Introduction
Both miRNAs and lncRNAs play important roles in
regulating cellular processes (1).
It was found that lncRNA can influence the occurrence and
development of tumor by regulating expression of miRNAs, and will
cause dysfunction of protein encoding genes if expressed abnormally
(2). The expression profile of
lncRNAs in a variety of tumor cells differs from that in normal
cells, which may contribute to the tumor development (3). The study on the interaction between
miRNAs and lncRNAs will revolutionize our knowledge about cell
structural network and regulatory network, and bring immeasurable
scientific and clinical value.
ASLNC02525, is a highly expressed lncRNA in liver
cancer that we identified by comparing the expression profile of
lncRNAs in clinical liver cancer and adjacent tissues. To elucidate
the role of ASLNC02525 in liver cancer, we examined ASLNC02525
levels in three selected liver cancer cell lines, HepG2, QGY-7701
and SMMC-7721, by fluorescent quantitation, and found higher
expression of ASLNC02525 in cancer cells than in the normal liver
epithelial cells L-02. We knocked down ASLNC02525 in these cell
lines via gene interference, which significantly inhibited
proliferation and invasion, indicating an oncogene-like role of
ASLNC02525. To further explain the tumor-promoting action of
ASLNC02525, we silenced ASLNC02525 in HepG2, and screened the
differentially expressed proteins, among which one was twist1. We
wandered whether ASLNC02525 directly regulates twist1. The results
revealed that silencing ASLNC02525 only increased protein level but
not mRNA level of twist1. At the same time, RIP experiment proved
that ASLNC02525 did not bind to twist1 protein directly. We
searched for the miRNAs having potential to bind to 3′-UTR
(untranslated regions) of twist1 as well as the ones that
theoretically bind to ASLNC02525, and found hsa-miRNA-489-3p, which
has one theoretical binding site on 3′-UTR of twist1 and four sites
on ASLNC02525. Thus, we progressed to validate whether
hsa-miRNA-489-3p is associated with the effect of ASLNC02525 on
twist1 protein.
The twist family of basic helix-loop-helix
transcription factors are highly conserved in nucleotide and amino
acid sequences, and known for their crucial role in embryonic
development (4,5). Nowadays, as one of the hotspots in
cancer research, twists have been proved to contribute to the
initiation and development of a multitude of cancers (6–10).
It can affect a host of signaling pathways, such as PI3K/AKT, STAT3
and ras pathways (10,11), playing a role of an oncogene
(12–16).
In the present study, we validated that high
expression of ASLNC02525 impaired the negative regulation of twist1
by hsa-miRNA-489-3p like an RNA sponge. The high expression of
ASLNC02525 canceled the inhibition of twist1 by hsa-miRNA-489-3p,
and silencing of ASLNC02525 recovered the negative regulation, so
the twist1 protein was decreased, subsequently inhibiting the
proliferation and invasion of liver cancer cells, which provides a
novel thought and theoretical basis for gene therapy of liver
cancer.
Materials and methods
Cell culture
HepG2, QGY-7701, SMMC-7721 and L-02 cells, obtained
from the Cell Bank of the Chinese Academy of Sciences (Shanghai,
China), were maintained in RPMI-1640 (Invitrogen, Carlsbad, CA,
USA) supplemented with 10% fetal calf serum (FCS; Invitrogen). 293T
cells, purchased from the American Type Culture Collection (ATCC;
Rockville, MD, USA) were maintained in Dulbecco's minimum essential
medium (DMEM) supplemented with 10% FBS. All these adherent cells
were passaged by 0.25% trypsin digestion (Invitrogen) and incubated
in an atmosphere of 5% CO2 at 3°C.
Construction of vectors
Construction of pSIH1-shRNA-ASLNC02525
plasmid
A siRNA sequence complementarily binding to
ASLNC02525 was chosen. The target sequences of siRNA
(5′-GTCTGTGTCGCCCGTCTGTG-3′) are homologous to ASLNC02525,
respectively. The oligonucleotide templates of these shRNAs were
chemically synthesized and cloned into the linear pSIH1-H1-copGFP
shRNA Vector (System Biosciences, Palo Alto, CA, USA) which was
obtained through digestion by BamHI and EcoRI
(Takara, Dalian, China) and purification by agarose gel
electrophoresis. An invalid siRNA sequence
(5′-GAAGCCAGATCCAGCTTCC-3′) was used as an NC (negative control).
Sequencing was used to confirm the vectors constructed
(pSIH1-shRNA-ASLNC02525 and pSIH1-NC).
Construction of luciferase reporter
vectors
Total RNA was extracted from 293 cells (ATCC),
reverse-transcribed into cDNA, and 2 µl of the reaction
product subsequently was used as a template for PCR. Primers were
designed that targeted the 3′-UTR of the twist1 gene such that
flanking XbaI restriction sites were introduced into the 224
bp (base-pair) PCR product containing the 5′-GATGTCA-3′
hsa-miRNA-489-3 target site. The forward and reverse primer
sequences were 5′-GCTCTAGACAGCAGGGCCGGAGACCTA-3′ and 5′-GC
TCTAGAGGTGCCGCTGCCCGTCTGGGAA-3′, respectively. PCR reaction
conditions were as follows: 35 cycles of a 94 denaturing step for
30 sec, a 55 annealing step for 30 sec, and a 72 elongation step
for 10 sec. The PCR product was digested with XbaI (Takara)
and cloned into the pGL3-promoter luciferase reporter vector
(Promega, Madison, MI, USA) to generate the vector
pGL3-Pro-WT-twist. The hsa-miRNA-489-3p target site in the
pGL3-Pro-WT-twist vector was mutated from 5′-GATGTCA-3′ to
5′-GCAGTAT-3′ to construct the mutated reporter vector,
pGL3-Pro-MT-twist. The products of all cloning and mutagenesis
reactions were confirmed by the DNA sequencing. Endotoxin-free DNA
was prepared in all cases. The hsa-miRNA-489-3p mimic
(5′-GUGACAUCACAUAUACGGCAGC-3′), the hsa-miRNA-489-3p inhibitor
(5′-GCUGCCGUAUAUGUGAUGUCAC-3), and negative control miRNA
(miRNA-NC, 5′-UAGAUCGCAGCUUGGAACCACA-3′) were all chemically
synthesized (Invitrogen).
