Introduction
Tumor necrosis factor-related apoptosis-inducing
ligand (TRAIL) is a transmembrane protein expressed on surface of
several types of immune cells (1,2).
TRAIL is a member of tumor necrosis factor (TNF) family, and has
recently received much attention because it selectively induces
apoptosis in various cancer types without toxic effect to normal
cells, independently of growth rate and p53 status of the cancer
cells (2). TRAIL triggers
apoptosis upon binding to its death receptors, DR4/TRAIL-R1 or
DR5/TRAIL-R2, leading to activation of caspase-8 which further
activates caspase-3. In some cell types, TRAIL-induced apoptosis
requires contribution of the mitochondrial pathway activated by
caspase-8-mediated Bid cleavage (1). Clinical trials, using recombinant
soluble TRAIL or agonistic monoclonal antibodies (mAb) targeting
TRAIL receptors, are being investigated for cancer therapy, either
as monotherapy or in combination with other drugs (3). The appearance of TRAIL resistance has
been observed in many cancer cell lines and primary tumor cells,
and may be due to defects in TRAIL receptor signaling or
dysregulation of anti-apoptotic proteins controlled by survival
signaling pathways which are occasionally over-activated in cancer
(4). Currently, there is much
interest in searching for natural products that overcome TRAIL
resistance for use during TRAIL treatment (5).
Propolis (bee glue) is a resinous material collected
from various plants by honeybees for use as a defense against
pathogenic microorganisms, and is attractive for pharmacological
research as a rich source of bioactive compounds (6). Propolis extract has been shown to
enhance TRAIL-induced apoptosis (7). However, the bioactivity and chemical
composition of propolis varies with geographic location (6,8,9).
Chrysin (5,7-dihydroxyflavone) is a flavonoid found in some
propolis extracts and in plants (10). There is increasing evidence of
potential benefits of chrysin as a pharmacological agent. Diverse
anticancer effects and mechanisms have been reported for chrysin,
including inducing apoptosis by Akt inhibition (11), inducing cell cycle arrest through
p38-MAPK activation (12), and
enhancing cytotoxicity of chemotherapeutic drugs via cellular
glutathione depletion (13,14).
Chrysin was shown to sensitize HCT-116 human colorectal cancer
cells to TNFα-induced apoptosis, by inhibiting TNFα-induced NF-κB
activation and preventing expression of FLIPL
anti-apoptotic protein which is an NF-κB target gene (15). Chrysin appears to enhance
TRAIL-induced apoptosis in variety of human cancer cell lines
originating from different organs including colorectal cancer,
cervical cancer, hepatocellular carcinoma and nasopharyngeal
carcinoma, suggesting that chrysin may have potential as a TRAIL
sensitizer (16). However, until
now, the mechanism for TRAIL sensitization effect of chrysin is
still not known.
The mechanism of a TRAIL sensitizing agent should be
clear before further study on using it in combination treatment
with TRAIL or agonistic mAb targeting TRAIL receptors in clinical
trials. Therefore, this study investigated the mechanism underlying
the TRAIL sensitization effect of chrysin in TRAIL resistant human
cancer cell lines, by determining the involvement of NF-κB
activation, glutathione depletion, and levels of anti-apoptotic
proteins in TRAIL sensitization, and further exploring the
contributing signaling pathways.
Materials and methods
Propolis extraction and isolation of
compounds
Propolis was collected from a hive of Apis
mellifera from Chiangmai province, Thailand, in August 2006. A
voucher specimen (SWU 0212) was deposited at the Faculty of
Pharmacy, Srinakharinwirot University, Nakhon Nayok province,
Thailand. Thai propolis (1 kg) was extracted by sonication with
methanol (2 liters, 90 min × 3) at room temperature. Removal of
methanol yielded the methanol extract (517 g). Part of the methanol
extract (140 g) was subjected to silica gel column chromatography
using dichloromethane, with increasing concentration of methanol,
give 16 fractions. Chrysin (732 mg) and tectochrysin (314 mg) were
isolated by crystallization from fractions. Chrysin and
tectochrysin were identified by comparison of physical and
spectroscopic data with authentic standard and literature values
(17,18).
Reagents and antibodies
Recombinant human TRAIL and recombinant human tumor
necrosis factor (TNF)-α were purchased from Peprotech (London, UK).
Curcumin was obtained from Fluka Chemika (Steinheim, Switzerland).
N-acetylcysteine (NAC) and
3-(4,5-dimethythiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT),
and chrysin were purchased from Sigma-Aldrich (St. Louis, MO).
STAT3 specific inhibitor (cucurbitacin-I) and IKK-2 inhibitor IV
were obtained from Calbiochem (San Diego, CA). Lipofectamine 2000
reagent and hygromycin B were purchased from Invitrogen (Carlsbad,
CA). Test compounds were dissolved in dimethyl sulfoxide (DMSO) and
kept as a stock solution at −20°C. The final concentration of DMSO
was kept below 0.2% throughout the study. Primary antibodies
specific to caspase-3, poly (ADP-ribose) polymerase (PARP), Mcl-1,
Bcl-XL, XIAP, survivin, FLIP, STAT3 and phosphorylated form of
STAT3 (Tyr705), Akt (Ser473), ERK1/2
(Thr202/Tyr204), p38
(Thr180/Tyr182), JNK
(Thr183/Tyr185), p65 NF-κB
(Ser536) were obtained from Cell Signaling Technology
(Beverly, MA). Primary antibodies specific to actin, pan 14-3-3,
Akt, ERK1/2, p38, JNK and p65 NF-κB were obtained from Santa Cruz
Biotechnology (Santa Cruz, CA).
Cell culture
A549 human lung adenocarcinoma and HeLa human
cervical carcinoma are TRAIL resistant cell lines (19). Cell lines were obtained from the
American Type Culture Collection (ATCC, Rockville, MD). A549 was
cultured in RPMI-1640 supplemented with 10% fetal bovine serum
(FBS), 100 U/ml of penicillin and 100 μg/ml of streptomycin.
HeLa was cultured in DMEM supplemented with 2 mM L-glutamine, 10%
FBS and the antibiotics. The cells were maintained at 37°C in a
humidified atmosphere of 5% CO2.
Cell viability assay
Cell viability after treatment was determined by MTT
method (20). Briefly, cells in
96-well plates (100 μl/well) were pretreated by adding
medium (100 μl) containing test compound to the wells, and
incubated for 30 min, then aliquots of TRAIL in medium (5
μl) were added to each well (final concentration of 100
ng/ml for A549 cells, or 200 ng/ml for HeLa cells), and further
incubated for 24 h. Then, the wells were replaced and incubated
with fresh culture media containing MTT (0.5 mg/ml) for 2 h at
37°C. Finally, the media were removed and DMSO (100 μl) was
added to the wells, and absorbance was measured at 550 nm in a
microplate reader, subtracted with absorbance at 650 nm. The number
of viable cells was determined from the absorbance. Assays were
performed in triplicate wells. Data were expressed as percent
viability compared with control.
Establishment of stable NF-κB reporter
cell line
A549 cells were transfected with
pGL4.32[luc2P/NF-κB-RE/Hygro] vector (Promega, Madison, WI)
using Lipofectamine 2000 reagent according to manufacturer’s
protocol. After transfection for 24 h, the transfected cells were
selected by sub-culturing into new flasks containing RPMI-1640
supplemented with 10% FBS and 800 μg/ml hygromycin B and
maintained in this media over 2 weeks to obtain the stable
transfected cells (A549/NF-κB).
NF-κB transcriptional activity assay
NF-κB transcriptional activity was determined by
luciferase reporter gene assay. A549/NF-κB cells were seeded into a
96-well plate (100 μl/well), and left overnight. Cells were
pretreated with test compound (100 μl) for 30 min, followed
by adding 5 μl of medium containing TNFα or TRAIL, and
further incubated for 3 h. Cells were then lysed with 25 μl
of passive lysis buffer (Promega), and 20 μl of cell lysate
was mixed with 50 μl of luciferase assay reagent (Promega).
Luminescence was determined using a luminometer (Atto, Tokyo,
Japan).
