Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the fifth
leading cause of cancer death in Japan (1). This disease usually shows a poor
prognosis because of the rapid progression and development of
distant metastasis by the time of diagnosis (2,3). In
addition, this tumor is resistant to conventional chemotherapy and
radiation therapy. Effective technologies such as positron emission
tomography (PET) and endoscopic ultrasound-guided fine needle
aspiration-biopsy (EUS-FNAB) can improve the detection of PDAC.
However, the availability of these methods is restricted due to
their high costs or difficulty of technique. On the other hand, the
serum levels of carcinoembryonic antigen (CEA) and carbohydrate
antigen 19-9 (CA19-9) have been used as useful markers for the
presence of PDAC (4). However,
their sensitivity and specificity for the early detection of PDAC
is not sufficient. Therefore, an easy and accurate method for the
detection of PDAC is required to improve its poor prognosis.
MicroRNAs (miRNAs) are small non-coding RNAs
composed of 18–25 nucleotides that regulate the translation of
specific genes through binding to the 3′-untranslated regions
(UTRs) of their target mRNAs. Since first discovered by Lee et
al in 1993 (5), accumulating
evidence has revealed that miRNAs play important roles in the
generation or development of various cancers. In PDAC tissues, a
number of studies have identified multiple aberrantly expressed
miRNAs, including miR-21 (6–9),
miR-155 (7–9), miR-146a (10), miR-196a (11), miR-196b (12), miR-200a/b/c (13–15),
miR-221 and miR-222 (7,16). Interestingly, miRNAs are stably
detectable in the plasma/serum since they are protected from RNase
activity (17–20) by microvesicles like exosomes
(21–23), forming protein complexes with Ago2
(24), and lipoprotein complexes
(25). The stability of miRNAs in
body fluids suggests that circulating miRNAs could be useful
diagnostic markers in various cancers. Recently, it has been
reported that increased or decreased expression of circulating
miRNAs, such as miR-18a, miR-21, miR-155, miR-196a, and miR-200a/b,
miR-210, miR-221, are correlated with the development of PDAC
(20,26–29)
(Table I). Among these miRNAs, the
correlation between tumor progression and the expression of miR-21
has been well documented in PDAC patients (26,30–37).
 | Table IExperimentally validated circulating
miRNA in PDAC. |
Table I
Experimentally validated circulating
miRNA in PDAC.
| miRNAs | Authors/(Ref.) | Expression in
PDAC |
|---|
| miR-21 | Wang, et al
(26), Liu, et al (34), Ali, et al (35), Liu, et al (36), Kong, et al (37) | Upregulation |
| miR-210 | Wang, et al
(26), Ho, et al (28), Liu, et al (36) | Upregulation |
| miR-155 | Wang, et al
(26), Kon, et al (37) | Upregulation |
| miR-196a | Wang, et al
(26), Liu, et al (36), Kong, et al (37) | Upregulation |
| miR-221 | Kawaguchi, et
al (29), Ali, et al
(35) | Upregulation |
| miR-200a, 200b | Li, et al
(14) | Upregulation |
| miR-18a | Morimura, et
al (27) | Upregulation |
| miR-181a,181b | Liu, et al
(36) | Upregulation |
| miR-20a, 24, 25,
99a, 185, 191 | Liu, et al
(34) | Upregulation |
| miR-1290, 24, 134,
378, 146a, 484, -628-3p, 1825 | Li, et al
(53) | Upregulation |
| let-7d,
miR-146a | Ali, et al
(35) | Downregulation |
We previously performed comprehensive analysis of
miRNA expression profiles in PDAC and non-invasive intraductal
papillary-mucinous neoplasm (IPMN) (GSE29542) to explore the miRNAs
involved in the invasive growth of PDAC. We identified miR-126 and
miR-197 as differentially expressed miRNAs in PDAC and revealed
that these miRNAs contributed to the invasion of PDAC cells
(38,39). We also found miR-483-3p as one of
the upregulated miRNAs along with miR-21 and-197 in PDAC by
re-reviewing this comprehensive analysis. miR-483-3p, which is
located within intron 2 of the IGF2 locus, is overexpressed in a
variety of tumors such as malignant mesothelioma, Wilms’ tumor
tissues, colon, breast and liver cancers (40,41).
In addition, the expression of miR-483-3p is strongly enhanced in
PDAC tissues and suppresses the expression of DPC4/Smad4 (42). However, little is known about the
plasma expression of miR-483-3p and its association with the
clinicopathological features in PDAC patients. Therefore, we tested
whether evaluation of the plasma miR-483-3p level would be useful
for detecting PDAC, and investigated its possible use as a
biomarker by comparing it with plasma miR-21 and conventional tumor
marker (CEA and CA19-9) levels.
Materials and methods
Microdissection from tissue samples
Ten PDAC and 13 IPMN tissue samples were obtained
from patients who underwent surgical resection in Miyagi Cancer
Center Hospital from 2010 to 2012. Histologic diagnosis of each
sample was performed by two pathologists who were not informed
about the present study. Histologically, cancer duct cells and
non-tumor cells (mixed with acinar cells, inflammatory duct cells
and stromal cells) from PDAC tissues (n=10), and non-invasive IPMN
cells (adenoma and carcinoma in situ, n=13) were
microdissected and subjected to RNA extraction. These
paraffin-embedded tissues were cut into 10-μm sections and ~10
sequential regions from the same paraffin block were microdissected
using a Leica CIR MIC system (Leica Microsystems, Wetzkar,
Germany). Total RNA including miRNA was extracted using the Recover
All Total Nucleic Acid Isolation kit (Ambion, Austin, TX, USA),
according to the manufacturer’s instructions.
Patients and blood samples
Between April 2011 and March 2013, the plasma
samples of 32 PDAC patients, 30 healthy controls (HC) and 12 IPMN
patients were collected at Miyagi Cancer Center. Patient
characteristics with respect to age, sex, and stages of disease are
described in Table II. The stage
of PDAC was assessed according to the Union for International
Cancer Control (UICC) Classification (43). All patients were pathologically
diagnosed as having PDAC using surgical (n=8) or biopsy specimens
[EUS-FNA (n=20), endoscopic transpapillary biopsy (n=3) and biopsy
from duodenal invasion (n=1)]. As a control, plasma samples were
collected from 30 HC. HC were 22 medical staff members and 8
hospitalized patients with benign disease such as asymptomatic
cholecystolithiasis or choledocholithiasis. They underwent medical
examinations and did not have any pancreatic disease. All IPMN
patients were the branch duct type of IPMNs (BD-IPMN). In this
study, the definition of BD- IPMN was based on imaging as a
grapelike multilocular cystic lesion without mural nodules
communicating with the MPD by computed tomography (CT), magnetic
resonance cholangiopancreatography (MRCP), endoscopic
ultrasonography (EUS), and/or endoscopic retrograde
cholangiopancreatography (ERCP).
