Introduction
Pituitary adenomas (PAs), also known as pituitary
neuroendocrine tumours (PitNETs), are among the three primary
intracranial tumours, with an incidence of ~17.1% among
intracranial neoplasms, while the prevalence of clinically
symptomatic cases is estimated at ~1 in 1,100 individuals (1). Based on hormone hypersecretion
status, PitNETs are classified as either a clinically
non-functioning PitNET (NF-PitNET) or a hormone-secreting PitNET
(2). According to the 2022 World
Health Organization classification, which incorporates cell
lineage, hormone composition, and specific histological and
immunohistochemical (IHC) features, PitNETs are categorised into
subtypes, including prolactin-secreting (PRL-PitNET), growth
hormone-secreting (GH-PitNET), and adrenocorticotropic
hormone-secreting (ACTH-PitNET) tumours, among others (3). PRL-PitNET is the most prevalent
subtype, constituting ~53% of all PitNETs (1).
PRL-PitNET is currently the only PitNET type for
which pharmacotherapy is the preferred management approach.
Dopamine receptor agonists (DAs), including bromocriptine (BRC) and
cabergoline (CAB), are first-line treatments for PRL-PitNET
(4,5). Although most patients with a
PRL-PitNET achieve excellent outcomes with DA therapy, 20-30%
exhibit resistance to BRC, and 5-15% are resistant to CAB (6,7).
Previous studies have identified decreased expression or activity
of the dopamine D2 receptor (DRD2) as a primary cause of drug
resistance in PRL-PitNET (8-11); however, the specific mechanisms
underlying this remain incompletely understood. As PitNETs are
predominantly benign tumours, reports of gene mutations are
infrequent (12). Evidence
suggests that dysregulation of epigenetic modifications may be a
primary driver of PitNETs pathogenesis (13). Therefore, investigating
alterations in post-transcriptional regulatory mechanisms is
clinically significant for the targeted therapy of PitNETs.
Non-coding RNAs (ncRNAs) can target and regulate
signalling pathways, playing major roles in physiological and
pathological processes. Circular RNAs (circRNAs) are a class of
endogenous ncRNA characterised by a covalently closed loop
structure, generated through back-splicing of precursor mRNA
(14). This circular structure
confers resistance to exonuclease degradation, resulting in greater
stability compared with linear RNAs. Accumulating evidence
indicates that circRNAs play critical roles in numerous
tumour-related processes, including formation, drug resistance,
progression and relapse. To date, the association between circRNAs,
drug resistance in PRL-PitNET, and DRD2 expression/activity has not
been elucidated.
Exosomes (30-150 nm in diameter) are nanoscale
membrane vesicles derived from endosomal multivesicular bodies and
released into the extracellular environment. Previous studies
demonstrated that exosomal circRNAs exhibit high stability and
tissue specificity, playing crucial roles in neoplasm processes
such as drug resistance (15-17), diagnosis (18-20), treatment (21,22), prognosis (23,24), immunity (25), metabolism (26) and progression (27-29). For example, bone abnormalities
associated with GH-PitNET may be partially mediated by exosomes
carrying miR-21-5p, which targets PDCD4 to modulate the Smad7/Runx2
pathway (30). Additionally,
Zhang et al (31) found
that exosome-delivered lncRNA H19 reduced CAB resistance in GH3
cell lines, and plasma exosomal H19 served as a biomarker for
predicting CAB response in patients with a PRL-PitNET. However,
whether and how exosome-transmitted circRNAs induce CAB resistance
in PRL-PitNET remains unknown. In our previous study using gene
microarray assays (32) and
experimental analysis, circOMA1 was shown to be associated with
PRL-PitNET pathogenesis (33).
In the present study, circOMA1 was shown to be upregulated in
drug-resistant PRL-PitNET cells, and silencing circOMA1
substantially increased CAB sensitivity in PRL-PitNET cells.
Mechanistically, circOMA1 functioned as a cytoplasmic miR-145-5p
sponge, promoting Kelch-repeat and BTB domain-containing protein 7
(KBTBD7)-mediated DRD2 ubiquitination and consequently attenuating
CAB-induced autophagy.
Materials and methods
Patient samples
Human prolactinoma specimens were obtained from
patients undergoing an endoscopic endonasal transsphenoidal
approach. Immediately after removal, the tumour tissue was placed
in an ice box at 4°C and transferred to the laboratory within 2 h
for research or stored at −80°C. In addition, whole blood was
collected from 219 patients with PAs from the elbow vein. The
present study was performed in accordance with the ARRIVE
guidelines. All participants provided written informed consent, and
the study was approved by the Ethics Committee of the First
Affiliated Hospital of Sun Yat-sen University [approval no.
(2020)090; Guangzhou, China]. DA resistance was defined as a lack
of normalisation of prolactin (PRL) serum levels or associated
volume reduction (maximum diameter reduction ≥30%) after treatment
with standard DA doses (7.5-10 mg BRC daily or 2.0 mg CAB weekly)
for at least 6 months. The patient's postoperative blood samples
were collected at 1-7 days after surgery. Tissues from 6 cases of
drug-sensitive PitNET (PRL-PitNET-S) and 14 cases of drug-resistant
PitNET (PRL-PitNET-R) were collected. The clinical-pathological
information is shown in Tables SI
and SII.
Cell culture, plasmids and reagents
MMQ rat prolactinoma cells were obtained from ATCC
(cat. no. CRL-10609) and cultured in DMEM/F12 (Gibco; Thermo Fisher
Scientific, Inc.) supplemented with 10% FBS (Shanghai ExCell
Biology, Inc.), 100 U/ml penicillin, and 100 μg/ml
streptomycin (Nanjing KeyGen Biotech Co., Ltd.). 293T cells were
obtained from ATCC (cat. no. CRL-11268) and cultured in DMEM
supplemented with 10% FBS, 100 U/ml penicillin, and 100 mg/ml
streptomycin. Cells were maintained in a humidified incubator
supplied with 5% CO2 at 37°C or stored in liquid
nitrogen in serum-free cell freezing medium (cat. no. CS0401;
GenKern Biotechnology).
Exosome isolation
MMQ cells stably overexpressing exogenous circOMA1
using a lentiviral vector were established (termed circOM), with
cells transduced with an empty lentiviral vector serving as the
control (termed circNC). The circular structure of circOMA1 was
identified by amplification with junction-spanning primers and
Sanger sequencing. The exosomes in the supernatant from circOM or
circNC cells were extracted by differential ultracentrifugation.
Briefly, the cells were cultured in complete DMEM/F12, and when
required, the media replaced with serum-free DMEM/F12 medium for 24
or 48 h. The supernatant was collected and centrifuged through
2,000 × g for 20 min at 4°C, followed by 10,000 × g for 30 min.
After discarding the sediments, the supernatant was filtered
through 0.22-μm filters (MilliporeSigma), followed by
ultracentrifugation at 110,000 × g for 70 min through a Type 70Ti
rotor at 4°C (Beckman Coulter, Inc.). Subsequently, the pellets
were resuspended in PBS and purified by ultracentrifugation at
110,000 × g for 70 min at 4°C. Subsequently, the exosomes were
resuspended in PBS for co-culture or experimental use, or stored at
−80°C. All centrifugation conditions were at 4°C.
Extraction of genomic DNA (gDNA) and DNA
electrophoresis
gDNA Extraction from circOM cells was performed
using the SteadyPure Universal Genomic DNA Extraction Kit (cat. no.
AG21009; Accurate Biology) according to the manufacturer's
protocol. The product was then amplified by qPCR and visualised on
a 2% agarose gel using Safe Green (Biosharp Life Sciences). The DNA
polymerase used was included in SYBR Green Premix Pro Taq HS qPCR
Kit (cat. no. AG11701; Accurate Biology). The sequences of forward
and reverse primers were as follows: CircOMA1 forward,
5'ACCCAAGATGCCAGAATGGT-3' and reverse, 5'-TTGATGACAGCCCCGTGAG-3';
OMA1 forward, 5'-CGGTTCCTCTCTTGTTGA-3' and reverse,
5'-GTAGCTTGCTCCTTCCTG-3'; GAPDH forward, 5'-GCGAGATCCCTCCAAAAT-3'
and reverse, 5'-GTCCTTCCACGATACCAA-3'. The thermocycling conditions
were as follows: Preheat at 95°C for 30 sec, followed by 40 cycles
of 95°C for 5 sec and 60°C for 30 sec.
Actinomycin D and RNase R assay
circOM cells were treated with 0.5 μg/ml
actinomycin D (cat. no. HY-17559; MedChemExpress), and RNA was
extracted at different time points (0, 6, 12, 18 and 24 h). Reverse
transcription-quantitative (RT-qPCR) was used to detect the
stability of RNA. For RNase R assays, equal quantities of total RNA
(1 μg) with or without 3 U/μg RNase R (Guangzhou
Geneseed Biotech. Co., Ltd.) were incubated at 37°C for 15 min and
70°C for 10 min. Subsequently, RT-qPCR was used to detect the
abundance of target genes, and the products were visualised by DNA
electrophoresis on a 2% agarose gel.
RNA isolation and RT-qPCR
Total RNA was extracted from MMQ and 293T cells,
clinical samples, and exosomes using TRIzol® reagent
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. Nuclear and cytoplasmic RNA was
extracted using a Nuclear/cytoplasmic separation kit (cat. no.
BB-36021; Bestbio; https://www.beibokit.com/). Reverse transcription of
total RNA into cDNA was performed using PrimeScript™ RT MasterMix
(Takara Bio, Inc.) or a miRNA first-strand cDNA synthesis kit
(Accurate Biology) according to the manufacturer's instructions.
qPCR was performed using Genious 2X SYBR Green Fast qPCR MIX
(ABclonal Biotech Co., Ltd.). The thermocycling conditions were as
follows: Preheat at 95°C for 30 sec, followed by 40 cycles of 95°C
for 5 sec and 60°C for 30 sec. To quantify circRNA, mRNA, and miRNA
expression levels, the 2−ΔΔCq method was used (34), with GAPDH or U6 as the internal
control to normalise target gene expression. U6 and GAPDH were used
as nuclear and cytoplasmic controls, respectively. The experiment
was performed at least three times. Primer sequences are listed in
Table SIII.
Western blot analysis
Total protein was extracted from circOM, 293T cells,
PRL-PitNETs tissues, and exosomes using RIPA lysis buffer (Jiangsu
CoWin Biotech Co., Ltd.), quantified using BCA (Jiangsu CoWin
Biotech Co., Ltd.), and then 20-50 μg of protein was loaded
on 8, 10, or 12% SDS-gels by SDS-PAGE, and subsequently transferred
to a 0.22-μm or 0.45-μm PVDF membrane
(MilliporeSigma). Subsequently, membranes were blocked with 5%
skimmed milk for 1 h and incubated overnight with the primary
antibody at 4°C. Finally, the membranes were incubated in
horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG (H+L)
or HRP-conjugated goat anti-mouse IgG (H+L) secondary antibody at
room temperature for 2 h. Signals were visualised using an
ImageQuant Las4000mini with enhanced chemiluminescence reagent
(MilliporeSigma). Information on the antibodies used is listed in
Table SIV. ImageJ v1.8.0
(National Institutes of Health) was used for densitometric
analysis.
5-EdU assay
The sterile coverslips were coated with 100
μg/ml polylysine (Beyotime Institute of Biotechnology) in
24-well plates, and 1×105 cells were added for overnight
culture. Then, 10 μM EdU solution (Beyotime Institute of
Biotechnology) was added to the cells and incubated for 2 h at room
temperature. Subsequently, the cells were fixed with 4%
paraformaldehyde for 20 min at room temperature, washed, and
permeabilised with 0.3% Triton X-100. Next, a click-reaction was
performed according to the protocols (cat. no. C0075S; Beyotime
Institute of Biotechnology). Hoechst 33342 was used to stain the
nucleus for 10 min at room temperature. Images were observed using
confocal laser microscopy.
Immunoblotting and co-immunoprecipitation
(Co-IP)
For the Co-IP assay, cells were lysed using 500
μl IP lysis buffer with PMSF (cat. no. abs955; Absin) after
washing with pre-cooled PBS. The lysate was centrifuged at 14,000 ×
g for 10 min at 4°C and incubated with the indicated primary
antibody or rabbit anti-IgG antibody as a negative control. Then,
the compound was gently rotated overnight at 4°C. After incubation,
5 μl Protein A and 5 μl Protein G were added, and the
mixture was gently mixed at 4°C for 3 h. The centrifuged particles
were washed 4 times with a washing buffer, 1X SDS buffer was added,
and the sample was boiled at 100°C for 5 min for western
blotting.
Cycloheximide (CHX) assay
CHX (a protein synthesis inhibitor) and MG132 (a
proteasome inhibitor) were purchased from MedChemExpress. circOM,
circNC or MMQ cells were treated with 100 μg/ml CHX alone or
combined with 20 μM MG132 for 0, 4, 8, or 12 h, after which,
western blotting was performed.