Lentivirus packaging
One day before the transfection, 293TN cells were
seeded into 10-cm dishes (Corning, Inc., Corning NY, USA). A total
of 2 µg of each pSIH1-shRNA-ASLNC02525 vector or pSIH1-NC
and 10 g pPACK Packaging Plasmid mix (System Biosciences) were
co-transfected using Lipofectamine 2000 (Invitrogen) in accordance
with the manufacturer's protocol. The medium was replaced with
Dulbecco's modified Eagle's medium (DMEM) plus 1% fetal bovine
serum (FBS). Forty-eight hours later, the supernatant was harvested
and then was cleared by centrifugation at 5000 × g at 4°C for 5
min, and passed through a 0.45-µm PVDF membrane (Millipore,
Billerica, MA, USA). The titer of virus was determined by gradient
dilution. The packaged lentiviruses were named as
Lv-shRNA-ASLNC02525 and Lv-NC.
Assessment of ASLNC02525,
hsa-miRNA-489-3p and twist1 protein in liver cancer specimens and
liver cancer cell lines
Five pairs of liver cancer and para-carcinoma
tissues (Huashan Hospital, Shanghai, China), as well as HepG2,
QGY-7701, SMMC-7721 and L-02 cells (1×107 each), were
collected, followed by total RNA extraction and real-time PCR for
measurement of ASLNC02525 and hsa-miRNA-489-3p and total protein
extraction and western blotting for twist1.
Effect of silencing ASLNC02525 on
hsa-miRNA-489-3p and twist1 protein
HepG2, QGY-7701, SMMC-7721 and L-02 in logarithmic
phase were seeded to 6-well plates at 5×105 cells/well.
One day later, viral solution was added at a MOI of 10. The
infection efficiency was evaluated by observing and analyzing the
fluorescent mark 72 h after the infection. In addition, total RNA
and protein were isolated from the cells and subjected to real-time
PCR and western blotting for ASLNC02525 and twist1 protein,
respectively.
Luciferase reporter experiment
We used TargetScan (http://www.targetscan.org/) to predict whether an
hsa-miRNA-489-3p binding site exists within the 3′-UTR of human
twist1 mRNA (NM_000474.3). The results showed that a seven-base
hsa-miRNA-489-3p seed sequence is present in the 3′-UTR of twist1
mRNA. The same tool was used to predict the binding sites of
hsa-miRNA-489-3p on ASLNC02525.
A suspension of 293 cells in logarithmic phase
growth was prepared and the number of viable cells counted using a
hemocytometer in conjunction with trypan blue staining. The cells
were seeded into 6-well plates at a concentration of
2×105 cells/well and maintained in DMEM supplemented
with 10% FCS at 37°C for 24 h in a 5% CO2 atmosphere.
The transfection of plasmid DNA and RNA was performed using
Lipofectamine 2000 (Invitrogen). Transfection of cells with pGL-TK
(100 ng) served as a reference for luciferase detection. Luciferase
activity was measured using the Dual-luciferase reporter assay
system (Promega) 48 h after the transfection.
The experiment to observe the effect of ASLNC02525
depletion on the inhibition of luciferase by hsa-miR489-3p mimics
was carried out in HepG2 cells, the plasmid transfection and
luciferase activity assay were the same as that used in validation
of the target site.
Cellular proliferation assay
Increased proliferation is one indicator in liver
cancer cells whether ASLNC02525 knockdown could inhibit
proliferation. HepG2, QGY-7701 and SMMC-7721 cells infected with
recombinant lentiviruses (Lv-shRNA-ASLNC02525) for 72 h,
trypsinized, and seeded into 96-well plates at a density of
1×104 cells/well. The cells were cultured under normal
conditions and cell viability was examined using Cell Counting
kit-8 (CCK-8; Dojindo Laboratories, Inc., Kumamoto, Japan) at 24-,
48- and 72-h time-points. Briefly, 10 µl of CCK-8 solution
was added, and the cells were then cultured under normal conditions
for an additional 4 h before the measurement of absorbance at 450
nm.
Cell invasion assay
Cell invasion experiments were performed using the
QCM™ 24-well Fluorimetric Cell Invasion Assay kit (ECM554; Chemicon
International, Temecula, CA, USA) according to the manufacturer's
instructions. The kit uses an insert polycarbonate membrane with an
8-µm pore size. The insert was coated with a thin layer of
EC Matrix™ that occluded the membrane pores and blocked migration
of noninvasive cells. Culture medium (500 µl) supplemented
with 10% FBS was used as chemoattractant. Cells that migrated and
invaded the underside of the membrane were fixed in 4%
paraformaldehyde. The invading cells were stained by DAPI, and the
number was then determined by fluorescence and reported as relative
fluorescence units (RFUs).
Examination of effects of ASLNC02525 on
twist1 and downstream proteins
PI3K, STAT3 and E-cadherin levels in HepG2, QGY-7701
and SMMC-7721 cells were assessed using western blotting 72 h after
the infection with Lv-NC or Lv-shRNA-ASLNC02525.
Real-time PCR
Total RNA was isolated with TRIzol reagent
(Invitrogen) according to the manufacturer's instructions and
reversely transcribed into cDNA using the M-MLV Reverse
Transcriptase (Takara Bio, Inc., Shiga, Japan) and oligo(dT)18
primer (Takara Bio). The following specific primers were used in
quantitative PCR of human ASLNC02525 and β-actin: ASLNC02525:
5′-TTCCAGTGACTCCACGTGC-3′ and 5′-AACTTTGGGCCTGTGCCGAAGGGT-3′; and
β-actin: 5′-CCTGTACGCCAACACAGTGC-3′ and 5′-ATACTCCTGC
TTGCTGATCC-3′. The lengths of amplified products were 219 and 211
bp, respectively. Real-time PCR was performed using SYBR Premix Ex
Taq™ kit (Takara) and TP800 System (Takara Bio). cDNA from 200 ng
total RNA was used as the template. The PCR reactions was carried
out under the following conditions: 40 cycles of denaturation at
95°C for 10 sec, annealing at 60°C for 20 sec and extension at 72°C
for 20 sec. The mRNA levels of twist1 were normalized using the
ΔΔCt method, to the expression of an endogenous housekeeping gene,
β-actin.
The expression of hsa-miRNA-489-3p was analyzed with
the 2−ΔΔCt method. For each sample, triplicate
determinations were performed, and mean values were adopted for
further calculations. All values were normalized to an endogenous
U6 control. The PCR primers for mature hsa-miRNA-489-3p or U6 were
designed as follows: hsa-miRNA-489-3p sense,
5′-GTGACATCACATATACGGCAGC-3′ and reverse, 5′-GT
CGTATCCAGTGCGTGTCGTG-3′; U6 sense, 5′-GTGCTC GCTTCGGCAGCACAT-3′ and
reverse, 5′-TACCTTGCGA AGTGCTTAAAC-3′.
