Introduction
Lung cancer is the leading cause of cancer-related
mortality worldwide, with a higher rate of mortality each year
compared with colon, breast and prostate cancers (1-4).
Targeted therapy has been used for the clinical treatment of lung
cancer and therapies targeting epidermal growth factor receptor
(EGFR) or vascular endothelial growth factor (VEGF) have been shown
to be effective, as well as traditional chemotherapy (5-8).
Cyclooxygenase (COX), an essential enzyme in prostaglandin (PG)
synthesis, represents another potential target. PGs are involved in
various physiological and pathogenic processes, such as
inflammatory reactions, gastro-intestinal cytoprotection, and
ulceration, hemostasis and thrombosis (9-15).
The COX enzyme has two isoforms, COX-1 and COX-2. COX-1 is
constitutively expressed in most cells and tissues, and produces
PGs involved in the maintenance of the gastric mucosa, the
regulation of renal blood flow and platelet aggregation. By
contrast, COX-2 can be induced by various pro-inflammatory and
mitogenic stimuli (9-11).
The inhibition of COX-1 causes side-effects, such as
gastrointestinal tract bleeding and kidney failure (13,16-18).
By contrast, COX-2 inhibitors are widely used as they have few
side-effects. Celecoxib, a COX-2 inhibitor, induces cell cycle
arrest, inhibits tumor growth, suppresses tumor angiogenesis and
induces apoptosis in several types of cancer. Such responses to
celecoxib have been investigated in cells that highly express COX-2
(15,18,19).
However, Ramer et al reported that the expression of COX-2
was paradoxically increased by celecoxib, which activated
peroxisome proliferator-activated receptor (PPAR)γ (20). The increased COX-2 expression
induced by celecoxib may modulate local inflammatory responses and
COX-2 may thus be a potential target of sustained celecoxib
treatment.
Exosomes, small vesicles composed of a lipid bilayer
membrane, are found in most biological fluids (21-23).
Several characteristics of exosomes have been explored, including
their small size (50-100 nm), cup-shaped morphology and specific
protein markers (24-27). The major role of exosomes is the
delivery of bioactive molecules between cells (21,28).
Recently, exosomes have been tested in the treatment of cancers.
Cancer-derived exosomes can be used as biomarkers for cancer
diagnosis as they are enriched with proteins, mRNAs and miRNAs
related to cancer (21,24,25,28).
Of note, in this study, we found that celecoxib increased COX-2
expression in both the cytoplasm and exosomes in lung cancer
cells.
In the present study, we investigated whether
adjacent cells may acquire exosomes containing COX-2 from lung
cancer cells following treatment with celecoxib, focusing on the
intercellular responses of cells neighboring lung cancers related
to exosome transport.
Materials and methods
Cell culture and celecoxib treatment
The human lung cancer carcinoma cell lines, A549,
H460 and 11-18, the human monocytic cell line, THP-1, the Burkitt's
lymphoma B cell line, Raji, human primary umbilical vein
endothelial cells (HUVECs) and the human umbilical vein cell line,
EA.hy926, were all purchased from the American Type Culture
Collection (ATCC, Manassas, VA, USA). All the cells, apart from the
HUVECs and EA.hy926 cells were maintained in RPMI-1640 medium
(HyClone, Logan, UT, USA). The HUVECs were maintained with F-12K
medium containing endothelial growth factors and the EA.hy926 cells
was maintained in DMEM supplemented with 10% fetal bovine serum
(FBS), with 100 U/ml penicillin and 100 µg/ml streptomycin
(all from HyClone) at 37°C in a humidified 5% CO2
atmosphere.
To determine the optimal concentration, the cells
were incubated with celecoxib (cat. no. C251000; Toronto Research
Chemicals, Toronto, ON, Canada) at 0, 25, 50, 75 or 100 µM
for 16 h, and 75 µM for 0, 2, 4, 6, 8 or 12 h. To perform
further experiment with minimally affected cell growth inhibition,
75 µM of celecoxib was treated in lung cancer cells for 16
h.
Monocyte preparation
Peripheral blood mononuclear cells (PBMCs) from
human blood were isolated by Ficoll-Paque (Amersham Life Science,
Buckinghamshire, England) density-gradient centrifugation. Monocyte
purification was achieved using a monocyte isolation kit (Miltenyi
Biothec, Bergisch Gladhach, Germany). PBMCs were combined with FcR
blocking reagent and biotin-antibody cocktail and incubated for 10
min at 4°C. Subsequenlty, anti-biotin MicroBeads were added and
incubated for 15 min at 4°C. The cells were centrifuged at 300 × g,
4°C for 10 min after washing and resuspended with buffer. The cell
suspension was loaded into columns and rinsed with buffer. The
cells that passed through were unlabeled cells, representing the
enriched monocyte fraction. This study was approved by the
Institutional Bioethics Review Board at the College of Medicine,
Inje University, Busan, Korea and all donors provided informed
consent for the use of their samples in this study.
Analysis of cytotoxicity
Cell viability was examined by WST-1 assay (Takara,
Shiga, Japan). The cells were seeded in 96-well plates
(5×103 cells/well) and incubated for 24 h. The cells
were treated with various concentrations of exosomes overnight.
WST-1 then was added to each well, and the plates were incubated
for 1 h and the absorbance was measured using a microplate reader
(Multiskan FC model; Thermo Scientific, Waltham, MA, USA) at a
wavelength of 450 nm.
RT-PCR
Total RNA was isolated using TRIzol reagent
(Invitrogen, Carlsbad, CA, USA) according to the manufacturer's
instructions. Briefly, the harvested cells were mixed with TRIzol
reagent and chloroform. The mixture was centrifuged at 12,000 × g
for 15 min at 4°C. The supernatant was discarded and washed with
75% ethanol in DEPC-treated water. The pellet containing RNA was
resuspended with DEPC-treated water. RT-premix (Bioneer, Daejeon,
Korea) was used for cDNA synthesis from 2 µg of total RNA.
The mixture of RNA and oligo(dTs) was combined with RT-premix and
incubated for 1 h at 42°C and 5 min at 95°C. cDNA was amplified
using primers the following primers: COX-2 forward, GCT CCT ACC CAC
GCA GAT TT and reverse, AGA CGC CAT TTG GAT TGG GT; and β-actin
forward, ATC CAC GAA ACT ACC TTC AA and reverse, ATC CAC ACG GAG
TAC TTG C and Tenuto PCR premix (Enzynomics, Daejeon, Korea) in a
PCR Thermal Cycler (Takara). PCR products were analyzed by gel
electrophoresis in TAE buffer through 1% agarose gels. The
densities of bands were analyzed using ImageJ 1.51K software
[National Institutes of Health (NIH), Bethesda, MD, USA].
