Introduction
Pituitary tumors are among the most common
intracranial neoplasms, accounting for 10-15% of all intracranial
tumors (1). Although most
pituitary tumors are benign, they cause complications such as
abnormal hormone secretion, visual field defect and headache, which
impair the quality of life of patients (2). Current treatment strategies for
pituitary tumors include surgical resection, medical therapy and
radiotherapy. However, issues such as high recurrence rate, drug
resistance and side effects persist (3-5).
Therefore, exploring novel molecular targeted therapies for
pituitary tumors is clinically important.
Astragaloside IV (AS-IV) is among the primary active
components of Astragalus membranaceus, a traditional Chinese
herb (6). AS-IV has been
confirmed to possess pharmacological activity, including
anti-inflammatory, antioxidant, immunomodulatory and antitumor
effects (7). AS-IV provides
significant antitumor effects in various tumor models, such as lung
(8), liver (9) and breast cancer (10). However, whether AS-IV inhibits
pituitary tumors and the underlying molecular mechanism remain
unclear. Tubulin β-4B isotype (TUBB4B) is a key member of the
tubulin family and is involved in critical biological processes
such as cell mitosis (11,12), cytoskeleton formation (13) and intracellular transport
(14). TUBB4B is abnormally
expressed in non-small cell lung cancer, Behçet's uveitis, and
colorectal cancer and is associated with tumor proliferation,
invasion and poor prognosis (15-17). In addition, stathmin 1 (STMN1), a
microtubule-destabilizing protein, regulates cell cycle progression
by modulating microtubule dynamics (18). Its expression is enhanced in
triple-negative breast cancer, hepatic fibrosis, and esophageal
cancer and is often associated with the activation of the ERK/MAPK
signaling pathway (19). U0126
is an inhibitor of the ERK/MAPK pathway and can suppress ERK
pathway activation by inhibiting the activity and phosphorylation
of MEK1/2 (20).
The present study aimed to investigate the effects
of AS-IV on the proliferation and apoptosis of pituitary tumor
cells and explore its antitumor mechanism by targeting TUBB4B to
regulate the STMN1/ERK signaling pathway. Through cell experiments,
transcriptome sequencing, molecular and dynamics simulation and
nude mouse xenograft models, the present study aimed to verify the
key role of the AS-IV-regulated TUBB4B-STMN1-ERK axis in driving
the development and progression of pituitary tumors, which may
provide a novel theoretical basis and potential strategy for
targeted therapy of pituitary tumors.
Materials and methods
Materials
The suppliers for all materials are provided in
Table I.
 | Table ISuppliers of materials. |
Table I
Suppliers of materials.
| Name | Supplier | Cat. no. |
|---|
| GH3 complete
medium | Boster Biological
Technology | ZYPYG0264 |
| Ham's F-12K | Thermo Fisher
Scientific, Inc. | 21127030 |
| Horse serum | Thermo Fisher
Scientific, Inc. | 16050122 |
| Puromycin | MCE | HY-K1057 |
| Fluorescence
microscope | Nikon
Corporation | ECLIPSE Ts2R |
| Total RNA kit
I | Omega Bio-Tek,
Inc. | R6834-01 |
| NanoDrop One | Thermo Fisher
Scientific, Inc. | 840-317400 |
| cDNA Synthesis
kit | Thermo Fisher
Scientific, Inc. | K1622 |
| UltraSYBR Mixture
PCR | Cwbio | CW0957H |
| Gentier 96R
RT-qPCR | Xi'an Tianlong
Science and Technology Co., Ltd. | TL22R221038785 |
| BALB-c nude
mice | Hunan SJA
Laboratory Animal Co., Ltd. | Not applicable |
|
Paraformaldehyde | Biosharp Life
Sciences | BL539A |
| Triton X-100 | Beijing Solarbio
Science & Technology Co., Ltd. | T8200 |
| TUNEL | Beyotime
Biotechnology | C1086 |
| DAPI | Beyotime
Biotechnology | C1005 |
| EdU Imaging
kit | APeXBIO | K1075 |
| Crystal violet | Beijing Solarbio
Science & Technology Co., Ltd. | G1062 |
| Annexin
V-PE/7-AAD | BD Biosciences | BD559763 |
| NovoCyte | ACEA Biosciences,
Inc. | None |
| RIPA | Beyotime
Biotechnology | P0013B |
| NP-40 lysis
buffer | Beijing Solarbio
Science & Technology Co., Ltd. | N8032 |
| TBS | Wuhan Servicebio
Technology Co., Ltd. | G0001 |
| Goat anti-rabbit
IgG (H+L) secondary antibody, HRP | Thermo Fisher
Scientific, Inc. | C31460100 |
| Goat anti-Mouse IgG
(H+L) secondary antibody, HRP | Thermo Fisher
Scientific, Inc. | 31430 |
| TUBB4B
antibody | Boster Biological
Technology | BM4264 |
| PCNA antibody | Boster Biological
Technology | BM3888 |
| Bcl-2 antibody | Abmart
Pharmaceutical Technology Co., Ltd. | T40056 |
| Bax antibody | Abmart
Pharmaceutical Technology Co., Ltd. | T40051 |
| Pro-caspase3
antibody | Boster Biological
Technology | M00334-9 |
| Cleaved caspase3
antibody | Abmart
Pharmaceutical Technology Co., Ltd. | TA7022 |
| Total PARP
antibody | Abmart
Pharmaceutical Technology Co., Ltd. | P79881 |
| Cleaved PARP
antibody | Abmart
Pharmaceutical Technology Co., Ltd. | T55035 |
| STMN1 antibody | Abmart
Pharmaceutical Technology Co., Ltd. | PA4263 |
| p-STMN1
antibody | Abmart
Pharmaceutical Technology Co., Ltd. | TP56510 |
| cPLA2 antibody | Abmart
Pharmaceutical Technology Co., Ltd. | PA3288 |
| p-cPLA2
antibody | Abmart
Pharmaceutical Technology Co., Ltd. | PA3287 |
| CCND1 antibody | Boster Biological
Technology | PB0403 |
| ERK antibody | Abmart
Pharmaceutical Technology Co., Ltd. | T40071 |
| p-ERK antibody | Abmart
Pharmaceutical Technology Co., Ltd. | T40072 |
| JNK antibody | Abmart
Pharmaceutical Technology Co., Ltd. | T40073 |
| p-JNK antibody | Abmart
Pharmaceutical Technology Co., Ltd. | T40074 |
| GAPDH antibody | Boster Biological
Technology | A00227 |
| Tween-20 | Beyotime
Biotechnology | ST825 |
| BSA | BioFroxx;
neoFroxx | 4240GR100 |
| Rapid blocking
solution | Epizyme | PS108P |
| Stripping
buffer | Cwbio | CW0056M |
| ECL solution | Biosharp
Lifesciences | BL520A/B |
| Goat anti-rabbit
IgG (H+L) Secondary antibody, Alexa Fluor-488 | Wuhan Sanying
Biotechnology | SA00013-2 |
| Phalloidin | Suzhou UYiLandi
Biotechnology Co., Ltd. | YP0052 |
| Hoechst-33342 | MCE | HY-15559A |
| HRP-conjugated goat
anti-mouse/rabbit IgG | Wuhan Servicebio
Technology Co., Ltd. | TBAG0036 |
| Hematoxylin | Wuhan Servicebio
Technology Co., Ltd. | G1004 |
| Eosin | Wuhan Servicebio
Technology Co., Ltd. | G1001 |
| Citric acid | Wuhan Servicebio
Technology Co., Ltd. | G1201 |
| Endogenous
peroxidase blocking buffer | Beyotime
Biotechnology | P0100A |
| IHC secondary
antibody | Wuhan Servicebio
Technology Co., Ltd. | G1303 |
| DAB | Wuhan Servicebio
Technology Co., Ltd. | G1212 |
| U0126 | MCE | HY-12031A |
Clinical samples
A total of three human pituitary tumor specimens
were collected (November 2024 to June 2025) during neurosurgery at
the First Affiliated Hospital of Shihezi University, Shihezi,
China. The present study was approved by the Scientific and
Technological Ethics Committee of the First Affiliated Hospital of
Shihezi University (approval no. KJ2024-476-01). Patients with
pituitary tumors confirmed by preoperative computed tomography and
postoperative pathology were included Exclusion criteria were as
follows: Patients with incomplete clinical data; patients who
undergone radiotherapy or chemotherapy prior to surgery; samples
with poor tissue preservation or insufficient RNA/protein quality.
All patients signed a written informed consent form. Patient
information is summarized in Table
II.
 | Table IIPatient information. |
Table II
Patient information.
| No. | Age, years | Sex | Knosp grade | Subtype of
Pituitary Tumors |
|---|
| 1 | 46 | Male | IV | GH |
| 2 | 53 | Male | IV | GH |
| 3 | 37 | Female | II | P |
Cell culture
Rat pituitary tumor cell lines (GH3 and MMQ) were
obtained from Wuhan Procell Biotechnology Co., Ltd. Both GH3 and
MMQ cells were cultured in a GH3-specific complete medium (82.5%
Ham's F-12K medium, 15% horse serum, 2.5% FBS and 1%
penicillin-streptomycin) in a sterile, humidified incubator at 37°C
with 95% air and 5% CO2. GH3 cells grew in a loosely
adherent manner, while MMQ cells grew in suspension. When
confluency reached 70-80%, the cells were passaged at a ratio of
1:3 to maintain exponential growth.
Sequencing and bioinformatics
analysis
AS-IV was purchased from MedChemExpress (purity,
99.93%) and dissolved in DMSO according to the manufacturer's
instructions. GH3 cells were divided into the AS-IV treatment group
(n=6) and DMSO-treated negative control group (NC, n=3).
Transcriptomics sequencing was performed at Hangzhou Lianchuan
Biotechnology Co., Ltd. RNA was isolated using TRIzol (Invitrogen;
Thermo Fisher Scientific, Inc.). The total RNA was quantified and
quality-controlled using a NanoDrop ND-1000 (NanoDrop). rRNA was
removed using the Epicentre Ribo-Zero Gold Kit (Illumina, Inc.),
and cDNA was synthesized from the fragmented RNA using reverse
transcriptase (Invitrogen SuperScript™ II Reverse Transcriptase,
cat. no. 1896649). Then, E. coli DNA polymerase I (NEB, cat.
no. m0209) and RNase H (NEB, cat. no. m0297) were used for
double-strand synthesis to convert the composite double-stranded
RNA-DNA hybrid into double-stranded DNA. The double-stranded DNA
was digested with UDG (NEB, cat. no. m0280), followed by PCR
(pre-denaturation at 95°C for 3 min; 8 cycles of denaturation at
98°C for 15 sec each; annealing at 60°C for 15 sec; extension at
72°C for 30 sec, followed by a final extension at 72°C for 5 min)
to generate a library consisting of fragments ranging from 300±50
bp. Finally, the library was sequenced using Illumina Novaseq™ 6000
(LC Bio Technology Co., Ltd.) in a paired-end (PE150) sequencing
mode. The library concentration was measured using a Qubit
fluorometer (Thermo Fisher Scientific, Inc.) and a High Sensitivity
DNA Chip on a Bioanalyzer 2100 (Agilent Technologies), with the
final loading concentration adjusted to 2 nM for sequencing.
Low-quality sequences in the raw sequencing data
were filtered out via Cutadapt (v1.9; cutadapt.readthedocs.io/).
The processed data were used for transcriptome assembly and
quantitative analysis. Differentially expressed genes (DEGs) were
screened with the limma package (v3.5x; bioinf.wehi.edu.au/limma/) in R (v4.2.1; r-project.org/) with the threshold defined as
fold-change (FC) ≥2 or ≤0.5 and q-value <0.05
(|log2FC|≥1 and q<0.05). Gene Ontology (GO)
enrichment analysis (geneontology.org/), Gene Set Enrichment Analysis
(GSEA; gsea-msigdb.org/), Kyoto Encyclopedia of
Genes and Genomes (KEGG) pathway enrichment (kegg.jp/) and
protein-protein interaction (PPI) analysis (string-db.org/) were performed using WebGestalt
(webgestalt.org/). Partial bioinformatics data
were obtained from the Gene Expression Omnibus (GEO) database
(accession no. GSE136781; ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE136781) and
the STRING database (cn.string-db.org; TUBB4B and STMN1).
Lentivirus-mediated TUBB4B overexpression
(OE) and knockdown (KD)
The lentiviral vectors used for OE included
LV5(EF-1α/GFP&Puro)-TUBB4B (OE-TUBB4B) and LV5
(EF-1α/GFP&Puro)-NC (OE-shNC; both 1×109 TU/ml). The
lentiviral vectors used for knockdown LV3-GFP&Puro-TUBB4B-141
(KD1-TUBB4B), LV3-GFP&Puro-TUBB4B-199 (KD2-TUBB4B) and
LV3-GFP&Puro-NC (KD-shNC; all 1×108 TU/ml) and
polybrene were purchased from Suzhou GenePharma Co., Ltd. The
sequences of lentiviral vectors are listed in Table III. The lentiviral packaging
system used in this study is a third-generation system. Lentiviral
particles were prepared in 293T cells (purchased from GenePharma).