 | Table IIThe characteristics of PDAC patients,
IPMN patients and healthy controls (HC). |
Table II
The characteristics of PDAC patients,
IPMN patients and healthy controls (HC).
| PDAC (N=32) | IPMN (N=12) | HC (N=30) |
|---|
| Age |
| Mean ± SD
(range) | 70.6±8.7
(48–89) | 74.6±8.8
(61–89) | 44.5±19.1
(21–85) |
| Gender |
| Male | 22 | 6 | 11 |
| Female | 10 | 6 | 19 |
| Stagea | I/II/III/IV | 1/8/10/13 | |
Blood plasma samples and RNA
extraction
Blood samples were collected from patients and
controls in sodium EDTA-Na tubes and were immediately centrifuged
at 3500 rpm for 10 min at room temperature. Then the plasma
supernatant was collected and stored at −80°C until further
analysis. Total RNA containing small RNA was extracted from 300 μl
plasma samples using a mirVana PARIS kit (Ambion), undiluted into
100 μl of preheated (95°C) elution solution according to the
manufacturer’s instructions. To normalize sample-to-sample
variation in the RNA isolation step, 2 μl of synthetic C.
elegans miR-39, which lacks sequence homology to human miRNAs,
was added to each denatured sample.
The detection of miRNAs
The amount of miRNAs was quantified in duplicate via
quantitative RT-PCR using the human TaqMan MicroRNA Assay kit
(Applied Biosystems, Foster City, CA, USA). The reverse
transcription reaction was carried out with a TaqMan MicroRNA
Reverse Transcription kit (Applied Biosystems) in 15 μl containing
5 μl of RNA extract, 0.15 μl of 100 mM dNTPs, 1.5 μl of 10× reverse
transcriptase buffer, 1 μl of Multiscribe reverse transcriptase (50
U/μl), 0.19 μl of RNase inhibitor (20 U/μl), 3 μl of gene-specific
primer, and 4.16 μl of Nuclease-free water. For the synthesis of
cDNA, reaction mixtures were incubated at 16°C for 30 min, at 42°C
for 30 min, and at 85°C for 5 min, and then held at 4°C. Next, 15
μl of cDNA solution was amplified with 25 μl of TaqMan 2× Universal
PCR Master Mix with no AmpErase UNG (Applied Biosystems), 2.5 μl of
gene-specific primers/ probe, and 7.5 μl of nuclease-free water in
a final volume of 50 μl. Quantitative RT-PCR was run on a
LightCycler® 480 real-time PCR system (Roche), and
reaction mixtures were incubated at 95°C for 10 min, followed by 40
cycles of 95°C for 15 sec, and 60°C for 1 min. Cycle threshold (Ct)
values were calculated with the LightCycler 480 for Software
Version 1.5 (Roche). The relative expression of the mature miRNAs
was calculated using the comparative CT (2−2ΔΔCt) method
with miR-16 and RNU6B for plasma and tissue samples, respectively,
as the endogenous control to normalize the data (44,45).
Statistical analysis
Student’s t-test or Mann-Whitney U test was used to
evaluate differences in the miRNA expression between PDAC or IPMN
cases and controls. Receiver operating characteristic (ROC) curves
were constructed and the area under the curve (AUC) was calculated
to evaluate the sensitivity and specificity for predicting cases
and controls based on the expression of each individual miRNA and
their combinations. Fisher’s exact test or Pearson’s χ2
test was used to determine if there was a significant association
between the clinicopathological factors and the relative plasma
expression levels of miRNAs. Survival rate was estimated by the
Kaplan-Meier method. The difference in survival rates between the
groups was tested for significance using the log-rank test. The
overall survival was defined as the period from the date of
diagnosis for PDAC to the date of death or last follow-up. All
tests of statistical significance were two-sided. A P-value of
<0.05 was considered statistically significant. All statistical
analyses were done using Excel 2010 software.
Ethics
The Institutional Review Board of the Miyagi Cancer
Center (MCC) approved this study protocol, and written informed
consent was obtained from each patient (permit MCC-24-38).
Results
The expression of miR-483-3p as well as
miR-21 is enhanced in PDAC cells compared to IPMN and normal
pancreatic cells
To validate the results of the previous
comprehensive analysis, we first examined the miR-483-3p and miR-21
expression levels in a new series of PDAC and IPMN tissues. The
results for these miRNAs, normalized with RNU6B are shown in
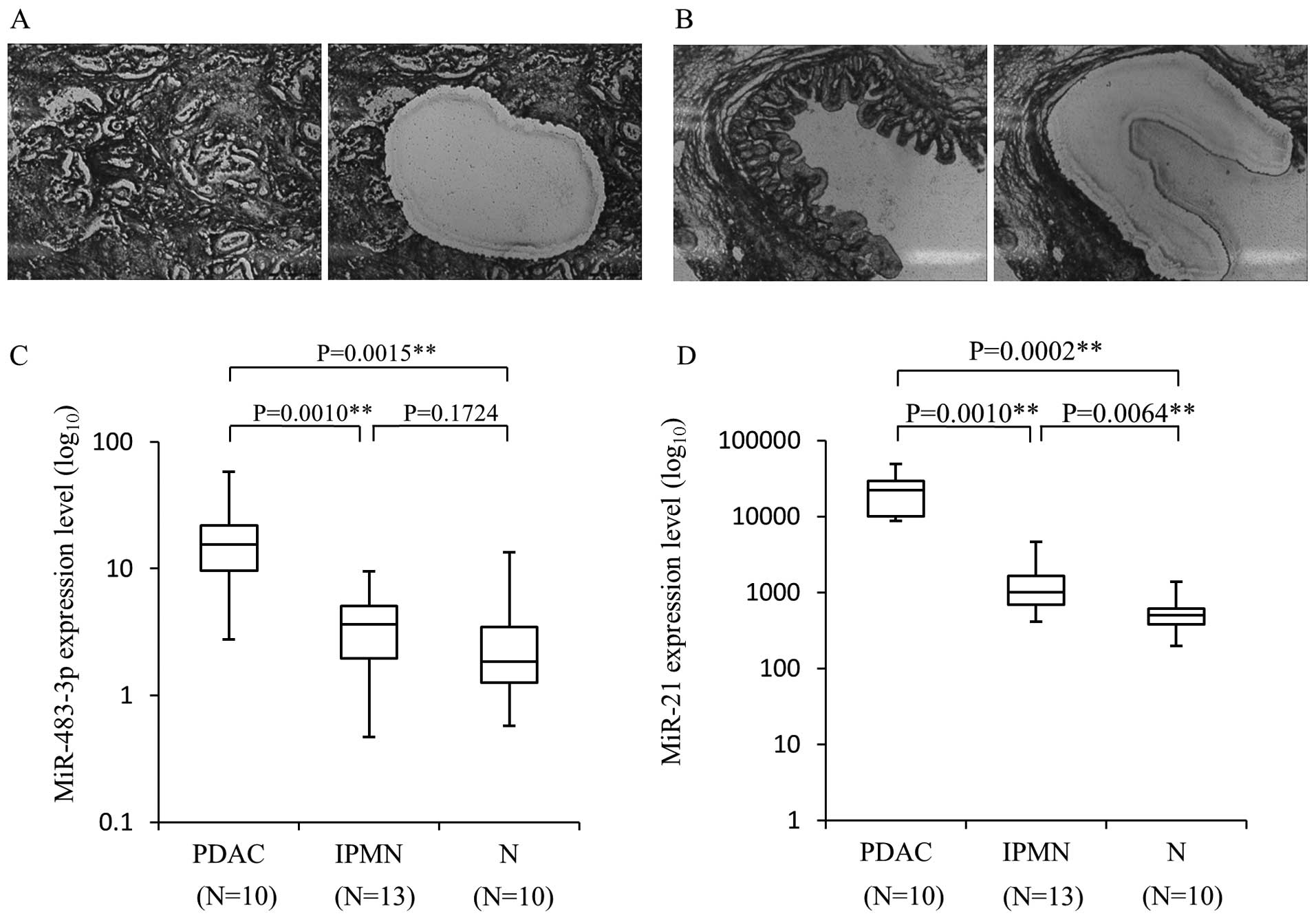
Fig. 1. In microdissected PDAC
cells, the mean level of miR-483-3p expression was 19.36±5.23
[miR-483-3p/RNU6B, mean ± standard error (SE)], and this expression
level was significantly higher than that of non-tumor cells
(3.26±1.22, P=0.0015). The miR-483-3p expression in PDAC cells was
also significantly higher than in IPMN lesions (4.07±0.75,
P=0.0010). Although there was no statistically significant
difference between IPMN cells and non-tumor cells, the miR-483-3p
level tended to be slightly higher in the IPMN samples than in the
normal tissue, suggesting that the increased expression of
miR-483-3p occurs in the transition from benign to malignant.