Western blot analysis
The cells were harvested, rinsed with
phosphate-buffered saline (PBS) (Welgene, Gyeongsan, Korea) and
lysed in lysis buffer (Invitrogen) supplemented with protease
inhibitor cocktail (Sigma, St. Louis, MO, USA). The cells were
incubated on ice for 10 min and centrifuged for 10 min at 15,000 ×
g. Protein quantity was determined by BCA assay (Thermo Scientific)
and then separated on 10% sodium dodecyl sulfate
(SDS)-polyacrylamide gels overlaid with a 4% polyacrylamide
stacking gel. The proteins were transferred onto a nitrocellulose
membrane after separation. The membranes were blocked with 5% skim
milk for 1 h at room temperature and washed 3 times with
Tris-buffered saline containing 0.1% Tween-20 (TBST, pH 7.4) for 10
min each. Each membrane was incubated with primary antibodies to
COX-2 (1:1,000 dilution, cat. no. sc-12282; Cell Signaling
Technology, Beverly, MA, USA), CD63 (1:500 dilution, cat. no.
sc-15363) or β-actin (1:2,000 dilution, cat. no. sc-47778) (both
from Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA) in 0.1%
TBST at 4°C overnight. The membranes were then washed with 0.1%
TBST 3 times for 10 min and incubated with secondary antibody
[anti-rabbit IgG [1:5,000 dilution, cat. no. 7074; Cell Signaling
Technology) for COX-2 and CD63; anti-mouse IgG (1:5,000 dilution,
cat. no. 7076; Cell Signaling Technology) for β-actin] for 1 h at
room temperature. Following incubation, each membrane was washed
with 0.1% TBST 3 times. β-actin or CD63 was detected as an internal
standard or exosome marker, respectively. Signals were detected by
chemiluminescence (Fuji Photo Film Co., Tokyo, Japan). The
densities of the bands were analyzed using ImageJ 1.51K software
(NIH).
Isolation of exosomes and culture with
exosomes
Exosomes were isolated using an Exospin exosome
purification kit (Cell Guidance Systems, St. Louis, MO, USA)
according to the manufacturer's instructions. The H460 and 11-18
cells were cultured in 150-mm dishes and cell culture supernatant
was collected by centrifugation at 300 × g for 10 min. The
supernatant was transferred into new tubes and centrifuged at
20,000 × g at 4°C for 30 min in a high-speed centrifuge. The
collected supernatant was transferred into new tubes and buffer A
(provided with the kit) was added. The mixture was incubated at 4°C
for overnight after mixing well. The following day, the mixture was
centrifuged at 20,000 × g at 4°C for 1 h. Exosome-containing
pellets were resuspended with PBS following the aspiration of the
supernatant. Spin columns were prepared and spun down at 50 × g for
30 sec. Pellets containing exosomes were loaded into the tops of
columns and centrifuged at 50 × g for 60 sec. The columns were
placed into 1.5-ml tube and rinsed with PBS or medium applied at
the top and centrifuged at 50 × g for 60 sec.
The purified exosomes were used to examine cell
viability, the transport of COX-2, and the production of cytokines
in cells uptaking exosomes. To examine cell viability, various
concentration (100, 250 or 500 µg/ml) of exosomes were
incubated with the THP-1 cells for 24 h. For the measurements of
the COX-2 concentration and the production of cytokines, exosomes
(75 or 150 µg/ml) were used to treat the THP-1, human
primary monocytes, HUVECs, EA.hy926 or Raji cells for 16 h.
Transmission electron microscopy
Exosomes were purified using an Exospin kit and
resuspended in a small volume of PBS. Exosomes were fixed with 2.5%
glutaraldehyde solution, washed and post-fixed in 1% osmium
tetroxide. The pellets were then dehydrated through an ethanol
series followed by acetone, and then embedded in epoxy resin.
Ultrathin sections were contrasted with uranyl acetate and lead
citrate. Transmission electron microscopy JEM-1200 EX II (Jeol,
Tokyo, Japan) was used to view these sections at 80 kV.
Enzyme immunoassay (EIA) and
enzyme-linked immunosorbent assay (ELISA)
To measure the concentrations of PGE2 and
VEGF, cell culture supernatants were examined using an EIA kit
(Cayman Chemical Co., Ann Arbor, MI, USA) and an ELISA kit (R&D
Systems, Minneapolis, MN, USA) according to the instructions
provided by the respective manufacturers. The kit contains the
required solutions and all antibodies, and concentrations and
dilution ratios followed the instructions. Briefly, PGE2
standard solution and culture supernatants were added to plates
with goat polyclonal anti-mouse IgG. PGE2 AChE tracer
and PGE2 monoclonal antibodies (provided with the kit)
were added followed by incubation for 18 h at 4°C. Ellman's reagent
(provided with the kit) was added to the wells after rinsing 5
times with washing buffer and the plates were incubated for 1 h.
The absorbance was measured with a microplate reader (Multiskan FC
model, Thermo Scientific) at 405 nm.
To coat the plates, VEGF capture antibody (provided
with the kit) was added to the wells followed by incubation
overnight. Each plate was then washed and blocked for 1 h. Standard
solutions and culture supernatants were added after washing. VEGF
detection antibody (provided with the kit) was then added followed
by incubation for 2 h, followed by the addition of streptavidin-HRP
and incubation for 20 min in indirect light and washing of each
plate three times. Substrate solutions were added followed by
incubation for 20 min in indirect light. Stop solution was added
and the optical density of each well was determined using a
microplate reader (Multiskan FC model; Thermo Scientific) set to
450 nm.
Tube formation assay
Tube formation assay was conducted in 96-well plates
coated with 50 µl of growth factor-reduced Matrigel (BD
Biosciences, San Jose, CA, USA) according to the manufacturer's
instructions. The EA.hy926 cells (1.0×104 cells/well)
were treated with filtered THP-1 supernatant (following incubation
with exosomes carrying high concentrations of COX-2) or control
medium for an additional 16 h. Tube formation was photographed
under a microscope (iRIS; Logos Biosystems, Anyang, Korea). The
numbers of tubes in an identical area were manually counted and
statistically analyzed using the Student's t-test. A P-value
<0.05 was considered to indicate a statistically significant
difference.
