Introduction
One of the most fundamental causes of cancer-related
mortality and morbidity among males is prostate cancer (PCa)
(1), with higher incidence rates
observed in more developed countries (2,3). The
etiology of PCa is complex and is dependent on a wide range of
factors, such as age, genetics, lifestyle and environmental
conditions (4,5). Although some new methods for the
diagnosis and treatment of cancer have been developed, they are
still in the experimental stage (6-8). The
primary treatment for PCa is androgen deprivation therapy, which is
associated with curative efficacy for those cases with early-stage
disease (9,10). However, PCa is generally associated
with poor outcomes attributable to poor screening efforts, low
rates of successful curative treatment and high rates of
therapeutic resistance. There is thus an urgent need to more
clearly define the mechanistic basis for this disease, in order to
identify novel biomarkers for improve patient outcomes.
The dynamic, reversible
N6-methyladenosine (m6A) modification is the
most commonly identified epigenetic alteration present in
eukaryotic RNAs (11). Several
enzymes regulate the m6A modification (12), and the dysregulation of this
process has been found to be associated with oncogenesis and
malignant tumor behaviors owing to alterations in the expression
levels of specific tumor suppressor genes or oncogenes within human
cells (13,14). Long non-coding RNAs (lncRNAs) are
transcripts of >200 nucleotides in length that have no coding
potential (15). lncRNAs can
regulate critical cellular processes, including autophagy,
apoptosis, differentiation, invasion and proliferation (16,17).
Functionally, these lncRNAs can interact with RNAs and proteins to
alter RNA splicing, RNA stability, microRNAs (miRNAs/miRs)
activity, histone modifications, transcription, or translational
activity (18). It has been
demonstrated that lncRNAs are able to function as competing
endogenous RNAs (ceRNAs), which can bind to specific miRNAs in a
sequence-dependent manner, thereby sequestering them and indirectly
regulating target gene expression (19,20).
Furthermore, miRNAs can in turn regulate lncRNA expression
(21). However, additional
explorations are necessary for the meticulous understanding of the
interaction among m6A modification, lncRNAs, miRNAs and
the progression of PCa.
The expression of lncRNA plasmacytoma variant
translocation 1 (PVT1), located at 8q24.21 (22), is upregulated in PCa and multiple
types of cancer (23,24). Despite such an upregulation, the
specific mechanisms whereby PVT1 influences oncogenic activity have
not yet been fully explored. Zhao et al (25) reported that STAT3 promoted PVT1
transcription by binding to specific sequences within its promoter
region. Chen et al (26)
found that the ALKBH5-mediated m6A modification of PVT1
contributed to the tumorigenesis of osteosarcoma. However, to date,
at least to the best of our knowledge, no study has determined
whether there is a similar association between the m6A
modification status and PVT1 overexpression in PCa. Simiar to other
lncRNAs, PVT1 is able to function as a ceRNA (27). In a previous study, PVT1 was shown
to suppress miR-30d-5p, thereby enhancing gallbladder cancer cell
proliferation and metastatic activity (28). Contrarily, another study reported
the interaction of PVT1 with miR-619-5p within PCa cells in a
manner conducive to tumor growth and gemcitabine resistance
(29). Additional research,
however, is essential to fully elucidate the miRNA-dependent
mechanisms whereby PVT1 influences PCa cell behavior.
In the present study, it was hypothesized that PVT1
dysregulation may influence PCa onset and progression. Through a
series of assays, it was ultimately found that PVT1 was capable of
promoting PCa progression, in part by serving as a ceRNA for
miR-27b-3p, thereby indirectly promoting bloom syndrome protein
(BLM) upregulation. In addition, the methyltransferase 3
(METTL3)-mediated m6A modification of PVT1 promoted its
upregulation, influencing the PCa progression through the
miR-27b-3p/BLM axis.
Materials and methods
Bioinformatics analysis
On the basis of the Oncomine (www.oncomine.org) and Gene Expression Omnibus (GEO;
https://www.ncbi.nlm.nih.gov/geo/)
databases, the differential expression of PVT1 between PCa tumors
compared with healthy prostate tissues was analyzed. Based on The
Cancer Genome Atlas (TCGA) database, the associations between PVT1
expression with sample type, Gleason scores, objective response
rate and overall survival in PCa were assessed; moreover, the
diagnostic and prognostic values of PVT1 were analyzed using
receiver operating characteristic curve (ROC) analysis, which
determined the area under curve (AUC).
The miRanda (https://tools4mirs.org/software/target_prediction/miranda/),
miRwalk (http://mirwalk.umm.uni-heidelberg.de/), miRDB
(https://mirdb.org/), RNA22 (https://cm.jefferson.edu/rna22/) and TargetScan
(https://www.targetscan.org/vert_80/)
databases were exploited to predict the target genes of miRNA.
Potential miRNA binding partners for PVT1 were identified using the
miRanda and the LncBase Predicted (ver. 2) online databases. The
miRDB platform was applied to predict the target genes of
miRNA.
Clinical samples
Tumor specimens and paired paracancerous tissues
from patients with PCa were obtained from the Affiliated Hospital
of Zunyi Medical University (Zunyi, China). A total of 12 samples
were collected, including three paracancerous tissues, three benign
prostate hyperplasia (BPH) tissues, three tissues with Gleason
scores ≤7 and three tissues with Gleason scores >7. The
clinicopathological data of the patients are presented in Table I. The study protocol was approved
by the Ethics Committee of Affiliated Hospital of Zunyi Medical
University [Compliance Medical Ethics Review (2021) no. 1-037].
Written informed consent was obtained from all participants prior
to enrollment.
 | Table IBaseline characteristics of the
patients in the present study. |
Table I
Baseline characteristics of the
patients in the present study.
| Tissue type | Case no. | Age, years | PSA | Volume (mm) | Gleason | Transfer |
|---|
| BPH | 1 | 81 | 0.962 | 53×56×48 | - | - |
| 2 | 83 | 1.232 | 61×54×60 | - | - |
| 3 | 70 | 1.442 | 50×50×48 | - | - |
| PCa | 4 | 76 | 5.47 | 45×48×38 | 3+3 | No |
| 5 | 70 | 2.23 | 35×46×29 | 3+3 | No |
| 6 | 78 | 1.28 | 34×43×23 | 3+3 | No |
| 7 | 71 | 68.48 | 35×35×37 | 4+5 | Yes |
| 8 | 78 | 525.56 | 32×47×30 | 4+3 | Yes |
| 9 | 75 | 118.1 | 51×78×47 | 4+5 | Yes |
Cells and cell culture
LNCaP and PC3 human PCa cells and RWPE-2 cell lines
were acquired from the American Type Culture Collection (ATCC) and
cultured in a Dulbecco's modified Eagle's medium (DMEM) containing
penicillin/streptomycin (Corning Inc.) and 10% fetal bovine serum
(FBS; Gibco; Thermo Fisher Scientific, Inc.) in a 5% CO2
humidified incubator at 37°C.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
TRIzol reagent was used for the extraction of total
RNA using a reverse transcription kit (TransGen Biotech Co., Ltd.)
according to the manufacturer's instructions, and it was
subsequently used to prepare cRNA. qPCR was performed on a
LightCycler System 2.0 (Roche Diagnostics GmbH) with the following
thermocycling conditions: Initial pre-denaturation for 3 min at
94°C, followed by 40 cycles of 95°C with 10 sec, 60°C for 10 sec
and 76°C for 20 sec; the evaluation of the solubility curve was
performed at 95°C for 5 sec and 65°C for 1 min, which was followed
by cooling at 42°C for 30 sec. All experiments were performed in
triplicate, and the relative gene expression was calculated using
the 2−ΔΔCq method (30). All the primers used for RT-qPCR are
listed in Table SI.
Transfection assay
PVT1 or METTL3 knockdown was achieved using two PVT1
or METTL3-specific shRNAs (PVT1-sh1 and PVT1-sh2, METTL3-sh1 and
METTL3-sh2), with a non-targeting shRNA (NC-sh) serving as a
control (D03003, GenePharma Co., Ltd.). For PVT1 or BLM
overexpression, the cDNA encoding PVT1 or BLM was reinforced via
PCR and subcloned into the pcDNA3.1 vector (Invitrogen; Thermo
Fisher Scientific, Inc.) to construct the pcDNA-PVT1 vector, with
an empty vector serving as a negative control (pcDNA-NC). The
miR-27b-3p mimic (5′-UUC ACA GUG GCU AAG UUC UGC-3′) and the
appropriate negative control mimic (NC mimic, 5′-UUU GUA CUA CAC
AAA AGU ACU G-3′) were obtained from Guangzhou RiboBio Co., Ltd.
The cells plated in six-well plates at a density of
1×106 cells/well were transfected with the
aforementioned shRNAs, plasmids, or mimics (PVT1-specific shRNAs
and NC-sh, 50 nM; pcDNA-PVT1, pcDNA-BLM, and pcDNA-NC, 2 µg;
miR-27b-3p mimic and NC mimic, 50 nM) using Lipofectamine
2000® (Invitrogen; Thermo Fisher Scientific, Inc.)
according to the manufacturer's instructions. The cells were then
used in subsequent experiments at 48-72 h post-transfection, and
RT-qPCR was employed to verify the efficiency of transfection. All
shRNA sequences are listed in Table
SII.
Tumor xenograft assays
A total of 20 male 4-week-old BALB/c nude mice
(weighing 18-22 g) were obtained from Zunyi Medical University and
maintained under specific pathogen-free conditions (temperature,
24±2°C; humidity, 50±15%) with free access to food and water in
laminar airflow cabinets. Following 1 week for adaptation, a total
of 20 mice were randomly divided into four groups to establish a
mouse xenograft model. Briefly, the mice were subcutaneously
implanted with control tumor cells or those that had been stably
transfected with lenti-NC-sh or lenti-sh-PVT1 and pcDNA-BLM vector
(1×107 cells) in 200 µl phosphate-buffered saline
(PBS). After 7 days, tumor growth was controlled and tumor volume
(mm3) was measured as follows: Tumor volume=(length x
width2)/2. To ensure compliance with humane endpoints,
the tumor diameter of the xenografts did not exceed 2 cm and the
tumor volume was <4,400 mm3. The mice were sacrificed
6 weeks later by cervical dislocation, and tumors were therefore
harvested and weighed. The death of the mice was verified by signs
of no breathing and no corneal reflex. The Guidelines for the Care
and Use of Laboratory Animals were used to design all experiments,
which received confirmation from the Ethics Committee of the
Affiliated Hospital of Zunyi Medical University
(ZMU21-2205-002).
All animal welfare considerations were taken,
including efforts to minimize suffering and distress or special
housing conditions. Animal health and behavior were monitored every
day. According to the AVMA Guidelines on Euthanasia, if any
ulceration or infection occurs in the tumor site, or the existed
tumor interfere with eating or free movement, the animals should be
euthanized.
Cell proliferation assay
Cell Counting Kit-8 (CCK-8) and colony formation
assays were employed to assess cell proliferation. In CCK-8 assay,
cells were seeded into 96-well plates (density, 2,000 cells/well)
and incubated for 24, 48 and 72 h at 37°C, and 10 µl CCK-8
reagent (Dojindo Laboratories, Inc.) were then added, followed by
an additional incubation for 1 h at 37°C. The absorbance was then
measured at 450 nm.
Clonogenic assay
The PCa cells were cultured in DMEM supplemented
with 10% FBS (1,000 cells/ml), and then seeded into 6-cm culture
plates for 2-3 weeks. The culture medium was then changed and the
cells were rinsed three times with PBS. Following fixation with 4%
paraformaldehyde (PFA) for 15 min at room temperature, the cells
were stained with Giemsa (Beyotime Institute of Biotechnology) for
20 min at room temperature, and all colonies consisting of >50
cells were counted under a microscope (Leica Microsystems GmbH)
(31).