Similarly, the expression of miR-21 was significantly higher in
PDAC [23150.61±4493.76 (miR-21/RNU6B, mean ± SE)] than IPMN
(1413.05±319.18, P=0.0001) and non-tumor cells (580.45±109.06,
P=0.0002). On the other hand, miR-21 expression was significantly
higher in IPMN compared to non-tumor cells, suggesting that miR-21
contributes to an early step of pancreatic tumorigenesis.
Plasma expression level of miR-483-3p and
-21 is increased in PDAC patient
Next, we hypothesized that higher miR-483-3p
expression in primary carcinoma cells would influence the plasma
levels of miR-483-3p in PDAC patients. To determine the quantities
of plasma miRNAs, we used cell-miR-39 as internal control miRNAs,
and miR-16 was used for the normalization. Plasma miR-483-3p
expression was calculated from the comparative Ct method by
quantitative RT-PCR. Using this assay, circulating miR-483-3p was
detectable in all plasma samples from 32 PDAC patients, 30 HC and
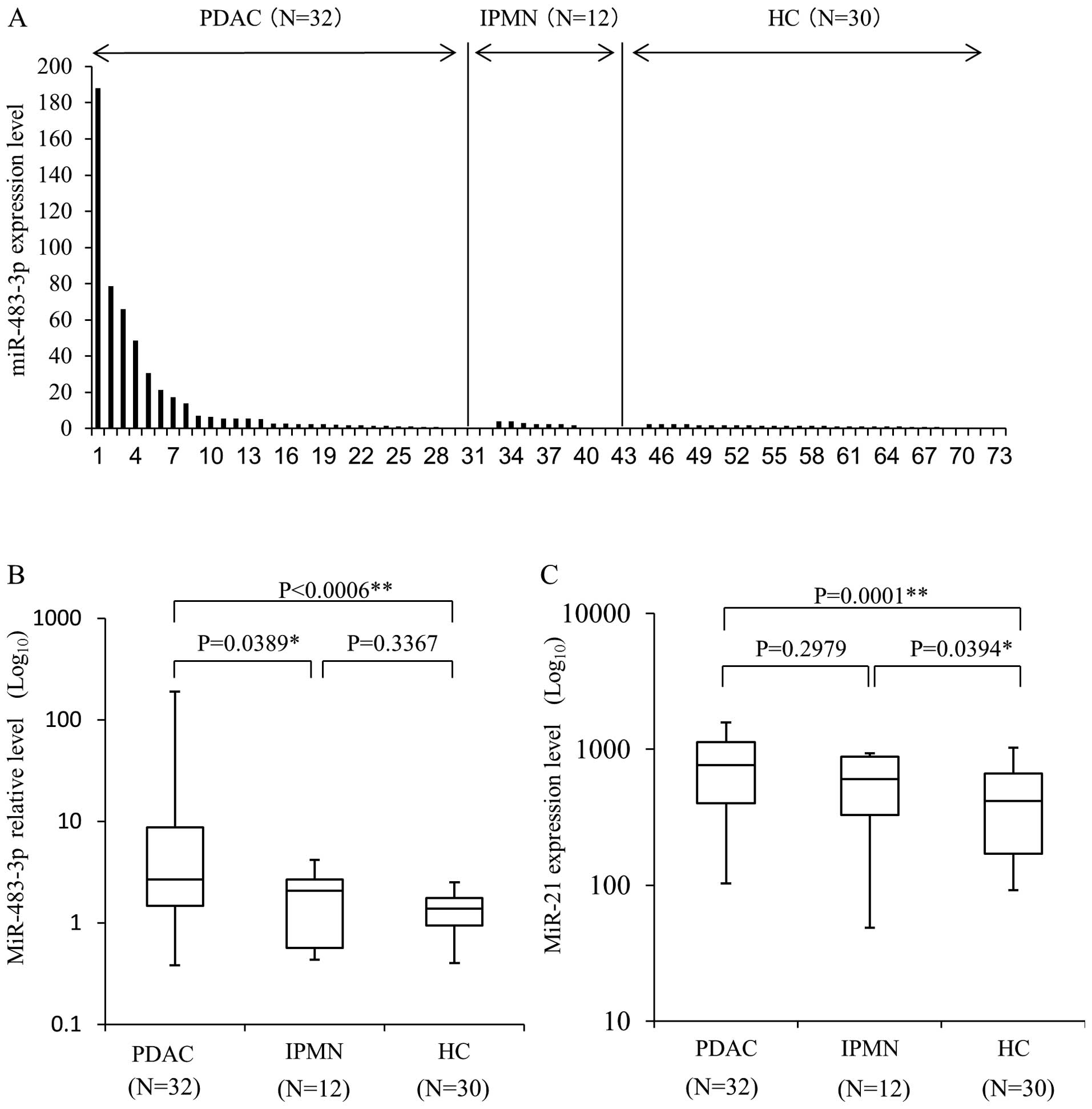
12 IPMN patients (Fig. 2A). The
mean level of plasma miR-483-3p expression in the PDAC patients
[16.50±6.49 (miR-483-3p/miR-16, mean ± SE)] was significantly
higher than in HC (1.37±0.11, P=0.0006) and IPMN patients
(1.90±0.39, P=0.0389) (Fig. 2B).