Statistical analysis
The densities of all the bands were measured using
ImageJ 1.51K software (NIH), and the relative ratio to the internal
standard was calculated. Two-sided Student's t-tests were used to
evaluate differences between 2 groups. For comparison of
viabilities between more than 2 groups, data were analyzed using
one-way ANOVA (with Tukey's multiple comparison test). The data
from ELISA and tube formation data are presented as the means ± SD.
A P-value <0.05 was considered to indicate a statistically
significant difference.
Results
Celecoxib paradoxically increases COX-2
expression in lung cancer
Cytotoxicity was examined when the lung cancer cell
lines were treated with celecoxib. The A549, H460 and 11-18 lung
cancer cell lines were incubated with various concentrations of
celecoxib. The concentration of celecoxib leading to the 50%
suppression of cell viability was estimated to be ~50 µM in
the H460 and 11-18 cells, but >100 µM in the A549 cells.
We observed that higher concentrations of celecoxib led to a
greater decrease in the viability of the lung cancer cells (data
not shown).
To observe the effects of celecoxib on COX-2
expression in lung cancer cells following treatment with celecoxib
at various concentrations and treatment durations, the 3 lung
cancer cell lines, A549, H460 and 11-18, were treated with 25, 50,
75 or 100 µM of celecoxib for 6 h. COX-2 expression was then
detected by western blot analysis. The expression of COX-2 was
elevated in a dose-dependent manner (Fig. 1A), as well as in a time-dependent
manner when the cells lines were treated with 75 µM of
celecoxib for 2, 4, 6, 8 or 12 h. A marked increase in COX-2
expression was observed after 4 h (Fig. 1B). COX-2 mRNA expression was
immediately increased following treatment with celecoxib (Fig. 1C).
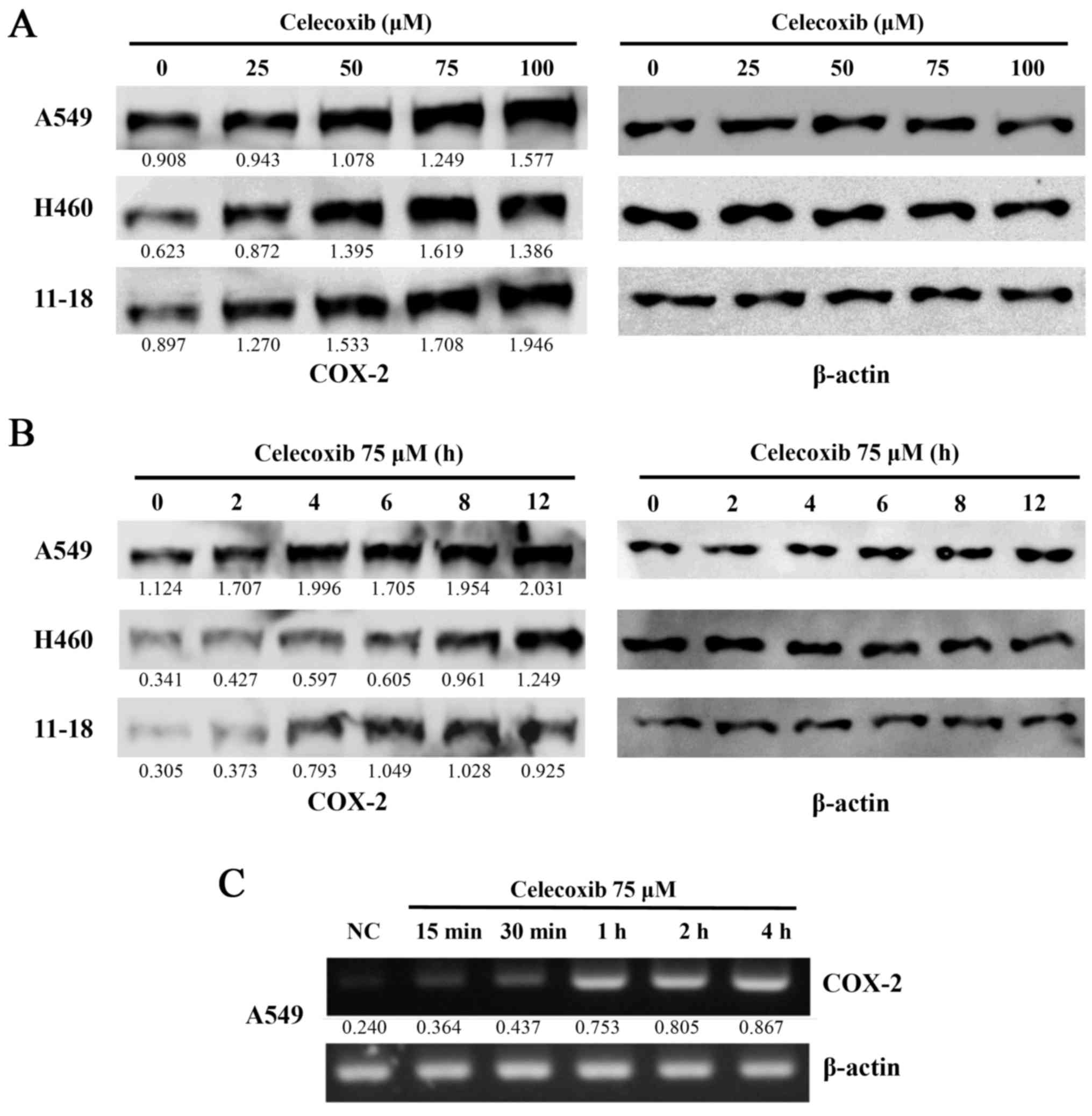 | Figure 1Cell viability and cyclooxygenase-2
(COX-2) expression following the treatment of lung cancer cell
lines with celecoxib. (A) The lung cancer cell lines, A549, H460
and 11-18, were treated with celecoxib (0, 25, 50, 75 and 100
µM) for 16 h. (B) The A549, H460 and 11-18 cells were also
treated with celecoxib at 75 µM for 0, 2, 4, 6, 8 and 12 h.
The cells were then harvested and proteins were extracted. (A and
B) COX-2 protein expression was measured by western blot analysis.