For transfection, 10 μg of lentiviral transfer plasmids
(expressing TUBB4B or shRNA) were mixed with the packaging plasmids
pGag/Pol, pRev, and pVSV-G in a 4:2:2:1 ratio and transfected using
Lipofectamine 3000 (Thermo Fisher Scientific, Inc.) at 37°C for 48
h. After transfection, the supernatant containing lentiviral
particles was collected, filtered through a 0.45 μm filter,
and concentrated by ultracentrifugation. The viral titer was
determined by qPCR and expressed in TU/ml.
 | Table IIILentivirus-mediated RNA sequences for
stable transfection. |
Table III
Lentivirus-mediated RNA sequences for
stable transfection.
| Name | Sequence,
5'➔3' | Note |
|---|
| Vector |
TAATATCCCTCTTTAAGTGT | Non-targeting
sequence inserted into the lentiviral backbone JLVO-CAG-GFP-Apuro
(9771 bp) |
| OE-TUBB4B | NM_199094.3 (1338
bp) | Full-length coding
sequence of TUBB4B |
| shNC |
TTCTCCGAACGTGTCACGT | Scrambled
non-targeting shRNA |
| KD1-TUBB4B |
GCGACGAGCATGGCATTGATC | shRNA targeting
TUBB4B |
| KD2-TUBB4B |
GGAGCGCATCAACGTGTACTA | shRNA targeting
TUBB4B |
GH3/MMQ cells were seeded into 6-well plates at a
density of 2×105 cells/well. When cell confluency
reached 70%, lentiviral vectors and polybrene were added to 6-well
plates containing complete medium (MOI for the GH3 cell line was
~50, and for the MMQ cell line, ~100) and allow transfection to
proceed for 24-48 h at 37°C and 5% CO2. After 72 h, the
cells were selected with 1 μg/ml puromycin for 48 h at 37°C
and 5% CO2 to establish stably transfected cell lines,
and 1 μg/ml puromycin was used to maintain the continued
culture of stably transfected cells. The transfection efficiency of
the stably transfected cell lines was verified by reverse
transcription-quantitative (RT-q)PCR and western blotting. Cells
stably transfected for up to 2 weeks were used for subsequent
experiments.
RT-qPCR
Total RNA was extracted from the GH3/MMQ cells using
Total RNA kit I, and the RNA concentration was quantified with a
NanoDrop One device. cDNA was synthesized via one-step RT using the
RevertAid First Strand cDNA Synthesis kit following the
manufacturer's instructions. Target cDNA fragments were amplified
and detected using UltraSYBR Mixture PCR reagents on a Gentier 96R
RT-qPCR system (Xi'an Tianlong Technology Co., Ltd.) according to
the manufacturer's instructions. The thermocycling conditions were
as follows: initial denaturation at 95°C for 10 min, followed by 40
cycles of 95°C for 15 sec and 60°C for 60 sec). GAPDH was adopted
as the internal reference to normalize the relative expression of
target mRNAs, and the 2−∆∆Cq method (21) was employed for quantitative
analysis of relative nucleic acid expression. All primer sequences
are listed in Table IV.
 | Table IVPrimer sequences for reverse
transcription-quantitative PCR. |
Table IV
Primer sequences for reverse
transcription-quantitative PCR.
| Gene | Forward primer,
5'➔3' | Reverse primer,
5'➔3' |
|---|
| TUBB4B |
ATTGATCCCACTGGCACGTA |
TCACAACGTCCAACACCGAG |
| GAPDH |
GACATGCCGCCTGGAGAAAC |
AGCCCAGGATGCCCTTTAGT |
| CCNA1 |
TGAACAGGGGGACAGAGACA |
GAGTCAACCAGCATTGGGGA |
| CCNB1 |
ATTGCAGCTGGGGCTTTTTG |
AGAGATTCCTCCGTGTGGGA |
| CCND1 |
CAAGTGTGACCCGGACTGC |
GCAAGCCAGACCAGCTTCTT |
| CCNE1 |
CGTTTAAGCCCCCTGACCAT |
CACTTCTCCCGTGTCGTTGA |
| CDK1 |
TATCCCTCCTGGCCAGTTCA |
GGTACCACAGCGTCACTACC |
| CDK2 |
AAATCCGGCTCGACACTGAG |
TCCAGCAGCTTGACGATGTT |
| CDK4 |
GATGCGCCAGTTTCTAAGCG |
AGGGCCATCTGGTAGCTGTA |
| CDK6 |
GTGGACCTCTGGAGTGTTGG |
CCACGTCTGAACTTCCACGA |
BALB/c nude mouse model
establishment
The animal experiments were approved by the
Bioethics Committee of Shihezi University, Shihezi, China (approval
no. A2024-353). Thirty female BALB/c nude mice (age, 3-4 weeks old;
weight, 15-18 g at the start of the study) were purchased from
Helilai Co., Ltd. and housed in a specific pathogen-free
environment (temperature, 24±2°C; humidity, 50-60%; 12 h light/dark
cycle) with ad libitum access to sterilized food and water.
At the start of the experiment, each nude mouse was injected with
2×106 GH3 cells suspended in 100 μl PBS into the
left axilla. Mice in the Vector + AS-IV, OE-TUBB4B + AS-IV and
KD2-TUBB4B+AS-IV) with n=5 per subgroup, were intraperitoneally
injected with AS-IV at a half-maximal inhibitory concentration
(IC50) dose of 10-20 mg/kg daily, whereas the mice in
the control group, consisting of three subgroups (Vector,
OE-TUBB4B, and KD2-TUBB4B) with n=5 per subgroup (total n=15),were
intraperitoneally injected with the same volume of normal saline
daily. Humane endpoints, including tumor diameter exceeding 15 mm,
significant weight loss, and visible signs of tumor ulceration or
infection. The body weight and the length/width of the subcutaneous
tumors were measured every 3 days. The mice were euthanized by
cervical dislocation following 21 days of continuous feeding; death
was confirmed by the absence of a heartbeat, cessation of breathing
and fixed, dilated pupils. Tumor samples were weighed and stored
for further analysis.
Cell viability assay
GH3/MMQ cells were seeded into 96-well plates at a
density of 8×103 cells/well and cultured in a 37°C for
24 h. The cells were cultured in 100 μl GH3 complete medium
containing AS-IV at (0 to 140 μM in a 37°C, 5%
CO2 incubator for 0-72 h. At 0, 24, 48, and 72 h, 10
μl Cell Counting Kit 8 (CCK-8) reagent and 90 μl F12K
medium were added to each well, followed by incubation for 2 h. The
optical density at 450 nm was measured using a microplate
reader.
To investigate the mechanism of TUBB4B, GH3 cells
were transduced with lentivirus to establish OE-TUBB4B and Vector
stable cell lines. Following the manufacturer's instructions, the
cells were treated with 60 nM U0126 (a MEK/ERK pathway inhibitor)
at 37°C with 5% CO2 for 48 h, after which cell viability
was assessed using the CCK-8 assay.
TUNEL assay for apoptosis detection
GH3 cells were seeded into 24-well plates at a
density of 1×105 cells/well. After incubating for 24 h
at 37°C and 5% CO2, the GH3 complete medium was replaced
with GH3 complete medium containing AS-IV ranging from 0 to 160
μM, and the cells were cultured at a 37°C for 48 h. The
cells were washed three times with PBS and 500 μl 4%
paraformaldehyde was added for 15 min at room temperature. The
cells were permeabilized with 0.4% Triton X-100 for 10 min at room
temperature. A TUNEL apoptosis detection kit was used according to
the manufacturer's instructions. The nuclei were stained at room
temperature in the dark for 20 min with DAPI at a final
concentration of 1 μg/ml. Finally, the slides were mounted
with 50% glycerol in PBS, cells were observed using a fluorescence
microscope and the number of TUNEL-positive cells was counted using
ImageJ software (v1.54p; National Institutes of Health).
EdU assay for proliferation
detection
GH3 cells were seeded into 24-well plates at a
density of 1×105 cells/well. After incubating for 24 h
at 37°C, the medium was replaced with a GH3 complete medium
containing 112.3 μM AS-IV, and the cells were cultured at
37°C for 48 h. An EdU Imaging kit was used according to the
manufacturer's instructions to stain nuclei in the S phase. Nuclei
were stained at room temperature in the dark for 20 min with DAPI
at a final concentration of 1 μg/ml. The results were
observed using a fluorescence microscope, and the number of
EdU-positive cells was counted using ImageJ software.
Colony formation assay
GH3 cells were seeded into 6-well plates at a
density of 500 cells/well. The medium was changed every 3 days, and
the cells were cultured at 37°C for 12-15 days. At room
temperature, a total of 1 ml 4% paraformaldehyde was added to each
well to fix the cells for 15 min, followed by staining with 0.1%
crystal violet for 15 min. Excess crystal violet was removed by
washing with water. Colonies were defined as clusters containing
>50 cells. Images were captured using a light microscope, and
the number of colonies was counted using ImageJ software.
Flow cytometry for apoptosis and cell
cycle detection
The cells from the Vector, Vector + AS-IV,
OE-TUBB4B+AS-IV, and KD2-TUBB4B+AS-IV groups were collected,
resuspended in precooled PBS at a density of 1×106
cells/ml and centrifuged at 400 × g and 4°C for 5 min. The
supernatant was discarded, and the cells were resuspended in 200
μl PBS. A total of 5 μl Annexin V-PE and 5 μl
7-AAD apoptosis detection reagents were added and incubated at 4°C
in the dark for 30 min. Following addition of 300 μl PBS,
detection was performed using an ACEA NovoCyte (Agilent
Technologies) flow cytometer. The total number of cells undergoing
early and late apoptosis was calculated, using NovoExpress software
(v1.5.6; Agilent Technologies).
Western blotting
GH3/MMQ cells and tumors from the nude mice were
collected. Protein samples were extracted using RIPA lysis buffer,
and the protein concentration was quantified with an enhanced
Bicinchoninic Acid Protein Assay kit. 50 μg proteins were
loaded into each lane; after separation by 10% SDS-PAGE and
transferred to a 0.45 μm PVDF membrane. The PVDF membrane
was blocked with 5% (w/v) non-fat dry milk in Tris-buffered saline
containing 0.1% Tween-20 at room temperature for 1 h, incubated
with the primary antibody at 4°C overnight and incubated with the
corresponding horseradish peroxidase-conjugated secondary antibody
at room temperature for 1 h. Western blots were visualized using
ECL reagent, and analyzed using ImageJ software. Antibodies were as
follows: TUBB4B 1:1,000; PCNA 1:2,000; Bcl-2 1:1,000; Bax 1:1,000;
pro-caspase-3 1:1,000; cleaved-caspase-3 1:1,000; T-PARP 1:1,000;
cleaved-PARP 1:1,000; STMN1 1:500; p-STMN1 1:1,000; cPLA2 1:1,000;
p-cPLA2 1:500; CCND1 1:1,000; ERK 1:1,000; p-ERK 1:1,000; JNK
(1:1,000); p-JNK 1:1,000; GAPDH 1:20,000; Goat anti-rabbit
1:20,000; Goat anti-mouse 1:20,000.)
Hematoxylin and eosin (H&E) and
immunohistochemical (IHC) staining
Human pituitary and mouse tumor specimens were fixed
with a 4% paraformaldehyde solution at room temperature for 24 h,
followed by paraffin embedding and serial sectioning (thickness, 4
μm). For H&E staining, the sections were rehydrated and
stained with hematoxylin solution (0.5%) for 3-5 min, followed by
eosin Y solution (0.5%) for 15-20 sec at room temperature. For IHC
staining, the sections were rehydrated and antigen retrieval was
performed in high-temperature citric acid solution at 160°C for 10
min and washed with PBS. Sections were blocked with 3%
H2O2 at room temperature for 10 min, followed
by blocking with PBS containing 5% BSA at room temperature for 30
min. Primary antibodies against TUBB4B (1:100), PCNA (1:100), Bcl-2
(1:200), Bax (1:100) and cleaved-caspase-3 (1:200) were added at
4°C in the dark overnight. The sections were incubated with a
universal secondary antibody (HRP-conjugated goat anti-mouse/rabbit
IgG, Servicebio; catalog No. TBAG0036) at a dilution of 1:200 at
room temperature for 1 h. DAB chromogen was added for 20 min. For
nuclear counterstaining, the sections were stained with hematoxylin
at room temperature for 1-2 min. The sections were observed and
photographed using a light microscope (Olympus BX53).
Immunofluorescence staining
GH3 cells stably transfected with lentiviruses were
seeded into confocal dishes at a density of 1×105
cells/dish. Subsequently, cells in the Vector, OE-TUBB4B+AS-IV, and
KD2-TUBB4B+AS-IV groups were treated with AS-IV at a concentration
of 112.3 μM at 37°C for 48 h, fixed with 4% paraformaldehyde
at room temperature for 15 min, and permeabilized with 0.4% Triton
X-100 at room temperature for 10 min. The cells were stained with
phalloidin at room temperature in the dark for 20 min, placed in
blocking solution for 30 min, incubated with TUBB4B primary
antibody (1:100) at 4°C overnight, and then incubated with
fluorescent secondary antibody [goat anti-rabbit IgG (H+L),
conjugated with Alexa Fluor-488; 1:200] at room temperature for 1
h. The nuclei were stained with Hoechst 33342 (10 μg/ml) for
20 min at room temperature. Finally, images were captured using a
confocal microscope.
