|
1
|
Chilumula K, Saha PK, Muthyala T, Saha SC,
Sundaram V and Suri V: Prognostic role of uterine artery Doppler in
early- and late-onset preeclampsia with severe features. J
Ultrasound. Aug 24–2020.Epub ahead of print. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Dyess NF and Kinsella JP: Cardiovascular
implications for offspring born to mothers with preeclampsia. J
Pediatr. 228:11–12. 2021. View Article : Google Scholar
|
|
3
|
ACOG Practice Bulletin No. 202:
Gestational hypertension and preeclampsia. Obstet Gynecol.
133:12019.
|
|
4
|
Bakrania BA, George EM and Granger JP:
Animal models of preeclampsia: Investigating pathophysiology and
therapeutic targets. Am J Obstet Gynecol. Nov 23–2020.Epub ahead of
print. View Article : Google Scholar
|
|
5
|
Jia Y, Xie H, Zhang J and Ying H:
Induction of TGF-β receptor I expression in a DNA
methylation-independent manner mediated by DNMT3A downregulation is
involved in early-onset severe preeclampsia. FASEB J.
34:13224–13238. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Reddy M, Fenn S, Rolnik DL, Mol BW, da
Silva Costa F, Wallace EM and Palmer KR: The impact of the
definition of preeclampsia on disease diagnosis and outcomes: A
retrospective cohort study. Am J Obstet Gynecol.
224:217.e1–217.e11. 2021. View Article : Google Scholar
|
|
7
|
Saghafi N, Pourali L, Ghavami Ghanbarabadi
V, Mirzamarjani F and Mirteimouri M: Serum heat shock protein 70 in
preeclampsia and normal pregnancy: A systematic review and
meta-analysis. Int J Reprod Biomed. 16:1–8. 2018.PubMed/NCBI
|
|
8
|
D'Ambra E, Capauto D and Morlando M:
Exploring the regulatory role of circular RNAs in neurodegenerative
disorders. Int J Mol Sci. 20:54772019. View Article : Google Scholar :
|
|
9
|
Bai S, Wu Y, Yan Y, Shao S, Zhang J, Liu
J, Hui B, Liu R, Ma H, Zhang X and Ren J: Construct a
circRNA/miRNA/mRNA regulatory network to explore potential
pathogenesis and therapy options of clear cell renal cell
carcinoma. Sci Rep. 10:136592020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Dai J, Zhuang Y, Tang M, Qian Q and Chen
JP: CircRNA UBAP2 facilitates the progression of colorectal cancer
by regulating miR-199a/VEGFA pathway. Eur Rev Med Pharmacol Sci.
24:7963–7971. 2020.PubMed/NCBI
|
|
11
|
Deepthi K and Jereesh AS: An ensemble
approach for CircRNA-Disease association prediction based on
Autoencoder and deep neural network. Gene. 762:1450402020.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gong G, Han Z, Wang W, Xu Q and Zhang J:
Silencing hsa_circRNA_0008035 exerted repressive function on
osteosarcoma cell growth and migration by upregulating
microRNA-375. Cell Cycle. 19:2139–2147. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen D, Ma W, Ke Z and Xie F: CircRNA
hsa_circ_100395 regulates miR-1228/TCF21 pathway to inhibit lung
cancer progression. Cell Cycle. 17:2080–2090. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhang J, Liu H, Hou L, Wang G, Zhang R,
Huang Y, Chen X and Zhu J: Circular RNA_LARP4 inhibits cell
proliferation and invasion of gastric cancer by sponging miR-424-5p
and regulating LATS1 expression. Mol Cancer. 16:1512017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang Y, Zhang J, Li J, Gui R, Nie X and
Huang R: CircRNA_014511 affects the radiosensitivity of bone marrow
mesenchymal stem cells by binding to miR-29b-2-5p. Bosn J Basic Med
Sci. 19:155–163. 2019.PubMed/NCBI
|
|
16
|
Ding C, Yi X, Wu X, Bu X, Wang D, Wu Z,
Zhang G, Gu J and Kang D: Exosome-mediated transfer of circRNA
CircNFIX enhances temozolomide resistance in glioma. Cancer Lett.
