Introduction
Musculoskeletal sarcomas are relatively rare
malignancies of bone and soft tissues with unknown etiology,
ill-defined pathogenesis and rapid progression. Sarcomas are indeed
highly aggressive tumors and induce devastating consequences both
in terms of morbidity and mortality (1,2). The
life expectancy of sarcoma patients has not significantly changed
over the past 30 years despite the use of non-specific aggressive
multidrug chemotherapy regimens in addition to surgery and/or
radiation therapy and the recent adoption of biological approaches
based on the identification of molecular targets has not remarkably
modified the outcome (3). It is
likely that current preclinical models are insufficient to describe
the complexity of sarcomagenesis and, accordingly, to identify
effective targets for therapy.
Over the last decades, the discovery of a
tumor-initiating cell population in human acute myeloid leukemia
(4,5) has led to the introduction of the
cancer stem cell (CSC) concept instead of the classical stochastic
model to describe tumor development. This hypothesis postulates
that cancers, like normal adult tissues, contain a minority of
quiescent uncommitted cells that reside at the apex of the
neoplastic cell hierarchy and originate from deregulated endogenous
stem cells or an aberrant reprogramming of committed cells
(6). According to this hypothesis,
CSCs drive cancer development as well as its progression, response
to therapy, and metastatization (7). The pioneering works on leukemia stem
cells prompted further research into a wide variety of solid
malignancies, including breast, brain, colon, pancreas, melanoma,
ovary and prostate cancers, where CSCs have been identified and
characterized (8–14). The isolation assay for CSCs is
based on their ability to grow forming suspended spherical, clonal
colonies. To date, this sphere-forming assay is still the most
widely used method (8,10,15–17).
The capacity to grow as floating spheres has also been exploited
for the isolation of putative CSCs from sarcomas. In particular,
sphere cultures from sarcomas, so-called ‘sarcospheres’, have been
isolated from continuous cell lines (18–22)
or tissue samples (23,24) by the sphere assay. Other
investigators have used indirect assays, based on detoxification
systems that are known to be active in stem cells, like the
Hoechst33342 dye exclusion test (25) or the aldehyde dehydrogenase 1
(ALDH1) activity (26). Recently,
it has been reported that limited oxygen availability, reproducing
the physiological tumor microenvironment, increase the CSCs
fraction and promote a more undifferentiated state (27). Therefore, the control of the
culture condition could be a crucial aspect for the isolation of
CSCs. Here, starting from a number of fresh tissue samples, we
verified the efficacy of the sphere system assay to isolate sphere
cultures from human sarcomas and analysed the stem cell-related
properties of the resulting cultures in terms of gene expression
and in vivo tumorigenic potential. We then exposed
sarcospheres to a hypoxic environment (1% O2), and
compared their behaviour to that of cells grown at 21%
O2.
Materials and methods
Sample collection and sphere system
assay
Fresh biopsies from 49 patients (25 males and 24
females, age 6–85 years) with a histological diagnosis of sarcoma
were prospectically obtained between 2009 and 2012. All samples
were collected after a signed informed consent and following the
institutional ethics committee approval. Tumor and patient data are
summarized in Table I. Samples
were subjected to mechanical and enzymatic digestion to obtain
single cell suspensions. Cells were maintained at 37°C in a
humidified 5% CO2 atmosphere in DMEM:F12 medium with
progesterone (20 nM), putresceine (10 μg/ml), sodium
selenite (30 nM), apo-transferrin (100 μg/ml), and insulin
(25 μg/ml) (Sigma-Aldrich, St. Louis, MO) in low-attachment
flasks (Nunc, Penfield, NY). Fresh human EGF (20 ng/ml) and bFGF
(10 ng/ml) (PeproTech, Rocky Hill, NJ) were added twice a week
until cells started to grow forming floating aggregates. Cultures
were expanded by mechanical dissociation of spheres, followed by
re-plating of cells and residual cell aggregates in complete
medium.
 | Table I.Tumor and patient
characteristics. |
Table I.
Tumor and patient
characteristics.
| Tumor
histotype | Case nos. | Gender | Age range |
|---|
| Osteosarcoma (OS)
(n=11) | 1–11 | Female (n=6) | 8–71 |
| Male (n=5) | 13–71 |
| Chondrosarcoma (CS)
(n=9) | 12–20 | Female (n=6) | 40–78 |
| Male (n=3) | 25–77 |
| Ewing’s sarcoma
(ES) (n=2) | 21–22 | Male (n=2) | 55–58 |
| Liposarcoma (LS)
(n=18) | 23–40 | Female (n=10) | 17–83 |
| Male (n=8) | 40–76 |
| Rhabdomyosarcoma
(RMS) (n=2) | 41–42 | Female (n=1) | 6 |
| Male (n=1) | 10 |
| Other (n=7) | 43–49 | Female (n=1) | 30 |
| Male (n=6) | 51–85 |
Cell lines
In order to obtain parental tumor cell cultures,
single cells obtained from biopsies after enzymatic dissociation
were seeded in Iscove’s modified Dulbecco’s medium (IMDM,
Invitrogen, Carlsbad, CA), plus 20 U/ml penicillin, 100 mg/ml
streptomycin and 10% fetal bovine serum (FBS). The embryonic
carcinoma cell line NT2/D1 was purchased from American Type Culture
Collection (ATCC, Manassas, VA) and cultured in DMEM/high glucose
(Euroclone, Milan, Italy) plus 10% FBS (Hyclone, Logan, UT). Cells
were incubated at 37°C in a humidified 5% CO2
atmosphere. For the hypoxia experiments, the RD rhabdomyosarcoma
(RMS) cell line was purchased from ATCC.
Real-time PCR
Total RNA was extracted from sarcospheres by using
the NucleoSpin RNA II (Qiagen GmbH, Hilden, Germany), and reverse
transcribed. The expression of mRNA for OCT3/4, Nanog, and SOX2 was
evaluated using a Light Cycler instrument (Roche Diagnostics,
Indianapolis, IN) and the Universal Probe Library (Roche Applied
Science). Probes and primers were selected using web-based assay
design software (ProbeFinder http://www.rocheapplied-science.com). Sequences and
primers are reported in Table II.