Molecular docking and dynamics
simulation
AutoDockTools software (v1.5.7; http://autodock.scripps.edu/) was used to preprocess
the conformations of AS-IV and TUBB4B and appropriate docking sites
were selected. AutoDock Vina software (v1.1.2; vina.scripps.edu/) was used to select the optimal
molecular docking conformation. PyMOL software (v2.3.0; pymol.org/) and LigPlot+ software (v2.2.8;
ebi.ac.uk/thornton-srv/software/LigPlus/) were used for analysis
and visualization.
Amber24 software (ambermd.org/)
was used for molecular dynamics simulation purposes, with the
ff19SB force field and Optimal Point Charge (OPC) water model
selected. The AS-IV-TUBB4B complex was placed in a cubic water box.
The cutoff distance for electrostatic and van der Waals
interactions was set to 1.0 nm, the time step was 2 femtosecond
(fs) and the particle-mesh Ewald method (22) used for long-range correction of
electrostatic interactions. Energy minimization was performed,
followed by 200 ps Microcanonical ensemble (NVE) equilibrium
dynamics and 100 ps number of particles, pressure and temperature)
equilibrium dynamics. The V-rescale method was used for temperature
coupling of the thermal bath system (23) and the Parrinello-Rahman method
was used for pressure control (24). Finally, 100 nsec molecular
dynamics sampling was performed. Indicators such as root mean
square fluctuation (RMSF), root mean square deviation (RMSD),
radius of gyration (Rg) and solvent-accessible surface area (SASA)
were calculated using Amber modules (AmberTools24; https://ambermd.org), Cpptraj (v5.1.0; github.com/Amber-MD/cpptraj) and Python scripts
(v3.10; https://www.python.org).
Cell thermal shift assay (CETSA)
GH3 cells in the logarithmic growth phase were
treated with 112.3 μM AS-IV at 37°C for 24 h in the shNC,
OE-TUBB4B and KD2-TUBB4B groups. Following trypsin digestion, the
cells were resuspended and aliquoted at 1×106 cells/well
into eight EP tubes. The tubes were heated for 10 min across a
temperature gradient (35-70°C). NP-40 lysis buffer was added and
the sample was centrifuged at 4°C, 20,000 × g for 20 min. The
protein supernatant was collected. The TUBB4B protein expression
was detected via western blot analysis as aforementioned. Melting
curves were plotted by analyzing changes in TUBB4B solubility or
abundance at different temperatures.
Statistical analysis
Statistical analysis was performed using GraphPad
Prism 10 (Dotmatics). All experiments were repeated at least three
times, and all data are expressed as the mean ± SD. One- or two-way
ANOVA followed by Dunnett's t test was used for comparisons.
P<0.05 was considered to indicate a statistically significant
difference.
Results
AS-IV inhibits proliferation and promotes
apoptosis of GH3/MMQ cells in a concentration- and time-dependent
manner
CCK-8 assay demonstrated that AS-IV significantly
inhibited the activity of GH3 and MMQ cells in a concentration- and
time-dependent manner (Fig.
1D-K). The chemical structure of AS-IV is shown in Fig. 1A. The IC50 values
detected after 48 h AS-IV treatment in GH3 and MMQ cells were 112.3
and 247.7 μM, respectively (Fig. 1B and C). To ensure consistency in
subsequent experiments, 48 h was selected as the treatment duration
and IC50 values were used as the drug concentration for
AS-IV intervention.
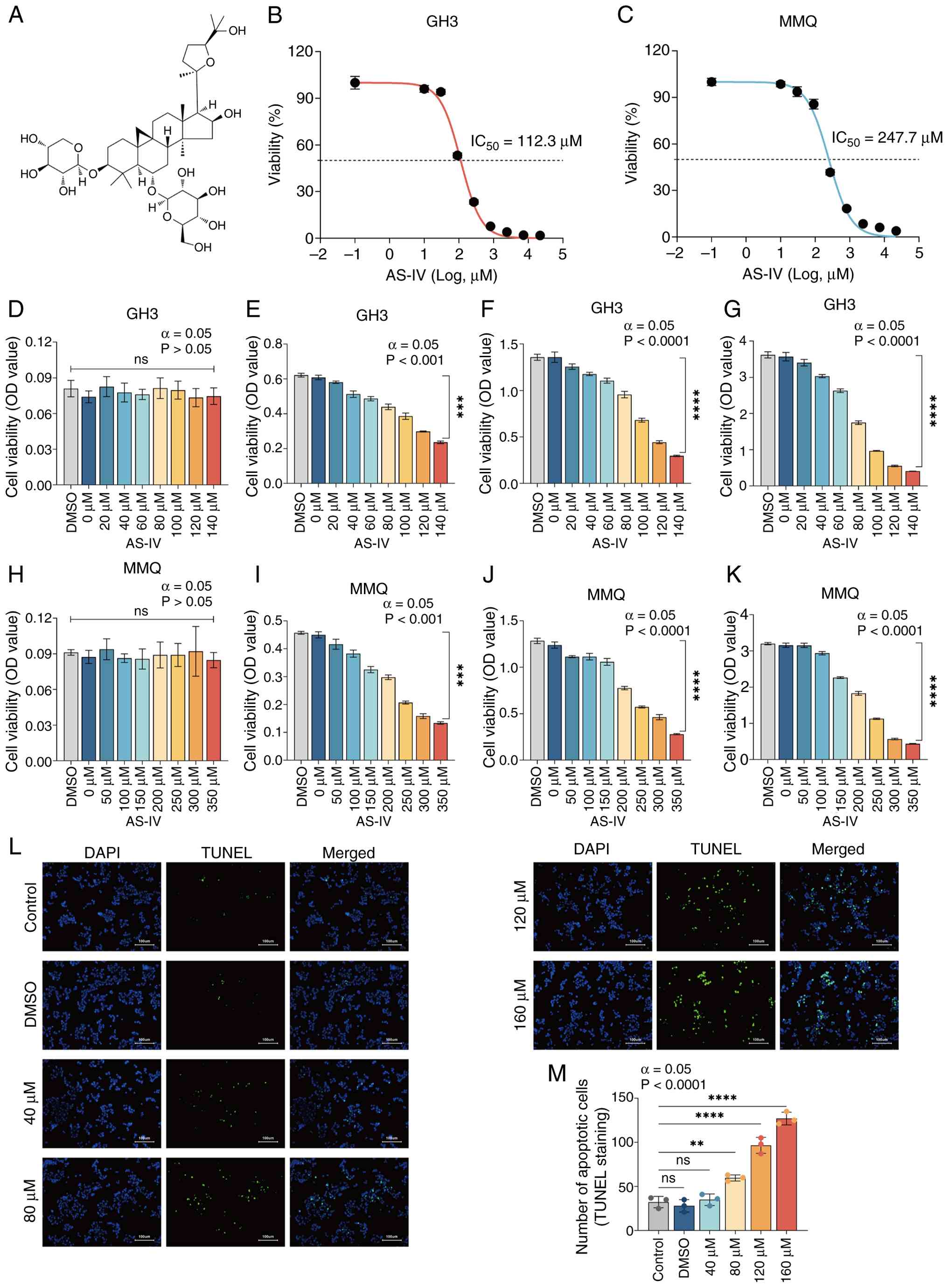 | Figure 1AS-IV inhibits the proliferation and
promotes the apoptosis of GH3/MMQ cells in a concentration- and
time-dependent manner. (A) Chemical structure of AS-IV.
IC50 of AS-IV in (B) GH3 and (C) MMQ cells following 48
h treatment. Viability of the GH3 cells treated with 0-140
μM AS-IV at (D) 0, (E) 24, (F) 48 and (G) 72 h. Viability of
MMQ-treated cells treated with 0-350 μM AS-IV at (H) 0, (I)
24, (J) 48 and (K) 72 h. (L) TUNEL assay was used to assess the
apoptotic effect of AS-IV on GH3. (M) Number of apoptotic cells in
each group. ****P<0.0001, ***P<0.001,
**P<0.01 vs. control. ns, not significant; AS-IV,
astragaloside IV; IC50, half-maximal inhibitory
concentration; OD, optical density. |
TUNEL assay was performed to confirm that AS-IV
promotes apoptosis in GH3 cells in a concentration-dependent manner
(Fig. 1L). The apoptosis rate of
the GH3 cells treated with 160 μM AS-IV was significantly
greater than that of the control cells (Fig. 1M). Western blot analysis of
apoptosis-associated proteins (Fig.
S1A) revealed that the expression of cleaved-PARP (Fig. S1C) significantly increased in
AS-IV-treated cells. The expression of the 113 kDa total PARP
isoform decreased but that of the 89 kDa form increased, however
there was no significant difference in total protein levels
(Fig. S1B and C). Western
blotting revealed that pro-caspase3 protein levels remained largely
unchanged in AS-IV-treated cells, whereas the relative expression
of cleaved-caspase3 protein (Fig.
S1D-F) was significantly elevated. Transmission electron
microscopy revealed increased numbers of vesicles, apoptotic
bodies, fragmented mitochondrial tubular networks and loss of
mitochondrial cristae fusion in AS-IV-treated cells (Fig. S1G). These findings confirmed
that AS-IV induces pituitary tumor cell apoptosis.
Transcriptome sequencing reveals
significant downregulation of TUBB4B in AS-IV-treated GH3 cells and
high expression of TUBB4B in pituitary tumors
To explore the key signals and related pathways
underlying the AS-IV-mediated inhibition of GH3 cell proliferation,
single-cell sequencing of AS-IV-treated GH3 cells was performed.
Compared with the NC group, 931 up- and 829 downregulated DEGs were
identified (Fig. 2A and B). GO
enrichment analysis in the biological process category (Fig. 2C) revealed that the functions of
the DEGs were enriched in 'DNA replication' and 'cell cycle'. A
heatmap of the transcriptomic DEGs (Fig. 2D) revealed that the tubulin
family (which synthesizes the cytoskeleton) (25), especially TUBB, was significantly
downregulated in the AS-IV group. GSEA revealed that genes were
significantly downregulated in 'mitotic cell cycle',
'cyclin-dependent protein kinase holoenzyme complex',
'double-stranded RNA binding' and 'structural constituent of
cytoskeleton' (Fig. 2E-H). As
these enriched gene sets are associated with tubulin-associated
functions, it was hypothesized that there was a potential link
between cytoskeletal organization and RNA binding, with TUBB4B
possibly serving as a bridging molecule. To explore this
hypothesis, we selected the gene lists of 'double-stranded RNA
binding' and 'structural constituent of cytoskeleton' to construct
a Venn diagram (Fig. 2I), which
revealed that TUBB4B was present in the intersection. It was
hypothesized that TUBB4B may be a key gene mediating AS-IV-induced
inhibition of pituitary tumor proliferation and promotion of
apoptosis. GEO data demonstrated that TUBB4B expression in
pituitary tumor tissues was significantly higher than that in
normal pituitary tissue (Fig.
2J), which was consistent with the IHC staining results of
pituitary tumor tissue from patients (Fig. 2K). These results confirm that
TUBB4B is highly expressed in pituitary tumors.
