Introduction
Myelination is the production of the myelin sheath
that surrounds the axons of nerve cells; it occurs through
reciprocal interactions between the glial cells and the axons that
they ensheath (1). Schwann cells
(SCs) in the peripheral nervous system (PNS) wrap their membranes
spirally around long segments of axons. Subsequently, myelinated
fibers with multilayered sheaths are developed, something that
allows the propagation of rapid impulses (2,3).
Demyelination is a hallmark of numerous neurological diseases, such
as Charcot-Marie-Tooth disease, which is a PNS hereditary disorder
affecting 1/2,500 individuals (1,4).
The mechanism of remyelination in neurological diseases is similar
to developmental myelination (2),
therefore it is possible to identify potential therapeutic
treatments for neurological diseases by studying the process of
normal myelination in the PNS.
Several previous studies have investigated the genes
and pathways associated with peripheral myelination focusing on SC
plasticity, polarization, function of the neuregulin (NRG), and
extracellular matrix and cytoskeletal signals (2,3,5).
These reviews revealed that NRG1 served a key role in myelination
by regulating the majority of SC biological processes (6). NRG1 bounded to the epidermal growth
factor receptor and subsequently activated several
secondary-messenger cascades, including the phosphoinositide
3-kinase/phosphatidylinositol 3,4,5 trisphosphate/protein kinase B
(AKT) (7), intracellular calcium
(8) and mitogen-activated protein
kinase signaling pathways (9). In
addition, the proto-oncogene Wnt/β-catenin (10), endocytic (11) and hedgehog signaling pathways
(12) were also reported to be
associated with peripheral myelination. However, further
investigation is required to clarify the mechanisms of peripheral
myelination.
MicroRNAs (miRs) serve a post-transcriptional role
in PNS myelination (13). The
majority of previous studies in this area have focused on the
function of miRs in peripheral remyelination following nerve injury
(14–18). However, the functional roles of
miRs are rarely studied in developmental peripheral myelination.
miR-29a has been identified to prevent the expression of peripheral
myelin protein 22, which is a disease-associated protein in
myelinating SCs (19). miR-318
has been identified as a suppressor of certain myelination
inhibitors, including sex determining region Y-box 2, Jun
proto-oncogene and cyclin D1 (20). Let-7 miR promotes the expression
of positive transcriptional factor early growth response protein 2
through suppression of the myelination inhibitor neurogenic locus
notch homolog protein 1 (21).
These findings reveal only a small part of the mechanisms of miR in
PNS myelination.
Kangas et al (22) presented the genome-wide RNA
expression microarray data at different stages of PNS myelination
using a dorsal root ganglion (DRG) explant culture. Amongst the
differentially expressed genes (DEGs), enriched Kyoto Encyclopedia
of Genes and Genomes (KEGG) pathways and Gene Ontology (GO) terms
associated with PNS differentiation were identified. Several
previous studies have focused on the potential role of miRs in
peripheral myelination (23,24). However, studies on the
interactions between DEGs and miRs based on genome expression
profiles in regards to peripheral myelination are rare. In the
present study, the microarray dataset GSE60345 (21) was downloaded, and DEGs were
analyzed and subjected to KEGG pathway and GO term enrichment
analyses. Several miRs were identified as being associated with
peripheral myelination based on the miR-target gene regulatory
network analysis.
Materials and methods
Data acquisition
The GSE60345 microarray dataset (21) gene expression profiles were
obtained from the National Center for Biotechnology Information
Gene Expression Omnibus database (ncbi.nlm.nih.gov/geo/) (25). The microarray dataset GSE60345 was
originally obtained using a SurePrint G3 Mouse GE 8×60K Microarray
kit (cat. no. 028005; Agilent Technologies, Inc., Santa Clara, CA,
USA). The organism studied was Mus musculus. A total of 18
samples of DRG explant cultures were collected at 0 (n=6), 10 (n=6)
and 22 days (n=6) following switching to myelination medium from
embryonic day 13.5 C57BL/6J mouse embryos. The different
time-points that the samples were collected at represented the
different stages of the myelination process; i) premyelination (0
days), ii) early myelination (10 days); and iii) advanced
myelination (22 days).
Data preprocessing and DEG statistical
analysis
The raw data was normalized using the limma package
(version 3.10.3) of R software (26). The gene expression matrix was
divided into two groups of 10 day vs. 0 day and 22 day vs. 0 day.
The P-value of each gene was calculated using the non-paired t-test
in the limma package software and the P-value was adjusted using
the Benjamini-Hochberg method. According to the cut-off value of
adjacent P<0.01 and log2 fold change ≥2, the DEGs of
each group were identified. The upregulated and downregulated genes
were obtained by pooling the DEGs of the two groups.
GO term and KEGG pathway enrichment
analyses
The upregulated and downregulated genes were
subjected to GO term (27) and
KEGG pathway enrichment analysis. The analyses were conducted with
Fisher's exact test using the 'mRNA enrichment' module in the
Multifaceted Analysis Tool for Human Transcriptome (biocloudservice.com). The cut-off value was set as
P<0.05.
Protein-protein interactions (PPI)
network
The interaction of proteins encoded by the DEGs was
predicted using the Search Tool for the Retrieval of Interacting
Genes version 10.0 (28). The PPI
score (medium confidence) was set at 0.4. Cytoscape version 3.2.0
software (29) was used to
visualize the PPI network. The modules consisted of the 200 nodes
with the highest degree values.
miR-target gene regulatory network
analysis
The validated miRs that were identified to regulate
DEGs from the two groups were downloaded from a validated gene-miR
interaction information retrieval system, miRWalk version 2.0
(30). Cytoscape software was
used to construct the miR-target gene regulatory network.
