Introduction
The prevalence of rheumatoid arthritis (RA), a
systemic autoimmune disease that induces systemic complications and
progressive disability, is rising steadily (1,2).
Generally, the prevalence of RA worldwide is ~1% of the total adult
population (1,2). The characteristics of RA include
progressive joint destruction and disability, and it is associated
with high prevalence rates of comorbidities including
cardiovascular diseases and other extra-articular and systemic
diseases (3). Compared with the
normal population, a decreased life expectancy in patients with RA
has been demonstrated (4).
Therefore, the development of an effective treatment for RA is
critical for the improvement of the health of these patients. The
aims of the treatments of RA include the suppression of
inflammation, remission of severe symptoms, prevention of organ and
joint damage and improvement of physical function (4,5).
However, RA treatment is costly with respect to the surgical
procedures, medications including biologics and indirect costs
(5). For example, at present the
approaches for the treatment of RA include the use of
anti-rheumatic drugs, but a good response has only been observed in
a minority of patients (5).
Therefore, it is necessary to optimize the treatment strategies to
ameliorate the clinical and socioeconomic effects of RA.
Although the exact immunological mechanism of RA
remains unclear, genetic and environmental factors have been
demonstrated to contribute to the pathogenesis of RA (6,7).
Several previous studies have suggested that RA fibroblast-like
synoviocytes (RA-FLS) serve a vital role in RA development by
various mechanisms (6-8). Firstly, RA-FLS produce
proinflammatory cytokines, including interleukin-6 (IL-6) and tumor
necrosis factor-α (TNF-α), achieving a perpetual state of
inflammation and cartilage destruction (1-3).
Secondly, as a uniquely aggressive phenotype, RA-FLS exhibit
significant levels of invasiveness into the extracellular matrix
and exacerbate joint damage (8).
The excessively proliferated RA-FLS increase oxygen consumption and
hypoxia in the local microenvironment, resulting in a series of
events including proinflammatory cell infiltration, synovium
inflammation and degradation of cartilage (9). These processes occur simultaneously,
and develop a positive feedback loop, resulting in the promotion of
RA progression (9,10). Therefore, targeting RA-FLS may
improve the clinical outcomes of RA treatment, and the induction of
proliferation inhibition and apoptosis of RA-FLS represents a
promising approach in the treatment of RA (8,10).
Ribonucleotide reductase M2 (RRM2) is a crucial
protein that regulates the formation of deoxyribonucleotides;
therefore, it is important for the repair and synthesis of DNA
(11). Inhibition of RRM2
significantly inhibits cellular proliferation and induces cell
apoptosis (12,13). A previous study suggested that
changes in RRM2 expression levels may exhibit a marked effect on
tumor metastasis and progression, making it a promising cancer
therapeutic target (14).
Gemcitabine and GTI-2040 are 2 RRM2 inhibitors and have undergone a
clinical trial for cancer therapy (trial no. NCT00078962) (13). Gao et al (14) confirmed that the expression of
RRM2 has been identified to be significantly increased in liver
cancer compared with in non-liver cancer tissue, and RRM2
suppression may significantly inhibit the proliferation and
migration of liver cancer cells, suggesting that RRM2 is a
promising target in the therapy of liver cancer. As RRM2
suppression has been validated to decrease the levels of cellular
proliferation and induce cellular apoptosis (14), we hypothesized that the inhibited
expression of RRM2 in RA-FLS may markedly suppress the
proliferation and increase apoptosis in RA-FLS. Nevertheless, to
the best of our knowledge, there has been limited data on the
effect of RRM2 suppression in RA until now.
Genetic therapy, also termed gene therapy, which is
the removal or alteration of genes in cells for therapeutic
purposes, represents a promising method for treating various
diseases, including RA (15,16). For the treatment of RA, the gene
therapy approach may confer potential effects through the specific
delivery of various gene products. At present, a number of studies
have been performed to investigate the therapeutic efficacy of gene
therapy in animal models (15,16). Tomita et al (16) demonstrated that the treatment of
rats with collagen-induced arthritis with nuclear factor
kappa-light-chain-enhancer of activated B cells decoy
oligodeoxynucleotides-loaded liposomes led to an amelioration of
arthritis. Notably, gene therapy using small interfering RNA
(siRNA) represents an elegant approach with great promise for the
treatment of various diseases, and is also a promising alternative
to chemotherapy, which is associated with several side effects
(17-20). As siRNA molecules alone cannot
cross cellular barriers to reach the targets inside the cells,
formulations based on nanoparticles have been demonstrated to
significantly increase the cellular delivery of siRNA and
facilitate siRNA-based therapy in clinical settings (17-20). In previous studies, we
successfully constructed a targeted liposome-polycation-DNA complex
(LPD) conjugated with the epidermal growth factor receptor antibody
to deliver siRNA efficiently to cancer cells and observed marked
gene silencing activity and anticancer efficacy (17-19).
Cell permeable peptides (CPP), also termed protein
transduction domains, are a diverse class of peptides with dozens
of amino acids that enhance the cellular uptake of various
substances including nanoparticles (21,22). CyLoP-1 is a cationic CPP rich in
cysteine and has been demonstrated by Jha et al (23) to exhibit the efficient cellular
delivery of various agents. In order to efficiently deliver RRM2
siRNA to RA-FLS and inhibit their proliferation, the present study
developed a cell permeable peptide-conjugated
liposome/protamine/DNA/RRM2 siRNA complex (CCP-LPDR) as a
nanoparticle-based drug delivery system for RA-FLS. We hypothesize
that CCP-LPDR may deliver RRM2 siRNA to RA-FLS effectively;
achieving increased therapeutic effect in RA-FLS. The in
vitro targeting, gene silencing activity and cellular
apoptosis-inducing activity of CCP-LPDR in RA-FLS were then
examined by the flow cytometry and reverse transcription
quantitative polymerase chain reaction (RT-qPCR). The present study
demonstrated that CCP-LPDR efficiently delivered RRM2 siRNA to
RA-FLS and achieved an improved therapeutic efficacy against RA-FLS
compared with the non-targeted control.
Materials and methods
Reagents
Grade X protamine, calf thymus DNA and cholesterol
were purchased from Sigma-Aldrich; Merck KGaA (Darmstadt, Germany).
The following reagents were purchased from Avanti Polar Lipids
(Alabaster, AL, USA): 1,2-dioleoyl-3-trimethylammonium-propane
(DOTAP, a cationic lipid); functional PEGylated lipids with a
maleimide group,
1,2-distearoyl-sn-glycero-3-phospho-ethanolamine-N-[maleimide
(polyethylene glycol)-2000] (DSPE-PEG-Mal); and,
1,2-dioleoyl-sn-glycero-3-phosphoeth-anolamine-N-carboxyfluorescein
(CFPE, a fluorescent lipid). GL Biochem (Shanghai) Ltd., (Shanghai,
China) synthesized the cell permeable peptides, CRWRWKCCKK (CyLoP-1
CCP). Shanghai GenePharma Co. Ltd. (Shanghai, China) synthesized
and provided the following siRNAs: Negative control (NC) siRNA,
fluorescent siRNA (FAM-siRNA) and RRM2 siRNA. The sequences of
siRNAs are summarized in Table I.
Dojindo Molecular Technologies, Inc. (Kumamoto, Japan) provided the
cellular proliferation kit Cell Counting Kit-8 (CCK-8), and the
Reverse Transcription System kit was provided by Promega
Corporation (Madison, WI, USA). Thermo Fisher Scientific, Inc.
(Waltham, MA, USA) provided the TRIzol® reagent and
SYBR™-Green PCR Master Mix. R&D Systems, Inc. (Minneapolis, MN,
USA) provided the ELISA kits to measure the concentration of TNF-α
(cat. no. DTA00C) and IL-6 (cat. no. D6050). Ultra-4 centrifugal
filter devices were purchased from EMD Millipore (Billerica, MA,
USA).
 | Table IqPCR primers and siRNA. |
Table I
qPCR primers and siRNA.