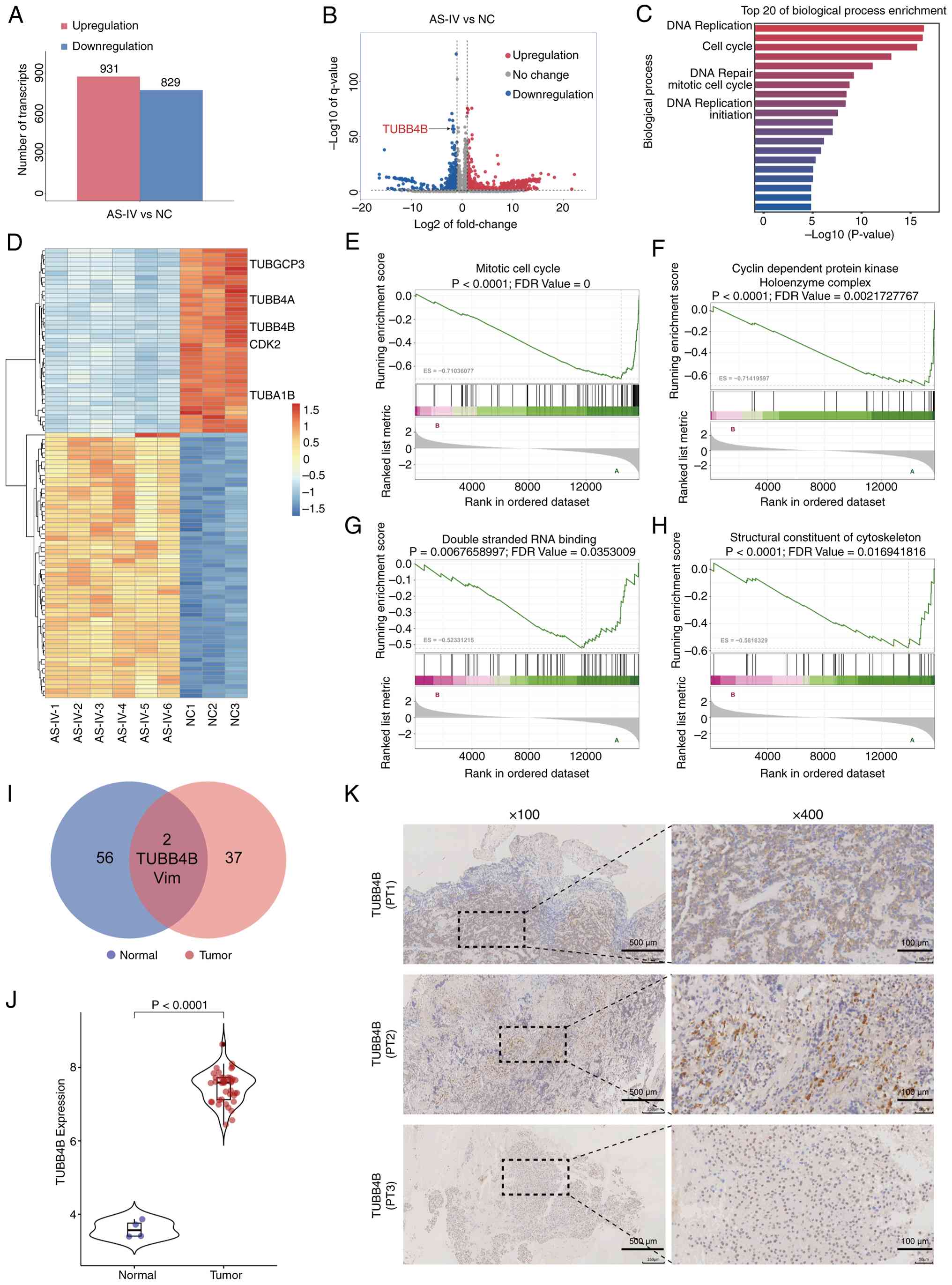 | Figure 2Transcriptomic sequencing reveals
that the TUBB4B gene is significantly downregulated in GH3 cells
treated with AS-IV and TUBB4B is highly expressed in pituitary
tumors. (A) Number of DEGs between the AS-IV and NC group. (B)
Volcano plot showing the distribution of DEGs. (C) Gene Ontology
enrichment analysis for the biological processes. (D) Heatmap of
the expression of significant DEGs. Gene set enrichment analysis
demonstrated that downregulated DEGs were enriched in (E) 'mitotic
cell cycle', (F) 'cyclin-dependent protein kinase holoenzyme
complex', (G) 'double-stranded RNA binding' and (H) 'structural
constituent of cytoskeleton' (ES<-0.5; P<0.05; FDR<0.25).
(I) Venn diagram of genes enriched in 'double-stranded RNA binding'
and 'structural constituent of cytoskeleton' showing that the
TUBB4B gene is at the intersection. (J) Expression of TUBB4B in
normal human pituitary glands and pituitary tumors; TUBB4B was
highly expressed in pituitary tumors (data from GSE136781,
P<0.0001). (K) Immunohistochemical staining of TUBB4B in tissue
samples from PTs with pituitary neuroendocrine tumors. TUBB4B,
tubulin β4B class IVb; AS-IV, Astragaloside IV; DEG, differentially
expressed genes; NC, negative control; ES, enrichment score; FDR,
false discovery rate; PT, pituitary tumor; Vim, vimentin. |
TUBB4B promotes proliferation and
inhibits apoptosis in pituitary tumor cells
Stably transfected cell lines with
lentivirus-mediated OE-TUBB4B or knockdown were established
(Fig. 3A). The transfection
efficiency was verified by RT-qPCR and western blotting (Fig. 3B, C and E-G). The results
confirmed the successful establishment of stably transfected cell
lines. In both cell lines, the knockdown efficiency of KD2-TUBB4B
was greater than that of KD1-TUBB4B; therefore, the KD2-TUBB4B
stable cell line was selected for subsequent experiments.
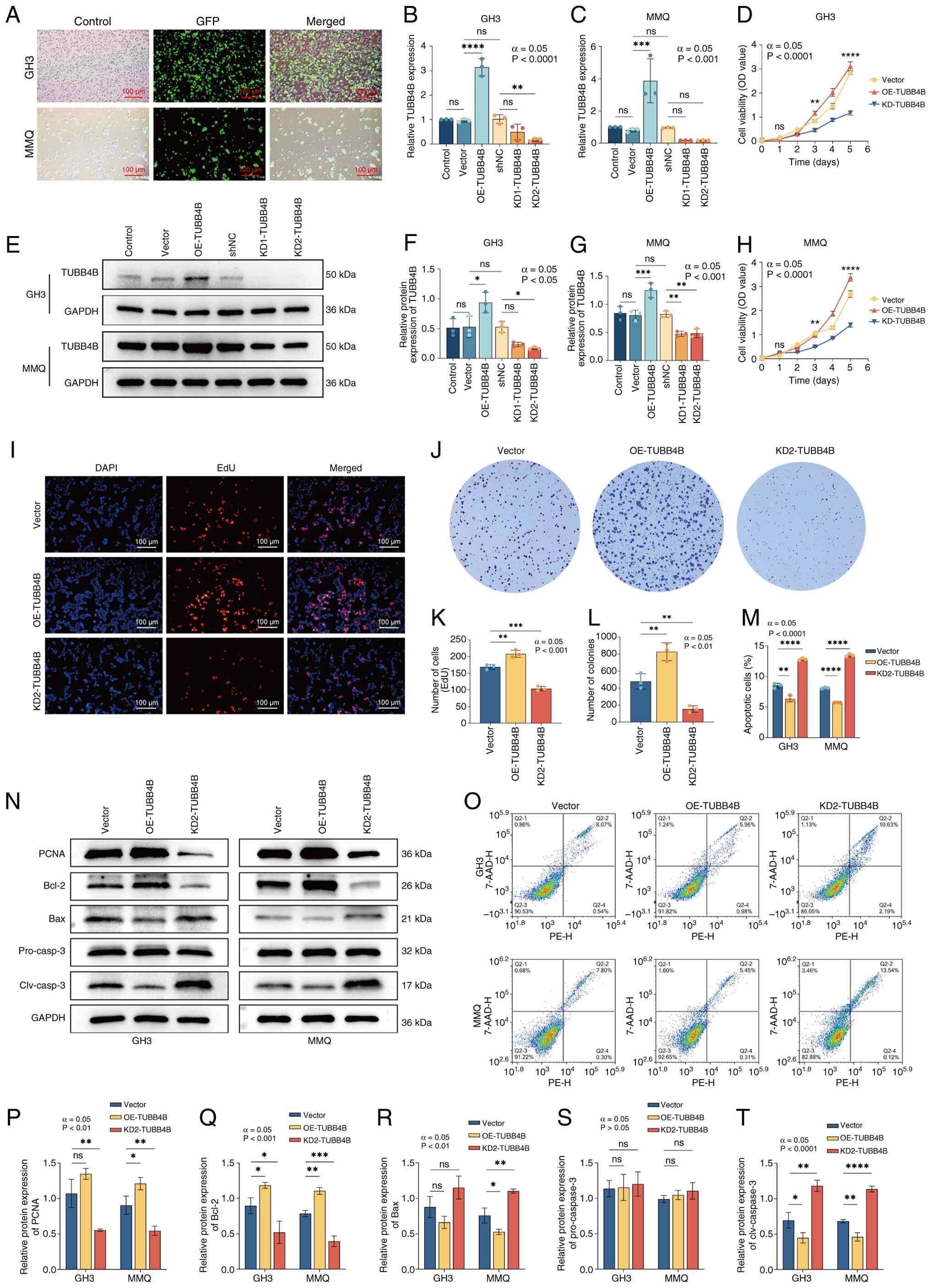 | Figure 3TUBB4B promotes the proliferation and
inhibits the apoptosis of pituitary tumor cells. (A) Fluorescence
images of the GH3/MMQ cells transfected with lentivirus. Reverse
transcription-quantitative PCR was used to assess expression
efficiency of TUBB4B in lentivirus-transfected (B) GH3 and (C) MMQ
cell lines. (D) Cell Counting Kit-8 assay was used to assess the
viability of GH3 cells. (E) Western blotting was used to assess
expression efficiency of TUBB4B in lentivirus-transfected (F) GH3
and (G) MMQ cell lines. (H) Cell Counting Kit-8 assay was used to
assess the viability of MMQ cells. (I) EdU staining was used to
assess the effect of TUBB4B on the proliferation of GH3 cells. (J)
Colony formation of GH3 cells. (K) EdU-positive cells. (L) Number
of colonies. (M) Number of apoptotic GH3/MMQ cells. (N) Western
blot bands (O) Flow cytometry. Western blotting was used to assess
protein expression of (P) PCNA, (Q) Bcl-2, (R) Bax, (S)
pro-caspase-3 and (T) clv caspase-3 in GH3 and MMQ cell lines.
****P<0.0001, ***P<0.001,
**P<0.01, *P<0.05 vs. vector. TUBB4B,
tubulin β4B class IVb; OE, overexpression; shNC, short hairpin RNA
negative control; KD, knockdown; OD, optical density; clv, cleaved;
ns, not significant. |
Cell viability was assessed with the CCK-8 assay. In
both cell lines, viability in the OE-TUBB4B group was greater than
that in the shNC group, which was significantly greater than that
in the KD2-TUBB4B group (Fig. 3D and
H). EdU assays revealed that the number of proliferating GH3
cells (in the S phase of DNA synthesis; red fluorescence) was
highest in the OE-TUBB4B group and lowest in the KD2-TUBB4B group
(Fig. 3I and K). Colony
formation assay (Fig. 3J and L)
revealed that the number of colonies in the OE-TUBB4B group was
significantly greater than that in the shNC group, which was
greater than that in the KD2-TUBB4B group. Flow cytometry confirmed
that OE-TUBB4B decreased the proportion of apoptotic cells, whereas
KD-TUBB4B increased the percentage of apoptotic cells in both cell
lines (Fig. 3M and O). Western
blotting was used to verify the expression of
proliferation-associated protein (PCNA) and apoptosis-related
proteins (Bcl-2, Bax, pro-caspase-3 and cleaved-caspase-3) in both
cell lines (Fig. 3N and P-T).
OE-TUBB4B cells had increased expression of PCNA and antiapoptotic
protein (Bcl-2) and decreased expression of proapoptotic proteins
(Bax and cleaved-caspase-3); in KD2-TUBB4B cells, decreased
expression of PCNA and Bcl-2 was observed, along with increased
expression of Bax and cleaved-caspase-3. Cells in the OE-TUBB4B
group exhibited increased expression of PCNA and Bcl-2 and
decreased expression of Bax and cleaved caspase-3; in contrast,
cells in the KD2-TUBB4B group exhibited decreased expression of
PCNA and Bcl-2, and increased expression of Bax and cleaved
caspase-3, confirming the regulatory role of TUBB4B. In summary,
TUBB4B promotes the proliferation and inhibits the apoptosis of
pituitary tumor cells.
AS-IV exerts an inhibitory effect on
pituitary tumors by targeting TUBB4B
To verify that AS-IV inhibits pituitary tumors by
targeting TUBB4B, molecular conformations of TUBB4B and AS-IV were
generated. The surface electrostatic potential of the TUBB4B
protein is shown in Fig. 4A, and
molecular docking was performed between AS-IV and TUBB4B (Fig. 4B). The binding energies of
multiple TUBB4B protein conformations with AS-IV indicated a high
degree of fit in the binding pocket and a binding energy of −8.2
kcal/mol (Fig. 4C). These
findings confirmed that AS-IV and TUBB4B had strong binding
activity and bind under natural conditions. On the basis of the
molecular docking results, a 100 nsec molecular dynamics simulation
was performed on the AS-IV-TUBB4B complex to investigate its
binding stability and dynamic behavior. RMSF (Fig. 4D) of the AS-IV-TUBB4B complex
fluctuated at residues 56, 80, 175, 277-282 and 427, which may be
associated with the function of TUBB4B affected by AS-IV; the RMSD
(Fig. 4E) fluctuated in the
early stage (0-20 nsec) but stabilized at 20-100 nsec, indicating
that the structure of the complex was relatively stable; the Rg
value (Fig. 4F) remained stable
at ~2.1 nm, indicating that the structure of the complex was
compact and the SASA (Fig. 4G)
of the entire system was relatively stable. Free energy landscape
of the complex demonstrated that the conformation of the
AS-IV-TUBB4B complex was stable when the Rg values ranged from 2.07
to 2.09 nm and the RMSD was 0.03-0.17 nm (Fig. 4H).
