Introduction
Osteosarcoma is a common primary malignant tumor
derived from the musculoskeletal system with high rates of
recurrence and metastasis. Therapeutic options have significantly
improved over time, especially in terms of chemotherapy and
limb-sparing surgery. The 5-year survival rate of patients with
osteosarcoma increased from 20 to 70-80% after the introduction of
neo-adjuvant chemotherapy (1).
However, to the best of our knowledge, further improvements to the
5-year survival rate have not been reported for nearly 30 years,
mainly due to distant metastasis and recurrence. Conventional
treatments used for advanced osteosarcoma, such as surgery,
chemotherapy, radiotherapy or combinations of these, have had
little effect on the long-term survival of patients with this
disease (2). New drugs are needed
in order to improve the outcomes of advanced osteosarcoma.
Molecular chaperones are a class of proteins that
mediate the assembly or disassembly of other macromolecular
structures through catalytic or non-catalytic routes. Heat shock
proteins (HSPs) are the major type of chaperones, which are
expressed in response to elevated temperatures or other cellular
stresses and prevent or correct damage caused by heat-related
misfolding (3). HSPs,
particularly HSP70 and HSP90, play an important role in
cancer-related molecular events by controlling kinases, signaling
proteins, cell-cycle regulators and steroid receptors (4). Stress-induced phosphoprotein 1
(STIP1/STI1 or p60), also called HSP70/HSP90 organizing protein
(HOP), is a 66-kDa protein and the most extensively studied
multi-domain co-chaperone. STIP1, which mediates the activity of
the HSP70/HSP90 chaperone heterocomplex, contains a nuclear
localization signal regulated by cell cycle kinase phosphorylation,
and undergoes nuclear-cytoplasmic shuttling considered to be
influenced by cell cycle status and cell cycle kinases (5,6).
Compared with normal cells, cancer cells can sustain higher
metabolic rates through multiple intrinsic mechanisms and signal
transduction pathways. The role of STIP1 in cancer remains unclear,
but there is increasing evidence that STIP1 may regulate multiple
cancer-related biological processes, including tumor formation,
metastasis and drug resistance in many types of cancer cells such
as pancreatic cancer and gastric cancer (7-9).
A possible mechanism underlying tumor formation,
multidrug resistance, recurrence and metastasis is suggested by the
cancer stem/stem-like cells (CSCs) theory, according to which a
sub-population of cancer cells with self-renewal and
differentiation capacities drive tumor proliferation (10). Previous studies have demonstrated
that there is a group of CSCs among various cancer cell types,
including melanoma, hematopoietic, brain, breast, pancreatic, colon
and prostate cancer (10,11). Gibbs et al (12) isolated and identified CSCs from
osteosarcoma cells obtained from primary biopsies by using the
cell-sphere formation method in serum-free suspension medium
containing growth factors. Human osteosarcoma cell lines contain
CD133-positive (CD133+) CSC populations, which exhibit
increased production of ATP and lactate dehydrogenase under
hypoxia, and have high chemoresistance and metastatic potential
(13-16). Li et al (17) isolated CD133+
osteosarcoma cells, which exhibited similar properties to CSCs,
from the MG63 cell line by magnetic activated cell sorting
(MACS).
In the present study, CD133+ CSCs were
isolated and identified from the human osteosarcoma MG63 cell line.
Furthermore, RNA interference technology was used to knock down
STIP1 expression in CSCs isolated from the MG63 cells, and to
examine the molecular mechanisms involved in osteosarcoma CSC
invasion and migration.
Materials and methods
Cell lines and culture
The human osteosarcoma MG63 cell line was purchased
from the China Center for Type Culture Collection at Wuhan
University. Cells were cultured in DMEM/F12 (HyClone; GE Healthcare
Life Sciences) supplemented with 10% FBS (Gibco; Thermo Fisher
Scientific, Inc.) in a 5% CO2 humidified incubator at
37°C. Cells were digested with 2.5 g/l trypsin (Gibco; Thermo
Fisher Scientific, Inc.) for 3-5 min at 37°C and passaged once
every 5-6 days.
Isolation of CSCs from the osteosarcoma
MG63 cell line
The process of osteosarcoma CSC isolation was
conducted as reported previously (12,13,18,19). MG63 cells were harvested by
trypsinization, detached, centrifuged for 5 min at 1,200 × g at
room temperature and resuspended in 1 ml PBS (cat. no. PYG0021;
Wuhan Boster Biological Technology, Ltd.) containing 1% BSA (cat.
no. AR1006; Wuhan Boster Biological Technology, Ltd.) at
1×107 cells per 500 µl, then incubated with an
anti-CD133 antibody (cat. no. sc-30220; 1:200; Santa Cruz
Biotechnology, Inc.) at 37°C for 15 min. After two washes with PBS
containing 1% BSA, the cells were labeled with micro-beads
(Miltenyi Biotec GmbH) at 37°C for 15 min. The cells were
resuspended and poured into a separation column (LS Columns;
Miltenyi Biotec GmbH), after which CD133+ cells were
retained in the column and flushed out with 2 ml PBS containing 1%
BSA with the aid of a plunger.
Subsequently, the CD133+ cells were
plated in ultra-low-attachment culture plates (Corning Life
Sciences) in serum-free medium, consisting of DMEM/F12, 20
µg/l human epidermal growth factor (PeproTech, Inc.), 1X N2
supplement (Invitrogen; Thermo Fisher Scientific, Inc.), 2 mmol/l
L-glutamine, 4 U/l insulin (both Sigma-Aldrich; Merck KGaA),
1×105 U/l penicillin and 1×105 U/l
streptomycin. The pH value was adjusted to 7.2-7.5. Additional
fresh serum-free medium was added once every other day. After
CD133+ CSC spheres that contained >50 cells formed,
the CD133+ CSCs were collected and dissociated by
trypsinization, then passaged in serum-free medium at a ratio of
1:2-1:4.
Flow cytometry analysis
CD133+ CSCs were resuspended to
single-cell suspensions, washed twice with PBS, and incubated with
antibodies against CD44 (cat. no. sc-65265; 1:100; Santa Cruz
Biotechnology, Inc.), CD105 (cat. no. sc-53999; 1:100; Santa Cruz
Biotechnology, Inc.) and CD133 (1:100) in 0.5% BSA and PBS for 60
min at 4°C. After washing, samples were incubated with a secondary
antibody (1:150; Goat Anti-Mouse IgG H+L Secondary Antibody-FITC
Conjugate for CD44 and CD105; cat. no. BA1101; Goat Anti-Rabbit IgG
H+L Secondary Antibody-FITC Conjugate for CD133; cat. no. BA1105,
Wuhan Boster Biological Technology, Ltd.), in 0.5% BSA and PBS for
15 min at 37°C. Finally, the samples were analyzed by flow
cytometry using FACSort with CellQuest, ModFit LT for Mac V2.0
software (BD Biosciences) after washing twice. Each experiment was
performed in triplicate.