RFP-GFP-LC3 assay
circNC or circOM cells were seeded into sterile
6-well plates and incubated for 1 day. RFP-GFP-LC3 plasmids were
transfected into cells using Liposome® 3000 (Thermo
Fisher Scientific, Inc.) using 3 μg of RFP-GFP-LC3 plasmid
at room temperature. After 48 h, the media was replaced, and the
cells were treated with CAB. The following day, cells were fixed
with 4% paraformaldehyde for 20 min and nuclei were stained using 1
μg/ml DAPI for 10 min at room temperature. Subsequently,
cells were imaged using a confocal microscope.
Lentivirus and stable cell line
construction
The sgRNA primer sequences of KBTBD7 were designed
by CHOPCHOP (chopchop.cbu.uib.no/). sgRNA1 is designed to target
positions 2067-2086bp of the target gene, while sgRNA2 targets
positions 570-589 bp. The sgRNA sequences (sgRNA1 forward,
5'-CACCGAGTATATGAATACGACACTA-3' and reverse,
5'-AAACTAGTGTCGTATTCATATACTC-3'; and sgRNA2 forward,
5'-CACCGCTTCAAGAGCATGTTCACAG-3' and reverse,
5'-AAACCTGTGAACATGCTCTTGAAGC-3') were synthesised by Beijing Qingke
Biotechnology Co., Ltd. Phosphorylate and oligos (sgRNAs) were
inserted into a LentiCRISPRv2 vector (cat. no. 52961; Addgene,
Inc.), linearised by restriction endonuclease using BsmBI
(cat. no. R0739; New England BioLabs, Inc.) using T4 ligase. 293T
cells were plated in a 10-cm culture dish and cultured until 80%
confluent. The constructed target plasmid LentiCRISPRv2 and
packaging plasmids psPAX2 (cat. no. 12260; Addgene, Inc.) and
pMD2.G (cat. no. 12259; Addgene, Inc.) were transfected using
Liposome® 3000 with a ratio of 5 μg: 5 μg:
5 μg. After 48 h of transfection, the supernatant containing
lentiviral particles (generation system used, 2nd; MOI used to
infect cells, 20) from 293T cells was collected and filtered
through a 0.45-μm filter (MilliporeSigma) for subsequent MMQ
cell infection. After 6 h of infection with lentiviral particles,
the media from the MMQ cells was replaced with fresh media. After
48 h, the level of endogenous protein was then verified by western
blotting, and puromycin (2 μg/ml) was added to select for
successfully transfected cells for 1 week. Finally, a stable
gene-knockdown MMQ cell line was obtained via the limited dilution
method.
Cell viability and colony formation
assays
For the cell activity assay, 3×103 cells
were plated in 96-well plates and incubated at specified time
points with or without CAB (cat. no. HY-15296; MedChemExpress). The
minimum number of cells considered to form a colony was 50.
Subsequently, Cell Counting Kit-8 (CCK-8) solution (Dojindo
Molecular Technologies, Inc.) was added (10 μl/well), and
the mixture was incubated at room temperature for 2 h. Absorbance
was measured using a microplate reader (Tecan Group, Ltd.) at 450
nm. The cells were seeded into 24-well plates (100 cells/well)
coated with 100 μg/ml polylysine (Beyotime Institute of
Biotechnology). After culturing for 2 weeks, the cells were fixed
with 4% paraformaldehyde (Biosharp Life Sciences) for 20 min and
stained with 1% crystal violet (MilliporeSigma) for 15 min at room
temperature. Images of the colonies were captured using a
Fluorescent (enzyme-linked) spot analyser (S6 ULTRA; Cellular
Technology Limited).
Luciferase reporter assay
A total of 1×104 MMQ cells/well were
plated in 96 wells. When confluence reached 70-80%, miR-145-5p
mimics or miR-NC (50 nmol/l) and fluorescent reporter plasmid (50
ng/well, pmirGLO constructs with WT or mutated target sequence)
were co-transfected using Lipofectamine® 3000
(Invitrogen; Thermo Fisher Scientific, Inc.). After 24 h of
transfection, firefly luciferase activity was normalised to
Renilla luciferase activity using the luciferase reporter
assay system (Promega Corporation).
Immunofluorescence assays
Cells were washed three times with PBS, then cells
or tissues were fixed with 4% paraformaldehyde for 20 min at room
temperature and permeabilised with 0.2% Triton X-100 for 10 min at
room temperature. Subsequently, after blocking with 10% goat serum
(MilliporeSigma) at room temperature for 1 h, the cells or tissues
were incubated with corresponding primary antibodies overnight at
4°C. Next, cells or tissues were incubated with a fluorescent
secondary antibody at room temperature for 1 h, followed by
staining with 1 μg/ml DAPI for 5 min. Each step required
three 15 min washes with PBS. Fluorescence images were obtained
using a confocal laser scanning microscope.
RNA-FISH
The circOMA1 hybridisation probe was designed and
synthesised by Shanghai GenePharma, Co., Ltd. RNA-FISH was
performed on circOM cells and tissues using an in-situ
hybridisation kit (Exon Biotech Inc.), according to the
manufacturer's protocol. Images were obtained on a Nikon C2
laser-scanning confocal microscope. The FISH probe was labelled
with digoxigenin, and the sequence was
5'-CACTTGACTACCTGAGGTATAAG-3'.
Vector construction and cell
transduction
circOMA1 small interfering (si)RNA
(5'-ACCUCAGGUAGUCAAGUGATT-3'), miR-145-5p mimic
(5'-GUCCAGUUUCCCAGGAAUCCCU-3'), miR-145-5p inhibitor
(5'-AGGGAUUCCUGGGAAAACUGGAC-3'), and the corresponding negative
controls were designed and synthesised by Beijing Qingke
Biotechnology Co., Ltd. A total of 4×104 cells/well
(circOM or circNC) were plated in a 24-well plate, when the cell
density was ~80%, and DMEM/F12 medium without
penicillin/streptomycin was added. The miR-145-5p mimic (30 nM) and
miR-145-5p inhibitor (50 nM) were transfected using
Lipofectamine® 3000 (cat. no. L3000-015, Thermo Fisher
Scientific, Inc.), and the control group was set up by transfecting
the control vector. After 24 h of transfection at room temperature,
subsequent experiments were performed immediately.
Ubiquitination assay
CircOM or circNC cells were lysed in RIPA lysis
buffer and centrifuged at 10,000 × g for 10 min at 4°C.
Supernatants containing 500 μg total protein were
transferred to fresh 1.5-ml tubes and incubated with the
appropriate anti-DRD2 primary antibody or control IgG with gentle
rotation for 2 h at 4°C. A total of 20 μl of Protein A/G
magnetic beads was added, and the mixtures were rotated overnight
at 4°C. Immunocomplexes were collected by centrifugation at 1,000 ×
g for 5 min at 4°C, washed four times with cold RIPA buffer,
resuspended in 40 μl 1X SDS sample buffer, and boiled for 5
min. Ubiquitin conjugation of DRD2 was detected by western
blotting.
Haematoxylin-eosin and IHC staining
The xenograft tumours were removed, fixed with 4%
paraformaldehyde at room temperature, paraffin-embedded and then
sectioned (4-μm thick). Sections were dewaxed with xylene
and hydrated (by sequential immersion in 100, 95, 85, and 70%
ethanol for 5 min per solution), then stained with H&E or
antigen retrieval (citrate buffer solution) at room temperature.
After the endogenous peroxidase was blocked with 3% hydrogen
peroxide for 10 min, and blocked using 10% goat serum for 30 min at
room temperature, samples were incubated with primary antibody
overnight at 4°C. Finally, the sections were incubated with DAB
(Jiangsu CoWin Biotech Co., Ltd.) and observed under a fluorescence
microscope (DMI4000B; Leica Microsystems GmbH).
Co-culture assay
circOM or circNC cells were co-cultured with the
parental cells using a 0.4-μm Transwell chamber (Corning,
Inc.) for 5-7 days. After co-culture, the sensitivity of parental
cells to CAB was measured using a CCK-8 assay. Separately, 30
μg/ml exosomes from the supernatant of circOM or circNC
cells were co-cultured with parental cells for 24, 48, or 72 h.
Subsequently, parental cells were collected for subsequent
experimental studies.
Exosome labelling
Exosomes from circOM or circNC cell supernatants
were labelled using a diluted PKH67 (a fluorescent dye, Beijing
Fluorescence Biotechnology Co., Ltd.) to ensure a final
concentration of 4 μM. Cells were incubated for 5 min and
gently mixed. Subsequently, 10% FBS without exosomes containing an
excess of dye was diluted with PBS and centrifuged at 4°C, 110,000
× g for 70 min to remove any unbound dye. The supernatant was
discarded, and the exosomes were washed twice in PBS by
centrifugation at 110,000 × g.
Nanoparticle tracking analysis (NTA)
NTA of exosomes derived from circOM or circNC cell
supernatant was performed using a Flow Nanoanalyzer (NanoFCM, U30)
to measure the concentration, size, and distribution of
exosomes.
Transmission electron microscopy
(TEM)
circOM or circNC cells were treated with 2.5%
glutaraldehyde at 4°C. After washing with PBS three times (5 min
per wash), cells were fixed using 1% osmium tetroxide for 2 h at
4°C and washed as aforementioned. Samples were dehydrated with a
gradient of ethanol solutions (50, 70, 80, 90, and 100% for 10 min
each), followed by 100% ethanol for another 10 min. Samples were
dehydrated twice in acetone for 10 min each, then infiltrated with
acetone and embedding solution at 3:1 for 30 min and 1:1 for 4 h,
followed by overnight incubation in pure embedding solution at 4°C.
After embedding, polymerisation was performed at 37°C for 24 h,
followed by 60°C for 48 h. Ultrathin sections of 50-70 nm were
obtained and double-stained with 3% uranyl acetate for 30 min and
lead citrate for 15 min at room temperature. For the exosome assay,
samples were purified from cellular supernatant or plasma via
differential ultracentrifugation, and 20 μl of the
suspension was placed on a clean slide. A copper grid was floated
on the droplet for 2 min to allow for sample adsorption, after
which excess liquid was removed using filter paper to leave a thin
film. The grid was then floated on a droplet of 1% phospho-tungstic
acid for 2 min, wicked with filter paper, and dried passively at
room temperature. All samples were observed and imaged using a TEM
(Tecnai G2 Spirit Twin) at an accelerating voltage of 100 kV.
Xenograft mouse model
A total of 20 female old BALB/c nude mice (4-6 weeks
old; weighing 14-16 g) were purchased from the East Campus of Sun
Yat-sen University and raised in specific pathogen-free conditions,
with a 12/12-h light/dark cycle, with the temperature maintained at
26°C and humidity maintained at 50%. All the animal experiments
were approved by the Animal Ethics Committee of Sun Yat-sen
University (approval no. SYSU-IACUC-MED-2024-B014). circOMA1 was
transfected into MMQ cells using a lentiviral vector (circOM) or a
blank vector (circNC). A total of 1×106 MMQ cells,
circOM, or circNC cells were mixed in DMEM/F12 medium and high
concentration Matrigel at a ratio of 1:1 (Shanghai Yeasen
Biotechnology Co., Ltd.) and were respectively injected
subcutaneously into the right flank or left flank of female BALB/c
nude mice (the total injection volume was 100 μl, containing
1×106). The volume of xenograft tumours was recorded
every 3 days using calliper measurements of length and width, and
the formula volume=length × width/2. After 15 days, the mice were
euthanised using 150 mg/kg sodium pentobarbital, the tumours were
excised, and the weights of the neoplasms were measured
immediately. Based on age and weight, BALB/c nude mice were matched
and assigned to groups with corresponding identification numbers.
Tumour volume and weight were measured in BALB/c nude mice by
investigators blinded to group assignment and numbering. If a
tumour in a BALB/c nude mouse exhibited extensive ulceration or
reached a maximum diameter of >15 mm, the mouse was euthanised
ahead of schedule. For exosome transmission resistance assays,
3×106 MMQ cells were mixed with DMEM/F12 medium and a
high-concentration matrix at a 1:1 ratio, inoculated subcutaneously
into nude mice, and randomly divided into two groups after tumour
formation; the total injection volume was 100 μl. The number
of nude mice in each group was 5. On day 10, 1,1'-Dioctad
ecyl-3,3,3',3'-tetramethylindodicarbocyanine perchlorate (cat. no.
KGMP0025; Nanjing KeyGen Biotech Co., Ltd.) labelled exosomes
derived from circOM or circNC (100 μg/mouse, tail vein
injection) combined with CAB (0.5 mg/Kg, intraperitoneal injection)
were administered. To reduce discomfort, a human insulin syringe
with a thinner needle was used. After 24 h, the distribution of
exosomes in the subcutaneous tumour, brain, and pituitary gland of
nude mice was observed using a microscopic living imaging system
(cat. no. IVM-MS2; IVM Technology). Additionally, macroscopic
live-imaging technology was used to observe exosome distribution in
the hearts, livers, spleens, lungs and kidneys of nude mice (IVIS
Spectrum). Exosomes and CAB were injected twice a week for 2 weeks.