Cell migration and invasion assays
Wound healing assays were employed for measuring
cell migration as previously described (32). Briefly, the cells were seeded in
six-well plates at a confluency of 100%, and a 20 µl pipette
tip was then used to generate a linear scratch wound in the cell
surface. The cells were then further cultured for 48 h in
serum-free medium. Wound closure was then imaged using an Olympus
1X71 camera system (Olympus Corporation) at 0 and 48 h
post-wounding.
Transwell assays were used to assess cell invasion.
Briefly, the cells were added to Transwell inserts (8-µm
pore size, 5×104 cells/well), and DMEM containing 20%
FBS was added to the lower chamber. The cells were then incubated
for 24 h at 37°C, in which a cotton swab was utilized for the
elimination of non-invasive cells, and the remaining cells were
fixed with 4% PFA for 20 min at room temperature, stained with
hematoxylin (Beyotime Institute of Biotechnology) for 15 min at
room temperature, and counted under a light microscope (Olympus
Corporation).
Western blot analysis
RIPA buffer (Beyotime Institute of Biotechnology)
with protease inhibitor cocktail (Roche Diagnostics Inc.) was used
for the extraction of proteins from cells. The protein
concentration was determined using a BCA kit (Beyotime Institute of
Biotechnology). Proteins were subsequently separated through 10%
sodium dodecyl sulfate-polyacrylamide gel electrophoresis
(SDS-PAGE), transferred onto polyvinylidene difluoride (PVDF)
membranes (Millipore Sigma), and the blots were blocked with 5%
non-fat milk in TBST prior to a 12-h incubation with antibodies
specific for rabbit cyclin D1 (1:1,000, cat. no. 55506), CDK6
(1:1,000, cat. no. 13331), CDK4 (1:1,000, cat. no. 12790), Bax
(1:1,000, cat. no. 5023), Bcl-2 (1:1,000, cat. no. 4223), cleaved
caspase-3 (1:1,000, cat. no. 9664), cleaved caspase-9 (1:1,000,
cat. no. 7237), E-cadherin (1:1,000, cat. no. 3195), N-cadherin
(1:1,000, cat. no. 13116), Vimentin (1:1,000, cat. no. 5741S),
METTL3 (1:1,000, cat. no. 96391), BLM (1:1,500, cat. no. 2742),
c-Myc (1:1,000, cat. no. 9402), or Slug (1:1,000, cat. no. 9585S)
(all form Cell Signaling Technology, Inc.) for 12 h at 4°C.
Anti-mouse β-actin (1:5,000, cat. no. 81115-1-RR, ProteinTech
Group, Inc.) served as a loading control, of which incubation
conditions were as follows: 12 h at 4°C. Subsequently, the
membranes were incubated with appropriate species-specific
secondary antibodies (1:5,000, cat. no. A0208, Beyotime Institute
of Biotechnology) for 2 h at room temperature before the
visualization of protein bands using an ECL Western Blotting
Substrate kit (cat. no. ab65623, Abcam).
Immunohistochemical staining
Paraffin-embedded tissues sections were
deparaffinized, exposed to antigen retrieval, treated for blocking
endogenous peroxidase activities, blocked using goal serum (Gibco;
Thermo Fisher Scientific, Inc.), and incubated with antibodies
specific for BLM (1:200; cat. no. 15161-1-AP, ProteinTech Group,
Inc.), proliferating cell nuclear antigen (PCNA; 1:200, cat. no.
10205-2-AP; ProteinTech Group, Inc.), cleaved caspase-9 (1:200;
cat. no. 9509, Cell Signaling Technology, Inc.), or N-cadherin
(1:400; cat. no. 13116; Cell Signaling Technology, Inc.) overnight
at 4°C, followed by 2 h-incubation with an appropriate secondary
antibody (1:4,000; cat. no. ab205718; Abcam) at room temperature,
and imaging using a phase-contrast microscope (cat. no. DMi1, Leica
Microsystems GmbH).
Dual-luciferase reporter assay
Bioinformatics tools were employed to estimate the
complementary sites of binding for miR-27b-3p within PVT1 and the
BLM 3′-UTR. WT or mutated (MUT) versions of the predicted
miR-27b-3p binding site were cloned into the pmirGLO vector
(Promega Corporation) to prepare a reporter construct. The PC3 and
LNCap cells were subsequently co-transfected with 2 µg of
these reporter constructs and either 50 nM of miR-27b-3p mimics or
corresponding controls using Lipofectamine 2000®
(Invitrogen; Thermo Fisher Scientific, Inc.). Moreover, at 48 h
post-transfection, using a dual-luciferase reporter assay system
(Promega Corporation), the Firefly fluorescence was normalized to
the Renilla fluorescence to quantify the luciferase
activity.
RNA immunoprecipitation (RIP)
The RIP assay was performed using a Thermo Fisher
RIP kit (Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. Briefly, RIP lysis buffer was utilized
to lyse the cells, followed by lysate incubation with human
anti-AGO2 (1:1,000, cat. no. 2897, Cell Signaling Technology,
Inc.), anti-BLM (1:1,500, cat. no. 2742, Cell Signaling Technology,
Inc.), or control mouse anti-IgG (1:2,000, cat. no. 7076, Cell
Signaling Technology, Inc.) antibodies. RNA was then accumulated
and assessed using RT-qPCR.
RNA pull-down assay
RNA pull-down assay was performed using synthesized
PVT1 as a probe to detect miR-27b-3p and BLM by RT-qPCR. First, the
DNA fragment with the whole PVT1 sequence was amplified by PCR
using a T7-containing primer. The amplified products were then
cloned into pRCR8 (Invitrogen; Thermo Fisher Scientific, Inc.). The
biotin-labeled PVT1 was transcribed with the Biotin RNA Labeling
Mix In Vitro (Roche Diagnostics) and T7 RNA polymerase
(Roche Diagnostics). Subsequently, the samples were treated with
RNase-free DNase I (Roche Diagnostics) and recycled using the QIA
quick Nucleotide Removal kit (Qiagen Inc.). They were then purified
using the RNeasy Mini kit (Qiagen Inc.). The PC3 and LNCap cells
were transfected with pcDNA-PVT1 and pcDNA-NC prior to gathering
for a 10-min incubation with specific cell lysates at room
temperature. The cell lysates were mixed with biotin-labeled PVT1
(Wuhan Boster Biological Technology, Ltd.,) and then incubated for
1 h at 4°C. Subsequently, streptavidin agarose beads (Invitrogen;
Thermo Fisher Scientific, Inc.) were added into each binding
reaction according to the manufacturer's protocol and the mixture
was incubated for 1 h at 25°C. After washing with the wash buffer,
biotin-coupled RNA complex was pulled down by centrifugation at
3,000 × g for 5 min at 4°C. Elution buffers (50 µl) were
used to elute the complex. The eluate was collected, precipitated
RNA was extracted using TRIzol® reagent (MilliporeSigma)
and enriched miR-27b-3p and BLM were assessed using RT-qPCR.
Methylated RNA immunoprecipitation
Total RNA extracted from the PCa cells was
immunoprecipitated using anti-m6A (cat. no. ab151230,
Abcam) and a Magna methylated RNA immunoprecipitation (MeRIP)
m6A kit (cat. no. 17-10499, MilliporeSigma) based on the
provided instructions, following the chemical fragmentation of this
RNA to ~100 nucleotides in length. M6A-containing RNA
enrichment was then assessed using RT-qPCR.
RNA stability analysis
To block new RNA synthesis, the polymerase II
inhibitor, α-amanitin (100 nM, MilliporeSigma) was added to the
culture medium for treating the transfected cells. At the
designated time points (0, 2 and 4 h), the cells were collected and
lysed to detect the RNA levels of PVT1 using RT-qPCR analysis.
RNA-fluorescence in situ hybridization
(FISH) and subcellular fractionation
The subcellular localization of PVT1 was identified
using the FISH technique. In line with the Ribo™ lncRNA FISH Probe
Mix (Red) (Guangzhou RiboBio Co., Ltd.), the specific methods used
were as follows: A slide was placed into a 24-well culture plate.
The PC3 and LNCap cells were taken and seeded at 6×104
cells/well, reaching ~80% cell confluency. The slides were then
removed and fixed with 1 ml 4% PFA. After being treated with
protease K, glycine and ethyl phthalide reagent, the cells were
joined with 250 µl prehybridization solution and incubated
at 42°C for 1 h. The pre-hybridization solution was removed and the
21-nt FISH probes (Guangzhou RiboBio Co., Ltd., were then added
Cy3-aaaGTG AGT AGT CGG ACG GAG GA) at 37°C overnight. The cells
were hybridized overnight at 42°C. For immunofluorescence assay,
the fixed cells were incubated with the anti-Cy3 antibody (cat. no.
ab52060, Abcam) in blocking buffer (1X PBS, 0.1% Triton, and 2
mg/ml bovine serum albumin) for 30 min PBS plus with Tween-20
(PBST) for 5 min each and incubated with the secondary antibody in
blocking buffer for a further 30 min at the room temperature,
followed by DAPI staining (cat. no. ab104139, Abcam) diluted with
PBST. The cells were then observed and photographed under a
fluorescence microscope (Olympus Corporation).
Cellular fractionation was performed as follows:
Briefly, the cells were washed with ice-cold PBS, collected, spun
down and re-suspended in ice-cold buffer I (10 mM Hepes, pH 8.0,
1.5 mM MgCl2, 10 mM KCl, 1 mM DTT) supplemented with protease
inhibitor cocktail, followed by incubation for 15 min on ice to
allow the cells to swell. Igepal-CA630 (MilliporeSigma) was then
added at a final concentration of 1% (use 10% stock solution)
followed by vortexing for 10 sec. Nuclei were collected by
centrifuging 2~3 min at maximum speed 21,100 × g). The resultant
supernatant was cytosolic fraction. Nuclei were then lysed in ice
cold buffer II (20 mM Hepes, pH 8.0, 1.5 mM MgCl2, 25% (v/v)
glycerol, 420 mM NaCl, 0.2 mM EDTA, 1 mM DTT) supplemented with
protease inhibitor cocktail followed by vigorous rotation at 4°C
for 30 min and centrifugation for 15 min at 12,000 × g at 4°C. The
resultant supernatant was nuclear fraction. Both cytosolic and
nuclear RNAs were extracted by Phenol-Chloroform-Isoamyl Alcohol
mixture (cat. no. 77618, MilliporeSigma) followed by RT-qPCR
analysis.
Cycloheximide (CHX) half-life assay
At 48 h post-transfection, 20 µg/ml CHX
(MilliporeSigma) were added to the cell medium. At the designated
time points (0, 2, 4 and 8 h), the cells were collected and lysed
to detect the protein levels of BLM using western blot
analysis.
Statistical analysis
SPSS18.0 (IBM, Inc.) and GraphPad Prism 7.0
(GraphPad Software Inc.) software were used to analyze the data.
The analyses were repeated in triplicate. Survival outcomes were
compared using Kaplan-Meier plots and the log-rank test. Pearson's
correlation analysis was performed to analyze the expression of
PVT1 and its potential target miRNAs. Quantitative data are
expressed as the mean ± standard deviation (SD) and analyzed using
a Student's t-test or one-way analysis of variance followed by
Tukey's post hoc test using GraphPad Prism software. A two-sided
P-value <0.05 was considered to indicate a statistically
significant difference.
Results
The upregulation of PVT1 expression is
related to poor survival outcomes of patients with PCa
Initially, the Oncomine and GEO databases (33-42)
were used to evaluate PVT1 expression, and it was revealed that the
PVT1 expression level was significantly upregulated in PCa tumors
compared with healthy prostate tissues (Fig. 1A and B). In TCGA database, the PVT1
expression level was significantly related to the sample type and a
higher Gleason score (Fig. 1C).