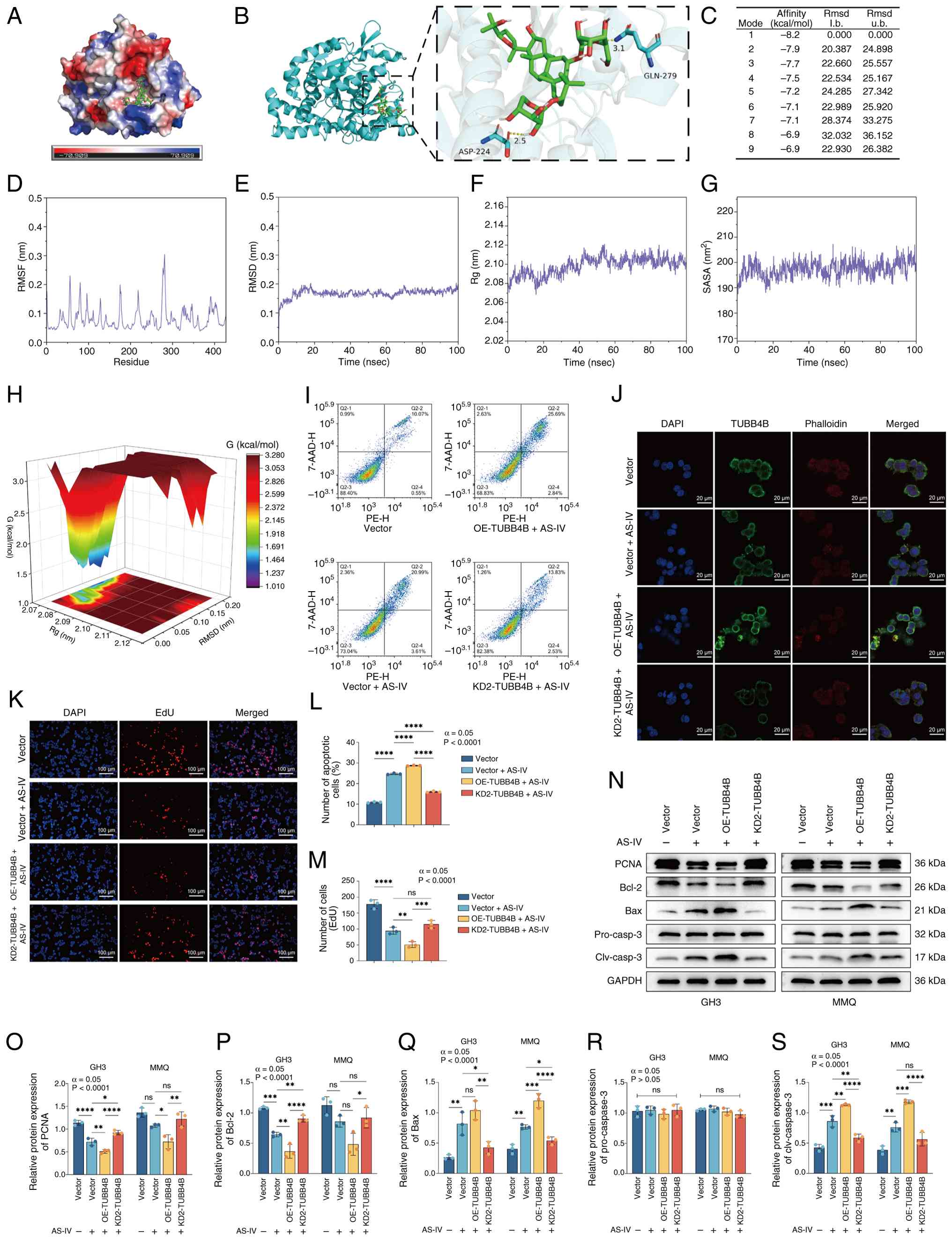 | Figure 4AS-IV exerts an inhibitory effect on
pituitary tumors by targeting and binding TUBB4B. (A) Surface
electrostatic potential of the TUBB4B protein. (B) Molecular
docking between AS-IV (green) and TUBB4B (blue). Yellow dashed
lines represent hydrogen bonds. (C) Molecular docking binding
energies of AS-IV and TUBB4B in different spatial conformations.
(D) RMSF, (E) RMSD, (F) Rg and (G) SASA between TUBB4B protein
residues and the AS-IV solution in the experimental system. (H)
Free energy landscape showing that the conformation of the
AS-IV-TUBB4B complex is dominant and more stable when the Rg was
2.07-2.09 nm and the RMSD was 0.03-0.17 nm. (I) Flow cytometry was
used to assess apoptosis of GH3 cells. (J) Immunofluorescence of
GH3 cells following AS-IV treatment. (K) EdU staining was used to
assess proliferation of GH3 cells. (L) Proportion of apoptotic GH3
cells. (M) Number of EdU-positive cells. (N) Western blotting was
used to assess the protein expression of (O) PCNA, (P) Bcl-2, (Q)
Bax, (R) pro-caspase-3 and (S) cleaved-caspase-3 in the GH3/MMQ
cell lines. ****P<0.0001, ***P<0.001,
**P<0.01, *P<0.05. AS-IV, Astragaloside
IV; TUBB4B, tubulin beta 4B class IVb; RMSF, root mean square
fluctuation; RMSD, Root Mean Square Deviation; Rg, radius of
Gyration; SASA, Solvent-Accessible Surface Area; OE,
overexpression; KD, knockdown; l. b., lower bound; u. b., upper
bound; clv, cleaved; ns, not significant. |
The present study demonstrated the targeted
interaction between AS-IV and TUBB4B by verifying whether TUBB4B
regulation directly influences the pharmacological efficacy of
AS-IV. Immunofluorescence demonstrated the shNC + AS-IV group
exhibited partial cell shrinkage, nuclear condensation and blurred
or fragmented cytoskeletal structures (Fig. 4J). In the OE-TUBB4B + AS-IV
group, some nuclei were markedly lysed and the cytoskeletal
components were fragmented and degraded in the cytoplasm,
indicating that OE-TUBB4B enhances the inhibitory effect of AS-IV
on pituitary tumors. Conversely, in the KD2-TUBB4B + AS-IV group,
although nuclei shrank, the cytoskeletal structure remained intact,
suggesting that KD-TUBB4B attenuated the inhibitory effect of AS-IV
on pituitary tumors. The present study validated the targeted
interaction between AS-IV and the TUBB4B protein via CETSA. TUBB4B
protein gradually denatured with increasing temperature (Fig. S2A). In AS-IV-treated cells, the
thermal denaturation temperature of the TUBB4B protein increased.
Compared with that in the shNC group (Fig. S2B), the melting curve in the
OE-TUBB4B group shifted to the right (Fig. S2C), whereas the KD2-TUBB4B group
showed the opposite trend (Fig. S2D
and E), indicating direct binding of AS-IV to TUBB4B. The
present study determined the effect of TUBB4B on AS-IV drug
sensitivity via viability assays. In the shNC group, the
IC50 was 112.8 μM, reflecting baseline
sensitivity, whereas in the OE-TUBB4B group, the IC50
was 60.4 μM and that in the KD2-TUBB4B group was 292.1
μM (Fig. S2F). The cells
overexpressing TUBB4B exhibited increased sensitivity to AS-IV,
whereas KD-TUBB4B expression decreased AS-IV sensitivity. Flow
cytometry (Fig. 4I and L), EdU
assay (Fig. 4K and M) and
western blot analysis were performed for both cell lines (Fig. 4N-S). The cells in the OE-TUBB4B +
AS-IV group proliferated more slowly and exhibited a higher
apoptosis rate than those in the Vector + AS-IV group; in contrast,
cells in the KD2-TUBB4B+AS-IV group proliferated more rapidly and
exhibited a lower apoptosis rate compared to the Vector + AS-IV
group. These results collectively confirmed that OE-TUBB4B enhanced
AS-IV efficacy, whereas KD-TUBB4B attenuated it, indicating that
TUBB4B modulates cellular sensitivity to AS-IV. These findings
collectively demonstrated that AS-IV targeted TUBB4B to exert its
inhibitory effect on pituitary tumors.
OE-TUBB4B upregulates STMN1 and activates
the ERK pathway
To clarify the mechanism by which TUBB4B affects the
proliferation and apoptosis of pituitary tumors, the cell cycle of
GH3 cells was analyzed. OE-TUBB4B accelerated the G1-to-S phase
transition (Fig. 5A and B).
RT-qPCR was used to assess cell cycle-related protein expression
(Fig. 5C); expression levels of
CCND1, CDK4 and CDK6 were significantly increased in the OE-TUBB4B
group, consistent with the KEGG pathway enrichment (Fig. 5D). PPI analysis (Fig. 5E) revealed that the TUBB4B
protein interacted with STMN1, which was consistent with the data
from the STRING database (Fig.
5F). Therefore, the present study focused on the regulation of
the ERK/MAPK pathway by the STMN1 protein.
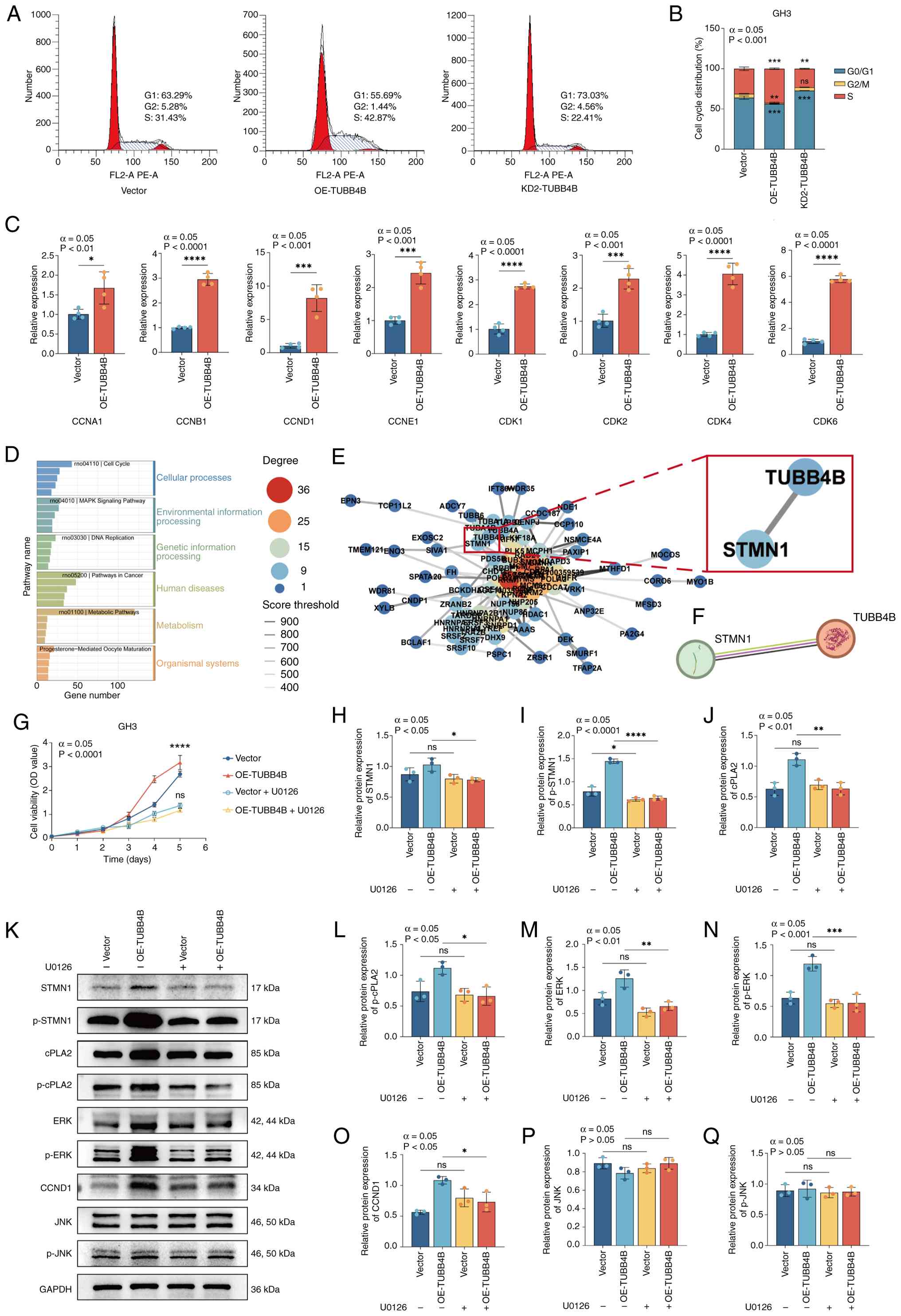 | Figure 5OE-TUBB4B upregulates STMN1 and
activates the ERK pathway. (A) Effect of TUBB4B on GH3 cell cycle
progression. (B) Cell cycle distribution. (C) Reverse
transcription-quantitative PCR was used to assess the effects of
OE-TUBB4B. (D) Transcriptomic sequencing showing Kyoto Encyclopedia
of Genes and Genomes pathway enrichment of differentially expressed
genes. (E) Transcriptomic sequencing showing the protein-protein
interaction network of proteins that interact with TUBB4B (red box
indicates key pathway proteins that interact with TUBB4B). (F)
Interaction diagram between the TUBB4B and STMN1 proteins from
STRING. (G) Cell Counting Kit-8 assay was used to assess the
viability of GH3 cells treated with U0126. In GH3 cell lines
treated with U0126, the protein expression levels of (H) STMN1, (I)
p-STMN1 (I), and cPLA2 (J) were assessed by western blotting (K),
and levels of p-cPLA2 (L), ERK (M), p-ERK (N), CCND1 (O), JNK (P),
and p-JNK (Q) were also assessed. ****P<0.0001,
***P<0.001, **P<0.01,
*P<0.05 vs. vector. OE, overexpression; TUBB4B,
tubulin beta 4B class IVb; STMN1, stathmin 1; p-, phosphorylated;
cPLA2, cytosolic phospholipase A2; CCND1, cyclin D1; KD, knockdown;
OD, optical density; ns, not significant. |
To verify whether changes in STMN1 expression
directly relied on TUBB4B regulation, immunofluorescence
colocalization of the TUBB4B and STMN1 protein was performed
(Fig. S3A). Upon OE of TUBB4B,
the green (TUBB4B) and the red fluorescence signal (STMN1)
increased. Following KD of TUBB4B expression, the fluorescence
signals of both TUBB4B and STMN1 simultaneously decreased, with a
corresponding decrease in colocalization. These results indicate
that STMN1 expression was associated with TUBB4B levels and that
STMN1 colocalized within the microtubule network. Western blot
analysis demonstrated OE-TUBB4B increased expression of the STMN1,
p-STMN1, ERK and p-ERK proteins (Fig. S3B), whereas KD-TUBB4B decreased
expression (Fig. S3C-F). These
findings indicate that STMN1 served as an effector molecule in the
TUBB4B downstream pathway and was associated with ERK pathway
activation.