We also evaluated the expression level of plasma miR-21, which was
previously shown to be overexpressed in the plasma and tissue of
PDAC (26,36–43).
The mean expression level of plasma miR-21 (Fig. 2C) was significantly higher in the
PDAC group [765.25±64.6 (miR-21/miR-16 ± SE)] than HC (414.54±44.7,
P=0.0001). The plasma expression level of miR-21 in IPMN patients
was also higher than in HC (P=0.0394), and no differences were seen
between PDAC and IPMN (P=0.2979). While no difference was observed
in the plasma miR-21 expression level between PDAC and IPMN
patients, miR-483-3p expression in the plasma of PDAC patients was
significantly higher than in that of IPMN patients. This result
indicates that plasma miR-483-3p measurement has the potential to
distinguish PDAC not only from healthy individuals, but also from
the patient of IPMN.
The utility of measuring miR-483-3p and
-21 in plasma for discriminating PDAC from healthy controls and/or
IPMN patients
To apply the expression value of plasma miRNAs for
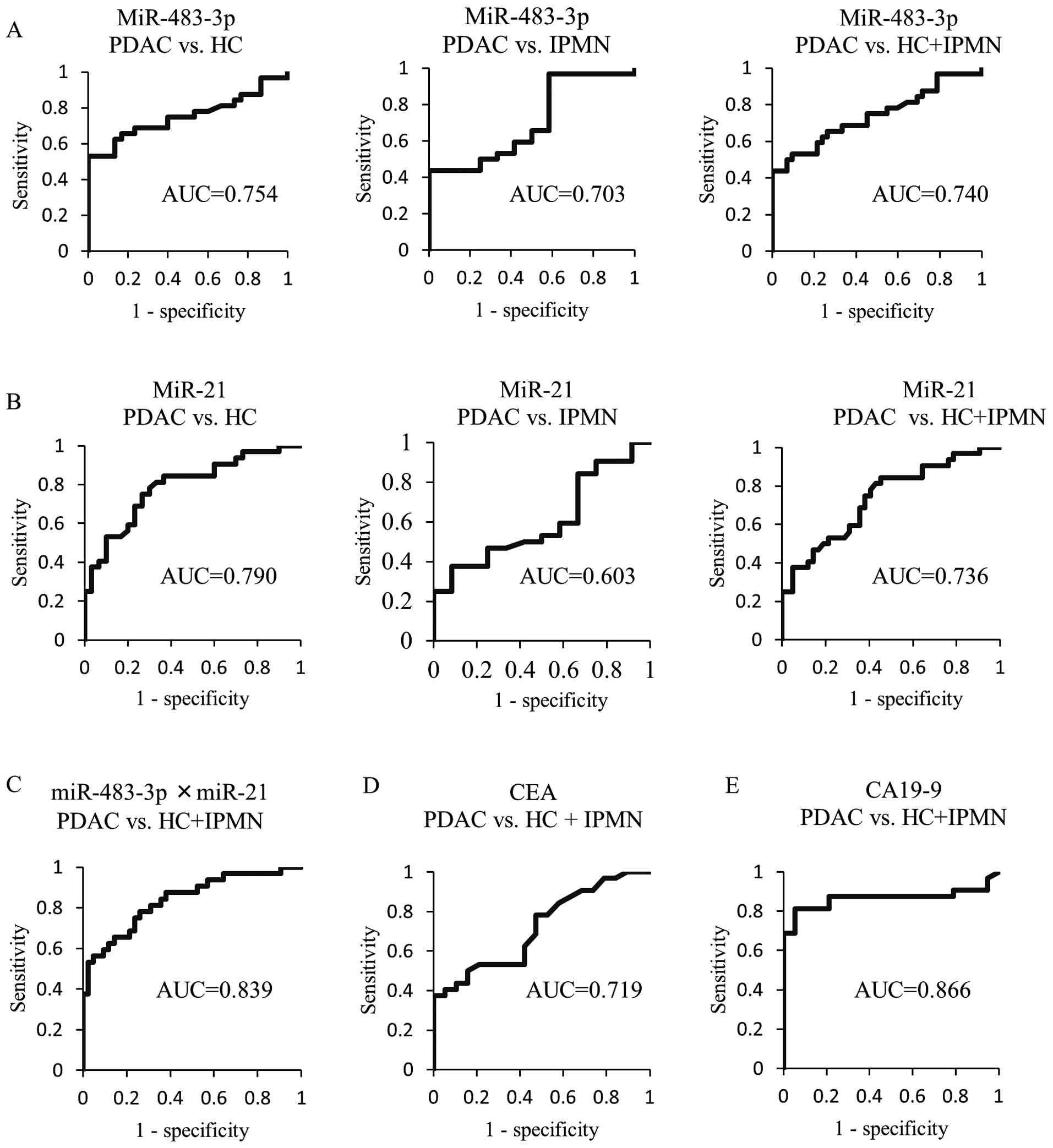
the diagnosis of PDAC, we then analyzed the ROC curves. The ROC
curves for differentiating between PDAC and HC, and/or IPMN based
on the expression level of miRNAs (miR-483-3p or miR-21) are shown
in Fig. 3. The AUC of miR-483-3p
(0.754) was slightly lower than that of miR-21 (0.790) for
differentiating PDAC from HC. On the other hand, for discriminating
PDAC from IPMN patients, the AUC of miR-483-3p (0.703) was higher
than that of miR-21 (0.603), In addition, the AUC value based on
the expression level of plasma miR-483-3p (0.740) and miR-21
(0.736) to distinguish PDAC patients from non-cancer individuals
(HC and IPMN) was similar. Consequently, we compared the ability of
measuring the plasma miR-483-3p and miR-21 to predict PDAC with
that of the conventional tumor markers CEA and CA19-9. The
respective AUC values of plasma miR-483-3p and miR-21 were higher
than that of CEA (0.719), but lower than that of CA19-9 (0.866).
Interestingly, the AUC (0.839) from the ROC curves for the
combination of miR-483-3p and miR-21 was similar to that of CA19-9
for discriminating patients with PDAC from non-cancer
individuals.