A representative example of 5 independent experiments is shown. (C)
COX-2 mRNA expression at the indicated time points was detected by
RT-PCR. The numbers under the bands indicate the ratio between
densities of marked bands and β-actin. A representative example of
3 independent experiments is shown. |
Celecoxib increases COX-2 expression in
both the cytoplasm and exosomes
The H460 cells were cultured in medium supplemented
with exosome-free FBS, and exosomes were then isolated.
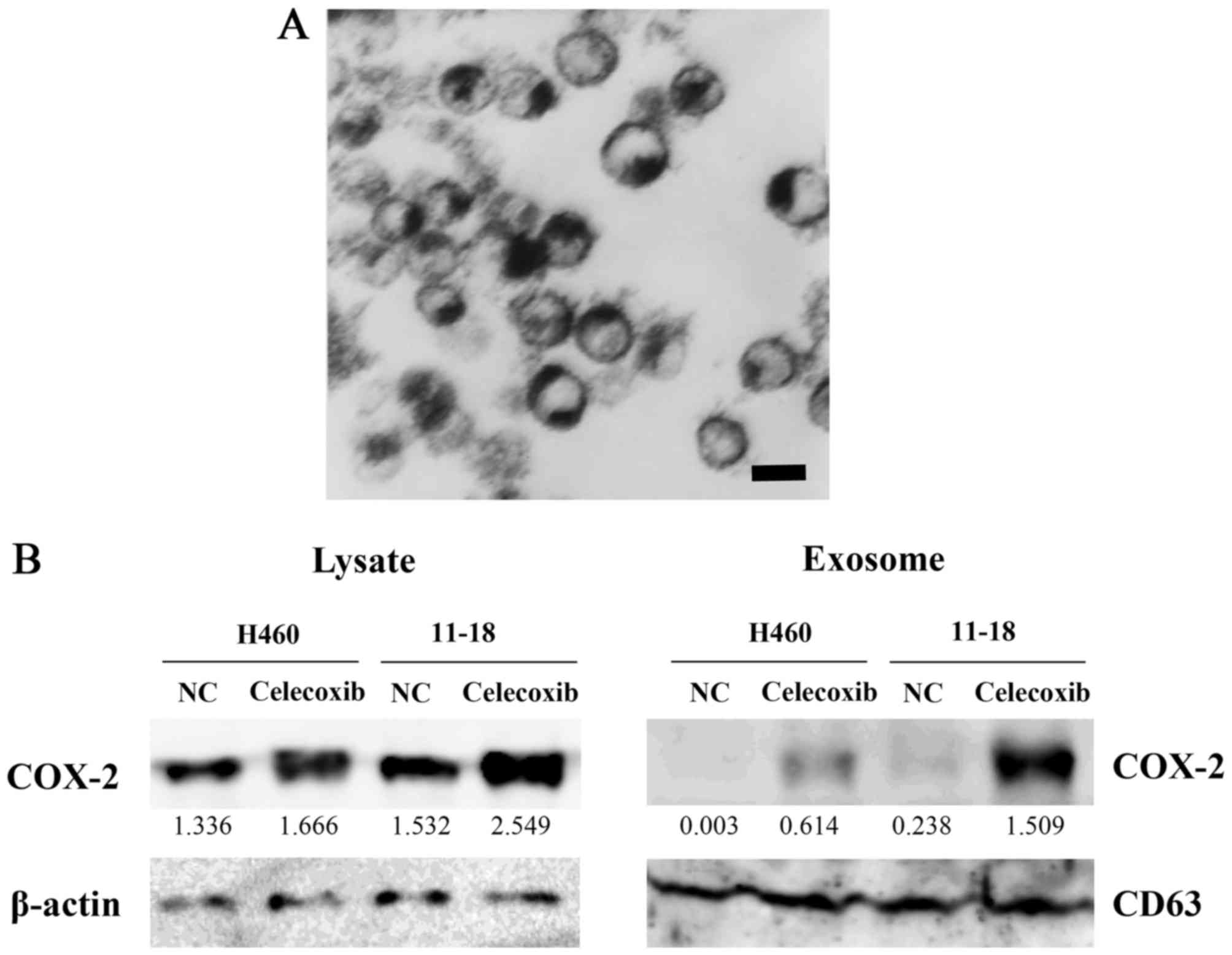
Transmission electron microscopy revealed that purified exosomes
were ~100 nm in size and had a dish-like shape (Fig. 2A).
The H460 and 11-18 cells were incubated in
exosome-free FBS with 75 µM of celecoxib for 16 h, and cell
lysates and exosomes were then prepared from cells and
supernatants, respectively. The expression level of COX-2 was
detected by western blot analysis. COX-2 expression was increased
in the cell lysates and exosomes collected from the
celecoxib-treated cells compared to the control cells. The
concentration of COX-2 in exosomes from the 11-18 cells was much
higher than that from the H460 cells. CD63 was used as an exosome
marker and β-actin was used as internal standard for cell lysates
(Fig. 2B).
COX-2 protein is transferred to THP-1
cells through exosomes from celecoxib-treated lung cancer
cells
Exosomes were isolated from celecoxib-treated and
untreated 11-18 cells. Cell viability was determined using the
monocytic leukemia cell line, THP-1, to confirm that there was no
change in the cell response by substances derived from other cells.