Transient transfections with small
interfering RNA (siRNA)
CD133+ CSCs were seeded into 6-well
plates at 4×104 cells/well. Three pre-designed siRNAs
(Shanghai GenePharma Co., Ltd.) were chosen for the knockdown of
STIP1 (STIP1-homo-643, STIP1-homo-941, STIP1-homo-1315) in addition
to the negative control (NC) siRNA (Table I). After overnight culture, the
medium was changed and the cells were transfected with siRNA or NC,
each at a final volume of 5 µl Lipofectamine®
2000 (Invitrogen; Thermo Fisher Scientific, Inc.) and final
concentration of 250 pmol siRNA in a total volume of 500 µl
Opti-MEM (Gibco; Thermo Fisher Scientific, Inc.). All procedures
were performed according to the Lipofectamine® 2000
manufacturer's protocol. The transfection efficiency was detected
by fluorescence microscopy (Olympus BX53) after 24-h culture. Cells
cultured for 48-72 h were trypsinized and used in subsequent
experiments.
 | Table ISequences of the siRNAs used in the
present study. |
Table I
Sequences of the siRNAs used in the
present study.
| siRNA | Sequence
(5′-3′) |
|---|
| STIP1-homo-643 | Sense:
GGGCACGAAACUACAAGAUTT |
| Antisense:
AUCUUGUAGUUUCGUGCCCTT |
| STIP1-homo-941 | Sense:
GCAGCGGUAUACUUUGAAATT |
| Antisense:
UUUCAAAGUAUACCGCUGCTT |
|
STIP1-homo-1315 | Sense:
CCCGAAAGAUGCCAAAUUATT |
| Antisense:
UAAUUUGGCAUCUUUCGGGTT |
| Negative
control | Sense:
UUCUCCGAACGUGUCACGUTT |
| Antisense:
ACGUGACACGUUCGGAGAATT |
Reverse transcription-quantitative PCR
(RT-qPCR) analysis
After the transfection of siRNA for 72 h, total RNA
was isolated from the cells using TRIzol™ (Invitrogen; Thermo
Fisher Scientific, Inc.) according to the manufacturer's protocol.
The RNA was quantified by spectrophotometry at an optical density
of 260 nm. First-strand cDNA was generated from 2 µg total
RNA using a cDNA Synthesis kit (Invitrogen; Thermo Fisher
Scientific, Inc.), according to the manufacturer's protocol, in a
total reaction volume of 2 µl. The primer sequences were
obtained from qPrimerDepot (http://primerdepot.nci.nih.gov/) and PrimerBank
(http://pga.mgh.harvard.edu/primerbank/), purchased
from Invitrogen; Thermo Fisher Scientific, Inc., and are listed in
Table II. GAPDH was used as the
reference gene. qPCR was carried out using the SYBR Green qPCR
Master mix (2X; Roche Diagnostics) and an iQ Real-Time system
(Bio-Rad Laboratories, Inc.) equipped with a 96-well optical
reaction plate in a total reaction volume of 20 µl. All qPCR
reactions were run in triplicate, and the mean threshold cycle
value was used for the determination of mRNA levels. The
thermocycling conditions were as follows: Initial denaturation step
at 95°C for 5 min, followed by 40 cycles at 94°C for 20 sec, 58°C
for 20 sec and 72°C for 20 sec, and a final extension step at 72°C
for 5 min. At the end of the reactions, the baseline and threshold
values for the amplifications were set using iCycler5 software
version 3.1 (Bio-Rad Laboratories, Inc.), and the relative
expression of mRNA was calculated using the comparative
2−ΔΔCq method (20).
 | Table IISequences of primers used in the
present study. |
Table II
Sequences of primers used in the
present study.
| Gene | Primer sequence
(5′-3′) | Product length,
bp |
|---|
| STIP1 | F:
AACGCGGAGCGGACGGATTC | 101 |
| R:
ACGCTCAGGGCCTTGTTGCC | |
| GAPDH | F:
CAATGACCCCTTCATTGACC | 106 |
| R:
GACAAGCTTCCCGTTCTCAG | |
Cell proliferation assays
CD133+ CSCs were seeded into 96-well
plates at a density of 5,000 cells/well in 100 µl DMEM/F12
supplemented with 10% FBS. Cell proliferation was detected 24 and
48 h after transfection, using the Cell Counting Kit-8 (CCK-8;
Beyotime Institute of Biotechnology). A total of 10 µl CCK-8
reagent was added to each well on the 96-well plate, which was then
placed in a CO2 incubator for 2 h to react. Following
incubation, the absorbance was measured on a plate reader using a
450 nm filter. Each assay was performed in six replicates.
Wound-healing assays
CD133+ CSCs were seeded into 6-well
plates using DMEM/F12 supplemented with 10% FBS and formed evenly
distributed monolayers. Then the medium was replaced with
serum-free DMEM/F12 medium and serum starvation continued
overnight. The monolayers were mechanically scratched using a
pipette tip to create a wound under the guidance of a sterilized
ruler. Cells were washed twice with PBS and fresh serum-free
DMEM/F12 medium was added in order to observe wound healing. Each
assay was performed in six replicates. Images were captured 24 and
48 h after incubation under an Olympus BX53 light microscope
(Olympus Corporation) at ×100 magnification The relative wound
distance was calculated as the percentage of the distance in
different groups at 24 and 48 h to the corresponding distance at 0
h (21).
Matrigel invasion assays
The upper chambers of Transwell plates were coated
with 100 µl Matrigel (BD Biosciences), then the
CD133+ CSCs (NC group and STIP1-siRNA group) at the cell
density of 1×105 cells/well were added to the upper
chambers in 200 µl serum-free medium, placed in a 24-well
plate, and 500 µl DMEM/F12 supplemented with 10% FBS was
added to the lower chambers. Five replicates were performed for
each group. Following 24 h incubation at 37°C, the non-invading
cells that remained on the upper membranes were removed by
swabbing. The invading cells on the underside of the filters were
fixed with 4% paraformaldehyde for 30 min, stained with 0.1%
crystal violet for 10 min at room temperature, and then images were
captured under light microscopy (magnification, ×200) for
quantification. The extent of cell invasion was quantified as the
number of invading cells in the STIP1-siRNA group divided by the
number of invading cells in the NC group.