Results
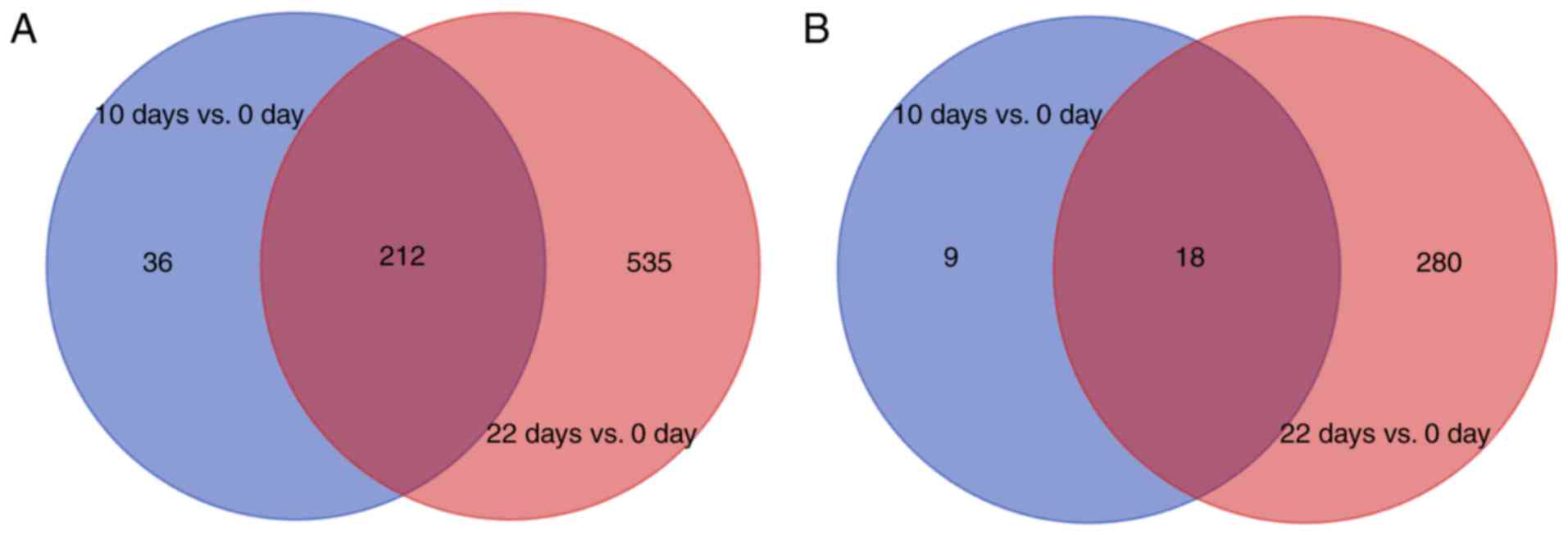
Identification of DEGs
The gene expression data was normalized following
preprocessing. A total of 276 DEGs, including 248 upregulated and
28 downregulated genes were obtained in the 10 day vs. 0 day group
(Fig. 1). By contrast, 1,045 DEGs
were identified in the 22 day vs. 0 day group comprised of 747
upregulated and 298 downregulated genes (Fig. 1). There were a higher number of
upregulated genes compared with downregulated genes, and the 22 day
vs. 0 day group contained more genes than the 10 day vs. 0 day
group. The Venn diagram revealed that the overlapping upregulated
and downregulated genes between the two groups were 212 and 18,
respectively (Fig. 1). A total of
783 upregulated genes and 307 downregulated genes were identified
by pooling the DEGs of the two groups.
GO term and KEGG pathway enrichment
analyses
A total of 26 pathways were significantly enriched
in the upregulated DEGs and 5 pathways were significantly enriched
in the downregulated DEGs. The top 10 enriched pathways according
to their P-value are listed in Table
I. The complement and coagulation cascades, cytokine-cytokine
receptor interaction, cell adhesion molecules, natural killer (NK)
cell mediated cytotoxicity and B cell receptor (BCR) signaling
pathways were significantly enriched by the upregulated DEGs.
Pathways significantly enriched by the downregulated DEGs included
the cell cycle, oocyte meiosis and the cellular tumor antigen p53
(p53) signaling pathway.
 | Table IKyoto Encyclopedia of Genes and
Genomes pathways that the DEGs were significantly enriched in. |
Table I
Kyoto Encyclopedia of Genes and
Genomes pathways that the DEGs were significantly enriched in.
A, Upregulated
genes
|
|---|
| Pathway ID no. | Pathway name | No. of DEGs
enriched in pathway | P-value |
|---|
| mmu04610 | Complement and
