
|
Mortellaro CM: Pathophysiology of
osteoarthritis. Vet Res Commun. 27(Suppl 1): S75–S78. 2003.
View Article : Google Scholar
|
|
3
|
Martel-Pelletier J: Pathophysiology of
osteoarthritis. Osteoarthritis Cartilage. 12(Suppl A): S31–S33.
2004. View Article : Google Scholar
|
|
4
|
Mandelbaum B and Waddell D: Etiology and
pathophysiology of osteoarthritis. Orthopedics. 28(Suppl 2):
s207–s214. 2005.PubMed/NCBI
|
|
5
|
Felson DT, Lawrence RC, Dieppe PA, et al:
Osteoarthritis: new insights. Part 1: the disease and its risk
factors. Ann Intern Med. 133:635–646. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lawrence RC, Felson DT, Helmick CG, et al:
Estimates of the prevalence of arthritis and other rheumatic
conditions in the United States. Part II. Arthritis Rheum.
58:26–35. 2008. View Article : Google Scholar
|
|
7
|
Naumann A, Dennis JE, Awadallah A, et al:
Immunochemical and mechanical characterization of cartilage
subtypes in rabbit. J Histochem Cytochem. 50:1049–1058. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wong M and Carter DR: Articular cartilage
functional histomorphology and mechanobiology: a research
perspective. Bone. 33:1–13. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Burgeson RE, Hebda PA, Morris NP and
Hollister DW: Human cartilage collagens. Comparison of cartilage
collagens with human type V collagen. J Biol Chem. 257:7852–7856.
1982.PubMed/NCBI
|
|
10
|
Eyre D: Collagen of articular cartilage.
Arthritis Res. 4:30–35. 2002. View
Article : Google Scholar
|
|
11
|
Poole AR, Kojima T, Yasuda T, Mwale F,
Kobayashi M and Laverty S: Composition and structure of articular
cartilage: a template for tissue repair. Clin Orthop Relat Res.
391:S26–S33. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Knudson CB and Knudson W: Cartilage
proteoglycans. Semin Cell Dev Biol. 12:69–78. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Cawston TE and Wilson AJ: Understanding
the role of tissue degrading enzymes and their inhibitors in
development and disease. Best Pract Res Clin Rheumatol.
20:983–1002. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Plaas A, Osborn B, Yoshihara Y, et al:
Aggrecanolysis in human osteoarthritis: confocal localization and
biochemical characterization of ADAMTS5-hyaluronan complexes in
articular cartilages. Osteoarthritis Cartilage. 15:719–734. 2007.
View Article : Google Scholar
|
|
15
|
Wu W, Billinghurst RC, Pidoux I, et al:
Sites of collagenase cleavage and denaturation of type II collagen
in aging and osteoarthritic articular cartilage and their
relationship to the distribution of matrix metalloproteinase 1 and
matrix metalloproteinase 13. Arthritis Rheum. 46:2087–2094. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bartel DP: MicroRNAs: genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Cordes KR and Srivastava D: MicroRNA
regulation of cardiovascular development. Circ Res. 104:724–732.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Fabian MR, Sonenberg N and Filipowicz W:
Regulation of mRNA translation and stability by microRNAs. Annu Rev
Biochem. 79:351–379. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Liu N and Olson EN: MicroRNA regulatory
networks in cardiovascular development. Dev Cell. 18:510–525. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang DZ: MicroRNAs in cardiac development
and remodeling. Pediatr Cardiol. 31:357–362. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhao Y and Srivastava D: A developmental
view of microRNA function. Trends Biochem Sci. 32:189–197. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Farh KK, Grimson A, Jan C, et al: The
widespread impact of mammalian MicroRNAs on mRNA repression and
evolution. Science. 310:1817–1821. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Chendrimada TP, Gregory RI, Kumaraswamy E,
et al: TRBP recruits the Dicer complex to Ago2 for microRNA
processing and gene silencing. Nature. 436:740–744. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lee Y, Ahn C, Han J, et al: The nuclear
RNase III Drosha initiates microRNA processing. Nature.