The correlation between plasma miRNAs and
clinicopathological factors
Finally, we evaluated the correlation between the
plasma level of miRNAs and clinicopathological factors in 32 PDAC
patients (Table III). We judged
the miR-483-3p level as ‘high expression’ when it was higher than a
cut-off value of 5. There was no significant relationship between
the plasma miR-483-3p level and clinical factors with overall
survival. The plasma miR-21 expression level was associated with
advanced stage (P=0.023), metastasis to lymph node (P=0.007) and
liver (P<0.001) when the cut-off value was defined as 850, as
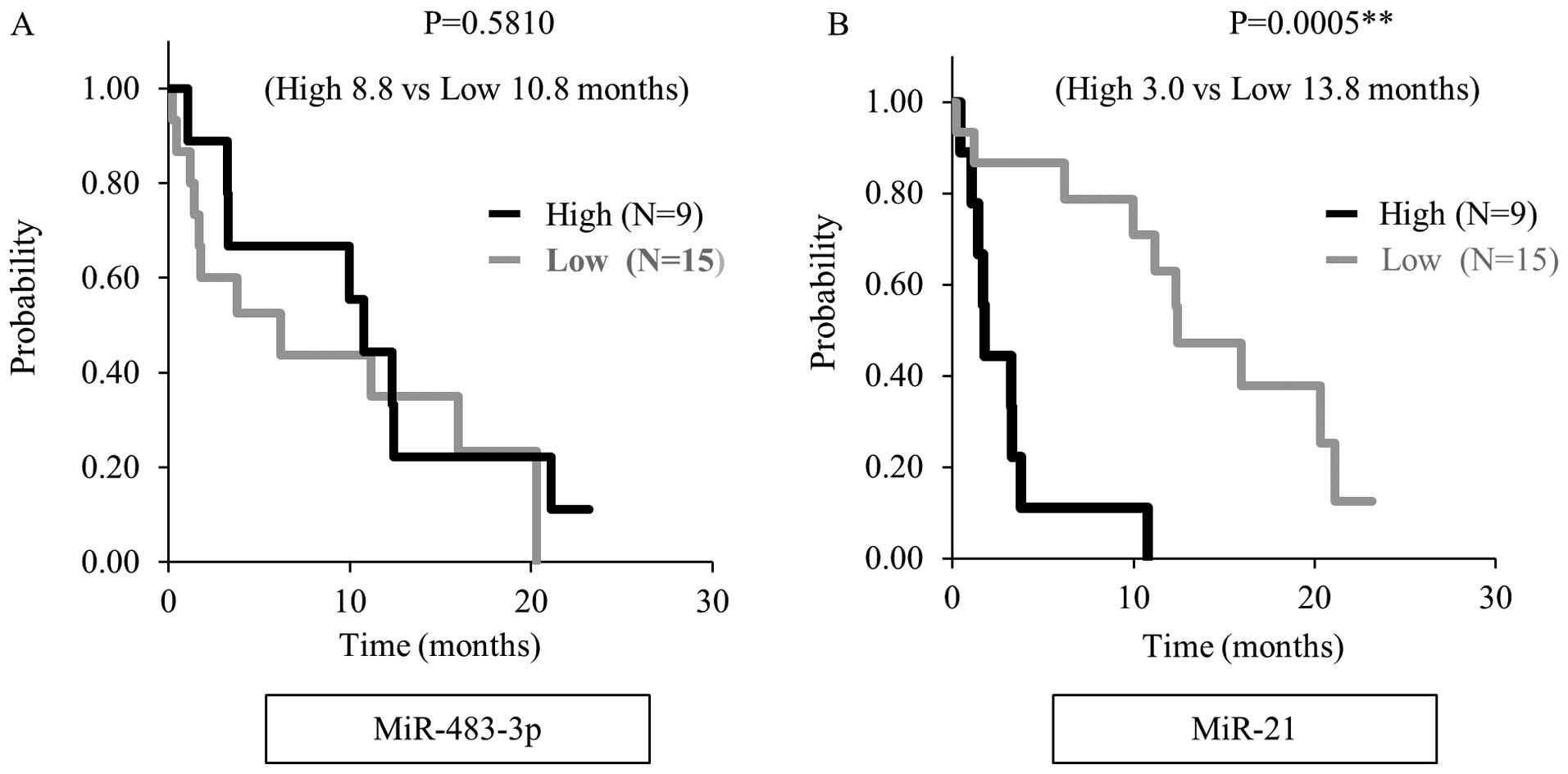
shown in Table IV. Moreover,
overall survival was significant shorter in the high miR-21
expression group than in the low expression group when we analyzed
by the Kaplan-Meier method in 24 unresectable PDAC patients, (3.0
months vs 13.8 months, P<0.001) (Fig. 4).
 | Table IIIThe association of miR-483-3p
expression level with clinicopathological factors in PDAC. |
Table III
The association of miR-483-3p
expression level with clinicopathological factors in PDAC.
| Low expression
(N=18) | High expression
(N=14) | P-value |
|---|
| Age |
| <70 | 8 | 8 | 0.476 |
| ≥70 | 10 | 6 | |
| Gender |
| Male | 12 | 10 | 0.773 |
| Female | 6 | 4 | |
| Diabetes
mellitus |
| Yes | 9 | 8 | 0.688 |
| No | 9 | 6 | |
| Location |
| Head | 7 | 7 | 0.530 |
| Body-tail | 11 | 7 | |
| Tumor size
(mm) |
| Mean ± SD | 45.6±22.0 | 38.3±12.6 | 0.424 |
| T
classificationa |
| T1–T2 | 7 | 8 | 0.305 |
| T3–T4 | 11 | 6 | |
| N
classificationa |
| Yes | 5 | 4 | 0.960 |
| No | 13 | 10 | |
| Liver
metastasis |
| Yes | 5 | 4 | 0.960 |
| No | 13 | 10 | |
| Ascites |
| Yes | 2 | 2 | 0.788 |
| No | 16 | 12 | |
| Stagea |
| I–III | 10 | 9 | 0.618 |
| IV | 8 | 5 | |
| Operation |
| Yes | 3 | 5 | 0.217 |
| No | 15 | 9 | |
 | Table IVThe association of miR-21 expression
level with clinicopathological factors in PDAC. |
Table IV
The association of miR-21 expression
level with clinicopathological factors in PDAC.