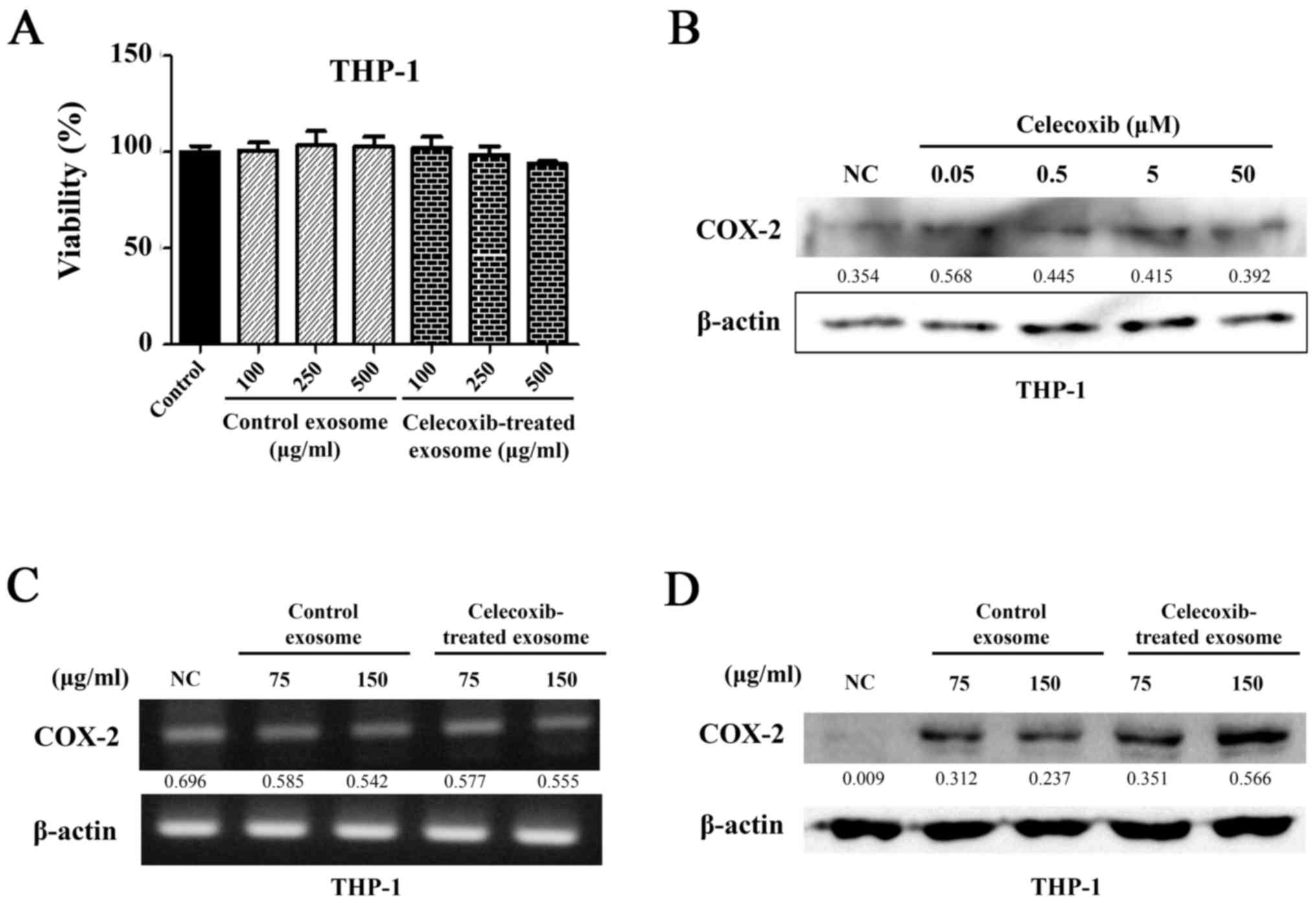
The THP-1 cells were seeded on 96-well plates and then 100, 250 and
500 µg/ml of exosomes were added. WST-1 was added following
incubation for 24 h and detected using a microplate reader. In all
wells, the suppression of cell viability was not observed following
treatment with exosomes from the control and celecoxib-treated
cells (P>0.05) (Fig. 3A).
As a small quantity of celecoxib carried in exosomes
can directly affect THP-1 cells, the THP-1 cells were treated with
celecoxib at concentrations of 0.05, 0.5, 5 and 50 µM for 16
h and the concentration of COX-2 was detected by western blot
analysis (Fig. 3B). Direct
incubation with celecoxib did not increase COX-2 protein expression
in THP-1 cells.
To determine whether exosomes can transfer COX-2 to
other cells, exosomes isolated from the supernatant of the 11-18
cells were added to THP-1 cells at 75 and 150 µg/ml.
Following incubation with exosomes, RNA was extracted from the
THP-1 cells and RT-PCR was performed. The same mRNA expression
level of COX-2 was observed in all samples (Fig. 3C). However, different protein
levels of COX-2 were observed, as shown by western blot analysis.
COX-2 expression was higher in the cells treated with exosomes from
the celecoxib-treated 11-18 cell supernatant. A greater
concentration of COX-2 was detected in the cells treated with 150
µg/ml of exosomes than in those treated with 75 µg/ml
(Fig. 3D). Thus, COX-2 protein
appeared to have been transferred to the THP-1 cells. Exosomes did
not transfer COX-2 mRNA, and did not stimulate the THP-1 cells.
COX-2 in exosomes is transferred to other
cell types, as well as THP-1
To determine whether COX-2 concentration can be
