Introduction
Osteoarthritis is a common joint disease affecting
elderly individuals that induces joint pain and stiffness, thus
severely affecting the quality of life of these individuals
(1). The incidence rate of
osteoarthritis among Asians aged ≥65 years is predicted to more
than double from 6.8% in 2008 to 16.2% in 2040. In addition, due to
an increase in the incidence of sports-associated trauma, the
average age of patients with osteoarthritis is estimated to
decrease (2). As a chronic joint
disease, osteoarthritis is often characterized by articular
cartilage degeneration and osteophyte formation. The lack of an
articular cartilage blood supply, neural control and lymphatic
vessels render the repair of this type of damage difficult
(3). To date, the precise
pathogenesis and molecular mechanisms responsible for the
development and progression of osteoarthritis remain unclear; thus,
there is no effective method yet available for the prevention or
treatment of osteoarthritis. The risk factors of osteoarthritis
include previous joint injury, abnormal joint or limb development
and genetic background, as well as being overweight or having a job
which requires the lifting of heavy objects, in which the
mechanical stress on the joints and low-grade inflammation cause
degenerative joint disease, such as osteoarthritis (4). Therefore, further studies on the
molecular pathogenesis of osteoarthritis may aid in the development
of novel approaches with which to prevent and control this disease
in order to improve the quality of life of affected
individuals.
Recently, microRNAs (miRNAs or miRs) in the
cartilage under conditions of homeostasis and osteoarthritis have
gained increasing attention. miRNAs are a class of endogenous
non-coding RNAs of 18-24 nucleotides in length that
post-transcriptionally regulate the expression of protein-coding
genes. miRNAs can recognize and bind to the 3′-untranslated region
of gene transcripts (mRNA) through base complementary pairing to
guide the RNA-induced silencing complex and degrade the target mRNA
or repress translation of the target mRNA (5-7).
Previous studies have demonstrated that both miR-140 and miR-146a
are involved in the pathogenesis of osteoarthritis, thus suggesting
that miRNAs play an important role in cartilage homeostasis
(8-12). A more recent study revealed that
miR-1 inhibited the ossification of the posterior longitudinal
ligament via the MALAT1/miR-1/Cx43 regulatory axis in patient
ligament samples (13). However,
the role of miR-1 in the development of osteoarthritis remains
unknown. A recent study demonstrated that miR-1 was highly
expressed in the hypertrophic zone of growth plate cartilage, and
regulated chondrocyte phenotypes during growth plate development
(14). Therefore, it was
hypothesized that miR-1 regulates chondrocyte homeostasis. Previous
studies have revealed that both miR-1 and Indian hedgehog (Ihh) are
distributed in the pre-hypertrophic region of cartilage tissues
(11,14,15). This region is involved in
chondrocyte hypertrophy, suggesting that miR-1 and Ihh play a role
in chondrocyte homeostasis.
Indeed, Ihh is involved in cartilage degeneration
and osteoarthritis progression (16). It has been demonstrated that
chondrocytes undergo hypertrophic differentiation in the early
stages of osteoarthritis (17,18), while Ihh functions to regulate
chondrocyte hypertrophy, thereby achieving endochondral
ossification (19-21). It was previously demonstrated that
the disruption of the Ihh signaling pathway in vivo was able
to attenuate osteoarthritis progression in a transgenic mouse
Ihhfl/fl model of osteoarthritis induced by surgery
(22), while Ipriflavone was able
to reduce cartilage degeneration in rats by blocking Ihh signaling
(23). Thus, the upregulation of
the Ihh pathway plays an important role in osteoarthritis
progression, whereas the inhibition of the Ihh pathway attenuates
cartilage degradation.
In the present study, the levels of miR-1 and Ihh
were first assessed in the tibial plateau of humans with or without
osteoarthritis, and a transgenic mouse model of osteoarthritis was
then established after subjecting
Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1 transgenic
mice to anterior cruciate ligament transection (ACLT) (24,25). The effects of miR-1 expression in
mice on the regulation of Ihh, glioma-associated oncogene homolog
(Gli)1, Gli2, Gli3, smoothened homolog (Smo), MMP-13, collagen type
X (Col10), Col2a1 and Aggrecan (Acan) expression were also
analyzed.
Materials and methods
Human cartilage tissues
The present study was approved (2019YX260) by the
Institutional Ethics Committee of the Second Hospital of Shanxi
Medical University (Taiyuan, China), and all patients provided
informed consent. Cartilage tissues (n=20) were obtained from the
cartilage samples of the tibial plateau during total knee
arthroplasty of patients with osteoarthritis who were diagnosed
according to the American Rheumatism Association Criteria for
osteoarthritis (26).
Histologically, these cartilage samples exhibited severe damage and
were harvested from the medial region of the tibial plateau, while
the relatively normal cartilage was harvested from the normal
appearing non-loaded area of the tibial plateau of the same patient
as a control (normal). These cartilage tissue samples after harvest
were ground in liquid nitrogen using a mortar and pestle and used
for reverse transcription-quantitative PCR (RT-qPCR), while the
articular cartilage tissue sections were processed and stained by
Safranin O/fast green or immunohistochemistry.
RT-qPCR
The whole-knee cartilage was first dissected with a
scalpel and then ground in liquid nitrogen, and total cellular RNA
was isolated from the human and mouse samples (for mouse samples,
please see below) using TRIzol™ reagent (Invitrogen; Thermo Fisher
Scientific, Inc.), according to the manufacturer's instructions.
The cartilage samples from 3 mice were pooled together, and 3
pooled samples per group were used for RNA isolation. These RNA
samples were then subjected to reverse transcription into
complementary DNA (cDNA) using PrimeScript™ RT Master Mix (Takara
Bio, Inc.). Total cellular miRNA was isolated from these human and
mouse cartilage samples using the miRNeasy Mini kit (Qiagen),
according to the manufacturer's protocol, and reverse transcribed
into cDNA using the MiScript Reverse Transcription kit (Qiagen).
rRNA 18s and U6 were used as internal controls for mRNA and miRNA,
respectively. The stem-loop primers for miR-1 were purchased from
Qiagen. These cDNA samples were then subjected to qPCR
amplification using the TB Green™ Premix Ex Taq™ II kit (Takara)
with the Applied Biosystems™ QuantStudio™ 6 Flex real-time PCR
system (Applied Biosystems; Thermo Fisher Scientific, Inc.). The
qPCR conditions were as follows: Pre-incubation of samples at 50°C
for 2 min and 95°C for 10 min, and then 40 cycles of denaturation
at 95°C for 10 sec, annealing at 55°C for 30 sec, and extension at
72°C for 30 sec. The level of each transcript was quantified by
using the threshold cycle (Ct) 2−ΔΔCq method (27). The primer sequences are presented
in Table I.
 | Table IPrimers for used for RT-qPCR. |
Table I
Primers for used for RT-qPCR.