Detection of protein contents in cells or
tissues
The total protein was extracted from the cells using
M-PER mammalian protein extraction reagent (Pierce, Rockford, IL,
USA) or from tissues using T-PER tissue protein extraction reagent
(Pierce). Equal amounts of protein (25 µg per lane)
estimated by a bicinchoninic acid (BCA) protein assay kit (Pierce)
were loaded onto (11%) SDS-PAGE gels and transferred onto
nitrocellulose membranes. The blots were probed with a monoclonal
antibody against human twist 1 (1:300), PI3K (1:500), STAT3
(1:600), E-cadherin (1:350) and β-actin (1:1,200) (Santa Cruz
Biotechnology, Santa Cruz, CA, USA), followed the secondary
HRP-conjugated anti-mouse/rabbit antibody (Santa Cruz
Biotechnology). After washing, the bands were detected by
chemiluminescence and imaged with X-ray film. β-actin was used as
an endogenous reference for normalization.
Statistical analysis
All data are expressed as mean ± SD, and were
analyzed by the one-way ANOVA. Least significant difference (LSD)
was used for multiple comparisons between any two means. P-values
<0.05 were considered statistically significant. All statistical
analyses were performed using SPSS 13.0 software.
Results
Assessment of ASLNC02525,
hsa-miRNA-489-3p and twist1 protein in liver cancer tissues
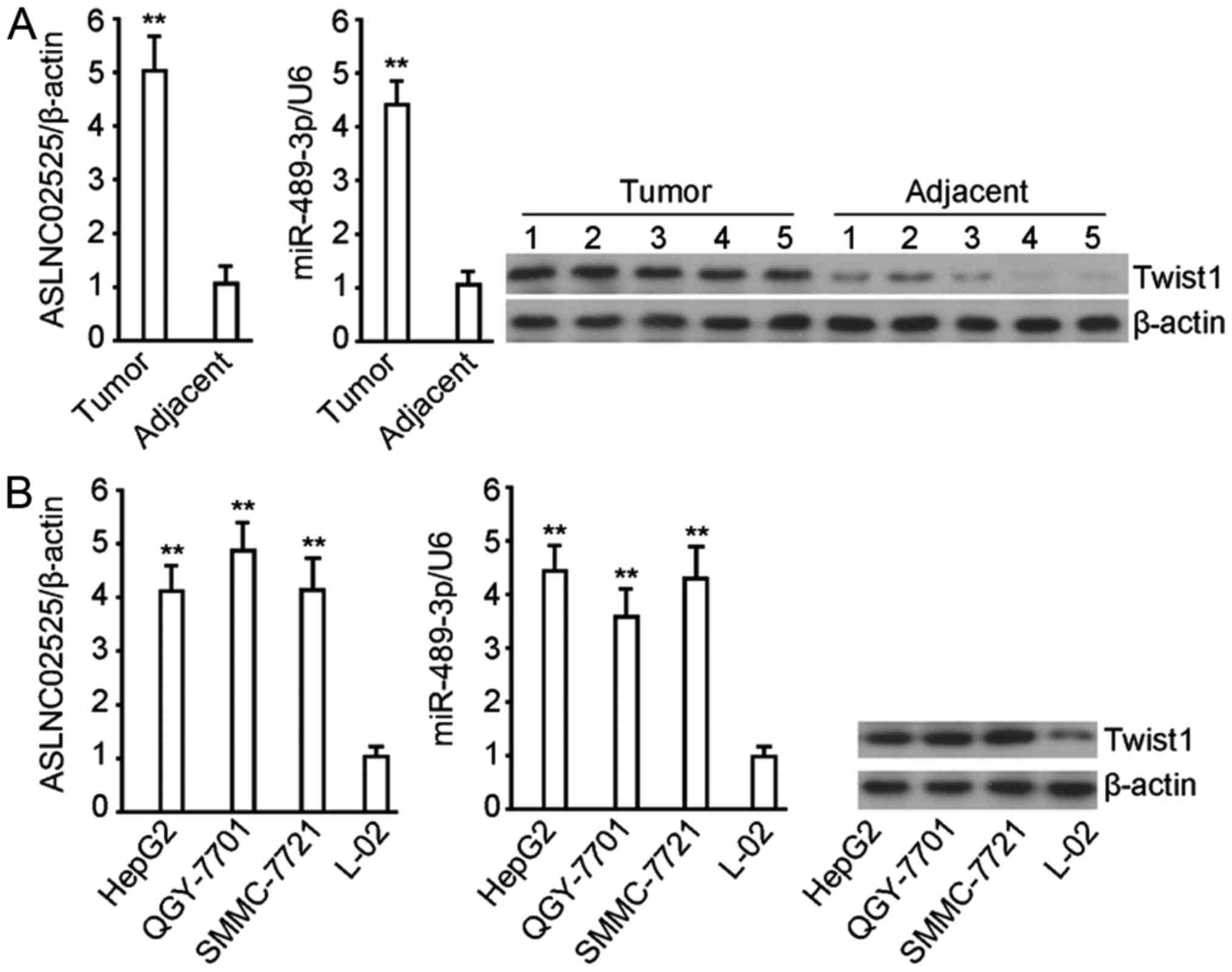
The expression of twist1 was higher in liver cancer
than in the adjacent tissue (P<0.01) (Fig. 1A), so were ASLNC02525 and
hsa-miRNA-489-3p. Compared with normal liver cell line L-02, the
three cancer cell lines exhibited higher twist1, as well as
ASLNC02525 and hsa-miRNA-489-3p (P<0.01) (Fig. 1B). The finding that twist1, a
predicted target gene of hsa-miRNA-489-3p, was positively
correlated with hsa-miRNA-489-3p in tissue and liver cancer cells
raised the question whether hsa-miRNA-489-3p regulates twist1,
which we investigated in following experiments.
