Head and neck cancer is the most common cancer type
globally and may include the nasopharynx, larynx, pharynx and oral
cavity (1,2). Oral cancer may be categorized into
buccal carcinoma, gingival carcinoma, maxillary sinus carcinoma,
tongue cancer and carcinoma of the floor of the mouth, 90% of which
are squamous cell carcinoma (1,3).
According to 2020 statistics, ~177,757 patients died of cancer in
these parts within the oral cavity (4). In general, the occurrence and
development of oral cancer are related to various factors,
including excessive smoking, drinking, betel nut chewing and other
external factors, gene mutation, human papillomavirus infection,
epigenetic modification and other internal factors (1,5-9).
With the in-depth study of epigenetics, RNA modification has been
indicated to be involved in various physiological processes,
including cell proliferation (10). Furthermore, it is closely
associated with the pathological processes of cancer (11).
Human epigenetics include DNA methylation, histone
modification and RNA modification, and are closely related to
various physiological activities, such as cell transcription and
differentiation, and have a critical role in gene expression and
regulation (12,13). Certain epigenetic changes are
related to oral squamous cell carcinoma (OSCC) (6). RNA modification is a chemical
modification in cells that may efficiently and specifically
regulate the gene expression and function of biological
macromolecules (14). It is
suggested that >170 kinds of RNA modifications have been
identified (15). The most common
RNA modifications include N6-methyladenosine
(m6A), m7G, m1A and
m5C, which have different roles in cells (14).
Human epigenetic mechanisms involve DNA methylation,
histone modification and RNA modification. These modifications
directly participate in gene expression and regulate biological
growth (13). RNA modification is
an integral part of the epigenetic mechanism and is closely
associated with the normal function of RNAs (21). RNA modifications mainly occur on
transfer RNA (tRNA) and non-coding RNA (15). Common RNA modifications include
m6A, m7G, m1A and m5C.
m7G modification in mRNA may be related to protein
translation (22). m1A
modification increases the structural stability of tRNA and induces
precise tRNA folding (23,24). m5C may affect the
translation accuracy of mRNA and regulate tRNA stability (25). m6A modification involves
the methylation of the sixth nitrogen atom on the base A of RNA
molecules (26). It is the most
abundant chemical modification within eukaryotic mRNA modification
(27). Meanwhile, m6A
modification also exists on long intergenic non-coding RNAs,
primary microRNAs and ribosomal RNA (28-30).
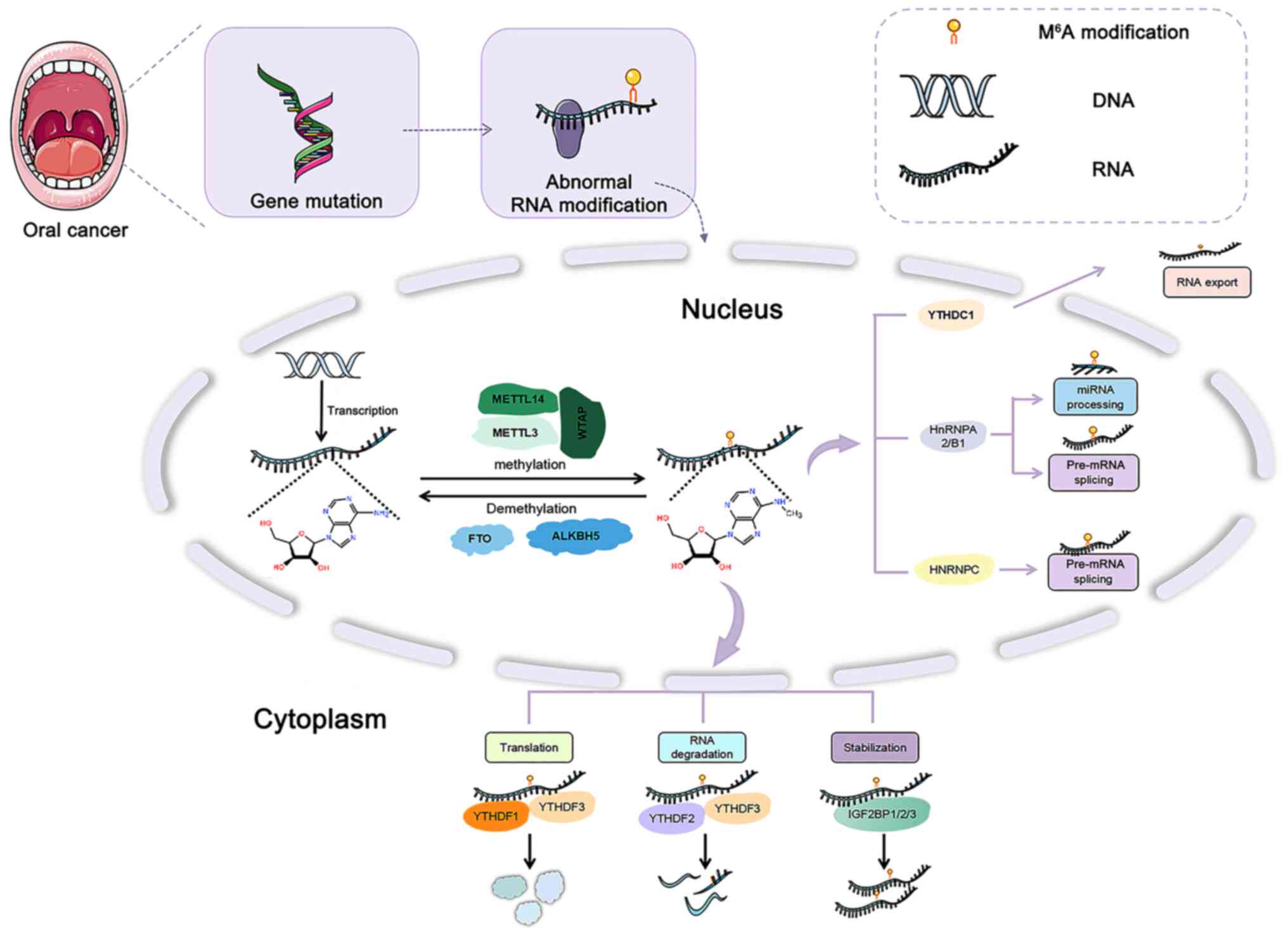
m6A modification is dynamic and reversible, including
the joint action of several catalytic enzymes (27,31).
