|
1
|
Hideshima T, Bergsagel PL, Kuehl WM and
Anderson KC: Advances in biology of multiple myeloma: Clinical
applications. Blood. 104:607–618. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ge F, Tao S, Bi L, Zhang Z and Zhang X:
Proteomics: Addressing the challenges of multiple myeloma. Acta
Biochim Biophys Sin (Shanghai). 43:89–95. 2011. View Article : Google Scholar
|
|
3
|
Minnema MC, van der Spek E, van de Donk NW
and Lokhorst HM: New developments in the treatment of patients with
multiple myeloma. Neth J Med. 68:24–32. 2010.PubMed/NCBI
|
|
4
|
Hussein MA: Multiple myeloma: Most common
end-organ damage and management. J Natl Compr Canc Netw. 5:170–178.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Doshi M, Lahoti A, Danesh FR, Batuman V
and Sanders PW; American Society of Nephrology Onco-Nephrology
Forum: Paraprotein-Related Kidney Disease: Kidney Injury from
Paraproteins-What Determines the Site of Injury? Clin J Am Soc
Nephrol. 11:2288–2294. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Rajkumar SV, Dimopoulos MA, Palumbo A,
Blade J, Merlini G, Mateos MV, Kumar S, Hillengass J, Kastritis E,
Richardson P, et al: International Myeloma Working Group updated
criteria for the diagnosis of multiple myeloma. Lancet Oncol.
15:e538–e548. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Avet-Loiseau H, Malard F, Campion L,
Magrangeas F, Sebban C, Lioure B, Decaux O, Lamy T, Legros L,
Fuzibet JG, et al Intergroupe Francophone du Myélome: Translocation
t(14;16) and multiple myeloma: Is it really an independent
prognostic factor? Blood. 117:2009–2011. 2011. View Article : Google Scholar
|
|
8
|
Corre J, Munshi N and Avet-Loiseau H:
Genetics of multiple myeloma: Another heterogeneity level? Blood.
125:1870–1876. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Smadja NV, Fruchart C, Isnard F, Louvet C,
Dutel JL, Cheron N, Grange MJ, Monconduit M and Bastard C:
Chromosomal analysis in multiple myeloma: Cytogenetic evidence of
two different diseases. Leukemia. 12:960–969. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Tremblay D: Novel targets in multiple
myeloma. Am J Hematol Oncol. 12:18–25. 2017.
|
|
11
|
Hanbali A, Hassanein M, Rasheed W, Aljurf
M and Alsharif F: The evolution of prognostic factors in multiple
myeloma. Adv Hematol. 2017:48126372017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Alaoui-Jamali MA and Xu YJ: Proteomic
technology for biomarker profiling in cancer: An update. J Zhejiang
Univ Sci B. 7:411–420. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Gholami AM, Hahne H, Wu Z, Auer FJ, Meng
C, Wilhelm M and Kuster B: Global proteome analysis of the NCI-60
cell line panel. Cell Rep. 4:609–620. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Aslam B, Basit M, Nisar MA, Khurshid M and
Rasool MH: Proteomics: Technologies and their applications. J
Chromatogr Sci. 55:182–196. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chanukuppa V, Taware R, Chatterjee T,
Sharma S, More TH, Taunk K, Kumar S, Santra MK and Rapole S:
Current Understanding of the Potential of Proteomics and
Metabolomics Approaches in Cancer Chemoresistance: A Focus on
Multiple Myeloma. Curr Top Med Chem. 18:2584–2598. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Amaya M, Baer A, Voss K, Campbell C,
Mueller C, Bailey C, Kehn-Hall K, Petricoin E III and Narayanan A:
Proteomic strategies for the discovery of novel diagnostic and
therapeutic targets for infectious diseases. Pathog Dis.