Western blot analysis
Briefly, total protein was extracted from cells
using RIPA lysis buffer containing a freshly added proteinase
inhibitor cocktail and a phosphatase inhibitor cocktail (all Wuhan
Boster Biological Technology, Ltd.). Protein concentrations were
quantified using a BCA Protein Assay kit (Thermo Fisher Scientific,
Inc.). Protein samples were denatured at 100°C for 10 min and then
preserved at -20°C for later use. The proteins (50 µg/lane)
were separated via SDS-PAGE (12% gel), and then transferred onto
polyvinylidene fluoride (PVDF) membranes (Millipore), which were
treated with TBS-0.05% Tween-20 (TBS-T) blocking buffer (cat. no.
AR0179; Wuhan Boster Biological Technology, Ltd.) for 1 h at room
temperature. The PVDF membranes were first probed with primary
antibodies (1:1,000 dilution) overnight at 4°C, and then re-probed
with a horseradish peroxidase-conjugated secondary antibody
(1:1,000-1:2,000 dilution; Wuhan Boster Biological Technology,
Ltd.) for 1.5 h at 37°C. Antibodies against the following proteins
were used: ERK1/2 (cat. no. 9102; 1:1,000; Cell Signaling
Technology, Inc.), phosphorylated (p-)ERK1/2 (cat. no. 9101;
1:1,000; Cell Signaling Technology, Inc.), Akt (cat. no. AA326;
1:1,000; Beyotime Institute of Biotechnology), p-Akt (Ser473; cat.
no. AA326; 1:1,000; Beyotime Institute of Biotechnology), STIP1
(cat. no. 4464; 1:1,000; Cell Signaling Technology, Inc.), matrix
metalloproteinase (MMP)-2 (cat. no. sc-13594; 1:500; Santa Cruz
Biotechnology, Inc.), MMP-9 (cat. no. sc-12759; 1:500; Santa Cruz
Biotechnology, Inc.), tissue inhibitor of metalloproteinase
(TIMP)-2 (cat. no. sc-21735; 1:500; Santa Cruz Biotechnology,
Inc.,), and GAPDH (cat. no. A00227; 1:1,000; Wuhan Boster
Biological Technology, Ltd.,). After washing the PVDF membranes
three times with TBS-T, chemiluminescence emission was visualized
using a SuperSignal West Femto Substrate Trial kit (Thermo Fisher
Scientific, Inc.). Each western blot image presented is from the
same lysate and is representative of three separate experiments.
The relative intensity of MMP-2, MMP-9 and TIMP-2 was normalized to
the corresponding GAPDH intensity, and that of p-Akt and p-ERK1/2
was normalized to the corresponding total Akt and ERK1/2 intensity,
respectively. The signal intensities of bands were quantified using
ImageJ software 1.52a (National Institutes of Health).
Statistical analysis
Quantitative data are presented as the mean ±
standard error of the mean. Statistical analyses were performed
using SPSS 15.0 statistics software (SPSS, Inc.). One-way ANOVA
followed by Tukey's post hoc test for multiple comparisons was used
to assess the differences among groups containing one variable,
whereas a two-way ANOVA followed by the Bonferroni post hoc test
was used to assess the differences among groups containing two
variables. P<0.05 was considered to indicate a statistically
significant difference for all analyses.
Results
Isolation and identification of human
osteosarcoma CD133+ CSCs
CD133+ cells were isolated via MACS from
a population of MG63 cells in the logarithmic phase of growth, and
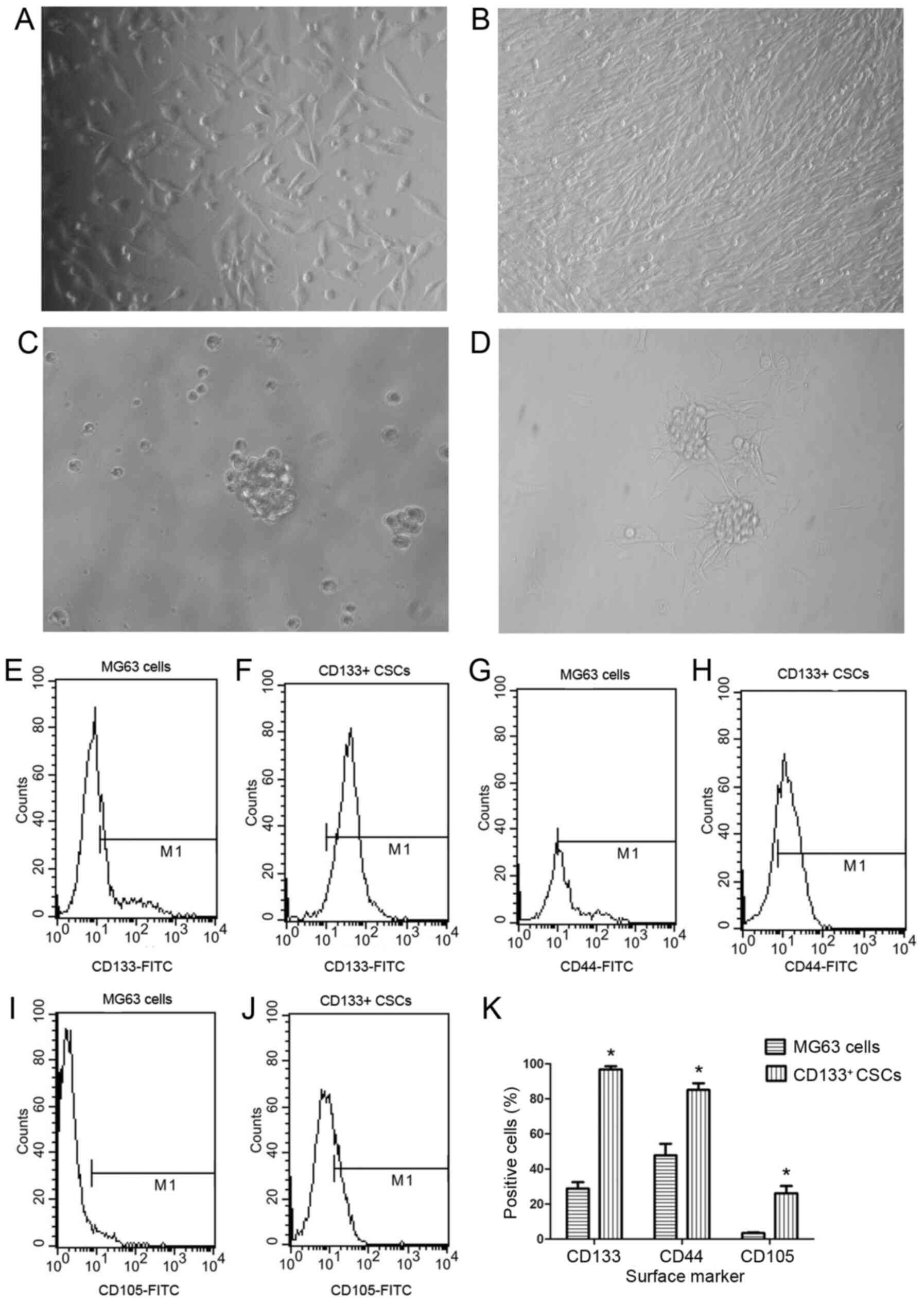
then inoculated in culture bottles (Fig. 1A). When observed under a light
microscope, CD133+ MG63 cells appeared irregular,
fibroblast-like and had a large nucleus (Fig. 1B). When the cells were cultured in
serum-free medium for 7-10 days, CD133+ osteosarcoma CSC
spheres formed; these were characterized by a round shape, a
regular border, and a cell number of ≥50 per sphere (Fig. 1C). Compared with a previous study
(19), microscopy indicated that
CD133+ osteosarcoma CSC spheres did not vary greatly in
size and shape, and closely resembled the spheres formed by MG63
cells; however the quantity of the former was larger than the
latter.