Methyltransferase, demethylase and methylated reading protein are
the main components that affect the stability, splicing and
translation of mRNA (27,31) (Fig.
1).
Oral cancer is a crucial component of head and neck
solid tumors. METTL3 and B-cell-specific Moloney murine leukemia
virus integration site 1 (BMI1) expression are upregulated, and
when the METTL3 gene is knocked down, BMI1 expression is reduced,
and the proliferation, migration and invasion abilities among oral
cancer cells become inhibited (54). Further studies have indicated that
METTL3 was able to facilitate OSCC development by promoting the
m6A methylation of BMI1 (54). Furthermore, the m6A
demethylase FTO may regulate the oral cancer cell cycle and promote
progression by regulating the expression of Cyclin D1 (75). m6A modifications have
multiple roles and may control oral cancer progression (Table II).
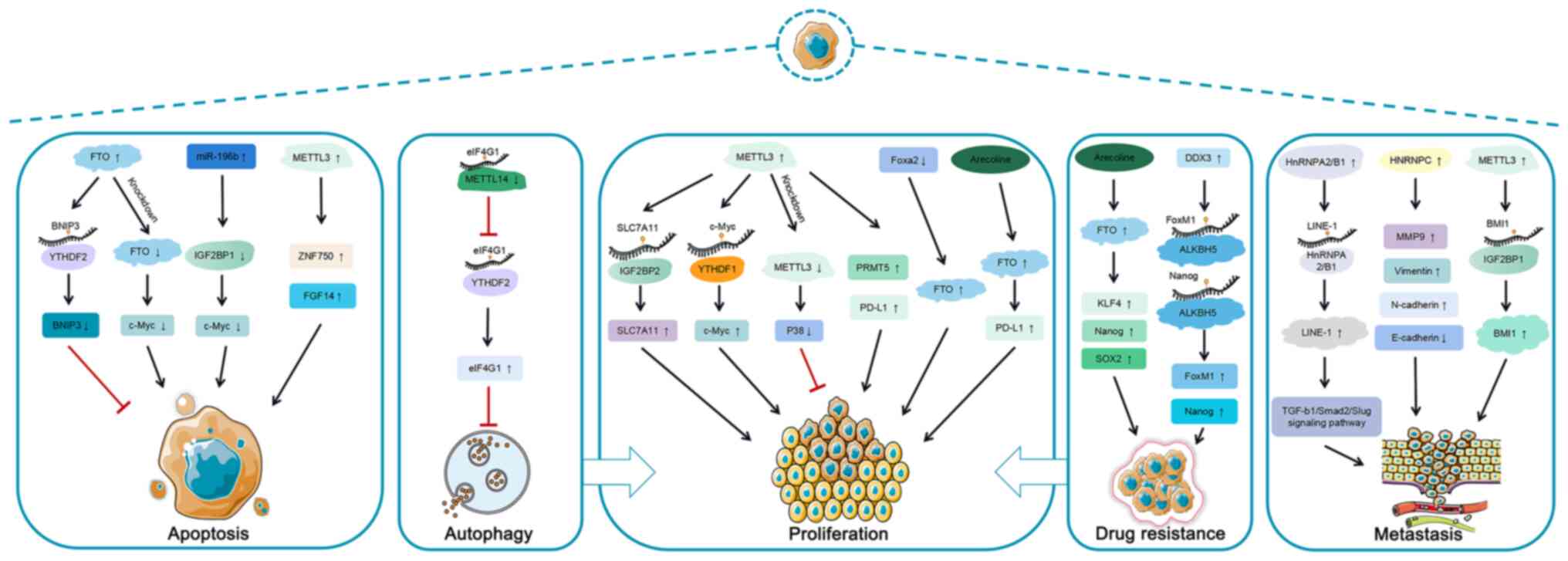
A characteristic of cancer cells is their resistance
against cell death. Cell death includes apoptosis, autophagic cell
death, ferroptosis, necroptosis and pyroptosis (79,80).
Cell death is involved in the progression of multiple malignancies
and it is closely associated with m6A modifications
(81). Therefore, the interaction
between cell death and m6A modification has been
elaborated in oral cancer.
Apoptosis is the programmed death of cells
controlled by certain genes to maintain the internal environment
stability of cells (82). Cancer
features malignant proliferation and less apoptosis (82). Therefore, cancer treatment includes
cell proliferation, metastasis and apoptosis as the therapeutic
targets. Furthermore, the expression of m6A demethylase
FTO is upregulated and significantly inhibits cell apoptosis
(83). Further study indicated
that Bcl-2/adenovirus E1B 19kDa interacting protein 3 (BNIP3) is
the downstream target of FTO-mediated m6A modification.
FTO regulates m6A demethylation of BNIP3 and induces its
degradation via the YTHDF2-independent mechanism. The inhibition of
FTO expression leads to the promotion of BNIP3 expression,
increasing apoptosis of breast cancer cells and inhibiting their
proliferation (83). In
nasopharyngeal carcinoma, inhibition of METTL3 expression promoted
ZNF750 expression and then upregulated FGF14 expression, promoting
cancer cell apoptosis (72).
m6A reader IGF2BP1 in hepatocellular carcinoma is the
target gene of miR-196b (84).
Furthermore, miR-196b overexpression may inhibit the expression of
IGF2BP1 and reduce the expression level of c-Myc. Thus, it promotes
apoptosis of hepatocellular carcinoma cells (84). The above evidence indicates that
m6A modification may affect the apoptosis of various
tumor cells, thus inhibiting tumor progression; it is essential to
further clarify the relationship between m6A and
apoptosis.
Autophagy is a regular type of physiological
activity in eukaryotic cells involving the degradation of
organelles, proteins and other substances transferred to lysosomes.
It is associated with various diseases, including
neurodegenerative, inflammatory and autoimmune conditions, as well
as cancer (85,86). Autophagy has a complex role in the
development of tumors. It may produce protective autophagy to
promote tumor growth and cytotoxic autophagy to inhibit tumor
growth (87,88). Furthermore, autophagy influences
cell behavior (89,90). Thus, autophagy is closely related
to cell death and proliferation (91).
Ferroptosis is an iron-dependent form of cell death
with a potential application in cancer therapy (92). A recent study suggested that
m6A modification may be involved in the process of
ferroptosis in oral cancer (93).