altered in other cells through exosomes, several types of cells
were treated with exosomes. Human monocytes were isolated from
human blood. PBMCs were isolated from blood by Ficoll-Plaque using
density-gradient centrifugation and monocytes were collected using
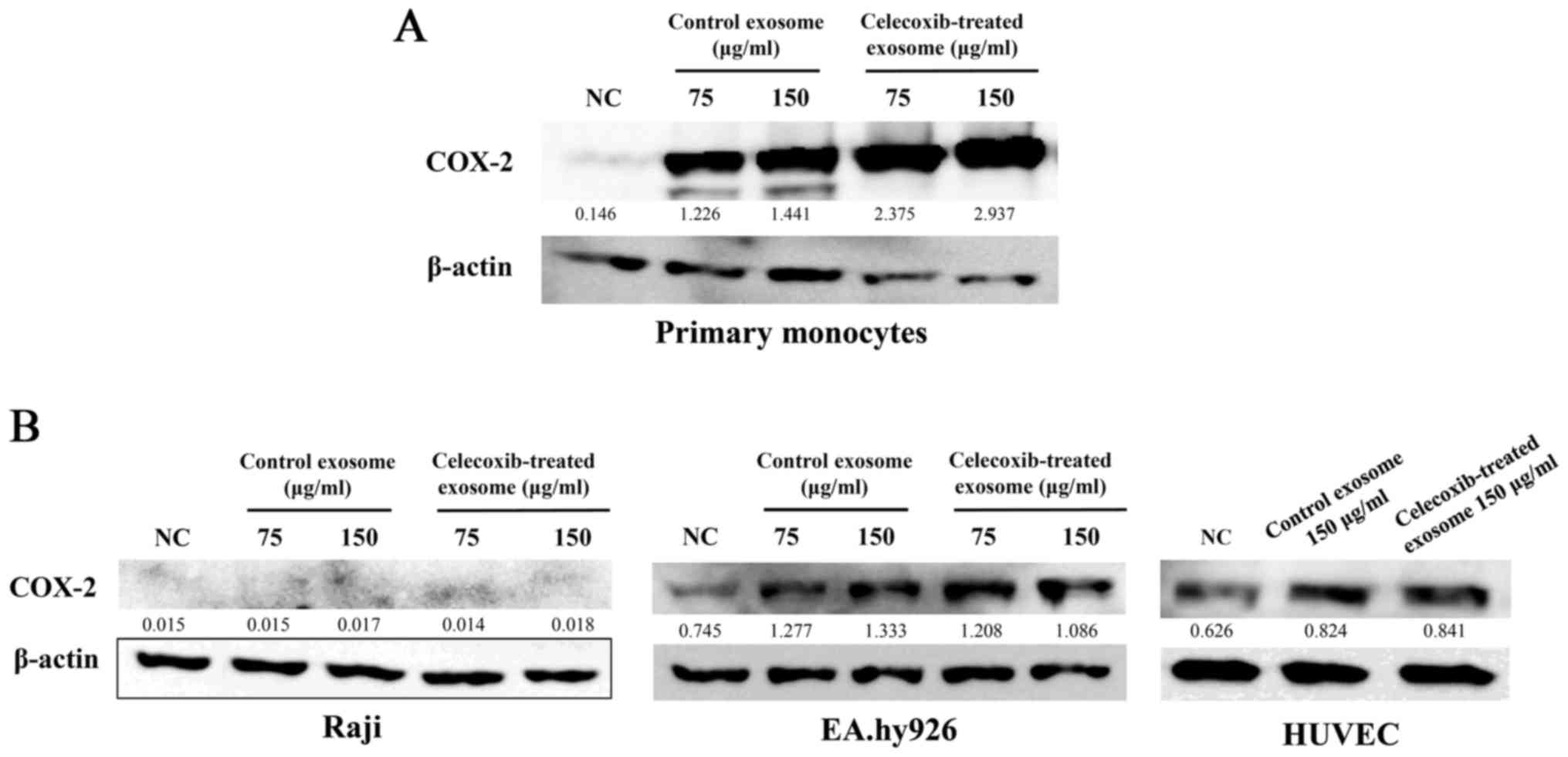
a monocyte isolation kit. Exosomes were added to human monocytes
for 16 h and COX-2 protein expression was detected by western blot
analysis. As in the monocytic leukemia cell line, THP-1, the
concentration of COX-2 increased in the human primary monocytes
following treatment with exosomes from the celecoxib-treated 11-18
cell supernatant (Fig. 4A).
To examine whether exosomes from lung cancer cells
transfer COX-2 to the same lung cancer type, exosomes from the
11-18 cells were added to the 11-18 cells. However, the
concentration of COX-2 was not altered (data not shown).
Exosomes isolated from the celecoxib-treated 11-18
cell supernatant were added to Raji cells (a Burkitt's lymphoma B
cell line), EA.hy926 (a human umbilical vein cell line) and HUVECs
(a human primary umbilical vein endothelial cell line). The
concentration of COX-2 was not higher in the Raji cells, but
differed in the EA.hy926 cells and HUVECs (Fig. 4B).
The production of PGE2 and
VEGF is increased by the transfer of COX-2 through exosomes
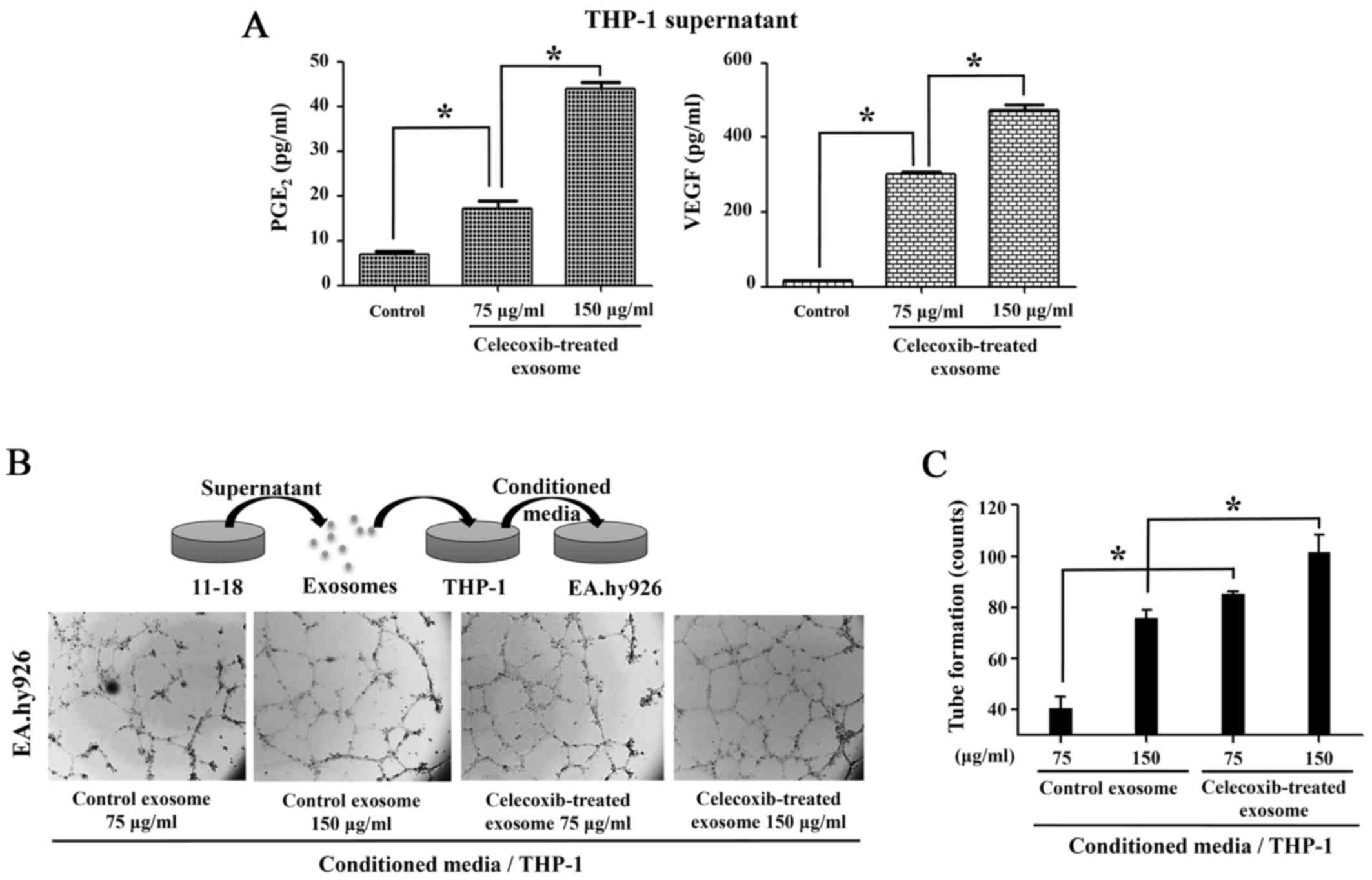
To measure the function of COX-2 transferred from
exosomes, a PGE2 EIA assay was performed. The cell
culture supernatant collected following treatment of the THP-1
cells with exosomes from celecoxib-treated 11-18 cells for 16 h
revealed that PGE2 production increased following
incubation with the exosomes (Fig.