coagulation cascades | 31 |
4.43×10−22 |
| mmu04060 | Cytokine-cytokine
receptor interaction | 32 |
5.84×10−8 |
| mmu05330 | Allograft
rejection | 15 |
1.24×10−7 |
| mmu05332 | Graft-vs.-host
disease | 15 |
1.24×10−7 |
| mmu04940 | Type I diabetes
mellitus | 15 |
3.76×10−7 |
| mmu04662 | B cell receptor
signaling pathway | 16 |
1.47×10−6 |
| mmu04514 | Cell adhesion
molecules | 22 |
2.92×10−6 |
| mmu04650 | Natural killer cell
mediated cytotoxicity | 19 |
4.96×10−6 |
| mmu05322 | Systemic lupus
erythematosus | 17 |
8.62×10−6 |
| mmu05320 | Autoimmune thyroid
disease | 14 |
1.16×10−5 |
B, Downregulated
genes
|
|---|
| Pathway ID no. | Pathway name | No. of DEGs
enriched in pathway | P-value |
|---|
| mmu04110 | Cell cycle | 13 |
2.73×10−9 |
| mmu04114 | Oocyte meiosis | 11 |
1.38×10−7 |
| mmu04914 |
Progesterone-mediated oocyte
maturation | 8 |
1.77×10−5 |
| mmu04115 | p53 signaling
pathway | 5 |
4.68×10−3 |
| mmu04020 | Calcium signaling
pathway | 6 |
4.02×10−2 |
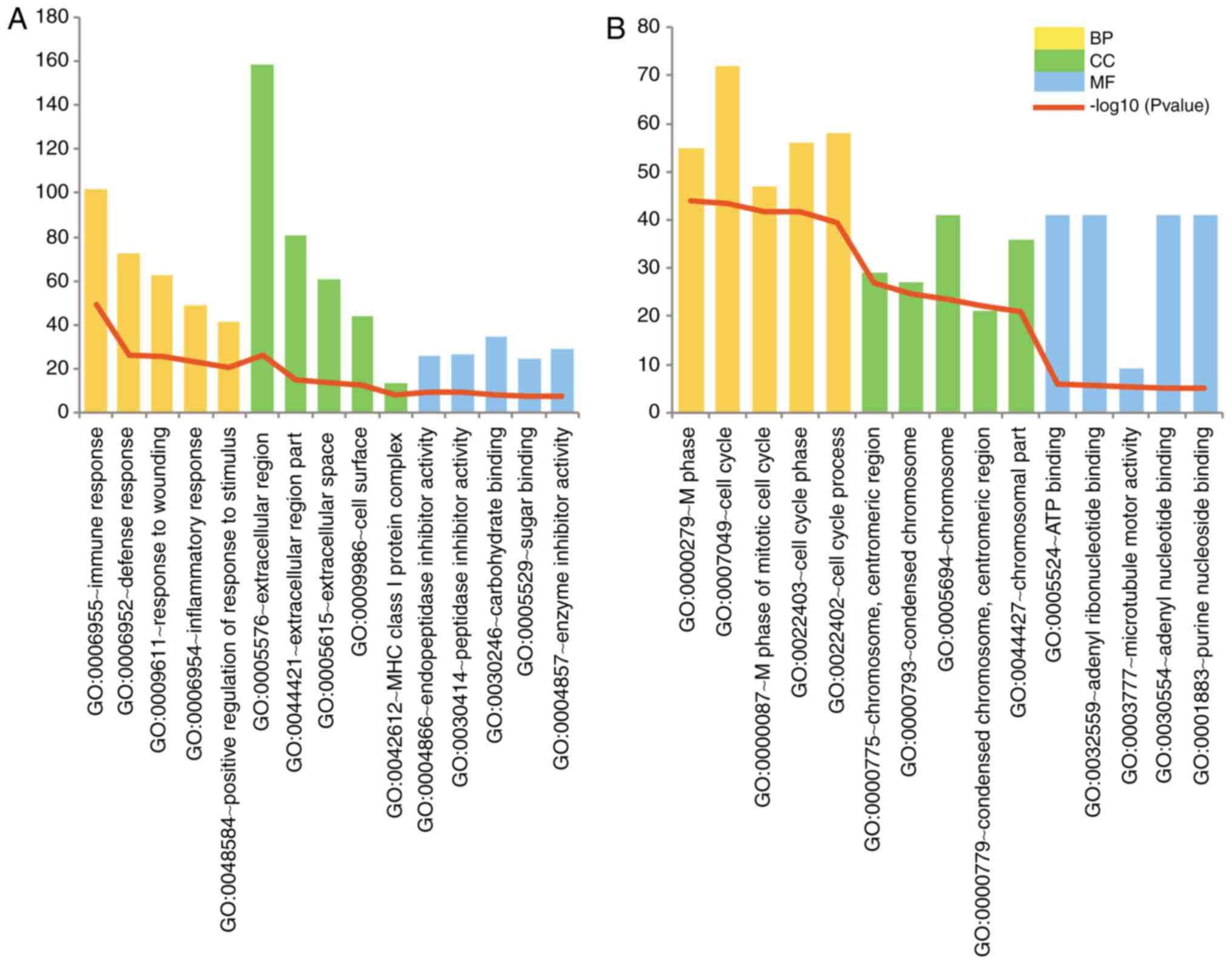
The top 5 GO terms in the biological process,
molecular function and cellular component categories according to
their P-values are illustrated in Fig. 2. The upregulated DEGs were mainly
associated with immune response processes and extracellular region
components. The cell cycle process was significantly enriched in
the downregulated DEGs.
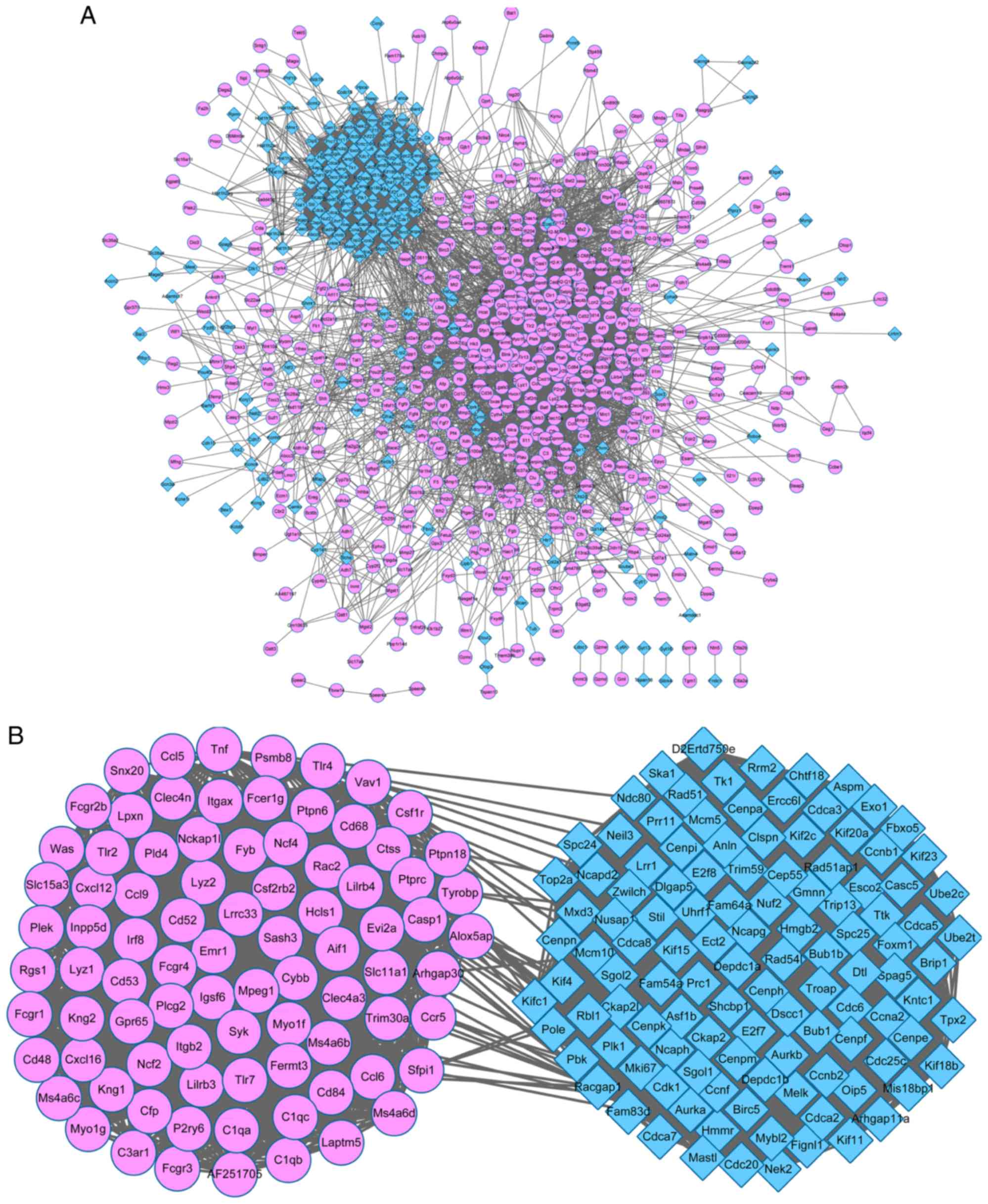
PPI network
A PPI network with 812 nodes and 13,141 edges was
constructed using all of the DEGs (Fig. 3A). The network revealed that the
majority of the nodes gathered together with a complex
connectivity, indicating that the proteins were closely associated
with each other. A PPI sub-network was subsequently constructed
from the 200 DEGs with the highest PPI degree value (Fig. 3B). The upregulated and
downregulated DEGs were gathered together. The 200 DEGs with the