The results were normalized to GADPH or β-actin according to the
2−ΔΔCT method (28).
 | Table II.Sequences and primers used for
real-time PCR. |
Table II.
Sequences and primers used for
real-time PCR.
| Gene | Accession no. | Forward primer | Reverse primer |
|---|
| OCT3/4 | NM_002701.4 |
CTTCGCAAGCCCTCATTTC |
GAGAAGGCGAAATCCGAAG |
| Nanog | NM_024865.2 |
ATGCCTCACACGGAGACTGT |
AGGGCTGTCCTGAATAAGCA |
| SOX2 | NM_003106.3 |
GGGGGAATGGACCTTGTATAG |
GCAAAGCTCCTACCGTACCA |
| GADPH | NM_002046.3 |
AGCCACATCGCTCAGACAC |
GCCCAATACGACCAAATCC |
| β-actin | NM_001101.2 |
CCAACCGCGAGAAGATGA |
CCAGAGGCGTACAGGGATAG |
Generation and validation of
sarcosphere-derived xenografts
Animal experiments were approved by the Service for
Biotechnology and Animal Welfare of the Istituto Superiore di
Sanità in Rome, Italy and authorized by the Italian Ministry of
Health (Decree no. 217/2010-B). To evaluate the in vivo
tumorigenesis, sarcospheres were dissociated into single cells,
mixed with an equal volume of growth factor-containing Matrigel (BD
Bioscience, Franklin Lakes, NJ) and then 5×104 cells
were injected in the flank of 4-6-week-old female NOG mice (Charles
River Laboratories, Wilmington, MA). When tumor diameter reached a
maximum of 10 mm, mice were sacrificed and the tumor tissue was
then collected for further investigation.
Xenograft analysis
Hematoxylin and eosin (H&E) staining was
performed to analyse xenograft morphology in comparison to the
original tumors. Immunohistochemistry (IHC) was carried out on
paraffin-embedded sections to analyse the expression of desmin,
myogenin and muscle actin. Primary antibodies anti-desmin (cat. no.
D33), anti-myogenin (cat. no. F5D) and anti-muscle actin (cat. no.
HHF35) were from Dako (Dako Denmark A/S, Glostrup, Denmark). The
presence of the RMS-specific fusion transcript PAX3-FKHR was
evaluated by conventional PCR (Invitrogen). Briefly, RNA was
extracted from fresh biopsies using a modified method including
TRIzol reagent (Invitrogen) and the RNeasy Mini kit (Qiagen GmbH)
and reverse transcribed. The positive case control consisted of
alveolar RMS, which was analysed by sequencing to confirm the
specific translocations. Forward and reverse primers were:
5′-CCGACAGCAGCTCTGCCTAC-3′, 5′-TGAACTTGCTGTGTAGGGACAG-3′. The PCR
products were resolved on 3% agarose gel electrophoresis followed
by Gel Star staining (Lonza, Rockland, ME). The sequencing was
performed by Bio-Fab Research (http://www.biofabresearch.it/index2.html, Rome,
Italy). Computer analysis of sequences was performed by basic local
alignment search tool (BLAST) sequence similarity searches using
the National Center of Biotechnology Information Database
(NCBI).
Hypoxia induction
The RD cell line was maintained as monolayer in
complete IMDM. To obtain sarcospheres, RD were then cultured in
anchorage-independent condition as previously described, until the
formation of floating spheres, indicated as RDsph. Hypoxic
atmosphere (1% O2) condition was created by using a
modular incubator chamber (Billups-Rothenberg, Inc., Del Mar,
CA).
Sphere characteristics under hypoxia
To evaluate the sphere formation efficiency, single
cells derived from RDsph were seeded at 500 cells/well in
low attachment 6-well plates at 21 and 1% O2. After 10
days, the total number of spheres was manually counted. To analyse
the sphere proliferation and size, and the expression of stem
cell-related genes, cells derived from RDsph were seeded in
low attachment 6-well plates (20,000 cells/ml) and incubated at 21
and 1% O2 for 10 days. Cell growth was evaluated at 3
and 10 days by a dye exclusion viability assay. The radius of each
tumor spheroid was measured using NIS-Elements Microscope Imaging
Software (Nikon, Tokyo, Japan) and used to calculate the volume (V
= 4/3 π r3). Real-time PCR, as previously described,
evaluated the expression of OCT3/4, Nanog and SOX2 at 21 or 1%
O2.
Statistical analysis
Due to the small number of observations, data were
considered as not normally distributed, and the non-parametric
Mann-Whitney U test was used and p<0.05 was considered
significant. Statistical analysis was performed with the StatView™
5.0.1 software (SAS Institute Inc., Cary, NC). Values were
expressed as means ± SEM.
