|
1
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chen W, Zheng R, Zeng H, Zhang S and He J:
Annual report on status of cancer in China, 2011. Chin J Cancer
Res. 27:2–12. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Gao Y, Hu N, Han X, Giffen C, Ding T,
Goldstein A and Taylor P: Family history of cancer and risk for
esophageal and gastric cancer in Shanxi, China. BMC Cancer.
9:2692009. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Song Y, Li L, Ou Y, Gao Z, Li E, Li X,
Zhang W, Wang J, Xu L, Zhou Y, et al: Identification of genomic
alterations in oesophageal squamous cell cancer. Nature. 509:91–95.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Png KJ, Halberg N, Yoshida M and Tavazoie
SF: A microRNA regulon that mediates endothelial recruitment and
metastasis by cancer cells. Nature. 481:190–194. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Isozaki Y, Hoshino I, Akutsu Y, Hanari N,
Mori M, Nishimori T, Murakami K, Akanuma N, Takeshita N, Maruyama
T, et al: Usefulness of microRNA-375 as a prognostic and
therapeutic tool in esophageal squamous cell carcinoma. Int J
Oncol. 46:1059–1066. 2015.
|
|
9
|
Spizzo R, A1meida MI, Colombatti A and
Calin GA: Long non-coding RNAs and cancer: A new frontier of
translational research? Oncogene. 3 l:4577–4587. 2012. View Article : Google Scholar
|
|
10
|
Necsulea A, Soumillon M, Warnefors M,
Liechti A, Daish T, Zeller U, Baker JC, Grützner F and Kaessmann H:
The evolution of lncRNA repertoires and expression patterns in
tetrapods. Nature. 505:635–640. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yang L, Duff MO, Graveley BR, Carmichael
GG and Chen LL: Genome-wide characterization of non-polyadenylated
RNAs. Genome Biol. 12:R162011. View Article : Google Scholar
|
|
12
|
Caley DP, Pink RC, Trujillano D and Carter
DR: Long non-coding RNAs, chromatin, and development. Sci World J.
10:90–102. 2010. View Article : Google Scholar
|
|
13
|
Mercer TR, Dinger ME and Mattick JS: Long
non-coding RNAs: Insights into functions. Nat Rev Genet.
10:155–159. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Guttman M, Amit I, Garber M, French C, Lin
MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, et al:
Chromatin signature reveals over a thousand highly conserved large
non-coding RNAs in mammals. Nature. 458:223–227. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yuan JH, Yang F, Wang F, Ma JZ, Guo YJ,
Tao QF, Liu F, Pan W, Wang TT, Zhou CC, et al: A long non-coding
RNA activated by TGF-β promotes the invasion-metastasis cascade in
hepatocellular carcinoma. Cancer Cell. 25:666–681. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gupta RA, Shah N, Wang KC, Kim J, Horlings
HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote cancer
metastasis. Nature. 464:1071–1076. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Qiu JJ, Lin YY, Ding JX, Feng WW, Jin HY
and Hua KQ: Long non-coding RNA ANRIL predicts poor prognosis and
promotes invasion/metastasis in serous ovarian cancer. Int J Oncol.
46:2497–2505. 2015.PubMed/NCBI
|
|
18
|
Yang H, Liu Z, Yuan C, Zhao Y, Wang L, Hu
J, Xie D, Wang L and Chen D: Elevated JMJD1A is a novel predictor
for prognosis and a potential therapeutic target for gastric
cancer. Int J Clin Exp Pathol. 8:11092–11099. 2015.PubMed/NCBI
|
|
19
|
Kim K, Jutooru I, Chadalapaka G, Johnson
G, Frank J, Burghardt R, Kim S and Safe S: HOTAIR is a negative
prognostic factor and exhibits pro-oncogenic activity in pancreatic
cancer. Oncogene. 32:1616–1625. 2013. View Article : Google Scholar
|
|
20
|
Martínez-Fernández M, Rubio C, Segovia C,
López-Calderón FF, Dueñas M and Paramio JM: EZH2 in Bladder Cancer,
a Promising Therapeutic Target. Int J Mol Sci. 16:27107–27132.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Gerstein MB, Kundaje A, Hariharan M, Landt
SG, Yan KK, Cheng C, Mu XJ, Khurana E, Rozowsky J, Alexander R, et
al: Architecture of the human regulatory network derived from
ENCODE data. Nature. 489:91–100. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Guttman M and Rinn JL: Modular regulatory
principles of large non-coding RNAs. Nature. 482:339–346. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wang J, Xie H, Ling Q, et al:
Coding-non-coding gene expression in intrahepatic
cholangiocarcinoma. Transl Res. 168:107–121. 2016. View Article : Google Scholar
|
|
24
|
Lu XF, Li EM, Du ZP, Xie JJ, Guo ZY, Gao
SY, Liao LD, Shen ZY, Xie D and Xu LY: Specificity protein 1
regulates fascin expression in esophageal squamous cell carcinoma
as the result of the epidermal growth factor/extracellular
signal-regulated kinase signaling pathway activation. Cell Mol Life
Sci. 67:3313–3329. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Yeo SY, Ha SY, Yu EJ, Lee KW, Kim JH and
Kim SH: ZNF282 (Zinc finger protein 282), a novel E2F1
co-activator, promotes esophageal squamous cell carcinoma.