Intravenous tail vein injections of exosomes and CAB administration
were performed by personnel who were not involved in the tumour
modelling procedure. If a BALB/c nude mouse exhibited extensive
ulceration or a tumour reached a diameter of >15 mm, the mouse
was euthanised ahead of schedule. At the end of the procedure, mice
were euthanised and tumours excised for subsequent analysis.
Bioinformatics prediction and database
analysis
The regulatory axis between miR-145-5p and the
target gene KBTBD7 was analyzed through TargetScan (https://www.targetscan.org/vert_72/), miRWalk
(http://mirwalk.umm.uni-heidelberg.de/) and miRDB
(https://mirdb.org/index.html). The
expression levels of plasma exosomal circOMA1 in healthy
individuals, benign conditions, and patients with different types
of tumors were analyzed using the exoRBase V2 database (http://www.exorbase.org/exoRBaseV2/help/toIndex).
Statistical analysis
GraphPad Prism (version 8.0, GraphPad Software,
Inc.; Dotmatics) was used for statistical analyses. Student's
t-test (parametric data) or a paired t-test (non-parametric data)
was used to analyse the differences in statistics between both
groups. IC50 calculations were performed, and bar charts
and line plots were generated using GraphPad. Data are presented as
the mean ± SEM. P<0.05 was considered to indicate a
statistically significant difference.
Results
CircOMA1 expression is upregulated in
drug-resistant PRL-PitNET
The authors' previous gene microarray assay
(32) and bioinformatics
analysis identified circOMA1 as closely associated with PitNETs
occurrence and development (33). To investigate its potential role
in PRL-PitNET resistance and progression, circOMA1 expression was
assessed in drug-sensitive prolactinoma (PRL-PitNET-S) and
drug-resistant prolactinoma (PRL-PitNET-R) tissues, which revealed
significantly higher circOMA1 expression in PRL-PitNET-R tissues
(Fig. 1A). Due to the scarcity
of human-derived PitNET cell lines, MMQ rat prolactinoma cells were
used. MMQ cells stably overexpressing exogenous circOMA1 using a
lentiviral vector were established (termed circOM), with cells
transduced with an empty lentiviral vector serving as the control
(termed circNC).
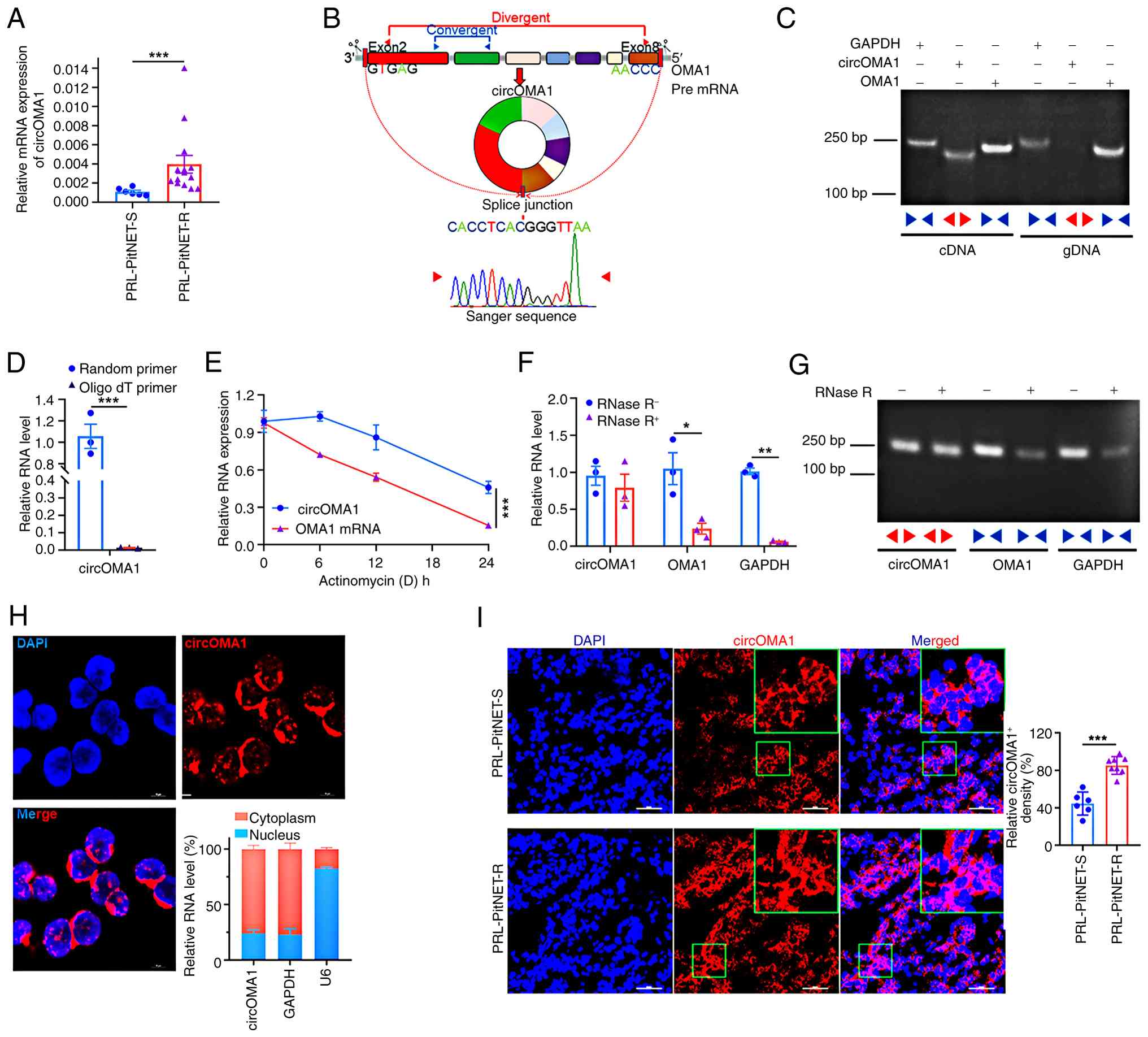
circOMA1 (hsa_circ_0002316) is a 1,381bp transcript
generated by back-splicing of exons 2-8 of the OMA1 gene. Its
head-to-tail splice junction was confirmed by RT-qPCR and Sanger
sequencing (Fig. 1B). Specific
amplification of the splice junction occurred only with cDNA
templates derived from circOM cells, not with genomic DNA (gDNA)
templates (Fig. 1C).
Furthermore, circOMA1 was amplified using random primers but not
oligo(dT) primers, confirming it lacked a polyadenylated tail
(Fig. 1D). To assess stability,
circOM cells were treated with actinomycin D (an RNA synthesis
inhibitor). circOMA1 demonstrated significantly greater stability
compared with linear OMA1 mRNA (Fig.
1E). Additionally, circOMA1 exhibited resistance to RNase R
digestion (which degrades linear RNA), while GAPDH and linear OMA1
mRNA were degraded, as confirmed by RNase R assay and agarose
electrophoresis (Fig. 1F and G).
Collectively, these results confirmed the successful generation of
MMQ cell lines overexpressing circOMA1 and validated its circular
nature and stability. To determine circOMA1 localisation, FISH and
subcellular fractionation were performed. Both methods revealed
predominant cytoplasmic distribution of circOMA1 in circOM cells
(Fig. 1H). Consistent with this,
FISH analysis of PRL-PitNET-S and PRL-PitNET-R tissues showed
circOMA1 localisation patterns similar to those observed in circOM
cells (Fig. 1I).
circOMA1 promotes cell proliferation and
resistance to CAB
To further confirm whether circOMA1 promoted CAB
resistance, cell viability was assessed 72 h after treatment with
increasing CAB concentrations (7.8125-250 μM). Dose-response
analysis revealed that circOMA1 overexpression significantly
increased CAB resistance, with the IC50 values of circOM
(85.04±4.96 μM) exceeding that of circNC controls
(51.17±3.81 μM) (Fig.
2A). For subsequent experiments, 50 μM CAB was used.
Furthermore, the selected drug concentration was similar to or
consistent with those used in previous studies (35-37). Notably, circOMA1 knockdown
reversed CAB resistance in circOM cells (Fig. 2B and C). Furthermore, CCK-8, EdU
incorporation and colony formation assays demonstrated that
circOMA1 overexpression enhanced proliferation and colony formation
in cells treated with CAB compared with the circNC cells (Fig. 2D-F and S1A). Given the pathognomonic
hyperprolactinemia in PRL-PitNET, the effect of circOMA1 on PRL
secretion was examined. circOMA1 overexpression elevated both PRL
mRNA and protein levels (Fig.
S1B-D). This phenotype correlated with increased expression of
pituitary-specific positive transcription factor 1 (Pit-1), the
master transcriptional regulator of PRL in PRL-PitNET (38). Consistently, it was further found
that circOMA1 facilitated the expression of Pit-1 (Fig. S1E).
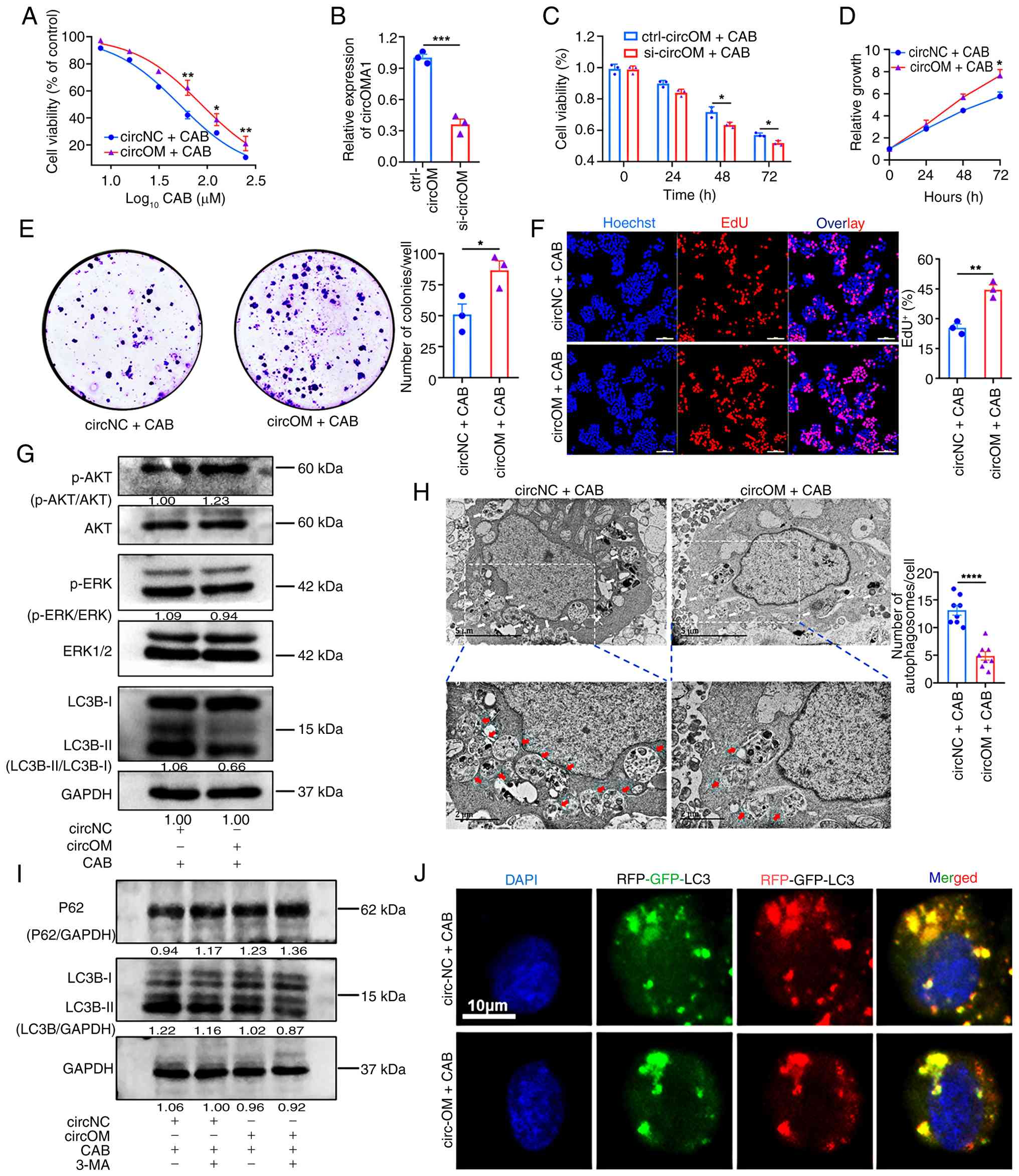 | Figure 2circOMA1promotes CAB resistance in
PRL-PitNET cells through regulation of the autophagy pathway. (A)
CCK-8 determination of IC50 in circOM and circNC cells
after 72 h of treatment with a gradient of CAB concentrations
(R2=0.8611, 0.7030, 0.9248). (B) circOMA1 expression
after 48 h of siRNA targeting. (C) Sensitivity of circOM cells to
50 μM CAB following circOMA1 knockdown
(R2=0.7891, 0.8357). (D-F) Representative images and
quantification (right panels) of CCK-8, colony formation, and EdU
assays used to assess the effect of circNC vs. circOM on CAB
sensitivity. Scale bar, 100 μm. (G) Western blot analysis of
LC3B, p-ERK1/2 and p-AKT in circNC and circOM cells after 72 h of
treatment with 50 μM CAB. (H) Representative transmission
electron microscopy images and quantification of autophagosomes in
circNC (n=8) and circOM (n=8) cells after 72 h of treatment with 50
μM CAB. Scale bars, 5 and 2 μm; arrows indicate
autophagosomes. (I) Western blot analysis of LC3B and p62 in circNC
and circOM cells. (J) Representative confocal images of autophagic
flux in circNC and circOM cells. Scale bar, 10 μm. Data are
presented as the mean ± SEM of at least three repeats. Differences
between groups were compared using an unpaired Student's t-test.