Results
Establishment of sarcosphere cultures
from human sarcomas
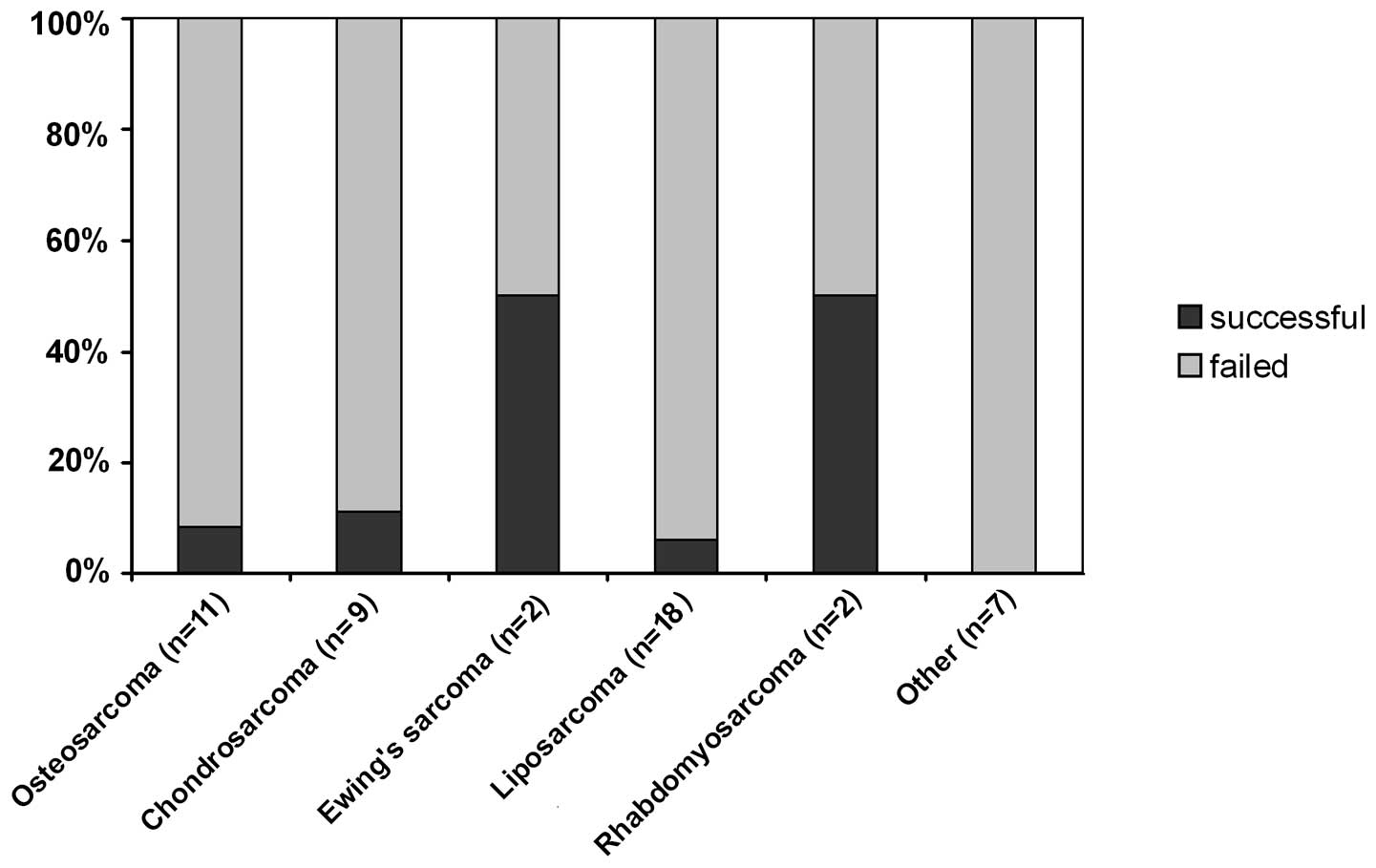
Using a sphere-forming assay, we established sphere
cultures in 5 out of 49 cases (10.2%) (Fig. 1). Details of tumor histotypes,
diagnosis and time needed for sphere formation are summarized in
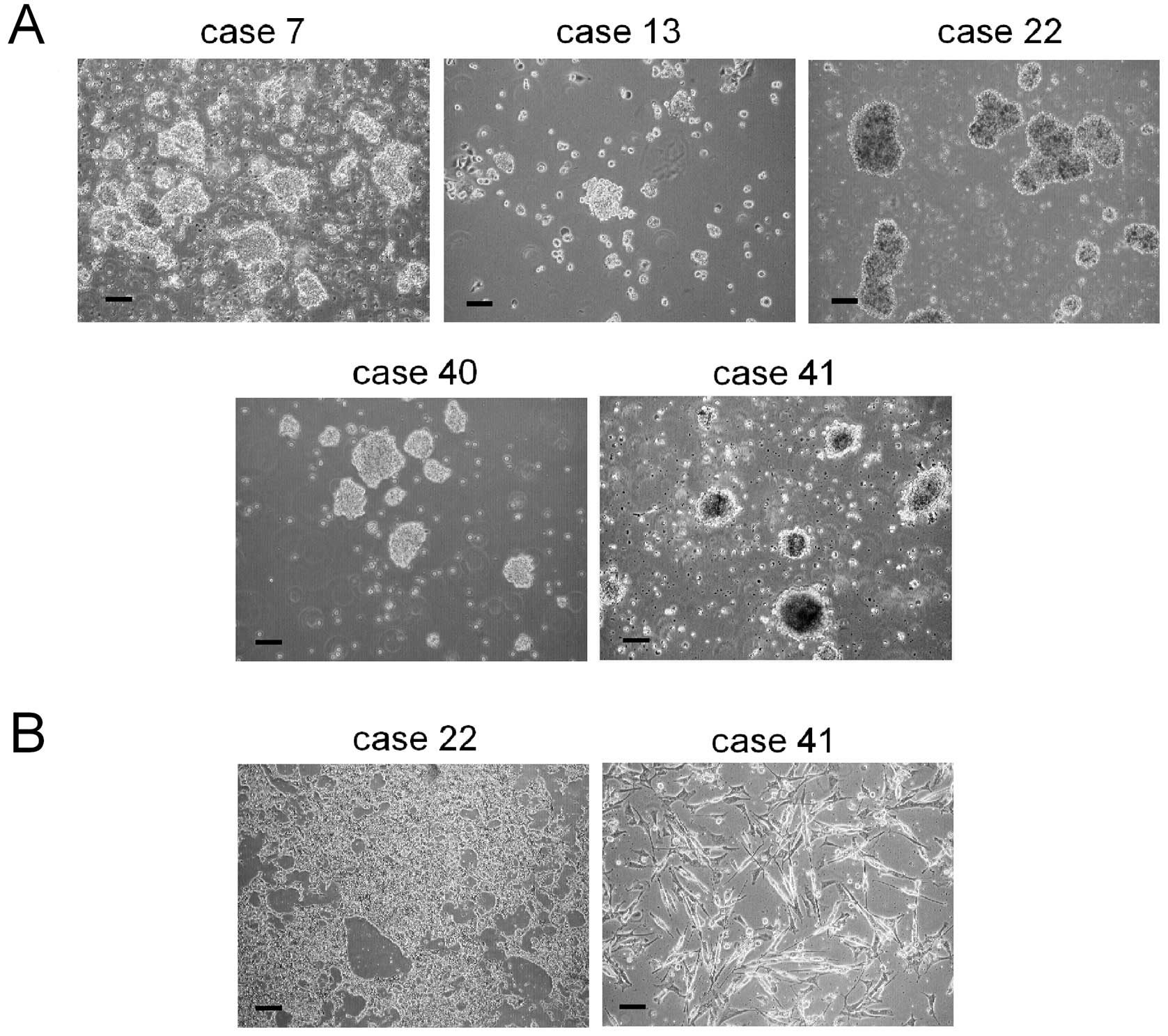
Table III. Established cultures
consisted in floating cell aggregates (Fig. 2A). We also obtained parental tumor
cell cultures from Ewing’s sarcoma (ES) and RMS samples (cases 22
and 41; Fig. 2B).