71:177–189. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Nagaraj NS, Singh OV and Merchant NB:
Proteomics: A strategy to understand the novel targets in protein
misfolding and cancer therapy. Expert Rev Proteomics. 7:613–623.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Moseley FL, Bicknell KA, Marber MS and
Brooks G: The use of proteomics to identify novel therapeutic
targets for the treatment of disease. J Pharm Pharmacol.
59:609–628. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ma TZ, Piao Z, Jin SY and Kwak YG:
Differential expression of serum proteins in multiple myeloma. Exp
Ther Med. 17:649–656. 2019.PubMed/NCBI
|
|
20
|
Bai J, Yang Y, Wang J, Zhang L, Wang F and
He A: Variability of serum novel serum peptide biomarkers
correlates with the disease states of multiple myeloma. Clin
Proteomics. 16:172019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Fernando RC, de Carvalho F, Mazzotti DR,
Evangelista AF, Braga WMT, de Lourdes Chauffaille M, Leme AFP and
Colleoni GWB: Multiple myeloma cell lines and primary tumors
proteoma: Protein biosynthesis and immune system as potential
therapeutic targets. Genes Cancer. 6:462–471. 2015.
|
|
22
|
Sasikala P, Harsha C, Bajaj J, Sharma R,
Pandey A and Krishna S: Quantitative Proteomic Profiling Unravels
Dynamic Changes in the Myeloma Cell Proteome Treated with Valproic
Acid (VPA). Blood. 118:18472011. View Article : Google Scholar
|
|
23
|
Chanukuppa V, Paul D, Taunk K, Chatterjee
T, Sharma S, Kumar S, Santra MK and Rapole S: XPO1 is a critical
player for bortezomib resistance in multiple myeloma: A
quantitative proteomic approach. J Proteomics. 209:1035042019.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tripathy S: The role of serum protein
electrophoresis in the detection of multiple myeloma: An experience
of a corporate hospital. J Clin Diagn Res. 6:1458–1461. 2012.
|
|
25
|
Tosi P, Tomassetti S, Merli A and Polli V:
Serum free light-chain assay for the detection and monitoring of
multiple myeloma and related conditions. Ther Adv Hematol. 4:37–41.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Gerecke C, Fuhrmann S, Strifler S,
Schmidt‑Hieber M, Einsele H and Knop S: The diagnosis and treatment
of multiple myeloma. Dtsch Arztebl Int. 113:470–476.
2016.PubMed/NCBI
|
|
27
|
Rajkumar SV and Kyle RA: Multiple myeloma:
diagnosis and treatment. In: Mayo Clinic Proceedings; 80. Elsevier;
pp. 1371–1382. 2005
|
|
28
|
Gajendra S, Jha B, Goel S, Sahni T, Sharma
R, Shariq M, Jaiswal S and Sachdev R: Leishman and Giemsa stain: A
new reliable staining technique for blood/bone marrow smears. Int J
Lab Hematol. 37:774–782. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2-ΔΔCT method. Methods. 25:402–408. 2001. View Article : Google Scholar
|
|
30
|
Gajbhiye A, Dabhi R, Taunk K,
Jagadeeshaprasad MG, RoyChoudhury S, Mane A, Bayatigeri S,
Chaudhury K, Santra MK and Rapole S: Multipronged quantitative
proteomics reveals serum proteome alterations in breast cancer
intrinsic subtypes. J Proteomics. 163:1–13. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Takata T, Ishigaki Y, Shimasaki T,
Tsuchida H, Motoo Y, Hayashi A and Tomosugi N: Characterization of
proteins secreted by pancreatic cancer cells with anticancer drug
treatment in vitro. Oncol Rep. 28:1968–1976. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tennant JR: Evaluation of the trypan blue
technique for determination of cell viability. Transplantation.