Considering the association between patient clinicopathological
characteristics and the PVT1 expression level in patients with PCa,
ROC curves suggested that PVT1 was accompanied by a high
sensitivity when it was employed for the diagnosis of PCa (AUC,
0.903; Fig. 1D). Kaplan-Meier
curves also suggested that a higher PVT1 expression level was
related to more unfavorable survival outcomes of patients with PCa
(Fig. 1E). In time-dependent ROC
curve analyses, the AUC values corresponding to the 1-, 3- and
5-year survival rates were 0.990, 0.769 and 0.769, respectively
(Fig. 1F). These findings thus
indicated that PVT1 may be a prognostic biomarker for PCa.
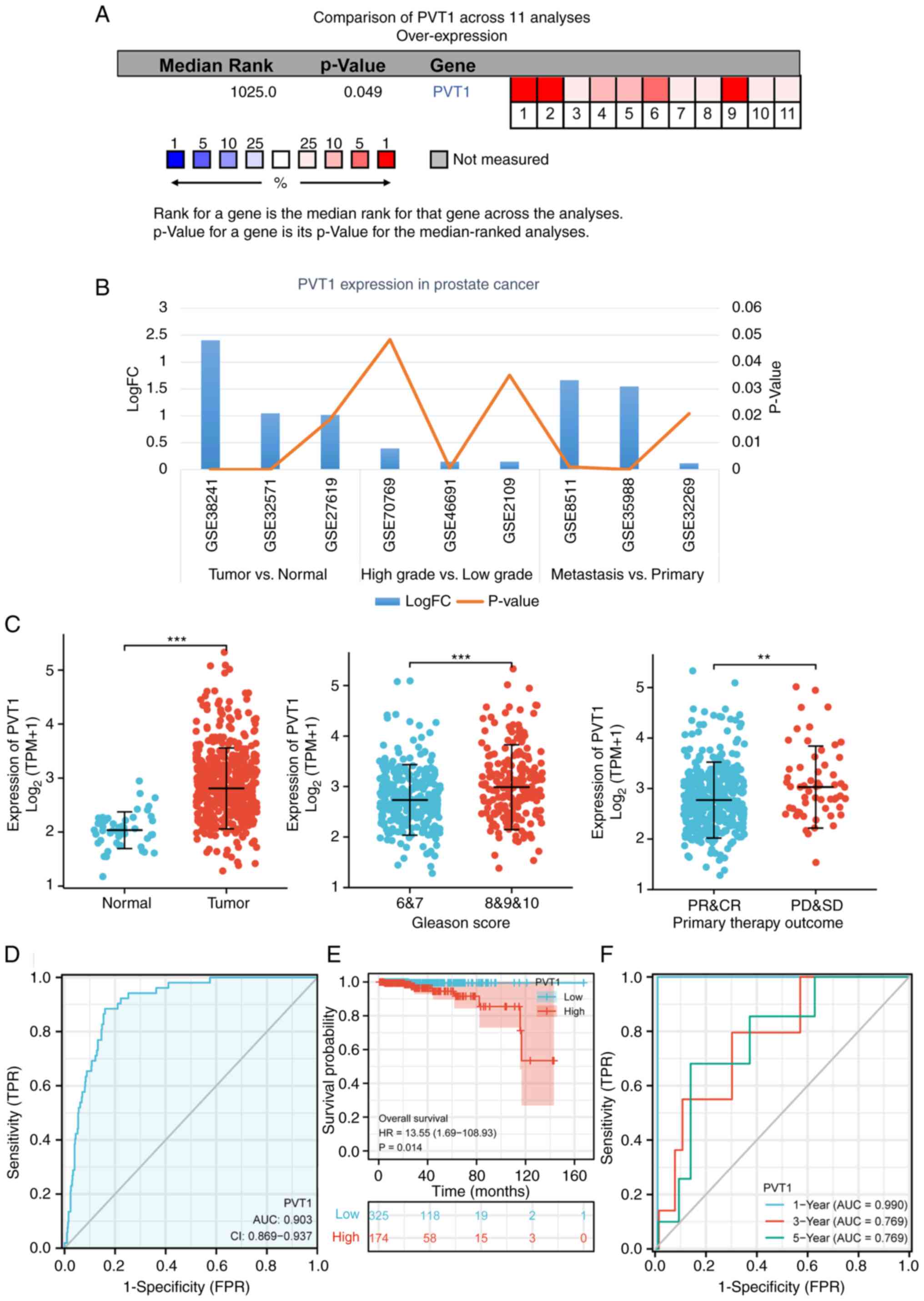 | Figure 1Analysis of the PVT1 expression level
in PCa tissues and cells. (A) PVT1 expression level in PCa and
normal tissues was detected using the Oncomine database. These
datasets, in Arabic numerical order, were on prostate carcinoma vs.
normal (33), prostate carcinoma
vs. normal (34), prostate
carcinoma vs. normal (35),
prostate carcinoma vs. normal (36), prostate carcinoma vs. normal
(37), prostate carcinoma vs.
normal (38), benign prostatic
hyperplasia stroma vs. normal (39), prostate carcinoma epithelia vs.
normal (39), prostate carcinoma
vs. normal (40), prostate
carcinoma vs. normal (41),
prostate carcinoma vs. normal (42). (B) The PVT1 expression level was
compared between tumors and control tissues, and it was dependent
on the sample type or histological grade. (C) PVT1 expression level
in PCa and paired para-cancerous tissue samples from TCGA-PRAD
database. (D) ROC curves corresponding to the PVT1 expression level
in TCGA cohort. (E) Kaplan-Meier curves exploring the association
between PVT1 expression level and overall survival of patients with
PCa. (F) Time-dependent ROC curve-based analyses of the 1-, 3- and
5-year survival rates. **P<0.01 and
***P<0.001. PVT1, plasmacytoma variant translocation
1; PCa, prostate cancer; TCGA, The Cancer Genome Atlas; PRAD,
prostate adenocarcinoma; CR, complete response; PR, partial
response; SD, stable disease; PD, progressive disease; AUC, area
under the curve; HR, hazard ratio. |
METTL3-mediated m6A
modification promotes the upregulation of PVT1 expression
The potential functional role of PVT1 in PCa was
assessed by analyzing its expression level in PCa tissue. The PVT1
expression level was markedly upregulated in PCa tumors compared
with paired paracancerous tissues or BPH samples (Fig. 2A). Similarly, the PVT1 expression
level was upregulated in the LNCap and PC3 PCa cell lines compared
with the control RWPE-2 cells (Fig.
2B).
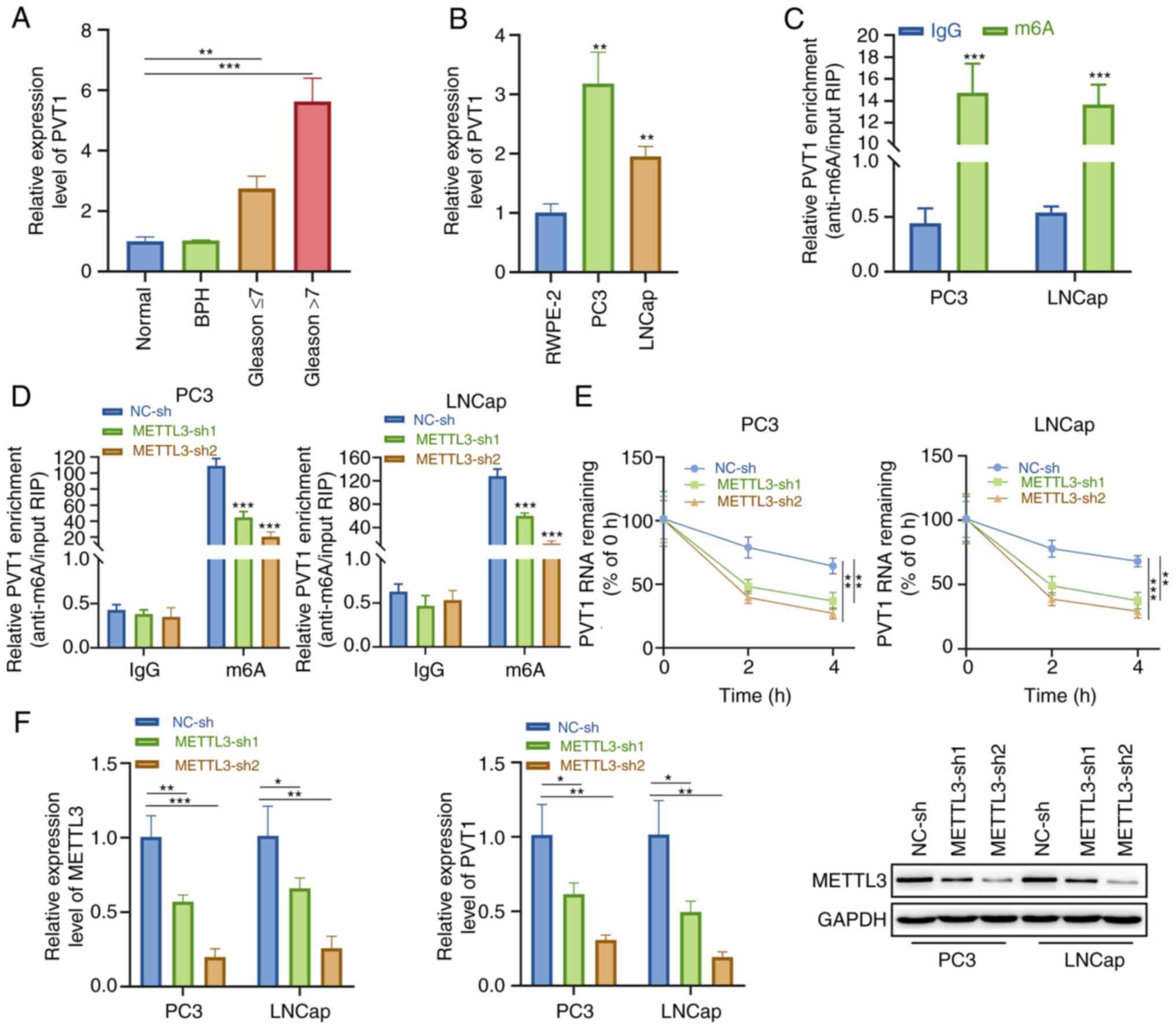 | Figure 2METTL3-mediated m6A
modification promotes PVT1 upregulation. (A) PVT1 upregulation was
observed in PCa tissues compared with healthy or BPH tissues as
determined using RT-qPCR. (B) PVT1 expression level in RWPE-2, PC3
and LNCap cells was measured using RT-qPCR. (C) Anti-m6A
antibody was used to conduct RIP assay. (D) Lentiviruses expressing
shRNAs specific for METTL3 or corresponding negative control
constructs were used to transduce PC3 and LNCap cells, and PVT1
expression level was then assessed following MeRIP-qPCR analysis
conducted with anti-m6A. (E) α-amanitin (50 mM) was used to treat
the PCa cells, in which METTL3 had or had not been knocked down,
and RT-qPCR was used to assess PVT1 stability over time compared
with baseline. (F) RT-qPCR and western blot analysis were
respectively employed for detecting the expression levels of PVT1
and METTL3 in PC3 and LNCap cells, in which METTL3 had been knocked
down. Data were from triplicate measurement and are presented as
the mean ± SD. *P<0.05, **P<0.01 and
***P<0.001. m6A, METTL3, methyltransferase
3; N6-methyladenosine; PCa, prostate cancer; RT-qPCR,
reverse transcription-quantitative polymerase chain reaction. |
Epigenetic alterations are critical regulators of
the processes governing PCa progression (43,44).
In the present study, RIP assay was thus conducted to explore the
association between the m6A modification status and PVT1
upregulation using anti-METTL3 antibody, which is specific for the
most prominent m6A-modifying enzyme. METTL3
precipitation resulted in a noticeable PVT1 enrichment compared
with control IgG precipitation in the tested PCa cell lines
(Fig. 2C), whereas this effect was
reversed following METTL3 knockdown, in which PVT1 enrichment was
attenuated compared with control IgG (Fig. 2D), suggesting a potential role of
METTL3 as a regulator of PVT1 expression level. To further confirm
this potential regulatory association, a lentivirus encoding a
METTL3-specific shRNA (METTL3-sh) was used to knockdown this enzyme
in LNCap and PC3 cells. When α-amanitin was used to treat these two
cell lines in order to disrupt the synthesis of RNA, METTL3
knockdown was found to significantly reduce the PVT1 half-life
(Fig. 2E). METTL3 knockdown was
successfully confirmed using qPCR and western blot analysis, and
when PVT1 the expression level was assessed in these cells, it was
found to be downregulated and positively associated with the METTL3
expression level (Fig. 2F),
confirming the existence of interactions between PVT1 and METTL3.