In this report, immunological analyses indicated differential
expression of m6A in high-risk and low-risk groups of
oral squamous carcinoma patients. Furthermore, a prognostic model
based on eight ferroptosis lncRNAs was able to provide a prognostic
assessment and immunological analysis for patients with OSCC
(93). This indicates that
ferroptosis has a critical role in oral cancer progression in which
m6A was involved.
Rapid and uncontrolled proliferation is the most
basic and essential characteristic of cancer (102). Most abnormal proliferation of
cancer cells is associated with the expression changes of a series
of genes or activating signal pathways through epigenetic
modification (103). In
particular, m6A modification is closely related to the
proliferation of cancer cells (42). It was indicated that the expression
of programmed cell death 1 ligand (PD-L1) is upregulated in
patients with OSCC. Furthermore, m6A eraser FTO promotes
the expression of PD-L1 by mediating m6A modification
and MYC activity and upregulating PD-L1 to promote cell
proliferation (19). It is well
known that betel nut chewing is a risk factor for oral cancer.
Furthermore, arecoline exposure may significantly upregulate FTO,
MYC and PD-L1 in OSCC (19).
Another study reported that knockdown of transcription factor
forkhead box (Fox)a2 was able to negatively regulate FTO expression
and promote cell proliferation in OSCC (60). METTL3 is significantly expressed in
OSCC and may stimulate solute carrier family 7 member 11 (SLC7A11)
expression through m6A-mediated IGF2BP2 binding, thus
facilitating OSCC proliferation (20). The study also observed that
triptolide may inhibit OSCC progression by inhibiting the
METTL3-SLC7A11 axis (20).
Furthermore, another study indicated that knocking down the
expression of METTL3 impaired the stem cell-like activity in OSCC
cells (104). It may reduce the
m6A level, downregulate p38 expression and inhibit the
cells' proliferation ability (104). METTL3 also enhances the stability
of c-Myc through YTHDF1-mediated m6A modification and
promotes the occurrence and development of OSCC (105). Previous report observed that
METTL3 may enable the expression of protein arginine
methyltransferase 5 (PRMT5) and PD-L1, thus facilitating OSCC
proliferation (106). METTL3 may
also promote OSCC proliferation by promoting m6A
methylation of BMI1 (54). To
date, the studies on the effect of m6A on cell
proliferation involving OSCC were primarily focused on METTL3 and
FTO. m6A is able to promote proliferation through
various regulatory mechanisms, indicating a complex effect of
m6A modification on OSCC.
Most tumors have the characteristics of invasion
and metastasis. EMT has a critical role in cancer metastasis. EMT
is a cellular process involving cells losing their epithelial
characteristics and acquiring mesenchymal characteristics (107). It has several biological
functions during the process of tumor metastasis. Its occurrence
markers usually refer to the loss of the epithelial marker
E-cadherin and upregulation of the interstitial marker Vimentin
(107).
Chemotherapy resistance is a complex problem in
OSCC treatment. It is a defensive mechanism of tumor cells to
maintain their homeostasis. The inducement of chemotherapy
resistance includes gene mutation, gene amplification and
epigenetic changes (108).
Furthermore, cancer stem cells (CSCs) are also important for drug
resistance in tumors (109).
Previous studies have revealed that arecoline-treated OSCC cells
may upregulate FTO expression and enhance their resistance to
cisplatin, a cancer chemotherapy drug. By contrast, the mRNA and
protein levels of tumor stem cell pluripotent transcription factors
Nanog, SOX2 and Kruppel-like factor 4 (KLF4) are all upregulated
(60). However, downregulating the
expression level of FTO may increase the sensitivity of OSCC cells
toward cisplatin (60). In
addition, the expression levels of the pluripotent transcription
factors videlicet Nanog, SOX2 and KLF4 in tumor stem cells
decreased to varying degrees, rendering FTO a potential therapeutic
target for cisplatin resistance in OSCC (60). Another m6A demethylase,
ALKBH5, has also been indicated to be closely associated with
chemotherapy resistance in OSCC (110). A study has demonstrated that the
human RNA helicase DEAD-box helicase 3 (DDX3) expression is
upregulated in cisplatin-resistant OSCC cells. When DDX3 expression
is downregulated, the CSC marker is also downregulated in
chemotherapy-resistant OSCC cells. Furthermore, DDX3 regulates the
expression of the CSC transcription factors FoxM1 and Nanog,
through ALKBH5, thereby promoting cisplatin resistance in OSCC
(110). The emergence of
chemotherapy resistance challenges oral cancer treatment, which is
closely associated with various mechanisms, including
m6A. Elucidating the mechanisms of chemotherapy
resistance is of great significance in understanding oral cancer
development.
Immunotherapy has gradually become the focus of the
in-depth understanding of tumor immunology. Tumor cells
downregulate the expression of antigens on the cell surface and
escape immune surveillance through various mechanisms (111). Furthermore, m6A
modifications have essential roles during the generation of immune
responses (112). A previous
study indicated that, depending on m6A
regulation-related genes, a prognostic marker may effectively
determine the prognosis of HNSCC (113). This prognostic marker was
associated with immune cell infiltration in HNSCC (113). A recent study suggested that
METTL3 expression was increased in OSCC and inhibited
CD8+ T-cell activation (106). Furthermore, METTL3 was indicated
to regulate the expression of PRMT5 and PD-L1 through methylation
modification, thereby modulating OSCC immunity (106). Another study determined that in
OSCC, FTO was involved in the resistance to T-cell lethality by
regulating the MYC/PD-L1 signaling pathway (60). Thus, studying immune response and
m6A may provide novel therapeutic targets for
immunotherapy in oral cancer.
In-depth epigenetics studies may help reveal the
biological mechanisms of cancer and provide new targets for cancer
treatment. RNA modification has a critical regulatory role in
several cancers as an essential branch of epigenetics. It causes
changes in the expression of certain proteins in cells by
regulating the expression of various genes, leading to
carcinogenesis (16). For
instance, METTL3 promotes the maturation of miR221/222 to reduce
the expression of PTEN protein to encourage the proliferation of
bladder cancer cells (42). In
oral cancer, FTO promotes PD-L1 expression to facilitate cell
proliferation (19). Furthermore,
METTL3 promotes SLC7A11 expression through m6A-mediated
IGF2BP2 binding, thereby enabling OSCC proliferation (20). m6A modification is the
most common mRNA modification, with substantial research
significance (14). m6A
includes a variety of modifying enzyme components. m6A
writer catalyzes the methylation of m6A on RNA. The
recognition of m6A methylation by the reader affects the
splicing, output, degradation, translation and other biological
processes of mRNA. m6A eraser may remove these
modifications, resulting in a dynamic and reversible process
(16). In addition, m6A
modification affects miRNAs, the processing of miRNAs, and the
biological function of lncRNAs and promotes the translation of
circRNAs (16). m6A
modification may have a direct or indirect regulatory role in
numerous intracellular activities.