425:415–419. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Gregory RI, Yan KP, Amuthan G, et al: The
Microprocessor complex mediates the genesis of microRNAs. Nature.
432:235–240. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Denli AM, Tops BB, Plasterk RH, Ketting RF
and Hannon GJ: Processing of primary microRNAs by the
Microprocessor complex. Nature. 432:231–235. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Small EM and Olson EN: Pervasive roles of
microRNAs in cardiovascular biology. Nature. 469:336–342. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ghayor C, Chadjichristos C, Herrouin JF,
et al: Sp3 represses the Sp1-mediated transactivation of the human
COL2A1 gene in primary and de-differentiated chondrocytes. J Biol
Chem. 276:36881–36895. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zhang Z, Kang Y, Zhang H, et al:
Expression of microRNAs during chondrogenesis of human
adipose-derived stem cells. Osteoarthritis Cartilage. 20:1638–1646.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Cancedda R, Descalzi Cancedda F and
Castagnola P: Chondrocyte differentiation. Int Rev Cytol.
159:265–358. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Yang B, Guo H, Zhang Y, Chen L, Ying D and
Dong S: MicroRNA-145 regulates chondrogenic differentiation of
mesenchymal stem cells by targeting Sox9. PLoS One. 6:e216792011.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Martinez-Sanchez A, Dudek KA and Murphy
CL: Regulation of human chondrocyte function through direct
inhibition of cartilage master regulator SOX9 by microRNA-145
(miRNA-145). J Biol Chem. 287:916–924. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ning G, Liu X, Dai M, Meng A and Wang Q:
MicroRNA-92a upholds Bmp signaling by targeting noggin3 during
pharyngeal cartilage formation. Dev Cell. 24:283–295. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ohgawara T, Kubota S, Kawaki H, et al:
Regulation of chondrocytic phenotype by micro RNA 18a: involvement
of Ccn2/Ctgf as a major target gene. FEBS Lett. 583:1006–1010.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Soullier S, Jay P, Poulat F, Vanacker JM,
Berta P and Laudet V: Diversification pattern of the HMG and SOX
family members during evolution. J Mol Evol. 48:517–527. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wright E, Hargrave MR, Christiansen J, et
al: The Sry-related gene Sox9 is expressed during chondrogenesis in
mouse embryos. Nat Genet. 9:15–20. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Bi W, Deng JM, Zhang Z, Behringer RR and
de Crombrugghe B: Sox9 is required for cartilage formation. Nat
Genet. 22:85–89. 1999. View
Article : Google Scholar
|
|
39
|
Ikeda T, Kawaguchi H, Kamekura S, et al:
Distinct roles of Sox5, Sox6, and Sox9 in different stages of
chondrogenic differentiation. J Bone Miner Metab. 23:337–340. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Bell DM, Leung KK, Wheatley SC, et al:
SOX9 directly regulates the type-II collagen gene. Nat Genet.
16:174–178. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Zhang P, Jimenez SA and Stokes DG:
Regulation of human COL9A1 gene expression. Activation of the
proximal promoter region by SOX9. J Biol Chem. 278:117–123. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Liu Y, Li H, Tanaka K, Tsumaki N and
Yamada Y: Identification of an enhancer sequence within the first
intron required for cartilage-specific transcription of the
alpha2(XI) collagen gene. J Biol Chem. 275:12712–12718. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Sekiya I, Tsuji K, Koopman P, et al: SOX9
enhances aggrecan gene promoter/enhancer activity and is
up-regulated by retinoic acid in a cartilage-derived cell line,
TC6. J Biol Chem. 275:10738–10744. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Tew SR, Li Y, Pothacharoen P, Tweats LM,
Hawkins RE and Hardingham TE: Retroviral transduction with SOX9
enhances re-expression of the chondrocyte phenotype in passaged
osteoarthritic human articular chondrocytes. Osteoarthritis
Cartilage. 13:80–89. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Cucchiarini M, Thurn T, Weimer A, Kohn D,
Terwilliger EF and Madry H: Restoration of the extracellular matrix
in human osteoarthritic articular cartilage by overexpression of
the transcription factor SOX9. Arthritis Rheum. 56:158–167. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Dai L, Zhang X, Hu X, Zhou C and Ao Y:
Silencing of microRNA-101 prevents IL-1beta-induced extracellular
matrix degradation in chondrocytes. Arthritis Res Ther.