highest PPI degree values were subsequently subjected to KEGG
pathway analysis. A total of 28 pathways were significantly
enriched in the upregulated DEGs and 5 pathways were significantly
enriched in the downregulated DEGs. Table II lists the top 10 pathways
enriched by the DEGs according to their P-value. The leishmaniasis,
osteoclast differentiation, NK cell mediated cytotoxicity and BCR
signaling pathways were significantly enriched by the upregulated
DEGs, including the Fc γ receptor (FCGR), ras-related C3 botulinum
toxin substrate 2 (RAC2) and
1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase γ-2
(PLCG2). The downregulated DEGs were significantly enriched in the
cell cycle, oocyte meiosis and the p53 signaling pathway.
 | Table IITop 10 significantly enriched Kyoto
Encyclopedia of Genes and Genomes pathways of the top 200 DEGs. |
Table II
Top 10 significantly enriched Kyoto
Encyclopedia of Genes and Genomes pathways of the top 200 DEGs.
A, Upregulated
genes
|
|---|
| Pathway ID no. | Pathway name | No. of DEGs
enriched in pathway | P-value | Genes |
|---|
| mmu05140 | Leishmaniasis | 10 |
3.44×10−10 | PTPN6, TNF, NCF2,
NCF4, FCGR4, TLR2, TLR4, ITGB2, FCGR1, FCGR3 |
| mmu04380 | Osteoclast
differentiation | 12 |
6.01×10−10 | CYBB, TNF, FCGR2B,
NCF2, NCF4, PLCG2, FCGR4, FCGR1, CSF1R, FCGR3, SYK, TYROBP |
| mmu05150 | Staphylococcus
aureus infection | 9 |
1.23×10−9 | C1QA, C3AR1, C1QB,
FCGR2B, FCGR4, ITGB2, FCGR1, C1QC, FCGR3 |
| mmu04650 | Natural killer cell
mediated cytotoxicity | 11 |
1.68×10−9 | CD48, PTPN6, TNF,
RAC2, PLCG2, FCGR4, FCER1G, ITGB2, VAV1, SYK, TYROBP |
| mmu05152 | Tuberculosis | 12 |
2.10×10−8 | TNF, ITGAX, FCGR2B,
FCGR4, TLR2, FCER1G, TLR4, ITGB2, CTSS, FCGR1, FCGR3, SYK |
| mmu04666 | Fc γ R-mediated
phagocytosis | 9 |
8.34×10−8 | PTPRC, FCGR2B,
RAC2, PLCG2, INPP5D, FCGR1, VAV1, WAS, SYK |