CD133+ osteosarcoma CSCs have the ability
to form secondary CSC spheres that are similar to the primary CSC
spheres at a higher frequency. When CD133+ osteosarcoma
CSC spheres were cultured in serum-supplemented medium, the spheres
adhered to the wall and produced new cancer cells (Fig. 1D).
Certain stem cell surface markers, such as CD44,
CD105, CD133, and key characteristic molecules of pluripotent stem
cells, such as Stro-1 and Nanog, are considered as markers of
osteosarcoma CSCs (12,13,18,19,22). Thus, CD44, CD105 and CD133 were
selected as CSC markers for their identification by flow cytometry.
Only 28.78±3.67% of MG63 cells were CD133+, but the
proportion in CSCs was 96.82±1.83% (Fig. 1E, F and K). Among the
CD133+ CSCs, 85.13±3.84% were CD44+, compared
with 47.77±6.60% for MG63 cells (Fig.
1G, H and K). For CD105, the percentages were 26.18±4.13 and
3.46±0.44% in CSCs and MG63 cells, respectively (Fig. 1I-K).
Inhibition of STIP1 expression by
siRNA
Following transfection of siRNAs targeting STIP1,
efficiency was detected by fluorescence microscopy after 24 h
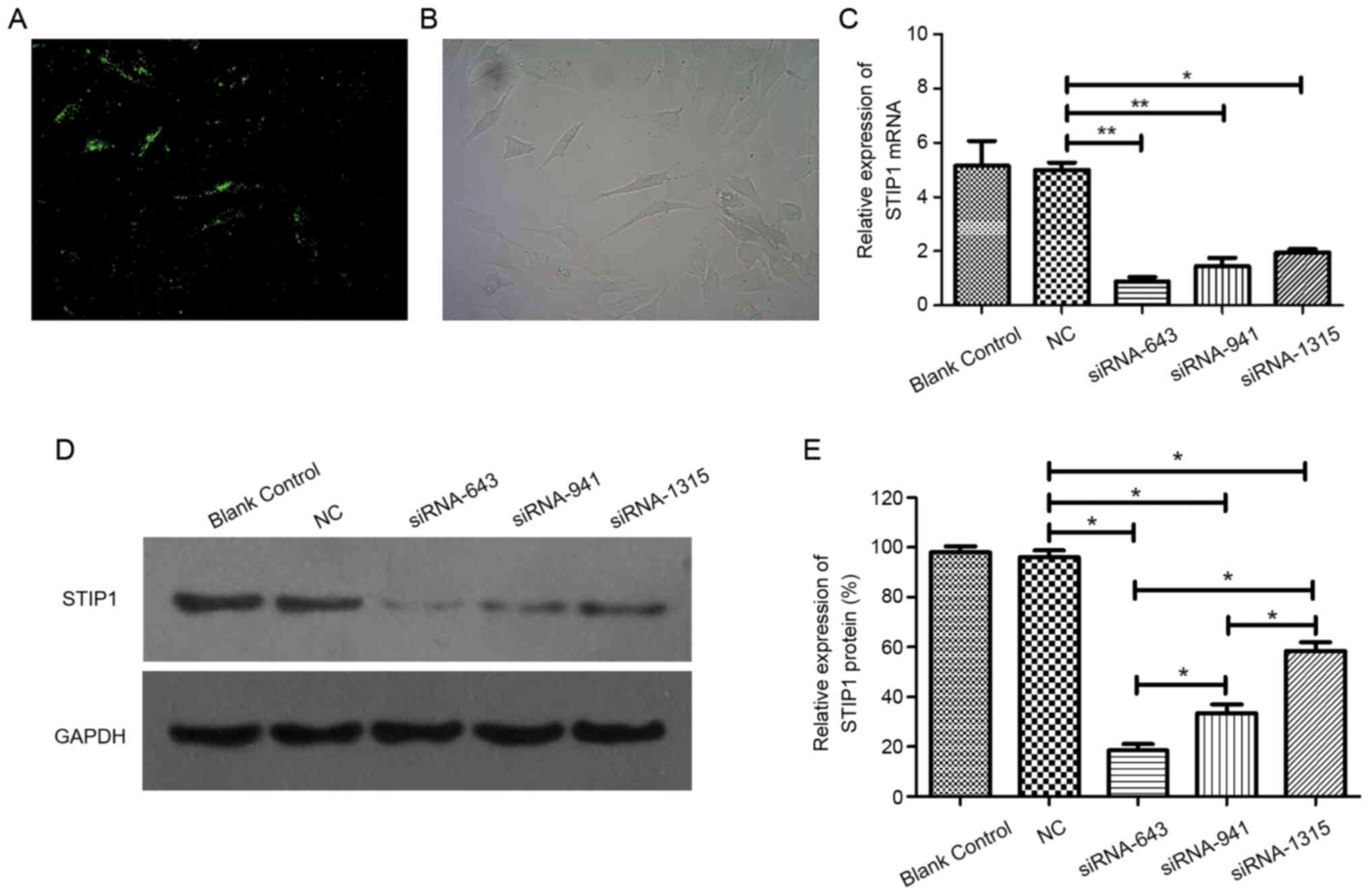
(Fig. 2A and B); the images
indicate that the transfection was successful. To evaluate whether
the STIP1 siRNAs inhibited STIP1 expression, the levels of STIP1
mRNA and protein in the CD133+ osteosarcoma CSCs were
analyzed using RT-qPCR and western blotting 48 h after
transfection. Both the mRNA (Fig.