Oral cancer progression is associated with various
cancer cell behaviors, such as cell proliferation, migration,
invasion and autophagy (114,115). m6A modification may
promote oral cancer progression by regulating m6A-related gene
expression to influence cell proliferation, metastasis and
aggression, and inhibiting autophagy (19,39,56,59,110). In addition, the abnormal
expression of m6A may lead to chemotherapy resistance in
oral cancer (60). Apoptosis is a
crucial cellular behavior among cancer cells (116). Studies have indicated that
m6A modification affected the fate of cancer cells by
regulating apoptosis (72,83,84).
For instance, IGF2BP1, FTO and METTL3 may induce apoptosis
(72,83,84).
Therefore, m6A modification and m6A-related
genes may affect cell behavior in oral cancer. Furthermore, it was
observed that certain drugs may target m6A to inhibit
oral cancer progression. Oxymatrine reduces the expression of CXC
motif chemokine receptor 4 by downregulating METTL3 and
m6A modification levels, thus inhibiting the progression
of OSCC (117). Triptolide
inhibits METTL3-mediated expression of SLC7A11, thereby suppressing
the malignancy of OSCC (20).
Allocryptopine reduces METTL3 expression and inhibits
m6A modification of patched receptor 1 and the
proliferation and EMT of OSCC through the m6A-mediated
Hedgehog signaling pathway (118). Therefore, m6A may be a
potential therapeutic target for oral cancer.
Not applicable.
HL and YW wrote the manuscript. HL, DW, WL, TX, SK,
MH, ZY, YG collected the references and prepared figures. All
authors reviewed the manuscript. All authors read and approved the
final manuscript. Data authentication is not applicable.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
Not applicable.
This work was supported by the Fundamental Research Funds for
the Jilin Province Department of Finance (grant no.
jcsz2021893-13), the Changchun Scientific and Technological
Development Program (grant no. 21ZY26), the Jilin Province
Scientific and Technological Development Program (grant nos.
20200801077GH, 20210204013YY, 20200504005YY and 20220505033ZP) and
Scientific Research Project of Traditional Chinese Medicine Bureau
of Guangdong Province (grant no. 20201236).
|
1
|
Chai AWY, Lim KP and Cheong SC:
Translational genomics and recent advances in oral squamous cell
carcinoma. Semin Cancer Biol. 61:71–83. 2020. View Article : Google Scholar
|
|
2
|
Huo XX, Wang SJ, Song H, Li MD, Yu H, Wang
M, Gong HX, Qiu XT, Zhu YF and Zhang JY: Roles of major RNA
adenosine modifications in head and neck squamous cell carcinoma.
Front Pharmacol. 12:7797792021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
D'souza S and Addepalli V: Preventive
measures in oral cancer: An overview. Biomed Pharmacother.
107:72–80. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Pickering CR, Zhang J, Yoo SY, Bengtsson
L, Moorthy S, Neskey DM, Zhao M, Ortega Alves MV, Chang K, Drummond
J, et al: Integrative genomic characterization of oral squamous
cell carcinoma identifies frequent somatic drivers. Cancer Discov.
3:770–781. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mascolo M, Siano M, Ilardi G, Russo D,
Merolla F, Rosa G and Staibano S: Epigenetic disregulation in oral
cancer. Int J Mol Sci. 13:2331–2353. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Goldenberg D, Lee J, Koch WM, Kim MM,
Trink B, Sidransky D and Moon CS: Habitual risk factors for head
and neck cancer. Otolaryngol Head Neck Surg. 131:986–993. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Guha N, Warnakulasuriya S, Vlaanderen J
and Straif K: Betel quid chewing and the risk of oral and
oropharyngeal cancers: A meta-analysis with implications for cancer
control. Int J Cancer. 135:1433–1443. 2014. View Article : Google Scholar
|
|
9
|
Herrero R, Castellsagué X, Pawlita M,
Lissowska J, Kee F, Balaram P, Rajkumar T, Sridhar H, Rose B,
Pintos J, et al: Human papillomavirus and oral cancer: The
International Agency for research on cancer multicenter study. J
Natl Cancer Inst. 95:1772–1783. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang W: mRNA methylation by NSUN2 in cell
proliferation. Wiley Interdiscip Rev RNA. 7:838–842. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Delaunay S and Frye M: RNA modifications
regulating cell fate in cancer. Nat Cell Biol. 21:552–559. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Bonasio R, Tu S and Reinberg D: Molecular
signals of epigenetic states. Science. 330:612–616. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ling C and Rönn T: Epigenetics in human
obesity and type 2 diabetes. Cell Metab. 29:1028–1044. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Barbieri I and Kouzarides T: Role of RNA
modifications in cancer. Nat Rev Cancer. 20:303–322. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Machnicka MA, Milanowska K, Osman Oglou O,
Purta E, Kurkowska M, Olchowik A, Januszewski W, Kalinowski S,
Dunin-Horkawicz S, Rother KM, et al: MODOMICS: A data-base of RNA
modification pathways-2013 update. Nucleic Acids Res. 41(Database
Issue): D262–D267. 2013. View Article : Google Scholar
|
|
16
|
Wang T, Kong S, Tao M and Ju S: The
potential role of RNA N6-methyladenosine in cancer progression. Mol
Cancer. 19:882020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Liu ZX, Li LM, Sun HL and Liu SM: Link
between m6A modification and cancers. Front Bioeng Biotechnol.
6:892018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jiang X, Liu B, Nie Z, Duan L, Xiong Q,
Jin Z, Yang C and Chen Y: The role of m6A modification in the
biological functions and diseases. Signal Transduct Target Ther.