| mmu04145 | Phagosome | 11 |
2.20×10−7 | CYBB, FCGR2B, NCF2,
NCF4, FCGR4, TLR2, TLR4, ITGB2, CTSS, FCGR1, FCGR3 |
| mmu05133 | Pertussis | 8 |
6.04×10−7 | C1QA, C1QB, TNF,
IRF8, TLR4, ITGB2, CASP1, C1QC |
| mmu04664 | Fc epsilon RI
signaling pathway | 7 |
6.19×10−6 | TNF, RAC2, PLCG2,
FCER1G, INPP5D, VAV1, SYK |
| mmu04662 | B cell receptor
signaling pathway | 7 |
7.34×10−6 | PTPN6, FCGR2B,
RAC2, PLCG2, INPP5D, VAV1, SYK |
B, Downregulated
genes
|
|---|
| Pathway ID no. | Pathway name | No. of DEGs
enriched in pathway | P-value | Genes |
|---|
| mmu04110 | Cell cycle | 13 |
1.16×10−14 | CDK1, CDC6, RBL1,
TTK, CDC20, CDC25C, MCM5, CCNB1, CCNB2, PLK1, BUB1, BUB1B,
CCNA2 |
| mmu04114 | Oocyte meiosis | 8 |
1.53×10−7 | CDK1, PLK1, SGOL1,
BUB1, FBXO5, CDC20, AURKA, CDC25C |
| mmu04914 |
Progesterone-mediated oocyte
maturation | 7 |
8.23×10−7 | CCNB1, CDK1, CCNB2,
PLK1, BUB1, CDC25C, CCNA2 |
| mmu04115 | p53 signaling
pathway | 4 |
2.15×10−3 | CCNB1, CDK1, CCNB2,
RRM2 |
| mmu03460 | Fanconi anemia
pathway | 3 |
1.65×10−2 | BRIP1, UBE2T,
RAD51 |
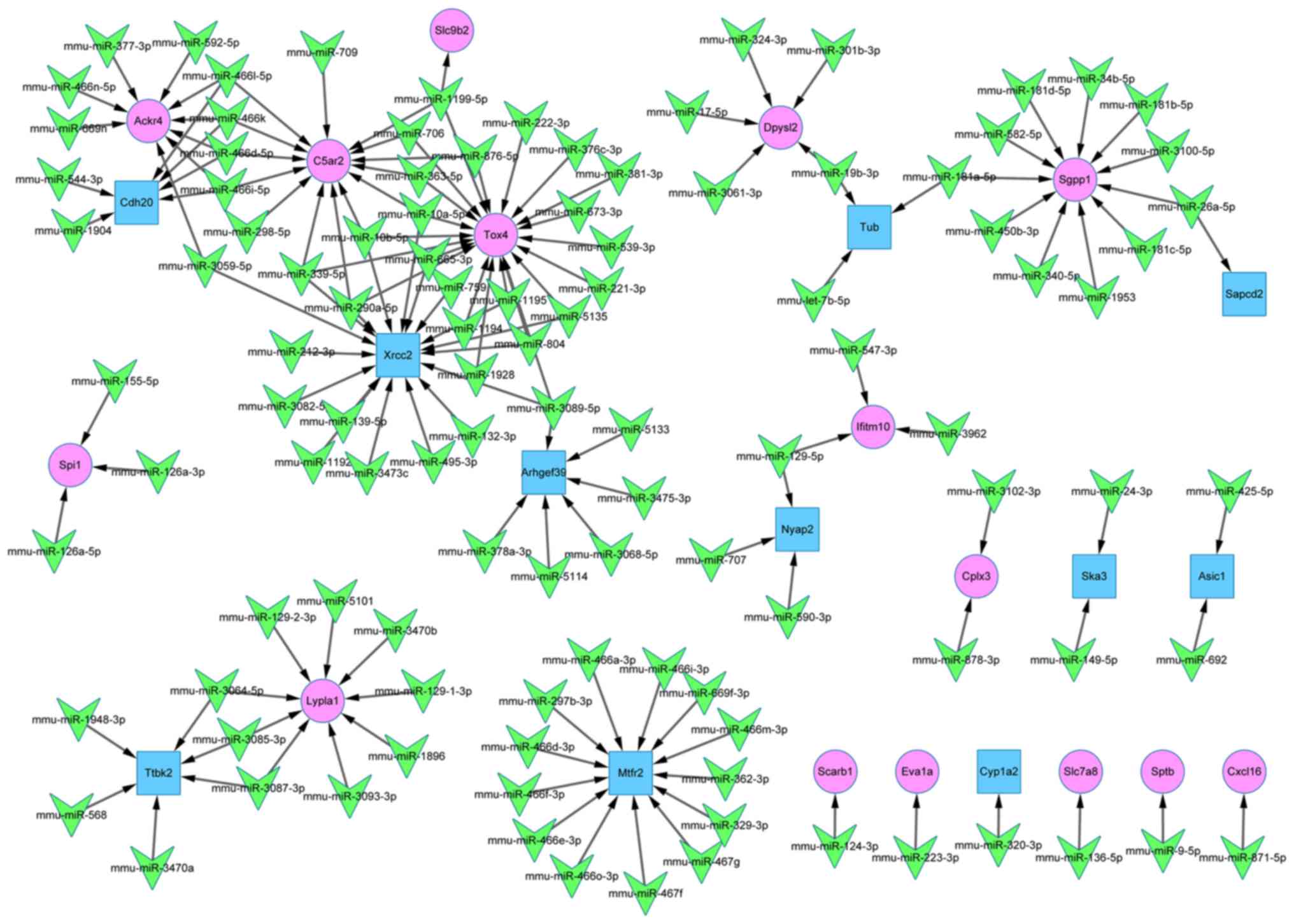
miR-target gene regulatory network
The miR-target gene regulatory network comprised of
135 nodes and 144 edges, as shown in Fig. 4. The association between 109 miRs
and 26 DEGs (15 upregulated and 11 downregulated) was clearly
illustrated. According to their connectivity degree value, the top
10 genes and miRs are listed in Table III. miR-10b-5p, miR-10a-5p and
miR-339-5p were identified as having a high connectivity degree
value. The validated target genes of these miRs were TOX high
mobility group box family member 4 (Tox4), DNA repair protein XRCC2
(Xrcc2) and C5a anaphylatoxin chemotactic receptor C5a2
(C5ar2).
 | Table IIITop 10 miRs and DEGs according to
their degree value. |
Table III
Top 10 miRs and DEGs according to
their degree value.