Effect of silencing ASLNC02525 via
lentiviral approach on hsa-miRNA-489-3p and twist1 protein in liver
cancer cells
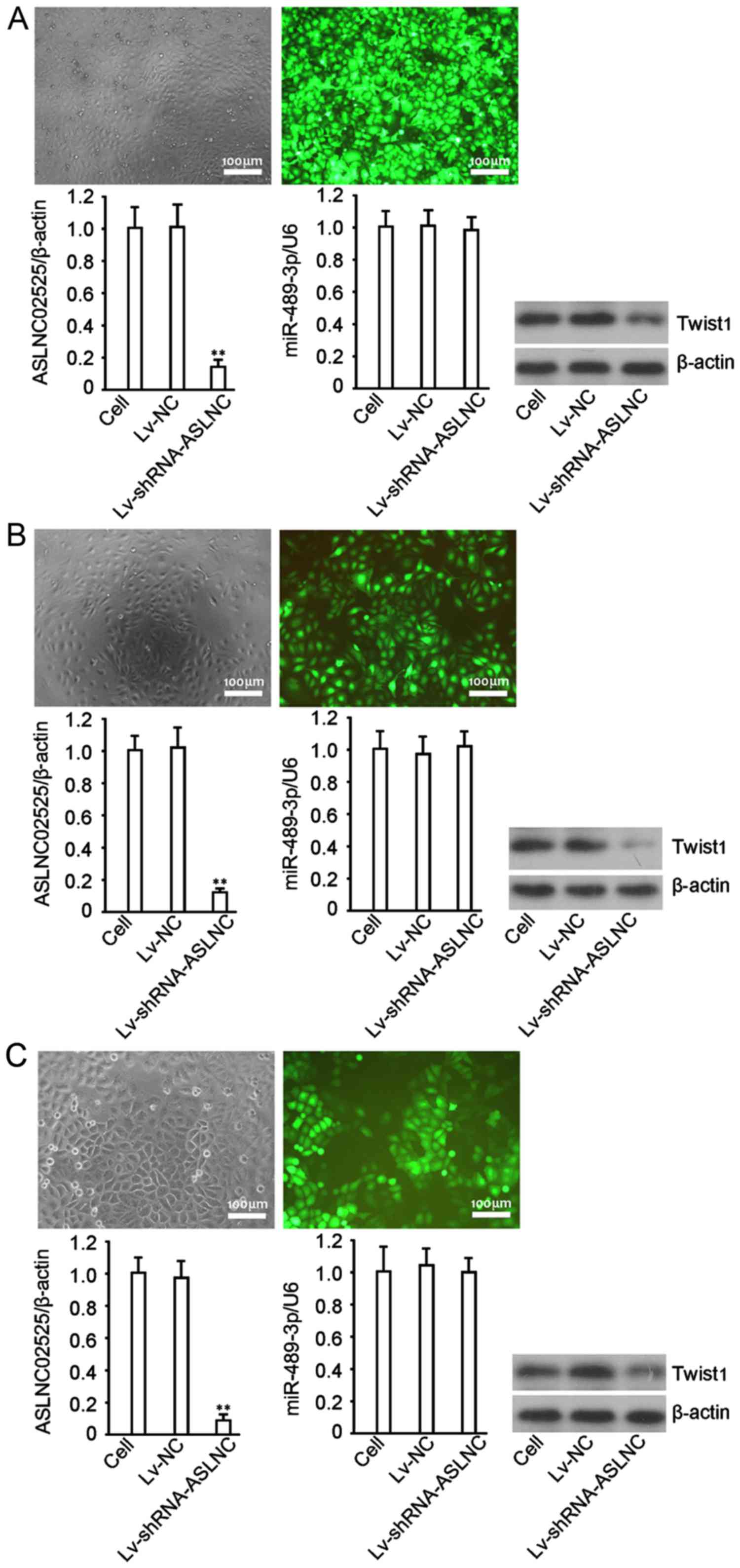
Recombinant lentiviruses, Lv-NC and
Lv-shRNA-ASLNC02525, were used to infect HepG2, QGY-7701 and
SMMC-7721. Green fluorescent protein (GFP) was detected in most of
the cells 72 h after the infection, and the proportion of
GFP-expressing cells suggested that the gene delivery efficiency
was >90% in all three cell lines (Fig. 2, left panels). ASLNC02525 was
significantly decreased by Lv-shRNA-ASLNC02525 (P<0.05)
(Fig. 2, middle panels). No change
in hsa-miRNA-489-3p was observed, but twist1 protein was
significantly decreased (P<0.01) (Fig. 2, right panel). The finding is of
importance: in the presence of high ASLNC02525 expression,
hsa-miRNA-489-3p was not correlated to twist1 protein expression
but in the absence of ASLNC02525, hsa-miRNA-489-3p was negatively
correlated to twist1 protein.
Luciferase experiments
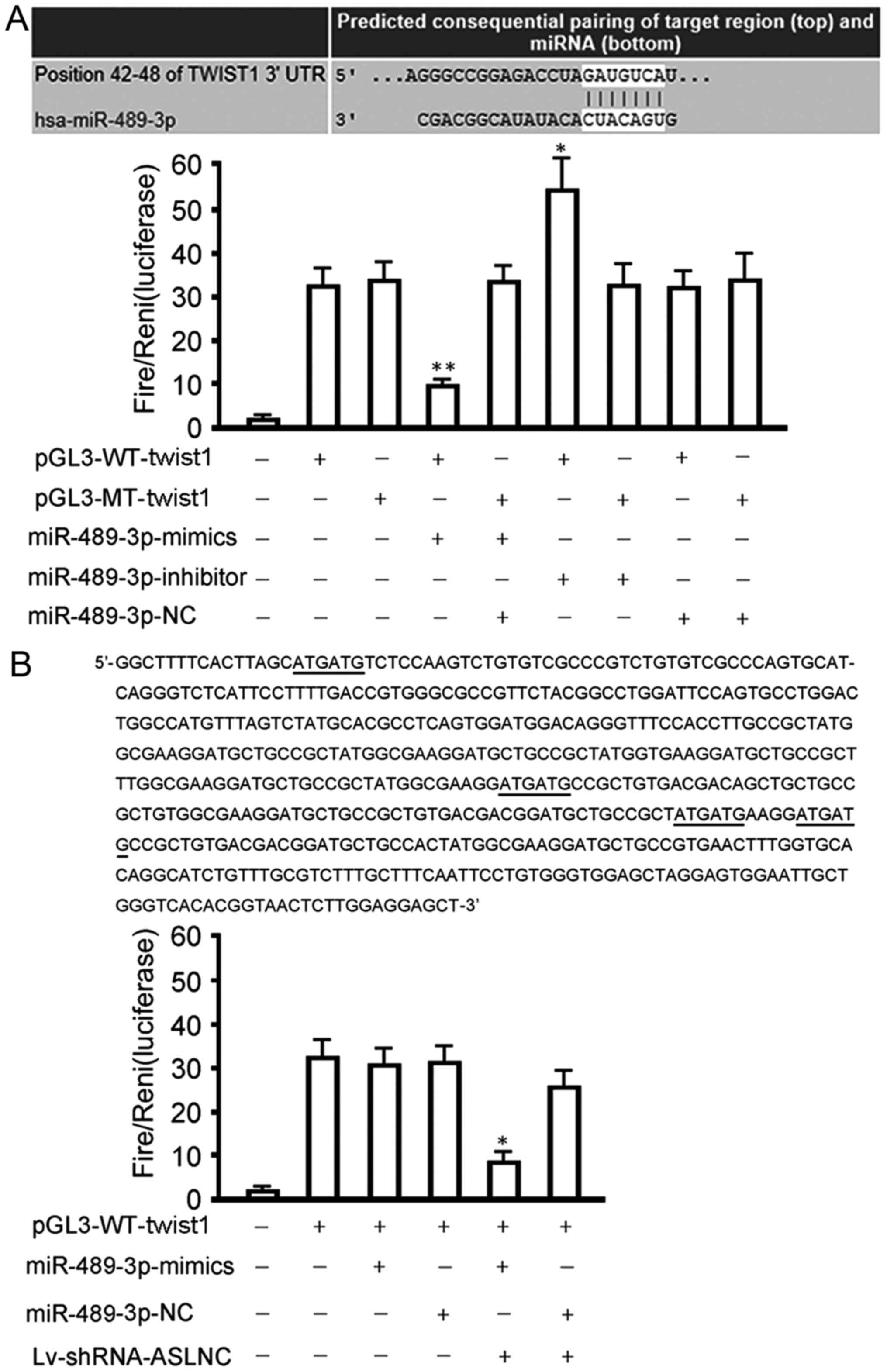
Our bioinformatics analysis identified a seven-base
hsa-miRNA-489-3p seed sequence in the 3′-UTR of twist1 mRNA
(Fig. 3A). We therefore
constructed luciferase reporter vectors to verify whether this site
represents a valid hsa-miRNA-489-3p target. Reporter vectors were
generated that contained the wild-type twist1 3′-UTR or a variant
in which the hsa-miRNA-489-3p target site within the 3′-UTR had
been mutated. Both reporter constructs expressed luciferase at a
high level (Fig. 2A). However, the
miRNA-489-3p mimic significantly inhibited luciferase activity in
cells transfected with the reporter vector encoding the wild-type