6:742021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li X, Chen W, Gao Y, Song J, Gu Y, Zhang
J, Cheng X and Ai Y: FTO regulates arecoline-exposed oral cancer
immune response through PD-L1. Cancer Sci. 113:2962–2973. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Xu L, Li Q, Wang Y, Wang L, Guo Y, Yang R,
Zhao N, Ge N, Wang Y and Guo C: m6A methyltransferase
METTL3 promotes oral squamous cell carcinoma progression through
enhancement of IGF2BP2-mediated SLC7A11 mRNA stability. Am J Cancer
Res. 11:5282–5298. 2021.
|
|
21
|
Shi H, Wei J and He C: Where, when, and
how: Context-dependent functions of RNA methylation writers,
readers, and eras. Mol Cell. 74:640–650. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Malbec L, Zhang T, Chen YS, Zhang Y, Sun
BF, Shi BY, Zhao YL, Yang Y and Yang YG: Dynamic methylome of
internal mRNA N7-methylguanosine and its regulatory role
in translation. Cell Res. 29:927–941. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Saikia M, Fu Y, Pavon-Eternod M, He C and
Pan T: Genome-wide analysis of N1-methyl-adenosine modification in
human tRNAs. RNA. 16:1317–1327. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Dominissini D, Nachtergaele S,
Moshitch-Moshkovitz S, Peer E, Kol N, Ben-Haim MS, Dai Q, Di Segni
A, Salmon-Divon M, Clark WC, et al: The dynamic
N(1)-methyladenosine methylome in eukaryotic messenger RNA. Nature.
530:441–446. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Trixl L and Lusser A: The dynamic RNA
modification 5-methylcytosine and its emerging role as an
epitranscriptomic mark. Wiley Interdiscip Rev RNA. 10. pp.
e15102019, View Article : Google Scholar
|
|
26
|
Zhong H, Tang HF and Kai Y:
N6-methyladenine RNA modification (m6A): An emerging
regulator of metabolic diseases. Curr Drug Targets. 21:1056–1067.
2020. View Article : Google Scholar
|
|
27
|
Roundtree IA, Evans ME, Pan T and He C:
Dynamic RNA modifications in gene expression regulation. Cell.
169:1187–1200. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yao ZT, Yang YM, Sun MM, He Y, Liao L,
Chen KS and Li B: New insights into the interplay between long
non-coding RNAs and RNA-binding proteins in cancer. Cancer Commun
(Lond). 42:117–140. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Alarcón CR, Lee H, Goodarzi H, Halberg N
and Tavazoie SF: N6-methyladenosine marks primary microRNAs for
processing. Nature. 519:482–485. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Du Y, Hou G, Zhang H, Dou J, He J, Guo Y,
Li L, Chen R, Wang Y, Deng R, et al: SUMOylation of the m6A-RNA
methyltransferase METTL3 modulates its function. Nucleic Acids Res.
46:5195–5208. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
van Tran N, Ernst FGM, Hawley BR, Zorbas
C, Ulryck N, Hackert P, Bohnsack KE, Bohnsack MT, Jaffrey SR,
Graille M and Lafontaine DLJ: The human 18S rRNA m6A
methyltransferase METTL5 is stabilized by TRMT112. Nucleic Acids
Res. 47:7719–7733. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Liu J, Yue Y, Han D, Wang X, Fu Y, Zhang
L, Jia G, Yu M, Lu Z, Deng X, et al: A METTL3-METTL14 complex
mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem
Biol. 10:93–95. 2014. View Article : Google Scholar :
|
|
33
|
Wang X, Huang J, Zou T and Yin P: Human
m6A writers: Two subunits, 2 roles. RNA Biol.
14:300–304. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang P, Doxtader KA and Nam Y: Structural
basis for cooperative function of Mettl3 and Mettl14
methyltransferases. Mol Cell. 63:306–317. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wang X, Feng J, Xue Y, Guan Z, Zhang D,
Liu Z, Gong Z, Wang Q, Huang J, Tang C, et al: Structural basis of
N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature.
534:575–578. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhou KI and Pan T: Structures of the m(6)A
methyltransferase complex: Two subunits with distinct but
coordinated roles. Mol Cell. 63:183–185. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Ping XL, Sun BF, Wang L, Xiao W, Yang X,
Wang WJ, Adhikari S, Shi Y, Lv Y, Chen YS, et al: Mammalian WTAP is
a regulatory subunit of the RNA N6-methyladenosine
methyltransferase. Cell Res. 24:177–189. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Liu S, Li Q, Li G, Zhang Q, Zhuo L, Han X,
Zhang M, Chen X, Pan T, Yan L, et al: The mechanism of
m6A methyltransferase METTL3-mediated autophagy in
reversing gefitinib resistance in NSCLC cells by β-elemene. Cell
Death Dis. 11:9692020. View Article : Google Scholar
|
|
39
|
Wang F, Zhu Y, Cai H, Liang J, Wang W,
Liao Y, Zhang Y, Wang C and Hou J: N6-methyladenosine
methyltransferase METTL14-mediated autophagy in malignant
development of oral squamous cell carcinoma. Front Oncol.
11:7384062021. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Yue B, Song C, Yang L, Cui R, Cheng X,
Zhang Z and Zhao G: METTL3-mediated N6-methyladenosine modification
is critical for epithelial-mesenchymal transition and metastasis of
gastric cancer. Mol Cancer. 18:1422019. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Chen X, Xu M, Xu X, Zeng K, Liu X, Pan B,
Li C, Sun L, Qin J, Xu T, et al: METTL14-mediated
N6-methyladenosine modification of SOX4 mRNA inhibits tumor
metastasis in colorectal cancer. Mol Cancer. 19:1062020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Han J, Wang JZ, Yang X, Yu H, Zhou R, Lu
HC, Yuan WB, Lu JC, Zhou ZJ, Lu Q, et al: METTL3 promote tumor
proliferation of bladder cancer by accelerating pri-miR221/222
maturation in m6A-dependent manner. Mol Cancer. 18:1102019.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Sun T, Wu Z, Wang X, Wang Y, Hu X, Qin W,
Lu S, Xu D, Wu Y, Chen Q, et al: LNC942 promoting METTL14-mediated
m6A methylation in breast cancer cell proliferation and
progression. Oncogene. 39:5358–5372. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Wang X, Zhao BS, Roundtree IA, Lu Z, Han
D, Ma H, Weng X, Chen K, Shi H and He C: N(6)-methyladenosine
modulates messenger RNA translation efficiency. Cell.