*P<0.05, **P<0.01,
***P<0.001 and ****P<0.0001. CAB,
cabergoline; NC, negative control; circ, circular RNA; PRL-PitNET,
prolactin-secreting pituitary neuroendocrine tumour; siRNA, small
interfering RNA; CCK-8, Cell Counting Kit-8; LC3B,
microtubule-associated protein 1 light chain 3β; p-,
phosphorylated. |
circOMA1 induces CAB resistance by
attenuating autophagy and AKT dephosphorylation
Previous studies have demonstrated that CAB binds to
DRD2 with high affinity, primarily exerting its effects through
autophagy and inhibiting the AKT pathway (39). In the present study, it was
revealed that circOMA1 markedly promoted AKT pathway activation and
reduced the LC3-II/LC3-I ratio in circOM cells following CAB
treatment (Fig. 2G). Consistent
with this, TEM revealed a decrease in the number of autophagosomes
in circOM cells compared with circNC cells after CAB exposure
(Fig. 2H). To further
investigate autophagic flux, western blotting showed decreased
LC3-II/LC3-I and increased P62 (a late autophagy substrate) levels
in circOM cells. This effect was amplified by co-treatment with the
autophagy inhibitor 3-MA (Fig.
2I). Using the RFP-GFP-LC3 reporter assay (yellow puncta
represent autophagosomes and red puncta represent autolysosomes due
to GFP quenching in acidic lysosomes), results indicated fewer
yellow puncta in circOM cells (Fig.
2J). Therefore, circOMA1 induced CAB resistance by attenuating
autophagy and inhibiting the AKT pathway.
circOMA1 promotes the ubiquitination of
DRD2, thereby inducing CAB resistance by upregulating KBTBD7
Previous studies have established that reduced
expression or activity of DRD2 is a major cause of drug resistance
in PRL-PitNET (8-11). Thus, whether circOMA1 promoted
CAB resistance by regulating DRD2 was next assessed. The results
showed that DRD2 protein levels were decreased in circOM (Fig. 3A), but its mRNA levels were not
(Fig. S1F). Previous research
identified that KBTBD7 [a substrate adaptor for the CUL3-RING
ubiquitin (Ub) ligase complex] can specifically bind GABARAP
proteins, ubiquitylate TIAM1 (40), and promote the ubiquitination of
DRD2 (36). In the present
study, accelerated degradation of DRD2 in circOM cells was observed
following treatment with the protein synthesis inhibitor CHX
(Fig. 3B), while accelerated
degradation was reversed by co-treatment with the proteasome
inhibitor MG132 (Fig. 3C).
Furthermore, Co-IP assays revealed significantly increased
ubiquitination of DRD2 protein in circOM cells (Fig. 3D). Similarly, KBTBD7 protein
levels were markedly elevated in circOM (Fig. 3E). Subsequent immunofluorescence
analysis of MMQ cells demonstrated cytoplasmic co-localisation of
KBTBD7 and DRD2 (Fig. 3F), and
Co-IP confirmed their physical interaction (Fig. 3G). Knocking down circOMA1 in
circOM cells downregulated KBTBD7 and PRL expression while
concomitantly upregulating DRD2 levels (Fig. 3H). To further elucidate whether
circOMA1 regulated DRD2-mediated resistance specifically through
KBTBD7, KBTBD7 expression was knocked out using CRISPR/Cas9 in
circOM cells. This resulted in a significant increase in DRD2
expression (Fig. 3I and S2A-C), and a reversal of CAB
resistance (Fig. 3J).
Collectively, these findings indicated that circOMA1 induced CAB
resistance by enhancing KBTBD7 expression, which in turn promoted
DRD2 ubiquitination and degradation. Finally, in prolactinoma
tissues, KBTBD7 expression was markedly higher in CAB-resistant
(PRL-PitNET-R) cases than in controls, whereas DRD2 expression
showed the opposite trend (Fig.
3K). Immunofluorescence also confirmed their co-localisation in
the cytoplasm of these tumour tissues (Fig. 3L).
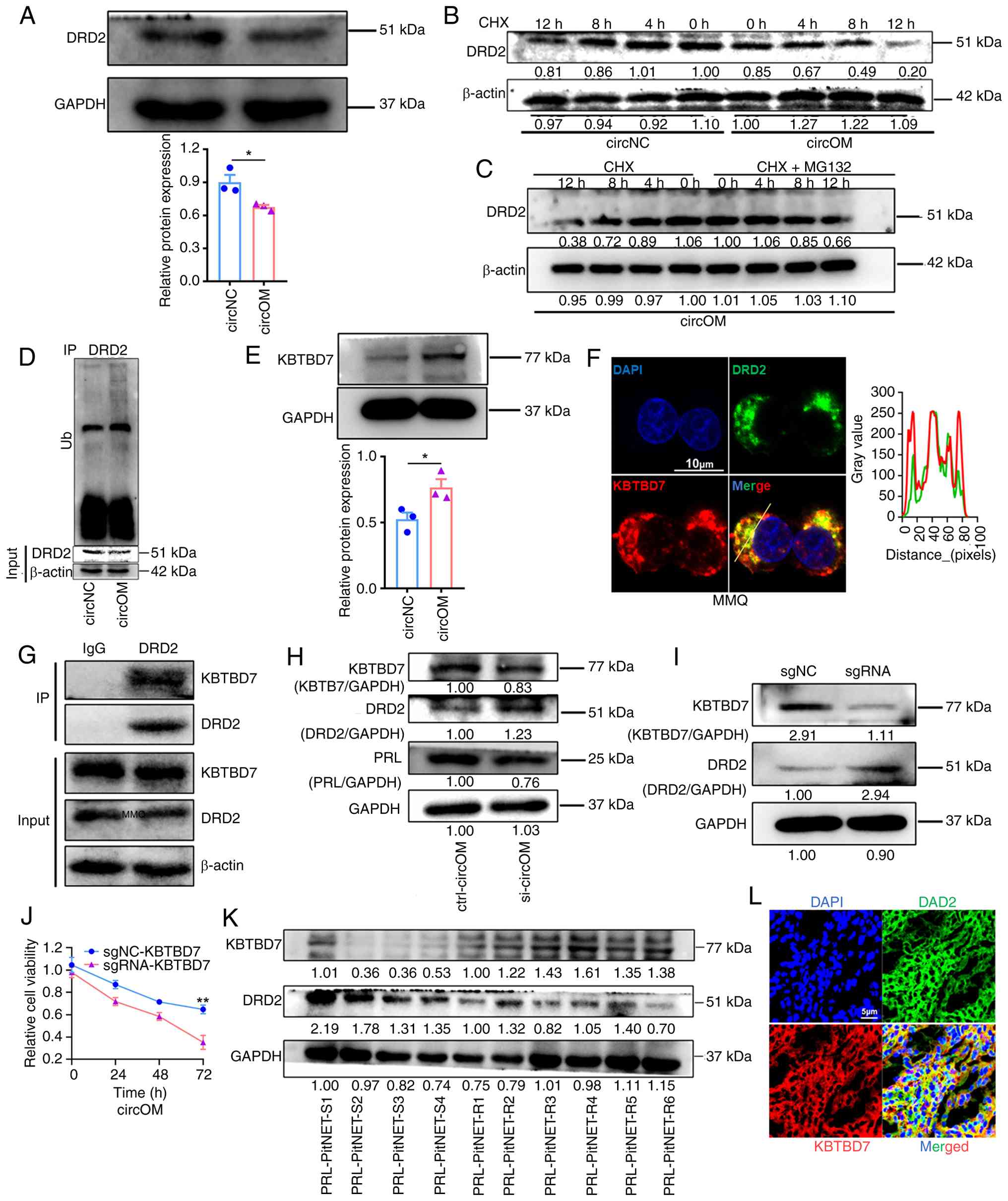 | Figure 3circOMA1 promotes ubiquitin-mediated
degradation of DRD2 and induces CAB resistance in MMQ cells by
upregulating KBTBD7. (A) Western blot detection of DRD2 protein in
circNC and circOM cells (top) with quantification (bottom). (B)
DRD2 protein levels in circNC and circOM cells were measured by
western blotting after treatment with 100 μg/ml CHX for
varying lengths of time. (C) Western blot analysis of DRD2 in
circOM cells at the indicated time points after CHX treatment with
or without treatment with 20 μM MG132. (D) Co-IP analysis of
DRD2 ubiquitination in circNC and circOM cells (green arrow
indicates DRD2). (E) Western blot and densitometric analysis of
KBTBD7 expression in circNC and circOM cells. (F) Representative
immunofluorescence images showing distribution and co-localisation
of DRD2 and KBTBD7 in MMQ cells; the line profile at right
indicates co-localisation. Scale bar, 10 μm. (G) Co-IP
demonstrating the interaction between DRD2 and KBTBD7 in MMQ cells.
(H) Immunoblot analysis of KBTBD7, DRD2 and PRL after circOMA1
knockdown for 72 h in circOM cells. (I) Immunoblot detection of
DRD2 in circOM cells after KBTBD7 knockout by CRISPR/Cas9. (J)
CCK-8 determination of circOM cell sensitivity to CAB after KBTBD7
knockdown (R2=0.9240). (K) Western blot detection of
KBTBD7 and DRD2 in PRL-PitNET-S (n=4) and PRL-PitNET-R (n=6)
tissues. (L) Representative immunofluorescence images showing
distribution and co-localisation of KBTBD7 and DRD2 in PRL-PitNET
tissues. Scale bar, 10 μm. Data are presented as the mean ±
SEM of at least three repeats. Differences between groups were
compared using a unpaired Student's t-test. *P<0.05
and **P<0.01. CAB, cabergoline; NC, negative control;
circ, circular RNA; PRL-PitNET, prolactin-secreting pituitary
neuroendocrine tumour; PRL-PitNET-S, PRL-PitNET-sensitive;
PRL-PitNET-R, PRL-PitNET-resistant; CHX, cycloheximide; MG132,
proteasome inhibitor; CRISPR/Cas9, clustered regularly interspaced
short palindromic repeats/CRISPR-associated protein 9; Co-IP,
co-immunoprecipitation; circ, circular RNA; DRD2, dopamine D2
receptor; KBTBD7, Kelch-repeat and BTB domain-containing protein
7. |
circOMA1 induces CAB resistance by
regulating DRD2 through the miR-145-5p/KBTBD7 axis
The authors' previous research demonstrated that
circOMA1 functioned as a sponge for miR-145-5p (33), and miR-145-5p expression was
significantly reduced in PRL-PitNET-R (41). Consistently, miR-145-5p levels
were lower in circOM (Fig. 4A),
whereas circOMA1 knockdown increased miR-145-5p expression
(Fig. 4B). This finding aligns
with the low miR-145-5p expression observed in PRL-PitNET-R
clinical samples (Fig. 4C).
Therefore, it was hypothesised that miR-145-5p may regulate KBTBD7
expression by targeting its 3' untranslated region (UTR).
Bioinformatics analysis using TargetScan (https://www.targetscan.org/vert_72/), miRWalk
(http://mirwalk.umm.uni-heidelberg.de/) and miRDB
(https://mirdb.org/index.html) identified
potential binding sites for miR-145-5p within the KBTBD7 3'UTR
(Fig. 4D). To validate this
interaction, dual-luciferase reporter plasmids containing either
the wild-type (WT) or mutated (Mut) KBTBD7 3'UTR sequence were
established (Fig. 4E and F;
Fig. S3). Co-transfection with
miR-145-5p significantly reduced luciferase activity for the WT
reporter but not the Mut reporter (Fig. 4G), confirming that miR-145-5p
specifically targeted the KBTBD7 3'UTR.