14:R2682012. View
Article : Google Scholar : PubMed/NCBI
|
|
47
|
Xu J, Kang Y, Liao WM and Yu L: MiR-194
regulates chondrogenic differentiation of human adipose-derived
stem cells by targeting Sox5. PLoS One. 7:e318612012. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Parvizi J, Zmistowski B, Berbari EF, et
al: New definition for periprosthetic joint infection: from the
Workgroup of the Musculoskeletal Infection Society. Clin Orthop
Relat Res. 469:2992–2994. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Hatakeyama Y, Nguyen J, Wang X, Nuckolls
GH and Shum L: Smad signaling in mesenchymal and chondroprogenitor
cells. J Bone Joint Surg Am. 85-A(Suppl 3): S13–S18.
2003.PubMed/NCBI
|
|
50
|
Pan Q, Yu Y, Chen Q, et al: Sox9, a key
transcription factor of bone morphogenetic protein-2-induced
chondrogenesis, is activated through BMP pathway and a CCAAT box in
the proximal promoter. J Cell Physiol. 217:228–241. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Denker AE, Nicoll SB and Tuan RS:
Formation of cartilage-like spheroids by micromass cultures of
murine C3H10T1/2 cells upon treatment with transforming growth
factor-beta 1. Differentiation. 59:25–34. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Lin EA, Kong L, Bai XH, Luan Y and Liu CJ:
miR-199a, a bone morphogenic protein 2-responsive MicroRNA,
regulates chondrogenesis via direct targeting to Smad1. J Biol
Chem. 284:11326–11335. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Liang ZJ, Zhuang H, Wang GX, et al:
MiRNA-140 is a negative feedback regulator of MMP-13 in
IL-1beta-stimulated human articular chondrocyte C28/I2 cells.
Inflamm Res. 61:503–509. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Buechli ME, Lamarre J and Koch TG:
MicroRNA-140 expression during chondrogenic differentiation of
equine cord blood-derived mesenchymal stromal cells. Stem Cells
Dev. 22:1288–1296. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Miyaki S, Nakasa T, Otsuki S, et al:
MicroRNA-140 is expressed in differentiated human articular
chondrocytes and modulates interleukin-1 responses. Arthritis
Rheum. 60:2723–2730. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Nicolas FE, Pais H, Schwach F, et al: mRNA
expression profiling reveals conserved and non-conserved miR-140
targets. RNA Biol. 8:607–615. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Pais H, Nicolas FE, Soond SM, et al:
Analyzing mRNA expression identifies Smad3 as a microRNA-140 target
regulated only at protein level. RNA. 16:489–494. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Nakamura Y, Inloes JB, Katagiri T and
Kobayashi T: Chondrocyte-specific microRNA-140 regulates
endochondral bone development and targets Dnpep to modulate bone
morphogenetic protein signaling. Mol Cell Biol. 31:3019–3028. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Song J, Kim D and Jin EJ: MicroRNA-488
suppresses cell migration through modulation of the focal adhesion
activity during chondrogenic differentiation of chick limb
mesenchymal cells. Cell Biol Int. 35:179–185. 2011. View Article : Google Scholar
|
|
60
|
Kim D, Song J, Kim S, Chun CH and Jin EJ:
MicroRNA-34a regulates migration of chondroblast and
IL-1beta-induced degeneration of chondrocytes by targeting EphA5.
Biochem Biophys Res Commun. 415:551–557. 2011. View Article : Google Scholar
|
|
61
|
Abouheif MM, Nakasa T, Shibuya H, Niimoto
T, Kongcharoensombat W and Ochi M: Silencing microRNA-34a inhibits
chondrocyte apoptosis in a rat osteoarthritis model in vitro.
Rheumatology (Oxford). 49:2054–2060. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Kim D, Song J and Jin EJ: MicroRNA-221
regulates chondrogenic differentiation through promoting
proteosomal degradation of slug by targeting Mdm2. J Biol Chem.