Immunoblot analysis
Immunoblot analysis was performed as previously
described (21). Briefly, whole
cell lysates were subjected to electrophoresis in 7.5 or 10%
SDS-PAGE, and electrophoretically transferred to Immobilon-P nylon
membrane (Millipore, Bedford, MA). The membranes were blocked with
BlockAce (Dainippon Pharmaceutical, Co. Ltd., Osaka, Japan) for at
least 2 h, and probed with the indicated primary antibodies
overnight, followed by horseradish peroxidase-conjugated secondary
antibodies (Dako, Glostrup, Denmark). Bands were visualized using
ECL reagents (Amersham Bioscience, Piscataway, NJ).
Quantitative reverse
transcriptase-polymerase chain reaction (qRT-PCR)
Isolation of total cellular RNA from cells was
performed using RNeasy mini kit (Qiagen, Valencia, CA), and 2
μg of the total cellular RNA was used to synthesize
first-strand cDNA by using SuperScript III Reverse Transcriptase
Kit (Invitrogen). Quantitative determination of gene expression was
performed by real-time PCR on a LightCycler® 2.0
Instrument (Roche). PCR primer sequences were obtained from
following references; human Mcl-1 (22); human β-actin (23). Mcl-1 primer (fwd) 5’-CGG TAA TCG
GAC TCA ACC TC-3’; and Mcl-1 primer (rev) 5’-CCT CCT TCT CCG TAG
CCA A-3’. β-actin primer (fwd) 5’-GAC CTG ACT GAC TAC CTC ATG A-3’;
and β-actin primer (rev) 5’-AGC ATT TGC GGT GGA CGA TGG AG-3’. The
PCR reaction mixture (20 μl) was composed of 1 μl of
5-fold diluted cDNA solution, 10 μl QuantiTect™ SYBR-Green
PCR Master mix (Qiagen), PCR primers and sterile distilled water.
An initial activation step at 95°C for 15 min was followed by 40
cycles comprising denaturation at 94°C/15 sec, annealing at 55°C/30
sec, and extension at 72°C/30 sec. Expression level of Mcl-1 gene
was normalized to β-actin gene, and calculated from crossing point
(Cp) value of the sample, relative gene expression level
= 2(Cpactin−
Cptarget). The relative change in gene
expression compared with untreated conditions was expressed as fold
change, calculated by 2−ΔΔCp method (24).
Transfection of small interfering
RNAs
Silencing of Mcl-1 expression in A549 cells was
performed as previously described (19). Human Mcl-1 Stealth RNAi™ siRNA
(HSS181042) was purchased from Invitrogen. Firefly luciferase (GL2)
siRNA was obtained from Hokkaido System Science Co. Ltd. (Sapporo,
Japan), and used as control siRNA (19). A549 cells were transfected with
siRNAs at a final concentration of 20 nM, using Lipofectamine 2000
reagent. Cells were treated with TRAIL at 24 h after
transfection.
Statistical analysis
Data are expressed as mean + SD, and analyzed by
Student’s two-tailed t-test to determine the significance of
differences between groups. A p-value of lower than 0.05 was
considered to be significant.
Results
Enhancement of TRAIL-induced apoptosis in
human cancer cells by Thai propolis extract and chrysin
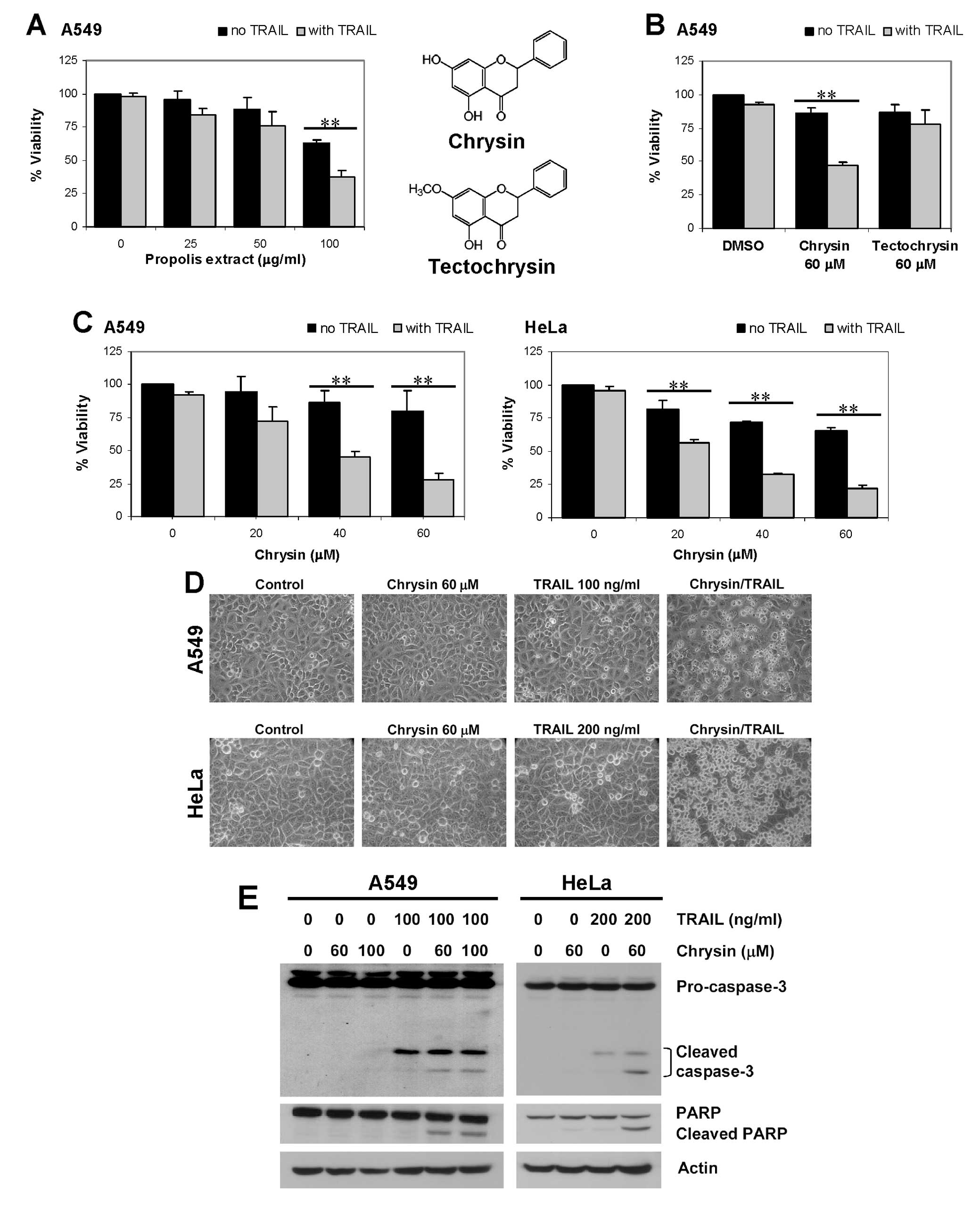
Initially, TRAIL sensitization effect of Thai
propolis extract was examined in a TRAIL resistant cell line, A549.
Pretreatment with propolis extract (25–100 μg/ml) for 30
min, followed by TRAIL stimulation for 24 h, enhanced TRAIL-induced
cell death. At the highest propolis concentration test (100
μg/ml), cell viability of the combination treatment was 38%,
compared with 63 and 98% viability with propolis alone and TRAIL
alone, respectively (Fig. 1A).
Chrysin and tectochrysin were found to be two major constituents of
the Thai propolis extract. Chrysin enhanced TRAIL-induced cell
death, with cell viability being 47% in the combination treatment,
compared to 86 and 92% viability with chrysin alone and TRAIL
alone, respectively, while tectochryin did not show significant
effect (Fig. 1B). TRAIL
sensitization effect of chrysin was observed in a dose-dependent
manner over the tested ranges (20–60 μM) in both A549 and
HeLa cell lines, which originated from different organs (Fig. 1C).
Hallmarks of cells undergoing apoptosis are cleavage
of procaspase-3 and poly(ADP-ribose) polymerase (PARP). Massive
cell death of A549 and HeLa cells could be observed as early as 3 h
after treatment with chrysin/TRAIL combination (Fig. 1D), concordant with enhancement of
apoptosis as clearly shown by increased cleavage of pro-caspase-3
and PARP (Fig. 1E). Our results
revealed that apoptosis-inducing effect of TRAIL was enhanced by
chrysin.