2C) and protein expression levels (Fig. 2D and E) of STIP1 were knocked down
in the STIP1 siRNA-transfected CSCs compared with those observed in
the NC siRNA-transfected and blank control (non-siRNA-transfected)
groups, siRNA-643 group showed the best inhibition effect on STIP,
siRNA-941 group showed a medium inhibition effect, and siRNA-1315
group showed the worst inhibition effect (Fig. 2C, D and E). In order to evaluate
the biological effects of interfering STIP1 and explore the
biological differences among different interference levels,
siRNA-643 and siRNA-1315 were used in subsequent experiments.
Inhibition of cell migration and invasion
ability following STIP1 knockdown
To determine whether STIP1 knockdown affected cell
proliferation, a CCK-8 assay was performed within 48 h of
transfection. No significant changes in cell proliferation were
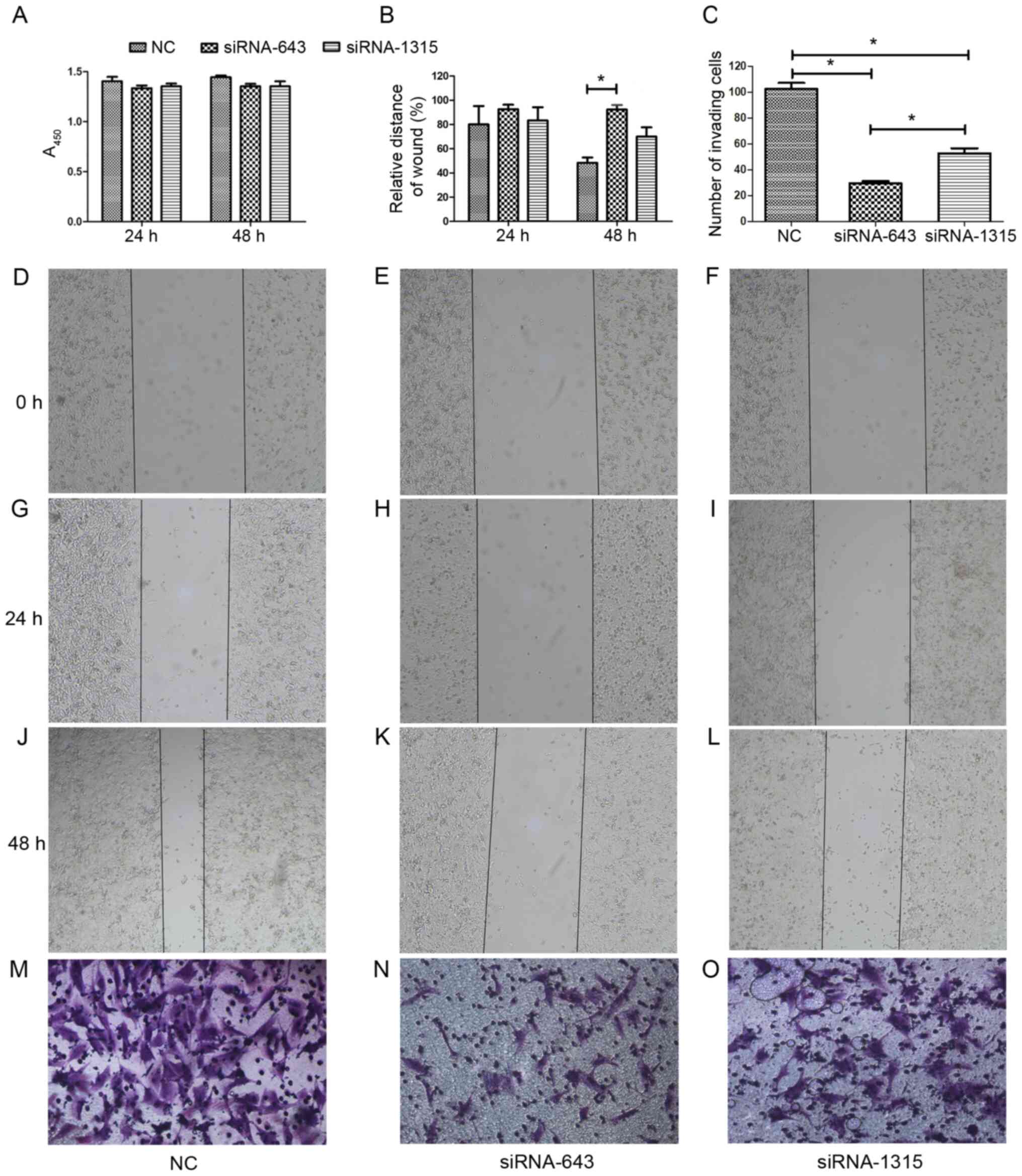
observed between the NC and siRNA groups at 24 and 48 h (Fig. 3A). To examine the effect of STIP1
on the migration ability of CD133+ osteosarcoma CSCs, a
wound-healing assay was performed, which demonstrated that the
wound healing ability of STIP1 siRNA-643 transfected CSCs were
weaker than those of the NC (P<0.05) at 48 h (Fig. 3B, D-I). To investigate the effect
of STIP1 on the invasion capability of CD133+
osteosarcoma CSCs, a Matrigel Transwell assay was carried out,
which confirmed that, following STIP knock-down, the number of CSCs
that invaded the lower chambers was significantly decreased
compared with that observed in the NC group (P<0.05).
Furthermore, the number of invading cells in the siRNA-643 group
was lower than that reported in the siRNA-1315 group (Fig. 3C and M-O). Overall, these results
suggest that STIP1 knockdown inhibits the migration and invasion
capabilities of CD133+ osteosarcoma CSCs.