| Species | Gene | DNA sequence |
|---|
| Mouse | Col2a1 |
5′-AAGGGACACCGAGGTTTCACTGG-3′ |
| |
5′-GGGCCTGTTTCTCCTGAGCGT-3′ |
| Col10 |
5′-GCCAGGAAAGCTGCCCCACG-3′ |
| |
5′-GAGGTCCGGTTGGGCCTGGT-3′ |
| Acan |
5′-CAGTGGGATGCAGGCTGGCT-3′ |
| |
5′-CCTCCGGCACTCGTTGGCTG-3′ |
| MMP-13 |
5′-GGACCTTCTGGTCTTCTGGC-3′ |
| |
5′-GGATGCTTAGGGTTGGGGTC-3′ |
| Ihh |
5′-CCACTTCCGGGCCACATTTG-3′ |
| |
5′-GGCCACCACATCCTCCACCA-3′ |
| Gli1 |
5′-GGTCCGGATGCCCACGTGAC-3′ |
| |
5′-TCCCGCTTGGGCTCCACTGT-3′ |
| Gli2 |
5′-TGGCAGCGATGGGCCTACCT-3′ |
| |
5′-GTGTGCTGCTGTTTGGC-3′ |
| Gli3 |
5′-CATGAACAGCCCTTTAAGAC-3′ |
| |
5′-TCATATGTGAGGTAGCACCA-3′ |
| Smo |
5′-CTCCTACTTCCACCTGCTCAC-3′ |
| |
5′-CAAAACAAATCCCACTCACAGA-3′ |
| Human | COL2A1 |
5′-TGAGGGCGCGGTAGAGACCC-3′ |
| |
5′-TGCACACAGCTGCCAGCCTC-3′ |
| COL10 |
5′-TGCCTCTTGTCAGTGCTAACC-3′ |
| |
5′-GCGTGCCGTTCTTATACAGG-3′ |
| Acan |
5′-CATTCACCAGTGAGGACCTCGT-3′ |
| |
5′-TCACACTGCTCATAGCCTGCTTC-3′ |
| MMP-13 |
5′-TGCTGCATTCTCCTTCAGGA-3′ |
| |
5′-ATGCATCCAGGGGTCCTGGC-3′ |
| Ihh |
5′-ATCATCTTCAAGGACGAGGAGA-3′ |
| |
5′-GGGCCTTTGACTCGTAATACAC-3′ |
| Gli1 |
5′-GAACCCTTGGAAGGTGATATGTC-3′ |
| |
5′-GGCAGTCAGTTTCATACACAGAT-3′ |
| Gli2 |
5′-GCGTGTTTACCCAATCCTGT-3′ |
| |
5′-GATGCTCCCTCAGAGTCCTG-3′ |
| Gli3 |
5′-CTTTGCAAGCCAGGAGAAAC-3′ |
| |
5′-TTGTTGGACTGTGTGCCATT-3′ |
| Smo |
5′-CCTTTGGCTTTGTGCTCATTACCTT-3′ |
| |
5′-CGTCACTCTGCCCAGTCAACCT-3′ |
| Mouse and
human | 18s |
5′-CGGCTACCACATCCAAGGAA-3′ |
| |
5′-GCTGGAATTACCGCGGCT-3′ |
Animals and care
The animal protocol of the present study was
approved (SYDL2019010) by the Institutional Animal Care and Use
Committee (IACUC) of The Second Hospital of Shanxi Medical
University (Taiyuan, China) and following the Guidelines of the
Care and Use of Laboratory Animals issued by the Chinese Council on
Animal Research. A total of 92 mice of 2-months age were maintained
in a specific pathogen-free (SPF) 'barrier' facility and housed
under controlled temperature and humidity and alternating 12-h
light and dark cycles. The mice received SPF mouse chow and were
provided with sterile drinking water ad libitum. During the
experiment, the behavior and health of the mice were monitored
daily and recorded every 3 days. The mice were anesthetized with
pentobarbital sodium by an intra-peritoneal injection (35 mg/kg).
Mice were then euthanatized using 100% CO2 when the
following humane endpoints were reached: i) The mice experienced
weight loss (15-20% of their original body weight rapidly
decreased); ii) anorexia (no food at all for 24-36 h); iii)
weakness (unable to eat and drink on their own); and iv) infection
of body organs (penicillin treatment not effective). Following
euthanasia, the death of the mice was confirmed by continuous
non-breathing for 2-3 min, no heartbeat and no blink reflex.
Transgenic mice and surgical induction of
osteoarthritis with ACLT
Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1 transgenic
mice with conditional miR-1 overexpression were generated at the
Department of Orthopedics, Warren Alpert Medical School, Brown
University and were provided by the Department of Orthopedics,
Warren Alpert Medical School, Brown University. Specifically,
Col2a1-Cre-ERT2 mice were mated with
GFPfl/fl-RFP-miR-1 mice to obtain next-generation mice
with the Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1
genotype. To stimulate tamoxifen-induced CreERT2 recombinase and
miR-1 expression in the cartilage, the mice were intraperitoneally
injected with tamoxifen (100 µg/g body weight/day for 5
consecutive days), as described in a previous study (11).
In the present study, 5 of wild-type C57BL/C mice
and 5 transgenic mice were utilized, at 2 months old. These mice
were photographed, X-ray detected, and the knee joints were then
harvested following euthanasia for Safranin O staining to assess
the phenotypic differences between the transgenic and wild-type
mice. Furthermore, 20 of transgenic mice were utilized, at 2 months
old, and were randomly divided into the tamoxifen (intraperitoneal
injection) or control group (intraperitoneal injection of corn oil;
n=10/group). The mice were euthanized at 3 months of age, and the
knee joints were then harvested for RT-qPCR to assess the levels of
miR-1 and Ihh. In addition, another 60 of these transgenic mice
were selected, aged 2 months old, and were randomly divided into
the following 4 groups (n=15/group): Non-tamoxifen + ACLT,
non-tamoxifen + Sham (sham-operated), tamoxifen + ACLT, tamoxifen +
Sham. Tamoxifen was intraperitoneally injected into 2-month-old
mice (100 µg/g body weight/day for 5 consecutive days),
while ACLT was conducted in 3-month-old mice. The surgery was
conducted on the right knees of the mice; 2 months later, these
mice were euthanized and the right hind limbs were harvested
immediately for further analyses.
Radiography
Small-animal X-ray radiography was performed to
detect the changes in the knee joints of these mice at 2 months
after surgery using a small-animal X-ray apparatus (UltraFocus,
Faxitron). In brief, the X-ray films were acquired at the
anteroposterior and lateral positions of the mice under anesthesia
(pentobarbital sodium by an intraperitoneal injection; 35 mg/kg),
and the exposure time and kV were set to the auto setting.
Histological evaluation
At 2 months after the different procedures, the knee
joint of the right hind limb was harvested and fixed in 4% buffered
paraformaldehyde for 48 h and decalcified in a 10%
ethylenediaminetetraacetic acid solution (pH 7.2) for 8 weeks.
These samples underwent routine tissue processing and were then
embedded in paraffin and sectioned in the coronal orientation using
a rotary microtome (Leica Microsystems GmbH) to provide
6-µm-thick coronal sections. Following deparaffinization in
xylene and rehydration in graded ethanol solutions and then in
H2O, the sections were stained with Safranin O/Fast
Green (Sigma-Aldrich; Merck KGaA). The sections were reviewed and
scored for cartilage degradation by 2 pathologists who were without
knowledge of the study groups, according to The International
Association of Osteoarthritis Research (OARSI) grading system
(28). The score ranged from 0 to
6, with a higher score indicating more severe degradation.
Immunohistochemistry
Paraffin-embedded sections of these mouse tissue
specimens were immunostained for the expression of Ihh, MMP-13,
Col10 and Col2a1. The deparaffinized and rehydrated sections were
incubated in 3% H2O2 in phosphate-buffered
saline (PBS) to block any endogenous peroxidase activity at room
temperature for 10 min. The sections were washed with PBS and
digested with 0.1% trypsin at 37°C for 30 min to expose antigens
that may be blocked by tissue fixation and processing.
Subsequently, the sections were incubated with 20% normal serum at
room temperature for 30 min and then with anti-mouse Col2a1 (1:100;
cat. no. ab34712; Abcam), anti-Col10 (1:100; cat. no. BA2023; Wuhan
Boster Biological Technology, Ltd.), anti-Ihh (1:50; cat. no.
bs-6624R; BIOSS), or anti-MMP-13 (1:100; cat. no. ab39012; Abcam)
at 4°C overnight. The sections were then washed 3 times with PBS
and were then incubated with a horseradish peroxidase-conjugated
secondary antibody (Stock solution; cat. no. PV-9001; ZSGB-BIO) at
37°C for 30 min; the color reaction was conducted by brief
incubation of the sections in 3,3′-diaminobenzidine solution. The
immunostained sections were reviewed and photographed under a Leica
DM6B microscope (Leica Microsystems GmbH) for quantitation.