TRAIL sensitization effect of chrysin is
not mediated by NF-κB inhibition
In addition to inducing apoptosis signaling, TRAIL
and TNFα receptors also activate survival signaling such NF-κB
which regulates expression of several anti-apoptotic proteins to
counteract the apoptosis signaling (1). Since chrysin was shown to sensitize
TNFα-induced apoptosis in human cancer cells by inhibiting NF-κB
activation (15), thus, we
examined effect of chrysin on TNFα- or TRAIL-induced NF-κB
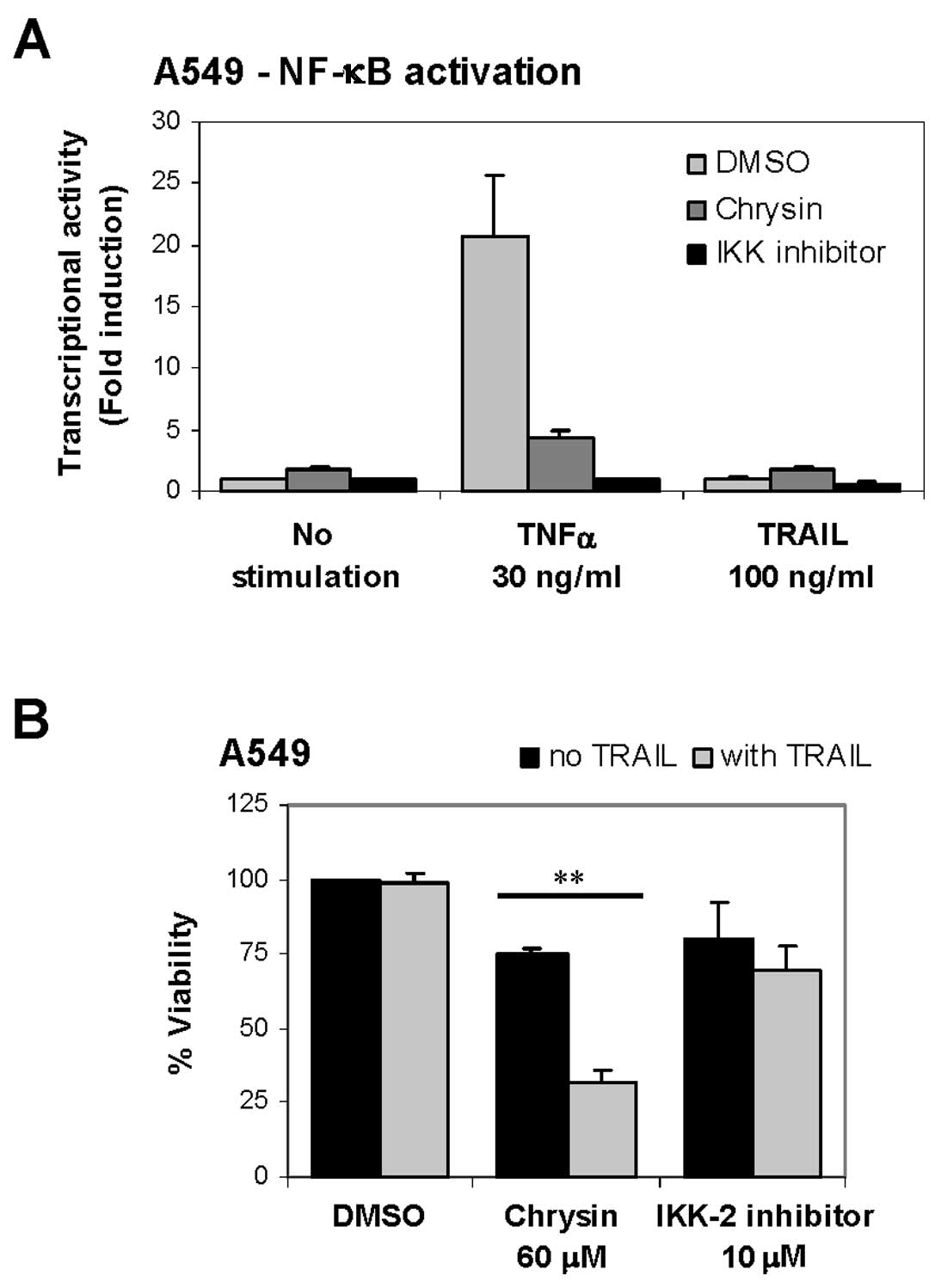
activation in A549/NF-κB stable cell line. Stimulation of TNFα for
3 h induced activation and increased transcriptional activity of
NF-κB by 20.8-fold compared to control (Fig. 2A). TNFα-induced NF-κB activation
was diminished by chrysin pretreatment (60 μM), with 79%
inhibition observed, while IKK-2 inhibitor pretreatment (10
μM) led complete inhibition (Fig. 2A). In contrast, TRAIL stimulation
slightly induced NF-κB activation by 1.1-fold, while NF-κB
activation was more increased by chrysin alone or chrysin/TRAIL
combination treatment, with 1.8- and 1.9-fold induction
respectively (Fig. 2A). These
results indicated that chrysin did not inhibit TRAIL-induced NF-κB
activation in A549 cells. Furthermore, inhibition of TRAIL-induced
NF-κB activation by IKK-2 inhibitor did not significantly enhance
cell death in A549 cells (Fig.
2B), indicating that inhibition of NF-κB activation is unlikely
to be the mechanism of chrysin for TRAIL sensitization.
Glutathione depletion does not contribute
to TRAIL sensitization effect of chrysin
Chrysin has been shown to sensitize A549 cells to
chemotherapeutic drugs by depleting intracellular glutathione
(13,14). N-acetylcysteine (NAC), a precursor
of glutathione synthesis, has been shown to reverse TRAIL
sensitization by curcumin (25).
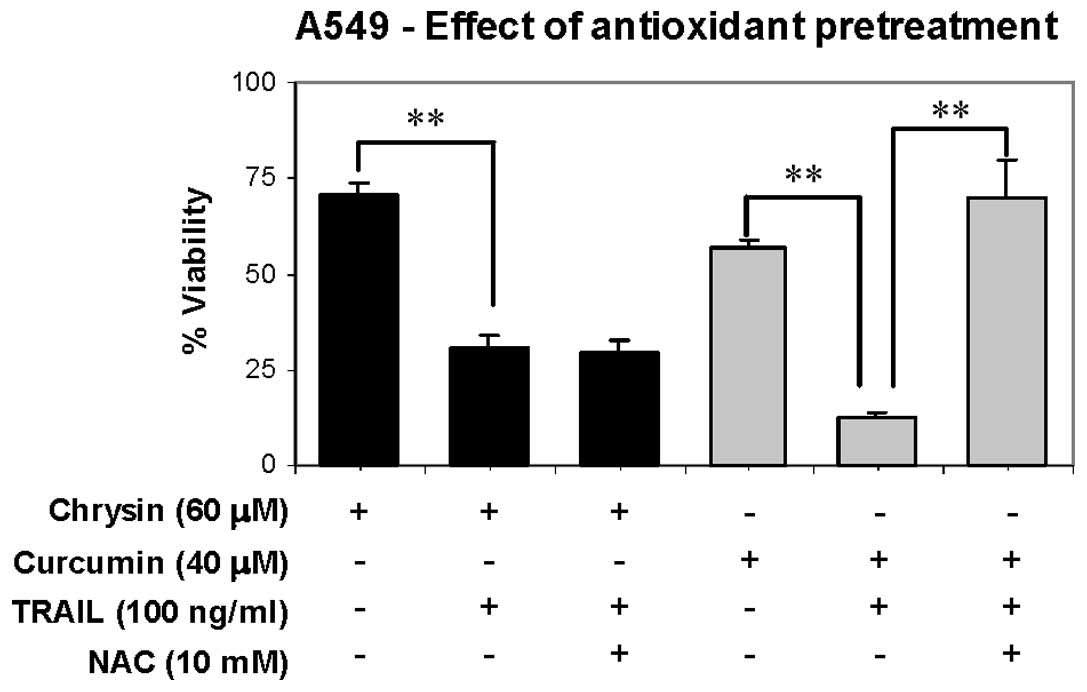
We evaluated the involvement of glutathione depletion in TRAIL
sensitization effect of chrysin by pretreatment of A549 cells with
NAC (10 mM) for 2 h, to increase intracellular glutathione level
before exposure to TRAIL combination treatment. If the cytotoxic
effect of the combination treatment is mediated by glutathione
depletion effect, the NAC pretreatment will abolish the cytotoxic
effect. The results showed that TRAIL sensitization effect of
chrysin was unaffected by NAC pretreatment, but TRAIL sensitization
effect of curcumin was potently reversed by NAC pretreatment
(Fig. 3). The results clearly
indicated that glutathione depletion does not contribute to the
TRAIL sensitization effect of chrysin.