CCK-8 assay revealed no significant difference in
cell viability between the OE-TUBB4B and the shNC group following
U0126 treatment (Fig. 5G).
Western blot analysis (Fig.
5H-Q) indicated that OE-TUBB4B led to increased expression of
p-STMN1, p-cPLA2, p-ERK and CCND1. Additionally, the expression of
total STMN1, cPLA2 and ERK proteins increased, and no significant
differences in the levels of the JNK and p-JNK protein were
detected. Following U0126 treatment, the STMN1, p-STMN1, cPLA2,
p-cPLA2, ERK, p-ERK, CCND1, JNK, and p-JNK protein levels in the
Vector + AS-IV group or the OE-TUBB4B+AS-IV group did not
significantly change. These findings indicated that the ERK/MAPK
pathway is involved in proliferation and that blocking this pathway
inhibited positive feedback. These findings also suggest that
OE-TUBB4B may promote pituitary tumor proliferation through the
activation of positive feedback among downstream proteins.
AS-IV inhibits proliferation and promotes
the apoptosis of xenograft tumors in vivo by targeting TUBB4B
To verify the reliability of the in vitro
cell experiments in vivo, a nude mouse subcutaneous
xenograft model was established using TUBB4B-regulated GH3 cell
lines (Fig. 6A). There was no
significant difference in body weight between any groups (Fig. 6B), which was attributed to the
benign nature of the GH3 cell line. Compared with the Vector group,
both tumor volume and weight were significantly increased in the
OE-TUBB4B group, whereas they were decreased in the KD2-TUBB4B
group (Fig. 6C and C). Compared
with the Vector + AS-IV group, both tumor volume and weight were
decreased in the OE-TUBB4B + AS-IV group, whereas they were
increased in the KD2-TUBB4B + AS-IV group. Overall, the trends in
tumor volume and weight were consistent across the groups. The
largest tumor was observed in the OE-TUBB4B group, with a maximum
tumor diameter of 12.7 mm, a volume of 892 mm3 and a
mass of 1,671 mg. Compared with the shNC group, the OE-TUBB4B group
presented notable blood sinuses and the tumor cells exhibited the
highest density, large in size, and have a high
nucleus-to-cytoplasm ratio, consistent with a state of high
proliferation; the KD2-TUBB4B and shNC + AS-IV group presented
partial nuclear fragmentation and cytoskeletal
fragmentation/degradation in tumor tissue; the OE-TUBB4B + AS-IV
group presented notable nuclear pyknosis/fragmentation and severe
apoptosis and necrosis of tumor tissue and the KD2-TUBB4B + AS-IV
group presented partial nuclear shrinkage but relatively regular
tissue morphology (Fig. 6E).
Western blotting (Figs. 6F, H-J
and S4A-C) and IHC staining
(Fig. 6G and K-O) results for
the xenograft tumor tissue were consistent with those of the in
vitro experiments; compared with the Vector group, the
OE-TUBB4B group exhibited increased expression of PCNA and Bcl-2
proteins and decreased expression of Bax and cleaved-caspase-3
proteins; while the KD2-TUBB4B group exhibited the opposite trends
in these protein expressions compared to the Vector group;
Following AS-IV intervention, the OE-TUBB4B+AS-IV group exhibited
lower PCNA and Bcl-2 protein expression compared to the Vector +
AS-IV group, while Bax and cleaved-caspase-3 protein expression was
higher; conversely, the KD2-TUBB4B+AS-IV group showed the opposite
trends in these protein expressions compared with the Vector +
AS-IV group. These results were consistent with the aforementioned
findings that OE-TUBB4B increased the effectiveness of AS-IV, while
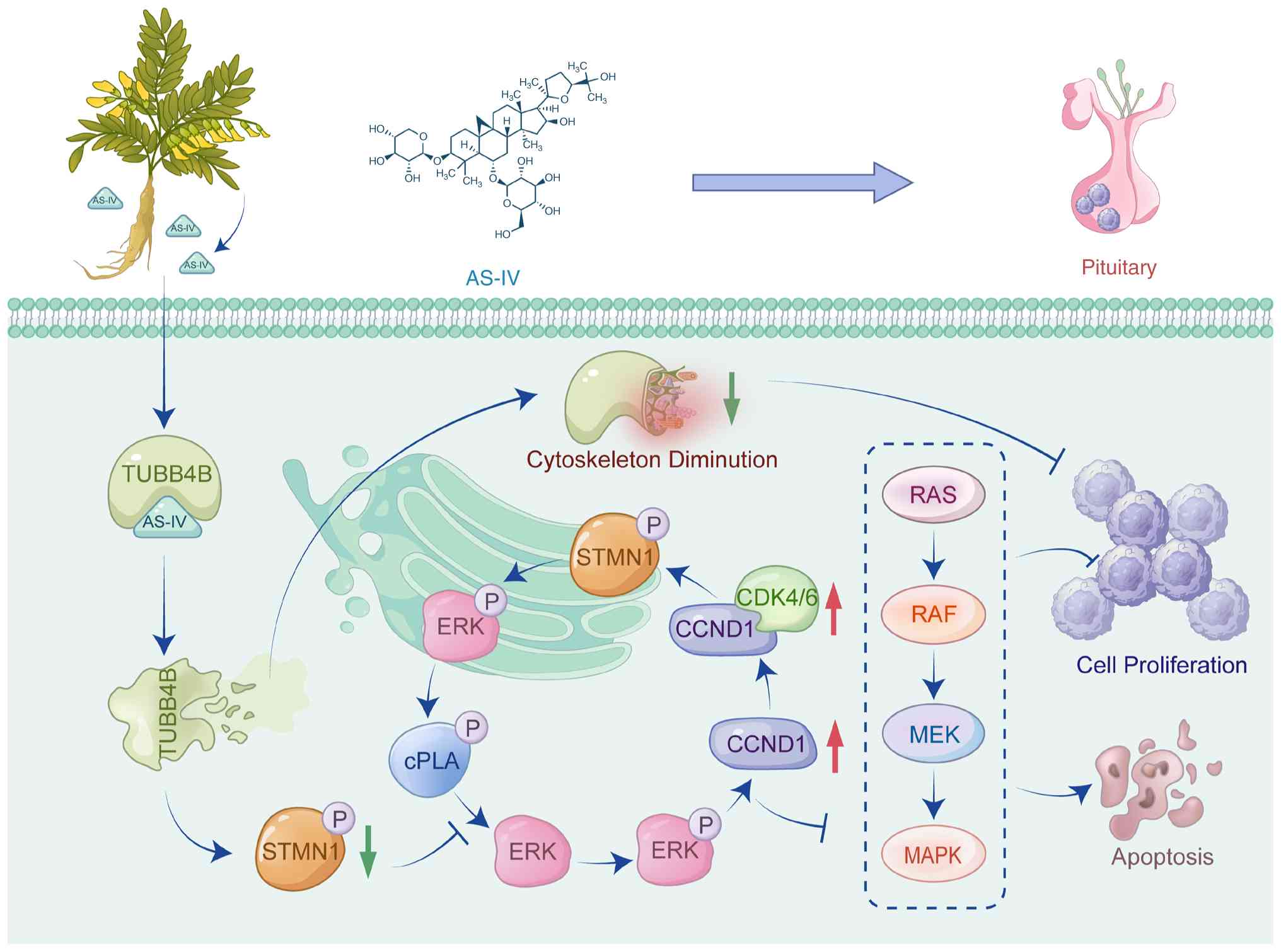
KD-TUBB4B attenuated effectiveness. Fig. 7 shows the schematic of the
overall mechanism by which AS-IV regulates proliferation and
apoptosis in pituitary tumors by targeting of TUBB4B. In summary,
AS-IV inhibited proliferation and promoted apoptosis of xenograft
tumors in animals by targeting of TUBB4B.
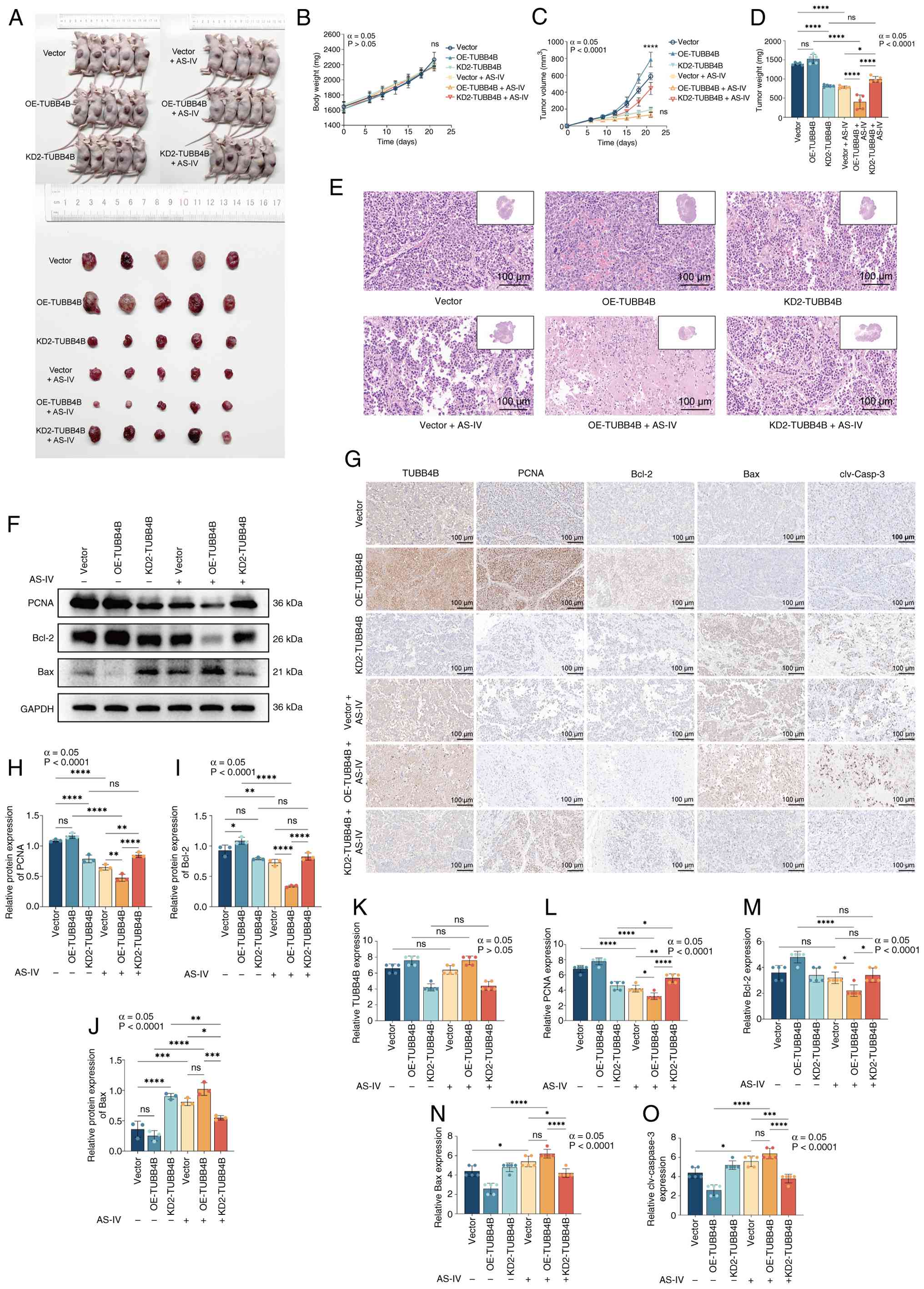 | Figure 6AS-IV inhibits proliferation and
promotes apoptosis of xenograft tumors in animals by targeting of
TUBB4B. (A) Xenograft tumors in nude mice. (B) Body weight and (C)
tumor volume growth curve of the nude mice. (D) Tumor weight. (E)
Representative hematoxylin and eosin staining images of tumors
(inset: 10X magnification). (F) Western blotting was used to assess
protein expression. (G) Representative IHC staining images of
TUBB4B, PCNA, Bcl-2, Bax and cleaved-caspase-3 proteins. Expression
levels of (H) PCNA, (I) Bcl-2 and (J) Bax. IHC staining scores for
(K) TUBB4B, (L) PCNA, (M) Bcl-2, (N) Bax and (O) clv-caspase-3
(Allred scoring system was used). ****P<0.0001,
***P<0.001, **P<0.01,
*P<0.05. AS-IV, astragaloside IV; TUBB4B, tubulin β4B
class IVb; IHC, immunohistochemistry; OE, overexpression; KD,
knockdown; ns, not significant; clv, cleaved. |
Discussion
The present study demonstrated that AS-IV
significantly inhibited pituitary tumor cell proliferation and
induced apoptosis by targeting TUBB4B and regulating the STMN1/ERK
signaling pathway. AS-IV inhibited pituitary tumor cell viability
in a concentration- and time-dependent manner, with its molecular
mechanism relying on high-affinity binding to TUBB4B and regulation
of the downstream STMN1/ERK pathway. To the best of our knowledge,
the present study is the first to reveal that TUBB4B OE enhances
AS-IV efficacy and TUBB4B regulation affects pituitary tumor cell
sensitivity to AS-IV. These findings not only demonstrate the
anti-pituitary tumor mechanism of AS-IV but also provide a novel
strategy for targeted drug therapy of pituitary tumors and a basis
for exploring multitarget regulation by active components in
traditional Chinese medicine.