A, qPCR primer
sequences
|
|---|
| Gene name Forward
(5′-3′) | Reverse
(5′-3′) | Product size,
bp |
|---|
| β-actin
CGTGGACATCCGTAAAGACC |
ACATCTGCTGGAAGGTGGAC | 209 |
| RRM2
TCTATGGCTTCCAAATTGCC |
GACACAAGGCATCGTTTCAA | 128 |
| TNF-α
CACCACTTCGAAACCTGGGA |
TGTAGGCCCCAGTGAGTTCT | 105 |
| IL-6
CTCAATATTAGAGTCTCAACCCCCA |
GAGAAGGCAACTGGACCGAA | 163 |
B, siRNA sequences
|
|---|
| Gene name | Forward
(5′-3′) | Reverse
(5′-3′) | Product size,
bp |
|---|
| NC |
UUCUCCGAACGUGUCACGUTT |
ACGUGACACGUUCGGAGAATT | – |
| RRM2 |
GAUUUAGCCAAGAAGUUCAGA |
UGAACUUCUUGGCUAAAUCGC | – |
Cell culture of human RA-FLS
The RIKEN BioResource Center (Tsukuba, Japan)
provided the human RA-FLS MH7A cell line. The cells were maintained
in 10 cm cultured dishes at 37°C in an incubator with a humidified
atmosphere of 5% CO2. The medium used for cell culture
was Dulbecco's modified Eagle's medium with 10% fetal bovine serum
(Thermo Fisher Scientific, Inc.). Cells in the exponential phase of
growth were used for subsequent experiments.
Fabrication of LPD complex
The development of liposomes was performed using a
protocol as previously described (Fig. 1) (11,15,16). Briefly, cationic liposomes (the
concentration of liposomes was 10 mM with a 1:1 molar ratio of
DOTAP and cholesterol) were developed by a thin film method based
on hydration. The fluorescent liposomes with CFPE was constructed
by initially adding 1% CFPE (molar ratio) to the lipid mixture
composed of cholesterol and DOTAP. Multi-layer liposomes (MLL) were
formed following the hydration of the lipid film with deionized
water. Subsequent to hydration, MLL were then extruded to form
single-layer liposomes composed of DOTAP/cholesterol. To prepare
the LPD, the initially prepared DOTAP/cholesterol liposomes (125
µl) were mixed with 1 mg/ml protamine (30 µl) to produce 'solution
I'. At total of 24 µg calf thymus DNA and 24 µg siRNA (20 µM) were
also mixed together to form 'solution II'. To form LPD, solutions I
and II were then mixed together. At 50°C, the LPD complex was mixed
with 40 µl DSPE-PEG-Mal micelles (10 mg/ml). For peptide
conjugation, 80 µg CCP was mixed with the DSPE-PEG-Mal-modified LPD
complex at 25°C for 6 h.
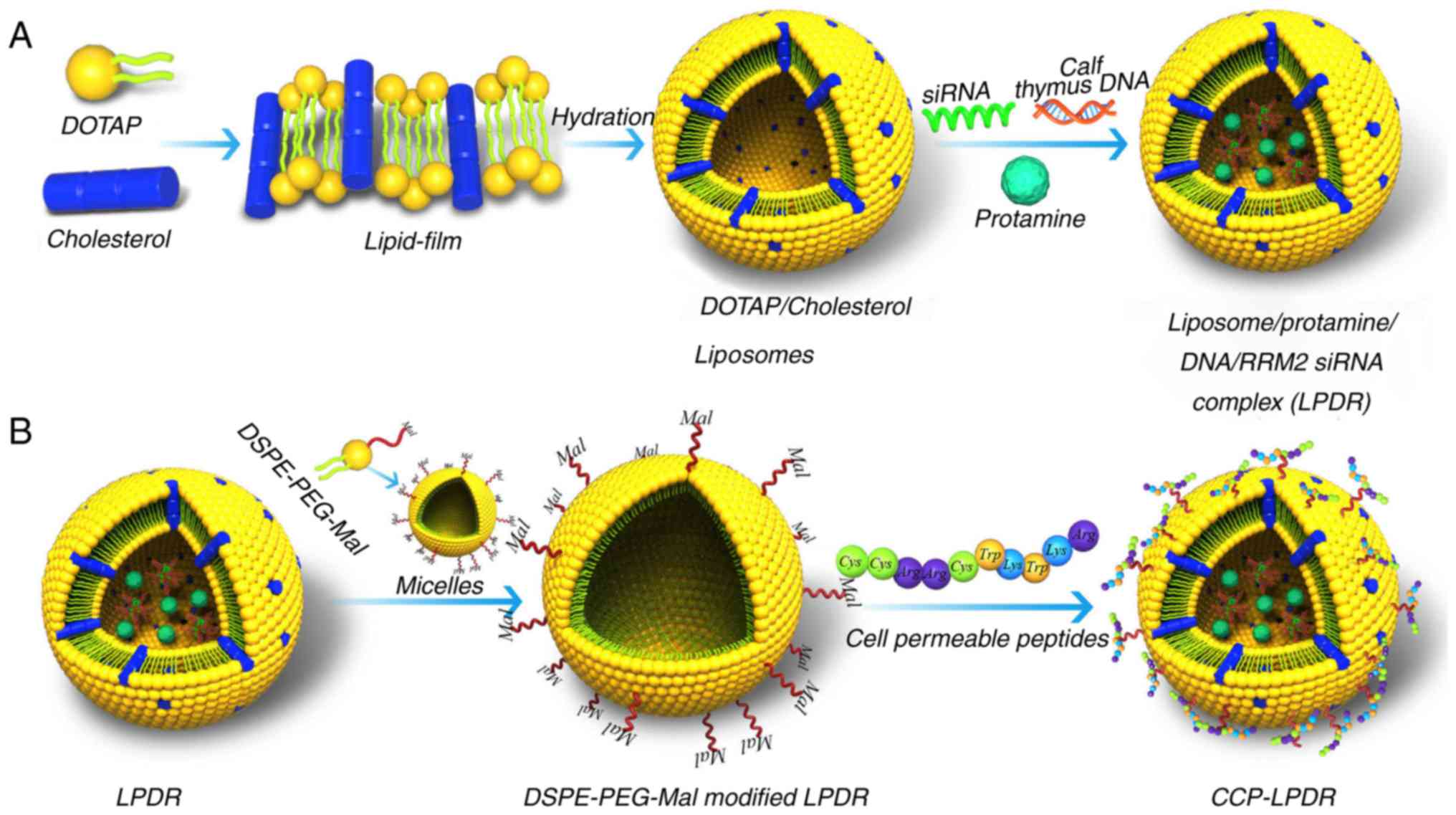 | Figure 1Development of cationic liposomes. (A)
The development of cationic liposomes comprising DOTAP and
cholesterol was performed using the thin film-based hydration
approach. DOTAP/cholesterol liposomes were formed following
hydration with water. Then, the LPDR was formed by mixing
DOTAP/cholesterol liposomes, RRM2 siRNA, protamine and calf thymus
DNA. (B) LPDR was inserted with the DSPE-PEG-Mal micelles, and CCP
were conjugated with the DSPE-PEG-Mal modified LPDR to form
CCP-LPDR. siRNA, small interfering RNA; CCP, cell permeable
peptides; RRM2, ribo-nucleotide reductase M2; DOTAP,
1,2-dioleoyl-3-trimethylammonium-propane; DSPE-PEG-Mal, functional
PEGylated lipids with a maleimide group,
1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[maleimide
(polyethylene glycol)-2000]; LPDR, liposome/protamine/DNA/RRM2
siRNA complex. |
Similar to CCP-LPDR, non-targeted LPD complexes
without CCP were developed without conjugation of any peptides. The
abbreviations used were follows: CCP-LPDR; LPDR; CCP-conjugated LPD
complex loaded with negative control siRNA (CCP-LPDN); and LPD
complex loaded with negative control siRNA (LPDN).
Size, encapsulation efficacy and
stability of cationic liposomes
The cationic liposomes (10 µl) were diluted with 1
ml deionized water, and the characteristics, including size,
polydispersity index (PDI) and zeta (ζ) potential of the liposomes
were determined with the Zeta sizer Nano S (Malvern Instruments,
Ltd., Malvern, UK). The siRNA encapsulation efficiency of the
liposomes was examined by an ultrafiltration-based approach:
Following the addition of the FAM-siRNA-loaded liposomes in the
Ultra-4 centrifugal filter devices, the centrifugation of the
filter devices was performed for 30 min at 3,500 × g and 25°C to
remove the unloaded siRNA. Following centrifugation, deionized
water was added. Subsequent to repeated ultrafiltration four times,
the unloaded FAM-siRNA were collected and quantified using the
calibration curve of FAM-siRNA with Microsoft Excel 2010 (Microsoft
Corporation, Redmond, WA, USA). The fluorescence for FAM-siRNA was
examined with the Synergy™ 4 (Biotek Instruments, Inc., Winooski,
VT, USA) at an excitation wavelength of 495 nm and emission
wavelength of 525 nm. The siRNA encapsulation efficiency, expressed
as percentage, was calculated using the following formula = [(The
mass of total siRNA - the mass of unloaded siRNA)/the mass of total
siRNA] × 100.