Mcl-1 downregulation mediates TRAIL
sensitization effect of chrysin
Many naturally occurring polyphenols enhance TRAIL
sensitization through downregulation of anti-apoptotic proteins
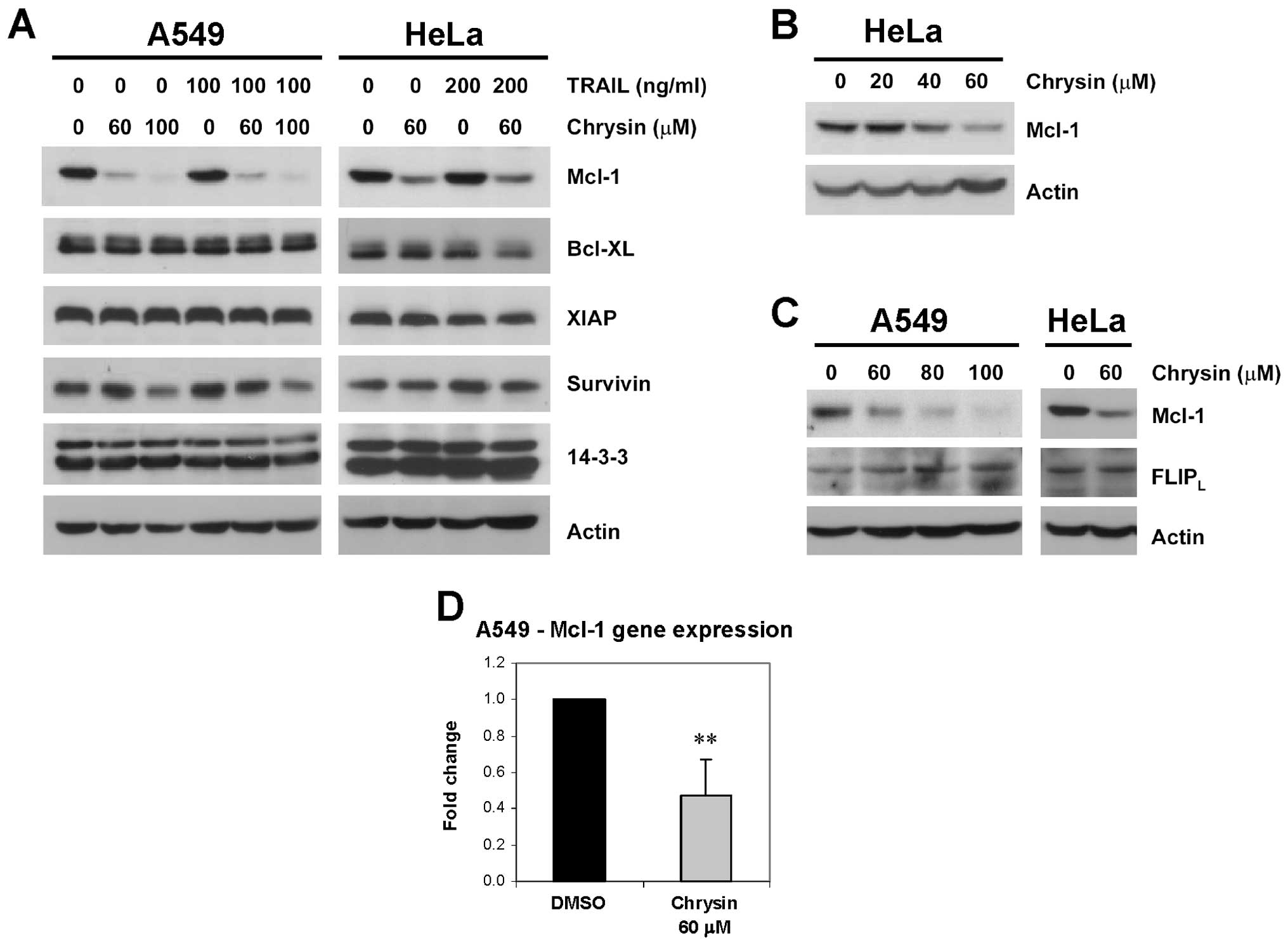
(26). We further examined effect
of chrysin on levels of several anti-apoptotic proteins, including
Mcl-1, Bcl-XL, XIAP, survivin and 14-3-3, in both A549 and HeLa
cells. The results showed that, at 3 h after treatment, chrysin
alone (60–100 μM) or in combination with TRAIL selectively
decreased Mcl-1 protein level, while the levels of other proteins
remained unaffected (Fig. 4A). The
dose-dependent decrease in Mcl-1 protein level by chrysin was
observed in both A549 and HeLa cell lines (Fig. 4A and B).
Effect of chrysin on another short half-life
anti-apoptotic protein, FLIP, was examined compared with Mcl-1. The
rapid turnover rate of both proteins in cells is mediated by the
proteasome (27). FLIP has 2
isoforms - FLIPL and FLIPS, but
FLIPS was not detected in our cell lines. Chrysin
treatment (60 μM) for 3 h decreased Mcl-1 level, while
FLIPL levels in both cell lines were not reduced
(Fig. 4C).
To explore the cause of Mcl-1 reduction by chrysin,
we determined mRNA level of Mcl-1 gene after treatment by using
qRT-PCR. Effect of chrysin on Mcl-1 gene expression was clearly
demonstrated, chrysin (60 μM) treatment for 3 h reduced
Mcl-1 mRNA level by 52% (Fig. 4D).
The results suggested that modulation of TRAIL sensitivity of
cancer cells by chrysin was occurred through downregulation of
Mcl-1 gene expression.