Fluorescence molecular tomography
(FMT)
FMT is a sensitive and quantitative technique that
provides 3-dimensional tissue images in vivo (29,30). The FMT 4000 In Vivo Imaging
System from PerkinElmer, Inc. was utilized to monitor the levels of
MMPs and cathepsins in the mouse knee joints at 2 months after
surgery. The mice received a single dose of MMPSense 645 FAST
Fluorescent Imaging Agent (10 µl, 0.4 nmol; PerkinElmer,
Inc.) and ProSense 750 FAST (10 µl, 0.4 nmol; PerkinElmer)
via tail vein injection at 24 h before imaging. MMPSense is able to
detect MMP-2, MMP-3, MMP-7, MMP-9, MMP-12 and MMP-13, while
ProSense can detect cathepsins B, L, S, K, V and D, as well as
plasmin. The concentrations of MMPs and PRO probes in the mouse
knee joints were calculated by using the region of the interest
method, and the data are expressed as the means ± standard
deviation (SD; n=5 mice per group).
Statistical analysis
The data are expressed at the means ± SD and
statistically analyzed by using SPSS software (version 19.0; IBM
Corporation) and Graph Prism Software (version 5.0; Graph Prism
Software, Inc.). The comparisons of 2 groups of samples were
analyzed by two-way analysis of variance followed by Tukey's test.
A Kruskal-Wallis analysis with post-hoc Dunn's test was performed
to test differences in OARSI grading score between groups. A
P-value ≤0.05 was considered to indicate a statistically
significant difference.
Results
Downregulation of miR-1 and upregulation
of Ihh levels in human osteoarthritic cartilage
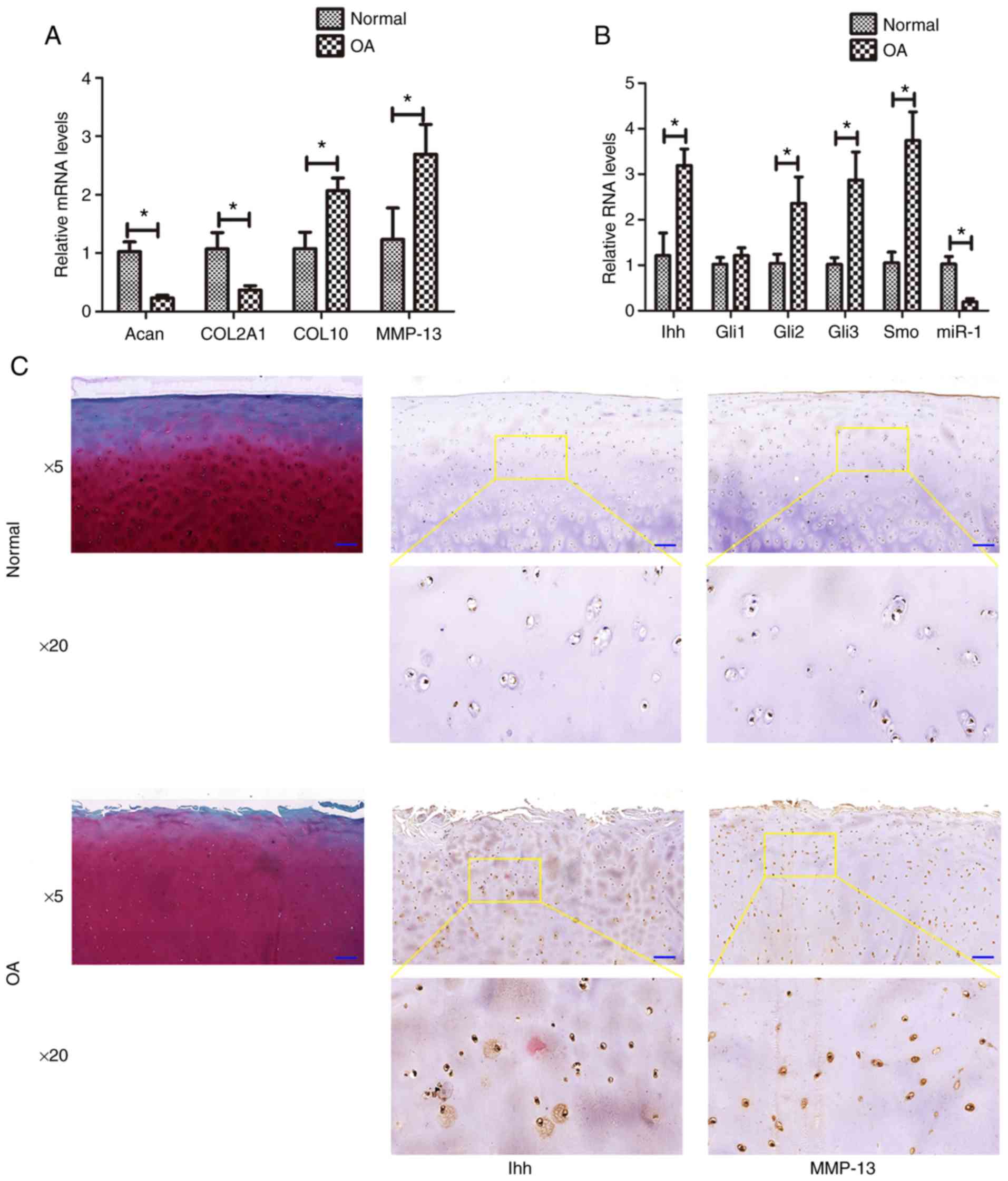
In the present study, the miR-1 and Ihh levels were
first assessed in osteoarthritic cartilage by RT-qPCR. The data
revealed that the normal samples expressed higher levels of miR-1
(hsa-miR-1-5p; P=0.009), COL2A1 (P= 0.016), and Acan (P= 0.025),
whereas they exhibited lower levels of Ihh (P=0.017), Gli2
(P=0.027), Gli3 (P=0.024), Smo (P=0.001), MMP-13 (P=0.005) and
COL10 (P=0.014), compared with those of the osteoarthritis group
(Fig. 1A and B). However no
marked differences were observed in the Gli1 level (P=0.058)
between the normal and diseased samples. The results of
immunohistochemistry revealed that the expression of Ihh and MMP-13
was markedly higher in the osteoarthritis group than in the normal
group (Fig. 1C).
Characteristics of transgenic mice
Subsequently,
Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1 mice were
successfully generated (Fig. 2A)
and the presence of miR-1 and Col2a1-Cre was confirmed by
genotyping (Fig. 2B). In
particular, prior to the tamoxifen injection, the transgenic and
wild-type mice at 2 months of age exhibited no differences in
phenotype, with the mice exhibiting a similar body type, X-ray
appearance and normal Safranin O staining (Fig. 2C). However, following the
tamoxifen injection, the transgenic mice exhibited lower levels of
Ihh (P=0.024), Gli2 (P=0.004), Gli3 (P=0.006), Smo (P=0.002),
MMP-13 (P=0.016) and Col10 (P=0.027); however, they exhibited
elevated levels of Col2a1 (P=0.006) and Acan (P=0.036) expression,
compared with those in the non-tamoxifen-injected mice; of note,
the level of Gli1 (P=0.009) did not differ significantly between
the groups. Moreover, the level of miR-1 (mmu-miR-1a-5p; P=0.004)
expression was evaluated, indicating the successful generation of
these transgenic mice (Fig. 2D and
E).