 | Table III.Established sarcosphere cultures from
human sarcomas. |
Table III.
Established sarcosphere cultures from
human sarcomas.
| Case no. | Diagnosis | Patient
sex/age | Site | Time for sphere
formation (weeks) |
|---|
| 7 | Osteosarcoma,
relapse | M/57 | Ilium | 3 |
| 13 | Chondrosarcoma | F/68 | Acetabulum | 16 |
| 22 | Extraskeletal
Ewing’s sarcoma pPNET | M/55 | Thigh | 2 |
| 40 | Liposarcoma | M/53 | Thigh | 8 |
| 41 | Alveolar
rhabdomyosarcoma, relapse | F/6 | Thigh | 2 |
Expression of stem cell-associated genes
in sarcospheres
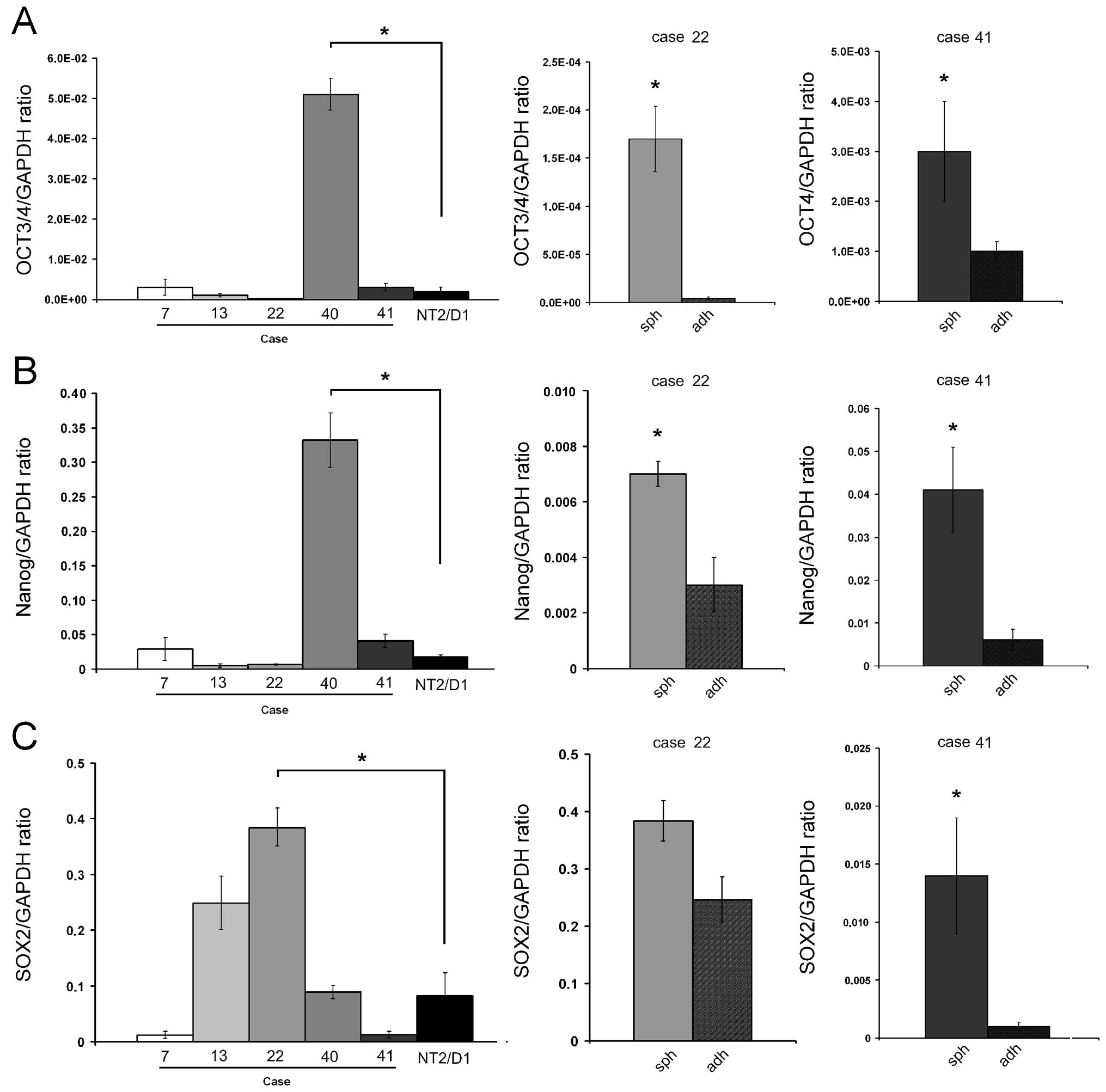
To assess stem cell-related properties, we analysed
the expression of OCT3/4, Nanog and SOX2 by real-time PCR. The
pluripotent embryonal cell line NT2/D1 was used as a positive
control. All sarcosphere cultures expressed consistent mRNA levels
for OCT3/4, Nanog and SOX2, which were significantly higher than
that of the positive control (cases 22 and 40). When compared to
adherent parental cells (cases 22 and 41), sarcospheres expressed
significantly higher levels of mRNA for all of the examined genes
(Fig. 3).