The evaluation of the stability of the liposomes was
performed as follows: Firstly, the liposomes were suspended in
various DMEM media, including PBS, 10% FBS or PBS with 20% FBS.
Then the solution was incubated at 25°C for 5 days. Each day, an
aliquot of liposomes was removed for the analysis of the change in
size of the cationic liposomes. An aliquot of liposomes (10 µl)
were diluted with 1 ml deionized water and the sizes of the
liposomes were determined with the Zetasizer Nano S (Malvern
Instruments, Ltd.) using the in-built software.
In vitro binding of liposomes with
RA-FLS
RA-FLS were plated in 48-well plates overnight. The
density of RA-FLS was 6×104 cells/well. Following this,
the old medium was discarded, and fresh medium was added to the
cell culture plates. The fresh medium contained the fluorescent
liposomes (FAM-siRNA loaded liposomes: Final FAM-siRNA
concentration, 200 nM; CFPE labeled liposomes: Final CFPE
concentration, 20 ng/ml). Subsequent to treatment with liposomes
for different lengths of times (1, 2, 4, 6, 12, 18 and 24 h), the
cells were washed with PBS, trypsinized, washed again, suspended in
PBS (0.3 ml) and examined using a flow cytometer (BD Biosciences,
San Jose, CA, USA). Data was analyzed using FlowJo (version 10;
FlowJo LLC, Ashland, OR, USA).
RT-qPCR
Using TRIzol® reagent (Thermo Fisher
Scientific, Inc.), the extraction of RNA from the RA-FLS was
performed following the manufacturer's protocol. The synthesis of
the first-strand cDNA was performed with the Reverse Transcription
System kit according to the manufacturer's protocol. The PCR was
performed with SYBR™ Green PCR Master Mix in a Roche Light Cycler
(Roche Diagnostics GmbH, Mannheim, Germany). The PCR thermocycler
conditions were described as follows: Denaturation for 2 min at
95°C, then 40 cycles of denaturation at 95°C for 3 sec, annealing
at 55°C for 10 sec and extension at 72°C for 25 sec. The
2ΔΔCq method was used to quantify the expression of Mrna
(24). The sequences of primers
used for β-actin, RRM2, TNF-α and IL-6 are summarized in Table I.
Western blot analysis
The extraction of proteins from the cells was
performed using radioimmunoprecipitation assay lysis buffer
(Beyotime Institute of Biotechnology, Haimen, China). The protein
concentration was determined by the Pierce BCA Protein Assay kit
(Thermo Fisher Scientific, Inc.). Each lane of 10% SDS-PAGE was
loaded with 50 µg protein. Proteins were then transferred onto
polyvinylidene fluoride membranes. The membranes were blocked with
10% bovine serum albumin (Thermo Fisher Scientific, Inc.) at 95°C
overnight. Following the transfer of the proteins to the membrane,
the following antibodies were added to the membrane: The primary
antibody was the mouse anti-human RRM2 mAb (1:1,000 dilution; cat.
no. ab57653; Abcam, Cambridge, MA, USA). The secondary antibody was
the horseradish peroxidase-conjugated goat anti-mouse IgG (1:1,500
dilution; ab97023; Abcam). The control antibody was GAPDH (cat. no.
ab9484; 1:1,000 dilution; Abcam). The visualization of the bands
was performed by the Amersham™ ECL Plus™ kit (GE Healthcare,
Chicago, IL, USA) and the Bio-Rad ChemiDoc XRS system with Quantity
One 1-D analysis software (version 4.6.8; Bio-Rad Laboratories,
Inc., Hercules, CA, USA).
Cell viability and apoptosis assays
The cell viability of RA-FLS following treatment
with liposomes was examined using a CCK-8 assay. RA-FLS were plated
for 16 h in 96-well cell culture plates. The density of cells was
1×104 cells/well. Following incubation overnight at
37°C, the old medium was removed, and the cells were exposed to
fresh medium containing liposomes (200 nM siRNA) for 6 h at 37°C.
Following this treatment, the drugs were discarded, and medium was
replaced by fresh medium. A total of 60 h later, the cell viability
of RA-FLS was measured following the protocol of the manufacturer
of the CCK-8 kit. The cell apoptosis assay was performed using the
Alexa Fluor® 488 Annexin V Apoptosis kit (Thermo Fisher
Scientific, Inc.), following the manufacturer's protocol. RA-FLS
were plated in 12-well cell culture plates overnight. The density
of RA-FLS was 2×105 cells/well. Following incubation at
37°C, the cells were then treated with liposomes (200 nM siRNA) for
6 h at 37°C. After 6 h, the drugs were removed, and medium was
replaced. A total of 60 h later, the cells were collected using
trypsinization and washed twice. Subsequent to washing, the
collected cells were re-suspended in an annexin-binding buffer.
Then, 5 µl propidium iodide (PI) and 5 µl Alexa Fluor®
488 Annexin V were added. After 15 min, 400 µl annexin-binding
buffer was added to the cells. The analysis of the cells was
performed with a flow cytometer (BD Biosciences). Data was analyzed
using FlowJo software (version 10).
Analysis of proinflammatory cytokines in
RA-FLS following liposome treatment
RA-FLS were plated in 12-well cell culture plates
overnight. The density was 2×105 cells/well. Then, the
RA-FLS were treated with liposomes (200 nM siRNA) for 6 h. The
drugs were removed 6 h later, and medium was replaced. A total of
60 h later, the protein and mRNA levels of cytokines were measured
using the IL-6 and TNF-α ELISA kits and RT-qPCR, respectively. The
samples (200 µl) were added to the ELISA plates and incubated for 2
h at 37°C. The sample was then aspirated and washed with PBS. Then,
200 µl conjugate was added to the plate and incubated for 4 h at
37°C. The substrate solution was then added, followed by the stop
solution. Finally, the absorbance of the wells at 450/540 nm was
examined with a microplate reader (Biotech ELx800 Universal; BioTek
Instruments, Inc.). The protein and mRNA levels of cytokines were
expressed as the percentage of the protein and mRNA levels of the
treated groups normalized to the untreated group.
Statistical analysis
The data was analyzed using SPSS (version 13; SPSS,
Inc., Chicago, IL, USA). A Student's unpaired t-test was used to
examine the differences between two groups, and a one-way analysis
of variance with a Student-Newman-Keuls post hoc test was performed
to examine the differences among ≥ three groups. P<0.05 was
considered to indicate a statistically significant difference.
Unless otherwise stated, all data in the present study are
presented as the mean ± standard deviation.
Results
Production and properties of the prepared
liposomes
Fig. 1 indicates
that the DOTAP/cholesterol liposomes were synthesized by hydrating
a lipid-film composed of DOTAP and cholesterol. Then, the
DOTAP/cholesterol liposomes were conjugated with RRM2 siRNA, calf
thymus DNA and protamine to form LPDR. The LPDR were then modified
with DSPE-PEG-Mal to prolong their circulation times. The grafted
PEG of DSPE-PEG-Mal was able to cover the surface of liposomes
efficiently and prevent the opsonization of liposomes. The CCP were
conjugated to the DSPE-PEG-Mal-modified LPDR to construct CCP-LPDR,
to increase the uptake of liposomes in RA-FLS.