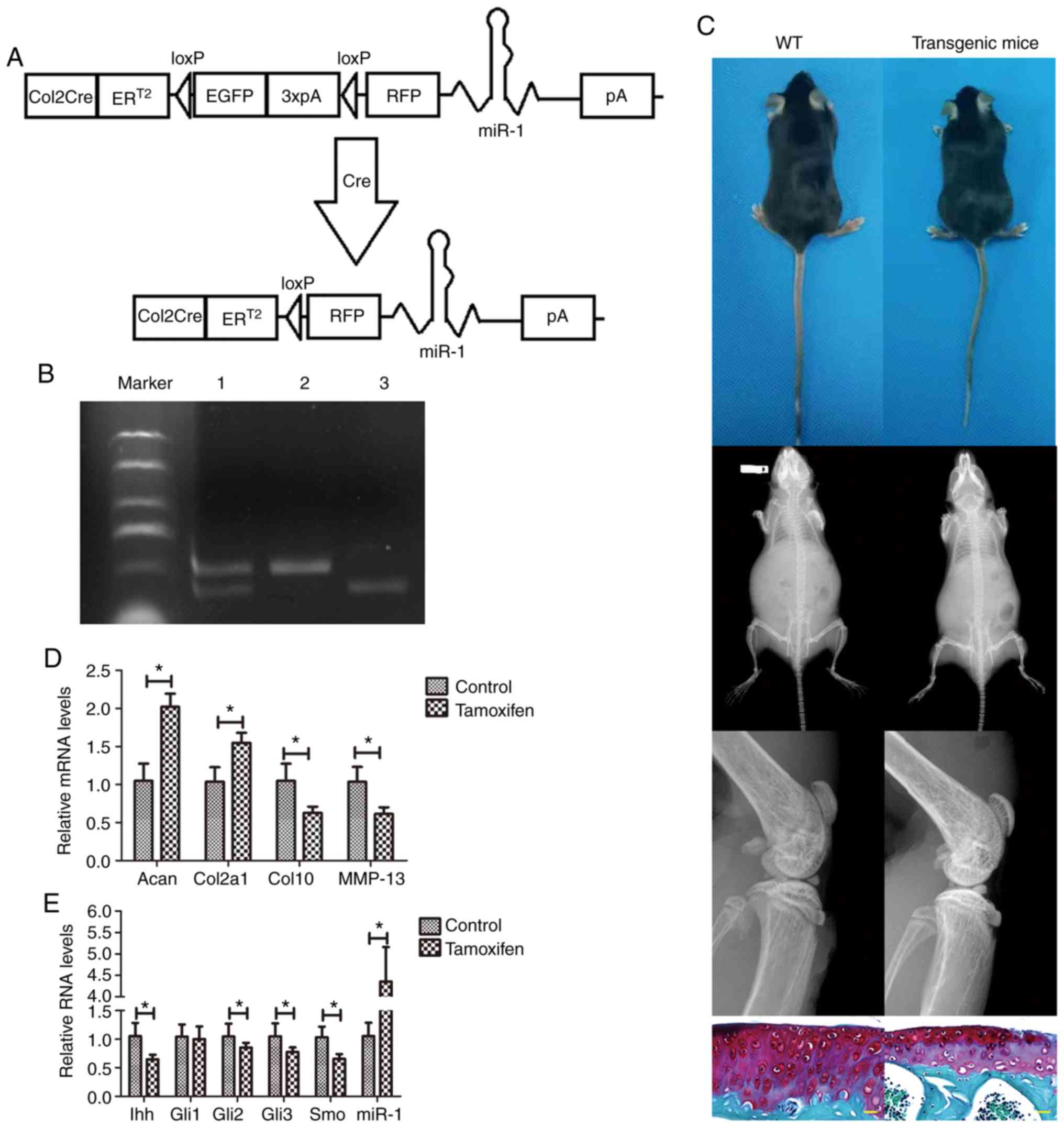 | Figure 2Transgenic mouse model of
osteoarthritis. (A) Illustration of the
Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1 mice. (B)
DNA gel electrophoresis showing the insertion of the loxP
restriction enzyme site in the miR-1 allele confirmed by PCR. Lane
1, miR-1 and Col2a1-Cre (300 bp and 450 bp); lane 2, Col2a1-Cre
(450 bp); lane 3, miR-1 (300 bp). (C) X-ray radiography and
Safranin O staining (yellow scale bar: 20 µm). (D and E)
RT-qPCR. Mouse cartilage samples from tamoxifen-treated mice and
control mice were collected 8 weeks after the surgery and subjected
to RT-qPCR analysis. *P<0.05. WT, wild-type; Acan,
Aggrecan; Col2a1, collagen, type II; Col10, collagen type X; MMP,
matrix metalloproteinase; Ihh, Indian hedgehog; Gli,
glioma-associated oncogene homolog; Smo, smoothened homolog. |
Transgenic mouse model of
osteoarthritis
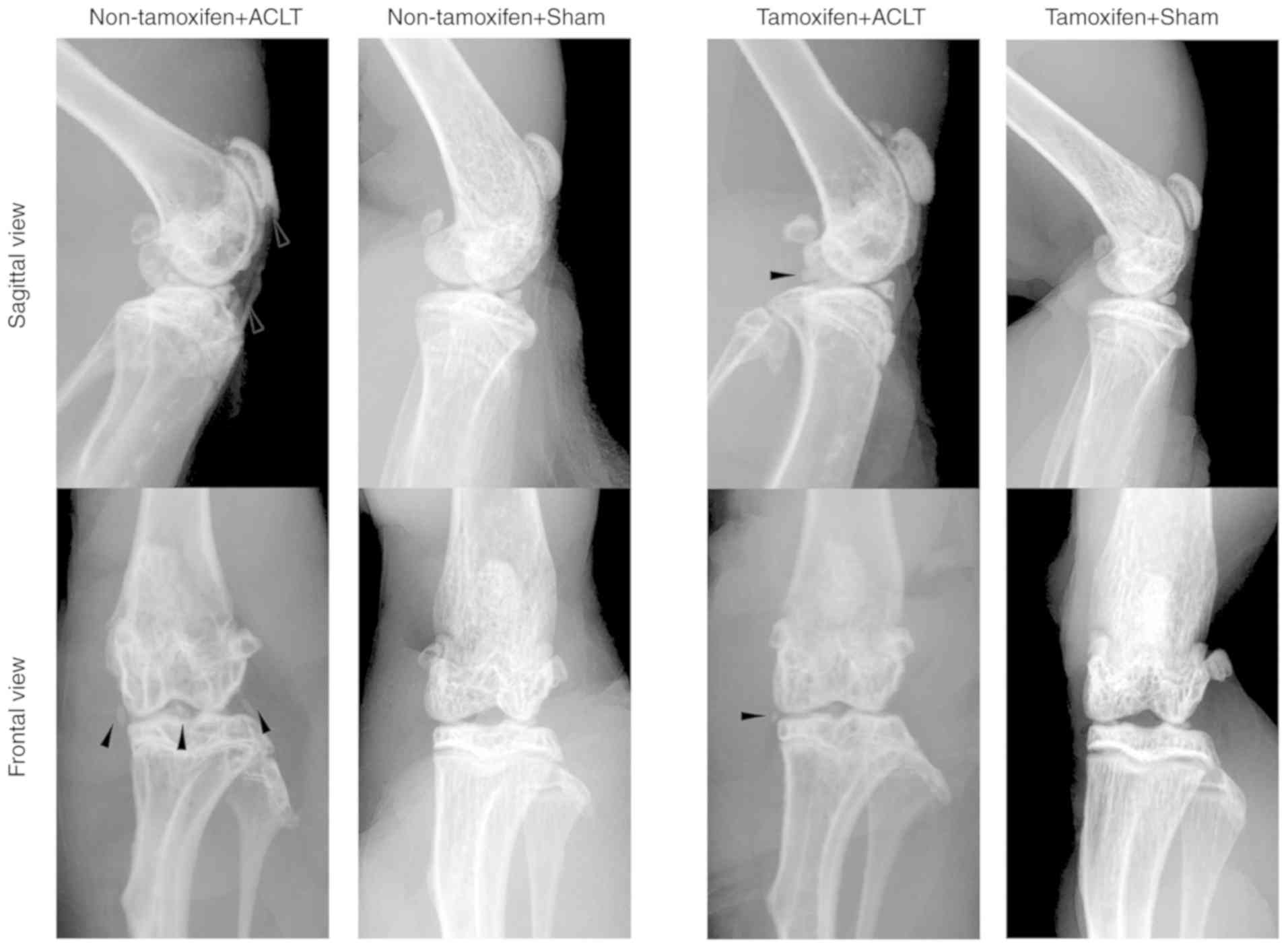
A transgenic mouse model of osteoarthritis was then
established by subjecting mice to ACLT. At 2 months after ACLT,
X-ray examination of the knee joints revealed that the patella,
tibial plateau and tibial intercondylar formed osteophytes in the
non-tamoxifen + ACLT group, and the joint space was narrowed; by
contrast, in the tamoxifen + ACLT group, although there were
osteophytes, there were no evident changes in these joints, and the
joint space was also normal (Fig.