161:1388–1399. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhou J, Wan J, Gao X, Zhang X, Jaffrey SR
and Qian SB: Dynamic m(6)A mRNA methylation directs translational
control of heat shock response. Nature. 526:591–594. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu
PJ, Liu C and He C: YTHDF3 facilitates translation and decay of
N6-methyladenosine-modified RNA. Cell Res. 27:315–328.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Hsu PJ, Zhu Y, Ma H, Guo Y, Shi X, Liu Y,
Qi M, Lu Z, Shi H, Wang J, et al: Ythdc2 is an
N6-methyladenosine binding protein that regulates
mammalian spermatogenesis. Cell Res. 27:1115–1127. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Wu B, Su S, Patil DP, Liu H, Gan J,
Jaffrey SR and Ma J: Molecular basis for the specific and
multivariant recognitions of RNA substrates by human hnRNP A2/B1.
Nat Commun. 9:4202018. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
He L, Li H, Wu A, Peng Y, Shu G and Yin G:
Functions of N6-methyladenosine and its role in cancer. Mol Cancer.
18:1762019. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Huang H, Weng H, Sun W, Qin X, Shi H, Wu
H, Zhao BS, Mesquita A, Liu C, Yuan CL, et al: Recognition of RNA
N6-methyladenosine by IGF2BP proteins enhances mRNA
stability and translation. Nat Cell Biol. 20:285–295. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Li Q, Ni Y, Zhang L, Jiang R, Xu J, Yang
H, Hu Y, Qiu J, Pu L, Tang J and Wang X: HIF-1α-induced expression
of m6A reader YTHDF1 drives hypoxia-induced autophagy and
malignancy of hepatocellular carcinoma by promoting ATG2A and ATG14
translation. Signal Transduct Target Ther. 6:762021. View Article : Google Scholar
|
|
52
|
Chen H, Yu Y, Yang M, Huang H, Ma S, Hu J,
Xi Z, Guo H, Yao G, Yang L, et al: YTHDF1 promotes breast cancer
progression by facilitating FOXM1 translation in an m6A-dependent
manner. Cell Biosci. 12:192022. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Liu T, Wei Q, Jin J, Luo Q, Liu Y, Yang Y,
Cheng C, Li L, Pi J, Si Y, et al: The m6A reader YTHDF1 promotes
ovarian cancer progression via augmenting EIF3C translation.
Nucleic Acids Res. 48:3816–3831. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Liu L, Wu Y, Li Q, Liang J, He Q, Zhao L,
Chen J, Cheng M, Huang Z, Ren H, et al: METTL3 promotes
tumorigenesis and metastasis through BMI1 m6A
methylation in oral squamous cell carcinoma. Mol Ther.
28:2177–2190. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Wang Y, Lu JH, Wu QN, Jin Y, Wang DS, Chen
YX, Liu J, Luo XJ, Meng Q, Pu HY, et al: LncRNA LINRIS stabilizes
IGF2BP2 and promotes the aerobic glycolysis in colorectal cancer.
Mol Cancer. 18:1742019. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Zhu F, Yang T, Yao M, Shen T and Fang C:
HNRNPA2B1, as a m6A reader, promotes tumorigenesis and
metastasis of oral squamous cell carcinoma. Front Oncol.
11:7169212021. View Article : Google Scholar
|
|
57
|
Huang GZ, Wu QQ, Zheng ZN, Shao TR, Chen
YC, Zeng WS and Lv XZ: M6A-related bioinformatics analysis reveals
that HNRNPC facilitates progression of OSCC via EMT. Aging (Albany
NY). 12:11667–11684. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang
Y, Yi C, Lindahl T, Pan T, Yang YG and He C: N6-methyladenosine in
nuclear RNA is a major substrate of the obesity-associated FTO. Nat
Chem Biol. 7:885–887. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Wang F, Liao Y, Zhang M, Zhu Y, Wang W,
Cai H, Liang J, Song F, Hou C, Huang S, et al: N6-methyladenosine
demethyltransferase FTO-mediated autophagy in malignant development
of oral squamous cell carcinoma. Oncogene. 40:3885–3898. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Li X, Xie X, Gu Y, Zhang J, Song J, Cheng
X, Gao Y and Ai Y: Fat mass and obesity-associated protein
regulates tumorigenesis of arecoline-promoted human oral carcinoma.
Cancer Med. 10:6402–6415. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Wang J, Qiao Y, Sun M, Sun H, Zie F, Chang
H, Wang Y, Song J, Lai S, Yang C, et al: FTO promotes colorectal
cancer progression and chemotherapy resistance via demethylating
G6PD/PARP1. Clin Transl Med. 12:e7722022.PubMed/NCBI
|
|
62
|
Tang B, Yang Y, Kang M, Wang Y, Wang Y, Bi
Y, He S and Shimamoto F: m6A demethylase ALKBH5 inhibits
pancreatic cancer tumorigenesis by decreasing WIF-1 RNA methylation
and mediating Wnt signaling. Mol Cancer. 19:32020. View Article : Google Scholar
|
|
63
|
Meyer KD, Saletore Y, Zumbo P, Elemento O,
Mason CE and Jaffrey SR: Comprehensive analysis of mRNA methylation
reveals enrichment in 3′ UTRs and near stop codons. Cell.
149:1635–1646. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Wang Y, Zhu GQ, Tian D, Zhou CW, Li N,
Feng Y and Zeng MS: Comprehensive analysis of tumor immune
microenvironment and prognosis of m6A-related lncRNAs in gastric
cancer. BMC Cancer. 22:3162022. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Guo Y, Wang R, Li J, Song Y, Min J, Zhao
T, Hua L, Shi J, Zhang C, Ma P, et al: Comprehensive analysis of
m6A RNA methylation regulators and the immune microenvironment to
aid immunotherapy in pancreatic cancer. Front Immunol.
12:7694252021. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Liu Z, Zhong J, Zeng J, Duan X, Lu J, Sun
X, Liu Q, Liang Y, Lin Z, Zhong W, et al: Characterization of the
m6A-associated tumor immune microenvironment in prostate cancer to
aid immunotherapy. Front Immunol. 12:7351702021. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Visvanathan A, Patil V, Arora A, Hegde AS,
Arivazhagan A, Santosh V and Somasundaram K: Essential role of
METTL3-mediated m6A modification in glioma stem-like
cells maintenance and radioresistance. Oncogene. 37:522–533. 2018.