The characteristics, including size, distribution, ζ
potential, polydispersity index (PDI) and encapsulation efficiency
of liposomes are summarized in Table
II. LPDR and LPDN were ~100 nm in size. Conjugation with CPP
did not have a significant effect on the liposome size, as observed
by the similar sizes of LPDR and CCP-LPDR, and those of LPDN and
CCP-LPDN. The low PDI (<0.2) of all the liposomes indicated that
the prepared liposomes had good homogeneity. All the liposomes
exhibited positive ζ potential of ~20 mV. The siRNA encapsulation
efficiency of LPD, CCP-LPDN and CCP-LPDR was >90% (Table II), indicating that the prepared
LPD complex was an efficient drug delivery system for siRNA.
 | Table IICharacteristics of
liposome-protamine-DNA-siRNA complex. |
Table II
Characteristics of
liposome-protamine-DNA-siRNA complex.
| Liposomes | Size, nm | Zeta potential,
mv | Polydispersity
index | Encapsulation
efficiency, % |
|---|
| CCP-LPDR | 135.32±9.35 | 18.92±7.54 | 0.15±0.06 | 93.45±2.87 |
| LPDR | 133.75±7.52 | 22.63±7.36 | 0.17±0.05 | 94.29±1.34 |
| CCP-LPDN | 142.82±8.13 | 21.93±6.27 | 0.18±0.03 | 92.63±3.18 |
| LPDN | 136.91±5.63 | 20.34±5.63 | 0.19±0.07 | 90.98±5.25 |
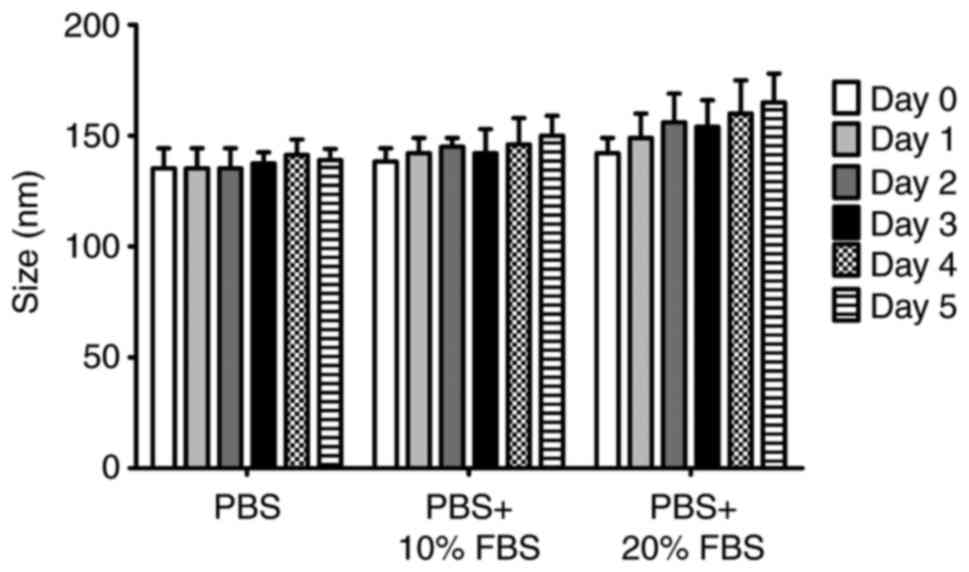
Data from the experiment to evaluate stability, in
which the liposomes were incubated in different media (PBS, PBS
with 10% FBS and PBS with 20% FBS) at 25°C for 5 days, is presented
in Fig. 2. Although an increase
of 10-20 nm in the size of the liposomes was observed when they
were in media containing FBS, the stability of liposomes was
markedly high during the entire incubation period, suggesting that
the serum did not significantly affect the stability of
liposomes.
In vitro binding of liposomes to
RA-FLS
To confirm whether CCP increased the targeting rate
of liposomes, fluorescent liposomes were incubated with RA-FLS.
Fig. 3A indicates that FAM-siRNA
CCP-LPDN exhibited a significantly increased transfection
efficiency compared with FAM-siRNA LPDN in RA-FLS in the stability
assay (P<0.05). With increases in time interval, the
transfection efficiency of FAM-siRNA LPDN also increased, and
plateaued at 18 h. Consistent with the FAM-siRNA assay, the in
vitro binding of RA-FLS with CFPE-labeled liposomes exhibited