Oncotarget. 5:12260–12272. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chen Y, Wang Y, Song H, Wang J, Yang H,
Xia Y, Xue J, Li S, Chen M and Lu Y: Expression profile of
apoptosis-related genes potentially explains early recurrence after
definitive chemoradiation in esophageal squamous cell carcinoma.
Tumour Biol. 35:4339–4346. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Rinn JL and Chang HY: Genome regulation by
long non-coding RNAs. Annu Rev Biochem. 81:145–166. 2012.
View Article : Google Scholar
|
|
28
|
Chen G, Wang Z, Wang D, Qiu C, Liu M, Chen
X, Zhang Q, Yan G and Cui Q: LncRNADisease: A database for
long-non-coding RNA-associated diseases. Nucleic Acids Res.
41D:D983–D986. 2013. View Article : Google Scholar
|
|
29
|
Gutschner T and Diederichs S: The
hallmarks of cancer: A long non-coding RNA point of view. RNA Biol.
9:703–719. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Huarte M: LncRNAs have a say in protein
translation. Cell Res. 23:449–451. 2013. View Article : Google Scholar :
|
|
31
|
Nagano T and Fraser P: No-nonsense
functions for long non-coding RNAs. Cell. 145:178–181. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hao Y, Wu W, Shi F, Dalmolin RJ, Yan M,
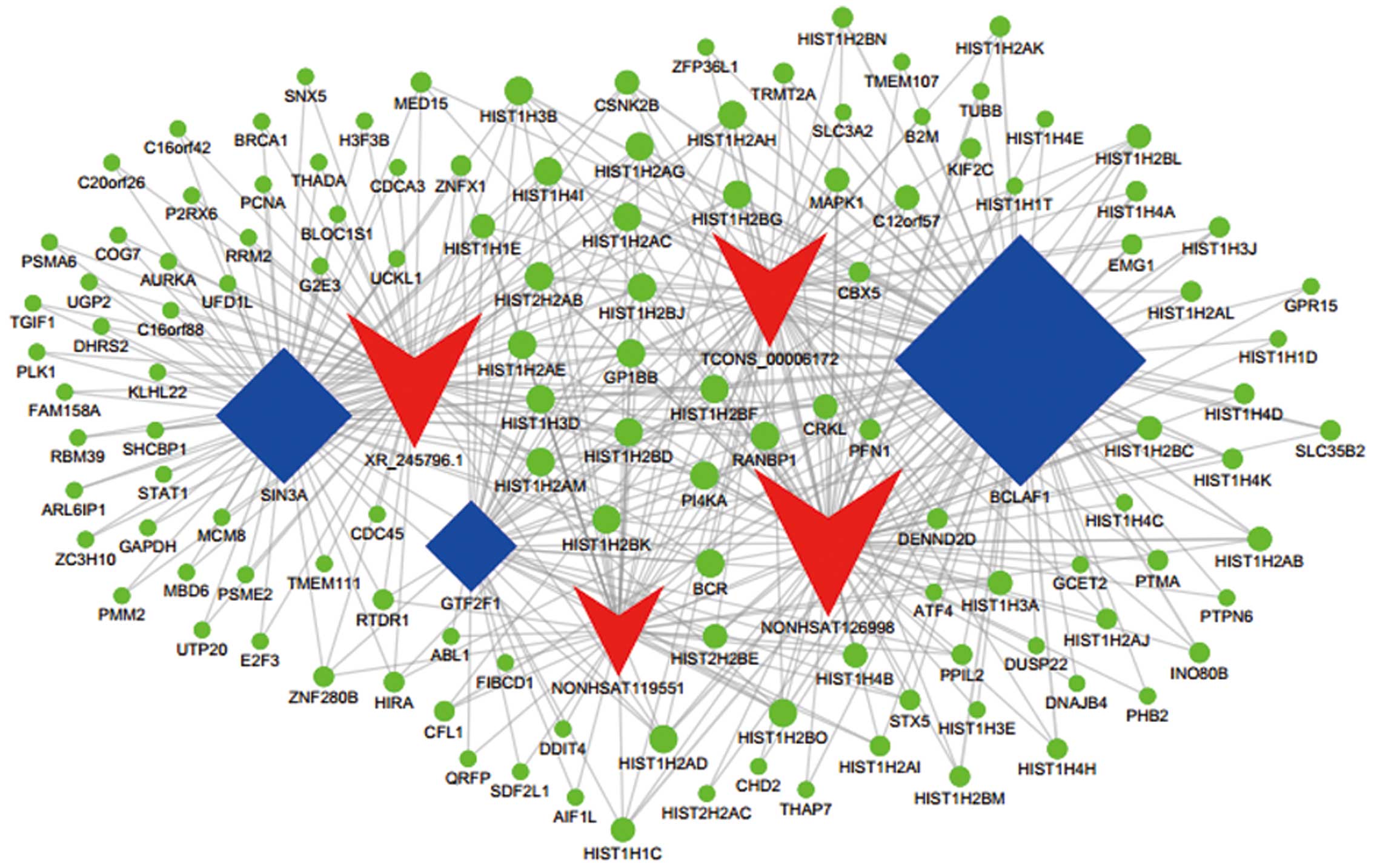
Tian F, Chen X, Chen G and Cao W: Prediction of long non-coding RNA
functions with co-expression network in esophageal squamous cell
carcinoma. BMC Cancer. 15:1682015. View Article : Google Scholar
|
|
33
|
Lee HK, Hsu AK, Sajdak J, Qin J and
Pavlidis P: Coexpression analysis of human genes across many
microarray data sets. Genome Res. 14:1085–1094. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Eisen MB, Spellman PT, Brown PO and
Botstein D: Cluster analysis and display of genome-wide expression
patterns. Proc Natl Acad Sci USA. 95:14863–14868. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: The Gene Ontology Consortium: Gene ontology: Tool for the
unification of biology. Nat Genet. 25:25–29. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wang KC, Yang YW, Liu B, Sanyal A,
Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta
RA, et al: A long non-coding RNA maintains active chromatin to
coordinate homeotic gene expression. Nature. 472:120–124. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Guenzl PM and Barlow DP: Macro lncRNAs: A
new layer of cis-regulatory information in the mammalian genome.
RNA Biol. 9:731–741. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Lee JT: Lessons from X-chromosome
inactivation: Long ncRNA as guides and tethers to the epigenome.
Genes Dev. 23:1831–1842. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Jiang Q, Wang J, Wang Y, Ma R, Wu X and Li
Y: TF2LncRNA: Identifying common transcription factors for a list
of lncRNA genes from ChIP-Seq data. BioMed Res Int.
2014:3176422014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Yang JH, Li JH, Jiang S, Zhou H and Qu LH:
ChIPBase: A database for decoding the transcriptional regulation of
long non-coding RNA and microRNA genes from ChIP-Seq data. Nucleic
Acids Res. 41(D1): D177–D187. 2013. View Article : Google Scholar :
|
|
41
|
Lopez-Pajares V, Qu K, Zhang J, Webster
DE, Barajas BC, Siprashvili Z, Zarnegar BJ, Boxer LD, Rios EJ, Tao
S, et al: A LncRNA-MAF:MAFB transcription factor network regulates
epidermal differentiation. Dev Cell. 32:693–706. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Li J, Chen Z, Tian L, Zhou C, He MY, Gao
Y, Wang S, Zhou F, Shi S, Feng X, et al: LncRNA profile study
reveals a three-lncRNA signature associated with the survival of
patients with oesophageal squamous cell carcinoma. Gut.
63:1700–1710. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Dulak AM, Stojanov P, Peng S, Lawrence MS,
Fox C, Stewart C, Bandla S, Imamura Y, Schumacher SE, Shefler E, et
al: Exome and whole-genome sequencing of esophageal adenocarcinoma
identifies recurrent driver events and mutational complexity. Nat
Genet. 45:478–486. 2013. View Article : Google Scholar : PubMed/NCBI
|