Overall, these results demonstrated that METTL3-mediated
m6A modifications may be associated with the
upregulation and stabilization of PVT1 in PCa cells.
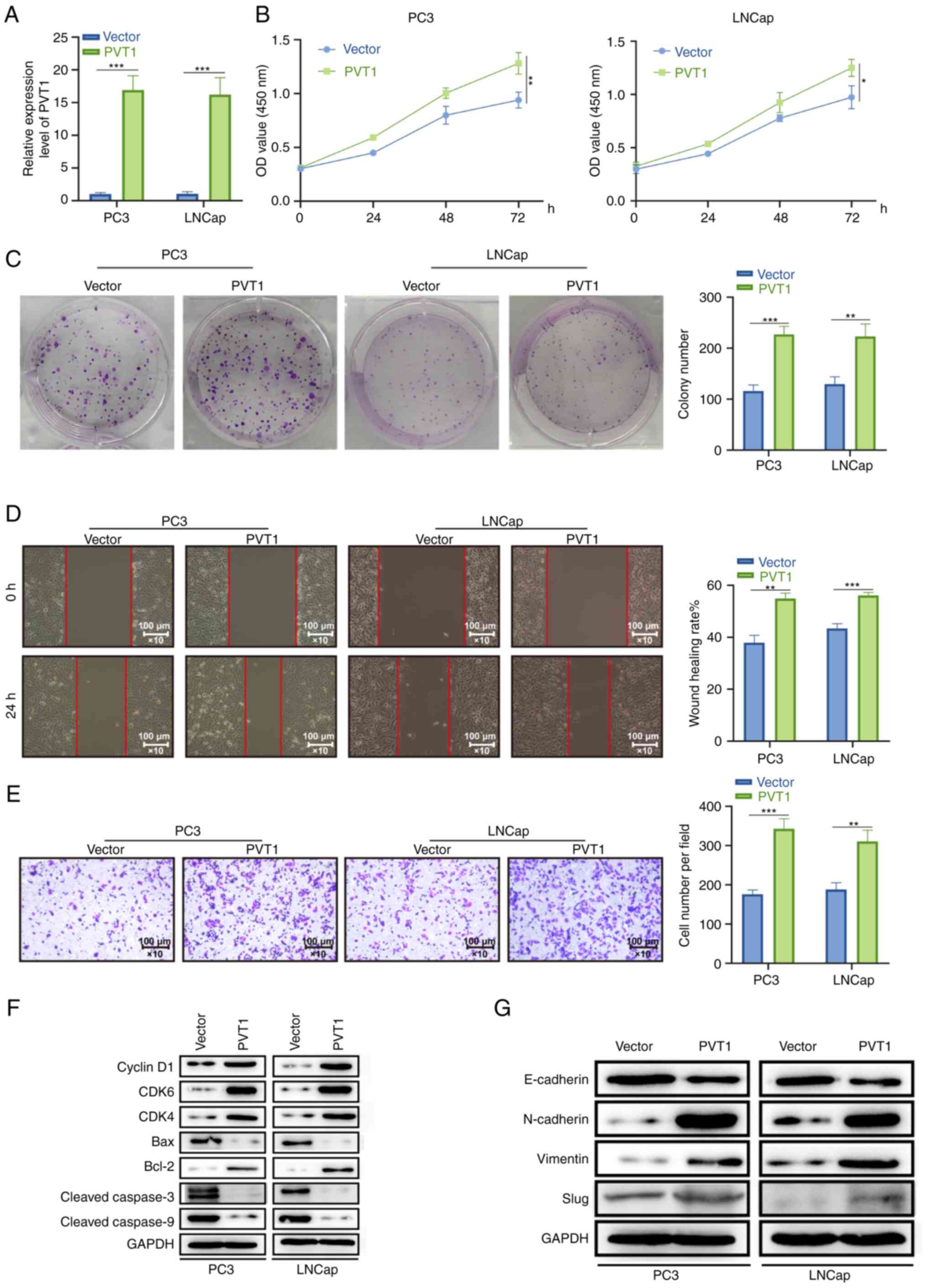
PVT1 overexpression enhances in vitro PCa
cell proliferation, apoptotic resistance, invasion and
migration
In order to explore the mechanisms through which
PVT1 can regulate PCa cell functionality, lentiviral vectors were
used to induce its overexpression in PC3 and LNCap cells, and
RT-qPCR was employed to confirm the transfection efficiency
(Fig. 3A). The overexpression of
this lncRNA was noted to be associated with significant increases
in PCA cell proliferation in CCK-8 and colony formation assays
(Fig. 3B and C). In Transwell and
wound healing assays, PVT1 overexpression was further found to
enhance the PCa cell migratory and invasive capabilities (Fig. 3D and E). When cell cycle- and
apoptosis-associated proteins were evaluated in these cells using
western blot analysis, the levels of G1/S-phase checkpoint proteins
(CDK4/6 and cyclin D1) were elevated following PVT1 overexpression,
as was consistent with the level of pro-survival oncogenic proteins
(Bcl-2), while the levels of pro-apoptotic proteins (cleaved
caspase-3/9 and Bax) were downregulated in these same cells
(Fig. 3F). The overexpression of
this lncRNA also inhibited the E-cadherin expression level, and
increased the protein levels of Slug, Vimentin and N-cadherin in
these PCa cells, which is consistent with the induction of
epithelial-mesenchymal transition (EMT) (Fig. 3G). These data underscore the
potential oncogenic role of PVT1 as a regulator of PCa cell
malignancy in vitro.
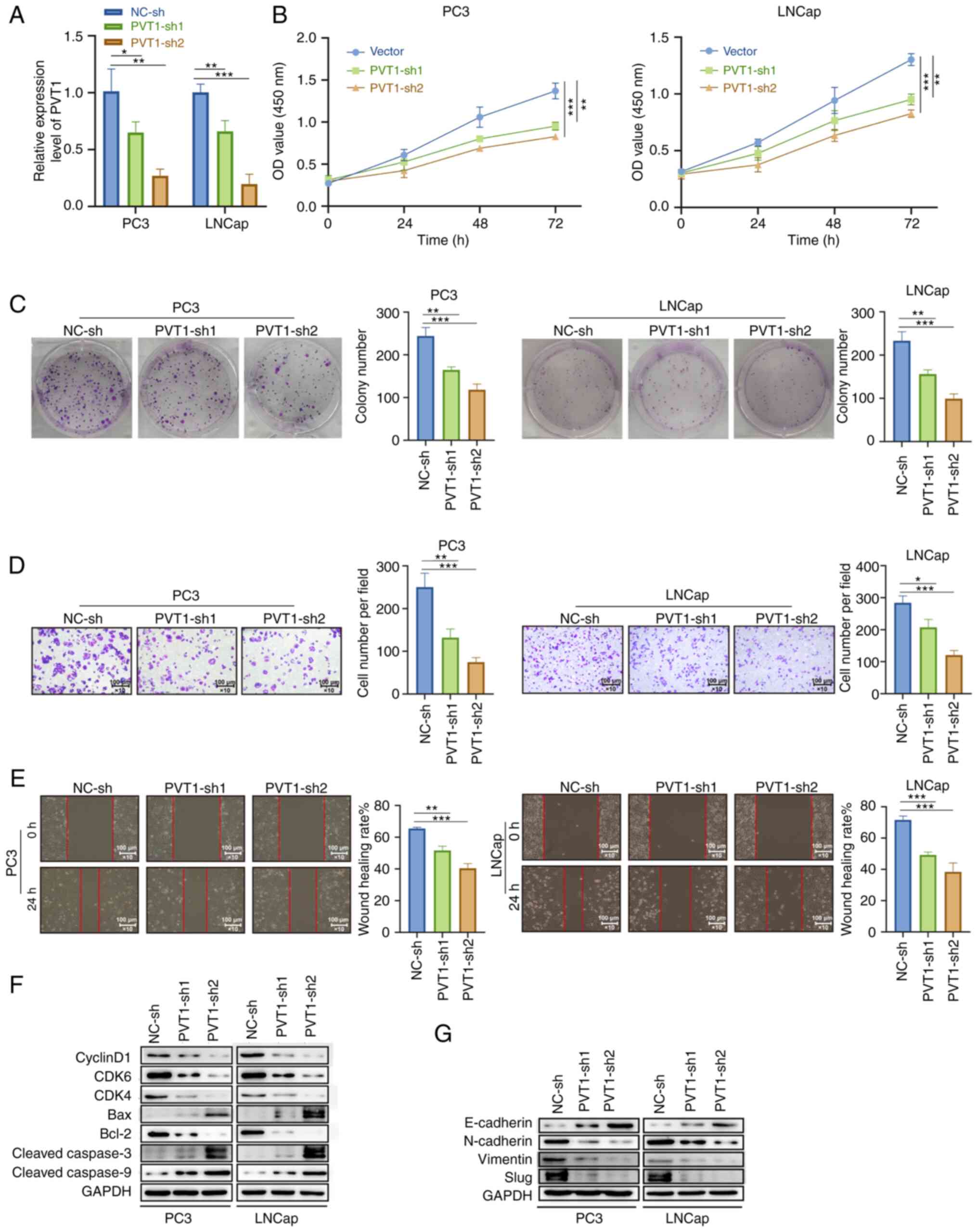
Knockdown of PVT1 inhibits in vitro PCa
cell proliferation, migration and invasion, whereas it induces cell
death
Subsequently, shRNA constructs specific for PVT1
(PVT1-sh1and PVT1-sh2) or corresponding control constructs (NC-sh)
were introduced into LNCap and PC3 cells, and PVT1 knockdown was
confirmed using RT-qPCR (Fig. 4A).
PVT1 knockdown suppressed the proliferation of these cells in both
colony formation and CCK-8 assays (Fig. 4B and C). Moreover, PVT1 knockdown
suppressed the invasive and migratory capabilities of the PCa cell
lines in Transwell and wound healing assays (Fig. 4D and E). Consistently, the
silencing of PVT1 also led to a marked decrease in the protein
levels of CDK4/6, cyclin D1 and Bcl-2 together with concomitant
increases in the levels of apoptosis-related proteins, such as
cleaved caspase-3/9 and Bax (Fig.
4F). The silencing of PVT1 further enhanced the E-cadherin
expression level, while it markedly suppressed the expression
levels of Slug, Vimentin and N-cadherin compared with the control
group (Fig. 4G). These results
indicated that the knockdown of PVT1 decreased PCa cell
malignancy.