479:1–12. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang X, Zhang H, Yang H, Bai M, Ning T,
Deng T, Liu R, Fan Q, Zhu K, Li J, et al: Exosome-delivered circRNA
promotes glycolysis to induce chemoresistance through the
miR-122-PKM2 axis in colorectal cancer. Mol Oncol. 14:539–555.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Xiao G, Huang W, Zhan Y, Li J and Tong W:
CircRNA_103762 promotes multidrug resistance in NSCLC by targeting
DNA damage inducible transcript 3 (CHOP). J Clin Lab Anal.
34:e232522020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Bian L, Zhi X, Ma L, Zhang J, Chen P, Sun
S, Li J, Sun Y and Qin J: Hsa_circRNA_103809 regulated the cell
proliferation and migration in colorectal cancer via
miR-532-3p/FOXO4 axis. Biochem Biophys Res Commun. 505:346–352.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhao Y, Zheng R, Chen J and Ning D:
CircRNA CDR1as/miR-641/HOXA9 pathway regulated stemness contributes
to cisplatin resistance in non-small cell lung cancer (NSCLC).
Cancer Cell Int. 20:2892020. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hu X, Ao J, Li X, Zhang H, Wu J and Cheng
W: Competing endogenous RNA expression profiling in pre-eclampsia
identifies hsa_circ_0036877 as a potential novel blood biomarker
for early pre-eclampsia. Clin Epigenetics. 10:482018. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ou Y, Liu M, Zhu L, Deng K, Chen M, Chen H
and Zhang J: The expression profile of circRNA and its potential
regulatory targets in the placentas of severe pre-eclampsia. Taiwan