3′-UTR (32.43±3.69 vs. 9.01±1.08; P<0.01), while the
miRNA-489-3p inhibitor significantly increased luciferase activity
in these cells (33.78±4.02 vs. 51.84±11.02; P<0.05). Conversely,
in cells transfected with the reporter vector encoding the mutated
hsa-miRNA-489-3p target site, neither the miRNA-489-3p mimic nor
the miRNA489-3p inhibitor had any observable effect on luciferase
activity (P>0.05). Co-transfection of miRNA-489-3p-NC
(non-targeting control) had no effect on the luciferase activity of
either of the vectors (P>0.05). These results verified the
presence of an hsa-miR-489-3p target site in the 3′-UTR of twist1
mRNA and demonstrated that binding of hsa-miR-489-3p to this target
site downregulates twist1 expression. Notably, miRNA-489-3p mimics
lost its inhibition on the activity of luciferase expressed by
wild-type (WT) luciferase reporter vector in HepG2 cells, and
regained the inhibition after ASLNC02525 silencing (Fig. 3B). Taken together, these data
suggest that ASLNC02525 overexpression impaired negative regulation
of twist1 by hsa-miRNA-489-3p.
Effect of ASLNC02525 depletion on
proliferation and invasion of hepatoma carcinoma cells
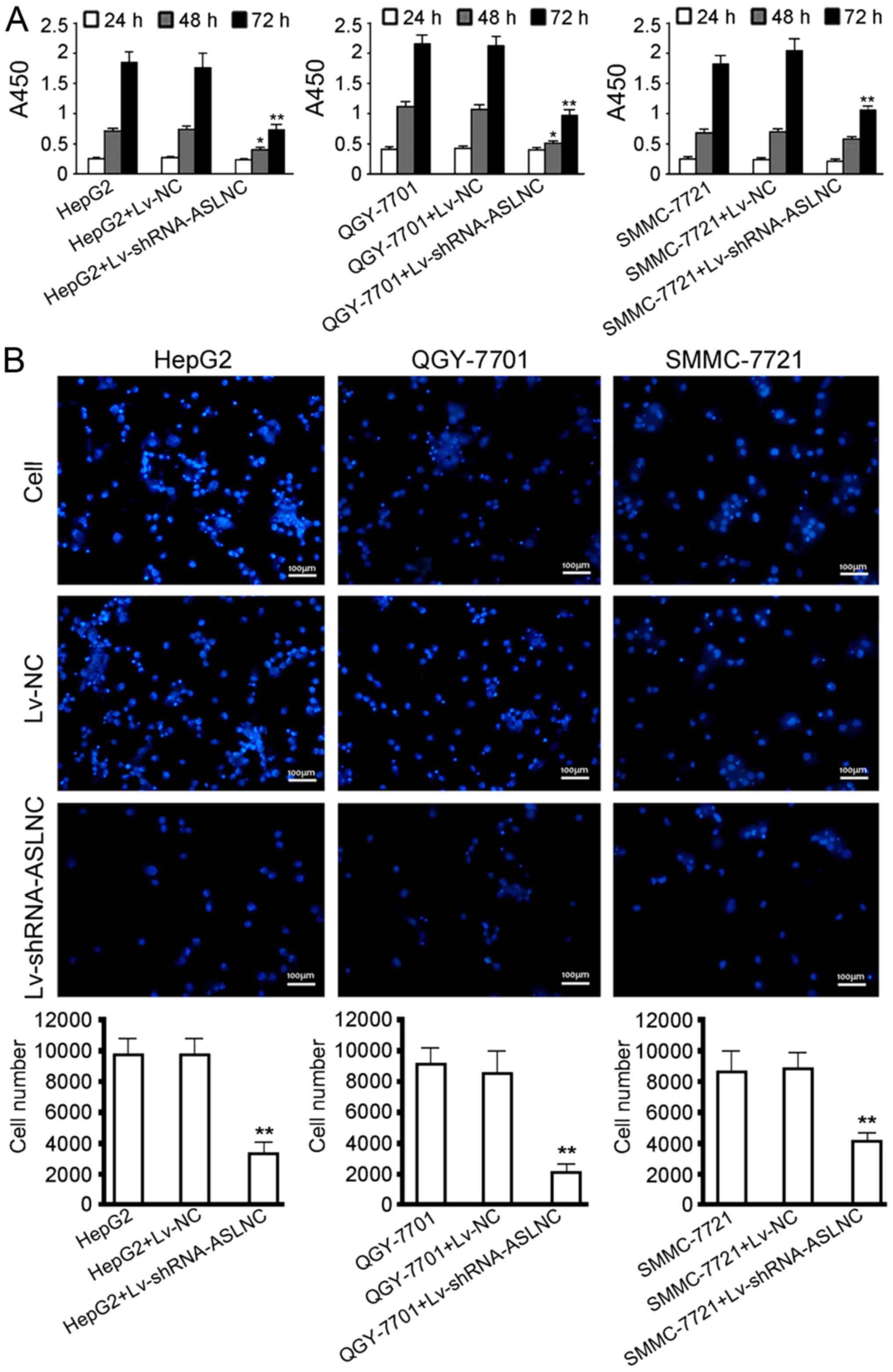
As suggested by cell proliferation assays (Fig. 4A), ASLNC02525 depletion effectively
inhibited the proliferation of HepG2, QGY-7701 and SMMC-7721 cells
(P<0.01) 72 h after the infection. The invasion assays (Fig. 4B) suggest that ASLNC02525 silencing
significantly reduced the invasion of these three cell lines,
reflected by decrease in the numbers of cells which passed through
the basement membrane (P<0.05).