3). In addition, there was no obvious osteophyte formation in
the 2 sham-operated groups.
Subsequently, the articular cartilage was stained
with Safranin O for the visualization of glycosaminoglycans.
Cartilage surface damage, as well as weaker Safranin O staining was
observed in the non-tamoxifen + ACLT group, compared with that in
the tamoxifen + ACLT group (Fig.
4A). Histologically, there was an intact cartilage surface and
less fibrosis in the tamoxifen + ACLT group. The OARSI grading
score was significantly higher in the non-tamoxifen + ACLT group
(2.52±0.24), compared with the tamoxifen + ACLT group (0.57±0.14;
P<0.05; Fig. 4B), indicating
more severe degradation and damage in the non-tamoxifen + ACLT
group.
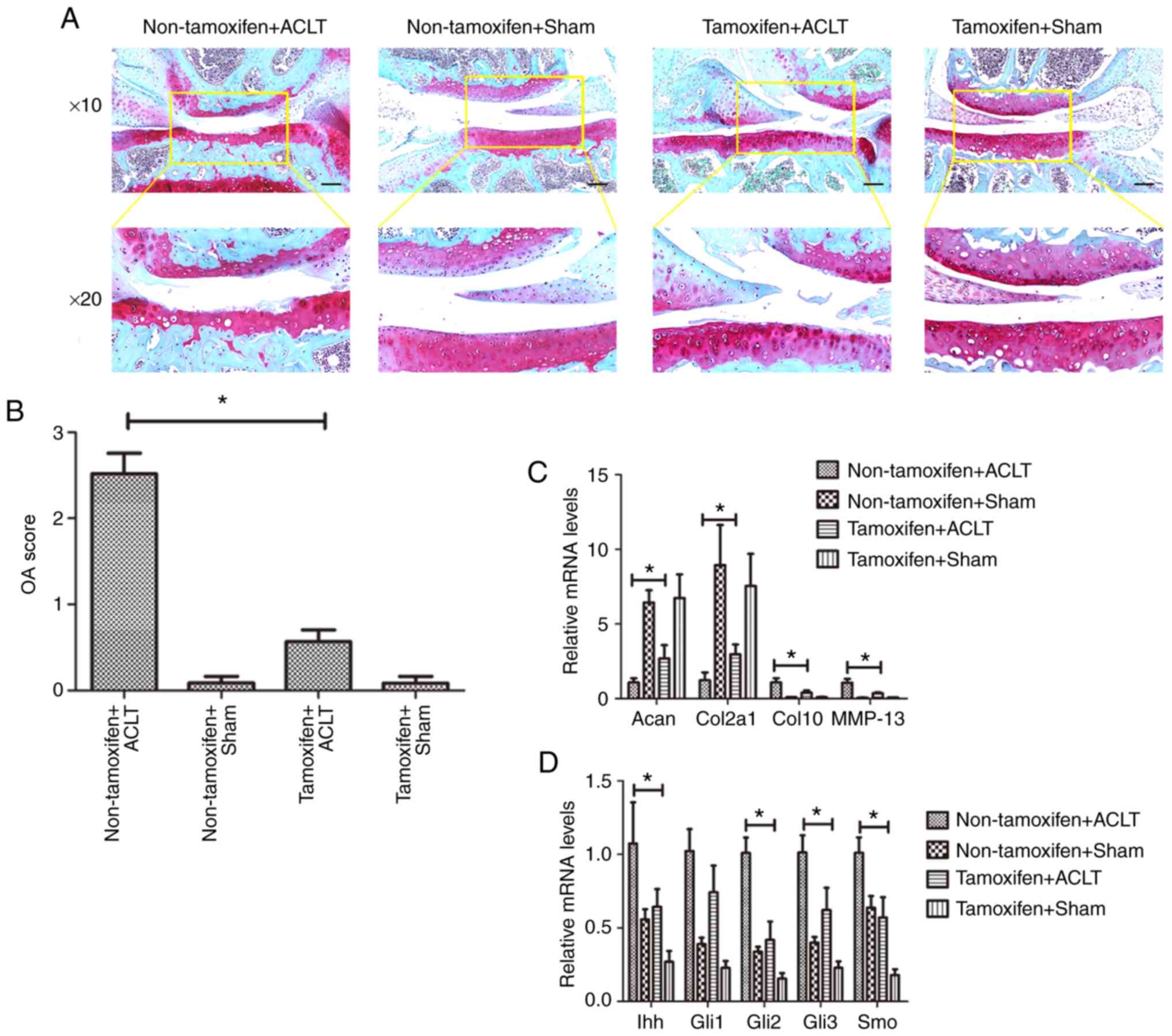 | Figure 4Effect of miR-1 overexpression on the
inhibition of disease progression in the transgenic mouse model of
osteoarthritis. (A) Safranin O staining. Mouse tissues were
collected at 8 weeks after ACLT and subjected to Safranin O
staining (black scale bar: 100 µm). The images the bottom
panels are magnified images of the are in the yellow boxes in the
top panels. (B) OARSI scores. (C and D) RT-qPCR. Mouse tissues were
collected at 8 weeks after ACLT and subjected to RT-qPCR analysis.