In vivo tumorigenic potential
To provide further evidence that these cells
represented a putative CSC population, single cells derived from
chondrosarcoma (CS) and RMS sarcospheres (cases 13 and 41) were
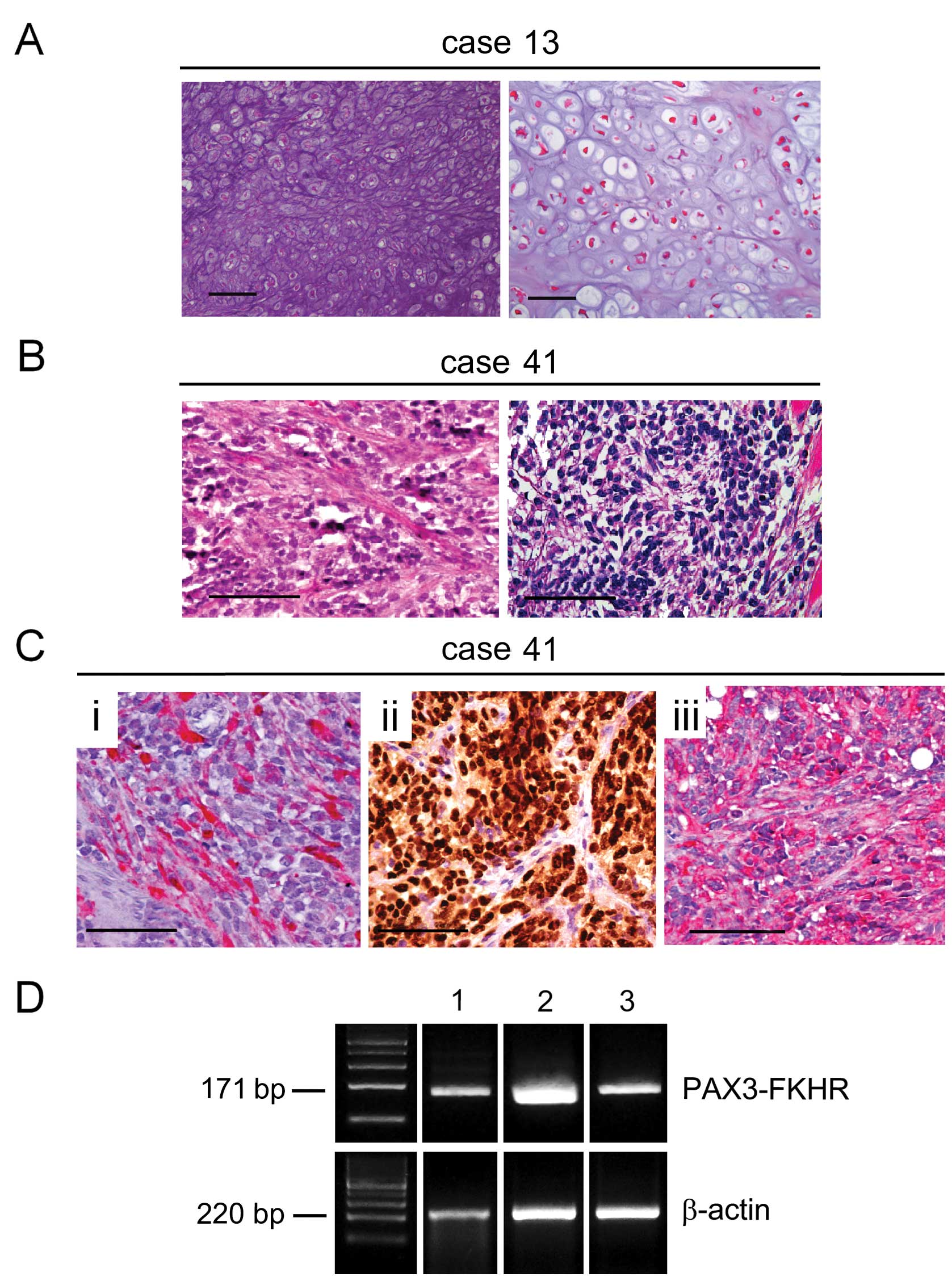
subcutaneously injected into NOG mice. H&E staining of
xenograft obtained from CSCs of CS (case 13) could be related to a
grade 2–3 CS with abundant chondroid matrix and lipid droplets, as
observed in the original biopsy (Fig.
4A). H&E staining for RMS xenograft (case 41) showed small
tumor cells with scant cytoplasm and regular nuclei, arranged in
clumps outlined by hyalinised fibrous septa, resembling the
original tumor pattern (Fig. 4B).
Immunostaining of RMS xenograft for desmin, myogenin and
muscle-actin also confirmed the original tumor phenotype (Fig. 4C). Moreover, RT-PCR analysis
revealed the presence of mRNA transcript for PAX3-FKHR in RMS
sarcospheres, the same alteration detected in the original tumor
(Fig. 4D). All products were
confirmed by DNA sequencing (data not shown).