| Low expression
(N=22) | High expression
(N=10) | P-value |
|---|
| Age |
| <70 | 13 | 3 | 0.127 |
| ≥70 | 9 | 7 | |
| Gender |
| Male | 14 | 8 | 0.355 |
| Female | 8 | 2 | |
| Diabetes
mellitus |
| Yes | 11 | 6 | 0.599 |
| No | 11 | 4 | |
| Location |
| Head | 10 | 4 | 0.773 |
| Body-tail | 12 | 6 | |
| Tumor size
(mm) |
| Mean ± SD | 38.8±15.8 | 50.3±22.5 | 0.137 |
| T
classificationa |
| T1–T2 | 12 | 3 | 0.197 |
| T3–T4 | 10 | 7 | |
| N
classificationa |
| Yes | 3 | 6 | 0.007c |
| No | 19 | 4 | |
| Liver
metastasis |
| Yes | 2 | 7 | <0.001c |
| No | 20 | 3 | |
| Ascites |
| Yes | 2 | 2 | 0.387 |
| No | 20 | 8 | |
| Stagea |
| I–III | 16 | 3 | 0.023b |
| IV | 6 | 7 | |
| Operation |
| Yes | 7 | 9 | 0.186 |
| No | 15 | 1 | |
Plasma miR-483-3p expression level is
decreased after resection of PDAC
Since the plasma miR-21 level was not increased in
operable patients (Table IV), we
evaluated alterations in the plasma miR-483-3p level alone after
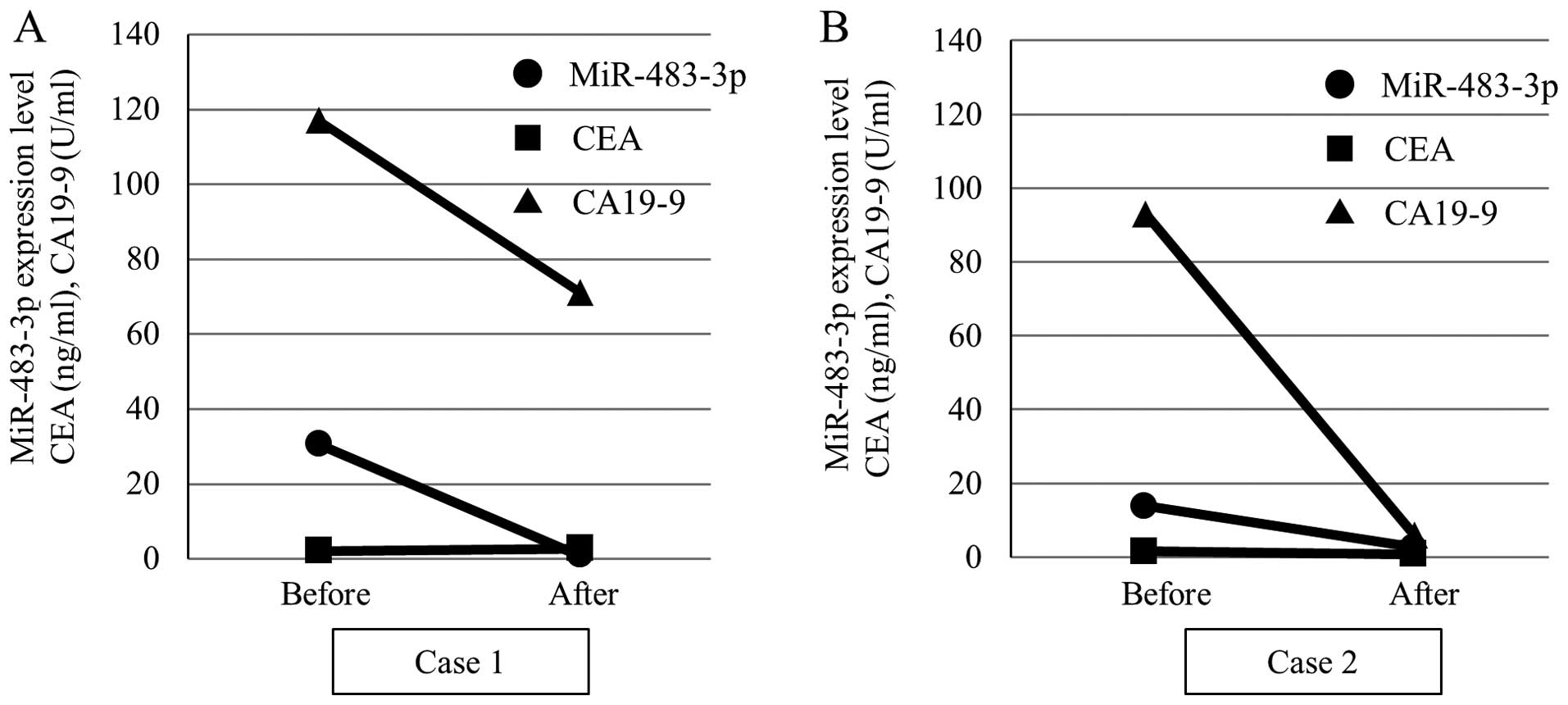
resection of the tumor tissues. Fig.
5 shows representative cases of the plasma expression level of
miR-483-3p and tumor markers before and after surgery. Case 1 was a
70-year-old female and case 2 was a 62-year-old female. In both
cases, pancreaticoduodenectomy was performed for the diagnosis of
pancreatic head cancer. By histological examination, the diagnosis
was pancreatic adenocarcinoma and the classification stage IIA
(T3N0M0) using UICC classification. As expected, a marked reduction
of the plasma miR-483-3p expression as well as the CA19-9 value was
observed after surgery, indicating that the expression of
miR-483-3p was strongly associated with the presence of PDAC.
Discussion
MiRNAs have been shown to play important roles in
the tumorigenesis and/or development of various cancers. Since it
exists stably in the plasma/serum, miRNA appears promising as a new
biomarker for the diagnosis and treatment of cancer. In PDAC
patients, numerous miRNAs including miR-21, miR-155, miR-196a and
miR-210 have been detected in the plasma/serum, and have been
suggested to be useful biomarkers for the diagnosis (6,7,9,26).
In the present study, we examined the expression of miR-483-3p as
well as miR-21 in the plasma of PDAC patient to assess whether
these miRNAs would be useful markers for the detection of PDAC and
for predicting the clinical status of this tumor. We clearly
demonstrated for the first time that the plasma miR-483-3p level
was significantly higher in PDAC patients compared to those of HC
and IPMN patients and that the AUC value of miR-483-3p for
discriminating PDAC from IPMN was higher than that of miR-21.
miR-483-3p has been shown to promote cell proliferation through
downregulation of its target gene, Smad4 in PDAC cells (42). Reportedly, the loss of Smad4
expression is frequently observed from the late stage of
carcinogenesis of PDAC, but infrequently in IPMN (46,47).
Taken together with our finding that miR-483-3p is expressed
specifically in PDAC patients without showing a relationship to the
clinical features, this miRNA may play a pivotal role in the
carcinogenesis rather than the development of PDAC through reducing
Smad4 expression.
Recent studies revealed that BD-IPMN is a risk
factor for PDAC since the occurrence of PDAC during the follow-up
period for BD-IPMN patients was not infrequent (48–50).
To date, there are no highly sensitive methods to predict the
occurrence of PDAC in IPMN patients. However, worsening diabetes
mellitus and abnormal serum CA19-9 levels have been demonstrated to
be useful to predict the development of PDAC in BD-IPMN patients
(51,52), and the sensitivity of CA19-9 to
predict concomitant PDAC was shown to be 45% (51). In the present study, the
sensitivity of the plasma miR-483-3p level to differentiate PDAC
from IPMN was 43.8%, similar to that of CA19-9, indicating that the
evaluation of this miRNA in plasma could also be a useful tool for
detecting the occurrence of PDAC in IPMN patients. The reduction of
the plasma miR-483-3p level by surgical resection of PDAC may
partially support this idea.