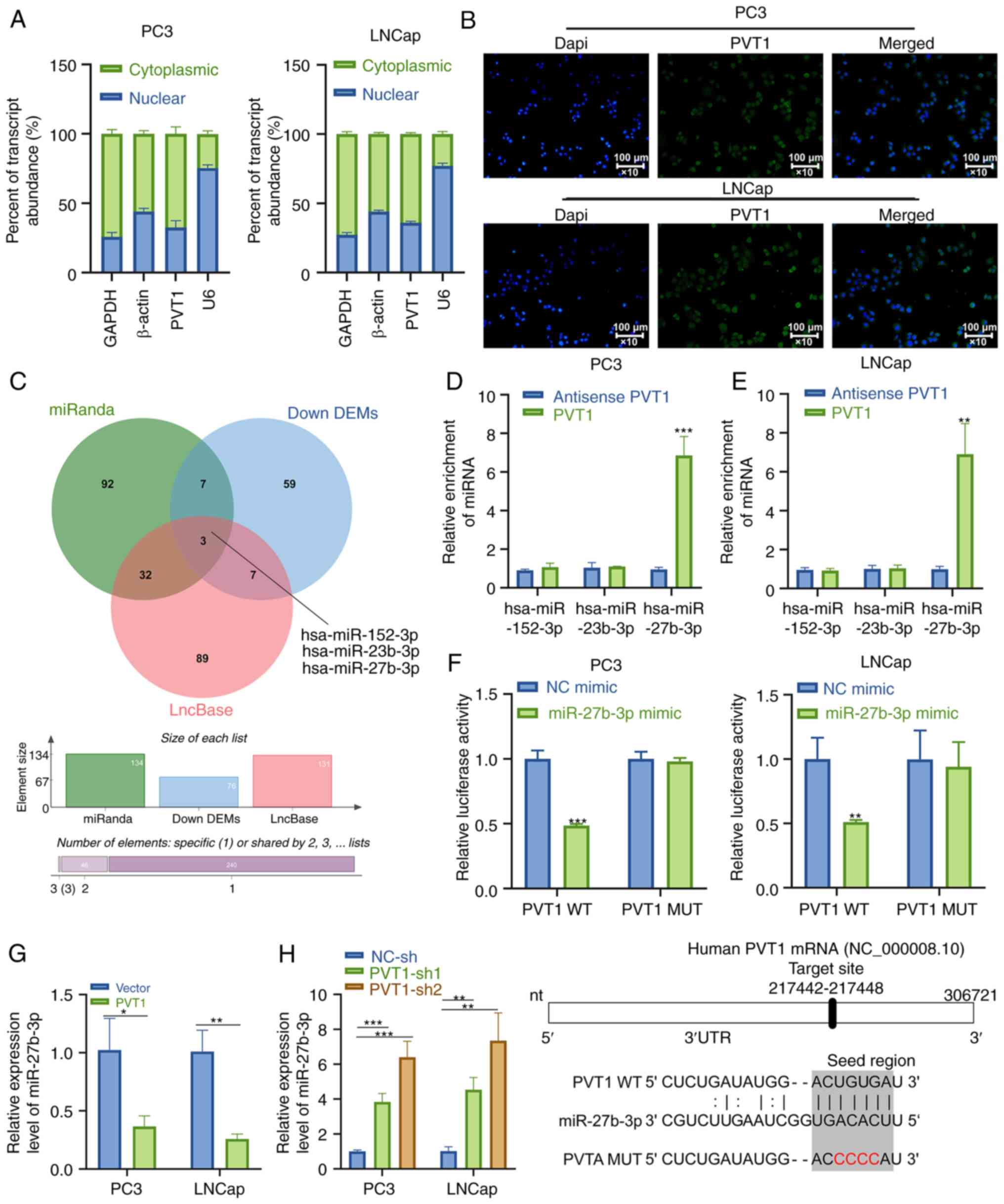
PVT1 functions as a ceRNA to sequester
miR-27b-3p within PCa cells
When functioning as ceRNAs, lncRNAs are able to
sequester miRNAs and thus indirectly promote the upregulation of
their target mRNAs. In the present study, PVT1 was found to be
primarily localized in the cytoplasm, rather than in the nucleus in
RNA-FISH and subcellular localization RT-qPCR assays (Fig. 5A and B), which is consistent with
the ability of PVT1 to primarily influence the characteristics of
PCa cells within the cytoplasm. Potential miRNA binding partners
for PVT1 were identified using miRanda and the LncBase Predicted
(ver. 2) online databases. Of the putative interacting miRNAs,
three were significantly downregulated in PCa samples in TCGA
database, and were thus considered as potential binding targets for
PVT1 in PCa (Fig. 5C). The present
study also observed the significant downregulation of miR-27b-3p,
miR-152-3p and miR-23b-3p expression in PCa tumors (Fig. S1A). According to the results of
Pearson's' correlation analysis, the associations between PVT1 and
each of these three miRNAs in TCGA PCa dataset were all consistent
with a potential ceRNA mechanism of action (Fig. S1B). Furthermore, RNA pull-down
assay verified that PVT1 could interact with miR-27b-3p in both PCa
cell lines tested (Fig. 5D and E).
A putative miR-27b-3p binding site was identified within PVT1, and
dual-luciferase reporter assay demonstrated that miR-27b-3p
overexpression was sufficient to suppress WT-PVT1 reporter activity
(Fig. 5F). In RT-qPCR analysis,
the miR-27b-3p expression level was significantly suppressed in
both LNCap and PC3 cells following PVT1 overexpression, whereas an
opposite effect was observed following PVT1 knockdown (Fig. 5G and H). Notably, miR-27b-3p did
not affect c-myc expression at both the mRNA and protein level
(Fig. S1C). The obtained outcomes
thus support the ability of PVT1 for a direct interaction, thereby
sequestering miR-27b-3p within PCa cells.
PVT1 controls the expression level of BLM
by functioning as a ceRNA for miR-27b-3p
Using the miRDB platform, four putative miR-27b-3p
target genes were identified, including ZNF124, SFXN4, EDRF1 and
BLM (Fig. 6A). Of these, BLM was
found to be inversely associated with the miR-27b-3p expression
level in PCa tissues (Fig. 6B).
When the BLM expression level was assessed in PCa samples from the
Oncomine database (33,38,42,45,46),
this gene was upregulated in PCa tumors (Fig. S1D). Western blot analysis and
RT-qPCR similarly confirmed the upregulation of BLM in PCa cell
lines (Fig. S2A). A higher BLM
expression level was associated with worse Gleason scores and worse
treatment outcomes (Fig. S1E).
Moreover, a higher BLM expression level was associated with more
undesirable disease-free survival outcomes in PCa cases (Fig. 6C). Following the overexpression of
miR-27b-3p (Fig. S2B), BLM mRNA
and protein levels in PCa cells were elevated (Fig. 6D). Moreover, following the
overexpression of BLM (Fig. S2C),
it was found that BLM promoted PVT1 expression (Fig. S2D). A putative binding site of
miR-27b-3p was identified within the BLM 3′-UTR, and the
dual-luciferase reporter assay demonstrated the ability of
miR-27b-3p to suppress BLM 3′-UTR-WT reporter plasmid activity in
PCa cells (Fig. 6E). RIP assay
revealed that miR-27b-3p co-precipitated with BLM, confirming the
binding association between these two RNAs (Fig. 6F). Moreover, the miR-27b-3p
expression level was significantly inhibited following PVT1
overexpression in LNCap and PC3 cells, whereas the same result was
not observed in the PVT1-MUT vector group. The BLM expression level
was markedly upregulated following PVT1 overexpression, rather than
in the PVT1-MUT vector group. These results thus support a role for
PVT1 as a regulator of BLM expression owing to its ability to
sequester miR-27b-3p.
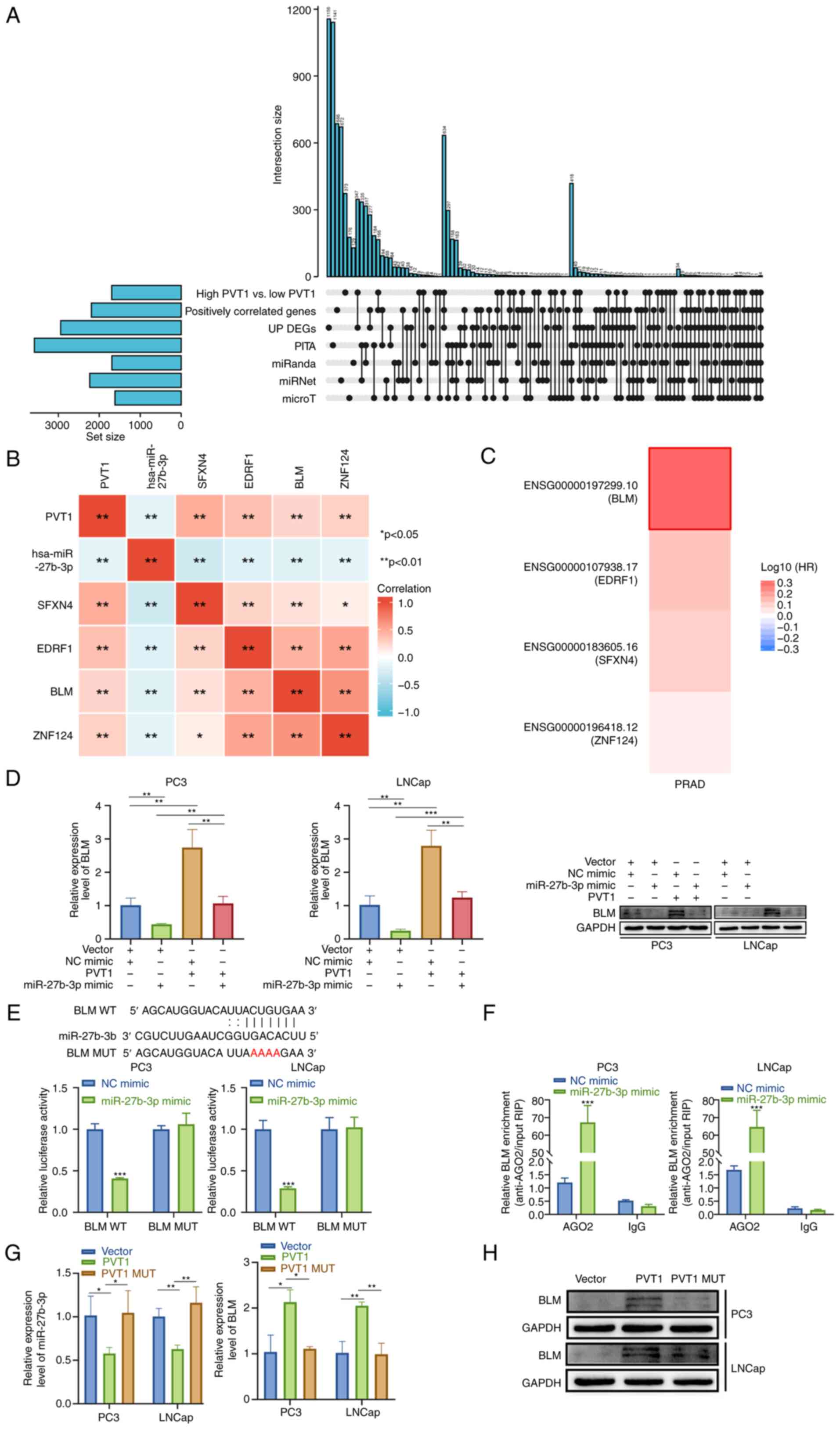 | Figure 6BLM is a miR-27b-3p target gene. (A)
The miRanda, miRwalk, miRDB, RNA22, and TargetScan databases were
employed to estimate possible miR-27b-3p target genes. (B)
Associations between the expression levels of PVT1 and miR-27b-3p,
and the identified target genes. (C) Analysis of disease-free
survival as a function of the expression levels of BLM, EDRF1,
SFXN4 and ZNF124. (D) reverse transcription-quantitative polymerase
chain reaction and western blot analysis were utilized to assess
the BLM expression level in LNCap and PC3 cells following PVT1
overexpression vector and miR-27b-3p mimic or NC mimic
co-transfection. (E) Luciferase activity in PCa cells was
quantified following miR-NC/miR-27b-3p mimic or BLM-WT/BLM-MT
co-transfection. (F) Interactions between miR-27b-3p and BLM were
assessed using RIP assay. (G) The expression levels of BLM and