285:26900–26907. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Dunn W, DuRaine G and Reddi AH: Profiling
microRNA expression in bovine articular cartilage and implications
for mechanotransduction. Arthritis Rheum. 60:2333–2339. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Magee C, Nurminskaya M, Faverman L, Galera
P and Linsenmayer TF: SP3/SP1 transcription activity regulates
specific expression of collagen type X in hypertrophic
chondrocytes. J Biol Chem. 280:25331–25338. 2005. View Article : Google Scholar
|
|
65
|
Kavurma MM and Khachigian LM: Sp1 inhibits
proliferation and induces apoptosis in vascular smooth muscle cells
by repressing p21WAF1/Cip1 transcription and cyclin
D1-Cdk4-p21WAF1/Cip1 complex formation. J Biol Chem.
278:32537–32543. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Deniaud E, Baguet J, Chalard R, et al:
Overexpression of transcription factor Sp1 leads to gene expression
perturbations and cell cycle inhibition. PLoS One. 4:e70352009.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Yang J, Qin S, Yi C, et al: MiR-140 is
co-expressed with Wwp2-C transcript and activated by Sox9 to target
Sp1 in maintaining the chondrocyte proliferation. FEBS Lett.
585:2992–2997. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Sumiyoshi K, Kubota S, Ohgawara T, et al:
Identification of miR-1 as a micro RNA that supports late-stage
differentiation of growth cartilage cells. Biochem Biophys Res
Commun. 402:286–290. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Guan YJ, Yang X, Wei L and Chen Q:
MiR-365: a mechanosensitive microRNA stimulates chondrocyte
differentiation through targeting histone deacetylase 4. FASEB J.
25:4457–4466. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Vega RB, Matsuda K, Oh J, et al: Histone
deacetylase 4 controls chondrocyte hypertrophy during
skeletogenesis. Cell. 119:555–566. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Tuddenham L, Wheeler G, Ntounia-Fousara S,
et al: The cartilage specific microRNA-140 targets histone
deacetylase 4 in mouse cells. FEBS Lett. 580:4214–4217. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Nicolas FE, Pais H, Schwach F, et al:
Experimental identification of microRNA-140 targets by silencing
and overexpressing miR-140. RNA. 14:2513–2520. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Zhong N, Sun J, Min Z, et al: MicroRNA-337
is associated with chondrogenesis through regulating TGFBR2
expression. Osteoarthritis Cartilage. 20:593–602. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Ham O, Song BW, Lee SY, et al: The role of
microRNA-23b in the differentiation of MSC into chondrocyte by
targeting protein kinase A signaling. Biomaterials. 33:4500–4507.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Iliopoulos D, Malizos KN, Oikonomou P and
Tsezou A: Integrative microRNA and proteomic approaches identify
novel osteoarthritis genes and their collaborative metabolic and
inflammatory networks. PLoS One. 3:e37402008. View Article : Google Scholar
|
|
76
|
Glasson SS, Askew R, Sheppard B, et al:
Deletion of active ADAMTS5 prevents cartilage degradation in a
murine model of osteoarthritis. Nature. 434:644–648. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Stanton H, Rogerson FM, East CJ, et al:
ADAMTS5 is the major aggrecanase in mouse cartilage in vivo and in
vitro. Nature. 434:648–652. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Miyaki S, Sato T, Inoue A, et al:
MicroRNA-140 plays dual roles in both cartilage development and
homeostasis. Genes Dev. 24:1173–1185. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Zhang M, Liu L, Xiao T and Guo W:
Detection of the expression level of miR-140 using realtime
fluorescent quantitative PCR in knee synovial fluid of
osteoarthritis patients. Zhong Nan Da Xue Xue Bao Yi Xue Ban.