STIP1 knockdown affects the expression of
MMPs
Based on the association between MMPs and invasion
in cancer, changes in MMP-2 and MMP-9 expression levels were
examined in order to explore the effects of STIP1 on migration and
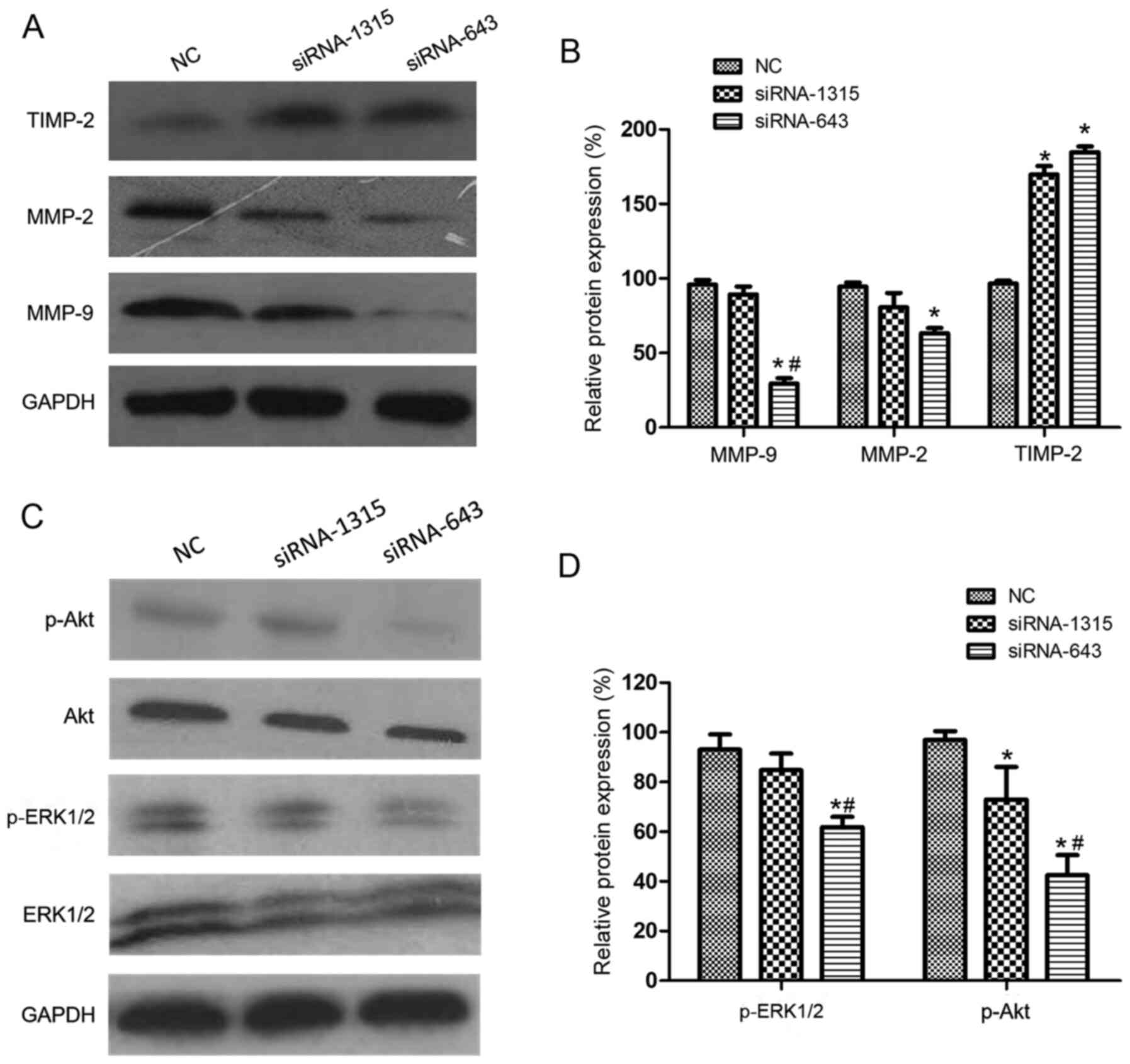
invasion. The results demonstrated that the siRNA-643-transfected
cells exhibited decreased MMP-2 and MMP-9 expression, and that
TIMP-2 protein levels were significantly increased in both cell
groups with STIP1 knock-down compared with the NC (Fig. 4A and B). These results were
consistent with the results of the migration and invasion assays in
CD133+ osteosarcoma CSCs.
STIP1 regulation of MMP expression may
involve the PI3K/Akt and ERK1/2 signaling pathways
The present study aimed to determine the potential
mechanism underlying the effect of STIP1 on CD133+
osteosarcoma CSC migration and invasion. Western blot analysis
confirmed that p-Akt was downregulated following STIP1 knockdown,
but the total Akt levels did not appear to be affected. In
addition, the ratio of p-ERK1/2 to total ERK1/2 was significantly
decreased in the siRNA-643 group compared with that in the NC
(Fig. 4C and D). Thus, these
results suggest that knockdown of STIP1 leads to a decrease in the
levels of the phosphorylated forms of Akt and ERK1/2, and
therefore, their activity.
Discussion
Based on current diagnostic guidelines and clinical
experience, standard chemotherapy has been shown to provide
significant benefits in terms of improving long-term survival
(1). However, improvement in
overall survival remains a challenge due to a large number of
patients with osteosarcoma developing resistance to
chemotherapeutic agents and undergoing metastasis (2). While a growing body of evidence
supports the CSC theory, some researchers believe that tumors are
derived from a single cancer-initiating cell with stem
cell/cell-like proper-ties, and that metastasis is attributable to
this sub-population of cancer cells (16). Furthermore, osteosarcoma CSCs are
responsible for the growth, metastasis and poor prognosis of
osteosarcoma; agents that may inhibit the proliferation and
differentiation of CSCs may provide new opportunities in the
prevention of tumor recurrence and metastasis (23,24). Previous studies have suggested
that CSCs may be isolated from cell lines using MACS or the
serum-free medium suspension cell sphere culture method (18,19). According to characteristic surface
markers and the self-renewal behavior of CSCs, the two methods were
combined in the present study to isolate the osteosarcoma CSCs in
order to improve the efficiency and purity of cell sorting.
CD133 is regarded as a marker of osteosarcoma CSCs
and a potential target for drug delivery. Killing CD133+
osteosarcoma CSCs may significantly inhibit osteosarcoma growth
(14,25). CD133 could also be used as a
sorting marker, as CD133+ osteosarcoma CSCs have high
expression of stemness, metastasis and drug-resistance markers
(13). Additionally, the results
of the present study showed that there were higher levels of CD44,
CD105 and CD133 expression in CD133+ osteosarcoma CSCs
that underwent the serum-free medium culture sorting method,
compared with MG63 cells, and suggest that the combined sorting
method can be used as an efficient tool for CSC isolation.