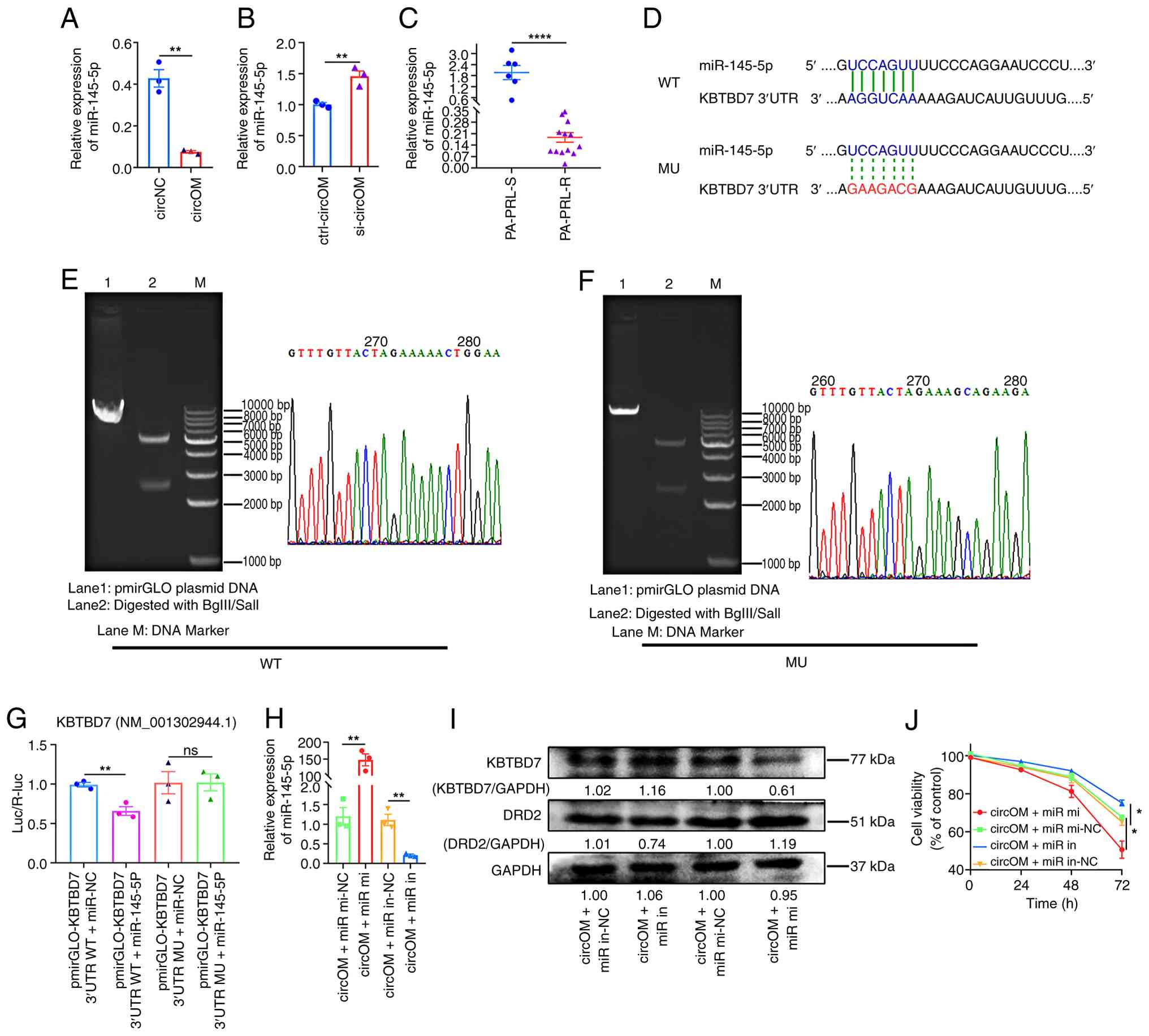 | Figure 4circOMA1 induces CAB resistance in
MMQ cells by regulating DRD2 via a miR-145-5p/KBTBD7 axis. (A)
RT-qPCR measurement of miR-145-5p levels in circNC and circOM
cells. (B) miR-145-5p expression in circOM cells measured using
RT-qPCR 48 h after circOMA1 knockdown. (C) RT-qPCR analysis of
miR-145-5p in PRL-PitNET-S (n=6) and PRL-PitNET-R (n=13) tissues.
(D-F) Bioinformatics prediction of miR-145-5p target sites in
KBTBD7 and schematic of WT and MU dual-luciferase reporter
constructs. (G) Dual-luciferase reporter assay confirming the
miR-145-5p binding site in the KBTBD7 3'-UTR. (H) RT-qPCR of
miR-145-5p in circOM cells 48 h after transfection with miR-145-5p
mimic (miR-mi), mimic negative control (miR-mi-NC), miR-145-5p
inhibitor (miR-in), or inhibitor negative control (miR-in-NC). (I)
Western blot analysis of KBTBD7 and DRD2 protein levels in circOM
cells 72 h after treatment with miR-145-5p mimic or inhibitor. (J)
Cell Counting Kit-8 assay assessing circOM cell sensitivity to 50
μM CAB following treatment with miR-145-5p mimic or
inhibitor (R2=0.8246, 0.8094). Data are presented as the
mean ± SEM of at least three repeats. Differences between groups
were compared using an unpaired Student's t-test.
*P<0.05, **P<0.01 and
****P<0.0001. ns, not significant; CAB, cabergoline;
NC, negative control; circ, circular RNA; PRL-PitNET,
prolactin-secreting pituitary neuroendocrine tumour; PRL-PitNET-S,
PRL-PitNET-sensitive; PRL-PitNET-R, PRL-PitNET-resistant; RT-qPCR,
reverse transcription-quantitative PCR; miR, microRNA; WT,
wild-type; MU, mutant; UTR, untranslated region; KBTBD7,
Kelch-repeat and BTB domain-containing protein 7. |
To further investigate whether circOMA1 induced CAB
resistance via the miR-145-5p/KBTBD7 axis regulating DRD2,
miR-145-5p levels were modulated in circOM cells. Overexpression of
miR-145-5p significantly downregulated KBTBD7, upregulated DRD2,
and enhanced CAB sensitivity in circOM cells. Conversely, knockdown
of miR-145-5p produced the opposite effects (Fig. 4H-J). These results demonstrated
that circOMA1 promoted CAB resistance by acting as a miR-145-5p
sponge, thereby alleviating miR-145-5p-mediated repression of
KBTBD7. Elevated KBTBD7 subsequently enhanced DRD2 ubiquitination
and degradation, ultimately leading to drug resistance.
circOMA1 promotes proliferation and CAB
resistance via exosomes
It has been previously demonstrated that
exon-derived circRNAs are primarily located in the cytoplasm and
are readily encapsulated into exosomes for secretion (42). Consistent with this, circOMA1,
derived from the reverse splicing of an exon, was found to be
predominantly distributed in the cytoplasm of MMQ cells using
RNA-FISH and subcellular fractionation (Fig. 1H). To investigate whether
circOMA1 was secreted via exosomes and whether it influenced
parental tumour cell proliferation, a bilateral tumour model was
established. MMQ cells were injected into the right flank of
4-6-week-old BALB/c nude mice, while circNC or circOM cells were
injected into the contralateral flank (Fig. 5A). circOMA1 expressing tumours
not only grew significantly faster but also promoted the growth of
the contralateral parental MMQ tumours and exhibited higher final
weights (Fig. 5B-D).
Furthermore, circOMA1 was detected in the contralateral parental
tumours opposite the circOM implants, but not in the
circNC-injected mice (Fig. 5E).
These findings suggest that circOMA1 may regulate tumour
progression in vivo via exosomal transfer.
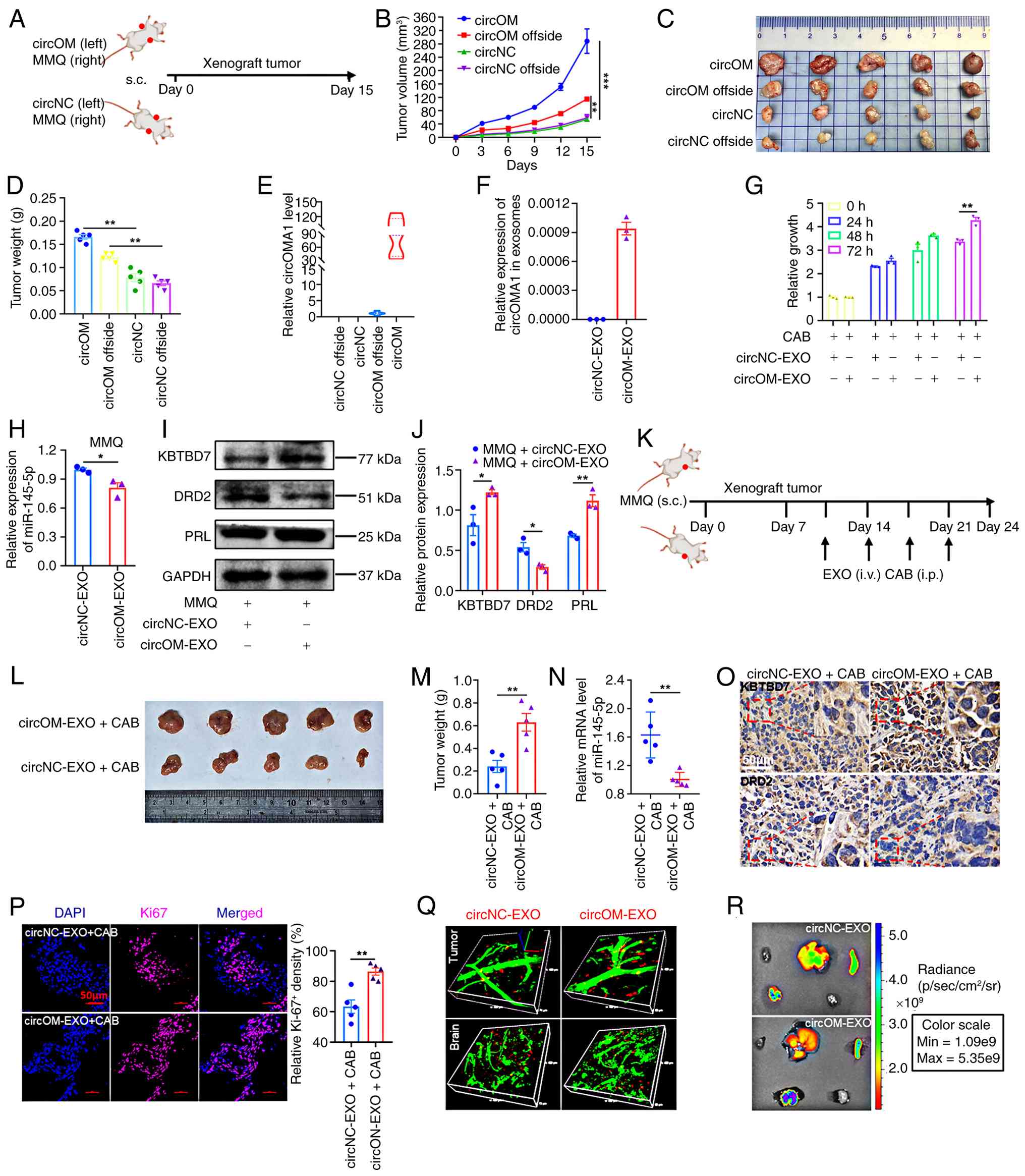 | Figure 5circOMA1 promotes parental cell
proliferation and CAB resistance through exosomes. (A) In
vivo pattern diagram of exosomes promoting parental cell
proliferation. (B) Growth curve of subcutaneous tumours in nude
mice. (C) Excised tumour images (n=5). (D) Mean tumour weight
(n=5). (E) circOMA1 expression in subcutaneous tumours of nude mice
detected using RT-qPCR (n=5). (F) RT-qPCR detection of circOMA1
expression in the exosomes from the supernatant of circNC or circOM
cells. (G) Cell Counting Kit-8 assay of MMQ cell sensitivity to CAB
when co-cultured with circNC-EXO or circOM-EXO (30 μg/ml of
exosomes). (H) RT-qPCR detection of miR-145-5p in MMQ cells after
24 h co-culture with circNC-EXO or circOM-EXO. (I and J) Western
blot detection and densitometric analysis of KBTBD7, DRD2 and PRL
expression in MMQ cells after 72 h co-culture with circNC-EXO or
circOM-EXO. (K) In vivo experimental diagram of exosome
circOMA1-mediated CAB resistance. (L) Excised tumour images (n=5).
(M) Mean tumour weight (n=5). (N) miR-145-5p expression in
subcutaneous tumours of nude mice (n=5). (O) Representative
immunohistochemistry images of KBTBD7 and DRD2 in subcutaneous
tumours of nude mice. Scale bar, 50 μm. (P) Representative
immunofluorescence image and quantification of Ki-67 expression in
the subcutaneous tumours of nude mice. Scale bar, 50 μm. (Q)
Representative microscopic distribution of exosomes in subcutaneous
tumours, and in the brain of nude mice by live imaging assay 24 h
after injection with 100 μg exosomes; green indicates blood
vessels; red indicates exosomes. (R) Representative macroscopic
distribution of exosomes in the heart, liver, spleen, lungs and
kidneys of nude mice by live imaging assay 24 h after injection
with 100 μg exosomes. Data are presented as the mean ± SEM
of at least three repeats. Differences between groups were compared
using an unpaired Student's t-test. *P<0.05,
**P<0.01 and ***P<0.001. CAB,
cabergoline; RT-qPCR, reverse transcription-quantitative PCR; EXO,
exosomes purified from supernatant; KBTBD7, Kelch-repeat and BTB
domain-containing protein 7; DRD2, dopamine receptor D2; PRL,
prolactin. |
Subsequently, a Transwell co-culture system was
used. MMQ cells were plated in the bottom chamber, while circOMA1
or circNC expressing cells were seeded in the upper chamber and
co-cultured for 5 days (Fig.