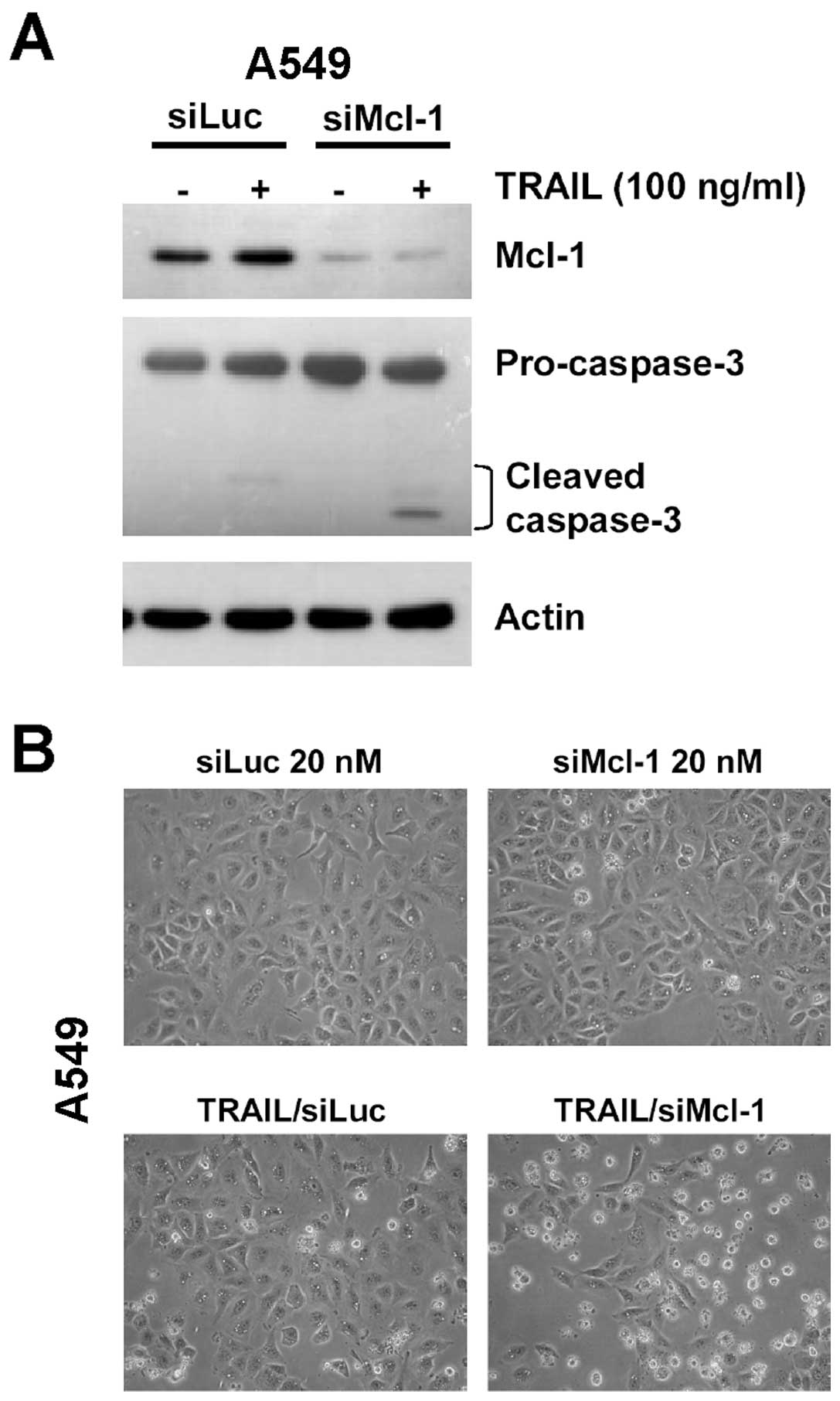
Silencing Mcl-1 expression is sufficient
to enhances TRAIL-induced apoptosis
To confirm that reduction of Mcl-1 level was able to
sensitize cells to TRAIL-induced apoptosis, siMcl-1 transfection
was used to silence Mcl-1 expression in A549 cells, while siLuc was
used as control siRNA. At 24 h after transfection, the level of
Mcl-1 protein was obviously decreased in siMcl-1 transfected cells
compared with siLuc transfected cells (Fig. 5A). After TRAIL stimulation for 3 h,
many apoptotic cells were observed in the siMcl-1 transfected cells
(Fig. 5B), correlating with
cleavage of pro-caspase-3, while the procaspase-3 cleavage was
faintly detected in TRAIL-stimulated siLuc transfected cells
(Fig. 5A). Taken together, the
results confirmed that TRAIL sensitization effect of chrysin was
mediated by Mcl-1 downregulation. Thus, we further investigated
mechanism of Mcl-1 modulation by chrysin.
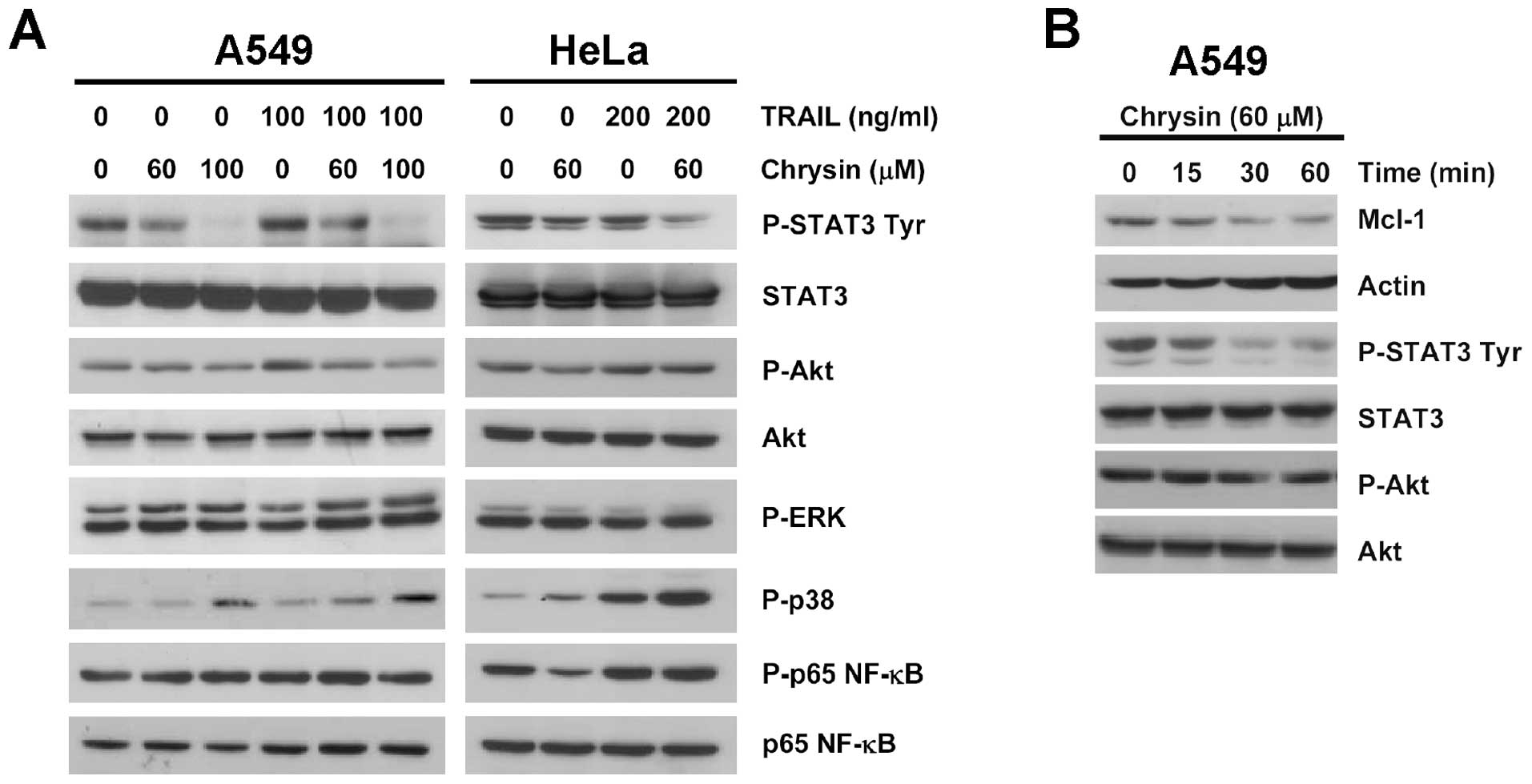
Chrysin downregulates Mcl-1 expression
through inhibiting STAT3 tyrosine phosphorylation
Mcl-1 gene expression is regulated by multiple
signaling pathways, including STAT3, Akt and p38 pathways (28). We examined the effects of chrysin
treatment (60–100 μM) for 3 h, either alone or in
combination with TRAIL, on several survival signaling pathways
including STAT3, Akt, ERK, p38 and NF-κB. Among the selected
pathways, marked inhibition was obtained only in constitutive STAT3
tyrosine phosphorylation (Tyr705), in both A549 and HeLa
cells (Fig. 6A). Tyr705
phosphorylation of STAT3 is necessary for DNA-binding and
transcriptional activity of STAT3 (29). Correlation between decreased STAT3
tyrosine phosphorylation and Mcl-1 down-regulation was observed in
A549 cells treated with chrysin in a time-dependent manner, while
Akt phosphorylation was not affected within the time-course
measured (Fig. 6B). The results
suggested that STAT3 was the target of chrysin for modulating level
of Mcl-1 expression.