A, DEGs
|
|---|
| Symbol | Regulation | Degree value |
|---|
| Tox4 | Up | 21 |
| Xrcc2 | Down | 18 |
| C5ar2 | Up | 14 |
| Mtfr2 | Down | 13 |
| Sgpp1 | Up | 11 |
| Ackr4 | Up | 9 |
| Lypla1 | Up | 9 |
| Ttbk2 | Down | 6 |
| Cdh20 | Down | 6 |
| Arhgef39 | Down | 6 |
B, miRs
|
|---|
| Symbol | Regulation | Degree value |
|---|
| mmu-miR-290-5p | – | 3 |
| mmu-miR-10a-5p | – | 3 |
| mmu-miR-339-5p | – | 3 |
| mmu-miR-10b-5p | – | 3 |
|
mmu-miR-4661-5p | – | 3 |
|
mmu-miR-466i-5p | – | 3 |
| mmu-miR-466k | – | 3 |
|
mmu-miR-466d-5p | – | 3 |
|
mmu-miR-1199-5p | – | 3 |
|
mmu-miR-3089-5p | – | 3 |
Discussion
Multiple peripheral nerve diseases have been
revealed to be closely associated with dysregulated myelination
(31). However, the mechanism of
dysregulated myelination has not been fully identified. In the
present study, an integrated bioinformatic analysis was performed
on the differential gene expression patterns throughout different
stages of myelination. A total of 783 upregulated and 307
downregulated genes were identified throughout the different stages
of peripheral myelination. The NK cell mediated cytotoxicity and
BCR signaling pathways were significantly enriched by the
upregulated DEGs, including FCGR, RAC2 and PLCG2. Pathways
significantly enriched by the downregulated DEGs included oocyte
meiosis, the cell cycle and the p53 signaling pathway. In addition,
miR-339-5p, miR-10a-5p and miR-10b-5p demonstrated a high degree
value in targeting Tox4, Xrcc2, C5ar2.
The NK cell mediated cytotoxicity and BCR signaling
pathways were significantly enriched in the upregulated DEGs among
the top 200 DEGs. NK cells are an essential type of cytotoxic
lymphocyte in the innate immune system, which have been revealed to
be associated with myelin loss (32). NK cells also serve a regulatory
role in multiple sclerosis, a disease characterized by chronic
demyelination (33). BCRs are
also a crucial component of the immune system and the BCR signaling
pathway has been implicated in the activation of nuclear factor
(NF)-κB, which serves an indispensable role in peripheral
myelination (34). The present
study revealed that these two pathways were significantly enriched
in the upregulated DEGs. The upregulation of genes in the NK cell
mediated cytotoxicity and the BCR signaling pathways suggested that
an improved immune response was closely associated with peripheral
myelination. In addition, several genes overlapped between these
two pathways, including FCGR, RAC2 and PLCG2. FCGR has previously
been identified in SCs, which may aid in the regulation of the
immune response (35). A recent
study reported that FCGR had a potential role in axonal injury
through the mediation of inflammation (36). RAC2 has been previously identified
as stimulating AKT activation in primary mast cells (37). AKT silencing was previously
reported to be associated with hypomyelination (38). The expression of PLCG2 was
revealed to be sensitive to ciliary neurotrophic factor (39), a cytokine which may promote
peripheral nerve regeneration (40). Consequently, the NK cell mediated
cytotoxicity and BCR signaling pathways may be associated with PNS
myelination through FCGR, RAC2 and PLCG2.
KEGG pathways significantly enriched by
downregulated DEGs among the top 200 DEGs were identified, such as
oocyte meiosis, the p53 signaling pathway and the cell cycle.
Oocyte meiosis is linked to the cell cycle and the p53/p21 axis is
associated with the cell cycle by regulating G1/S
transition (41). The cell cycle
is indispensable for glial cell proliferation, not only in PNS
myelination but in cell dedifferentiation in response to nerve
injury (42). The results of the
present study demonstrated that these three pathways were
downregulated, suggesting that glial cells were prone to
differentiation instead of proliferation following the initiation
of PNS myelination.
The miR-target gene regulatory network revealed that
three miRs (miR-339-5p, miR-10a-5p and miR-10b-5p) possessed a
relatively high degree value. miR-339-5p has been identified to
suppress the protein expression of β-secretase 1, which is a
positive regulator of myelination (43,44). Additionally, miR-339-5p was
reported to be associated with the NF-κB (45) and p53 signaling pathways (46) which are crucial to peripheral
myelination. miR-10a-5p and miR-10b-5p have been previously
reported to regulate the gene encoding brain-derived neurotrophic
factor, which may produce a long-term increase in PNS myelin
formation (47–49). In the present study, miR-339-5p,
miR-10a-5p and miR-10b-5p were identified to regulate Tox4, Xrcc2
and C5ar2, respectively. Tox4, also known as Lcp1, has previously
been revealed to bind with calcium and has been implicated in PNS
myelination (49). Xrcc2 encodes
a member of the RecA/DNA repair protein RAD51 homolog 1-(RAD51L1)
related protein family, in which hsRec2/Rad51L1 is included.
HsRec2/Rad51L1 may phosphorylate myelin basic protein, p53 and
cyclin E (50). C5ar2 encodes C5a
anaphylatoxin chemotactic receptor C5a2, which is also known as G
protein-coupled receptor 77 (GPR77). The function of GPR77 in
peripheral myelination is unclear, whereas another G
protein-coupled receptor, GPR126 (encoded by Adgrg6), has been
identified to be required for the initiation of SC myelination
(51). This suggests that C5ar2
may serve a similar role to Adgrg6. These findings suggest that
miR-339-5p, miR-10a-5p and miR-10b-5p may be associated with
peripheral myelination by targeting Tox4, Xrcc2, and C5ar2.
In conclusion, NK cell mediated cytotoxicity and the
BCR signaling pathway may be associated with peripheral myelination
by regulating FCGR, RAC2 and PLCG2. The indicated downregulation of
oocyte meiosis, the cell cycle and the p53 signaling pathway
suggests that SC proliferation decreases following the initiation
of PNS myelination. miR-339-5p, miR-10a-5p and miR-10b-5p serve a
potential role in the myelination of the PNS by regulating Tox4,
Xrcc2 and C5ar2. The results of the present study may provide
information to clarify the mechanisms of PNS myelination; however,
experimental validation is required to confirm these results.
Acknowledgments
The present study was supported by the key project
of Jinhua Municipal Science and Technology Bureau (grant no.
2016-3-008).