S4A). MMQ cells co-cultured with circOM cells exhibited
significantly increased resistance to CAB (Fig. S4B). Supporting the exosomal
involvement, the levels of circOMA1 in the conditioned medium
decreased upon treatment with the exosome inhibitor GW4869
(Fig. S4C).
Next, exosomes were isolated and purified from the
supernatant of circOM or circNC cells by differential
ultracentrifugation (Fig. S4D).
Using a nanoflow particle size analyser, the number and size of
exosomes were determined. NTA confirmed the presence of exosomes,
which predominantly ranged in diameter from 50-100 nm. Notably, the
average diameter of exosomes from circOM supernatant (circOM-EXO)
was larger than that from circNC supernatant (circOM-EXO) (Fig. S4E). Characterisation by flow
cytometry using established exosomal markers (43), showed that both circOM-EXO and
circNC-EXO expressed CD9 and CD63 (Fig. S4F), as well as ALG-2-interacting
protein X (ALIX), tumour susceptibility gene 101 protein (TSG101)
and HSP70 (Fig. S4G). TEM
revealed the typical cup-shaped morphology and lipid bilayer
structure of the isolated vesicles, with sizes consistent with NTA
measurements (Fig. S4H).
Critically, RT-qPCR showed circOMA1 expression specifically in
circOM-EXO, but not in circNC-EXO (Fig. 5F). Interestingly, NTA also
indicated that circOM cells secreted significantly more exosomes
than circNC cells (Fig.
S4I).
Given the established role of Rab family GTPases in
exosome biogenesis and release (44-46), their expression was examined;
only Rab27b was significantly upregulated in circOM cells compared
with circNC cells (Fig. S4J).
To assess exosome uptake by recipient cells, MMQ cells were
incubated with PKH67-labelled circOM-EXO or circNC-EXO for 24 h.
Laser confocal microscopy revealed that the labelled exosomes were
internalised by MMQ cells and primarily localised to the cytoplasm
(Fig. S5A). Flow cytometry
further confirmed the uptake of both circOM-EXO and circNC-EXO by
MMQ cells (Fig. S5B).
Finally, to determine if exosomal circOMA1 confers
CAB resistance, purified circOM-EXO or circNC-EXO were directly
co-cultured with MMQ cells. circOM-EXO significantly reduced the
sensitivity of parental MMQ cells to CAB (Fig. 5G). Consistent with this
functional transfer of resistance, EdU incorporation and colony
formation assays yielded similar results (Fig. S6A and B).
Exosomal circOMA1 attenuates autophagy
and AKT dephosphorylation to transmit resistance via the
miR-145-5p/KBTBD7/DRD2 axis
Next, it was investigated whether exosomal circOMA1
also transmitted CAB resistance to recipient MMQ cells by
regulating DRD2 through the miR-145-5p/KBTBD7 axis. First,
co-culture of MMQ cells with circOM-EXO for 24 h significantly
downregulated miR-145-5p levels in the recipient cells (Fig. 5H). Immunoblotting further
revealed that after 72 h of co-culture, circOM-EXO significantly
upregulated KBTBD7 and PRL protein expression while downregulating
DRD2 protein levels in MMQ cells (Fig. 5I and J). Consistent with the
protein data but contrasting with DRD2 mRNA (Fig. S6C), circOM-EXO upregulated Pit-1
mRNA levels (Fig. S6D).
Moreover, following 72 h of co-culture, circOM-EXO significantly
attenuated the CAB-induced dephosphorylation of AKT and autophagic
response in MMQ cells (Fig.
S6E-G).
To validate the in vivo relevance of this
exosome-mediated resistance transmission via the
miR-145-5p/KBTBD7/DRD2 axis, a xenograft tumour model was used. MMQ
cells were subcutaneously inoculated into the right flanks of
4-6-week-old female BALB/c nude mice. Starting at day 10, mice
received intravenous injections of circOM-EXO or circNC-EXO via the
tail vein twice weekly for 2 weeks, combined with CAB treatment.
Mice were euthanised on day 24 (Fig.
5K). The results demonstrated that circOM-EXO significantly
enhanced tumour resistance to CAB, resulting in larger tumours and
higher tumour weights in the circOM-EXO group compared with
controls (Fig. 5L and M).
Supporting the proposed axis, miR-145-5p levels were significantly
decreased (Fig. 5N), and IHC
revealed markedly elevated KBTBD7 protein expression alongside
reduced DRD2 protein in circOM-EXO-treated tumours (Fig. 5O). Further corroborating the
pro-tumorigenic role, circOM-EXO treatment considerably enhanced
tumour cell proliferation, as evidenced by higher Ki-67 expression
(Fig. 5P), and elevated levels
of Pit-1 and PRL (Fig. S6H and
I).
Finally, to track the biodistribution of
administered exosomes in vivo, DID-labelled circNC-EXO or
circOM-EXO were injected intravenously. Macro- and microscopic
in vivo imaging performed 24 h post-injection revealed
exosome enrichment in subcutaneous tumours, pituitary tissue and
brain tissue, with lower abundance in the brain than in pituitary
or tumours (Fig. S7A-C). For
enhanced spatial resolution of exosome localisation within tissues,
blood vessels were visualised using FITC-Dextran 2000. Microscopic
imaging clearly showed internalised exosomes within subcutaneous
tumours and brain tissue (Fig.
5Q). Macroscopic imaging of major organs confirmed strong
exosomal fluorescence signals in the liver, spleen and lungs, but
minimal signals in the heart and kidneys (Fig. 5R).
Exosomal circOMA1 can serve as a
potential prognostic target
A series of previous studies have established that
exosomal circRNAs can serve as potential biomarkers for tumour
prognosis (23,24) and progression (28,29). To investigate the potential of
exosomal circOMA1 as a prognostic biomarker in PitNETs, plasma
exosomes isolated from elbow venous blood of patients with PitNET
stratified by Knosp grade were assessed both pre- and
post-operatively. TEM confirmed the presence of exosomes exhibiting
the characteristic lipid bilayer membrane and cup-shaped
morphology. Crucially, RT-qPCR analysis revealed a significant
decrease in plasma exosomal circOMA1 levels following surgery
(Fig. 6A-D). Analysis of
clinical data also indicated that NF-PitNET had significantly
higher Knosp grades than PRL-PitNET, GH-PitNET and ACTH-PitNET
subtypes (Fig. 6E).
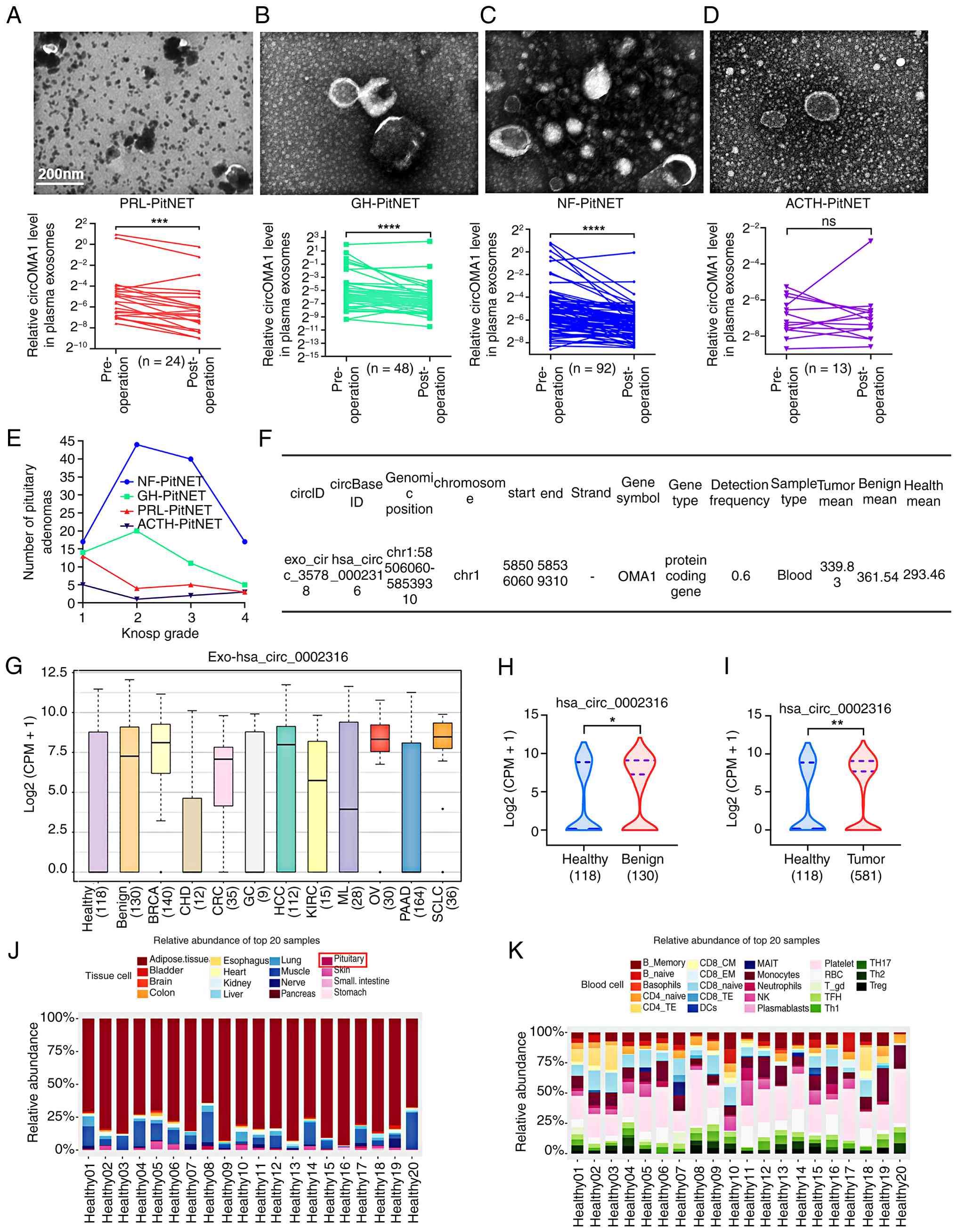 | Figure 6Exosomal circOMA1 can serve as a
potential target for the management and prognosis of PitNETs. (A)
Representative TEM images of plasma exosomes in patients with
PRL-PitNET (top) and preoperative vs. postoperative expression
levels of plasma exosomal circOMA1 (bottom) (n=24). (B)
Representative TEM images of plasma exosomes in patients with
GH-PitNET (top) and preoperative vs. postoperative expression
levels of plasma exosomal circOMA1 (bottom) (n=48). (C)
Representative TEM images of plasma exosomes in patients with
NF-PitNET (top) and preoperative vs. postoperative expression
levels of plasma exosomal circOMA1 (bottom) (n=92). (D)
Representative TEM images of plasma exosomes in patients with
ACTH-PitNET (top) and preoperative vs. postoperative expression
levels of plasma exosomal circOMA1 (bottom) (n=13). (E) Knosp grade
and corresponding number of PitNETs collected. (F) Expression
information of blood exosomal circOMA1 across populations in the
exosome database (http://www.exorbase.org/). (G) Specific expression of
blood exosomal circOMA1 in different population types. (H)
Difference in expression of blood exosomal circOMA1 between healthy
and benign populations. (I) Difference in expression of blood
exosomal circOMA1 between healthy and tumour populations. (J)
Relative expression of circOMA1 in tissue cells of 20 healthy
populations. (K) Relative expression of circOMA1 in blood cells of
20 healthy populations. Data are presented as the mean ± SEM of at
least three repeats. Differences between groups were compared using
a paired Student's t-test (A-D) or unpaired Student's t-test.
*P<0.05, **P<0.01,
***P<0.001 and ****P<0.0001. Scale bar,
200 nm. circRNA, circular RNA; PitNET, pituitary neuroendocrine
tumour; PRL-PitNET, prolactin-secreting PitNET; GH-PitNET, growth
hormone-secreting PitNET; NF-PitNET, nonfunctioning PitNET;
ACTH-PitNET, adrenocorticotropic hormone-secreting PitNET; RT-qPCR,
reverse transcription-quantitative PCR; ns, not significant; TEM,
transmission electron microscopy. |
To further assess the broader prognostic potential
of exosomal circOMA1, its relative expression abundance was
assessed using the exoRBase V2 database (47) (http://www.exorbase.org/exoRBaseV2/help/toIndex)
across healthy individuals, patients with various tumour types, and
individuals with benign conditions. Database analysis demonstrated
that the average expression level of exosomal circOMA1 in blood was
markedly elevated in both tumour-bearing populations and those with
benign conditions compared with healthy controls (Fig. 6F). Furthermore, specific tumour
types (with less variation observed among the three categories) and
benign conditions consistently showed considerably higher blood
exosomal circOMA1 levels than healthy populations (Fig. 6G). Subsequent detailed analysis
corroborated this elevation in tumour and benign populations
relative to healthy individuals (Fig. 6H and I). Analysis of circOMA1
expression in tissue cells from 20 healthy individuals revealed
relatively high abundance in the pituitary gland (Fig. 6J). Additionally, profiling
circOMA1 expression and distribution in blood cells from these
healthy donors indicated predominant enrichment in platelets
(Fig. 6K).