similar results (Fig. 3B). CFPE
CCP-LPND exhibited markedly increased transfection efficiency
compared with CFPE LPND with RA-FLS (P<0.05). Again, with
increase in time intervals, the transfection efficiency of CFPE
CCP-LPND increased, and at 18 h reached a plateau.
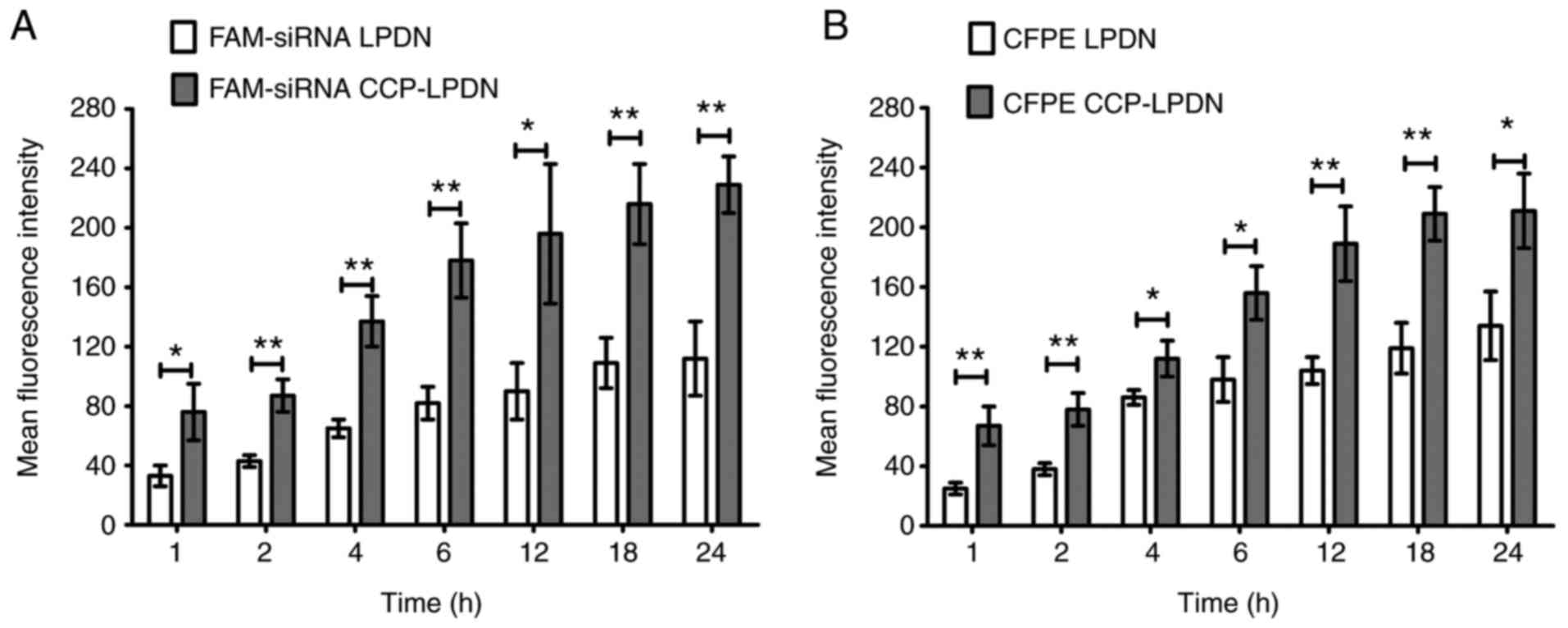 | Figure 3In vitro binding of liposomes
in RA-FLS. RA-FLS were incubated with florescent liposomes (200 nM
siRNA). Subsequently, the cells were washed to discard the
liposomes. Finally, RA-FLS were trypsinized, washed and suspended
in PBS. The analysis of the fluorescence was performed by flow
cytometry. (A) FAM-siRNA loaded liposomes; (B) CFPE-labeled
liposomes. *P<0.05; **P<0.01. Data are
presented as mean ± standard deviation (n=3). RA-FLS, rheumatoid
arthritis fibroblast-like synoviocytes; CFPE, fluorescent lipid,
1,2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-carboxyfluorescein;
siRNA, small interfering RNA; FMA-siRNA, fluorescent siRNA; CCP,
cell permeable peptides; LPD, liposome-polycation-DNA complex;
LPDN, LPD loaded with negative control siRNA; CCP-LPDN,
CCP-conjugated LPD loaded with negative control siRNA. |
Gene and protein silencing activity of
RRM2 by liposomes in RA-FLS
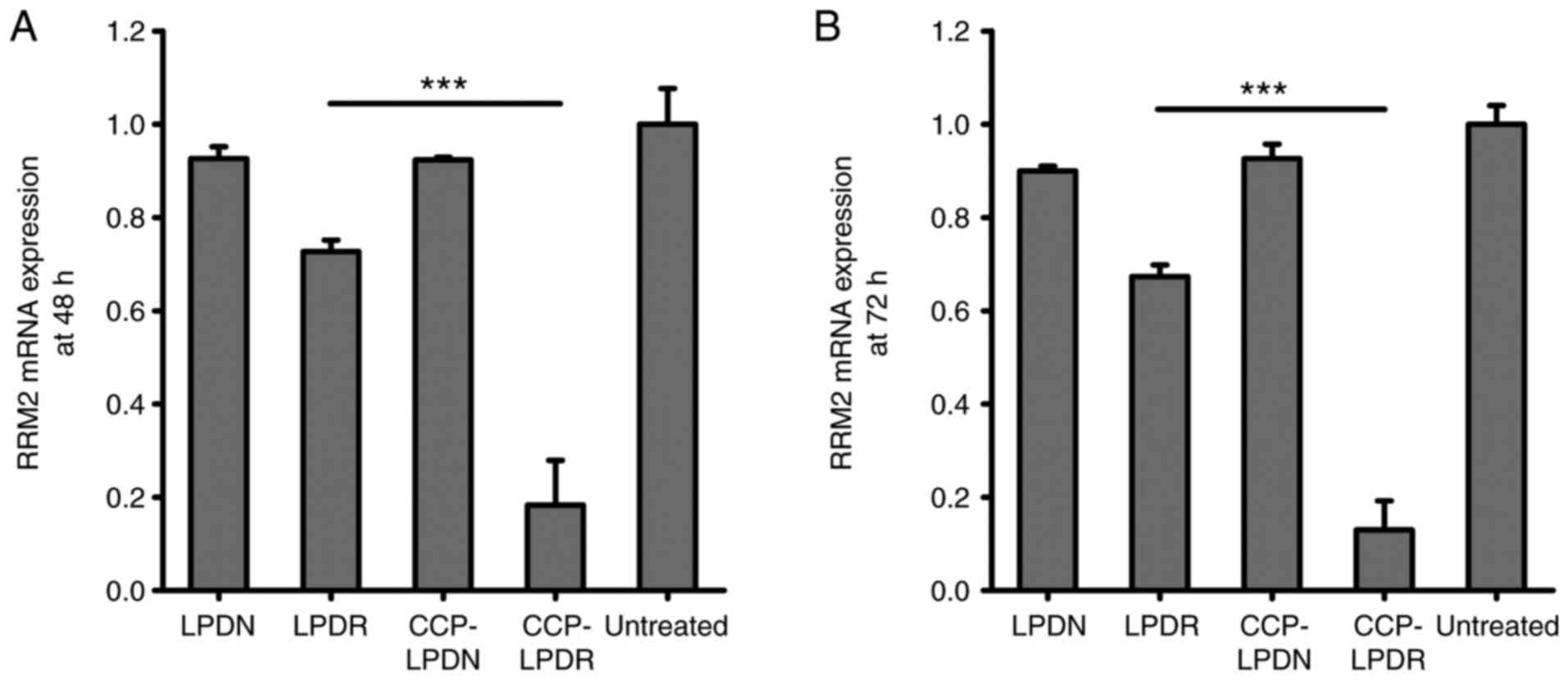
The gene and protein silencing activity of RRM2 by
liposomes was evaluated, and results are presented in Figs. 4 and 5. In the absence of RRM2 siRNA, LPDN and
CCP-LPDN barely affected RRM2 gene expression (Fig. 4). LPDR exhibited poor gene
silencing activity and inhibited the gene expression of RRM2 by
only ~25%. On the contrary, CCP-LPDR remarkably suppressed the RRM2
gene expression by ~80%, indicating an improved gene silencing
activity compared with that exhibited by LPDR (P<0.001). The
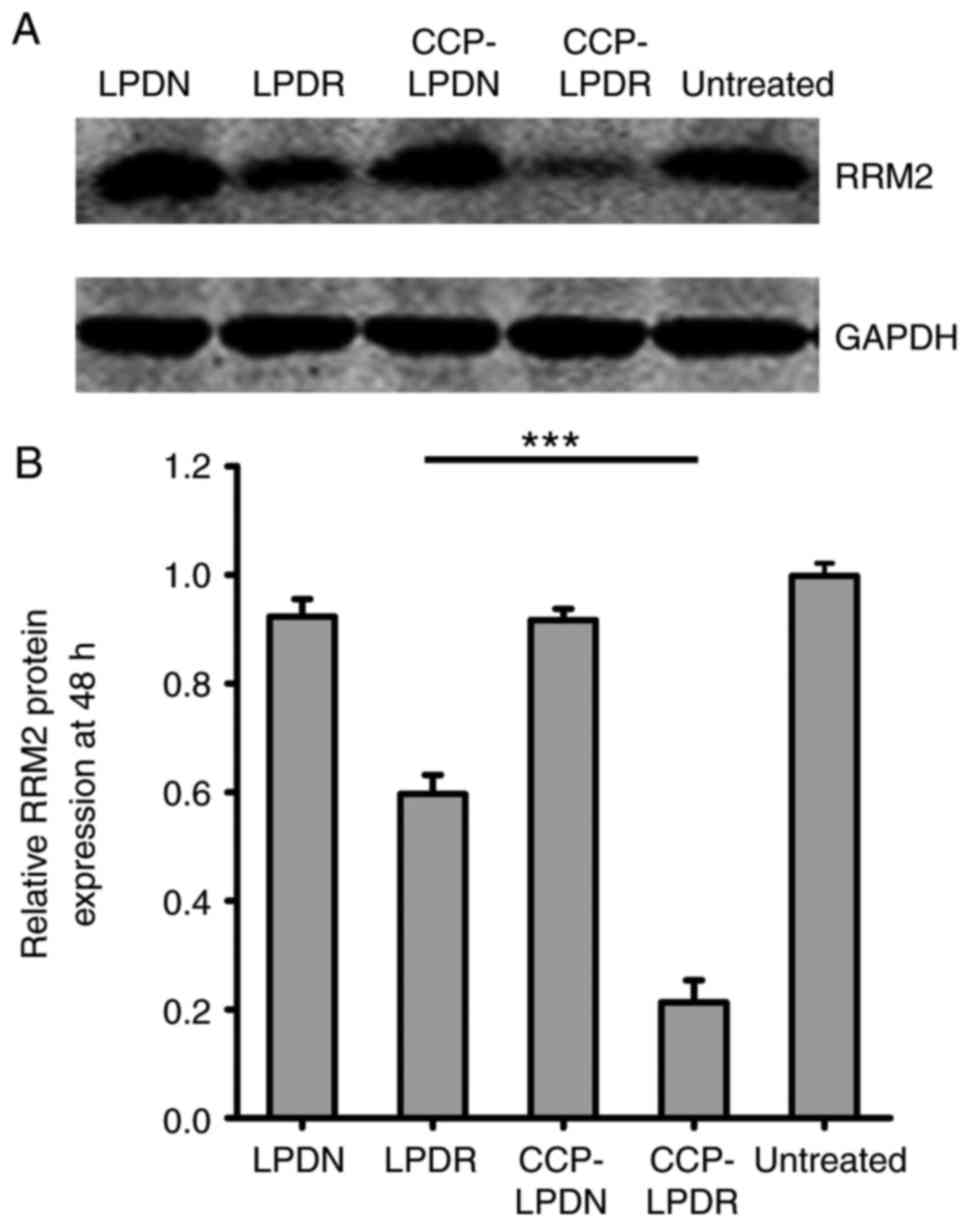
analysis of the expression of RRM2 protein obtained similar
results, as demonstrated in Fig.
5. CCP-LPDR inhibited the expression of RRM2 protein by ~80%,
whereas LPDR inhibited the expression of RRM2 protein by ~20%
(P<0.001), indicating that the RRM2 suppression by CCP-LPDR was
more efficient in comparison with LPDR. Therefore, it was confirmed
that CCP-LPDR exhibited improved RRM2 suppression activity in
RA-FLS in comparison with LPDR.