J Obstet Gynecol. 58:769–777. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
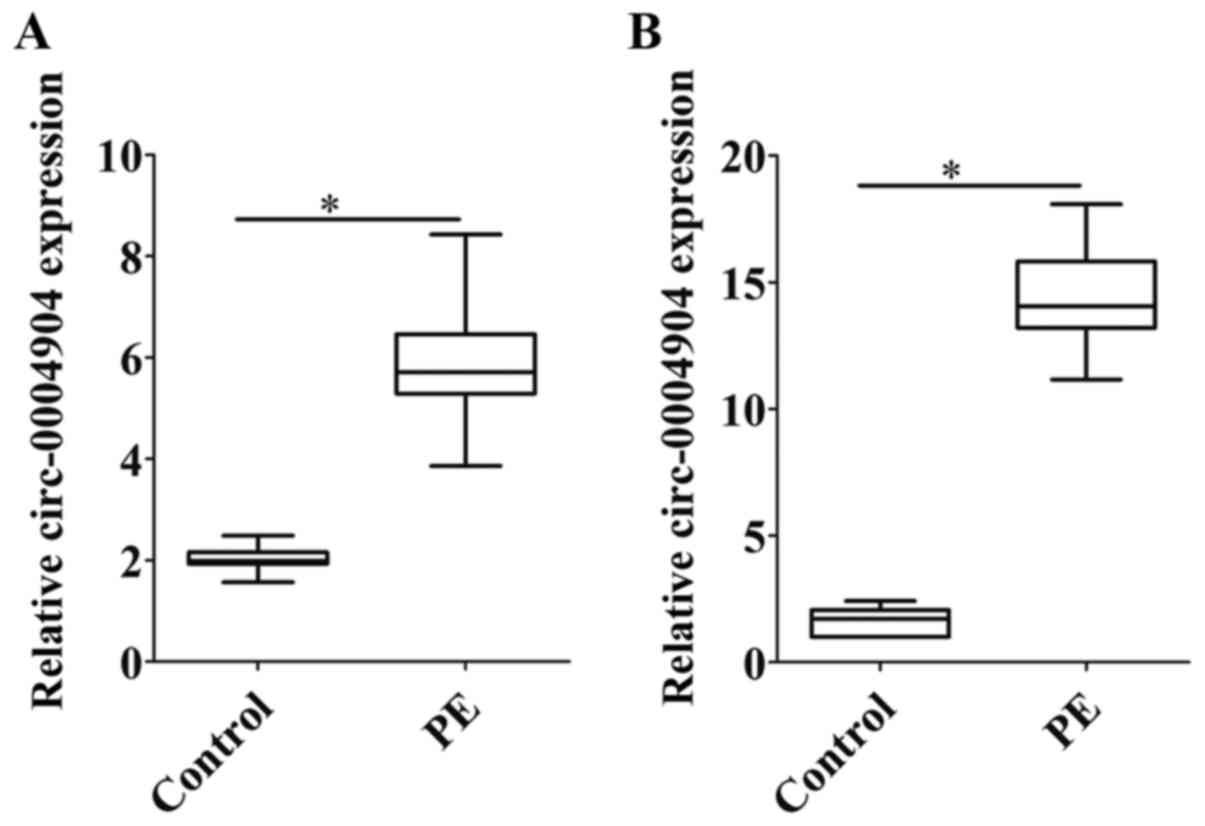
Jiang M, Lash GE, Zhao X, Long Y, Guo C
and Yang H: CircRNA-0004904, CircRNA-0001855, and PAPP-A: Potential
novel biomarkers for the prediction of preeclampsia. Cell Physiol
Biochem. 46:2576–2586. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Han D, Li J, Wang H, Su X, Hou J, Gu Y,
Qian C, Lin Y, Liu X, Huang M, et al: Circular RNA circMTO1 acts as
the sponge of microRNA-9 to suppress hepatocellular carcinoma
progression. Hepatology. 66:1151–1164. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
26
|
Wang Z, Yu Z, Wang GH, Zhou YM, Deng JP,
Feng Y, Chen JQ and Tian L: AURKB promotes the metastasis of
gastric cancer, possibly by inducing EMT. Cancer Manag Res.
12:6947–6958. 2020. View Article : Google Scholar :
|
|
27
|
Xu Y, Ye S, Zhang N, Zheng S, Liu H, Zhou
K, Wang L, Cao Y, Sun P and Wang T: The FTO/miR-181b-3p/ARL5B
signaling pathway regulates cell migration and invasion in breast
cancer. Cancer Commun (Lond). 40:484–500. 2020. View Article : Google Scholar
|
|
28
|
Shang A, Gu C, Wang W, Wang X, Sun J, Zeng
B, Chen C, Chang W, Ping Y, Ji P, et al: Exosomal circPACRGL
promotes progression of colorectal cancer via the
miR-142-3p/miR-506-3p-TGF-β1 axis. Mol Cancer. 19:1172020.
View Article : Google Scholar
|
|
29
|
Zhang HD, Jiang LH, Hou JC, Zhong SL, Zhou
SY, Zhu LP, Li J, Wang DD, Sun DW, Ji ZL and Tang JH: Circular RNA
hsa_circ_0052112 promotes cell migration and invasion by acting as
sponge for miR-125a-5p in breast cancer. Biomed Pharmacother.