miR-27b-3p in PCa cells following PVT1 overexpression vector or
PVT1-MUT treatment. (H) BLM expression level in PCa cells following
PVT1 overexpression vector or PVT1-MUT treatment as assessed using
western blot analysis. Data were from triplicate measurement and
are presented as the mean ± SD. *P<0.05,
**P<0.01 and ***P<0.001. BLM, bloom
syndrome protein; EDRF1, erythroid differentiation regulatory
factor 1; SFXN4, sideroflexin 4; ZNF124, zinc finger protein 124;
PVT1, plasmacytoma variant translocation 1; PCa, prostate
cancer. |
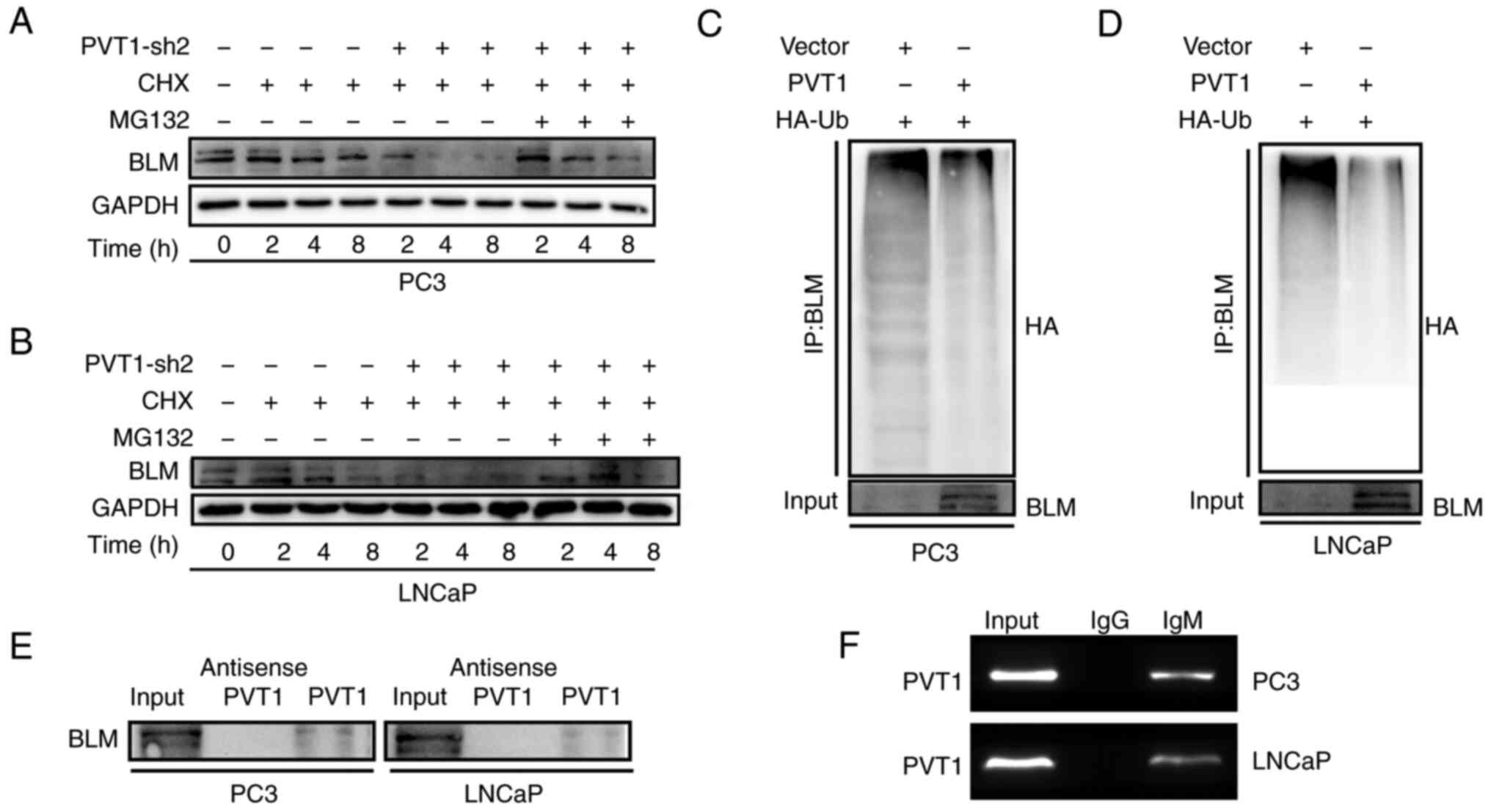
PVT1 directly stabilizes and interacts
with BLM
To determine whether PVT1 can alter the stability of
the BLM protein, CHX was used to treat the LNCap and PC3 cells, in
order to block translational activity, and BLM degradation was then
monitored over time. The BLM degradation rates significantly
decreased in the cells following PVT1 knockdown compared with the
control cells, while the BLM expression level in shPVT1-transfected
PCa cells increased, following MG132-mediated proteasome inhibition
(Fig. 7A and B). According to the
results of the immunoprecipitation assay, PVT1 also retained its
ubiquitination level (Fig. 7C and
D). RNA pull-down assay was additionally conducted, following
the incubation of LNCap and PC3 cell lysates with antisense PVT1 or
biotin-labeled PVT1, and revealed the ability of BLM and PVT1 to
directly interact within PCa cells (Fig. 7E). In addition, a RIP assay was
performed using anti-BLM, which similarly confirmed the ability of
BLM and PVT1 to interact together (Fig. 7F). These data thus provide evidence
for direct interactions between PVT1 and BLM, in which lncRNA may
partly function to reduce BLM ubiquitination, thereby improving its
stability.
PVT1 modulates the miR-27b-3p/BLM axis to
promote PCa progression
To examine the interaction between BLM and
miR-27b-3p in the context of the PVT1-mediated regulation of PCa
cell malignancy, a series of rescue experiments were performed.
Initially, RT-qPCR and western blot analysis were used to confirm
successful BLM upregulation in cells transfected with a BLM
overexpression vector (BLM-OE) (Fig.
8A). PC3 cells were then transfected with either BLM-OE or
empty vector control together with PVT1-sh2, and the malignant
activities of these cells were assessed. While PVT1 knockdown
reduced the colony formation and proliferative activity of these
cells (Fig. 8B and C), BLM
overexpression partially reversed these changes. Similarly, PVT1
knockdown impaired the PC3 migratory and invasive capabilities,
whereas BLM overexpression reversed these changes (Fig. 8D and E). Western blot analysis was
additionally performed to evaluate the expression levels of cell
cycle- and apoptosis-related proteins in these cells, and revealed
significant reductions in the expression levels of cyclin D1,
CDK4/6 and Bcl-2, following PVT1 knockdown that were reversed by
BLM overexpression (Fig. 8F). BLM
additionally reversed the PVT1 knockdown-mediated enhancement in
the E-cadherin expression level, and also reduced the expression
levels of Slug, Vimentin and N-cadherin (Fig. 8G). These data thus indicated that
PVT1 promoted PCa progression in part by inducing the
downregulation of BLM expression.
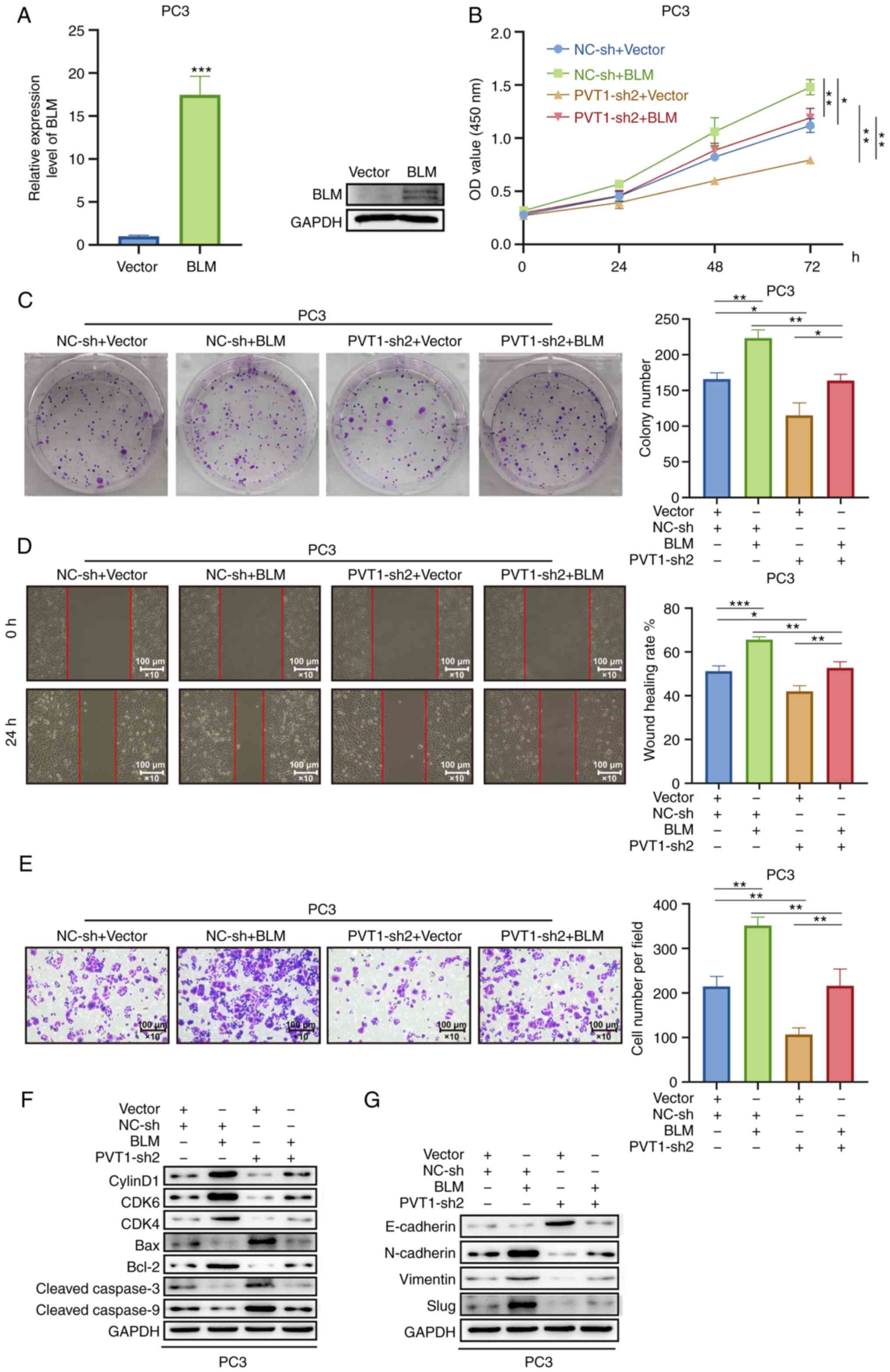 | Figure 8BLM overexpression reverses the
effects of PVT1 upregulation on tje proliferative, migratory and
invasive abilities of PCa cells. (A) BLM upregulation was detected
using reverse transcription-quantitative polymerase chain reaction
and western blot analysis in PC3 cells following BLM vector
introduction. (B-G) PVT1-sh2 and BLM overexpression or control
vectors were co-transfected into PC3 cells, of which the
proliferative, colony formation, migratory and invasive abilities
of PCa cells were assessed using (B) CCK-8, (C) colony formation,
(D) wound healing, and (E) Transwell assays, respectively. (F) Cell
cycle- and apoptosis-related protein expression levels in PC3 cells
were measured. (G) The expression levels of E-cadherin, N-cadherin,
vimentin and Slug in PC3 cells were measured using western blot
analysis. *P<0.05, **P<0.01 and
***P<0.001. BLM, bloom syndrome protein; PVT1,
plasmacytoma variant translocation 1; PCa, prostate cancer. |
PVT1 promotes PCa progression in vivo via
the miR-27b-3p/BLM pathway
Finally, the performance of PVT1 as a regulator of
PCa progression in vivo was assessed by subcutaneously
implanting nude mice with PC3 cells that had been transfected with
empty vector (Vector) or BLM-OE together with PVT1-sh2. Tumor
volume and weight were then monitored over time, which revealed
significantly a slower tumor growth and lower weight values
following PVT1 downregulation, while this outcome was reversed by
BLM overexpression (Fig. 9A-C).