Activity of liposomes on the cellular
proliferation and apoptosis in RA-FLS
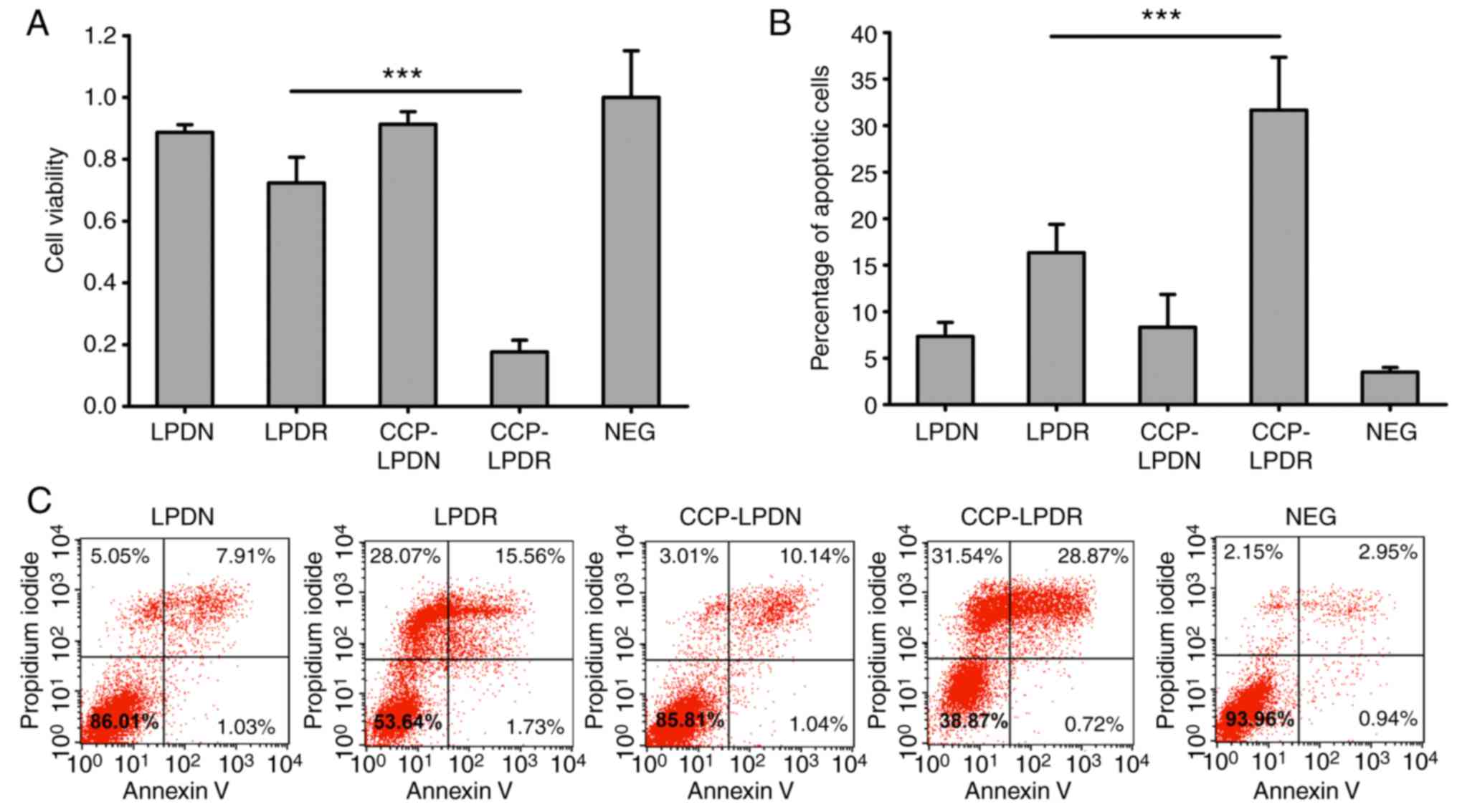
The effect of liposomes on the levels of
proliferation of RA-FLS was investigated using the CCK-8 assay
(Fig. 6A). As demonstrated in
Fig. 6A, LPDN and CCP-LPDN did
not affect the proliferation of RA-FLS. It is noteworthy that
CCP-LPDR exhibited significant inhibition of proliferation of
RA-FLS compared with LPDR (P<0.001). Similarly, LPDN and
CCP-LPDN did not induce significant apoptosis in RA-FLS, as
reflected by the fact that apoptotic cells constituted only ~5% of
the total population of cells following their treatment, which was
comparable to that observed in the untreated control group
(Fig. 6B and C). However, the
percentage of apoptotic cells was noticeably increased, and reached
~15%, following LPDR treatment, and CCP-LPDR treatment induced an
even higher percentage of apoptotic cells (~30%) compared with LPDR
(P<0.001), suggesting that CCP-LPDR exhibited the highest
efficiency in inducing apoptosis in RA-FLS.
Analysis of proinflammatory cytokines in
RA-FLS following liposome treatment
IL-6 and TNF-α are potent proinflammatory cytokines
secreted by RA-FLS, and they are pleiotropic cytokines with pivotal
roles in the pathophysiology of RA (25,26). They are used as valuable indexes
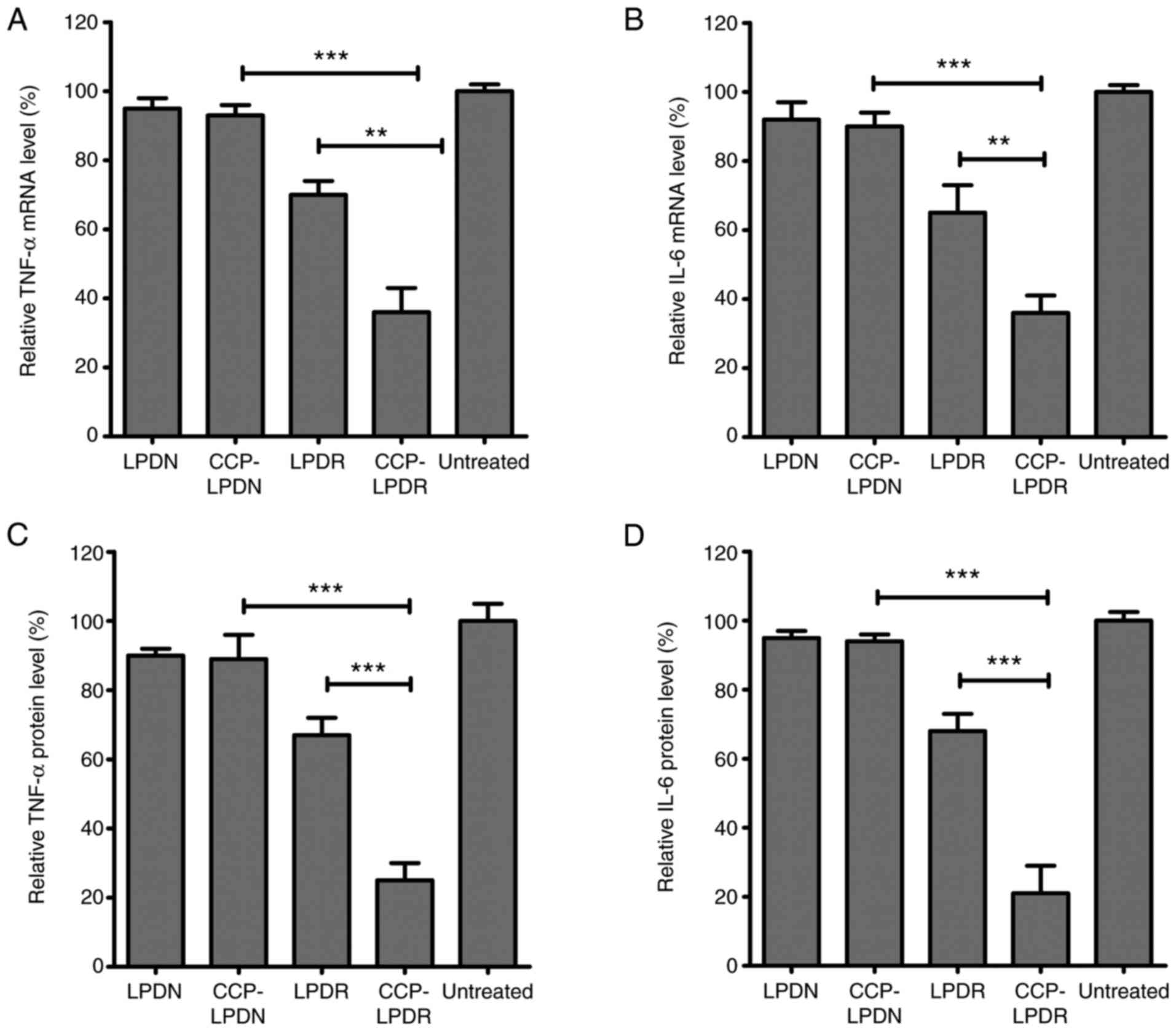
to evaluate the efficacy of treatment of RA (25,26). As indicated in Fig. 7, the levels of IL-6 and TNF-α in
the treated group were measured as the percentage of their mRNA and
protein levels normalized to the untreated group (Fig. 7). LPDN and CCP-LPDN affected RRM2
expression levels minimally, whereas LPDR and CCP-LPDR
significantly inhibited its expression in RA-FLS at mRNA and
protein levels. As demonstrated in Fig. 7A, CCP-LPDR induced a more
effi-cient inhibition of the TNF-α mRNA expression compared with
CCP-LPDN (P<0.001) and LPDR (P<0.01). Similarly, CCP-LPDR was
also more effective in inhibiting IL-6 mRNA expression compared
with CCP-LPDN (P<0.001) and LPDR (P<0.01) (Fig. 7B). With respect to the TNF-α and
IL-6 protein levels, similar results were achieved (Fig. 7C and D). CCP-LPDR was more
effective in inhibiting the TNF-α and IL-6 protein levels compared
with CCP-LPDN and LPDR (P<0.001), whereas LPDN and CCP-LPDN
minimally affected the protein levels of TNF-α and IL-6 in RA-FLS.