*P<0.05. Sham, sham-operated; ACLT, anterior cruciate
ligament transection; Acan, Aggrecan; Col2a1, collagen, type II;
Col10, collagen type X; MMP, matrix metalloproteinase; Ihh, Indian
hedgehog; Gli, glioma-associated oncogene homolog; Smo, smoothened
homolog. |
miR-1 inhibits Ihh expression and
cartilage catabolism, whereas it promotes anabolism in the
transgenic mouse model of osteoarthritis
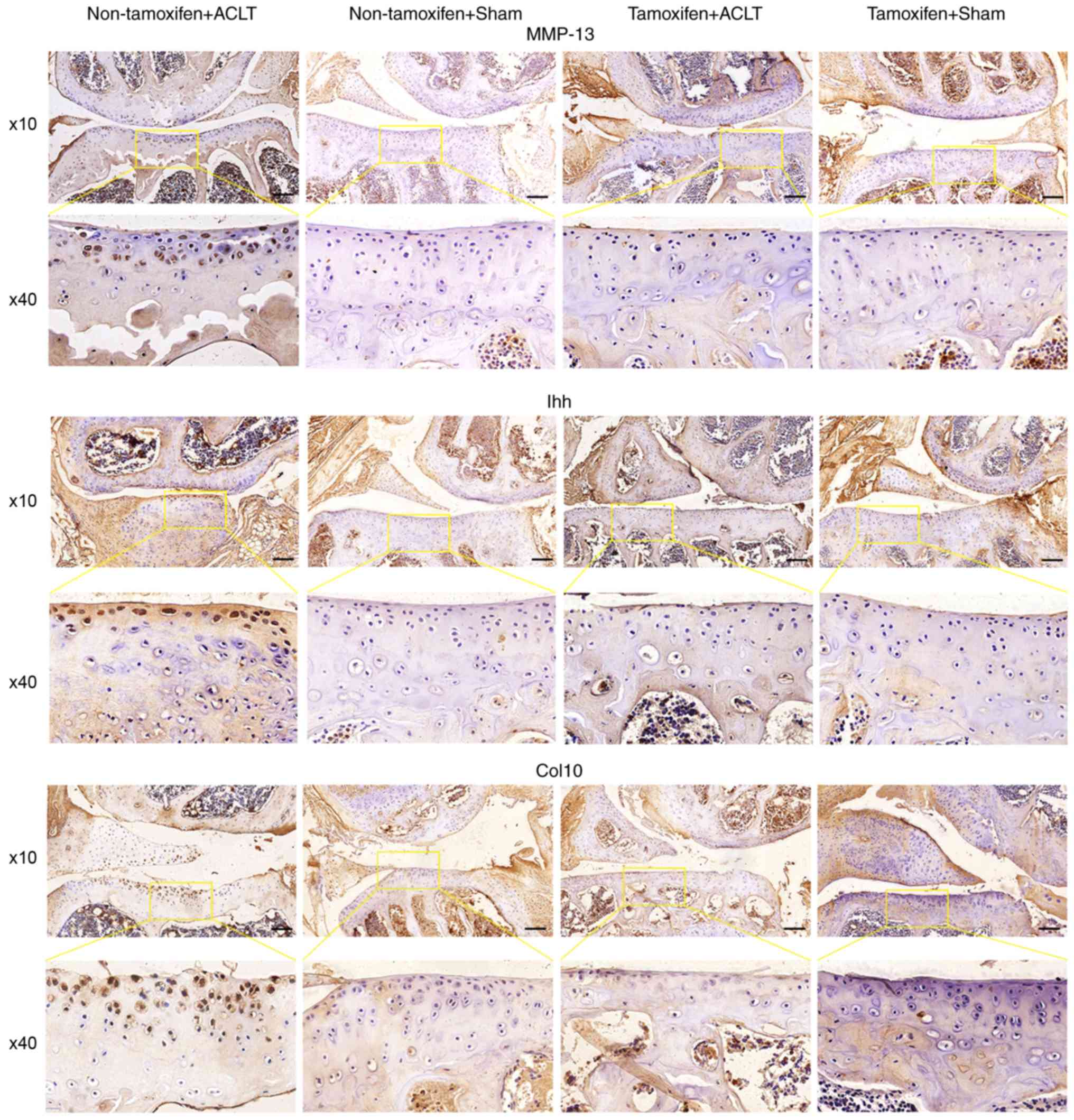
Immunohistochemistry revealed that the protein
expression of Ihh, Col10 and MMP-13 was markedly higher in the
joint tissues of the non-tamoxifen + ACLT group than in those of
the tamoxifen + ACLT group (Fig.
5). However, Col2a1 expression was lower in the joint tissues
of the non-tamoxifen + ACLT group than in those of the tamoxifen +
ACLT group (Fig. S1).
Furthermore, the RT-qPCR data confirmed the
immunohistochemical data (Fig. 4C and
D), indicating that the joint tissues of the tamoxifen + ACLT
group expressed lower levels of Ihh (P=0.026), Gli2 (P=0.028), Gli3
(P=0.004), Smo (P=0.001), MMP-13 (P=0.010) and Col10 (P=0.004);
however, they expressed higher levels of Col2a1 (P=0.012) and Acan
(P=0.035), compared with those of the non-tamoxifen + ACLT group;
the Gli1 (P=0.061) level did not differ significantly between the 2
groups.
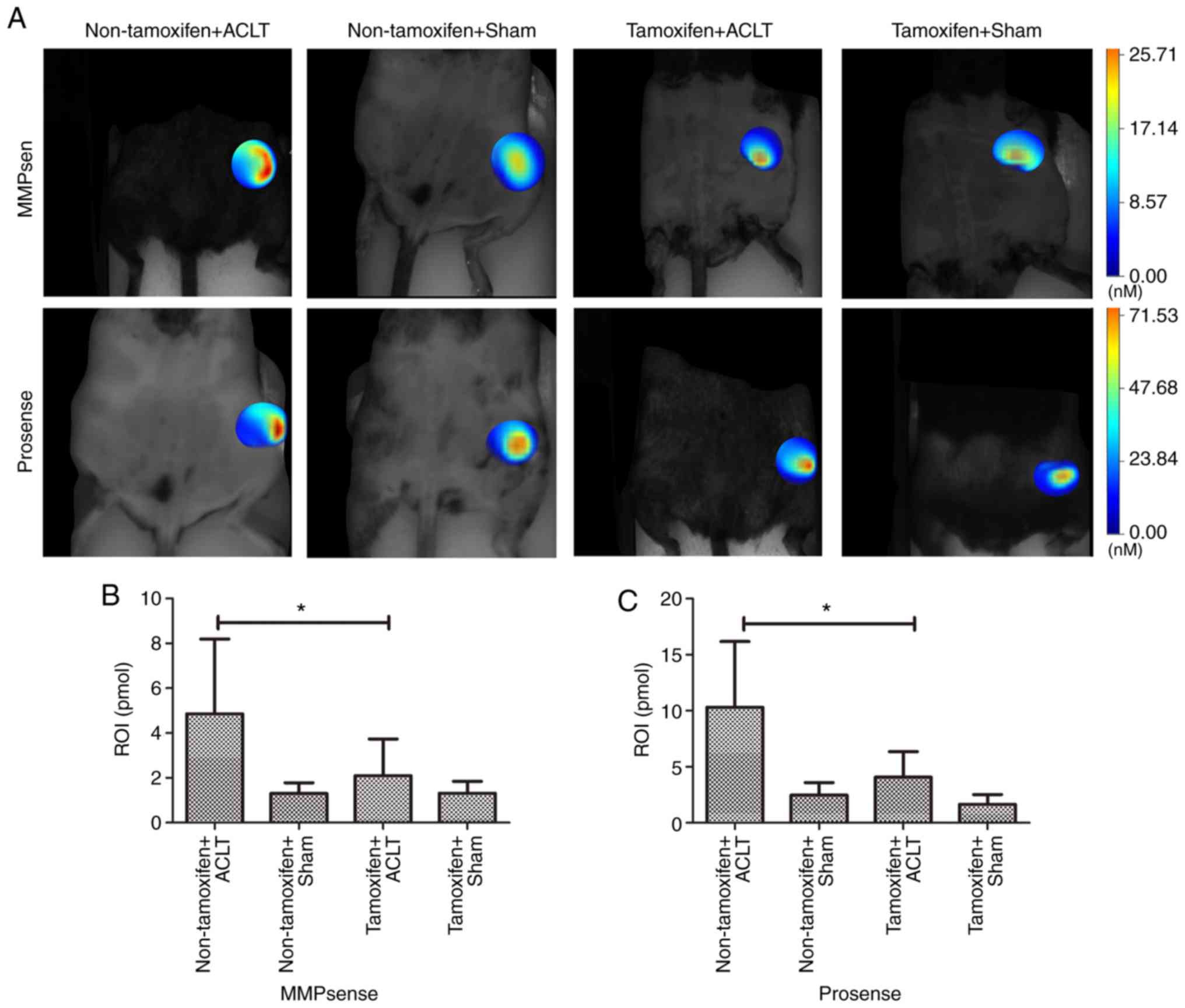
In addition, FMT with the appropriate probes was
used to monitor the levels of MMPs and cathepsins in the knees of
alive mice (Fig. 6). The data
revealed that the MMP level was significantly lower in the
tamoxifen + ACLT group (3.74±0.45 pmol) than in the non-tamoxifen +
ACLT group (8.20±1.51 pmol; P<0.001). Moreover, the cathepsin
level was also significantly lower (6.36±1.80 pmol) in the
tamoxifen + ACLT group than in the non-tamoxifen + ACLT group
(16.18±4.42 pmol; P=0.016), further confirming the
immunohistochemistry and RT-qPCR results.