Discussion
PRL-PitNET is the most common PitNET subtype and is
the only subtype for which pharmacological therapy (such as CAB) is
recommended as a first-line treatment. DAs normalise serum PRL
levels, restore gonadal axis function, and reduce tumour dimensions
in 80-90% of patients (48).
Although most PRL-PitNET patients benefit from DA therapy, a subset
exhibits resistance to DAs. Current clinical practice increasingly
favours (CAB, a second-generation DA) for PRL-PitNET treatment due
to its superior efficacy and reduced side-effect profile.
Furthermore, the extended half-life of CAB (63-109 h) allows for
convenient twice-weekly dosing (49). Int the present study, a
previously unrecognised role for circOMA1 in mediating and
propagating CAB resistance in PRL-PitNET was shown. These findings
suggest a novel potential combination therapy strategy for
overcoming CAB resistance in patients with a PRL-PitNET.
The biological functions of circRNAs include acting
as transcriptional regulators, miRNA sponges, modulators of RNA
stability, and interacting with RNA-binding proteins (50). A previous study by the authors
found that circOMA1 bound to DRD2 with a complex formation score
<0.5, suggesting a lack of stable binding capacity (33). In the present study, it was
demonstrated that circOMA1 indirectly regulated DRD2 expression by
sponging miR-145-5p, thereby influencing cellular sensitivity to
CAB. It was previously established that CAB activated DRD2 to
induce autophagy in MMQ cells, a process involving inhibition of
the AKT pathway (39). In the
present study, it was shown that circOMA1 attenuated autophagy and
AKT pathway inhibition, consequently inducing CAB resistance in MMQ
cells. Similarly, CAB also eliminated MMQ cells via the weak
induction of apoptosis and activation of the ERK pathway (39). Consistent with this, it was
observed that circOMA1-induced CAB resistance in MMQ cells involved
attenuated autophagy, accompanied by slight inhibition of the ERK
pathway. Although the majority of circRNAs function as non-coding
RNAs, a previous study revealed that certain circRNAs can be
translated into functional small peptides (33). Current research indicates circRNA
translation occurs under two primary conditions: CircRNA contains
both an open reading frame (ORF) and an internal ribosome entry
site (IRES); and the 5' UTR of the circRNA possesses an m6A
modification that facilitates translation (50). Sequence analysis revealed that
circOMA1 contained one ORF and two IRES elements, indicating its
potential for translation into functional peptides or proteins
(51,52). Although preliminary predictions
indicate that circOMA1 contains an ORF and an IRES, subsequent
steps require the construction of separated fluorescent tags and
tag antibodies targeting both flanking regions of the back-splicing
junction of circOMA1. First, immunofluorescence and western
blotting will be used to observe whether circOMA1 encodes a small
peptide after circularisation. Second, the tagged small peptide
will be identified by mass spectrometry using tag antibody
labelling, followed by in vitro synthesis and functional
phenotypic validation in both cellular and animal models.
Therefore, further investigation is warranted to determine whether
circOMA1 contributes to PRL-PitNET resistance and progression via
its translated products.
Immunoblotting and nanoflow cytometry analyses of
exosomal marker proteins revealed differential protein expression
profiles, consistent with prior research (53). In vivo experiments using
microscopic and macroscopic live-imaging techniques clearly
demonstrated significant enrichment of exosomes in tumours and the
pituitary gland, indicating their ability to readily cross the
blood-brain barrier. Conversely, exosome abundance in the brain
parenchyma was markedly lower, likely due to the restrictive nature
of the blood-brain barrier.
Exosomes represent an emerging, clinically
promising detection modality, often referred to as a 'liquid
biopsy.' This approach offers operational convenience and reduced
patient discomfort compared with traditional tissue biopsy. In the
present study, plasma exosomal circOMA1 levels generally decreased
significantly following surgery in patients with a PitNET.
Conversely, recurrent patients exhibited markedly elevated levels
prior to a second surgery. This pattern likely reflects the
reduction or absence of tumour-derived exosomal circOMA1
post-resection, with the decline being more pronounced in patients
with longer postoperative intervals. Statistical analysis was not
possible for certain patients due to the lack of pre- or
post-operative blood samples. Furthermore, elevated blood exosomal
circOMA1 expression was commonly observed across a range of other
tumour types, while exhibiting relatively high abundance in the
pituitary glands of healthy individuals. The relatively high
expression of circOMA1 in pituitary tissues, along with its
abnormally elevated expression in PRL-PitNETs, suggests that, in
rat prolactinoma models, delivering circOMA1 knockdown molecules
via exosomes may reduce circOMA1 expression in prolactinoma tissues
to a lower level, potentially allowing for improved tumour control.
Moreover, exoRBase database analysis reveals that circOMA1 is
highly abundant in human platelets in peripheral blood, suggesting
that platelet-derived exosomes may represent a feasible approach
for identifying prognostic markers.
The present study adopted a translational research
approach, starting from clinical observations in patients with
prolactinoma, performing rigorous cellular and animal experiments
to elucidate the underlying mechanism, and ultimately using
clinical samples for validation. It was found that exosomal
circOMA1 mediated drug resistance via the miR-145-5p/KBTBD7/DRD2
axis; however, several limitations remain: (i) Further validation
using primary prolactinoma cells is required to confirm the
functional relevance of the circOMA1/miR-145-5p/KBTBD7/DRD2 axis;
(ii) in cellular and animal experiments, while exosome-delivered
circOMA1 was shown to induce drug resistance, the potential
contribution of other exosomal cargo cannot be excluded; (iii)
while the sequences of circOMA1 and miR-145-5p in the rat MMQ cell
line matched those in humans, and the rat and human DRD2 proteins
share 95.71% similarity, potential species differences may still
affect circRNA-miRNA interactions, DRD2 regulation and exosome
biology; (iv) although the CAB concentration used in the in
vitro experiments were consistent with prior reports, its
extrapolation from cellular and animal models to clinical research
requires further optimisation and validation; and (v) the cohort
used to evaluate plasma exosomal circOMA1 as a prognostic biomarker
was relatively small. In addition, although evidence that circOMA1
drives resistance in prolactinomas through exogenous
overexpression/knockdown of circOMA1 was obtained, there is a lack
of further validation on circOMA1 mutations. Despite the
limitations, the present study nonetheless establishes that
exosome-transferred circOMA1 is involved in prolactinoma drug
resistance. Moving forward, multi-centre sample collection and
optimisation of primary cell culture protocols may help overcome
current challenges related to limited surgical samples and slow
cell proliferation. Although postoperative patients showed
significantly reduced plasma exosomal circOMA1 levels, further
studies with larger sample sizes and advanced approaches, such as
artificial intelligence-assisted analysis (based on the levels of
plasma exosomal circOMA1, along with clinical and pathological
diagnoses, machine learning was employed to establish baseline
thresholds for artificial intelligence-assisted diagnosis), are
warranted to strengthen these findings.
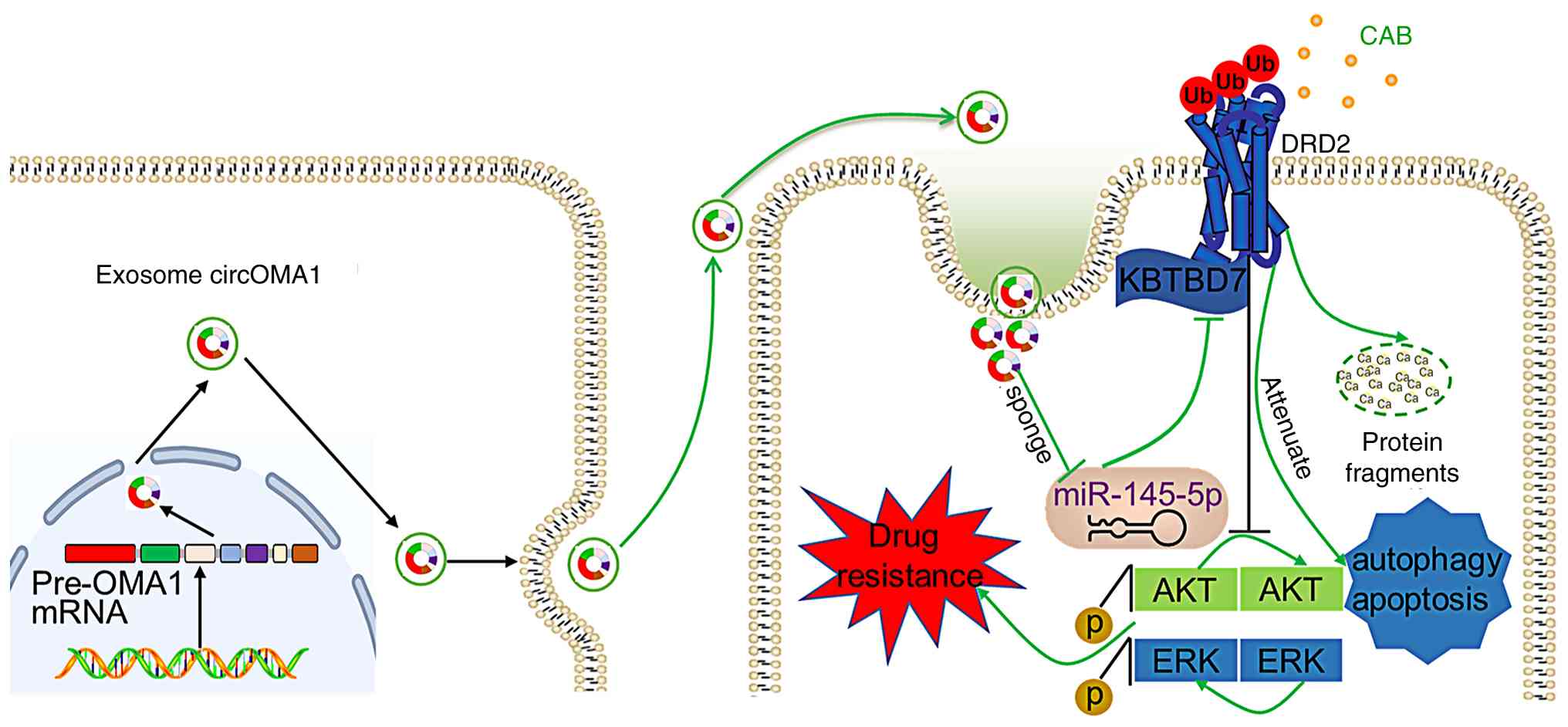
In conclusion, the results of the present study
demonstrated that exosomal circOMA1 is crucial in the development
and propagation of CAB resistance in PRL-PitNET. It was shown that
circOMA1 regulates DRD2 via the miR-145-5p/KBTBD2 axis, thereby
impairing downstream autophagy and inhibiting AKT pathway
suppression, ultimately leading to CAB resistance. Furthermore,
circOMA1 is transported by exosomes, facilitating the transmission
of CAB resistance among MMQ cells both in vitro and in
vivo (Fig. 7). Analysis of a
large cohort of PitNETs samples suggested the potential of exosomal
circOMA1 as a prognostic biomarker.
Supplementary Data
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
QR, DMZ, ZMW, SLZ, WLC and YHZ conceived the study.
QR and YHZ designed the study. QR, XLL, WYH and NW collected the
data. QR wrote the manuscript. QR, SKT and YHZ analysed and
interpretated the data. All authors read and approved the final
version of the manuscript. QR and YHZ confirm the authenticity of
all the raw data.
Ethics approval and consent to
participate
The present study was approved [approval no.
(2020)090] by the Ethical Committee of the First Affiliated
Hospital of Sun Yat-sen University (Guangzhou, China). Written
informed consent was obtained from all patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
The authors would like to thank Dt Zhaoni Wang (The
Third Affiliated Hospital of Sun Yat-sen University), for assisting
in the experimental process, and Dr Qingping Lan (Zhongshan School
of Medicine, Sun Yat-sen University) for providing plasmids.
Funding
The present study was supported by National Natural Science
Foundation of China (grant nos. 82203179 and 82470816), and the
Guangdong Provincial Natural Science Foundation (grant no.
2022A1515011265).
References
|
1
|
Tritos NA and Miller KK: Diagnosis and
management of pituitary adenomas: A review. JAMA. 329:1386–1398.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Trouillas J, Jaffrain-Rea ML, Vasiljevic
A, Raverot G, Roncaroli F and Villa C: How to classify the
pituitary neuroendocrine tumors (PitNET)s in 2020. Cancers (Basel).
12. pp. 5142020, View Article : Google Scholar
|
|
3
|
Villa C, Baussart B, Assié G, Raverot G
and Roncaroli F: The World Health Organization classifications of
pituitary neuroendocrine tumours: A clinico-pathological appraisal.
Endocr Relat Cancer. 30. pp. e2300212023, View Article : Google Scholar
|
|
4
|
Auriemma RS, Pirchio R, De Alcubierre D,
Pivonello R and Colao A: Dopamine agonists: From the 1970s to
today. Neuroendocrinology. 109:34–41. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Bevan JS, Webster J, Burke CW and Scanlon
MF: Dopamine agonists and pituitary tumor shrinkage. Endocr Rev.