Taken together, the expression levels of TNF-α and IL-6, the
proinflammatory cytokines, were markedly decreased in RA-FLS
following CCP-LPDR treatment.
Discussion
RA incurs high costs of treatment with respect to
surgical procedures, medications including biologics and indirect
costs. Therefore, optimized treatments are required for the
effective treatment of RA. Accumulating evidence has suggested that
RA-FLS serve a key role in the progression of RA by promoting the
production of proteases (6-8).
The present study developed CCP-LPDR, which efficiently delivered
RRM2 siRNA to RA-FLS, achieving an improved therapeutic efficacy
against RA-FLS compared with the non-targeted control.
The choice of target in gene therapy is crucial in
liposome-based gene therapy. In the present study, RRM2 was
selected as it is a crucial protein involved in DNA repair and
synthesis (11,12). Due to the substantial effect of
RRM2 on the development and metastasis of tumors, RRM2 inhibitors
(for example, GTI-2040 and gemcitabine) have been recruited for
clinical trials for various types of cancer (13). Our previous study validated that
RRM2 is a superior target for the treatment of liver cancer, and
following RRM2 suppression, the levels of migration and
proliferation of liver cancer cells were significantly inhibited
(14). The progression of RA is
similar to benign tumors, and the proliferation of RA-FLS and
abnormal synovium promote the progression of RA (6). Therefore, the inhibition of RA-FLS
proliferation was hypothesized to be a potential treatment for RA.
Considering that RRM2 serves a crucial role in proliferation of
RA-FLS, the present study aimed to inhibit their proliferation by
suppressing RRM2 expression in RA-FLS. To the best of our
knowledge, the present study is the first to demonstrate that,
following RRM2 suppression in RA-FLS by CCP-LPDR, the proliferation
of RA-FLS was significantly inhibited by ~80% compared with the
untreated control, and the level of apoptosis observed was ~30%
(Fig. 6). It is also the first
study to confirm that, by RRM2 suppression via CCP-LPDR,
significant inhibition of cellular proliferation and promotion in
cellular apoptosis of RA-FLS may be induced, suggesting that RRM2
is a good therapeutic target for RA.
Developing improved siRNA-loaded nanoparticles is a
critical step for the translation of siRNA-based therapies into
clinical settings (24). Cationic
liposomes serve as potential nanocarriers for siRNA delivery,
however, they are prone to fast clearance following uptake by
reticuloendothelial system (RES) (24). The attachment of opsonins to
cationic liposomes is the primary reason for a high affinity of RES
for unprotected cationic liposomes (25). Stealth liposomes with surface
grafted PEG is a practical approach to decrease the RES uptake, by
shielding the charge on the surface of the liposomes (26). In our previous studies
investigating breast and liver cancer, the LPD complexes prepared
were advanced PEGylated cationic liposomes, which exhibited a long
circulation time in vivo and accumulation in the body
(17-19). Therefore, the CCP-LPDR in the
present study was expected to avoid RES uptake in vivo. It
was demonstrated that the presence of CCP contributed significantly
to the uptake of CCP-LPDR in RA-FLS. The present study indicated
that the transfection efficiency of CCP-LPDR was markedly increased
in comparison with LPDN, achieving marked inhibition of RRM2 gene
and protein expression in RA-FLS. Subsequent to cell binding,
CCP-LPDR induced an increased inhibition of proliferation and
promotion of apoptosis in RA-FLS compared with LPDR, suggesting
that CCP may significantly improve the targeting and treatment
efficacy of CCP-LPDR in RA-FLS.
RA-FLS secrete IL-6 and TNF-α, which are potent
proinflammatory cytokines that promote the progression of RA
(27,28). It is noteworthy that the CCP-LPDR
method was more effective in the inhibition of the TNF-α and IL-6
protein levels in comparison with CCP-LPDN and LPDR, whereas LPDN
and CCP-LPDN barely affected the protein levels of TNF-α and IL-6
in RA-FLS. Therefore, TNF-α and IL-6, the proinflammatory
cytokines, were markedly decreased in RA-FLS following CCP-LPDR
treatment.
As FLS are required to be obtained by primary
culture from rats, a universal protocol of the culturing of primary
FLS from rats has not been successfully established. Therefore, the
effect of the treatment on normal FLS has not been assessed at
present. Nevertheless, examining the effect of the treatment on
normal FLS is important and will be incorporated into future study
if possible.
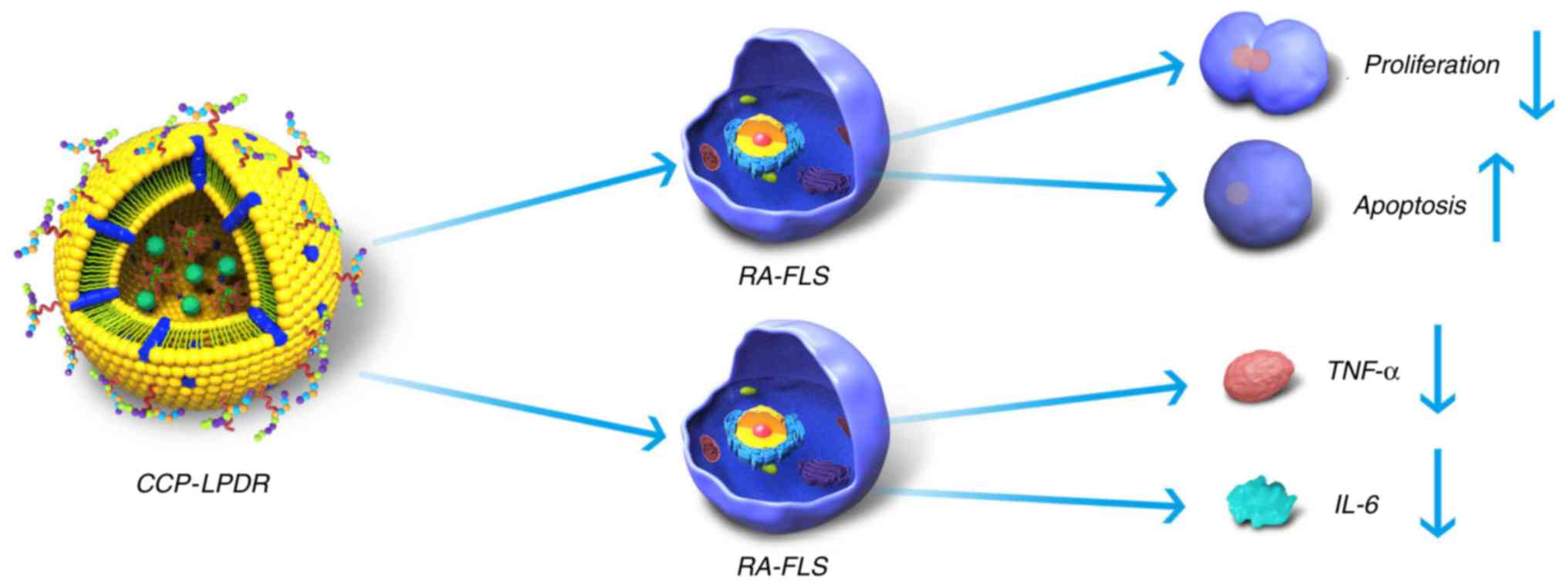
Taken together, the present study suggested the
mechanism of treatment efficacy of CCP-LPDR in RA (Fig. 8). Firstly, CCP-LPDR was targeted
to and internalized by RA-FLS. In the cytoplasm, CCP-LPDR released
RRM2 siRNA, achieving a combined therapeutic efficacy by increasing
the inhibition of proliferation and promoting apoptosis in RA-FLS.