2:685–694. 1964. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
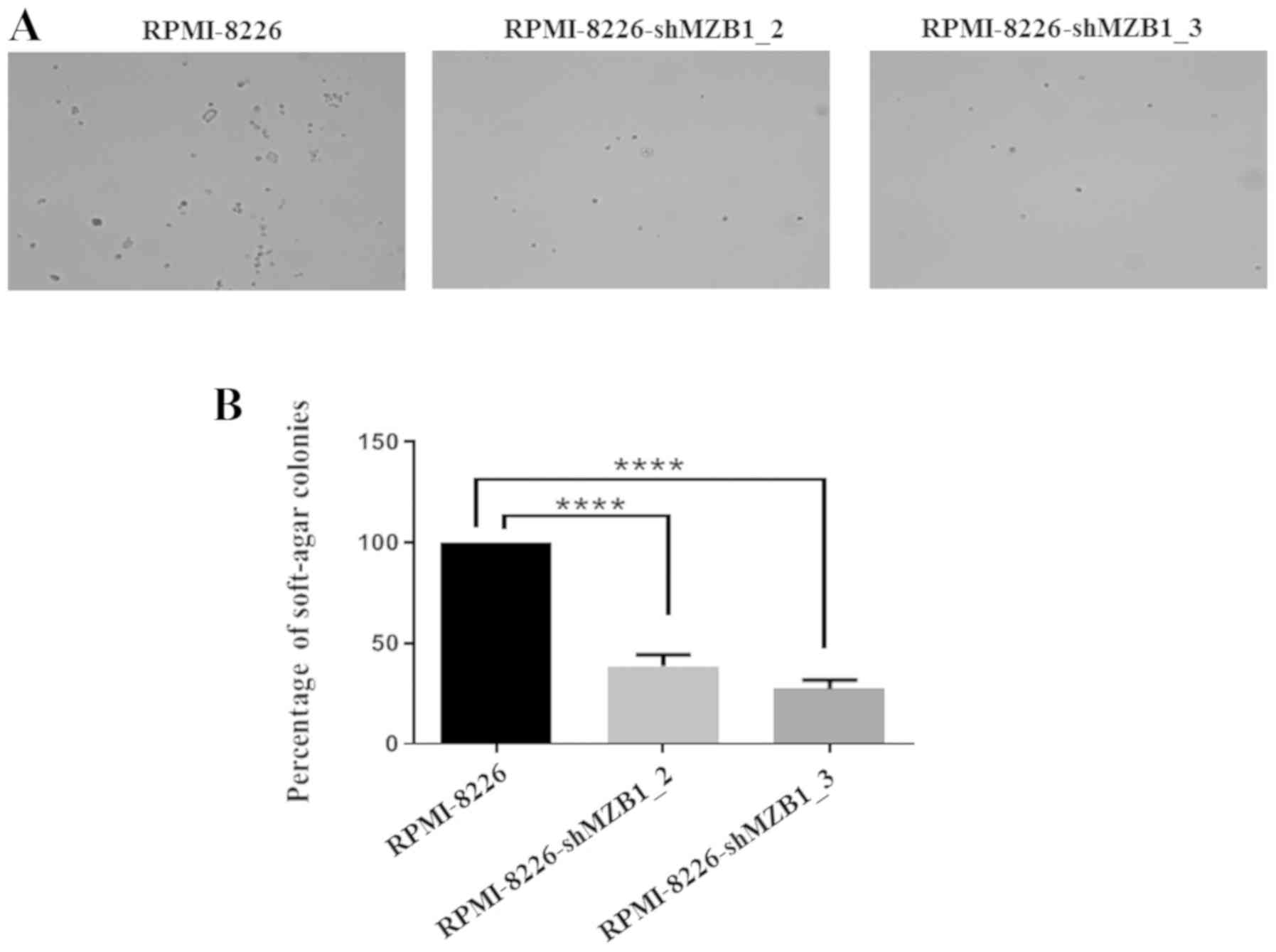
Courtenay VD: A soft agar colony assay for
Lewis lung tumour and B16 melanoma taken directly from the mouse.