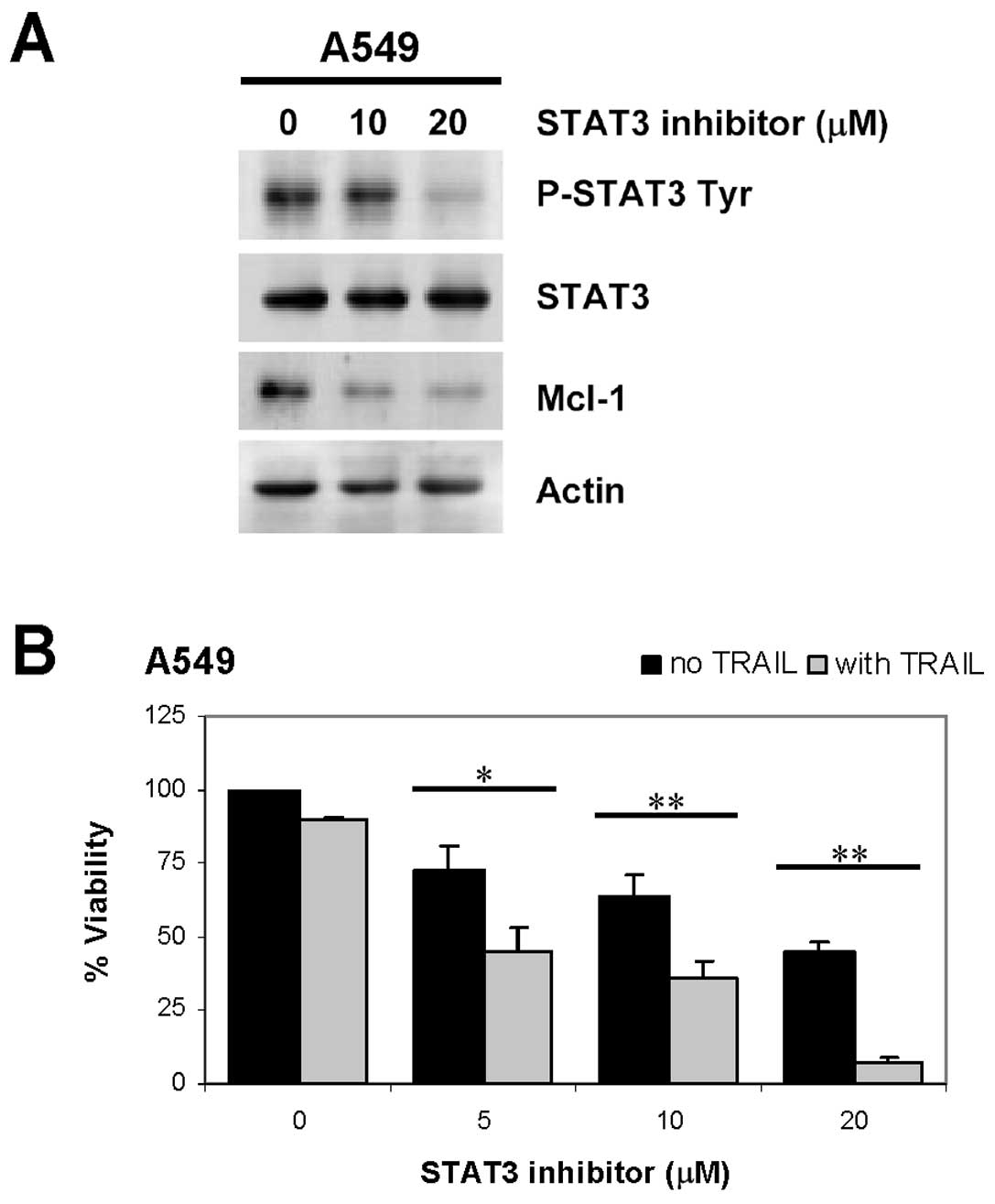
STAT3 specific inhibitor decreases Mcl-1
level and enhances TRAIL-induced cell death
To confirm that Mcl-1 expression is regulated by
STAT3, A549 cells was treated with a specific inhibitor of STAT3
(cucurbitacin-I) for 3 h, the result showed that the STAT3
inhibitor decreased both STAT3 tyrosine phosphorylation and Mcl-1
level (Fig. 7A). Furthermore, the
STAT3 inhibitor enhanced TRAIL-induced cell death (Fig. 7B) in a dose-dependent manner,
similar to that observed with chrysin treatment. Collectively, our
results indicated that the TRAIL sensitization effect of chrysin
was mediated by Mcl-1 down-regulation through inhibition of
constitutive STAT3 tyrosine phosphorylation.
Discussion
Induction of apoptosis via TRAIL receptor is a
promising targeted therapy, but this strategy also faces a similar
problem to the use of other chemotherapeutic drugs, because TRAIL
resistance is found in variety of cancers. The TRAIL sensitization
effect of chrysin was recently reported (16), but the underlying mechanism has not
been elucidated until now. In the present study, we demonstrated a
novel mechanism for apoptosis-sensitizing effect of chrysin that
overcomes TRAIL resistance of cancer cells.
We demonstrated that downregulation of Mcl-1
anti-apoptotic protein by STAT3 inhibition is the mechanism for
chrysin in enhancing the apoptosis-inducing effect of TRAIL. Mcl-1
is a Bcl-2 family anti-apoptotic protein, contributing to TRAIL
resistance by sequestering several proapoptotic proteins, such as
Bak, Bim and Puma, preventing their function to release cytochrome
c from mitochondria during TRAIL stimulation (30). STAT3 regulates anti-apoptotic genes
including Mcl-1, survivin and Bcl-XL (31). However, Mcl-1 differs from other
anti-apoptotic proteins by having a very short intracellular
half-life (∼40 min) and rapid turnover rate (32). This could explain the rapid
apoptosis induction by the chrysin/TRAIL combination, which could
be observed as early as 3 h after treatment (Fig. 1E). While chrysin treatment
decreased Mcl-1 level, the treatment did not change the level of
FLIPL, another short half-life anti-apoptotic protein
(33), indicating the specific
effect of chrysin. Results of silencing Mcl-1 expression by siRNA
confirmed that the decreased Mcl-1 level by chrysin was sufficient
to enhance apoptosis-inducing effect of TRAIL. The mechanism of
reversal of TRAIL resistance by STAT3/Mcl-1 downregulation was not
only found in our lung and cervical cancer cells, but was also
observed in variety of cancer types including breast cancer,
pancreatic cancer and hepatocellular carcinoma (34–36).
The reduction of Mcl-1 level by chrysin has been
observed in other cancer cell types including anaplastic thyroid
cancer and T-cell leukemia (37,38).
The report of Polier et al demonstrated that Mcl-1
downregulation by chrysin in T cell leukemia was a result of
inhibiting a transcription regulator, cyclin-dependent kinase 9
(CDK9), and subsequent reduction in Ser2 phosphorylation
of RNA polymerase II (RNA pol II), leading to decrease in Mcl-1
gene expression in T-cell leukemic cell line (37). The Ser2 phosphorylation
in the carboxy-terminal domain of RNA pol II by CDK9 activity is
required for the transcription elongation stage of mRNA synthesis
(39). It has been reported that,
during upregulation of STAT3 target genes,
Tyr705-phosphorylated STAT3 directly associated with
CDK9 in nucleus, and this complex recruited RNA pol II to promoter
site of the target genes, increasing Ser2
phosphorylation of the RNA pol II, allowing transcriptional
elongation of mRNA synthesis (40,41).
The implication of CDK9 and RNA pol II in expression regulation of
STAT3 target genes coordinates the findings of Polier et al
(37) and our results to depict
the molecular details of the mechanism of Mcl-1 downregulation by
chrysin via STAT3 inhibition.
Chrysin has been shown to inhibit Akt
phosphorylation in cancer cells (11,42).
However, we did not observe inhibition of Akt phosphorylation by
chrysin within the first 3 h after treatment (Fig. 6A), when the TRAIL-induced apoptosis
had already progressed (Fig. 1E),
and Mcl-1 level and STAT3 tyro-sine phosphorylation were
concomitantly decreased (Fig. 6B).
Our results suggested that Akt inhibition was not the primary
mechanism responsible for TRAIL sensitization effect of chrysin in
our cellular models.
Although, inhibition of NF-κB activation was the
mechanism of chrysin for enhancement of TNFα-induced apoptosis in
HCT116 colorectal cancer cells (15), chrysin did not inhibit
TRAIL-induced NF-κB activation in A549 cells. Moreover, chrysin
alone slightly increased NF-κB transcriptional activity (Fig. 2A), which was also observed in HeLa
cells (data not shown). This phenomenon is consistent with a
previous report showing that, in HeLa cells, chrysin induced DNA
binding activity of p65 NF-κB which could be reversed by NAC
pretreatment (43), indicating
involvement of a glutathione depletion effect in the increased DNA
binding activity of NF-κB by chrysin.