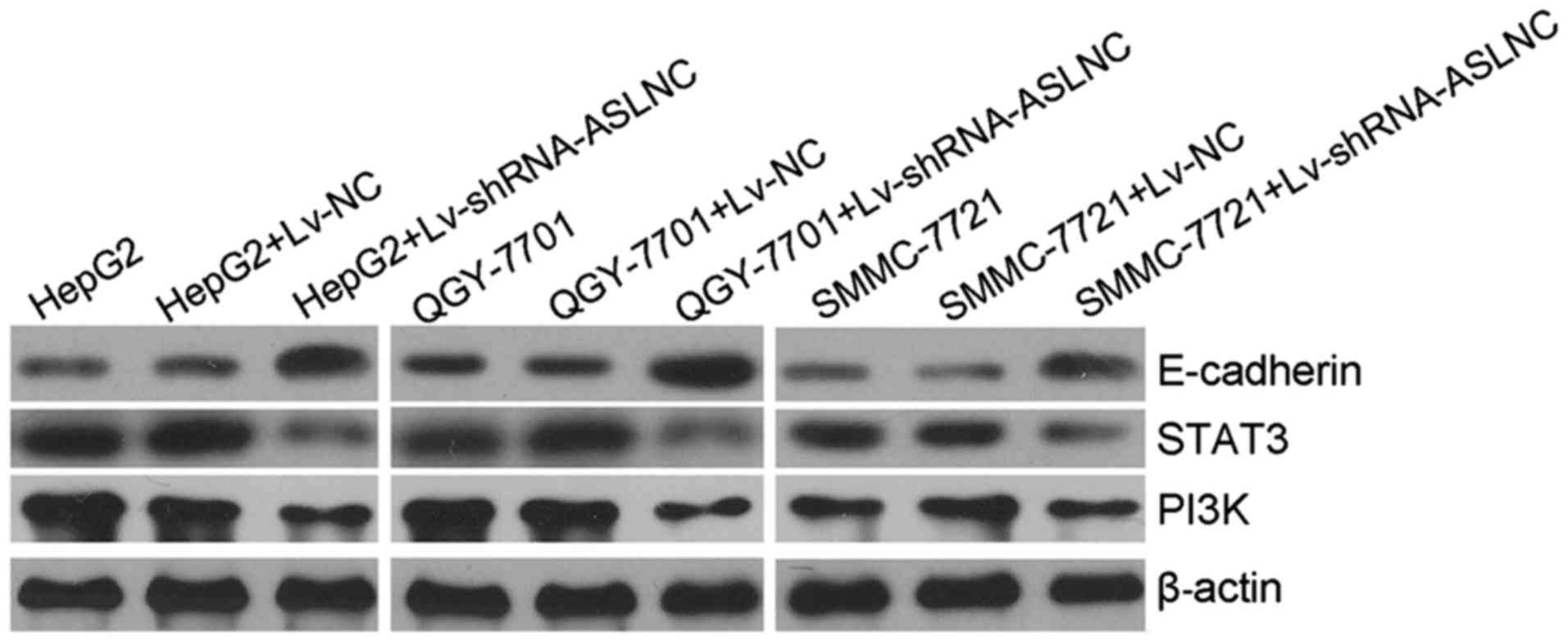
Effects of ASLNC02525 silencing on twist1
and its downstream proteins in liver cancer cells
We also assessed STAT3, PI3K and E-cadherin in the
ASLNC02525-depleted cells. The results (Fig. 5) showed that STAT3 and PI3K were
decreased and E-cadherin was increased by ASLNC02525 silencing but
not the Lv-NC. Decrease in STAT3 and PI3K is detrimental to cancer
and the increase in E-cadherin can inhibit epithelial-mesenchymal
transition (EMT) and reduce invasion of cancer cells.
Discussion
lncRNAs, endogenous RNAs longer than 200 nucleotides
discovered recently and attracting much interest, have been found
to have functions associated with various biological processes via
epigenetic modification. Increasingly major studies suggest
tangible involvement of lncRNA in cancer. Analysis of chromatin
reveals that there are over 1000 lncRNAs in mammals, which are less
conserved than the coding RNAs. As an important type of regulators,
lncRNAs exert their functions through a variety of ways. Although
they were firstly regarded as by-products by RNA polymerase II, or
transcriptional noise, recent studies have shown that lncRNAs are
associated with multiple biological processes such as chromosome
silencing, chromatin modification and transcriptional regulation
(17,18).
The interaction between lncRNAs and miRNAs has an
important influence on the onset and development of cancer
(19). miRNAs are able to regulate
lncRNAs in a targeted way: a study has shown that miR-21 targets
lncRNA GAS5 in addition to protein coding genes (20). lncRNAs can also affect the onset
and the development of cancer by regulating expression of miRNAs.
According to existing studies, lncRNAs regulate miRNAs through
three ways: i) to competitively combine to 3′-UTR of mRNAs with
miRNAs, so as to inhibit negative regulation by miRNAs. Pang et
al (21) found that an
anti-sense RNA complementary to BACE1 can complement to BACE1 mRNA,
competitively inhibiting the negative regulation of BACE1 by miRNA;
ii) to regulate target genes by forming pre-miRNAs after RNA
splicing and producing specific miRNAs; and iii) to act as
endogenous miRNA sponge to suppress miRNA function, so as to affect
malignant biological behavior of cancer cells (22). Abnormal lncRNA expression is found
in diversified diseases, and lncRNA expression profiles in a
variety of tumors are different from normal cells. The proportion
of lncRNAs in the total transcripts of genome is far larger than
that of encoding RNAs. In addition, lncRNAs play crucial roles in
the regulatory network though their interaction with DNA, RNA and
proteins. In addition to gene expression regulation, lncRNAs are
closely related to evolution of species, embryonic development,
metabolism and tumorigenesis. The evidence on involvement of
lncRNAs in diseases including cancers will provide basis and target
for diagnosis and treatment of diseases. Huang et al
(23) found lncRNA-Dreh can
inhibit hepatocellular carcinoma metastasis. Gabory et al
(24) demonstrated that lncRNA H19
is closely associated with the onset of tumors. Dysregulated
expression of lncRNA-MALAT1 is also found to link to cancer.