Br J Cancer. 34:39–45. 1976. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Shimizu Y, Meunier L and Hendershot LM:
pERp1 is significantly up-regulated during plasma cell
differentiation and contributes to the oxidative folding of
immunoglobulin. Proc Natl Acad Sci USA. 106:17013–17018. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Flach H, Rosenbaum M, Duchniewicz M, Kim
S, Zhang SL, Cahalan MD, Mittler G and Grosschedl R: Mzb1 protein
regulates calcium homeostasis, antibody secretion, and integrin
activation in innate-like B cells. Immunity. 33:723–735. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Mori S, Chang JT, Andrechek ER, Matsumura
N, Baba T, Yao G, Kim JW, Gatza M, Murphy S and Nevins JR:
Anchorage‑independent cell growth signature identifies tumors with
metastatic potential. Oncogene. 28:2796–2805. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
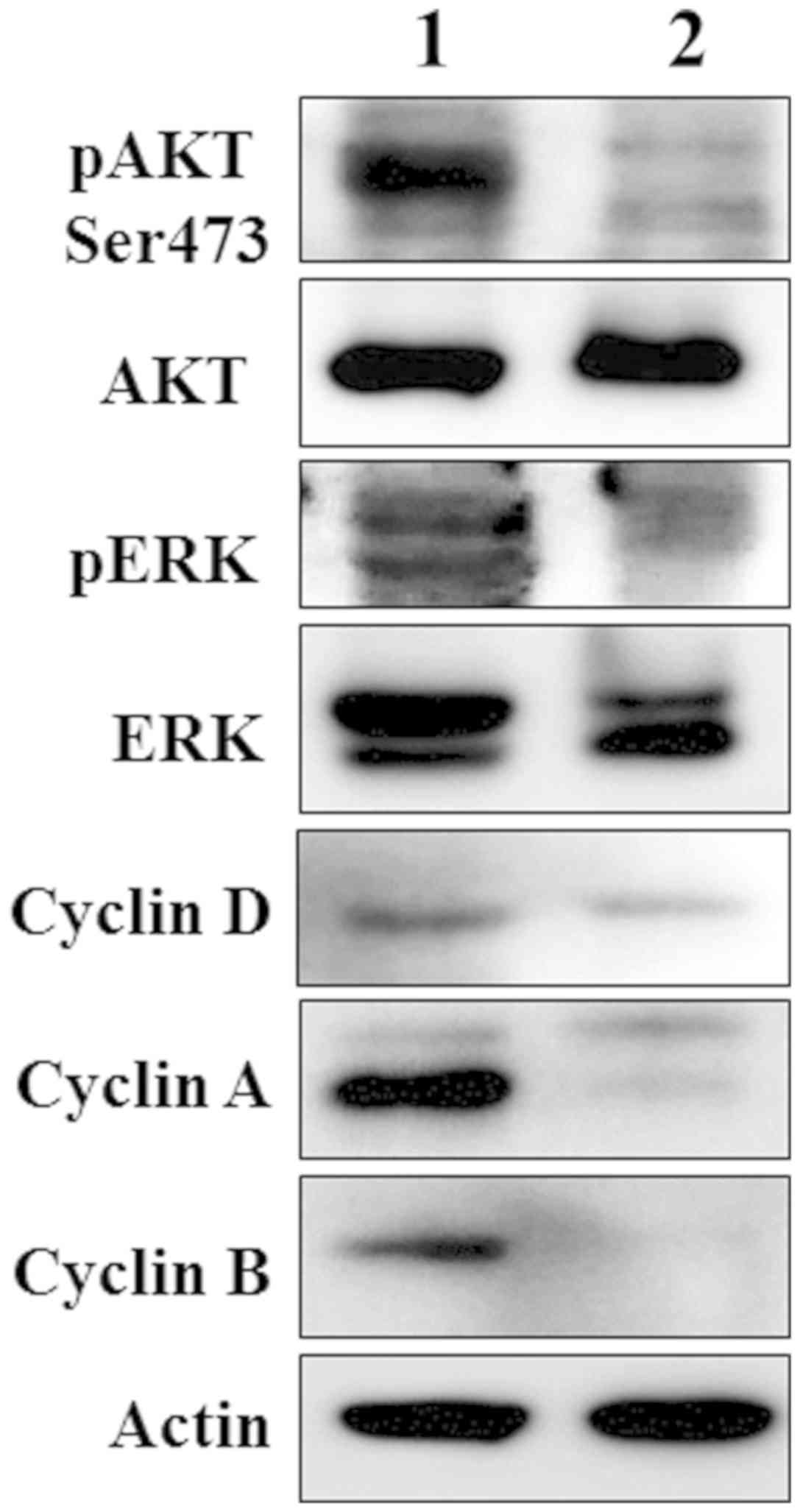
Younes H, Leleu X, Hatjiharissi E, Moreau
AS, Hideshima T, Richardson P, Anderson KC and Ghobrial IM:
Targeting the phosphatidylinositol 3-kinase pathway in multiple
myeloma. Clin Cancer Res. 13:3771–3775. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Jin Y and Dai Z: USO1 promotes tumor
progression via activating Erk pathway in multiple myeloma cells.