View Article : Google Scholar
|
|
68
|
Small W Jr, Bacon MA, Bajaj A, Chuang LT,
Fisher BJ, Harkenrider MM, Jhingran A, Kitchener HC, Mileshkin LR,
Viswanathan AN and Gaffney DK: Cervical cancer: A global health
crisis. Cancer. 123:2404–2412. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Zhou S, Bai ZL, Xia D, Zhao ZJ, Zhao R,
Wang YY and Zhe H: FTO regulates the chemo-radiotherapy resistance
of cervical squamous cell carcinoma (CSCC) by targeting β-catenin
through mRNA demethylation. Mol Carcinog. 57:590–597. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Sheng H, Li Z, Su S, Sun W, Zhang X, Li L,
Li J, Liu S, Lu B, Zhang S and Shan C: YTH domain family 2 promotes
lung cancer cell growth by facilitating 6-phosphogluconate
dehydrogenase mRNA translation. Carcinogenesis. 41:541–550. 2020.
View Article : Google Scholar
|
|
71
|
Ding RB, Chen P, Rajendran BK, Lyu X, Wang
H, Bao J, Zeng J, Hao W, Sun H, Wong AH, et al: Molecular landscape
and subtype-specific therapeutic response of nasopharyngeal
carcinoma revealed by integrative pharmacogenomics. Nat Commun.
12:30462021. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Zhang P, He Q, Lei Y, Li Y, Wen X, Hong M,
Zhang J, Ren X, Wang Y, Yang X, et al: m6A-mediated
ZNF750 repression facilitates nasopharyngeal carcinoma progression.
Cell Death Dis. 9:11692018. View Article : Google Scholar
|
|
73
|
Jin S, Li M, Chang H, Wang R, Zhang Z,
Zhang J, He Y and Ma H: The m6A demethylase ALKBH5 promotes tumor
progression by inhibiting RIG-I expression and interferon alpha
production through the IKKε/TBK1/IRF3 pathway in head and neck
squamous cell carcinoma. Mol Cancer. 21:972022. View Article : Google Scholar
|
|
74
|
Yu D, Pan M, Li Y, Lu T, Wang Z, Liu C and
Hu G: RNA N6-methyladenosine reader IGF2BP2 promotes lymphatic
metastasis and epithelial-mesenchymal transition of head and neck
squamous carcinoma cells via stabilizing slug mRNA in an
m6A-dependent manner. J Exp Clin Cancer Res. 41:62022. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Hirayama M, Wei FY, Chujo T, Oki S, Yakita
M, Kobayashi D, Araki N, Takahashi N, Yoshida R, Nakayama H and
Tomizawa K: FTO demethylates cyclin D1 mRNA and controls cell-cycle
progression. Cell Rep. 31:1074642020. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Vu LP, Pickering BF, Cheng Y, Zaccara S,
Nguyen D, Minuesa G, Chou T, Chow A, Saletore Y, MacKay M, et al:
The N6-methyladenosine (m6A)-forming enzyme
METTL3 controls myeloid differentiation of normal hematopoietic and
leukemia cells. Nat Med. 23:1369–1376. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Jiang F, Tang X, Tang C, Hua Z, Ke M, Wang
C, Zhao J, Gao S, Jurczyszyn A, Janz S, et al: HNRNPA2B1 promotes
multiple myeloma progression by increasing AKT3 expression via
m6A-dependent stabilization of ILF3 mRNA. J Hematol Oncol.
14:542021. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Han H, Fan G, Song S, Jiang Y, Qian C,
Zhang W, Su Q, Xue X, Zhuang W and Li B: piRNA-30473 contributes to
tumorigenesis and poor prognosis by regulating m6A RNA methylation
in DLBCL. Blood. 137:1603–1614. 2021. View Article : Google Scholar
|
|
79
|
Tang D, Kang R, Berghe TV, Vandenabeele P
and Kroemer G: The molecular machinery of regulated cell death.
Cell Res. 29:347–364. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Koren E and Fuchs Y: Modes of regulated
cell death in cancer. Cancer Discov. 11:245–265. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Zhi Y, Zhang S, Zi M, Wang Y, Liu Y, Zhang
M, Shi L, Yan Q, Zeng Z, Ziong W, et al: Potential applications of
N6-methyladenosine modification in the prognosis and treatment of
cancers via modulating apoptosis, autophagy, and ferroptosis. Wiley
Interdiscip Rev RNA. 13. pp. e17192022, View Article : Google Scholar
|
|
82
|
Wong RS: Apoptosis in cancer: From
pathogenesis to treatment. J Exp Clin Cancer Res. 30:872011.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Niu Y, Lin Z, Wan A, Chen H, Liang H, Sun
L, Wang Y, Li X, Xiong XF, Wei B, et al: RNA N6-methyladenosine
demethylase FTO promotes breast tumor progression through
inhibiting BNIP3. Mol Cancer. 18:462019. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Rebucci M, Sermeus A, Leonard E, Delaive
E, Dieu M, Fransolet M, Arnould T and Michiels C: miRNA-196b
inhibits cell proliferation and induces apoptosis in HepG2 cells by
targeting IGF2BP1. Mol Cancer. 14:792015. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Mizushima N and Levine B: Autophagy in
human diseases. N Engl J Med. 383:1564–1576. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Mizushima N and Komatsu M: Autophagy:
Renovation of cells and tissues. Cell. 147:728–741. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Dikic I and Elazar Z: Mechanism and
medical implications of mammalian autophagy. Nat Rev Mol Cell Biol.
19:349–364. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Amaravadi RK, Kimmelman AC and Debnath J:
Targeting autophagy in cancer: Recent advances and future
directions. Cancer Discov. 9:1167–1181. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Ferro F, Servais S, Besson P, Roger S,
Dumas JF and Brisson L: Autophagy and mitophagy in cancer metabolic
remodelling. Semin Cell Dev Biol. 98:129–138. 2020. View Article : Google Scholar
|
|
90
|
Li X, He S and Ma B: Autophagy and
autophagy-related proteins in cancer. Mol Cancer. 19:122020.
View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Chen Y and Gibson SB: Three dimensions of
autophagy in regulating tumor growth: Cell survival/death, cell
proliferation, and tumor dormancy. Biochim Biophys Acta Mol Basis
Dis. 1867:1662652021. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Lei G, Zhuang L and Gan B: Targeting
ferroptosis as a vulnerability in cancer. Nat Rev Cancer.
22:381–396. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Li T, Wang Y, Xiang X and Chen C:
Development and validation of a ferroptosis-related lncRNAs
prognosis model in oral squamous cell carcinoma. Front Genet.