Our results clearly showed that NAC pretreatment did
not abolish the TRAIL sensitization effect of chrysin, in contrast
to curcumin, another glutathione depleting agent (Fig. 3), suggesting that the glutathione
depletion effect did not contribute to TRAIL sensitization effect
of chrysin. A recent report showed the ability of chrysin to
activate AMP-activated protein kinase (AMPK), and this AMPK
activation mediated enhancement of doxorubicin cytotoxicity in A549
cells (44). The report also
demonstrated that AMPK activation led to inhibition of Akt
phosphorylation and NAC pretreatment could reverse the AMPK
activation. We suggest that the AMPK activation is unlikely to be
involved in TRAIL sensitization effect of chrysin, because our
results revealed that NAC pretreatment did not reverse TRAIL
sensitization effect of chrysin (Fig.
3). We suggest that mechanisms of the biological effects of
chrysin can be divided into glutathione depletion-dependent and
-independent mechanisms.
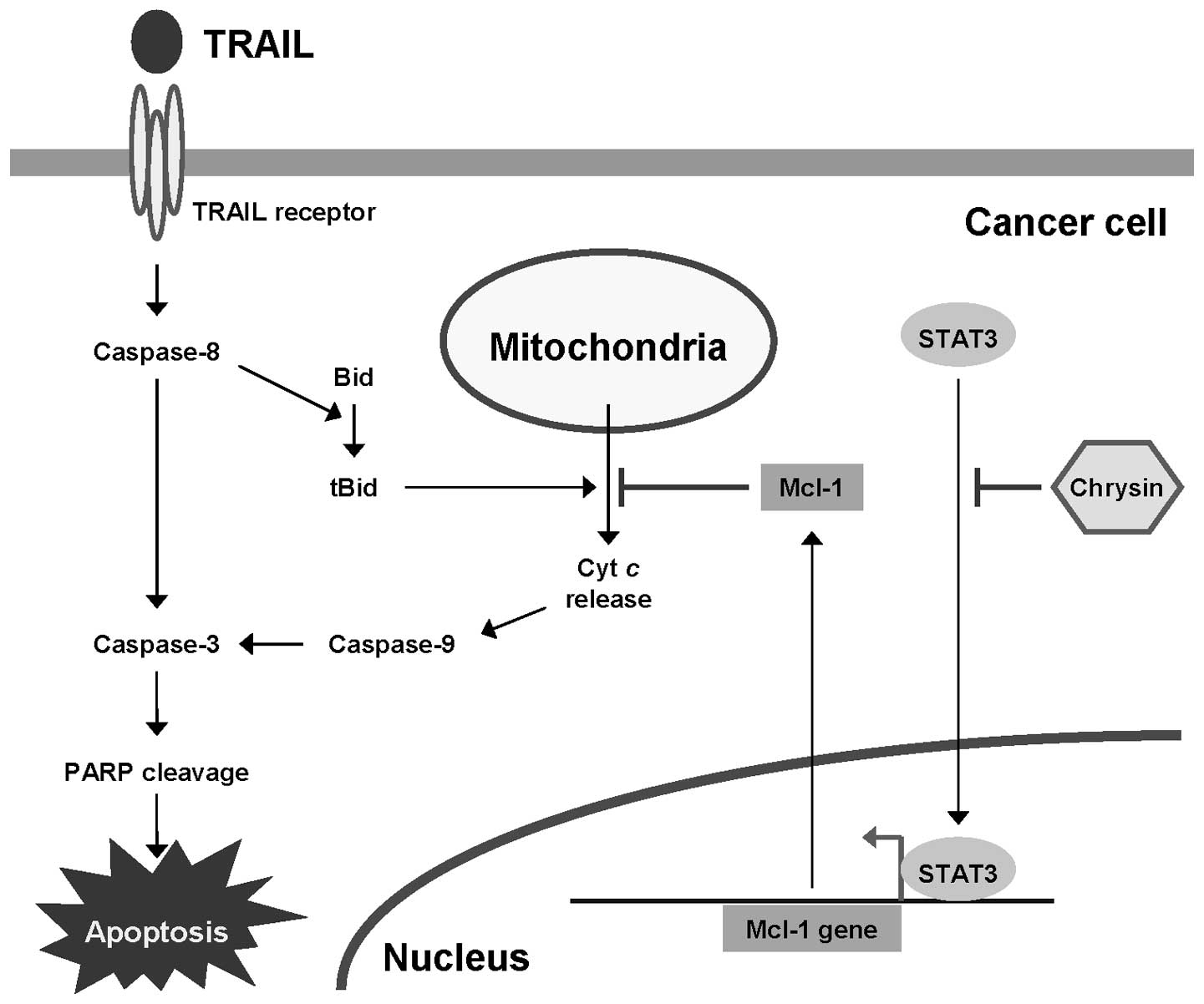
In conclusion, as depicted in Fig. 8, we have demonstrated that chrysin
can inhibit constitutive STAT3 activation, resulting in Mcl-1
downregulation, and this activity underlies the overcoming TRAIL
resistance of cancer cells by chrysin. Our study suggests the
potential of chrysin for use as a TRAIL sensitizer.
Acknowledgements
This study was supported by the
Thailand Research Fund (TRG5380026 to K.L.), a Japanese-Thai
Collaborative Scientific Research Fellowship (JSPS-NRCT; to K.L.,
S.A. and A.V.), and a research grant from the Chulabhorn Research
Institute. We would like to thank Ms. Yue Zhou, Dr Orawin
Prangsaengtong, Dr Rudee Surarit, Ms. Korakot Atjanasuppat, and Ms.
Khajeelak Chiablaem for their assistance.
References
|
1.
|
Kruyt FA: TRAIL and cancer therapy. Cancer
Lett. 263:14–25. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2.
|
Wang S: The promise of cancer therapeutics
targeting the TNF-related apoptosis-inducing ligand and TRAIL
receptor pathway. Oncogene. 27:6207–6215. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3.
|
Hellwig CT and Rehm M: TRAIL signaling and
synergy mechanisms used in TRAIL-based combination therapies. Mol
Cancer Ther. 11:3–13. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4.
|
Mahalingam D, Szegezdi E, Keane M, de Jong
S and Samali A: TRAIL receptor signalling and modulation: are we on
the right TRAIL? Cancer Treat Rev. 35:280–288. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5.
|
Ishibashi M and Ohtsuki T: Studies on
search for bioactive natural products targeting TRAIL signaling
leading to tumor cell apoptosis. Med Res Rev. 28:688–714. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Bankova V: Recent trends and important
developments in propolis research. Evid Based Complement Alternat
Med. 2:29–32. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7.
|
Szliszka E, Czuba ZP, Domino M, Mazur B,
Zydowicz G and Krol W: Ethanolic extract of propolis (EEP) enhances
the apoptosis-inducing potential of TRAIL in cancer cells.
Molecules. 14:738–754. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8.
|
Awale S, Shrestha SP, Tezuka Y, Ueda JY,
Matsushige K and Kadota S: Neoflavonoids and related constituents
from Nepalese propolis and their nitric oxide production inhibitory
activity. J Nat Prod. 68:858–864. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
9.
|
Awale S, Li F, Onozuka H, Esumi H, Tezuka
Y and Kadota S: Constituents of Brazilian red propolis and their
preferential cytotoxic activity against human pancreatic PANC-1
cancer cell line in nutrient-deprived condition. Bioorg Med Chem.
16:181–189. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10.
|
Hernandez J, Goycoolea FM, Quintero J, et
al: Sonoran propolis: chemical composition and antiproliferative
activity on cancer cell lines. Planta Med. 73:1469–1474. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
11.
|
Woo KJ, Jeong YJ, Park JW and Kwon TK:
Chrysin-induced apoptosis is mediated through caspase activation
and Akt inactivation in U937 leukemia cells. Biochem Biophys Res
Commun. 325:1215–1222. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12.
|
Weng MS, Ho YS and Lin JK: Chrysin induces
G1 phase cell cycle arrest in C6 glioma cells through inducing
p21Waf1/Cip1 expression: involvement of p38 mitogen-activated
protein kinase. Biochem Pharmacol. 69:1815–1827. 2005. View Article : Google Scholar
|
|
13.
|
Kachadourian R, Leitner HM and Day BJ:
Selected flavonoids potentiate the toxicity of cisplatin in human
lung adenocarcinoma cells: a role for glutathione depletion. Int J
Oncol. 31:161–168. 2007.
|
|
14.
|
Brechbuhl HM, Kachadourian R, Min E, Chan
D and Day BJ: Chrysin enhances doxorubicin-induced cytotoxicity in
human lung epithelial cancer cell lines: the role of glutathione.