5A). As PGE2 stimulates angiogenic VEGF production
in monocytes (29), VEGF ELISA was
performed using the supernatant of THP-1 cells treated with
exosomes. The production of both PGE2 and VEGF was
higher in the THP-1 cells treated with exosomes than in the
controls (Fig. 5A).
Following incubation with exosomes from the
celecoxib-treated lung cancer cells, conditioned medium of THP-1
was added to the EA.hy926 cells. In response to VEGF in medium,
tube formation was observed in the EA.hy926 cells, as in HUVECs.
The THP-1 cells incubated with exosomes from the celecoxib-treated
11-18 cells produced angiogenic factors, which triggered more tube
formation in the EA.hy926 cells than in the control exosomes
(Fig. 5B). Tube formation counts
were measured and analyzed, as shown in Fig. 5C.
Discussion
Lung cancer has the highest cancer mortality rate
worldwide. Efforts through surgery, radiotherapy and chemotherapy
to improve lung cancer therapy have increased the survival rate;
however, effective chemotherapeutic agents are still required. The
abnormalities of EGFR, KRAS and anaplastic lymphoma kinase (ALK)
genes are being targeted to lung cancer therapy. COX-2 is a
possible target of lung cancer. Celecoxib, a selective COX-2
inhibitor, has binding pockets that interact with COX-2 and disrupt
enzymatic activities. Actually, a high concentration of celecoxib
(>100 µM) effectively induced the death of lung cancer
cells (data not shown). In addition, celecoxib also regulates COX-2
expression. According to other studies, COX-2 transcription is
decreased by celecoxib in cells from gastric cancer, colon cancer
and breast cancer (30-32). However, this study demonstrated
that COX-2 expression was paradoxically increased following
treatment with celecoxib in lung cancer cells. Of note, other COX-2
specific inhibitors (etoricoxib and rofecoxib) did not increase
COX-2 expression, although a high concentration of sodium
salicylate slightly increased COX-2 levels (data not shown). Ramer
et al suggested the induction of COX-2 expression followed
by the activation of PPARγ in lung cancer cells (20). However, we observed that PPARγ
siRNA and GW9662 as a PPARγ antagonist did not regulate COX-2
expression in our experimental system (data not shown). It may be
that the mechanism of COX-2 induction in lung cancer cells is so
complex that it cannot be simply explained. The microarray analysis
of celecoxib-treated lung cancer cells revealed that not only
COX-2, but also numerous transcription factors were altered by
celecoxib (data not shown). Of note, since celecoxib in exosomes
may affect COX-2 expression in THP-1, the THP-1 cells were treated
with celecoxib; however, there was no change in the levels of COX-2
(Fig. 3B). Although THP-1 is also
a cancer cell line (such as A549, H460 and 11-18 cells), even 50
µM of celecoxib did not directly increase COX-2 expression.
We hypothesized that this was due to the fact that THP-1 originates
from monocytes and is associated with different cellular signaling
pathways. To elucidate the mechanisms responsible for the
paradoxical increase in COX-2 expression by celecoxib, further
studies using numerous transcriptional controls of COX-2 are
warranted.
Exosomes have recently been recognized as a system
of delivery. As exosomes can transfer molecules to other cells, the
study of exosomes has been carried out in various diseases
(23,25,26,28).
This study demonstrated that exosomes transfer COX-2 from lung
cancer cells to monocytes. Consequently, COX-2, which is delivered
through exosomes, increases PGE2 and VEGF production.
The mRNA level of COX-2 was not altered when the exosomes were
added to the THP-1 cells (Fig.
3C), demonstrating that the increase in COX-2 expression in
THP-1 cells was not due to transcriptional stimulation or transfer
of COX-2 mRNA by exosomes. Accordingly, proteins of COX-2 in
exosomes may have been directly transferred to the THP-1 cells. As
shown in Fig. 4, the concentration
of COX-2 was also somewhat elevated when the control exosomes were
added to other cells. As untreated 11-18 cells constitutively
express COX-2 (Fig. 1), their
exosomes can contain COX-2 and transfer it to other cells.
Celecoxib is used as an anti-pain and
anti-inflammatory drug and is considered a probable agent for the
adjuvant therapy of COX-2-positive cancers. Cancer cells highly
expressing COX-2 are believed to be a target of celecoxib. In this
study, we found that the expression of COX-2 was increased by
celecoxib in both COX-2-negative and -positive lung cancer cells.
The intrinsic functions of the induced expression of COX-2 in lung
cancer cells were not examined; however, we determined that great
amounts of COX-2 were loaded into exosomes from lung cancer cells,
where these could be delivered to adjacent or distant cells.
Monocyte uptake of COX-2 via exosomes led to the increased
production of PGE2 and VEGF, despite the presence of
celecoxib. Thus, the anti-inflammatory effects of celecoxib may be
diminished and may depend on the amount of exosome uptake and the
effective concentration of celecoxib within lesions. In addition,
after the administration of celecoxib ceases, previously secreted
exosomes remain and can act on other cells. Additional research is
required to explain the comprehensive effects on the whole body of
the exosomal transport of COX-2.
Taken together, the findings of the present study
demonstrate that celecoxib-induced COX-2 expression in lung cancer
cells is transferred to other cells by exosomes. These results
provide some indications that celecoxib modulates COX-2-positive
cancer cells and affects the microenvironments, including
inflammatory reactions.
Acknowledgments
This study was supported by the Basic Science
Research Program through NRF funded by the Ministry of Education
(NRF-2012R1A1A2006909, NRF-2015R1D1A1A01060152).
Abbreviations:
|
COX-2
|
cyclooxygenase-2
|
|
PG
|
prostaglandin
|
|
EGFR
|
epidermal growth factor receptor
|
References
|
1
|
Hirsch FR and Lippman SM: Advances in the
biology of lung cancer chemoprevention. J Clin Oncol. 23:3186–3197.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Keith RL: Lung cancer chemoprevention.