Biomed Pharmacother. 78:264–271. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Alao JP: The regulation of cyclin D1
degradation: Roles in cancer development and the potential for
therapeutic invention. Mol Cancer. 6:242007. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Desdouets C, Matesic G, Molina CA, Foulkes
NS, Sassone-Corsi P, Brechot C and Sobczak-Thepot J: Cell cycle
regulation of cyclin A gene expression by the cyclic AMP-responsive
transcription factors CREB and CREM. Mol Cell Biol. 15:3301–3309.
1995. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Hershko A: Mechanisms and regulation of
the degradation of cyclin B. Philos Trans R Soc Lond B Biol Sci.
354:1571–1575; discussion 1575-1576. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Landgren O and Weiss BM: Patterns of
monoclonal gammopathy of undetermined significance and multiple
myeloma in various ethnic/racial groups: Support for genetic
factors in pathogenesis. Leukemia. 23:1691–1697. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Rajkumar SV and Kyle RA: Treatment of
multiple myeloma and related disorders. Cambridge University Press;
2008, View Article : Google Scholar
|
|
44
|
Teras LR, DeSantis CE, Cerhan JR, Morton
LM, Jemal A and Flowers CR: 2016 US lymphoid malignancy statistics
by World Health Organization subtypes. CA Cancer J Clin.
66:443–459. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
van Anken E, Pena F, Hafkemeijer N,
Christis C, Romijn EP, Grauschopf U, Oorschot VM, Pertel T, Engels
S, Ora A, et al: Efficient IgM assembly and secretion require the
plasma cell induced endoplasmic reticulum protein pERp1. Proc Natl
Acad Sci USA. 106:17019–17024. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Miyagawa-Hayashino A, Yoshifuji H,
Kitagori K, Ito S, Oku T, Hirayama Y, Salah A, Nakajima T, Kiso K,
Yamada N, et al: Increase of MZB1 in B cells in systemic lupus
erythematosus: Proteomic analysis of biopsied lymph nodes.
Arthritis Res Ther. 20:132018. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Matsumura S, Imoto I, Kozaki K, Matsui T,
Muramatsu T, Furuta M, Tanaka S, Sakamoto M, Arii S and Inazawa J:
Integrative array‑based approach identifies MZB1 as a frequently
methylated putative tumor suppressor in hepatocellular carcinoma.
Clin Cancer Res. 18:3541–3551. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Kanda M, Tanaka C, Kobayashi D, Tanaka H,
Shimizu D, Shibata M, Takami H, Hayashi M, Iwata N, Niwa Y, et al:
Epigenetic suppression of the immunoregulator MZB1 is associated
with the malignant phenotype of gastric cancer. Int J Cancer.