Moreover, it is shown that lncRNAs are related to drug resistance
of liver cancers: CUDR lnRNA, 2.2 kb in length, located at 19p13.1,
can down-regulate caspase-3 and thus, promote growth, so as to
induce drug resistance (25,26).
Human twist gene is on chromosome 7, encoding a
transcription factor. It was first identified in Drosophila
in 1983, and found to be highly conservative between species: Mouse
twist has a 96% amino acid sequence homology to human twist, and
their DNA binding domains are 100% sequence conserved (4). Twist regulates gene transcription by
binding to E-box sequence on gene promoters (5). By inhibiting apoptosis,
downregulating p53, a key tumor suppressor gene, and promoting
angiogenesis, it can promote cancer development. Overexpression of
twist results in resistance to apoptosis in cancer cells and
increasing expression of twist in implanted MCF-7 breast cancer in
mice increased blood flow and vascular permeability (27). Twist also plays a pivotal role in
EMT, which is the onset of migration and invasion of a host of
cancers. Yang et al (28)
argued that as a functional basic helix-loop-helix transcription
factor, twist may bind to other helix-loop-helix proteins to form
dimers, which in turn activate or inhibit downstream targets, so as
to regulate E-cadherin. Therefore, upregulation of twist inhibits
E-cadherin, and induces EMT, so as to promote migration and
invasion of cancer cells. Furthermore, high expression of twist is
also linked to multidrug resistance in cancer cells (29,30).
We screened for differential lncRNAs in several
pairs of selected hepatocellular carcinoma and adjacent tissue by
using lncRNA chips. ASLNC02525, highly expressed in liver cancers,
came into view. We next examined ASLNC02525 in liver cancer tissues
and three hepatocellular carcinoma cell lines by fluorescent
quantitation, and the results were consistent with the screening,
that is, expression of ASLNC02525 in cancer tissues and cells was
higher than in para-carcinoma tissue and normal hepatic epithelial
cells. We knocked down ASLNC02525 in HepG2, QGY-7701 and SMMC-7721
cells by RNAi, and the results suggest that ASLNC02525 knockdown
inhibited the proliferation in these cells. We then sought to
explore what protein mediates the effects of ASLNC02525 on
proliferation and invasion in these cells. Thus, we subjected
normal and ASLNC02525 depleted cells to differential protein
screening by protein chips, and found twist1 (data not shown). The
silencing of ASLNC02525 in liver cancer cells showed a positive
correlation between twist1 expression and ASLNC02525 knockdown. We
proceeded to investigate whether ASLNC02525 depletion downregulates
twist1 expression or suppresses its activity to inhibit cancer by
RIP-Seq experiment (data not shown), and no twist1 mRNA was found
in the DNA sequence binding to twist1. The result that protein
expression but not transcription of twist1 was affected by
ASLNC02525 implied that there may be a post-transcription
regulation of twist1 changed by ASLNC02525 silencing. Since miRNAs
are common post-transcription regulation factor, we supposed that
abnormal expression of ASLNC02525 may break some regulation of
twist1 by a miRNA.
lncRNAs, as a competing endogenous RNA, interact
with miRNAs to regulate target genes, so as to play important roles
in the onset and development of cancers (31,32).
Therefore, we searched for potential miRNAs which may bind to both
ASLNC02525 and twist1 3′UTR, and found hsa-miRNA489-3p, which has,
respectively, four and one predictive binding site on ASLNC02525
and twist1 3′UTR. The hypothesis surfaced that high expression of
ASLNC02525 impairs the regulation of twist1 by hsa-miRNA-489-3p
like an RNA sponge, and results in cellular transformation, and the
depletion of ASLNC02525 recovers the negative regulation, so as to
suppress twist1, as well as cancer activity.
The experimental results proved our hypothesis: both
tissues and cells of liver cancer exhibited high levels of
ASLNC02525 and hsa-miRNA-489-3p, and silencing ASLNC02525 inhibited
proliferation and invasion of liver cancer cells, and reduced
twist1 protein, but no significant changes in hsa-miRNA-489-3p was
observed, indicating twist1 expression inhibition may mediate the
suppression of liver cancer cells by ASLNC02525 silencing. The
luciferase reporter assay showed that hsa-miRNA-489-3p bound to
twist1 3′UTR via its seed site and inhibited protein translation,
which was impaired by ASLNC02525 overexpression. In addition,
knockdown ASLNC02525 recovered the negative regulation of twist1 by
hsa-miRNA-489-3p. Although we proposed a preliminary explanation
for suppression of liver cancer by ASLNC02525 depletion, it
requires profound future study. In general, special structure is
required for competitive binding of lncRNA, so our next step is to
establish the structure of ASLNC02525 and the binding of its four
predicted sites to hsa-miRNA-489-3p, as well as whether the
inhibition is prevalent.
The importance of the study lies in providing some
insight into the potential for lncRNA as a target for gene therapy,
by demonstrating that lncRNA can suppress tumor inhibiting activity
of miRNAs via interrupting the regulation of some miRNA target
genes.
Acknowledgments
This work was supported by the National Clinical Key
Subject, the National Natural Science Foundation of China (grant
nos. 81072070, 81471241, 81171133, and 81271375) and the Natural
Science Foundation of Shanghai (grant no. 15ZR1405600).
References
|
1
|
Abdelmohsen K and Gorospe M: Noncoding RNA
control of cellular senescence. Wiley Interdiscip Rev RNA.
6:615–629. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Alaei-Mahabadi B and Larsson E: Limited
evidence for evolutionarily conserved targeting of long non-coding
RNAs by microRNAs. Silence. 4:42013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Schmitt AM and Chang HY: Long noncoding
RNAs in cancer pathways. Cancer Cell. 29:452–463. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Firulli AB and Conway SJ:
Phosphoregulation of Twist1 provides a mechanism of cell fate
control. Curr Med Chem. 15:2641–2647. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Atfi A and Baron R: p53 brings a new twist
to the Smad signaling network. Sci Signal. 1:pe332008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Barsky D and Venclovas C: DNA sliding
clamps: Just the right twist to load onto DNA. Curr Biol.
15:R989–R992. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bohlius J, Tonia T and Schwarzer G: Twist
and shout: One decade of meta-analyses of
erythropoiesis-stimulating agents in cancer patients. Acta
Haematol. 125:55–67. 2011. View Article : Google Scholar
|
|
8
|
Ai L, Kim WJ, Alpay M, Tang M, Pardo CE,
Hatakeyama S, May WS, Kladde MP, Heldermon CD, Siegel EM, et al:
TRIM29 suppresses TWIST1 and invasive breast cancer behavior.