Toxicol Appl Pharmacol. 258:1–9. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15.
|
Li X, Huang Q, Ong CN, Yang XF and Shen
HM: Chrysin sensitizes tumor necrosis factor-alpha-induced
apoptosis in human tumor cells via suppression of nuclear
factor-kappaB. Cancer Lett. 293:109–116. 2010. View Article : Google Scholar
|
|
16.
|
Li X, Wang JN, Huang JM, et al: Chrysin
promotes tumor necrosis factor (TNF)-related apoptosis-inducing
ligand (TRAIL) induced apoptosis in human cancer cell lines.
Toxicol In Vitro. 25:630–635. 2011. View Article : Google Scholar
|
|
17.
|
Wagner H, Chari VM and Sonnenbichler J:
13C-NMR-spektren natürlich vorkommender flavonoide. Tetrahedron
Lett. 17:1799–1802. 1976.(In German).
|
|
18.
|
Gaydou EM and Bianchini JP: Études de
composés flavoniques I. Synthèses et propriétés (UV, RMN du 13C) de
quelques flavones. Bull Soc Chim Fr II. 43–47. 1978.
|
|
19.
|
Choo MK, Kawasaki N, Singhirunnusorn P, et
al: Blockade of transforming growth factor-beta-activated kinase 1
activity enhances TRAIL-induced apoptosis through activation of a
caspase cascade. Mol Cancer Ther. 5:2970–2976. 2006. View Article : Google Scholar
|
|
20.
|
Lirdprapamongkol K, Kramb JP,
Suthiphongchai T, et al: Vanillin suppresses metastatic potential
of human cancer cells through PI3K inhibition and decreases
angiogenesis in vivo. J Agric Food Chem. 57:3055–3063. 2009.
View Article : Google Scholar
|
|
21.
|
Lirdprapamongkol K, Sakurai H, Suzuki S,
et al: Vanillin enhances TRAIL-induced apoptosis in cancer cells
through inhibition of NF-kappaB activation. In Vivo. 24:501–506.
2010.
|
|
22.
|
Yu C, Bruzek LM, Meng XW, et al: The role
of Mcl-1 downregulation in the proapoptotic activity of the
multikinase inhibitor BAY 43-9006. Oncogene. 24:6861–6869. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
23.
|
Unger RE, Krump-Konvalinkova V, Peters K
and Kirkpatrick CJ: In vitro expression of the endothelial
phenotype: comparative study of primary isolated cells and cell
lines, including the novel cell line HPMEC-ST1.6R. Microvasc Res.
64:384–397. 2002. View Article : Google Scholar
|
|
24.
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
25.
|
Jung EM, Lim JH, Lee TJ, Park JW, Choi KS
and Kwon TK: Curcumin sensitizes tumor necrosis factor-related
apoptosis-inducing ligand (TRAIL)-induced apoptosis through
reactive oxygen species-mediated upregulation of death receptor 5
(DR5). Carcinogenesis. 26:1905–1913. 2005. View Article : Google Scholar
|
|
26.
|
Jacquemin G, Shirley S and Micheau O:
Combining naturally occurring polyphenols with TNF-related
apoptosis-inducing ligand: a promising approach to kill resistant
cancer cells? Cell Mol Life Sci. 67:3115–3130. 2010. View Article : Google Scholar
|
|
27.
|
Bannerman DD, Tupper JC, Ricketts WA,
Bennett CF, Winn RK and Harlan JM: A constitutive cytoprotective
pathway protects endothelial cells from lipopolysaccharide-induced
apoptosis. J Biol Chem. 276:14924–14932. 2001. View Article : Google Scholar
|
|
28.
|
Akgul C: Mcl-1 is a potential therapeutic
target in multiple types of cancer. Cell Mol Life Sci.
66:1326–1336. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29.
|
Aggarwal BB, Kunnumakkara AB, Harikumar
KB, et al: Signal transducer and activator of transcription-3,
inflammation, and cancer: how intimate is the relationship? Ann NY
Acad Sci. 1171:59–76. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
30.
|
Meng XW, Lee SH, Dai H, et al: Mcl-1 as a
buffer for proapoptotic Bcl-2 family members during TRAIL-induced
apoptosis: a mechanistic basis for sorafenib (Bay 43-9006)-induced
TRAIL sensitization. J Biol Chem. 282:29831–29846. 2007. View Article : Google Scholar
|
|
31.
|
Hsieh FC, Cheng G and Lin J: Evaluation of
potential Stat3-regulated genes in human breast cancer. Biochem
Biophys Res Commun. 335:292–299. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
32.
|
Nijhawan D, Fang M, Traer E, et al:
Elimination of Mcl-1 is required for the initiation of apoptosis
following ultraviolet irradiation. Genes Dev. 17:1475–1486. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
33.
|
Poukkula M, Kaunisto A, Hietakangas V, et
al: Rapid turnover of c-FLIPshort is determined by its unique
C-terminal tail. J Biol Chem. 280:27345–27355. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
34.
|
Dong Y, Yin S, Li J, Jiang C, Ye M and Hu
H: Bufadienolide compounds sensitize human breast cancer cells to
TRAIL-induced apoptosis via inhibition of STAT3/Mcl-1 pathway.
Apoptosis. 16:394–403. 2011. View Article : Google Scholar
|
|
35.
|
Huang S and Sinicrope FA: Sorafenib
inhibits STAT3 activation to enhance TRAIL-mediated apoptosis in
human pancreatic cancer cells. Mol Cancer Ther. 9:742–750. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
36.
|
Chen KF, Tai WT, Liu TH, et al: Sorafenib
overcomes TRAIL resistance of hepatocellular carcinoma cells
through the inhibition of STAT3. Clin Cancer Res. 16:5189–5199.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
37.
|
Polier G, Ding J, Konkimalla BV, et al:
Wogonin and related natural flavones are inhibitors of CDK9 that
induce apoptosis in cancer cells by transcriptional suppression of
Mcl-1. Cell Death Dis. 2:e1822011. View Article : Google Scholar : PubMed/NCBI
|
|
38.
|
Phan T, Yu XM, Kunnimalaiyaan M and Chen
H: Antiproliferative effect of chrysin on anaplastic thyroid
cancer. J Surg Res. 170:84–88. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39.
|
Wang S and Fischer PM: Cyclin-dependent
kinase 9: a key transcriptional regulator and potential drug target
in oncology, virology and cardiology. Trends Pharmacol Sci.
29:302–313. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
40.
|
Giraud S, Hurlstone A, Avril S and
Coqueret O: Implication of BRG1 and cdk9 in the STAT3-mediated
activation of the p21waf1 gene. Oncogene. 23:7391–7398. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
41.
|
Hou T, Ray S and Brasier AR: The
functional role of an interleukin 6-inducible CDK9. STAT3 complex
in human gamma-fibrinogen gene expression. J Biol Chem.
282:37091–37102. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
42.
|
Fu B, Xue J, Li Z, Shi X, Jiang BH and
Fang J: Chrysin inhibits expression of hypoxia-inducible
factor-1alpha through reducing hypoxia-inducible factor-1alpha
stability and inhibiting its protein synthesis. Mol Cancer Ther.
6:220–226. 2007. View Article : Google Scholar
|
|
43.
|
Von Brandenstein MG, Ngum Abety A, Depping
R, et al: A p38-p65 transcription complex induced by endothelin-1
mediates signal transduction in cancer cells. Biochim Biophys Acta.
1783:1613–1622. 2008.PubMed/NCBI
|
|
44.
|
Shao JJ, Zhang AP, Qin W, Zheng L, Zhu YF
and Chen X: AMP-activated protein kinase (AMPK) activation is
involved in chrysin-induced growth inhibition and apoptosis in
cultured A549 lung cancer cells. Biochem Biophys Res Commun.
423:448–453. 2012. View Article : Google Scholar
|