Proc Am Thorac Soc. 9:52–56. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dela Cruz CS, Tanoue LT and Matthay RA:
Lung cancer: Epidemiology, etiology, and prevention. Clin Chest
Med. 32:605–644. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sandler AB and Dubinett SM: COX-2
inhibition and lung cancer. Semin Oncol. 31(Suppl 7): 45–52. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Dy GK and Adjei AA: Novel targets for lung
cancer therapy: Part I. J Clin Oncol. 20:2881–2894. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chen G, Kronenberger P, Teugels E, Umelo
IA and De Grève J: Targeting the epidermal growth factor receptor
in non-small cell lung cancer cells: The effect of combining RNA
interference with tyrosine kinase inhibitors or cetuximab. BMC Med.
10:282012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cuneo KC, Nyati MK, Ray D and Lawrence TS:
EGFR targeted therapies and radiation: Optimizing efficacy by
appropriate drug scheduling and patient selection. Pharmacol Ther.
154:67–77. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Herbst RS, Heymach JV and Lippman SM: Lung
cancer. N Engl J Med. 359:1367–1380. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Gasparini G, Longo R, Sarmiento R and
Morabito A: Inhibitors of cyclooxygenase 2: A new class of
anticancer agents? Lancet Oncol. 4:605–615. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Breyer MD, Harris RC and Matthew D:
Cyclooxygenase 2 and the kidney. Curr Opin Nephrol Hypertens.
10:89–98. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Frungieri MB, Calandra RS, Mayerhofer A
and Matzkin ME: Cyclooxygenase and prostaglandins in somatic cell
populations of the testis. Reproduction. 149:R169–R180. 2015.
View Article : Google Scholar
|
|
12
|
Vosooghi M and Amini M: The discovery and
development of cyclooxygenase-2 inhibitors as potential anticancer
therapies. Expert Opin Drug Discov. 9:255–267. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ghosh N, Chaki R, Mandal V and Mandal SC:
COX-2 as a target for cancer chemotherapy. Pharmacol Rep.
62:233–244. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Rizzo MT: Cyclooxygenase-2 in oncogenesis.
Clin Chim Acta. 412:671–687. 2011. View Article : Google Scholar
|
|
15
|
Grösch S, Maier TJ, Schiffmann S and
Geisslinger G: Cyclooxygenase-2 (COX-2)-independent
anticarcinogenic effects of selective COX-2 inhibitors. J Natl
Cancer Inst. 98:736–747. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Liggett JL, Zhang X, Eling TE and Baek SJ:
Anti-tumor activity of non-steroidal anti-inflammatory drugs:
Cyclooxygenase-independent targets. Cancer Lett. 346:217–224. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kismet K, Akay MT, Abbasoglu O and Ercan
A: Celecoxib: A potent cyclooxygenase-2 inhibitor in cancer
prevention. Cancer Detect Prev. 28:127–142. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wu T, Leng J, Han C and Demetris AJ: The
cyclooxygenase-2 inhibitor celecoxib blocks phosphorylation of Akt
and induces apoptosis in human cholangiocarcinoma cells. Mol Cancer
Ther. 3:299–307. 2004.PubMed/NCBI
|
|
19
|
Jendrossek V: Targeting apoptosis pathways
by Celecoxib in cancer. Cancer Lett. 332:313–324. 2013. View Article : Google Scholar
|
|
20
|
Ramer R, Walther U, Borchert P, Laufer S,
Linnebacher M and Hinz B: Induction but not inhibition of COX-2
confers human lung cancer cell apoptosis by celecoxib. J Lipid Res.
54:3116–3129. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Frydrychowicz M, Kolecka-Bednarczyk A,
Madejczyk M, Yasar S and Dworacki G: Exosomes - structure,
biogenesis and biological role in non-small-cell lung cancer. Scand
J Immunol. 81:2–10. 2015. View Article : Google Scholar
|
|
22
|
Miller IV and Grunewald TG: Tumour-derived
exosomes: Tiny envelopes for big stories. Biol Cell. 107:287–305.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Nazimek K, Bryniarski K, Santocki M and
Ptak W: Exosomes as mediators of intercellular communication:
Clinical implications. Pol Arch Med Wewn. 125:370–380.
2015.PubMed/NCBI
|
|
24
|
Zhang X, Yuan X, Shi H, Wu L, Qian H and
Xu W: Exosomes in cancer: Small particle, big player. J Hematol
Oncol. 8:832015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Milane L, Singh A, Mattheolabakis G,
Suresh M and Amiji MM: Exosome mediated communication within the
tumor microenvironment. J Control Release. 219:278–294. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hannafon BN and Ding WQ: Intercellular
communication by exosome-derived microRNAs in cancer. Int J Mol
Sci. 14:14240–14269. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhang J, Li S, Li L, Li M, Guo C, Yao J
and Mi S: Exosome and exosomal microRNA: Trafficking, sorting, and
function. Genomics Proteomics Bioinformatics. 13:17–24. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Roma-Rodrigues C, Fernandes AR and
Baptista PV: Exosome in tumour microenvironment: Overview of the
crosstalk between normal and cancer cells. Biomed Res Int.
2014:1794862014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Höper MM, Voelkel NF, Bates TO, Allard JD,
Horan M, Shepherd D and Tuder RM: Prostaglandins induce vascular
endothelial growth factor in a human monocytic cell line and rat
lungs via cAMP. Am J Respir Cell Mol Biol. 17:748–756. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Xiang HG, Xie X, Hu FQ, Xiao HB, Zhang WJ
and Chen L: Cyclooxygenase-2 inhibition as a strategy for treating
gastric adenocarcinoma. Oncol Rep. 32:1140–1148. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Fatima N, Yi M, Ajaz S, Stephens RM,
Stauffer S, Greenwald P, Munroe DJ and Ali IU: Altered gene
expression profiles define pathways in colorectal cancer cell lines
affected by celecoxib. Cancer Epidemiol Biomarkers Prev.
17:3051–3061. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Dai ZJ, Ma XB, Kang HF, Gao J, Min WL,
Guan HT, Diao Y, Lu WF and Wang XJ: Antitumor activity of the
selective cyclo-oxygenase-2 inhibitor, celecoxib, on breast cancer
in vitro and in vivo. Cancer Cell Int. 12:532012. View Article : Google Scholar
|