AS-IV, as the primary active component of the
traditional Chinese medicine A. membranaceus, has advantages
for tumor therapy because of its multitarget and low-toxicity
characteristics (26). Its
ability to inhibit tumor proliferation and invasion while promoting
tumor cell apoptosis has been demonstrated both in vitro and
in vivo (27,28). The present study revealed that
AS-IV demonstrates enhanced antitumor effects in
TUBB4B-overexpressing cells. Molecular docking and kinetic
simulations demonstrated that AS-IV bound to TUBB4B with high
affinity (binding energy, −8.2 kcal/mol) and formed a stable
complex. CETSA confirmed that AS-IV directly bound to TUBB4B and
enhanced its thermal stability. Thus, TUBB4B OE may increase the
number of intracellular targets available for AS-IV binding, thus
increasing the total number of drug-target interactions and overall
efficacy. This aligns with the characteristics of AS-IV as a
multitarget traditional Chinese medicine component, which exerts
stronger effects in target-enriched environments through
high-affinity binding (29,30). Second, TUBB4B regulation may be
mediated by altering the drug sensitivity of AS-IV.
TUBB4B-overexpressing cells exhibited heightened sensitivity to
AS-IV (lower IC50), whereas TUBB4B knockdown increased
resistance (higher IC50). These findings suggest that
TUBB4B expression may serve as a biomarker for AS-IV efficacy, with
highly expressing populations potentially benefitting more from
AS-IV treatment. Finally, the binding of AS-IV to TUBB4B may
disrupt normal interactions between TUBB4B and tubulin or its
downstream effector protein STMN1, thus interfering with
microtubule dynamics and cell cycle progression. As a key protein
in cytoskeletal architecture, TUBB4B directly influences
microtubule dynamics and mitotic processes and is associated with
cytoskeletal organization and intercellular gap junctions (31). In vivo experiments
confirmed that AS-IV did not significantly alter TUBB4B protein
expression levels, but its proliferation-promoting and
apoptosis-inhibiting effects were markedly blocked. These findings
suggested that following TUBB4B OE, increased formation of
dysfunctional TUBB4B-AS-IV complexes may exacerbate microtubule
dysfunction, leading to enhanced proliferation inhibition and
apoptosis induction. In summary, the phenomenon of TUBB4B OE
enhancing AS-IV efficacy does not contradict conventional targeted
therapy logic but reflects the unique mode of action of AS-IV
through binding and interfering with target proteins.
The present study used multiomics analysis and
experimental validation to demonstrate that TUBB4B activated the
ERK pathway by upregulating STMN1 expression, demonstrating its
pivotal role in the G1/S transition of the cell cycle. The MAPK
pathway comprises four primary branches: ERK, JNK, p38/MAPK and
ERK5 (32). Among these
pathways, the ERK/MAPK signaling pathway serves as the key network
that regulates cell proliferation, development and division
(33). STMN1, a key microtubule
depolymerizing protein, is upregulated in multiple types of tumor
and is associated with poor prognosis; its upregulation accelerates
the G1/S transition and promotes proliferation (34). For example, in gallbladder
carcinoma, elevated STMN1 expression inhibits tumor growth, induces
apoptosis and impairs mitosis (35). In pancreatic cancer, decreased
STMN1 expression decreases cell proliferation and invasiveness,
while high expression promotes distant metastasis and poor
differentiation (36). In
non-small cell lung cancer, STMN1 upregulation accelerates cell
proliferation, regulates microtubule stability to increase
migration and invasion and is associated with poor prognosis
(37). Inhibiting STMN1
expression and phosphorylation decreases cell proliferation,
migration and colony formation capacity (38). In the present study, PPI analysis
and immunofluorescence colocalization revealed a physical
interaction between TUBB4B and STMN1, with both proteins
colocalizing within the microtubule network. Western blotting
demonstrated TUBB4B expression significantly affected STMN1 and
p-STMN1 expression, accompanied by changes in ERK phosphorylation
levels. These findings suggested that TUBB4B may influence STMN1
expression and activity through protein interactions or stability
regulation. Following blockade of the ERK pathway with U0126, the
proliferation induced by TUBB4B OE was completely reversed, and
p-STMN1 and p-ERK levels were not increased, indicating that ERK
activation is essential for TUBB4B-mediated proliferation. However,
the present study did not modulate the STMN1 gene to validate
downstream pathways, nor has it used STMN1-specific inhibitors to
rule out the involvement of other pathways. Nevertheless, the
results of the KEGG pathway enrichment analysis and cell cycle
experiments support the existence of the TUBB4B-STMN1-ERK-cell
cycle regulatory axis. Mechanistically, TUBB4B may interact with
STMN1 to enhance its phosphorylation and activate the ERK pathway.
Activation of ERK increases the total protein expression of cPLA2
and its phosphorylation, which positively feeds back to activate
the ERK/MAPK pathway, thus promoting cell proliferation. The
effects of STMN1, cPLA2, and ERK pathway proteins have been shown
to occur through phosphorylation-mediated activation (39-41). However, the present study found
that following regulation by OE-TUBB4B, not only did their
phosphorylation levels increase, but their total protein expression
also rose. Therefore, it was hypothesized this pathway is subject
to feedback' regulation. Nevertheless, the specific mechanisms
underlying this regulation have not yet been thoroughly
investigated in this study.
Compared with established pituitary tumor drivers
such as pituitary-specific transcriptional factor 1, GATA binding
protein 2 and ubiquitin specific peptidase 8 (USP8) mutations,
TUBB4B is not widely recognized as a key target in pituitary tumors
(42-44). However, the present study
demonstrated that TUBB4B is highly expressed in pituitary tumors
and its regulation significantly affects tumor proliferation and
apoptosis, suggesting its potential value as a therapeutic target.
Notably, as TUBB4B is a key cytoskeletal protein, TUBB4B inhibitors
may exert broad-spectrum antitumor effects by disrupting processes
such as mitosis and cell migration. Unlike existing targeted
therapies for pituitary tumors, such as dopamine agonists (45) and somatostatin analogs (46), this mechanism may offer novel
therapeutic avenues for drug-resistant or refractory pituitary
tumors.
Notably, AS-IV has multiple targeted and
multi-pathway effects, which, while advantageous, also introduce
complexities and limitations. First, the efficacy of AS-IV varies
across different tumor types. Kong et al (47) reported an IC50 of 100
ng/ml for AS-IV on colorectal cancer cells (47), whereas Qiu et al (48) reported a value of 210 μM
for uterine leiomyoma cells. Similarly, in the present study, the
concentration of AS-IV was also relatively high. This may result
from differences in proliferation rate and cell cycle dependency
between benign and malignant tumors, variations in the integrity of
apoptosis or cell death mechanisms and disparities in the tumor
microenvironment and heterogeneity. Second, in addition to
regulating the TUBB4B-STMN1-ERK axis, AS-IV may exert antitumor
effects through pathways such as the PI3K/Akt (49), NF-κB (50) and TGF-β (51) pathways and induce immune
dysfunction (52) and allergic
reactions (53) via off-target
effects. These pathways and effects exhibit crosstalk with ERK,
collectively forming the network pharmacology basis of AS-IV.
However, the present study, we did not use drugs with different
chemical structures, nor did we conduct additional experiments
targeting the same receptor; therefore, we were unable to verify
whether off-target effects were present. Future research should
integrate proteomics and phosphoproteomics to elucidate the
multitarget regulatory landscape of AS-IV in pituitary tumors.
Finally, the present study primarily utilized the rat pituitary
tumor cell lines GH3 and MMQ, which effectively mimic tumor cell
behavior but differ from human pituitary tumor cells in terms of
genetic background, molecular phenotype and tumor microenvironment.
Future studies should validate the reliability of the
TUBB4B-STMN1-ERK axis using human cell lines or primary cell
cultures. Additionally, while the present study preliminarily
validated the upregulation of TUBB4B in pituitary tumors using
clinical samples, the sample size was small and lacked autopsy
pituitary specimens for comparative studies. Future studies should
incorporate large-scale independent cohort validation and
technologies such as tissue microarray to assess the feasibility of
TUBB4B as a prognostic biomarker or therapeutic target for
pituitary tumors. In summary, targeting TUBB4B by AS-IV may
represent a novel adjunctive therapeutic strategy rather than an
isolated intervention.
To the best of our knowledge, the present study is
the first to demonstrate that AS-IV inhibited pituitary tumor cell
proliferation and promoted apoptosis by targeting TUBB4B to
suppress the STMN1/ERK signaling pathway. Mechanistically, AS-IV
bound TUBB4B with high affinity, enhancing drug sensitivity by
blocking downstream STMN1/ERK signaling activation and arresting
the cell cycle at G1/S transition phase. TUBB4B OE enhanced AS-IV
efficacy, suggesting its potential as a biomarker for AS-IV
therapy. The present study not only revealed a novel mechanism
underlying the anti-pituitary tumor activity of AS-IV but also
provided theoretical support for therapeutic strategies targeting
the TUBB4B-STMN1-ERK axis in pituitary tumors.
Supplementary Data
Availability of data and materials
The data generated in the present study can be
accessed at China National GeneBank DataBase under accession number
ngb_07871 or at the following URL: db.cngb.org/data_resources/?query=CNP0008409.
Authors' contributions
JL conceived and designed the study, analyzed data
and wrote the manuscript. YQ performed molecular docking and
visualization. WZ conceived and designed the study. ZY provided
experimental guidance and technical support. YZ provided technical
support. JX analyzed and interpreted data. KX analyzed data. QL
conceived and designed the study and revised the manuscript. JL and
QL confirmed the authenticity of all the raw data. All authors have
read and approved the final manuscript.
Ethics approval and consent to
participate
The present study used anonymized human pituitary
tumor tissue and patient data from the First Affiliated Hospital of
Shihezi University, with written informed consent obtained from all
participants. The human research protocol was approved by the
Scientific and Technological Ethics Committee of the First
Affiliated Hospital of Shihezi University, Shihezi, China (approval
No.: KJ2024-476-01), while the animal experiments were approved by
the Bioethics Committee of Shihezi University, Shihezi, China
(approval no. A2024-353).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
The present study was supported by grants from 'Tianshan
Talents' Young and Middle-aged Medical and Health Backbone Talents
and the 'Science and Technology Program of the Corps' (grant no.
2022CB020).
References
|
1
|
Dai C, Kang J, Liu X, Yao Y, Wang H and
Wang R: How to classify and define pituitary tumors: Recent
advances and current controversies. Front Endocrinol (Lausanne).
12:6046442021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kong F, Cheng W and Zhan Q: Clinical study
on the selection of endoscopes and microscopes for transsphenoidal
surgery of non-aggressive pituitary macroadenoma and microadenoma
and the influencing factors of hyposmia after endoscopic
transsphenoidal surgery. Front Neurol. 15:13210992024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Raverot G, Ilie MD, Lasolle H, Amodru V,
Trouillas J, Castinetti F and Brue T: Aggressive pituitary tumours
and pituitary carcinomas. Nat Rev Endocrinol. 17:671–684. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Burman P, Casar-Borota O, Perez-Rivas LG
and Dekkers OM: Aggressive pituitary tumors and pituitary
carcinomas: From pathology to treatment. J Clin Endocrinol Metab.