Previous studies have demonstrated that STIP1 could
serve as a novel biomarker for poor prognosis and advanced cancer
progression in patients with malignant tumors, including ovarian
cancer, hepatocellular carcinoma, esophageal squamous cell
carcinoma, papillary thyroid carcinoma and gastric cancer (7,26).
Cell migration and invasion are important malignant characteristics
in cancer metastasis (27). It
has been reported that STIP1 may be associated with tumor
metastasis (7). Iglesia et
al (28) found that
glioblastoma stem-like cells have high expression levels of STIP1,
that STIP1 is able to modulate glioblastoma stem-like cell
proliferation and self-renewal, and that STIP1-knockdown affects
proliferative and tumorigenic capacity. It is well known that MMPs
play a key role in cell migration and invasion, in that cell
invasion involves the degradation of the basement membrane
extracellular matrix (27). Walsh
et al (8) showed that
STIP1 knockdown was able to inhibit invasion in pancreatic cancer
cells, and that this inhibition may be induced via the
downregulation of MMP-2. Furthermore, the same study revealed that
STIP1-siRNA did not alter levels of HSP70 or HSP90 expression,
whereas the expression levels of the HSP90 client proteins such as
HER2, Bcr-Abl, and proto-oncogenes c-MET and v-Src were decreased
with STIP1-siRNA transfection. In addition, STIP1 was able to act
as a transcription factor and bind to the MMP-9 promoter to enhance
MMP-9 transcriptional expression (29). In the present study, it was shown
that STIP1 knockdown was able to inhibit CD133+
osteosarcoma CSC migration and invasion, but did not directly
affect the ability of the cells to proliferate for 48 h, in
agreement with studies by Huang et al (27) and Walsh et al (8). Moreover, the results of the present
study showed that STIP1 knockdown reduced the expression of MMP-2
and MMP-9, and promoted the expression of TIMP-2.
To further the understanding of the molecular
mechanisms by which STIP1 knockdown inhibits CD133+
osteosarcoma CSC migration and invasion, western blotting was used
to investigate the association of STIP1 with the ERK1/2 and
PI3K/Akt pathways. The PI3K/Akt signaling pathway is important in
regulating the processes of cancer development, including tumor
formation, progression, multi-drug resistance, tumor invasion and
metastasis (19). Erlich et
al (30) reported that STIP1
promoted glioma proliferation through the MAPK and PI3K pathways.
Other signaling pathways, such as ERK1/2, Wnt/β-catenin, STAT3 and
JAK2, are triggered by STIP1 in various types of cancer (7,31).
The results of the present study suggest that STIP1 knockdown may
inhibit Akt and ERK1/2 phosphorylation, and subsequently inhibit
the activity of the PI3K/Akt and ERK1/2 pathways, in agreement with
the results reported by Wang et al (29).
In summary, the present study identified an
association between STIP1 and osteosarcoma CSC invasion, as well as
its potential molecular mechanisms. A better understanding of the
role of STIP1 in osteosarcoma could facilitate the development of
molecular targeted therapies able to reduce the metastasis rate and
improve overall patient survival.
Acknowledgments
No applicable.
Abbreviations:
|
STIP1
|
stress-induced phosphoprotein 1
|
|
HSP
|
heat shock proteins
|
|
HOP
|
HSP70/HSP90 organizing protein
|
|
CSCs
|
cancer stem/stem-like cells
|
|
p-Akt
|
phosphorylated Akt
|
|
MMP
|
matrix metalloproteinase
|
|
TIMP
|
tissue inhibitor of
metalloproteinase
|
|
siRNA
|
small interfering RNA
|
|
ERK1/2
|
extracellular regulated protein
kinases 1/2
|
Funding
This work was supported by the Hubei Province Health
and Family Planning Scientific Research Project (grant no.
WJ2017M058).
Availability of data and materials
The datasets used and/or analyzed in the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
JHW, YL and QL performed the experiments. JHW edited
and verified the manuscript. CG and HHX designed the study and
wrote the first draft of the manuscript. FJG provided the MG63
cells and performed in vitro experiments. XZ revised the
first draft of the article and repeated some of the experiments.
MSZ and HQ assisted with the statistics and survival data analysis.
TFC performed the experiments and participated in writing and
revising the manuscript. CG, JHW and HHX helped design the figures
and tables. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Heare T, Hensley MA and Dell'Orfano S:
Bone tumors: Osteosarcoma and Ewing's sarcoma. Curr Opin Pediatr.
21:365–372. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Xiao X, Wang W, Li Y, Yang D, Li X, Shen
C, Liu Y, Ke X, Guo S and Guo Z: HSP90AA1-mediated autophagy
promotes drug resistance in osteosarcoma. J Exp Clin Cancer Res.
37:2012018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ellis RJ: Molecular chaperones: Assisting
assembly in addition to folding. Trends Biochem Sci. 31:395–401.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wang Y and McAlpine SR: Regulating the
cytoprotective response in cancer cells using simultaneous
inhibition of Hsp90 and Hsp70. Org Biomol Chem. 13:2108–2116. 2015.
View Article : Google Scholar
|
|
5
|
Longshaw VM, Baxter M, Prewitz M and
Blatch GL: Knockdown of the co-chaperone Hop promotes extranuclear
accumulation of Stat3 in mouse embryonic stem cells. Eur J Cell
Biol. 88:153–166. 2009. View Article : Google Scholar
|
|
6
|
Longshaw VM, Chapple JP, Balda MS,
Cheetham ME and Blatch GL: Nuclear translocation of the Hsp70/Hsp90
organizing protein mSTI1 is regulated by cell cycle kinases. J Cell
Sci. 117(Pt 5): 701–710. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhai E, Liang W, Lin Y, Huang L, He X, Cai
S, Chen J, Zhang N, Li J, Zhang Q, et al: HSP70/HSP90-Organizing
protein contributes to gastric cancer progression in an autocrine
fashion and predicts poor survival in gastric cancer. Cell Physiol
Biochem. 47:879–892. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Walsh N, Larkin A, Swan N, Conlon K,
Dowling P, McDermott R and Clynes M: RNAi knockdown of Hop
(Hsp70/Hsp90 organising protein) decreases invasion via MMP-2 down
regulation. Cancer Lett. 306:180–189. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Baindur-Hudson S, Edkins AL and Blatch GL:
Hsp70/Hsp90 organising protein (hop): Beyond interactions with
chaperones and prion proteins. Subcell Biochem. 78:69–90. 2015.