Discussion
miRNAs post-transcriptionally regulate gene
expression and control a wide range of biological processes in
cells and tissues (31). miR-1
plays a role in myocyte proliferation and differentiation (32), as well as in the growth of growth
plate chondrocytes during development (14). In the present study, it was found
that miR-1 expression was low in human osteoarthritic joint tissues
and that miR-1 induction was able to suppress osteoarthritis in a
Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1 mouse
model of osteoarthritis. Furthermore, Ihh is a secreted protein
that is expressed in pre-hypertrophic chondrocytes (33), indicating that the upregulation of
Ihh can promote the hypertrophic phenotype and induce the
expression of typical hypertrophic markers, such as COL10 and
MMP-13 (10). Thus, upregulated
Ihh signaling can contribute to the development of osteoarthritis
(34-37). In the present study, it was
revealed that the miR-1-induced inhibition of Ihh (data not shown)
signaling was able to suppress the development of osteoarthritis in
a Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1 mouse
model of osteoarthritis. Specifically, it was found that Ihh
expression was low in normal human articular cartilage, whereas it
was increased in osteoarthritis-damaged cartilage. Ihh expression
was associated with the severity of osteoarthritis, in terms of
cartilage damage, thus confirming previous observations and
analyses of human samples (10,38). The expression of miR-1 was
inversely proportional to that of Ihh in human
osteoarthritis-damaged articular cartilage. Therefore, the
development of therapies that target Ihh may be a novel approach
for the treatment or prevention of osteoarthritis.
Indeed, the transgenic mouse model of osteoarthritis
is a useful tool with which to assess the effects of miR-1 on the
regulation of osteoarthritis development and treatment. In the
current study,
Col2a1-Cre-ERT2/GFPfl/fl-RFP-miR-1 mice were
obtained and generated and a transgenic mouse model of
osteoarthritis was then established following surgical ACLT. Mouse
cartilage was then resected for the assessment of osteoarthritis by
radiography, histology, gene expression and Safranin O staining.
The X-ray examination of the knee joints of mice at 2 months
following ACLT revealed that the patella, tibial plateau and tibial
intercondylar formed osteophytes in osteo-arthritic mice and that
the joint space was narrowed; whereas ACLT mice with miR-1
overexpression had no obvious changes in these joints or the joint
space. Safranin O staining of the articular cartilage revealed more
cartilage surface damage in the osteoarthritic mice compared with
that in the ACLT mice with miR-1 overexpression. Histologically,
the mice subjected to ACLT with miR-1 overexpression had an intact
cartilage surface with less fibrosis. These data clearly revealed
the role of miR-1 in the control of osteoarthritis development in
these mice.
The data of the present study also demonstrated that
the levels of Ihh and its downstream genes (such as Gli2, Gli3, and
Smo), MMP-13 and Col10 were lower in the miR-1-overexpressing
cartilage, whereas the levels of Col2a1 and Acan were higher. These
findings were further confirmed by the immunohistochemistry and FMT
data. FMT is an advanced, sensitive bioimaging method that provides
noninvasive, in-depth tissue imaging and quantification of in
vivo biological targets (10). In the present study, in
vivo FMT was performed and it was found that the expression of
MMPs and cathepsins was downregulated in miR-1-overexpressing mice
subjected to ACLT compared to that of osteoarthritic mice. These
findings were consistent with the immunohistochemistry and RT-qPCR
results.
However, the present study does have some
limitations; for example, the reason why there was no change in
Gli1 expression between the normal and osteoarthritic human samples
or between the tissues from osteoarthritic and miR-1-overexpressed
ACLT mice cannot be explained. Moreover, the direct miR-1 binding
or regulation of Ihh expression was not confirmed, although in
another study by the authors, bioinformatics analysis of miR-1
target genes using TargetScan (http://www.targetscan.org) and miRanda (http://www.microrna.org) revealed that there are two
specific binding sites (miRBas Accession no. #MIMAT0000416) between
miR-1 and the 3′-untranslated region of Ihh (unpublished data).
Further research using a luciferase reporter assay is required to
validate their direct interaction.
In conclusion, the present study revealed that miR-1
plays an important role in protection of the articular cartilage
from osteoarthritis-induced degeneration and is accompanied by the
downregulation of Ihh expression. Future studies are required to
further investigate miR-1 as a novel target for the prevention and
treatment of osteoarthritis.
Supplementary Data
Funding
The present study was funded by the National Science
Foundation for Young Scientists of China (grant no. 81601949), a
Key Research and Development Project of Shanxi Province (grant no.
201803D421066), a Returned Overseas Students Research Funding
Project of Shanxi Province (grant no. 2016-118), an Applied Basic
Research Project of Shanxi Province (grant no. 201601D102067), and
a Soft Science Research General Project of Shanxi Province (grant
no. 2017041083-3).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
PengcuiL, LW and XC conceived and designed the
research. XC and XG drafted the manuscript. XC, XG, TC, YG, SL and
PenghuaL performed the experiments. XC, XG, DS, CW and BL analyzed
the data. XC, DS, and PengcuiL edited the article. All authors read
and approved the final manuscript. All authors agree to be held
accountable for all aspects of the study.
Ethics approval and consent to
participate
The research protocol was approved by the Clinical
Research Ethics Committee of Shanxi Medical University (approval
no. 2019YX260), and the animal model experimental program was
approved by the Animal Experimental Ethics Committee of Shanxi
Medical University (approval no. SYDL2019010). Each participant was
required to obtain written informed consent before sample
collection.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgments
Not applicable.
References
|
1
|
Glyn-Jones S, Palmer AJ, Agricola R, Price
AJ, Vincent TL, Weinans H and Carr AJ: Osteoarthritis. Lancet.
386:376–387. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Miller ME, Rejeski WJ, Messier SP and
Loeser RF: Modifiers of change in physical functioning in older
adults with knee pain: The observational arthritis study in seniors
(OASIS). Arthritis Rheum. 45:331–339. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Arden N and Nevitt MC: Osteoarthritis:
Epidemiology. Best Pract Res Clin Rheumatol. 20:3–25. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Berenbaum F: Osteoarthritis as an
inflammatory disease (osteoarthritis is not osteoarthrosis!).
Osteoarthritis Cartilage. 21:16–21. 2013. View Article : Google Scholar
|
|
5
|
Araldi E and Schipani E: MicroRNA-140 and
the silencing of osteoarthritis. Genes Dev. 24:1075–1080. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Miyaki S, Sato T, Inoue A, Otsuki S, Ito
Y, Yokoyama S, Kato Y, Takemoto F, Nakasa T, Yamashita S, et al:
MicroRNA-140 plays dual roles in both cartilage development and
homeostasis. Genes Dev. 24:1173–1185. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhang X, Wang C, Zhao J, Xu J, Geng Y, Dai
L, Huang Y, Fu SC, Dai K and Zhang X: MiR-146a facilitates
osteoarthritis by regulating cartilage homeostasis via targeting
Camk2d and Ppp3r2. Cell Death Dis. 8:e27342017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Guan YJ, Li J, Yang X, Du S, Ding J, Gao
Y, Zhang Y, Yang K and Chen Q: Evidence that miR-146a attenuates
aging- and trauma-induced osteoarthritis by inhibiting Notch1,
IL-6, and IL-1 mediated catabolism. Aging Cell. 17:e127522018.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li YP, Wei XC, Li PC, Chen CW, Wang XH,
Jiao Q, Wang DM, Wei FY, Zhang JZ and Wei L: The role of miRNAs in
cartilage homeostasis. Curr Genomics. 16:393–404. 2015. View Article : Google Scholar
|
|
10
|
Bagga S, Bracht J, Hunter S, Massirer K,
Holtz J, Eachus R and Pasquinelli AE: Regulation by let-7 and lin-4
miRNAs results in target mRNA degradation. Cell. 122:553–563. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Eulalio A, Rehwinkel J, Stricker M,
Huntzinger E, Yang SF, Doerks T, Dorner S, Bork P, Boutros M and
Izaurralde E: Target-specific requirements for enhancers of
decapping in miRNA-mediated gene silencing. Genes Dev.