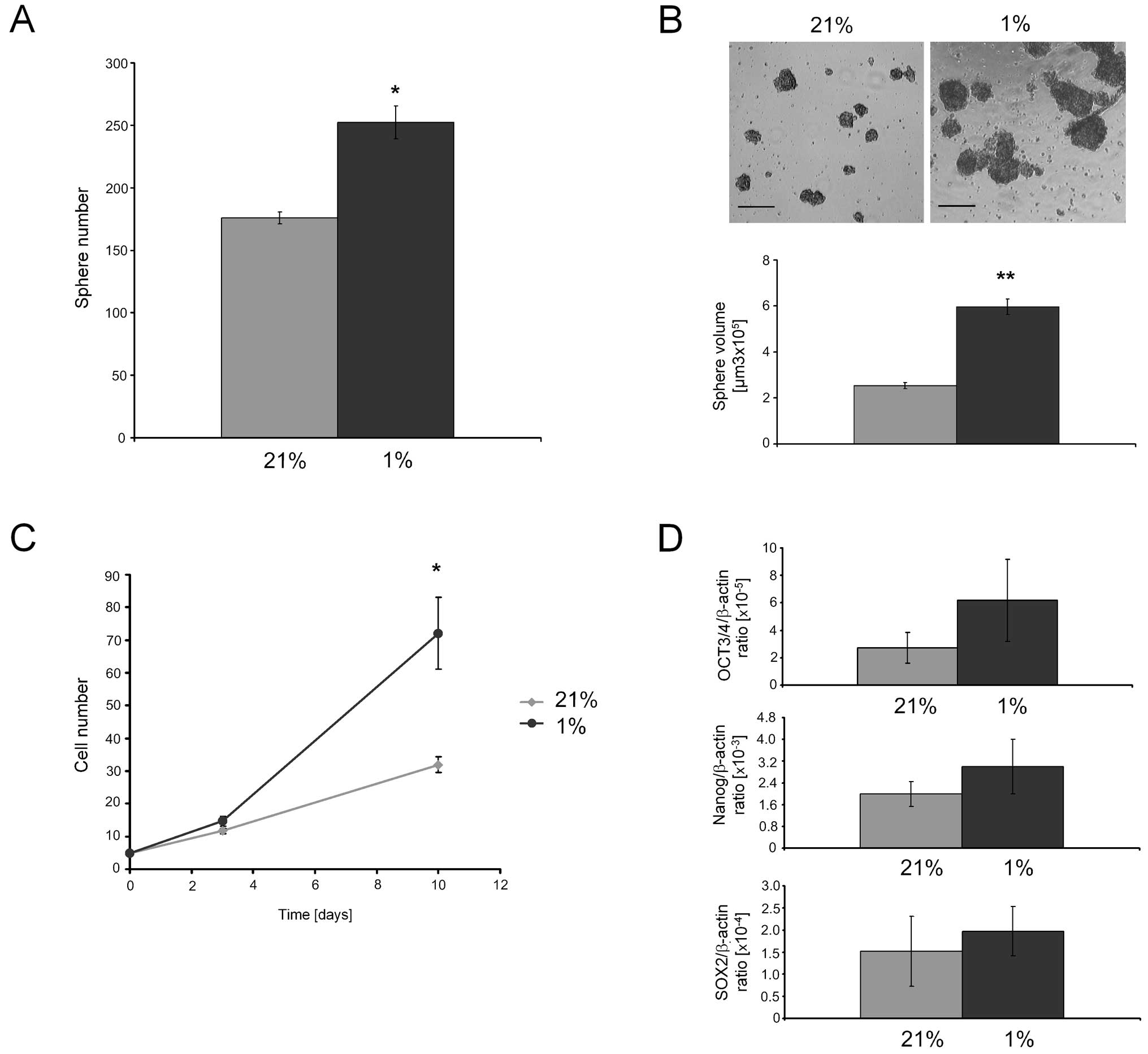
Culture under hypoxia
To evaluate if low oxygen levels could be suitable
for CSC spheres, we cultured spheroids obtained from RD cell line
at 21% (normoxia) or 1% O2 (hypoxia). Under hypoxia, the
number (Fig. 5A) and volume
(Fig. 5B) of RDsph were
significantly higher than in normoxia. Hypoxia also led to a
significant increase of the growth rate (Fig. 5C). Finally, the expression of mRNA
for stemness genes tended to increase under hypoxic conditions
compared to normoxia (Fig.
5D).
Discussion
The identification and characterization of CSCs has
led the way to novel perspectives for cancer treatment. In fact,
this minority subset of the tumor cell population is putatively
responsible for drug resistance, local relapse and metastasis,
crucial features of malignancies that directly impact on patient
survival. Although the clinical relevance of CSCs is yet to be
proved in solid cancers, different anti-leukemic stem cell agents
have already been used in ongoing clinical trials (29). Impressive research work has been
done in this field, giving evidence that the CSCs population
actually represents the fuel of the tumor and its eradication may
be expected to have remarkable clinical perspectives. It has been
shown that CSCs, like normal stem cells (30), are able to grow as free-floating
spheroidal aggregates in non-adherent conditions, as initially
observed in leukemias (4,5) and subsequently in solid tumors.
Thanks to this peculiarity, CSCs of brain and breast cancers, known
as neurospheres and mammospheres, respectively, have been
identified and are now subject of investigation for new targeted
therapies (8,9,31).
Similarly, CSC-enriched sphere cultures have been established and
characterised from other solid tumors (8–15).
Despite these promising advancements, the identification of CSCs in
sarcomas is still fraught with difficulties as a result of some
intrinsic peculiarities of these neoplasms (2,32,33),
including the unclear pathogenesis (34) and the lack of a general consensus
regarding the markers that define the putative mesenchymal
progenitors (35,36). Current data have been largely based
on continuous cell lines (18–21)
rather than on tissue specimens (23,24).
Indirect assays, such as the isolation of the side population, the
ALDH activity (25,26), and the expression of CD133 cell
marker (23,37) have been advocated as alternative
tools to identify sarcoma CSCs, but their reliability as stemness
indicators is still debated (33).