View Article : Google Scholar
|
|
10
|
Liu B, Ma W, Jha RK and Gurung K: Cancer
stem cells in osteosarcoma: Recent progress and perspective. Acta
Oncol. 50:1142–1150. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yang M, Yan M, Zhang R, Li J and Luo Z:
Side population cells isolated from human osteosarcoma are enriched
with tumor-initiating cells. Cancer Sci. 102:1774–1781. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gibbs CP, Kukekov VG, Reith JD,
Tchigrinova O, Suslov ON, Scott EW, Ghivizzani SC, Ignatova TN and
Steindler DA: Stem-like cells in bone sarcomas: Implications for
tumorigenesis. Neoplasia. 7:967–976. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Koka P, Mundre RS, Rangarajan R,
Chandramohan Y, Subramanian RK and Dhanasekaran A: Uncoupling
Warburg effect and stemness in CD133+ve cancer stem
cells from Saos-2 (osteosarcoma) cell line under hypoxia. Mol Biol
Rep. 45:1653–1662. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ni M, Xiong M, Zhang X, Cai G, Chen H,
Zeng Q and Yu Z: Poly(lactic-co-glycolic acid) nanoparticles
conjugated with CD133 aptamers for targeted salinomycin delivery to
CD133+ osteosarcoma cancer stem cells. Int J
Nanomedicine. 10:2537–2554. 2015. View Article : Google Scholar :
|
|
15
|
Yang J, Guo W, Wang L, Yu L, Mei H, Fang
S, Ji P, Liu Y, Liu G and Song Q: Cisplatin-resistant osteosarcoma
cells possess cancer stem cell properties in a mouse model. Oncol
Lett. 12:2599–2605. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Brown HK, Tellez-Gabriel M and Heymann D:
Cancer stem cells in osteosarcoma. Cancer Lett. 386:189–195. 2017.
View Article : Google Scholar
|
|
17
|
Li K, Li X, Tian J, Wang H, Pan J and Li
J: Downregulation of DNA-PKcs suppresses P-gp expression via
inhibition of the Akt/NF-κB pathway in CD133-positive osteosarcoma
MG-63 cells. Oncol Rep. 36:1973–1980. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gong C, Liao H, Guo F, Qin L and Qi J:
Implication of expression of Nanog in prostate cancer cells and
their stem cells. J Huazhong Univ Sci Technolog Med Sci.
32:242–246. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gong C, Liao H, Wang J, Lin Y, Qi J, Qin
L, Tian LQ and Guo FJ: LY294002 induces G0/G1 cell cycle arrest and
apoptosis of cancer stem-like cells from human osteosarcoma via
down-regulation of PI3K activity. Asian Pac J Cancer Prev.
13:3103–3107. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
21
|
Li J, Sun X, Wang Z, Chen L, Li D, Zhou J
and Liu M: Regulation of vascular endothelial cell polarization and
migration by Hsp70/Hsp90-organizing protein. PLoS One.
7:e363892012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Abarrategi A, Tornin J, Martinez-Cruzado
L, Hamilton A, Martinez-Campos E, Rodrigo JP, González MV, Baldini
N, Garcia-Castro J and Rodriguez R: Osteosarcoma: Cells-of-Origin,
cancer stem cells, and targeted therapies. Stem Cells Int.
2016:36317642016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Qi XT, Li YL, Zhang YQ, Xu T, Lu B, Fang
L, Gao JQ, Yu LS, Zhu DF, Yang B, et al: KLF4 functions as an
oncogene in promoting cancer stem cell-like characteristics in
osteosarcoma cells. Acta Pharmacol Sin. 40:546–555. 2019.
View Article : Google Scholar :
|
|
24
|
Chang Y, Zhao Y, Zhan H, Wei X, Liu T and
Zheng B: Bufalin inhibits the differentiation and proliferation of
human osteosarcoma cell line hMG63-derived cancer stem cells.
Tumour Biol. 35:1075–1082. 2014. View Article : Google Scholar
|
|
25
|
Chen F, Zeng Y, Qi X, Chen Y, Ge Z, Jiang
Z, Zhang X, Dong Y, Chen H and Yu Z: Targeted salinomycin delivery
with EGFR and CD133 aptamers based dual-ligand lipid-polymer
nanoparticles to both osteosarcoma cells and cancer stem cells.
Nanomedicine. 14:2115–2127. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Xu YW, Liu CT, Huang XY, Huang LS, Luo YH,
Hong CQ, Guo HP, Xu LY, Peng YH and Li EM: Serum Autoantibodies
against STIP1 as a potential biomarker in the diagnosis of
esophageal squamous cell carcinoma. Dis Markers. 2017:53840912017.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Huang L, Zhai E, Cai S, Lin Y, Liao J, Jin
H, Peng S, Xu L, Chen M and Zeng Z: Stress-inducible Protein-1
promotes metastasis of gastric cancer via Wnt/β-catenin signaling
pathway. J Exp Clin Cancer Res. 37:62018. View Article : Google Scholar
|
|
28
|
Iglesia RP, Prado MB, Cruz L, Martins VR,
Santos TG and Lopes MH: Engagement of cellular prion protein with
the co-chaperone Hsp70/90 organizing protein regulates the
proliferation of glioblastoma stem-like cells. Stem Cell Res Ther.
8:762017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wang HS, Tsai CL, Chang PY, Chao A, Wu RC,
Chen SH, Wang CJ, Yen CF, Lee YS and Wang TH: Positive associations
between upregulated levels of stress-induced phosphoprotein 1 and
matrix metalloproteinase-9 in endometriosis/adenomyosis. PLoS One.
13:e1905732018.
|
|
30
|
Erlich RB, Kahn SA, Lima FR, Muras AG,
Martins RA, Linden R, Chiarini LB, Martins VR and Moura Neto V:
STI1 promotes glioma proliferation through MAPK and PI3K path-ways.
Glia. 55:1690–1698. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tsai CL, Chao A, Jung SM, Tsai CN, Lin CY,
Chen SH, Sue SC, Wang TH, Wang HS and Lai CH: Stress-induced
phospho-protein-1 maintains the stability of JAK2 in cancer cells.
Oncotarget. 7:50548–50563. 2016. View Article : Google Scholar : PubMed/NCBI
|