The expression levels of BLM and PVT1 in tumor tissues from these
mice were detected using RT-qPCR, and it was found that the BLM
expression level was downregulated following PVT1 knockdown,
whereas BLM overexpression reversed this change in the tumor cells
(Fig. 9D). The immunohistochemical
staining of these tumors yielded consistent results, and
additionally indicated that there were fewer PCNA-positive tumor
cells in tumors, in which PVT1 was knocked down, while this outcome
was reversed by the overexpression of BLM (Fig. 9E). PVT1 knockdown additionally
increased the cleaved caspase-9 expression level and reduced the
N-cadherin expression level within these tumors, while BLM
overexpression exerted the opposite effect (Fig. 9E). These findings indicated that
PVT1 promoted PCa progression via the miR-27b-3p/BLM axis.
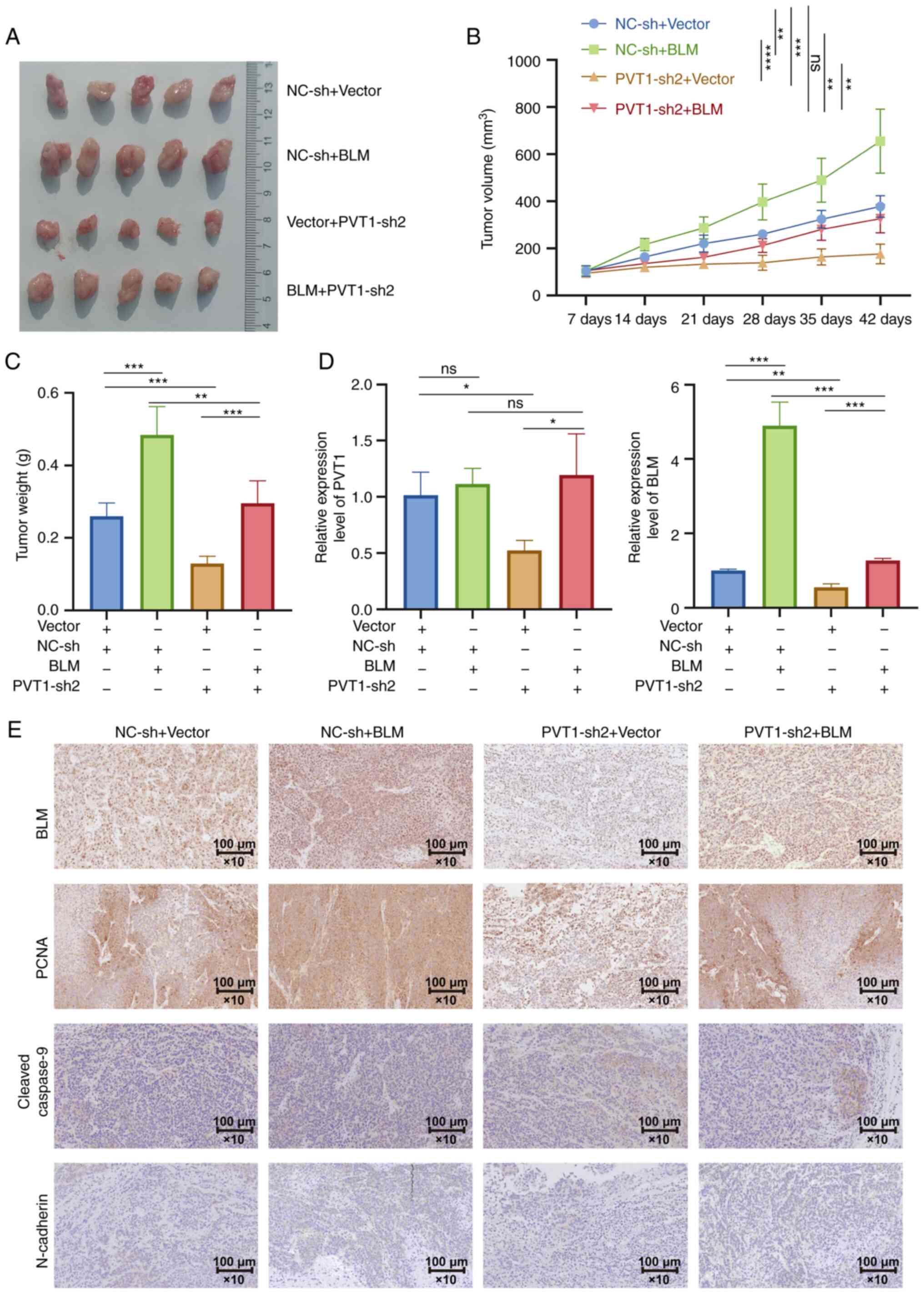 | Figure 9Knockdown of PVT1 suppresses PCa
tumor growth in vivo through mechanisms dependent on the
miR-27b-3p/BLM axis. (A) Tumors observed at 6 weeks
post-implantation in BALB/c nude mice in the indicated groups at
the experimental endpoint. (B) At 7 days following implantation,
tumor volume was monitored once per week. (C) Tumors were weighed
at the end of the experiment. (D) The expression levels of PVT1 and
BLM were monitored using reverse transcription-quantitative
polymerase chain reaction in isolated tumors. (E)
Immunohistochemical staining was employed to detect the expression
levels of N-cadherin, cleaved caspase-9, PCNA and BLM in xenograft
tumors (magnification, ×200; scale bars, 100 µm).
*P<0.05, **P<0.01 and
***P<0.001. ns, not significant; PVT1, plasmacytoma
variant translocation 1; PCa, prostate cancer; BLM, bloom syndrome
protein; PCNA, proliferating cell nuclear antigen. |
Discussion
As an important prognostic factor in the context of
tumor development and patient survival for various types of cancer,
PVT1 can regulate a range of biological activities, including
motility, invasion, proliferation and cell death (47,48).
In patients with PCa, the prognostic value of PVT1 has previously
been demonstrated (49-51), which is in line with the results of
the present study, indicating the upregulation of PVT1 in PCa tumor
samples and the association between the expression level of this
lncRNA, clinical staging and patient survival outcomes. Other
research groups have similarly reported that PVT1 upregulation is
predictive of a decreased patient overall survival, and it is
positively associated with lymph node metastasis, distant
metastasis, TNM staging and tumor size (24,51,52).
Overall, the obtained findings demonstrated that PVT1 may be a
prognostic biomarker for PCa, although further research regarding
the mechanistic basis for its dysregulation is warranted.
The m6A modification can regulate the
expression levels of several tumor-associated genes (13,53),
and the dysregulation of this process is strongly associated with
tumor progression (54). A complex
of methyltransferase comprising WTAP, METTL3 and other proteins
catalyzes the m6A modification of lncRNAs and other
targets within the nucleus (55).
Different patterns of m6A distributions have been
observed for mRNAs and lncRNAs, suggesting that distinct mechanisms
may govern these two modification processes, although additional
analysis are required to fully clarify this hypothesis (56,57).
The present study highlighted a novel PVT1-related regulatory
pathway in PCa cells. Through RIP and RNA pull-down experiments, it
was found that PVT1 was subjected to METTL3-mediated m6A
modification, with METTL3 ultimately suppressed PVT1 degradation,
resulting in a net increase in the expression level of this lncRNA.
Consequently, it was revealed that METTL3 knockdown suppressed PVT1
m6A modification, leading to the impairment of its
reader binding protein and a consequent increase in PVT1 turnover
owing to its co-localization with decay factors (58). Additional research, however, is
required to verify this hypothetical model.
Recent research has presented evidence for the
capability of lncRNAs to sequester specific miRNAs in a sponge-like
manner, thereby functioning as ceRNAs and altering the
functionality of these miRNAs (59). PVT1, for instance, has been
previously reported to bind to tumor suppressor miRNAs, thereby
upregulating the nucleolar protein 2 (NOP2) expression level, as
well as promoting PCa metastasis (60). Furthermore, PVT1 has been
demonstrated to enhance PCa cell growth in a miR-146a
methylation-dependent manner (61), in addition to promoting PCa
progression and EMT induction via the suppression of the
miR-187-mediated inhibition of Twist1 expression (62). Furthermore, PVT1 acts as a sponge
for miRNA-186-5p and positively regulates Twist1 to promote
prostate cancer progression (62).
PVT1 can regulate ferroptosis through miR-214-mediated TFR1 and p53
(63). PVT1 can promote
gallbladder cancer progression by regulating the miR-143/HK2 axis
(64). To clarify the mechanisms
whereby PCa influences PCa cell malignancy, the present study
conducted bioinformatics analyses, leading to the identification of
miR-27b-3p as an additional PVT1 target miRNA that has previously
been illustrated to play a tumor suppressive role in lung cancer
and esophageal squamous cell carcinoma (65,66).
Although the interaction of PVT1 and miR-27b-3p has been previously
reported (67,68), this finding was only demonstrated
in non-tumor cells. In the present study, RT-qPCR divulged a
negative regulatory association between PVT1 and miR-27b-3p, and
dual-luciferase reporter, pull-down and RIP assays additionally
confirmed the direct targeting association between these two RNAs
in PCa cells.
BLM controls essential intracellular processes,
including telomerase maintenance, DNA unwinding and DNA repair, in
addition to serving as a promising biomarker for PCa (69,70).
BLM has been reported to target the MYC proto-oncogene gene that
encodes c-MYC, thereby enabling it to function as a tumor
suppressor gene (71). In a
previous study, the authors found that miR-27b-3p could target BLM
(72). However, the finding was
only limited to cells and the binding of miR-27b-3p and BLM was
only confirmed by dual-luciferase reporter assay. In the present
study, by integrating prediction results from databases, the
expression level-based correlation of PVT1, miR-27b-3p and BLM in
PCa tissues from TCGA database, and the BLM expression level from
TCGA and Oncomine databases, BLM was detected as a miR-27b-3p
target. Furthermore, this association was confirmed using the
dual-luciferase reporter and AGO2 pull-down assays. Whether this
regulatory association is linked to the c-myc expression level,
however, warrants further investigation.
In conclusion, the findings of the present study
demonstrate that the METTL3/PVT1/miR-27b-3p/BLM axis may act as a
positive regulator of PCa progression. However, only the
METTL3methyltransferase was detected to mediate PVT1 m6A
modification, and additional research regarding this regulatory
epigenetic relationship is thus warranted.
Supplementary Data
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
BC and HX conceived the study and designed the
experiments. CL and HL contributed to data collection and confirm
the authenticity of all the raw data. GB and YZ performed the data
analysis and interpreted the results. BC wrote the manuscript. BC
and HX contributed to the critical revision of article. All authors
have read and approved the final manuscript.
Ethics approval and consent to
participate
The study protocol was approved by the Ethics
Committee of Affiliated Hospital of Zunyi Medical University
[Compliance Medical Ethics Review (2021) no. 1-037). Written
informed consent was obtained from all participants prior to
enrollment. The animal study protocol was approved by the Ethics
Committee of Affiliated Hospital of Zunyi Medical University
(ZMU21-2205-002).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgments
Not applicable.
Funding
The present study was financially supported by the National
Natural Science Foundation of China (grant no. 31860242).
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2020. CA Cancer J Clin. 70:7–30. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kimura T and Egawa S: Epidemiology of
prostate cancer in Asian countries. Int J Urol. 25:524–531. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Teo MY, Rathkopf DE and Kantoff P:
Treatment of advanced prostate cancer. Annu Rev Med. 70:479–499.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rebbeck TR: Prostate cancer genetics:
Variation by race, ethnicity, and geography. Semin Radiat Oncol.