Cancer Res. 74:4875–4887. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Allaya N, Khabir A, Sallemi-Boudawara T,
Sellami N, Daoud J, Ghorbel A, Frikha M, Gargouri A,
Mokdad-Gargouri R and Ayadi W: Over-expression of miR-10b in NPC
patients: Correlation with LMP1 and Twist1. Tumour Biol.
36:3807–3814. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ansieau S, Bastid J, Doreau A, Morel AP,
Bouchet BP, Thomas C, Fauvet F, Puisieux I, Doglioni C, Piccinin S,
et al: Induction of EMT by twist proteins as a collateral effect of
tumor-promoting inactivation of premature senescence. Cancer Cell.
14:79–89. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Fu J, Qin L, He T, Qin J, Hong J, Wong J,
Liao L and Xu J: The TWIST/Mi2/NuRD protein complex and its
essential role in cancer metastasis. Cell Res. 21:275–289. 2011.
View Article : Google Scholar
|
|
12
|
Grzegrzolka J, Biala M, Wojtyra P,
Kobierzycki C, Olbromski M, Gomulkiewicz A, Piotrowska A, Rys J,
Podhorska-Okolow M and Dziegiel P: Expression of EMT markers SLUG
and TWIST in breast cancer. Anticancer Res. 35:3961–3968.
2015.PubMed/NCBI
|
|
13
|
Ji H, Lu HW, Li YM, Lu L, Wang JL, Zhang
YF and Shang H: Twist promotes invasion and cisplatin resistance in
pancreatic cancer cells through growth differentiation factor 15.
Mol Med Rep. 12:3841–3848. 2015.PubMed/NCBI
|
|
14
|
Zhang H, Gong J, Kong D and Liu HY:
Anti-proliferation effects of Twist gene silencing in gastric
cancer SGC7901 cells. World J Gastroenterol. 21:2926–2936. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Kim K, Park EY, Yoon MS, Suh DS, Kim KH,
Lee JH, Shin DH, Kim JY, Sol MY and Choi KU: The role of TWIST in
ovarian epithelial cancers. Korean J Pathol. 48:283–291. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Han Z, Liu L, Liu Y and Li S: Sirtuin
SIRT6 suppresses cell proliferation through inhibition of Twist1
expression in non-small cell lung cancer. Int J Clin Exp Pathol.
7:4774–4781. 2014.PubMed/NCBI
|
|
17
|
Ponting CP, Oliver PL and Reik W:
Evolution and functions of long noncoding RNAs. Cell. 136:629–641.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Bergmann JH and Spector DL: Long
non-coding RNAs: Modulators of nuclear structure and function. Curr
Opin Cell Biol. 26:10–18. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Deng G and Sui G: Noncoding RNA in
oncogenesis: A new era of identifying key players. Int J Mol Sci.
14:18319–18349. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Pickard MR and Williams GT: Molecular and
cellular mechanisms of action of tumour suppressor GAS5 lncRNA.
Genes (Basel). 6:484–499. 2015. View Article : Google Scholar
|
|
21
|
Pang M, Xing C, Adams N, Rodriguez-Uribe
L, Hughs SE, Hanson SF and Zhang J: Comparative expression of miRNA
genes and miRNA-based AFLP marker analysis in cultivated tetraploid
cottons. J Plant Physiol. 168:824–830. 2011. View Article : Google Scholar
|
|
22
|
Brodersen P, Sakvarelidze-Achard L,
Schaller H, Khafif M, Schott G, Bendahmane A and Voinnet O:
Isoprenoid biosynthesis is required for miRNA function and affects
membrane association of ARGONAUTE 1 in Arabidopsis. Proc Natl Acad
Sci USA. 109:1778–1783. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Huang JF, Guo YJ, Zhao CX, Yuan SX, Wang
Y, Tang GN, Zhou WP and Sun SH: Hepatitis B virus X protein
(HBx)-related long noncoding RNA (lncRNA) down-regulated expression
by HBx (Dreh) inhibits hepatocellular carcinoma metastasis by
targeting the intermediate filament protein vimentin. Hepatology.
57:1882–1892. 2013. View Article : Google Scholar
|
|
24
|
Gabory A, Ripoche MA, Le Digarcher A,
Watrin F, Ziyyat A, Forné T, Jammes H, Ainscough JF, Surani MA,
Journot L, et al: H19 acts as a trans regulator of the imprinted
gene network controlling growth in mice. Development.
136:3413–3421. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lai MC, Yang Z, Zhou L, Zhu QQ, Xie HY,
Zhang F, Wu LM, Chen LM and Zheng SS: Long non-coding RNA MALAT-1
overexpression predicts tumor recurrence of hepatocellular
carcinoma after liver transplantation. Med Oncol. 29:1810–1816.
2012. View Article : Google Scholar
|
|
26
|
Tsang WP, Wong TW, Cheung AH, Co CN and
Kwok TT: Induction of drug resistance and transformation in human
cancer cells by the noncoding RNA CUDR. RNA. 13:890–898. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Mironchik Y, Winnard PT Jr, Vesuna F, Kato
Y, Wildes F, Pathak AP, Kominsky S, Artemov D, Bhujwalla Z, Van
Diest P, et al: Twist overexpression induces in vivo angiogenesis
and correlates with chromosomal instability in breast cancer.
Cancer Res. 65:10801–10809. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yang J, Mani SA, Donaher JL, Ramaswamy S,
Itzykson RA, Come C, Savagner P, Gitelman I, Richardson A and
Weinberg RA: Twist, a master regulator of morphogenesis, plays an
essential role in tumor metastasis. Cell. 117:927–939. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Al Saleh S1, Sharaf LH and Luqmani YA:
Signalling pathways involved in endocrine resistance in breast
cancer and associations with epithelial to mesenchymal transition
(Review). Int J Oncol. 38:1197–1217. 2011.PubMed/NCBI
|
|
30
|
Chen HF, Huang CH, Liu CJ, Hung JJ, Hsu
CC, Teng SC and Wu KJ: Twist1 induces endothelial differentiation
of tumour cells through the Jagged1-KLF4 axis. Nat Commun.
5:46972014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chen L, Wang W, Cao L, Li Z3 and Wang X:
Long non-coding RNA CCAT1 acts as a competing endogenous RNA to
regulate cell growth and differentiation in acute myeloid leukemia.
Mol Cells. 39:330–336. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Karreth FA and Pandolfi PP: ceRNA
cross-talk in cancer: When ce-bling rivalries go awry. Cancer
Discov. 3:1113–1121. 2013. View Article : Google Scholar : PubMed/NCBI
|