Interestingly, CSCs from Ewing’s sarcoma are involved in
chemoresistance (38). The
conclusive identification of the sarcoma CSC population and its
role in the pathogenic process could have a remarkable health
impact, especially for these tumors where conventional therapies
are often unsuccessful. In our study, we analysed the efficiency of
the sphere system assay for the isolation of CSC-enriched fractions
of sarcomas. For this purpose, we collected a number of human fresh
tissue biopsies, specifically 49 samples of bone and soft tissue
sarcomas. After a culturing period that, interestingly, was shorter
for samples derived from tumor relapses and for more aggressive
neoplasms, we were disappointed that we succeeded in obtaining
spheres only in a minority (10%) of cases, an isolation efficiency
that appears to be lower than that reported in other solid
neoplasms (39). However, it has
already been reported that experimental design variables, such as
the tumor stage of origin, could influence the isolation efficacy
(40). It is also possible that
the culture conditions for mesenchymal tumors are different from
those that have been originally defined as optimal for epithelial
tumors. Another reason for such unsatisfactory yield could be that
standard in vitro methods do not take into sufficient
account some peculiar characteristics of tumor environment. For
example, it is known that CSCs reside in a niche, both in the bone
marrow and in soft tissues, where the oxygen tension is
characteristically lower (3–4%) than that used for standard culture
conditions (21%) (41) and that
low oxygen levels are requested to maintain both normal and cancer
stem cell phenotype (42,43). It is also likely that within the
tumor microenvironment the behaviour of CSCs, including
proliferation and differentiation, is modulated by other stromal
elements, such as immune cells and tumor-associated fibroblasts.
The development of culture systems that more accurately
recapitulates the complexity of sarcoma microenvironment could
improve the ability to isolate, harvest and characterize sarcoma
CSCs for further investigations. To assess the stemness of cells
retrieved from sarcospheres, we analysed the expression of OCT3/4,
Nanog and SOX2, all markers generally associated to CSC (44) and found a significantly higher mRNA
level for all genes in CSCs as compared to parental cells. In a
case of alveolar RMS and another of CS we were also able to
demonstrate that cells retrieved from sarcospheres were able to
form tumors after subcutaneous injection in athymic mice. Notably,
the xenografts recapitulated the histologic pattern of the original
tumors, both in terms of morphology and surface markers and, in the
case of RMS, the transplanted cells originated from the
sarcospheres were also carrying the same, distinctive t(2;13)
translocation. To evaluate if a hypoxic culture environment could
improve the sphere system method efficacy, we also analysed the
effects of hypoxia on an RMS cell model, and found a strong
increase in the number, volume and growth of the spheres. The
control of oxygen levels also resulted in an increase in the
expression of the stem cell markers here considered. Our study
confirms that cells with CSC-like characteristics are present in
human sarcoma tissue samples. The CSCs component can be enriched by
the sphere system method and provide a reliable preclinical model
that can be used for future studies aimed at investigating crucial
aspects of sarcoma biology, such as resistance to treatments and
relapse. It is likely that the reproducibility of the method for
sarcoma CSCs may benefit from the adoption of culture conditions
that take into account the peculiarities of the stem niche and the
mutual interactions of CSCs and reactive elements within the tumor
microenvironment.
Acknowledgements
The authors would like to thank Lucy
Scioscia for the English revision of the manuscript. This study was
supported by grant FIRB: RBAP10447J from the Italian Ministry of
Education, Universities and Research.
References
|
1.
|
Luis ÁM, Aguilar DP and Martín JA:
Multidisciplinary management of soft tissue sarcomas. Clin Transl
Oncol. 12:543–553. 2010.
|
|
2.
|
Burningham Z, Hashibe M, Spector L and
Schiffman JD: The epidemiology of sarcoma. Clin Sarcoma Res.
2:13–16. 2012. View Article : Google Scholar
|
|
3.
|
Keedy VL: Treating metastatic soft-tissue
or bone sarcomas - potential role of ridaforolimus. Onco Targets
Ther. 5:153–160. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4.
|
Lapidot T, Sirard C, Vormoor J, et al: A
cell initiating human acute myeloid leukaemia after transplantation
into SCID mice. Nature. 17:645–648. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
5.
|
Bonnet D and Dick JE: Human acute myeloid
leukemia is organized as a hierarchy that originates from a
primitive hematopoietic cell. Nat Med. 3:730–737. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Reya T, Morrison SJ, Clarke MF and
Weissman IL: Stem cells, cancer, and cancer stem cells. Nature.
414:105–111. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
7.
|
Jordan CT, Guzman ML and Noble M: Cancer
stem cells. N Engl J Med. 355:1253–1261. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8.
|
Al-Hajj M, Wicha MS, Benito-Hernandez A,
Morrison SJ and Clarke MF: Prospective identification of
tumorigenic breast cancer cells. Proc Natl Acad Sci USA.
100:3983–3988. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
9.
|
Singh SK, Clarke ID, Terasaki M, Bonn VE,
Hawkins C, Squire J and Dirks PB: Identification of a cancer stem
cell in human brain tumors. Cancer Res. 63:5821–5828.
2003.PubMed/NCBI
|
|
10.
|
Ricci-Vitiani L, Lombardi DG, Pilozzi E,
Biffoni M, Todaro M, Peschle C and De Maria R: Identification and
expansion of human colon-cancer-initiating cells. Nature.
445:111–115. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11.
|
Li C, Heidt DG, Dalerba P, et al:
Identification of pancreatic cancer stem cells. Cancer Res.
67:1030–1037. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12.
|
Schatton T, Murphy GF, Frank NY, et al:
Identification of cells initiating human melanomas. Nature.
451:345–349. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13.
|
Zhang S, Balch C, Chan MW, et al:
Identification and characterization of ovarian cancer-initiating
cells from primary human tumors. Cancer Res. 68:4311–4320. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
14.
|
Lang Sh, Frame F and Collins A: Prostate
cancer stem cells. J Pathol. 217:299–306. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15.
|
Ponti D, Costa A, Zaffaroni N, et al:
Isolation and in vitro propagation of tumorigenic breast cancer
cells with stem/progenitor cell properties. Cancer Res.
65:5506–5511. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16.
|
Deleyrolle LP, Ericksson G, Morrison BJ,
et al: Determination of somatic and cancer stem cell self-renewing
symmetric division rate using sphere assays. PLoS One.