Furthermore, the levels of proinflammatory cytokines TNF-α and IL-6
were also markedly decreased by the treatment with CCP-LPDR in
RA-FLS.
The establishment of a potential treatment for
RA-FLS cells is a viable approach for the treatment of RA. In the
present study, a CCP-LPDR system was developed, which efficiently
delivered RRM2 siRNA to RA-FLS, obtaining increased therapeutic
efficacy with RA-FLS by increasing the levels of apoptosis and
inhibition of cellular proliferation and proinflammatory cytokines.
In conclusion, CCP-LPDR offered the possibility of suppressing
RA-FLS and therefore provided a potential therapeutic approach for
RA.
Funding
The present study was funded by the National Natural
Science Foundation of China (grant nos. 81771964 and 81472829).
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors' contributions
SF contributed to the design of the study and wrote
the manuscript. XiW performed the experiments. XuW and JS analyzed
the data. All authors have read and approved the final
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Authors' information
Dr Xin Wang, The First Department of Pain, Qingdao
Municipal Hospital, Qingdao, Shandong 266011; Dr Xueping Wang and
Dr Shiou Fu, The Second Department of Pain, Qingdao Municipal
Hospital, Qingdao, Shandong 266011; Dr Jin Sun, International Joint
Cancer Institute, Second Military Medical University, Shanghai
200433, P.R. China.
Acknowledgments
Not applicable.
References
|
1
|
Mclnnes IB and Schett G: The pathogenesis
of rheumatoid arthritis. N Engl J Med. 365:2205–2219. 2011.
View Article : Google Scholar
|
|
2
|
Glant TT, Mikecz K and Rauch TA:
Epigenetics in the pathogenesis of rheumatoid arthritis. BMC Med.
12:352014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dougados M, Soubrier M, Antunez A, Balint
P, Balsa A, Buch MH, Casado G, Detert J, El-Zorkany B, Emery P, et
al: Prevalence of comorbidities in rheumatoid arthritis and
evaluation of their monitoring: Results of an international,
cross-sectional study (COMORA). Ann Rheum Dis. 73:62–68. 2014.
View Article : Google Scholar :
|
|
4
|
Gabriel SE, Crowson CS, Kremers HM, Doran
MF, Turesson C, O'Fallon WM and Matteson EL: Survival in rheumatoid
arthritis: A population-based analysis of trends over 40 years.
Arthritis Rheum. 48:54–58. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kobelt G and Jönsson B: The burden of
rheumatoid arthritis and access to treatment: outcome and
cost-utility of treatments. Eur J Health Econ. 8(Suppl 2):
S95–S106. 2008. View Article : Google Scholar
|
|
6
|
Guo Q, Wang Y, Xu D, Nossent J, Pavlos NJ
and Xu J: Rheumatoid arthritis: Pathological mechanisms and modern
pharmacologic therapies. Bone Res. 6:152018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Boechat Nde O, Ogusku MM, Boechat AL and
Sadahiro A: Interaction between smoking and HLA-DRB1*04 gene is
associated with a high cardiovascular risk in Brazilian Amazon
patients with rheumatoid arthritis. PLoS One. 7:e415882012.
View Article : Google Scholar
|
|
8
|
Bartok B and Firestein GS: Fibroblast-like
synoviocytes: Key effector cells in rheumatoid arthritis. Immunol
Rev. 233:233–255. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Konisti S, Kiriakidis S and Paleolog EM:
Hypoxia - a key regulator of angiogenesis and inflammation in
rheumatoid arthritis. Nat Rev Rheumatol. 8:153–162. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Tang Q, Cui J, Tian Z, Sun J, Wang Z,
Chang S and Zhu S: Oxygen and indocyanine green loaded
phase-transition nanoparticle-mediated photo-sonodynamic cytotoxic
effects on rheumatoid arthritis fibroblast-like synoviocytes. Int J
Nanomedicine. 12:381–393. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Duxbury MS, Ito H, Zinner MJ, Ashley SW
and Whang EE: RNA interference targeting the M2 subunit of
ribonucleotide reductase enhances pancreatic adenocarcinoma
chemosensitivity to gemcitabine. Oncogene. 23:1539–1548. 2004.
View Article : Google Scholar
|
|
12
|
Zhang K, Hu S, Wu J, Chen L, Lu J, Wang X,
Liu X, Zhou B and Yen Y: Overexpression of RRM2 decreases
thrombspondin-1 and increases VEGF production in human cancer cells
in vitro and in vivo: Implication of RRM2 in angiogenesis. Mol
Cancer. 8:112009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Shao J, Zhou B, Chu B and Yen Y:
Ribonucleotide reductase inhibitors and future drug design. Curr
Cancer Drug Targets. 6:409–431. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gao J, Chen H, Yu Y, Song J, Song H, Su X,
Li W, Tong X, Qian W, Wang H, et al: Inhibition of hepatocellular
carcinoma growth using immunoliposomes for co-delivery of
adriamycin and ribonucleotide reductase M2 siRNA. Biomaterials.
34:10084–10098. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Roman-Blas JA and Jimenez SA: NF-kappaB as
a potential therapeutic target in osteoarthritis and rheumatoid
arthritis. Osteoarthritis Cartilage. 14:839–848. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tomita T, Takeuchi E, Tomita N, Morishita
R, Kaneko M, Yamamoto K, Nakase T, Seki H, Kato K, Kaneda Y, et al:
Suppressed severity of collagen-induced arthritis by in vivo
transfection of nuclear factor kappaB decoy oligodeoxynucleotides
as a gene therapy. Arthritis Rheum. 42:2532–2542. 1999. View Article : Google Scholar
|
|
17
|
Gao J, Liu W, Xia Y, Li W, Sun J, Chen H,
Li B, Zhang D, Qian W, Meng Y, et al: The promotion of siRNA
delivery to breast cancer overexpressing epidermal growth factor
receptor through anti-EGFR antibody conjugation by immunoliposomes.
Biomaterials. 32:3459–3470. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gao J, Yu Y, Zhang Y, Song J, Chen H, Li
W, Qian W, Deng L, Kou G, Chen J, et al: EGFR-specific PEGylated
immunoliposomes for active siRNA delivery in hepatocellular
carcinoma. Biomaterials. 33:270–282. 2012. View Article : Google Scholar
|
|
19
|
Gao J, Sun J, Li H, Liu W, Zhang Y, Li B,
Qian W, Wang H, Chen J and Guo Y: Lyophilized HER2-specific
PEGylated immunoliposomes for active siRNA gene silencing.
Biomaterials. 31:2655–2664. 2010. View Article : Google Scholar
|
|
20
|
Xu C, Lee SA and Chen X: RNA interference
as therapeutics for hepatocellular carcinoma. Recent Pat Anticancer
Drug Discov. 6:106–115. 2011. View Article : Google Scholar
|
|
21
|
Gao H, Zhang Q, Yu Z and He Q:
Cell-penetrating peptide-based intelligent liposomal systems for
enhanced drug delivery. Curr Pharm Biotechnol. 15:210–219. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Koren E and Torchilin VP: Cell-penetrating
peptides: Breaking through to the other side. Trends Mol Med.
18:385–393. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Jha D, Mishra R, Gottschalk S, Wiesmüller
KH, Ugurbil K, Maier ME and Engelmann J: CyLoP-1: A novel
cysteine-rich cell-penetrating peptide for cytosolic delivery of
cargoes. Bioconjug Chem. 22:319–328. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2−ΔΔC T method. Methods. 25:402–408.
2001. View Article : Google Scholar
|
|
25
|
Srirangan S and Choy EH: The role of
interleukin-6 in the pathophysiology of rheumatoid arthritis. Ther
Adv Musculoskelet Dis. 2:247–256. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Alonso-Ruiz A, Pijoan JI, Ansuategui E,
Urkaregi A, Calabozo M and Quintana A: Tumor necrosis factor alpha
drugs in rheumatoid arthritis: Systematic review and metaanalysis
of efficacy and safety. BMC Musculoskelet Disord. 9:522008.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lee H, Lytton-Jean AK, Chen Y, Love KT,
Park AI, Karagiannis ED, Sehgal A, Querbes W, Zurenko CS, Jayaraman
M, et al: Molecularly self-assembled nucleic acid nanoparticles for
targeted in vivo siRNA delivery. Nat Nanotechnol. 7:389–393. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Tseng YC, Mozumdar S and Huang L:
Lipid-based systemic delivery of siRNA. Adv Drug Deliv Rev.
61:721–731. 2009. View Article : Google Scholar : PubMed/NCBI
|