108:1585–1601. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Raverot G and Ilie MD: Immunotherapy in
pituitary carcinomas and aggressive pituitary tumors. Best Pract
Res Clin Endocrinol Metab. 36:1017122022. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Liang Y, Chen B, Liang D, Quan X, Gu R,
Meng Z, Gan H, Wu Z, Sun Y, Liu S and Dou G: Pharmacological
effects of astragaloside IV: A review. Molecules. 28:61182023.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Xia D, Li W, Tang C and Jiang J:
Astragaloside IV, as a potential anticancer agent. Front Pharmacol.
14:10655052023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Li L, Li G, Chen M and Cai R:
Astragaloside IV enhances the sensibility of lung adenocarcinoma
cells to bevacizumab by inhibiting autophagy. Drug Dev Res.
83:461–469. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhu Y and Lu F: Astragaloside IV inhibits
cell viability and glycolysis of hepatocellular carcinoma by
regulating KAT2A-mediated succinylation of PGAM1. BMC Cancer.
24:6822024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yu Y, Hao J, Wang L, Zheng X, Xie C, Liu
H, Wu J, Qiao S and Shi J: Astragaloside IV antagonizes the
malignant progression of breast cancer induced by macrophage M2
polarization through the TGF-β-regulated Akt/Foxo1 pathway. Pathol
Res Pract. 249:1547662023. View Article : Google Scholar
|
|
11
|
Dharmapal D, Jyothy A, Mohan A, Balagopal
PG, George NA, Sebastian P, Maliekal TT and Sengupta S: β-Tubulin
isotype, TUBB4B, regulates the maintenance of cancer stem cells.
Front Oncol. 11:7880242021. View Article : Google Scholar
|
|
12
|
Feng M, Wang K, Fu S, Wei H, Mu X, Li L
and Zhang S: Tubulin TUBB4B Is involved in spermatogonia
proliferation and cell cycle processes. Genes (Basel). 13:10822022.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yang Y, Boza-Serrano A, Dunning CJR,
Clausen BH, Lambertsen KL and Deierborg T: Inflammation leads to
distinct populations of extracellular vesicles from microglia. J
Neuroinflammation. 15:1682018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
McFadden JR, Tolete CDP, Huang Y,
Macnamara E, Sept D, Nesterova G, Gahl WA, Sackett DL and Malicdan
MCV: Clinical, genetic, and structural characterization of a novel
TUBB4B tubulinopathy. Mol Genet Metab Rep. 36:1009902023.PubMed/NCBI
|
|
15
|
Hu H, Zhang Y, Zhai H, Dong J, Zuo L, Guo
X and Wang C: P300 reduces TUBB4B expression to facilitate the
biological process of migration and invasion of non-small cell lung
cancer cells. Tissue Cell. 88:1023862024. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhang W, Zhang P, Pu Y, Chen Z, Su G, Deng
Y, Zhang Y, Ji Y, Huang Z, Zhou Q, et al: DNA
5-hydroxymethylcytosine landscape and transcriptional profile
highlight the TUBB4B-mediated Th17/Th1/Treg imbalance in Behçet's
uveitis. Invest Ophthalmol Vis Sci. 66:282025. View Article : Google Scholar
|
|
17
|
Sobierajska K, Ciszewski WM, Wawro ME,
Wieczorek-Szukała K, Boncela J, Papiewska-Pajak I, Niewiarowska J
and Kowalska MA: TUBB4B downregulation is critical for increasing
migration of metastatic colon cancer cells. Cells. 8:8102019.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Filipovich E, Gorodkova E, Shcherbakova A,
Asaad W, Popov S, Melnichenko G, Mokrysheva N and Utkina M: The
role of cell cycle-related genes in the tumorigenesis of adrenal
and thyroid neuroendocrine tumors. Heliyon. 11:e414572024.
View Article : Google Scholar
|
|
19
|
Liu B, Wang J, Wang G, Jiang W, Li Z, Shi
Y, Zhang J, Pei Q, Huang G, Wang L, et al: Hepatocyte-derived
exosomes deliver H2AFJ to hepatic stellate cells and promote liver
fibrosis via the MAPK/STMN1 axis activation. Int Immunopharmacol.
115:096052023. View Article : Google Scholar
|
|
20
|
Wang C, Zhang H, Xu F, Niu Y, Wu Y, Wang
X, Peng Y, Sun J, Liang L and Xu P: Substituted 3-benzylcoumarins
as allosteric MEK1 inhibitors: Design, synthesis and biological
evaluation as antiviral agents. Molecules. 18:6057–6091. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
22
|
Elking DM: A non-periodic particle mesh
Ewald method for radially symmetric kernels in free space. Comput
Phys Commun. 312:1097392025. View Article : Google Scholar
|
|
23
|
Bussi G, Donadio D and Parrinello M:
Canonical sampling through velocity rescaling. J Chem Phys.
126:0141012007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Hosseini AN and van der Spoel D: Martini
on the rocks: Can a coarse-grained force field model crystals? J
Phys Chem Lett. 15:1079–1088. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zocchi R, Compagnucci C, Bertini E and
Sferra A: Deciphering the tubulin language: Molecular determinants
and readout mechanisms of the tubulin code in neurons. Int J Mol
Sci. 24:27812023. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Liu X, Chu W, Shang S, Ma L, Jiang C, Ding
Y, Wang J, Zhang S and Shao B: Preliminary study on the
anti-apoptotic mechanism of astragaloside IV on radiation-induced
brain cells. Int J Immunopathol Pharmacol. 34:20587384209545942020.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hu T, Fei Z and Wei N: Chemosensitive
effects of astragaloside IV in osteosarcoma cells via induction of
apoptosis and regulation of caspase-dependent Fas/FasL signaling.
Pharmacol Rep. 69:1159–1164. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Liu J, Wang D, Ren N, Zhang L and Wang T:
Metabolites of Astragalus membranaceus and their pro-apoptotic and
cytotoxic activities: Insights into targeted metabolic pathways.
Front Pharmacol. 16:16479582025. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wu M, Li K, Wu J, Zhang Q, Ma X, Dai W,
Gao H, Ding X, Wang W and Xiao W: Astragaloside IV: A multipotent
phytochemical for treating fibrotic diseases (Review). Int J Mol
Med. 57:502026.
|
|
30
|
Xu L and Li Y, He Z, Chen W and Li Y:
Astragaloside IV ameliorates experimental autoimmune myasthenia
gravis through multi-target regulation of
immune-microbiota-metabolism network and ferroptosis inhibition.
Pathol Res Pract. Feb 24–2026.Epub ahead of print. View Article : Google Scholar
|
|
31
|
Sanzhaeva U, Wonsettler NR, Rhodes SB and
Ramamurthy V: TUBB4B is essential for the expansion of
differentiating spermatogonia. Sci Rep. 14:208892024. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Fang Z, Meng Q, Xu J, Wang W, Zhang B, Liu
J, Liang C, Hua J, Zhao Y, Yu X and Shi S: Signaling pathways in
cancer-associated fibroblasts: Recent advances and future
perspectives. Cancer Commun (Lond). 43:3–41. 2023. View Article : Google Scholar :
|
|
33
|
Guo YJ, Pan WW, Liu SB, Shen ZF, Xu Y and
Hu LL: ERK/MAPK signalling pathway and tumorigenesis. Exp Ther Med.
19:1997–2007. 2020.PubMed/NCBI
|
|
34
|
Zhang X, Ji J, Yang Y, Zhang J and Shen L:
Stathmin1 increases radioresistance by enhancing autophagy in
non-small-cell lung cancer cells. Onco Targets Ther. 9:2565–2574.
2016.PubMed/NCBI
|
|
35
|
Wang S, Su T, Tong H, Zhou D, Ma F, Ding
J, Hao Y, Shi W and Quan Z: Circβ-catenin promotes tumor growth and
Warburg effect of gallbladder cancer by regulating STMN1
expression. Cell Death Discov. 7:2332021. View Article : Google Scholar
|
|
36
|
Li J, Kong F, Wu K, Song K, He J and Sun
W: miR-193b directly targets STMN1 and uPA genes and suppresses
tumor growth and metastasis in pancreatic cancer. Mol Med Rep.
10:2613–2620. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zeng L, Lyu X, Yuan J, Chen Y, Wen H,
Zhang L, Shi J, Liu B, Li W and Yang S: STMN1 promotes tumor
metastasis in non-small cell lung cancer through
microtubule-dependent and nonmicrotubule-dependent pathways. Int J
Biol Sci. 20:1509–1527. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li M and Zhou Q: Prognostic value of STMN1
expression in non-small cell lung cancer: A Meta-analysis. Zhongguo
Fei Ai Za Zhi. 27:826–830. 2024.In Chinese.
|
|
39
|
Zhang Y, Wei S, Zhang Q, Zhang Y and Sun
C: Paris saponin VII inhibits triple-negative breast cancer by
targeting the MEK/ERK/STMN1 signaling axis. Phytomedicine.
130:1557462024. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lovrić J, Dammeier S, Kieser A, Mischak H
and Kolch W: Activated raf induces the hyperphosphorylation of
stathmin and the reorganization of the microtubule network. J Biol
Chem. 273:22848–22855. 1998. View Article : Google Scholar
|
|
41
|
Ye Y, Liu L, Feng Z, Liu Y, Miao J, Wei X,
Li H, Yang J, Cao X and Zhao J: The ERK-cPLA2-ACSL4 axis mediating
M2 macrophages ferroptosis impedes mucosal healing in ulcerative
colitis. Free Radic Biol Med. 214:219–235. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Wang X, Li J, Jiang C, Zhang C, Yuan L,
Zhang T, Liu Y, Ma S, Kang P, Li D, et al: Splicing factor FUS
facilitates the progression of PIT1-lineage PitNETs by upregulating
MDM2. Theranostics. 16:3032–3049. 2026. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Mitsutani M, Matsushita M, Yokoyama M,
Morita A, Hano H, Fujikawa T, Tagami T and Moriyama K: Growth
hormone directly stimulates GATA2 expression. Growth Horm IGF Res.
74:1015722024. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Zhou W, Zheng S, Ye L, Su T, Xie J, Jiang
L, Jiang Y, Zhong X, Wu L, Zhou W and Wang W: The USP8 mutational
status in combination with postsurgical cortisol levels for
predicting recurrence of cushing disease. J Clin Endocrinol Metab.
111:e156–e165. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Kaparounaki C, Ilie MD, De Alcubierre D,
Anagnostis P, Haidich AB, Isidori AM, Dekkers OM, Goulis DG and
Raverot G: Medical treatment in acromegaly: A network
meta-analysis. Eur J Endocrinol. 193:S83–S94. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
He D, Wang Q, Sheng Z and Li G:
Preoperative medical therapy for acromegaly: Current knowledge and
clinical significance. Front Endocrinol (Lausanne). 16:16360472026.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Kong P and Tang X, Liu F and Tang X:
Astragaloside IV regulates circ_0001615 and miR-873-5p/LASP1 axis
to suppress colorectal cancer cell progression. Chem Biol Drug Des.
103:e144232024. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Qiu T, Li D, Liu Y, Ren H, Yang X and Luo
W: Astragaloside IV inhibits the proliferation of human uterine
leiomyomas by targeting IDO1. Cancers (Basel). 14:44242022.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhang X, Zhang F, Li Y, Fan N, Zhao K,
Zhang A, Kang J, Lin Y, Xue X and Jiang X: Blockade of PI3K/AKT
signaling pathway by astragaloside IV attenuates ulcerative colitis
via improving the intestinal epithelial barrier. J Transl Med.
22:4062024. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Li M, Niu Y, Tian L, Zhang T, Zhou S, Wang
L, Sun J, Wumiti T, Chen Z, Zhou Q, et al: Astragaloside IV
alleviates macrophage senescence and d-galactose-induced bone loss
in mice through STING/NF-κB pathway. Int Immunopharmacol.
129:1115882024. View Article : Google Scholar
|
|
51
|
Li L, Wang Q, He Y, Sun L, Yang Y and Pang
X: Astragaloside IV suppresses migration and invasion of
TGF-β1-induced human hepatoma HuH-7 cells by regulating
Nrf2/HO-1 and TGF-β1/Smad3 pathways. Naunyn
Schmiedebergs Arch Pharmacol. 395:397–405. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Yu Z, Gao M, Wu X, Li J, Liu B, Wang R,
Tan J, Yu X and Geng L: Potential mechanisms and effects of
AFB1-induced asthma: A comprehensive analysis based on network
toxicology and molecular docking. PLoS One. 21:e03411722026.
View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Zou F, Du Q, Zhang Y, Zuo L and Sun Z:
Pseudo-allergic reactions induced by Chinese medicine injections: A
review. Chin Med. 18:1492023. View Article : Google Scholar : PubMed/NCBI
|