107:1342–1353. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Xin Y, Min P, Xu H, Zhang Z and Zhang Y
and Zhang Y: CD26 upregulates proliferation and invasion in keloid
fibroblasts through an IGF-1-induced PI3K/AKT/mTOR pathway. Burns
Trauma. 8:tkaa0252020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Panda AC, Dudekula DB, Abdelmohsen K and
Gorospe M: Analysis of circular RNAs using the web tool
circinteractome. Methods Mol Biol. 1724:43–56. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Xia L, Li F, Qiu J, Feng Z, Xu Z, Chen Z
and Sun J: Oncogenic miR-20b-5p contributes to malignant behaviors
of breast cancer stem cells by bidirectionally regulating CCND1 and
E2F1. BMC Cancer. 20:9492020. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wu YP, Lin XD, Chen SH, Ke ZB, Lin F, Chen
DN, Xue XY, Wei Y, Zheng QS, Wen YA and Xu N: Identification of
prostate cancer-related circular RNA through bioinformatics
analysis. Front Genet. 11:8922020. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Garikipati VNS, Verma SK, Cheng Z, Liang
D, Truongcao MM, Cimini M, Yue Y, Huang G, Wang C, Benedict C, et
al: Circular RNA CircFndc3b modulates cardiac repair after
myocardial infarction via FUS/VEGF-A axis. Nat Commun. 10:43172019.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Cornelius DC and Wallace K: Autophagy in
preeclampsia: A new target? EBioMedicine. 57:1028642020. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Nakashima A, Cheng SB, Ikawa M, Yoshimori
T, Huber WJ, Menon R, Huang Z, Fierce J, Padbury JF, Sadovsky Y, et
al: Evidence for lysosomal biogenesis proteome defect and impaired
autophagy in preeclampsia. Autophagy. 16:1771–1785. 2020.
View Article : Google Scholar
|
|
37
|
Zhao H, Gong L, Wu S, Jing T, Xiao X, Cui
Y, Xu H, Lu H, Tang Y, Zhang J, et al: The inhibition of protein
kinase C β contributes to the pathogenesis of preeclampsia by
activating autophagy. EBioMedicine. 56:1028132020. View Article : Google Scholar
|
|
38
|
Akcora Yildiz D, Irtegun Kandemir S,
Agacayak E and Deveci E: Evaluation of protein levels of autophagy
markers (Beclin 1 and SQSTM1/p62) and phosphorylation of cyclin E
in the placenta of women with preeclampsia. Cell Mol Biol