37:1210–1214. 2012.(In Chinese).
|
|
80
|
Tardif G, Hum D, Pelletier JP, Duval N and
Martel-Pelletier J: Regulation of the IGFBP-5 and MMP-13 genes by
the microRNAs miR-140 and miR-27a in human osteoarthritic
chondrocytes. BMC Musculoskelet Disord. 10:1482009. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Ukai T, Sato M, Akutsu H, Umezawa A and
Mochida J: MicroRNA-199a-3p, microRNA-193b, and microRNA-320c are
correlated to aging and regulate human cartilage metabolism. J
Orthop Res. 30:1915–1922. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Matsukawa T, Sakai T, Yonezawa T, et al:
MicroRNA-125b regulates the expression of aggrecanase-1 (ADAMTS-4)
in human osteoarthritic chondrocytes. Arthritis Res Ther.
15:R282013. View
Article : Google Scholar : PubMed/NCBI
|
|
83
|
Xu N, Zhang L, Meisgen F, et al:
MicroRNA-125b down-regulates matrix metallopeptidase 13 and
inhibits cutaneous squamous cell carcinoma cell proliferation,
migration, and invasion. J Biol Chem. 287:29899–29908. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Muramatsu F, Kidoya H, Naito H, Sakimoto S
and Takakura N: microRNA-125b inhibits tube formation of blood
vessels through translational suppression of VE-cadherin. Oncogene.
32:414–421. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Little CB, Barai A, Burkhardt D, et al:
Matrix metalloproteinase 13-deficient mice are resistant to
osteoarthritic cartilage erosion but not chondrocyte hypertrophy or
osteophyte development. Arthritis Rheum. 60:3723–3733. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Mapp PI and Walsh DA: Mechanisms and
targets of angiogenesis and nerve growth in osteoarthritis. Nat Rev
Rheumatol. 8:390–398. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Akhtar N, Rasheed Z, Ramamurthy S,
Anbazhagan AN, Voss FR and Haqqi TM: MicroRNA-27b regulates the
expression of matrix metalloproteinase 13 in human osteoarthritis
chondrocytes. Arthritis Rheum. 62:1361–1371. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Akhtar N and Haqqi TM:
MicroRNA-199a* regulates the expression of
cyclooxygenase-2 in human chondrocytes. Ann Rheum Dis.
71:1073–1080. 2012.
|
|
89
|
He L, He X, Lim LP, et al: A microRNA
component of the p53 tumour suppressor network. Nature.
447:1130–1134. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Chang TC, Wentzel EA, Kent OA, et al:
Transactivation of miR-34a by p53 broadly influences gene
expression and promotes apoptosis. Mol Cell. 26:745–752. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Jones SW, Watkins G, Le Good N, et al: The
identification of differentially expressed microRNA in
osteoarthritic tissue that modulate the production of TNF-alpha and
MMP13. Osteoarthritis Cartilage. 17:464–472. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Yamasaki K, Nakasa T, Miyaki S, et al:
Expression of MicroRNA-146a in osteoarthritis cartilage. Arthritis
Rheum. 60:1035–1041. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Li X, Gibson G, Kim JS, et al:
MicroRNA-146a is linked to pain-related pathophysiology of
osteoarthritis. Gene. 480:34–41. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Song J, Lee M, Kim D, Han J, Chun CH and
Jin EJ: MicroRNA-181b regulates articular chondrocytes
differentiation and cartilage integrity. Biochem Biophys Res
Commun. 431:210–214. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Swingler TE, Wheeler G, Carmont V, et al:
The expression and function of microRNAs in chondrogenesis and
osteoarthritis. Arthritis Rheum. 64:1909–1919. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Dudek KA, Lafont JE, Martinez-Sanchez A
and Murphy CL: Type II collagen expression is regulated by
tissue-specific miR-675 in human articular chondrocytes. J Biol
Chem. 285:24381–24387. 2010. View Article : Google Scholar
|
|
97
|
Steck E, Boeuf S, Gabler J, et al:
Regulation of H19 and its encoded microRNA-675 in osteoarthritis
and under anabolic and catabolic in vitro conditions. J Mol Med
(Berl). 90:1185–1195. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Hu F, Zhu W and Wang L: MicroRNA-203
up-regulates nitric oxide expression in temporomandibular joint
chondrocytes via targeting TRPV4. Arch Oral Biol. Nov 16–2012.(Epub
ahead of print). View Article : Google Scholar
|