139:2290–2298. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Herold T, Mulaw MA, Jurinovic V, Seiler T,
Metzeler KH, Dufour A, Schneider S, Kakadia PM, Spiekermann K,
Mansmann U, et al: High expression of MZB1 predicts adverse
prognosis in chronic lymphocytic leukemia, follicular lymphoma and
diffuse large B-cell lymphoma and is associated with a unique gene
expression signature. Leuk Lymphoma. 54:1652–1657. 2013. View Article : Google Scholar
|
|
50
|
Uhlén M, Fagerberg L, Hallström BM,
Lindskog C, Oksvold P, Mardinoglu A, Sivertsson Å, Kampf C,
Sjöstedt E, Asplund A, et al: Proteomics. Tissue-based map of the
human proteome. Science. 347:12604192015. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Cruz‑Rodriguez N, Combita AL, Enciso LJ,
Quijano SM, Pinzon PL, Lozano OC, Castillo JS, Li L, Bareño J,
Cardozo C, et al: High expression of ID family and IGJ genes
signature as predictor of low induction treatment response and
worst survival in adult Hispanic patients with B-acute
lympho-blastic leukemia. J Exp Clin Cancer Res. 35:642016.
View Article : Google Scholar
|
|
52
|
Peng M, Deng J, Zhou S, Tao T, Su Q and
Yang X and Yang X: The role of Clusterin in cancer metastasis.
Cancer Manag Res. 11:2405–2414. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Faiman B, Tariman JD, Mangan PA and Spong
J: Renal complications in multiple myeloma and related disorders:
Survivorship care plan of the IMF Nurse Leadership Board. Clin J
Oncol Nurs. 15:66–76. 2011. View Article : Google Scholar
|
|
54
|
Kim JE, Yoo C, Lee DH, Kim SW, Lee JS and
Suh C: Serum albumin level is a significant prognostic factor
reflecting disease severity in symptomatic multiple myeloma. Ann
Hematol. 89:391–397. 2010. View Article : Google Scholar
|
|
55
|
Zhu J, Wang M, Cao B, Hou T and Mao X:
Targeting the phosphatidylinositol 3-kinase/AKT pathway for the
treatment of multiple myeloma. Curr Med Chem. 21:3173–3187. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Xu H, Li J and Zhou ZG: NEAT1 promotes
cell proliferation in multiple myeloma by activating PI3K/AKT
pathway. Eur Rev Med Pharmacol Sci. 22:6403–6411. 2018.PubMed/NCBI
|
|
57
|
Ohguchi H, Harada T, Sagawa M, Kikuchi S,
Tai YT, Richardson PG, Hideshima T and Anderson KC: KDM6B modulates
MAPK pathway mediating multiple myeloma cell growth and survival.
Leukemia. 31:2661–2669. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Kubiczková L, Dúcka M, Sedlaříková L,
Kryukov F, Hájek R and Ševčíková S: Cyclins D in regulation and
dysregulation of the cell cycle in multiple myeloma. Klin Onkol.
26:313–318. 2013.In Czech. View Article : Google Scholar
|
|
59
|
Quinn J, Glassford J, Percy L, Munson P,
Marafioti T, Rodriguez-Justo M and Yong K: APRIL promotes
cell-cycle progression in primary multiple myeloma cells: Influence
of D-type cyclin group and translocation status. Blood.
117:890–901. 2011. View Article : Google Scholar
|
|
60
|
Ettari R, Pallio G, Pizzino G, Irrera N,
Zappalà M, Maiorana S, Di Chio C, Altavilla D, Squadrito F and
Bitto A: Non-covalent immunoproteasome inhibitors induce cell cycle
arrest in multiple myeloma MM.1R cells. J Enzyme Inhib Med Chem.
34:1307–1313. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Vizcaíno JA, Csordas A, del‑Toro N, Dianes
JA, Griss J, Lavidas I, Mayer G, Perez-Riverol Y, Reisinger F,
Ternent T, et al: 2016 update of the PRIDE database and its related
tools. Nucleic Acids Res. 44(D1): D447–D456. 2016. View Article : Google Scholar :
|