(Noisy-le-grand). 63:51–55. 2017. View Article : Google Scholar
|
|
39
|
Oh SY, Choi SJ, Kim KH, Cho EY, Kim JH and
Roh CR: Autophagy-related proteins, LC3 and Beclin-1, in placentas
from pregnancies complicated by preeclampsia. Reprod Sci.
15:912–920. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Gao L, Qi HB, Kamana KC, Zhang XM, Zhang H
and Baker PN: Excessive autophagy induces the failure of
trophoblast invasion and vasculature: Possible relevance to the
pathogenesis of preeclampsia. J Hypertens. 33:106–117. 2015.
View Article : Google Scholar
|
|
41
|
Chen ZH, Cao JF, Zhou JS, Liu H, Che LQ,
Mizumura K, Li W, Choi AM and Shen HH: Interaction of caveolin-1
with ATG12-ATG5 system suppresses autophagy in lung epithelial
cells. Am J Physiol Lung Cell Mol Physiol. 306:L1016–L1025. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Murrow L and Debnath J: Atg12-Atg3
coordinates basal autophagy, endolysosomal trafficking, and exosome
release. Mol Cell Oncol. 5:e10391912018. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
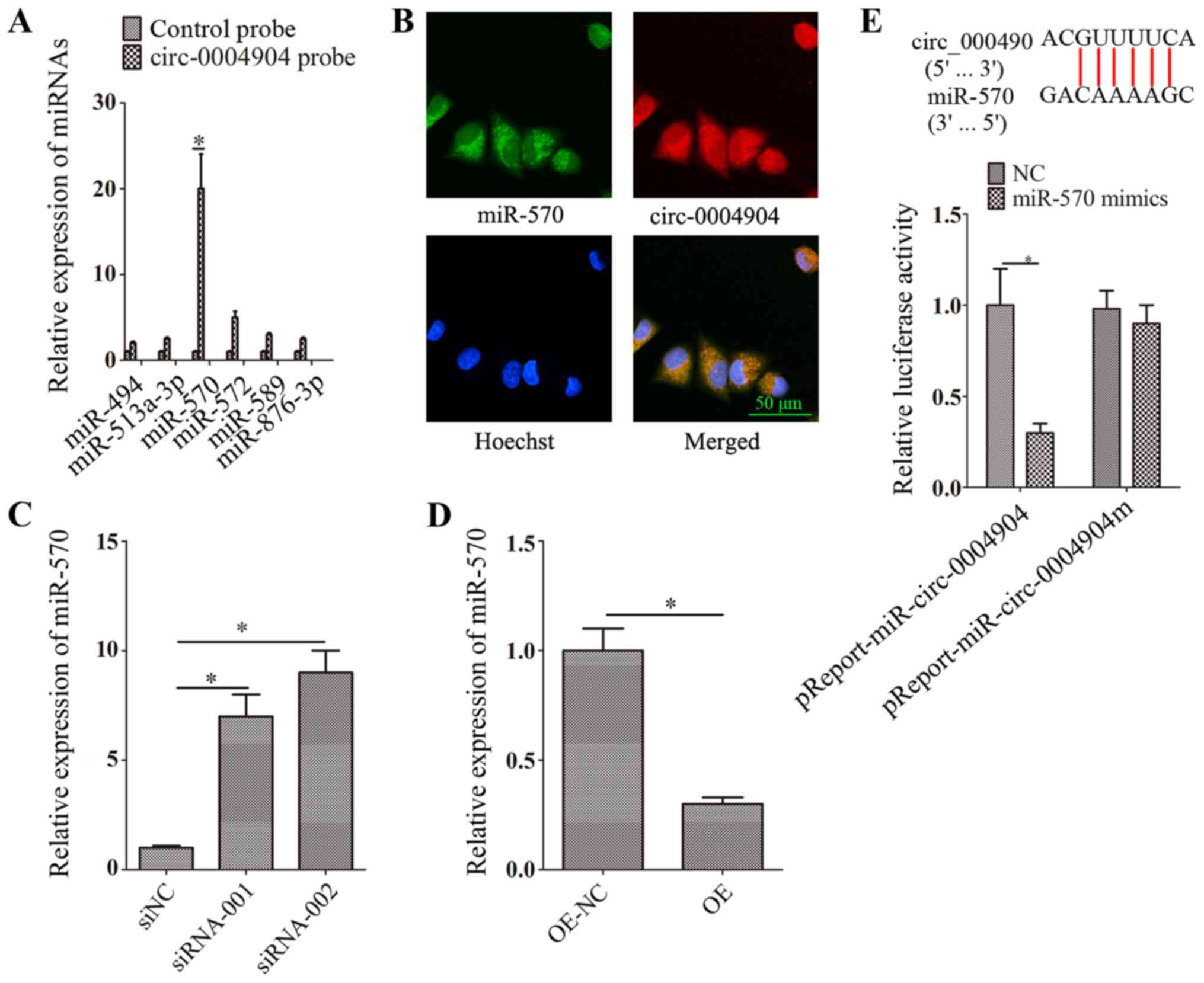
Bao X, Zhao L, Guan H and Li F: Inhibition
of LCMR1 and ATG12 by demethylation-activated miR-570-3p is
involved in the anti-metastasis effects of metformin on human
osteosarcoma. Cell Death Dis. 9:6112018. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Deng WG, Kawashima H, Wu G, Jayachandran
G, Xu K, Minna JD, Roth JA and Ji L: Synergistic tumor suppression
by coexpression of FUS1 and p53 is associated with down-regulation
of murine double minute-2 and activation of the apoptotic
protease-activating factor 1-dependent apoptotic pathway in human
non-small cell lung cancer cells. Cancer Res. 67:709–717. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Lin L, Wang Q, Xu F, Luo X, Xu J, Yan L,
Li Q and Hao H: BML-111, the lipoxin A4 agonist,
modulates VEGF or CoCl2-induced migration, angiogenesis
and permeability in tumor-derived endothelial cells. Immunol Lett.
230:27–35. 2021. View Article : Google Scholar
|
|
46
|
Sahay AS, Jadhav AT, Sundrani DP, Wagh GN,
Mehendale SS, Chavan-Gautam P and Joshi SR: VEGF and VEGFR1 levels
in different regions of the normal and preeclampsia placentae. Mol
Cell Biochem. 438:141–152. 2018. View Article : Google Scholar
|