References
|
1
|
Krajewski KM, Lewis RA, Fuerst DR,
Turansky C, Hinderer SR, Garbern J, Kamholz J and Shy ME:
Neurological dysfunction and axonal degeneration in
Charcot-Marie-Tooth disease type 1A. Brain. 6:1516–1527. 2000.
View Article : Google Scholar
|
|
2
|
Pereira JA, Lebrun-Julien F and Suter U:
Molecular mechanisms regulating myelination in the peripheral
nervous system. Trends Neurosci. 35:123–134. 2012. View Article : Google Scholar
|
|
3
|
Nave KA: Myelination and support of axonal
integrity by glia. Nature. 468:244–252. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ritter ST, Jense RJ and Davies JM:
Subarachnoid and peripheral nerve block in a patient with
Charcot-Marie-Tooth disease. Open J Anesthesiol. 3:44–47. 2013.
View Article : Google Scholar
|
|
5
|
Nave KA and Werner HB: Myelination of the
nervous system: Mechanisms and functions. Annu Rev Cell Dev Biol.
30:503–533. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Birchmeier C and Nave KA: Neuregulin-1, a
key axonal signal that drives Schwann cell growth and
differentiation. Glia. 56:1491–1497. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Taveggia C, Zanazzi G, Petrylak A, Yano H,
Rosenbluth J, Einheber S, Xu X, Esper RM, Loeb JA, Shrager P, et
al: Neuregulin-1 Type III determines the ensheathment fate of
axons. Neuron. 47:681–694. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kao SC, Wu H, Xie J, Chang CP, Ranish JA,
Graef IA and Crabtree GR: Calcineurin/NFAT signaling is required
for neuregulin-regulated schwann cell differentiation. Science.
323:651–654. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ye H, Jin YK, Dupree J, Tewari A,
Melendez-Vasquez C, Svaren J and Casaccia P: Yy1 as a molecular
link between neuregulin and transcriptional modulation of
peripheral myelination. Nat Neurosci. 13:1472–1480. 2010.
View Article : Google Scholar
|
|
10
|
Tawk M, Makoukji J, Belle M, Fonte C,
Trousson A, Hawkins T, Li H, Ghandour S, Schumacher M and Massaad
C: Wnt/beta-catenin signaling is an essential and direct driver of
myelin gene expression and myelinogenesis. J Neurosci.
31:3729–3742. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Stendel C, Roos A, Kleine H, Arnaud E,
Ozçelik M, Sidiropoulos PN, Zenker J, Schüpfer F, Lehmann U, Sobota
RM, et al: SH3TC2, a protein mutant in Charcot-Marie-Tooth
neuropathy, links peripheral nerve myelination to endosomal
recycling. Brain. 133:2462–2474. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yoshimura K and Takeda S: Hedgehog
signaling regulates myelination in the peripheral nervous system
through primary cilia. Differentiation. 83:S78–S85. 2012.
View Article : Google Scholar
|
|
13
|
Obernosterer G, Leuschner PJ, Alenius M
and Martinez J: Post-transcriptional regulation of microRNA
expression. RNA. 12:1161–1167. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wu D and Murashov AK: Molecular mechanisms
of peripheral nerve regeneration: Emerging roles of microRNAs.
Front Physiol. 4:552013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yi S, Yuan Y, Chen Q, Wang X, Gong L, Liu
J, Gu X and Li S: Regulation of Schwann cell proliferation and
migration by miR-1 targeting brain-derived neurotrophic factor
after peripheral nerve injury. Sci Rep. 6:291212016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Adilakshmi T, Sudol I and Tapinos N:
Combinatorial action of miRNAs regulates transcriptional and
post-transcriptional gene silencing following in vivo PNS injury.
PLoS One. 7:e396742012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yu B, Zhou S, Wang Y, Qian T, Ding G, Ding
F and Gu X: miR-221 and miR-222 promote Schwann cell proliferation
and migration by targeting LASS2 after sciatic nerve injury. J Cell
Sci. 125:2675–2683. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhou S, Gao R, Hu W, Qian T, Wang N, Ding
G, Ding F, Yu B and Gu X: MiR-9 inhibits Schwann cell migration by
targeting Cthrc1 following sciatic nerve injury. J Cell Sci.
127:967–976. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Verrier JD, Lau P, Hudson L, Murashov AK,
Renne R and Notterpek L: Peripheral myelin protein 22 is regulated
post-transcriptionally by miRNA-29a. Glia. 57:1265–1279. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yun B, Anderegg A, Menichella D, Wrabetz
L, Feltri ML and Awatramani R: MicroRNA-deficient Schwann cells
display congenital hypomyelination. J Neurosci. 30:7722–7728. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Gökbuget D, Pereira JA, Bachofner S,
Marchais A, Ciaudo C, Stoffel M, Schulte JH and Suter U: The
Lin28/let-7 axis is critical for myelination in the peripheral
nervous system. Nat Commun. 6:85842015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kangas SM, Ohlmeier S, Sormunen R,
Jouhilahti EM, Peltonen S, Peltonen J and Heape AM: An approach to
comprehensive genome and proteome expression analyses in Schwann
cells and neurons during peripheral nerve myelin formation. J
Neurochem. 138:830–844. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Verrier JD, Lau P, Hudson L, Murashov AK,
Renne R and Notterpek L: Peripheral myelin protein 22 is regulated
post-transcriptionally by miRNA-29a. Glia. 57:1265–1279. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Dugas JC, Cuellar TL, Scholze A, Ason B,
Ibrahim A, Emery B, Zamanian JL, Foo LC, Mcmanus MT and Barres BA:
Dicer1 and miR-219 Are required for normal oligodendrocyte
differentiation and myelination. Neuron. 65:597–611. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Barrett T, Suzek TO, Troup DB, Wilhite SE,
Ngau WC, Ledoux P, Rudnev D, Lash AE, Fujibuchi W and Edgar R: NCBI
GEO: Mining millions of expression profiles-database and tools.