Overexpression of plasma miR-21 was demonstrated in
PDAC and was correlated with a poor prognosis in this tumor
(26,35–37).
As had been shown in previous studies, we also demonstrated that
the plasma miR-21 level was increased in PDAC patients compared to
HC and associated with poor prognosis in unresectable PDAC
patients. In addition, we revealed the relationship between the
plasma miR-21 level and liver metastasis in PDAC patients. The
involvement of MiR-21 in the metastasis of PDAC cells was also
clarified in vitro. Moriyama et al showed that miR-21
precursor transfection promoted proliferation, invasion and
gemcitabine resistance in PDAC cells in vitro (30). Moreover, Kadera et al
reported that expression of miR-21 in tumor-associated fibroblasts
as well as PDAC cells enhanced the metastatic potential (33). These findings suggest that stromal
cells as well as metastatic PDAC cells are likely to be the origin
of plasma miR-21, and thus high plasma levels of this miRNA are
detected in patients with advanced stage PDAC.
Intriguingly, the AUC value for the diagnosis of
PDAC was increased to a level similar to that of CA19-9 when the
miR-483-3p and miR-21 plasma levels were combined. This is due to
mutually independent mechanisms that enhance the expression of the
respective miRNAs. As discussed above, miR-483-3p and miR-21 are
upregulated in the process of carcinogenesis and development,
respectively. Thus, the evaluation of miR-483-3p together with
miR-21 would forecast the status in patients with PDAC.
Acknowledgements
This study was supported by Grants-in-Aid for
Scientific Research (KAKENHI) (24591022) to K.S., Pancreas Research
Foundation of Japan to M.A. and The Naito Foundation to K.S.
References
|
1
|
Hirata K, Egawa S, Kimura Y, et al:
Current status of surgery for pancreatic cancer. Dig Surg.
24:137–147. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Vincent A, Herman J, Schulick R, Hruban RH
and Goggins M: Pancreatic cancer. Lancet. 378:607–620. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Satake K, Kanazawa G, Kho I, Chung YS and
Umeyama K: A clinical evaluation of carbohydrate antigen 19-9 and
carcinoembryonic antigen in patients with pancreatic carcinoma. J
Surg Oncol. 29:15–21. 1985. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lee RC, Feinbaum RL and Ambros V: The C.
elegans heterochronic gene lin-4 encodes small RNAs with antisense
complementarity to lin-14. Cell. 75:843–854. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Dillhoff M, Liu J, Frankel W, Croce C and
Bloomston M: MicroRNA-21 is overexpressed in pancreatic cancer and
a potential predictor of survival. J Gastrointest Surg.
12:2171–2176. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bloomston M, Frankel WL, Petrocca F, et
al: MicroRNA expression patterns to differentiate pancreatic
adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA.
297:1901–1908. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Habbe N, Koorstra JB, Mendell JT, et al:
MicroRNA miR-155 is a biomarker of early pancreatic neoplasia.
Cancer Biol Ther. 8:340–346. 2009. View Article : Google Scholar :
|
|
9
|
Ryu JK, Hong SM, Karikari CA, Hruban RH,
Goggins MG and Maitra A: Aberrant microRNA-155 expression is an
early event in the multistep progression of pancreatic
adenocarcinoma. Pancreatology. 10:66–73. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li Y, Vandenboom TG II, Wang Z, Kong D,
Ali S, Philip PA and Sarkar FH: miR-146a suppresses invasion of
pancreatic cancer cells. Cancer Res. 70:1486–1495. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang Y, Li M, Wang H, Fisher WE, Lin PH,
Yao Q and Chen C: Profiling of 95 microRNAs in pancreatic cancer
cell lines and surgical specimens by real-time PCR analysis. World
J Surg. 33:698–709. 2009. View Article : Google Scholar
|
|
12
|
Yu J, Li A, Hong SM, Hruban RH and Goggins
M: MicroRNA alterations of pancreatic intraepithelial neoplasias.
Clin Cancer Res. 18:981–992. 2012. View Article : Google Scholar :
|
|
13
|
Kent OA, Mullendore M, Wentzel EA, et al:
A resource for analysis of microRNA expression and function in
pancreatic ductal adenocarcinoma cells. Cancer Biol Ther.
8:2013–2024. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Li A, Omura N, Hong SM, et al: Pancreatic
cancers epigenetically silence SIP1 and hypomethylate and
overexpress miR-200a/200b in association with elevated circulating
miR-200a and miR-200b levels. Cancer Res. 70:5226–5237. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yu J, Ohuchida K, Mizumoto K, et al:
MicroRNA, hsa-miR-200c, is an independent prognostic factor in
pancreatic cancer and its upregulation inhibits pancreatic cancer
invasion but increases cell proliferation. Mol Cancer. 9:1692010.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lee EJ, Gusev Y, Jiang J, et al:
Expression profiling identifies microRNA signature in pancreatic
cancer. Int J Cancer. 120:1046–1054. 2007. View Article : Google Scholar
|
|
17
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chen X, Ba Y, Ma L, et al:
Characterization of microRNAs in serum: a novel class of biomarkers
for diagnosis of cancer and other diseases. Cell Res. 18:997–1006.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Filipowicz W, Bhattacharyya SN and
Sonenberg N: Mechanisms of posttranscriptional regulation by
microRNAs: are the answers in sight? Nat Rev Genet. 9:102–114.
2008. View
Article : Google Scholar : PubMed/NCBI
|
|
20
|
Mitchell PS, Parkin RK, Kroh EM, et al:
Circulating microRNAs as stable blood-based markers for cancer
detection. Proc Natl Acad Sci USA. 105:10513–10518. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hasselmann DO, Rappl G, Tilgen W and
Reinhold U: Extracellular tyrosinase mRNA within apoptotic bodies
is protected from degradation in human serum. Clin Chem.
47:1488–1489. 2001.PubMed/NCBI
|
|
22
|
Cocucci E, Racchetti G and Meldolesi J:
Shedding microvesicles: artefacts no more. Trends Cell Biol.
19:43–51. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kosaka N, Iguchi H, Yoshioka Y, Takeshita
F, Matsuki Y and Ochiya T: Secretory mechanisms and intercellular
transfer of microRNAs in living cells. J Biol Chem.
285:17442–17452. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Arroyo JD, Chevillet JR, Kroh EM, et al:
Argonaute2 complexes carry a population of circulating microRNAs
independent of vesicles in human plasma. Proc Natl Acad Sci USA.