21:2558–2570. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Valencia-Sanchez MA, Liu J, Hannon GJ and
Parker R: Control of translation and mRNA degradation by miRNAs and
siRNAs. Genes Dev. 20:515–524. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yuan X, Guo Y, Chen D, Luo Y, Chen D, Miao
J and Chen Y: Long non-coding RNA MALAT1 functions as miR-1 sponge
to regulate Connexin 43-mediated ossification of the posterior
longitudinal ligament. Bone. 127:305–314. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Li P, Wei X, Guan Y, Chen Q, Zhao T, Sun C
and Wei L: MicroRNA-1 regulates chondrocyte phenotype by repressing
histone deacetylase 4 during growth plate development. FASEB J.
28:3930–3941. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhou J, Wei X and Wei L: Indian Hedgehog,
a critical modulator in osteoarthritis, could be a potential
therapeutic target for attenuating cartilage degeneration disease.
Connect Tissue Res. 55:257–261. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lin AC, Seeto BL, Bartoszko JM, Khoury MA,
Whetstone H, Ho L, Hsu C, Ali SA and Alman BA: Modulating hedgehog
signaling can attenuate the severity of osteoarthritis. Nat Med.
15:1421–1425. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kirsch T, Swoboda B and Nah H: Activation
of annexin II and V expression, terminal differentiation,
mineralization and apoptosis in human osteoarthritic cartilage.
Osteoarthritis Cartilage. 8:294–302. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Pfander D, Swoboda B and Kirsch T:
Expression of early and late differentiation markers (proliferating
cell nuclear antigen, syndecan-3, annexin VI, and alkaline
phosphatase) by human osteoarthritic chondrocytes. Am J Pathol.
159:1777–1783. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhang C, Wei X, Chen C, Cao K, Li Y, Jiao
Q, Ding J, Zhou J, Fleming BC, Chen Q, et al: Indian hedgehog in
synovial fluid is a novel marker for early cartilage lesions in
human knee joint. Int J Mol Sci. 15:7250–7265. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Orfanidou T, Iliopoulos D, Malizos KN and
Tsezou A: Involvement of SOX-9 and FGF-23 in RUNX-2 regulation in
osteoarthritic chondrocytes. J Cell Mol Med. 13:3186–3194. 2009.
View Article : Google Scholar
|
|
21
|
Wei F, Zhou J, Wei X, Zhang J, Fleming BC,
Terek R, Pei M, Chen Q, Liu T and Wei L: Activation of Indian
hedgehog promotes chondrocyte hypertrophy and upregulation of
MMP-13 in human osteoarthritic cartilage. Osteoarthritis Cartilage.
20:755–763. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhou J, Chen Q, Lanske B, Fleming BC,
Terek R, Wei X, Zhang G, Wang S, Li K and Wei L: Disrupting the
Indian hedgehog signaling pathway in vivo attenuates surgically
induced osteoarthritis progression in Col2a1-CreERT2; Ihhfl/fl
mice. Arthritis Res Ther. 16:R112014. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Guo L, Wei X, Zhang Z, Wang X, Wang C, Li
P, Wang C and Wei L: Ipriflavone attenuates the degeneration of
cartilage by blocking the Indian hedgehog pathway. Arthritis Res
Ther. 21:1092019. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lorenz J and Grassel S: Experimental
osteoarthritis models in mice. Methods Mol Biol. 1194:401–419.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Jeon OH, Kim C, Laberge RM, Demaria M,
Rathod S, Vasserot AP, Chung JW, Kim DH, Poon Y, David N, et al:
Local clearance of senescent cells attenuates the development of
post-traumatic osteoarthritis and creates a pro-regenerative
environment. Nat Med. 23:775–781. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Altman R, Asch E, Bloch D, Bole G,
Borenstein D, Brandt K, Christy W, Cooke TD, Greenwald R, Hochberg
M, et al: Development of criteria for the classification and
reporting of osteoarthritis. Classification of osteoarthritis of
the knee. Diagnostic and therapeutic criteria committee of the
American rheumatism association. Arthritis Rheum. 29:1039–1049.
1986. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
28
|
Glasson SS, Chambers MG, Van Den Berg WB
and Little CB: The OARSI histopathology initiative-recommendations
for histological assessments of osteoarthritis in the mouse.
Osteoarthritis Cartilage. 18(Suppl 3): S17–S23. 2010. View Article : Google Scholar
|
|
29
|
Ntziachristos V, Bremer C and Weissleder
R: Fluorescence imaging with near-infrared light: New technological
advances that enable in vivo molecular imaging. Eur Radiol.
13:195–208. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Peterson JD, Labranche TP, Vasquez KO,
Kossodo S, Melton M, Rader R, Listello JT, Abrams MA and Misko TP:
Optical tomographic imaging discriminates between disease-modifying
anti-rheumatic drug (DMARD) and non-DMARD efficacy in collagen
antibody-induced arthritis. Arthritis Res Ther. 12:R1052010.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Selbach M, Schwanhausser B, Thierfelder N,
Fang Z, Khanin R and Rajewsky N: Widespread changes in protein
synthesis induced by microRNAs. Nature. 455:58–63. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Chen JF, Mandel EM, Thomson JM, Wu Q,
Callis TE, Hammond SM, Conlon FL and Wang DZ: The role of
microRNA-1 and microRNA-133 in skeletal muscle proliferation and
differentiation. Nat Genet. 38:228–233. 2006. View Article : Google Scholar
|
|
33
|
Young B, Minugh-Purvis N, Shimo T,
St-Jacques B, Iwamoto M, Enomoto-Iwamoto M, Koyama E and Pacifici
M: Indian and sonic hedgehogs regulate synchondrosis growth plate
and cranial base development and function. Dev Biol. 299:272–282.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Aigner T, Reichenberger E, Bertling W,
Kirsch T, Stoss H and von der Mark K: Type X collagen expression in
osteoarthritic and rheumatoid articular cartilage. Virchows Arch B
Cell Pathol Incl Mol Pathol. 63:205–211. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Clements DN, Carter SD, Innes JF, Ollier
WE and Day PJ: Analysis of normal and osteoarthritic canine
cartilage mRNA expression by quantitative polymerase chain
reaction. Arthritis Res Ther. 8:R1582006. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Little CB, Barai A, Burkhardt D, Smith SM,
Fosang AJ, Werb Z, Shah M and Thompson EW: Matrix metalloproteinase
13-deficient mice are resistant to osteoarthritic cartilage erosion
but not chondrocyte hypertrophy or osteophyte development.
Arthritis Rheum. 60:3723–3733. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Aigner T, Dietz U, Stoss H and von der
Mark K: Differential expression of collagen types I, II, III, and X
in human osteophytes. Lab Invest. 73:236–243. 1995.PubMed/NCBI
|
|
38
|
Tchetina EV, Squires G and Poole AR:
Increased type II collagen degradation and very early focal
cartilage degeneration is associated with upregulation of
chondrocyte differentiation related genes in early human articular
cartilage lesions. J Rheumatol. 32:876–886. 2005.PubMed/NCBI
|