13:220–240. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Molitch ME: Pharmacologic resistance in
prolactinoma patients. Pituitary. 8:43–52. 2005. View Article : Google Scholar
|
|
7
|
Maiter D: Management of dopamine
agonist-resistant prolactinoma. Neuroendocrinology. 109:42–50.
2019. View Article : Google Scholar
|
|
8
|
Wu ZB, Zheng WM, Su ZP, Chen Y, Wu JS,
Wang CD, Lin C, Zeng YJ and Zhuge QC: Expression of D2RmRNA
isoforms and ERmRNA isoforms in prolactinomas: Correlation with the
response to bromocriptine and with tumor biological behavior. J
Neurooncol. 99:25–32. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shimazu S, Shimatsu A, Yamada S, Inoshita
N, Nagamura Y, Usui T and Tsukada T: Resistance to dopamine
agonists in prolactinoma is correlated with reduction of dopamine
D2 receptor long isoform mRNA levels. Eur J Endocrinol.
166:383–390. 2012. View Article : Google Scholar
|
|
10
|
Bueno C, Trarbach EB, Bronstein MD and
Glezer A: Cabergoline and prolactinomas: Lack of association
between DRD2 polymorphisms and response to treatment. Pituitary.
20:295–300. 2017. View Article : Google Scholar
|
|
11
|
Pivonello C, Patalano R, Negri M, Pirchio
R, Colao A, Pivonello R and Auriemma RS: Resistance to dopamine
agonists in pituitary tumors: Molecular mechanisms. Front
Endocrinol (Lausanne). 12:7916332021. View Article : Google Scholar
|
|
12
|
Song ZJ, Reitman ZJ, Ma ZY, Chen JH, Zhang
QL, Shou XF, Huang CX, Wang YF, Li SQ, Mao Y, et al: The
genome-wide mutational landscape of pituitary adenomas. Cell Res.
26:1255–1259. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Jiang X and Zhang X: The molecular
pathogenesis of pituitary adenomas: An update. Endocrinol Metab
(Seoul). 28:245–254. 2013. View Article : Google Scholar
|
|
14
|
Dance A: Circular logic: Understanding
RNA's strangest form yet. Nature. 635:511–513. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hu K, Liu X, Li Y, Li Q, Xu Y, Zeng W,
Zhong G and Yu C: Exosomes Mediated Transfer of Circ_UBE2D2
Enhances the Resistance of Breast Cancer to Tamoxifen by Binding to
MiR-200a-3p. Med Sci Monit. 26:e9222532020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Pan Z, Zheng J, Zhang J, Lin J, Lai J, Lyu
Z, Feng H, Wang J, Wu D and Li Y: A novel protein encoded by
exosomal circATG4B Induces oxaliplatin resistance in colorectal
cancer by promoting autophagy. Adv Sci (Weinh). 9:e22045132022.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang X, Chen T, Li C, Li W, Zhou X, Li Y,
Luo D, Zhang N, Chen B, Wang L, et al: CircRNA-CREIT inhibits
stress granule assembly and overcomes doxorubicin resistance in
TNBC by destabilizing PKR. J Hematol Oncol. 15:1222022. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li L, Li W, Chen N, Zhao H, Xu G, Zhao Y,
Pan X, Zhang X, Zhou L, Yu D, et al: FLI1 exonic circular RNAs as a
novel oncogenic driver to promote tumor metastasis in small cell
lung cancer. Clin Cancer Res. 25:1302–1317. 2019. View Article : Google Scholar
|
|
19
|
Pan B, Qin J, Liu X, He B, Wang X, Pan Y,
Sun H, Xu T, Xu M, Chen X, et al: Identification of serum exosomal
hsa-circ-0004771 as a novel diagnostic biomarker of colorectal
cancer. Front Genet. 10:10962019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wen N, Peng D, Xiong X, Liu G, Nie G, Wang
Y, Xu J, Wang S, Yang S, Tian Y, et al: Cholangiocarcinoma combined
with biliary obstruction: An exosomal circRNA signature for
diagnosis and early recurrence monitoring. Signal Transduct Target
Ther. 9:1072024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Liu Y, Li Y, Zang J, Zhang T, Li Y, Tan Z,
Ma D, Zhang T, Wang S, Zhang Y, et al: CircOGDH is a penumbra
biomarker and therapeutic target in acute ischemic stroke. Circ
Res. 130:907–924. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang X, Xu Y, Ma L, Yu K, Niu Y, Xu X,
Shi Y, Guo S, Xue X, Wang Y, et al: Essential roles of exosome and
circRNA_101093 on ferroptosis desensitization in lung
adenocarcinoma. Cancer Commun (Lond). 42:287–313. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
He YD, Tao W, He T, Wang BY, Tang XM,
Zhang LM, Wu ZQ, Deng WM, Zhang LX, Shao CK, et al: A urine
extracellular vesicle circRNA classifier for detection of
high-grade prostate cancer in patients with prostate-specific
antigen 2-10ng/mL at initial biopsy. Mol Cancer. 20:962021.
View Article : Google Scholar
|
|
24
|
Wang J, Zhang Q, Zhou S, Xu H, Wang D,
Feng J, Zhao J and Zhong S: Circular RNA expression in exosomes
derived from breast cancer cells and patients. Epigenomics.
11:411–421. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Théry C, Ostrowski M and Segura E:
Membrane vesicles as conveyors of immune responses. Nat Rev
Immunol. 9:581–593. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Liu Y, Ma L, Hua F, Min Z, Zhan Y, Zhang W
and Yao J: Exosomal circCARM1 from spheroids reprograms cell
metabolism by regulating PFKFB2 in breast cancer. Oncogene.
41:2012–2025. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zheng R, Zhang K, Tan S, Gao F, Zhang Y,
Xu W, Wang H, Gu D, Zhu L, Li S, et al: Exosomal circLPAR1
functions in colorectal cancer diagnosis and tumorigenesis through
suppressing BRD4 via METTL3-eIF3h interaction. Mol Cancer.
21:492022. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ding L, Zheng Q, Lin Y, Wang R, Wang H,
Luo W, Lu Z, Xie H, Ren L, Lu H, et al: Exosome-derived circTFDP2
promotes prostate cancer progression by preventing PARP1 from
caspase-3-dependent cleavage. Clin Transl Med. 13:e11562023.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Yao X, Mao Y, Wu D, Zhu Y, Lu J, Huang Y,
Guo Y, Wang Z, Zhu S, Li X and Lu Y: Exosomal circ_0030167 derived
from BM-MSCs inhibits the invasion, migration, proliferation and
stemness of pancreatic cancer cells by sponging miR-338-5p and
targeting the Wif1/Wnt8/β-catenin axis. Cancer Lett. 512:38–50.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Xiong Y, Tang Y, Fan F, Zeng Y, Li C, Zhou
G, Hu Z, Zhang L and Liu Z: Exosomal hsa-miR-21-5p derived from
growth hormone-secreting pituitary adenoma promotes abnormal bone
formation in acromegaly. Transl Res. 215:1–16. 2020. View Article : Google Scholar
|
|
31
|
Zhang Y, Liu YT, Tang H, Xie WQ, Yao H, Gu
WT, Zheng YZ, Shang HB, Wang Y, Wei YX, et al: Exosome-Transmitted
lncRNA H19 inhibits the growth of pituitary adenoma. J Clin
Endocrinol Metab. 104:6345–6356. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Mao ZG, He DS, Zhou J, Yao B, Xiao WW,
Chen CH, Zhu YH and Wang HJ: Differential expression of microRNAs
in GH-secreting pituitary adenomas. Diagn Pathol. 5:792010.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Du Q, Hu B, Feng Y, Wang Z, Wang X, Zhu D,
Zhu Y, Jiang X and Wang H: circOMA1-Mediated miR-145-5p suppresses
tumor growth of nonfunctioning pituitary adenomas by targeting
TPT1. J Clin Endocrinol Metab. 104:2419–2434. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
35
|
Jian F, Sun Y, Sun Q, Zhang B and Bian L:
NEK2 regulates cellular proliferation and cabergoline sensitivity
in pituitary adenomas. J Cancer. 12:2083–2091. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Liu YT, Liu F, Cao L, Xue L, Gu WT, Zheng
YZ, Tang H, Wang Y, Yao H, Zhang Y, et al: The KBTBD6/7-DRD2 axis
regulates pituitary adenoma sensitivity to dopamine agonist
treatment. Acta Neuropathol. 140:377–396. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wu N, Zhu D, Li J, Li X, Zhu Z, Rao Q, Hu
B, Wang H and Zhu Y: CircOMA1 modulates cabergoline resistance by
downregulating ferroptosis in prolactinoma. J Endocrinol Invest.
46:1573–1587. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Scully KM, Jacobson EM, Jepsen K, Lunyak
V, Viadiu H, Carrière C, Rose DW, Hooshmand F, Aggarwal AK and
Rosenfeld MG: Allosteric effects of Pit-1 DNA sites on long-term
repression in cell type specification. Science. 290:1127–1131.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Tang C, Sun R, Wen G, Zhong C, Yang J, Zhu
J, Cong Z, Luo X and Ma C: Bromocriptine and cabergoline induce
cell death in prolactinoma cells via the ERK/EGR1 and AKT/mTOR
pathway respectively. Cell Death Dis. 10:3352019. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Genau HM, Huber J, Baschieri F, Akutsu M,
Dötsch V, Farhan H, Rogov V and Behrends C: CUL3-KBTBD6/KBTBD7
ubiquitin ligase cooperates with GABARAP proteins to spatially
restrict TIAM1-RAC1 signaling. Mol Cell. 57:995–1010. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Jian M, Du Q, Zhu D, Mao Z, Wang X, Feng
Y, Xiao Z, Wang H and Zhu Y: Tumor suppressor miR-145-5p sensitizes
prolactinoma to bromocriptine by downregulating TPT1. J Endocrinol
Invest. 42:639–652. 2019. View Article : Google Scholar
|
|
42
|
Lyu D and Huang S: The emerging role and
clinical implication of human exonic circular RNA. RNA Biol.
14:1000–1006. 2017. View Article : Google Scholar :
|
|
43
|
Lötvall J, Hill AF, Hochberg F, Buzás EI,
Di Vizio D, Gardiner C, Gho YS, Kurochkin IV, Mathivanan S,
Quesenberry P, et al: Minimal experimental requirements for
definition of extracellular vesicles and their functions: A
position statement from the International Society for Extracellular
Vesicles. J Extracell Vesicles. 3:269132014. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Savina A, Vidal M and Colombo MI: The
exosome pathway in K562 cells is regulated by Rab11. J Cell Sci.
115(Pt 12): 2505–2515. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Hsu C, Morohashi Y, Yoshimura S,
Manrique-Hoyos N, Jung S, Lauterbach MA, Bakhti M, Grønborg M,
Möbius W, Rhee J, et al: Regulation of exosome secretion by Rab35
and its GTPase-activating proteins TBC1D10A-C. J Cell Biol.
189:223–232. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Ostrowski M, Carmo NB, Krumeich S, Fanget
I, Raposo G, Savina A, Moita CF, Schauer K, Hume AN, Freitas RP, et
al: Rab27a and Rab27b control different steps of the exosome
secretion pathway. Nat Cell Biol. 12:19–30, sup 1-13. 2010.
View Article : Google Scholar
|
|
47
|
Qi Y, Xu R, Song C, Hao M, Gao Y, Xin M,
Liu Q, Chen H, Wu X, Sun R, et al: A comprehensive database of
exosome molecular biomarkers and disease-gene associations. Sci
Data. 11:2102024. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Glezer A and Bronstein MD: Prolactinomas.
Endocrinol Metab Clin North Am. 44:71–78. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Curran MP and Perry CM: Cabergoline: A
review of its use in the treatment of Parkinson's disease. Drugs.
64:2125–2141. 2004. View Article : Google Scholar
|
|
50
|
Pisignano G, Michael DC, Visal TH, Pirlog
R, Ladomery M and Calin GA: Going circular: History, present, and
future of circRNAs in cancer. Oncogene. 42:2783–2800. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Song R, Guo P, Ren X, Zhou L, Li P, Rahman
NA, Wołczyński S, Li X, Zhang Y, Liu M, et al: A novel polypeptide
CAPG-171aa encoded by circCAPG plays a critical role in
triple-negative breast cancer. Mol Cancer. 22:1042023. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Huang D, Zhu X, Ye S, Zhang J, Liao J,
Zhang N, Zeng X, Wang J, Yang B, Zhang Y, et al: Tumour circular
RNAs elicit anti-tumour immunity by encoding cryptic peptides.
Nature. 625:593–602. 2024. View Article : Google Scholar
|
|
53
|
Yim KHW, Krzyzaniak O, Al Hrout A, Peacock
B and Chahwan R: Assessing extracellular vesicles in human
biofluids using flow-based analyzers. Adv Healthc Mater.
12:e23017062023. View Article : Google Scholar : PubMed/NCBI
|