108:5003–5008. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Vickers KC, Palmisano BT, Shoucri BM,
Shamburek RD and Remaley AT: MicroRNAs are transported in plasma
and delivered to recipient cells by high-density lipoproteins. Nat
Cell Biol. 13:423–433. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang J, Chen J, Chang P, et al: MicroRNAs
in plasma of pancreatic ductal adenocarcinoma patients as novel
blood-based biomarkers of disease. Cancer Prev Res (Phila).
2:807–813. 2009. View Article : Google Scholar
|
|
27
|
Morimura R, Komatsu S, Ichikawa D, et al:
Novel diagnostic value of circulating miR-18a in plasma of patients
with pancreatic cancer. Br J Cancer. 105:1733–1740. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ho AS, Huang X, Cao H, Christman-Skieller
C, Bennewith K, Le QT and Koong AC: Circulating miR-210 as a novel
hypoxia marker in pancreatic cancer. Transl Oncol. 3:109–113. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kawaguchi T, Komatsu S, Ichikawa D, et al:
Clinical impact of circulating miR-221 in plasma of patients with
pancreatic cancer. Br J Cancer. 108:361–369. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Moriyama T, Ohuchida K, Mizumoto K, et al:
MicroRNA-21 modulates biological functions of pancreatic cancer
cells including theirproliferation, invasion, and chemoresistance.
Mol Cancer Ther. 8:1067–1074. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Giovannetti E, Funel N, Peters GJ, et al:
MicroRNA-21 in pancreatic cancer: correlation with clinical outcome
and pharmacologic aspects underlying its role in the modulation of
gemcitabine activity. Cancer Res. 70:4528–4538. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hwang JH, Voortman J, Giovannetti E, et
al: Identification of microRNA-21 as a biomarker for
chemoresistance and clinical outcome following adjuvant therapy in
resectable pancreatic cancer. PLoS One. 5:e106302010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kadera BE, Li L, Toste PA, Wu N, Adams C,
Dawson DW and Donahue TR: MicroRNA-21 in pancreatic ductal
adenocarcinoma tumor-associated fibroblasts promotes metastasis.
PLoS One. 8:e719782013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liu R, Chen X, Du Y, et al: Serum miRNA
expression profile as a biomarker in the diagnosis and prognosis of
pancreatic cancer. Clin Chem. 58:610–618. 2012. View Article : Google Scholar
|
|
35
|
Ali S, Almhanna K, Chen W, Philip PA and
Sarkar FH: Differentially expressed miRNAs in the plasma may
provide a molecular signature for aggressive pancreatic cancer. Am
J Transl Res. 28:28–47. 2010.
|
|
36
|
Liu J, Gao J, Du Y, et al: Combination of
plasma microRNAs with serum CA19-9 for early detection of
pancreatic cancer. Int J Cancer. 131:683–691. 2012. View Article : Google Scholar
|
|
37
|
Kong X, Du Y, Wang G, et al: Detection of
differentially expressed microRNAs in serum of pancreatic ductal
adenocarcinoma patients: miR-196a could be a potential marker for
poor prognosis. Dig Dis Sci. 56:602–609. 2011. View Article : Google Scholar
|
|
38
|
Hamada S, Satoh K, Fujibuchi W, et al:
MiR-126 acts as a tumor suppressor in pancreatic cancer cells via
the regulation of ADAM9. Mol Cancer Res. 10:3–10. 2012. View Article : Google Scholar
|
|
39
|
Hamada S, Satoh K, Miura S, et al: MiR-197
induces epithelial-mesenchymal transition in pancreatic cancer
cells by targeting p120 catenin. J Cell Physiol. 228:1255–1263.
2013. View Article : Google Scholar
|
|
40
|
Veronese A, Lupini L, Consiqlio J, et al:
Oncogenic role of miR-483-3p at the IGF2/483 locus. Cancer Res.
70:3140–3149. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Guled M, Lahti L, Lindholm PM, Salmenkivi
K, Bagwan I, Nicholson AG and Knuutila S: CDKN2A, NF2, and JUN are
dysregulated among other genes by miRNAs in malignant
mesothelioma-A miRNA microarray analysis. Genes Chromosomes Cancer.
48:615–623. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Hao J, Zhang S, Zhou Y, Hu X and Shao C:
MicroRNA 483-3p suppresses the expression of DPC4/Smad4 in
pancreatic cancer. FEBS Lett. 585:207–213. 2011. View Article : Google Scholar
|
|
43
|
Sobin LH, Gospodarowicz MK and Wittekind
C: TNM Cassification of Malignant Tumours. 7th edition.
Wiley-Blackwell; New York, NY: 2009
|
|
44
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Liang Y, Ridzon D, Wong L and Chen C:
Characterization of microRNA expression profiles in normal human
tissues. BMC Genomics. 8:1662007. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wilentz RE, Iacobuzio-Donahue CA, Argani
P, et al: Loss of expression of Dpc4 in pancreatic intraepithelial
neoplasia: evidence that DPC4 inactivation occurs late in
neoplastic progression. Cancer Res. 60:2002–2006. 2000.PubMed/NCBI
|
|
47
|
Biankin AV1, Biankin SA, Kench JG, et al:
Aberrant p16 (INK4A) and DPC4/Smad4 expression in intraductal
papillary mucinous tumours of the pancreas is associated with
invasive ductal adenocarcinoma. Gut. 50:861–868. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Yamaguchi K1, Ohuchida J, Ohtsuka T,
Nakano K and Tanaka M: Intraductal papillary-mucinous tumor of the
pancreas concomitant with ductal carcinoma of the pancreas.
Pancreatology. 2:484–490. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Uehara H, Nakaizumi A, Ishikawa O, et al:
Development of ductal carcinoma of the pancreas during follow-up of
branch duct intraductal papillary mucinous neoplasm of the
pancreas. Gut. 57:1561–1565. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Tanno S, Nakano Y, Koizumi K, et al:
Pancreatic ductal adenocarcinomas in long-term follow-up patients
with branch duct intraductal papillary mucinous neoplasms.
Pancreas. 39:36–40. 2010. View Article : Google Scholar
|
|
51
|
Ingkakul T1, Sadakari Y, Ienaga J, Satoh
N, Takahata S and Tanaka M: Predictors of the presence of
concomitant invasive ductal carcinoma in intraductal papillary
mucinous neoplasm of the pancreas. Ann Surg. 25:70–75. 2010.
View Article : Google Scholar
|
|
52
|
Kanno A, Satoh K, Hirota M, et al:
Prediction of invasive carcinoma in branch type intraductal
pappilary mucinous neoplasms of the pancreas. J Gastroenterol.
45:952–959. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Li A1, Yu J, Kim H, Wolfgang CL, Canto MI,
Hruban RH and Goggins M: MicroRNA array analysis finds elevated
serum miR-1290 accurately distinguishes patients with low-stage
pancreatic cancer from healthy and disease controls. Clin Cancer
Res. 19:3600–3610. 2013. View Article : Google Scholar : PubMed/NCBI
|