6:e158442011. View Article : Google Scholar : PubMed/NCBI
|
|
17.
|
Liu T, Cheng W, Lai D, Huang Y and Guo L:
Characterization of primary ovarian cancer cells in different
culture systems. Oncol Rep. 23:1277–1284. 2010.PubMed/NCBI
|
|
18.
|
Gibbs CP, Kukekov VG, Reith JD, et al:
Stem-like cells in bone sarcomas: implications for tumorigenesis.
Neoplasia. 7:967–976. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
19.
|
Fujii H, Honoki K, Tsujiuchi T, Kido A,
Yoshitani K and Takakura Y: Sphere-forming stem-like cell
populations with drug resistance in human sarcoma cell lines. Int J
Oncol. 34:1381–1386. 2009.PubMed/NCBI
|
|
20.
|
Wang L, Park P and Lin CY:
Characterization of stem cell attributes in human osteosarcoma cell
lines. Cancer Biol Ther. 8:543–552. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21.
|
Rainusso N, Man TK, Lau CC, et al:
Identification and gene expression profiling of tumor-initiating
cells isolated from human osteosarcoma cell lines in an orthotopic
mouse model. Cancer Biol Ther. 12:278–287. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22.
|
Basu-Roy U, Seo E, Ramanathapuram L, et
al: Sox2 maintains self renewal of tumor-initiating cells in
osteosarcomas. Oncogene. 31:2270–2282. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23.
|
Suvà ML, Riggi N, Stehle JC, et al:
Identification of cancer stem cells in Ewing’s sarcoma. Cancer Res.
69:1776–1781. 2009.
|
|
24.
|
Sette G, Salvati V, Memeo L, et al: EGFR
inhibition abrogates leiomyosarcoma cell chemoresistance through
inactivation of survival pathways and impairment of CSC potential.
PLoS One. 7:e468912012. View Article : Google Scholar : PubMed/NCBI
|
|
25.
|
Murase M, Kano M, Tsukahara T, et al: Side
population cells have the characteristics of cancer stem-like
cells/cancer-initiating cells in bone sarcomas. Br J Cancer.
101:1425–1432. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26.
|
Lohberger B, Rinner B, Stuendl N, et al:
Aldehyde dehydrogenase 1, a potential marker for cancer stem cells
in human sarcoma. PLoS One. 7:e436642012. View Article : Google Scholar : PubMed/NCBI
|
|
27.
|
Heddleston JM, Li Z, Lathia JD, Bao S,
Hjelmeland AB and Rich JN: Hypoxia inducible factors in cancer stem
cells. Br J Cancer. 102:789–795. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28.
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
29.
|
Felipe Rico J, Hassane DC and Guzman ML:
Acute myelogenous leukemia stem cells: from bench to bedside.
Cancer Lett. Jun 17–2012.(Epub ahead of print).
|
|
30.
|
Reynolds BA and Weiss S: Generation of
neurons and astrocytes from isolated cells of the adult mammalian
central nervous system. Science. 255:1707–1710. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
31.
|
Hallett RM, Kondratyev MK, Giacomelli AO,
Nixon AM, Girgis-Gabardo A, Ilieva D and Hassell JA: Small molecule
antagonists of the Wnt/β-catenin signaling pathway target breast
tumor-initiating cells in a Her2/Neu mouse model of breast cancer.
PLoS One. 7:e339762012.
|
|
32.
|
Gibbs CP Jr, Levings PP and Ghivizzani SC:
Evidence for the osteosarcoma stem cell. Curr Orthop Pract.
22:322–326. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
33.
|
Trucco M and Loeb D: Sarcoma stem cells:
do we know what we are looking for? Sarcoma. 2012 291705:May
9–2012.(Epub ahead of print). View Article : Google Scholar
|
|
34.
|
Galli R, Borello U, Gritti A, et al:
Skeletal myogenic potential of human and mouse neural stem cells.
Nat Neurosci. 3:986–991. 2000. View
Article : Google Scholar : PubMed/NCBI
|
|
35.
|
Toguchida J and Nakayama T: Molecular
genetics of sarcomas: applications to diagnoses and therapy. Cancer
Sci. 100:1573–1580. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
36.
|
Tolar J, Nauta AJ, Osborn MJ, et al:
Sarcoma derived from cultured mesenchymal stem cells. Stem Cells.
25:371–379. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
37.
|
Tirino V, Desiderio V, Paino F, et al:
Human primary bone sarcomas contain CD133+ cancer stem
cells displaying high tumorigenicity in vivo. FASEB J.
25:2022–2030. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
38.
|
Jiang X, Gwye Y, Russell D, et al: CD133
expression in chemo-resistant Ewing sarcoma cells. BMC Cancer.
10:116–127. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
39.
|
Hemmati HD, Nakano I, Lazareff JA,
Masterman-Smith M, Geschwind DH, Bronner-Fraser M and Kornblum HI:
Cancerous stem cells can arise from pediatric brain tumors. Proc
Natl Acad Sci USA. 100:15178–15183. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
40.
|
Gupta PB, Chaffer CL and Weinberg RA:
Cancer stem cells: mirage or reality? Nat Med. 5:1010–1012. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
41.
|
Mohyeldin A, Garzón-Muvdi T and
Quiñones-Hinojosa A: Oxygen in stem cell biology: a critical
component of the stem cell niche. Cell Stem Cell. 7:150–161. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
42.
|
Rosová I, Dao M, Capoccia B, Link D and
Nolta JA: Hypoxic preconditioning results in increased motility and
improved therapeutic potential of human mesenchymal stem cells.
Stem Cells. 26:2173–2182. 2008.PubMed/NCBI
|
|
43.
|
McCord AM, Jamal M, Shankavaram UT, Lang
FF, Camphausen K and Tofilon PJ: Physiologic oxygen concentration
enhances the stem-like properties of CD133+ human
glioblastoma cells in vitro. Mol Cancer Res. 7:489–497. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
44.
|
Okita K, Ichisaka T and Yamanaka S:
Generation of germline-competent induced pluripotent stem cells.
Nature. 448:313–317. 2008. View Article : Google Scholar : PubMed/NCBI
|