13:8479402022. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Gong Y, Fan Z, Luo G, Yang C, Huang Q, Fan
K, Cheng H, Jin K, Ni Q, Yu X and Liu C: The role of necroptosis in
cancer biology and therapy. Mol Cancer. 18:1002019. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Pasparakis M and Vandenabeele P:
Necroptosis and its role in inflammation. Nature. 517:311–320.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Li J, Huang S, Zeng L, Li K, Yang L, Gao
S, Guan C, Zhang S, Lao X, Liao G and Liang Y: Necroptosis in head
and neck squamous cell carcinoma: Characterization of
clinicopathological relevance and in vitro cell model. Cell Death
Dis. 11:3912020. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Shi J, Liu Z and Xu Q: Tumor necrosis
factor receptor-associated factor 6 contributes to malignant
behavior of human cancers through promoting AKT ubiquitination and
phosphorylation. Cancer Sci. 110:1909–1920. 2019.PubMed/NCBI
|
|
98
|
Lan H, Liu Y, Liu J, Wang X, Guan Z, Du J
and Jin K: Tumor-associated macrophages promote oxaliplatin
resistance via METTL3-mediated m6A of TRAF5 and
necroptosis in colorectal cancer. Mol Pharm. 18:1026–1037. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Yu P, Zhang X, Liu N, Tang L, Peng C and
Chen X: Pyroptosis: Mechanisms and diseases. Signal Transduct
Target Ther. 6:1282021. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Yue E, Tuguzbaeva G, Chen X, Qin Y, Li A,
Sun X, Dong C, Liu Y, Yu Y, Zahra SM, et al: Anthocyanin is
involved in the activation of pyroptosis in oral squamous cell
carcinoma. Phytomedicine. 56:286–294. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Wu L, Liu G, He YW, Chen R and Wu ZY:
Identification of a pyroptosis-associated long non-coding RNA
signature for predicting the immune status and prognosis in skin
cutaneous melanoma. Eur Rev Med Pharmacol Sci. 25:5597–5609.
2021.PubMed/NCBI
|
|
102
|
Deshpande A, Sicinski P and Hinds PW:
Cyclins and cdks in development and cancer: A perspective.
Oncogene. 24:2909–2915. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Thakur C and Chen F: Connections between
metabolism and epigenetics in cancers. Semin Cancer Biol. 57:52–58.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Xu T, Zhang W, Chai L, Liu C, Zhang S and
Xu T: Methyltransferase-like 3-induced N6-methyladenosine
upregulation promotes oral squamous cell carcinoma by through p38.
Oral Dis. Sep 3–2021.Epub ahead of print. View Article : Google Scholar
|
|
105
|
Zhao W, Cui Y, Liu L, Ma X, Qi X, Wang Y,
Liu Z, Ma S, Liu J and Wu J: METTL3 facilitates oral squamous cell
carcinoma tumorigenesis by enhancing c-Myc stability via
YTHDF1-mediated m6A modification. Mol Ther Nucleic
Acids. 20:1–12. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Ai Y, Liu S, Luo H, Wu S, Wei H, Tang Z,
Li X, Lv X and Zou C: METTL3 intensifies the progress of oral
squamous cell carcinoma via modulating the m6A amount of PRMT5 and
PD-L1. J Immunol Res. 2021:61495582021. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Pastushenko I and Blanpain C: EMT
transition states during tumor progression and metastasis. Trends
Cell Biol. 29:212–226. 2019. View Article : Google Scholar
|
|
108
|
Trédan O, Galmarini CM, Patel K and
Tannock IF: Drug resistance and the solid tumor microenvironment. J
Natl Cancer Inst. 99:1441–1454. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Shibue T and Weinberg RA: EMT, CSCs, and
drug resistance: The mechanistic link and clinical implications.
Nat Rev Clin Oncol. 14:611–629. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Shriwas O, Priyadarshini M, Samal SK, Rath
R, Panda S, Das Majumdar SK, Muduly DK, Botlagunta M and Dash R:
DDX3 modulates cisplatin resistance in OSCC through ALKBH5-mediated
m6A-demethylation of FOXM1 and NANOG. Apoptosis.
25:233–246. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Bellmunt J, Powles T and Vogelzang NJ: A
review on the evolution of PD-1/PD-L1 immunotherapy for bladder
cancer: The future is now. Cancer Treat Rev. 54:58–67. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Li N, Kang Y, Wang L, Huff S, Tang R, Hui
H, Agrawal K, Gonzalez GM, Wang Y, Patel SP and Rana TM: ALKBH5
regulates anti-PD-1 therapy response by modulating lactate and
suppressive immune cell accumulation in tumor microenvironment.
Proc Natl Acad Sci USA. 117:20159–20170. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Chen J, Lu T, Zhong F, Lv Q, Fang M, Tu Z,
Ji Y, Li J and Gong X: A signature of N6-methyladenosine
regulator-related genes predicts prognoses and immune responses for
head and neck squamous cell carcinoma. Front Immunol.
13:8098722022. View Article : Google Scholar
|
|
114
|
Shu CW, Weng JR, Chang HW, Liu PF, Chen
JJ, Peng CC, Huang JW, Lin WY and Yen CY: Tribulus terrestris fruit
extract inhibits autophagic flux to diminish cell proliferation and
metastatic characteristics of oral cancer cells. Environ Toxicol.
36:1173–1180. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Kumar VB, Lin SH, Mahalakshmi B, Lo YS,
Lin CC, Chuang YC, Hsieh MJ and Chen MK: Sodium danshensu inhibits
oral cancer cell migration and invasion by modulating p38 signaling
pathway. Front Endocrinol (Lausanne). 11:5684362020. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Balaji S, Terrero D, Tiwari AK, Ashby CR
Jr and Raman D: Alternative approaches to overcome chemoresistance
to apoptosis in cancer. Adv Protein Chem Struct Biol. 126:91–122.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Luo R, Xie L, Lin Y, Shao J and Lin Z:
Oxymatrine suppresses oral squamous cell carcinoma progression by
suppressing CXC chemokine receptor 4 in an m6A
modification decrease dependent manner. Oncol Rep. 48:1772022.
View Article : Google Scholar
|
|
118
|
Gong J, Wang C, Zhang F and Lan W: Effects
of allocryptopine on the proliferation and epithelial-mesenchymal
transition of oral squamous cell carcinoma through m6A mediated
hedgehog signaling pathway. J Environ Pathol Toxicol Oncol.
41:15–24. 2022. View Article : Google Scholar : PubMed/NCBI
|