Nucleic Acids Res. 33:D562–D566. 2005. View Article : Google Scholar
|
|
26
|
Smyth GK: Limma: Linear models for
microarray data. bioinformatics and computational biology solutions
using R and bioconductor. Gentleman R, Carey V, Dudoit S, Irizarry
R and Huber W: Springer; New York: pp. 397–420. 2005, View Article : Google Scholar
|
|
27
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: Gene ontology: Tool for the unification of biology. The Gene
Ontology Consortium. Nat Genet. 25:25–29. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Szklarczyk D, Franceschini A, Kuhn M,
Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork
P, et al: The STRING database in 2011: Functional interaction
networks of proteins, globally integrated and scored. Nucleic Acids
Res. 39:D561–D568. 2011. View Article : Google Scholar :
|
|
29
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Dweep H and Gretz N: miRWalk2.0: A
comprehensive atlas of microRNA-target interactions. Nat Methods.
12:6972015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Saporta MA and Shy ME: Inherited
peripheral neuropathies. Continuum. 17:294–315. 2013.
|
|
32
|
Ratelade J, Zhang H, Saadoun S, Bennett
JL, Papadopoulos MC and Verkman AS: Neuromyelitis optica IgG and
natural killer cells produce NMO lesions in mice without myelin
loss. Acta Neuropathol. 123:861–872. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Takahashi K, Aranami T, Endoh M, Miyake S
and Yamamura T: The regulatory role of natural killer cells in
multiple sclerosis. Brain. 127:1917–1927. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Suzuki H, Matsuda S, Terauchi Y, Fujiwara
M, Ohteki T, Asano T, Behrens TW, Kouro T, Takatsu K, Kadowaki T
and Koyasu S: PI3K and Btk differentially regulate B cell antigen
receptor-mediated signal transduction. Nat Immunol. 4:280–286.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Vedeler CA and Fitzpatrick-Kløve L:
Receptors for immunoglobulin G demonstrated on human peripheral
nerve fibres by electron microscopy. Neurosci Lett. 115:167–170.
1990. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kaida K: Antibodies to glycoconjugates in
autoimmune neuropathies. Clin Experimental Neuroimmunol. 6:387–394.
2015. View Article : Google Scholar
|
|
37
|
Yang FC, Kapur R, King AJ, Tao W, Kim C,
Borneo J, Breese R, Marshall M, Dinauer MC and Williams DA: Rac2
stimulates Akt activation affecting BAD/Bcl-XL expression while
mediating survival and actin function in primary mast cells.
Immunity. 12:557–568. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Cotter L, Ozçelik M, Jacob C, Pereira JA,
Locher V, Baumann R, Relvas JB, Suter U and Tricaud N: Dlg1-PTEN
interaction regulates myelin thickness to prevent damaging
peripheral nerve overmyelination. Science. 328:1415–1418. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Levaillant CJ, Sharma A, Muhling J,
Wheeler LP, Cozens GS, Hellström M, Rodger J and Harvey AR:
Significant changes in endogenous retinal gene expression assessed
1 year after a single intraocular injection of AAV-CNTF or
AAV-BDNF. Mol Ther Methods Clin Dev. 3:160782016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wan L, Xia R and Ding W: Short-term
low-frequency electrical stimulation enhanced remyelination of
injured peripheral nerves by inducing the promyelination effect of
brain-derived neurotrophic factor on Schwann cell polarization. J
Neurosci Res. 88:2578–2587. 2010.PubMed/NCBI
|
|
41
|
Mikule K, Delaval B, Kaldis P, Jurcyzk A,
Hergert P and Doxsey S: Loss of centrosome integrity induces
p38-p53-p21-dependent G1-S arrest. Nat Cell Biol. 9:160–170. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Jessen KR and Mirsky R: The origin and
development of glial cells in peripheral nerves. Nat Rev Neurosci.
6:671–682. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
43
|
Long JM, Ray B and Lahiri DK:
MicroRNA-339-5p down-regulates protein expression of β-site amyloid
precursor protein-cleaving enzyme 1 (BACE1) in human primary brain
cultures and is reduced in brain tissue specimens of Alzheimer
disease subjects. J Biol Chem. 289:5184–5198. 2014. View Article : Google Scholar
|
|
44
|
Willem M, Garratt AN, Novak B, Citron M,
Kaufmann S, Rittger A, DeStrooper B, Saftig P, Birchmeier C and
Haass C: Control of peripheral nerve myelination by the
beta-secretase BACE1. Science. 314:664–666. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang Y, Wei G, Di Z and Zhao Q:
miR-339-5p inhibits alcohol-induced brain inflammation through
regulating NF-κB pathway. Biochem Biophys Res Commun. 452:450–456.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Jansson MD, Damas ND, Lees M, Jacobsen A
and Lund AH: miR-339-5p regulates the p53 tumor-suppressor pathway
by targeting MDM2. Oncogene. 34:1908–1918. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Müller S: In silico analysis of regulatory
networks underlines the role of miR-10b-5p and its target BDNF in
huntington's disease. Transl Neurodegener. 3:172014. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Prins SA, Przybycien-Szymanska MM, Rao YS
and Pak TR: Long-term effects of peripubertal binge EtOH exposure
on hippocampal microRNA expression in the rat. PLoS One.
9:e831662014. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Tolwani RJ, Cosgaya JM, Varma S, Jacob R,
Kuo LE and Shooter EM: BDNF overexpression produces a long-term
increase in myelin formation in the peripheral nervous system. J
Neurosci Res. 77:662–669. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Havre PA, Rice M, Ramos R and Kmiec EB:
HsRec2/Rad51L1, a protein influencing cell cycle progression, has
protein kinase activity. Exp Cell Res. 254:33–44. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Monk KR, Naylor SG, Glenn TD, Mercurio S,
Perlin JR, Dominguez C, Moens CB and Talbot WS: A G protein-coupled
receptor is essential for schwann cells to initiate myelination.
Science. 325:1402–1405. 2009. View Article : Google Scholar : PubMed/NCBI
|