27:3–10. 2017. View Article : Google Scholar
|
|
5
|
Kaiser A, Haskins C, Siddiqui MM, Hussain
A and D'Adamo C: The evolving role of diet in prostate cancer risk
and progression. Curr Opin Oncol. 31:222–229. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Moammeri A, Abbaspour K, Zafarian A,
Jamshidifar E, Motasadizadeh H, Dabbagh Moghaddam F, Salehi Z,
Makvandi P and Dinarvand R: pH-Responsive, adorned nanoniosomes for
codelivery of cisplatin and epirubicin: Synergistic treatment of
breast cancer. ACS Appl Bio Mater. 5:675–690. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ramezani Farani M, Azarian M, Heydari
Sheikh Hossein H, Abdolvahabi Z, Mohammadi Abgarmi Z, Moradi A,
Mousavi SM, Ashrafizadeh M, Makvandi P, Saeb MR and Rabiee N: Folic
acid-adorned curcumin-loaded iron oxide nanoparticles for cervical
cancer. ACS Appl Bio Mater. 5:1305–1318. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sharifi E, Bigham A, Yousefiasl S, Trovato
M, Ghomi M, Esmaeili Y, Samadi P, Zarrabi A, Ashrafizadeh M,
Sharifi S, et al: Mesoporous bioactive glasses in cancer diagnosis
and therapy: Stimuli-responsive, toxicity, immunogenicity, and
clinical translation. Adv Sci (Weinh). 9:e21026782022. View Article : Google Scholar
|
|
9
|
Snow O, Lallous N, Singh K, Lack N, Rennie
P and Cherkasov A: Androgen receptor plasticity and its
implications for prostate cancer therapy. Cancer Treat Rev.
81:1018712019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yassin A, AlRumaihi K, Alzubaidi R,
Alkadhi S and Al Ansari A: Testosterone, testosterone therapy and
prostate cancer. Aging Male. 22:219–227. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang H, Shi X, Huang T, Zhao X, Chen W,
Gu N and Zhang R: Dynamic landscape and evolution of m6A
methylation in human. Nucleic Acids Res. 48:6251–6264. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yang Y, Hsu PJ, Chen YS and Yang YG:
Dynamic transcriptomic m6A decoration: Writers, erasers,
readers and functions in RNA metabolism. Cell Res. 28:616–624.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Sun T, Wu R and Ming L: The role of m6A
RNA methylation in cancer. Biomed Pharmacother. 112:1086132019.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang T, Kong S, Tao M and Ju S: The
potential role of RNA N6-methyladenosine in Cancer progression. Mol
Cancer. 19:882020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Boon RA, Jae N, Holdt L and Dimmeler S:
Long Noncoding RNAs: From clinical genetics to therapeutic targets?
J Am Coll Cardiol. 67:1214–1226. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Jiang R, Tang J, Chen Y, Deng L, Ji J, Xie
Y, Wang K, Jia W, Chu WM and Sun B: The long noncoding RNA lnc-EGFR
stimulates T-regulatory cells differentiation thus promoting
hepatocellular carcinoma immune evasion. Nat Commun. 8:151292017.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li J, Tian H, Yang J and Gong Z: Long
Noncoding RNAs regulate cell growth, proliferation, and apoptosis.
DNA Cell Biol. 35:459–470. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Paraskevopoulou MD and Hatzigeorgiou AG:
Analyzing MiRNA-LncRNA interactions. Methods Mol Biol.
1402:271–286. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ashrafizadeh M, Paskeh MDA, Mirzaei S,
Gholami MH, Zarrabi A, Hashemi F, Hushmandi K, Hashemi M, Nabavi N,
Crea F, et al: Targeting autophagy in prostate cancer: Preclinical
and clinical evidence for therapeutic response. J Exp Clin Cancer
Res. 41:1052022. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ashrafizaveh S, Ashrafizadeh M, Zarrabi A,
Husmandi K, Zabolian A, Shahinozzaman M, Aref AR, Hamblin MR,
Nabavi N, Crea F, et al: Long non-coding RNAs in the doxorubicin
resistance of cancer cells. Cancer Lett. 508:104–114. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Mirzaei S, Zarrabi A, Hashemi F, Zabolian
A, Saleki H, Ranjbar A, Seyed Saleh SH, Bagherian M, Sharifzadeh
SO, Hushmandi K, et al: Regulation of Nuclear Factor-KappaB (NF-κB)
signaling pathway by non-coding RNAs in cancer: Inhibiting or
promoting carcinogenesis? Cancer Lett. 509:63–80. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang Y, Tan Y, Wang H, Xu M and Xu L:
Long Non-Coding RNA plasmacytoma variant translocation 1 (PVT1)
enhances proliferation, migration, and epithelial-mesenchymal
transition (EMT) of pituitary adenoma cells by activating
β-catenin, c-Myc, and cyclin D1 expression. Med Sci Monit.
25:7652–7659. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wu H, Tian X and Zhu C: Knockdown of
lncRNA PVT1 inhibits prostate cancer progression in vitro and in
vivo by the suppression of KIF23 through stimulating miR-15a-5p.
Cancer Cell Int. 20:2832020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu J, Li Y, Zhang Q, Lv C, Wang M, Jiao Y
and Wang C: PVT1 expression is a predictor for poor survival of
prostate cancer patients. Technol Cancer Res Treat.
20:15330338209716102021. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhao J, Du P, Cui P, Qin Y, Hu C, Wu J,
Zhou Z, Zhang W, Qin L and Huang G: LncRNA PVT1 promotes
angiogenesis via activating the STAT3/VEGFA axis in gastric cancer.
Oncogene. 37:4094–4109. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chen S, Zhou L and Wang Y: ALKBH5-mediated
m6A demethylation of lncRNA PVT1 plays an oncogenic role
in osteosarcoma. Cancer Cell Int. 20:342020. View Article : Google Scholar
|
|
27
|
Qi X, Zhang DH, Wu N, Xiao JH, Wang X and
Ma W: ceRNA in cancer: Possible functions and clinical
implications. J Med Genet. 52:710–718. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Liu K and Xu Q: LncRNA PVT1 regulates
gallbladder cancer progression through miR-30d-5p. J Biol Regul
Homeost Agents. 34:875–883. 2020.PubMed/NCBI
|
|
29
|
Zhou C, Yi C, Yi Y, Qin W, Yan Y, Dong X,
Zhang X, Huang Y, Zhang R, Wei J, et al: LncRNA PVT1 promotes
gemcitabine resistance of pancreatic cancer via activating
Wnt/beta-catenin and autophagy pathway through modulating the
miR-619-5p/Pygo2 and miR-619-5p/ATG14 axes. Mol Cancer. 19:1182020.
View Article : Google Scholar
|
|
30
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Song YX, Sun JX, Zhao JH, Yang YC, Shi JX,
Wu ZH, Chen XW, Gao P, Miao ZF and Wang ZN: Non-coding RNAs
participate in the regulatory network of CLDN4 via ceRNA mediated
miRNA evasion. Nat Commun. 8:2892017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hu Y, Rao SS, Wang ZX, Cao J, Tan YJ, Luo
J, Li HM, Zhang WS, Chen CY and Xie H: Exosomes from human
umbilical cord blood accelerate cutaneous wound healing through
miR-21-3p-mediated promotion of angiogenesis and fibroblast
function. Theranostics. 8:169–184. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Arredouani MS, Lu B, Bhasin M, Eljanne M,
Yue W, Mosquera JM, Bubley GJ, Li V, Rubin MA, Libermann TA and
Sanda MG: Identification of the transcription factor single-minded
homologue 2 as a potential biomarker and immunotherapy target in
prostate cancer. Clin Cancer Res. 15:5794–5802. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Lapointe J, Li C, Higgins JP, van de Rijn
M, Bair E, Montgomery K, Ferrari M, Egevad L, Rayford W, Bergerheim
U, et al: Gene expression profiling identifies clinically relevant
subtypes of prostate cancer. Proc Natl Acad Sci USA. 101:811–816.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
LaTulippe E, Satagopan J, Smith A, Scher
H, Scardino P, Reuter V and Gerald WL: Comprehensive gene
expression analysis of prostate cancer reveals distinct
transcriptional programs associated with metastatic disease. Cancer
Res. 62:4499–4506. 2002.PubMed/NCBI
|
|
36
|
Luo JH, Yu YP, Cieply K, Lin F, Deflavia
P, Dhir R, Finkelstein S, Michalopoulos G and Becich M: Gene
expression analysis of prostate cancers. Mol Carcinog. 33:25–35.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Singh D, Febbo PG, Ross K, Jackson DG,
Manola J, Ladd C, Tamayo P, Renshaw AA, D'Amico AV, Richie JP, et
al: Gene expression correlates of clinical prostate cancer
behavior. Cancer Cell. 1:203–209. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Taylor BS, Schultz N, Hieronymus H,
Gopalan A, Xiao Y, Carver BS, Arora VK, Kaushik P, Cerami E, Reva
B, et al: Integrative genomic profiling of human prostate cancer.
Cancer Cell. 18:11–22. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Tomlins SA, Mehra R, Rhodes DR, Cao X,
Wang L, Dhanasekaran SM, Kalyana-Sundaram S, Wei JT, Rubin MA,
Pienta KJ, et al: Integrative molecular concept modeling of
prostate cancer progression. Nat Genet. 39:41–51. 2007. View Article : Google Scholar
|
|
40
|
Varambally S, Yu J, Laxman B, Rhodes DR,
Mehra R, Tomlins SA, Shah RB, Chandran U, Monzon FA, Becich MJ, et
al: Integrative genomic and proteomic analysis of prostate cancer
reveals signatures of metastatic progression. Cancer Cell.
8:393–406. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Welsh JB, Sapinoso LM, Su Al, Kern SG,
Wang-Rodriguez J, Moskaluk CA, Frierson HF Jr and Hampton GM:
Analysis of gene expression identifies candidate markers and
pharmacological targets in prostate cancer. Cancer Res.
61:5974–5978. 2001.PubMed/NCBI
|
|
42
|
Yu YP, Landsittel D, Jing L, Nelson J, Ren
B, Liu L, McDonald C, Thomas R, Dhir R, Finkelstein S, et al: Gene
expression alterations in prostate cancer predicting tumor
aggression and preceding development of malignancy. J Clin Oncol.
22:2790–2799. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wang R and Liu X: Epigenetic regulation of
prostate cancer. Genes Dis. 7:606–613. 2019. View Article : Google Scholar
|
|
44
|
Li J, Xie H, Ying Y, Chen H, Yan H, He L,
Xu M, Xu X, Liang Z, Liu B, et al: YTHDF2 mediates the mRNA
degradation of the tumor suppressors to induce AKT phosphorylation
in N6-methyladenosine-dependent way in prostate cancer. Mol Cancer.
19:1522020. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Grasso CS, Wu YM, Robinson DR, Cao X,
Dhanasekaran SM, Khan AP, Quist MJ, Jing X, Lonigro RJ, Brenner JC,
et al: The mutational landscape of lethal castration-resistant
prostate cancer. Nature. 487:239–243. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Liu P, Ramachandran S, Ali Seyed M,
Scharer CD, Laycock N, Dalton WB, Williams H, Karanam S, Datta MW,
Jaye DL and Moreno CS: Sex-determining region Y box 4 is a
transforming oncogene in human prostate cancer cells. Cancer Res.
66:4011–4019. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Ding C, Yang Z, Lv Z, DU C, Xiao H, Peng
C, Cheng S, Xie H, Zhou L, Wu J and Zheng S: Long non-coding RNA
PVT1 is associated with tumor progression and predicts recurrence
in hepatocellular carcinoma patients. Oncol Lett. 9:955–963. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Li KK, Lau KM and Ng HK: Signaling pathway
and molecular subgroups of medulloblastoma. Int J Clin Exp Pathol.
6:1211–1222. 2013.PubMed/NCBI
|
|
49
|
Lin HY, Callan CY, Fang Z, Tung HY and
Park JY: Interactions of PVT1 and CASC11 on prostate cancer risk in
african americans. Cancer Epidemiol Biomarkers Prev. 28:1067–1075.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Videira A, Beckedorff FC, daSilva LF and
Verjovski-Almeida S: PVT1 signals an androgen-dependent
transcriptional repression program in prostate cancer cells and a
set of the repressed genes predicts high-risk tumors. Cell Commun
Signal. 19:52021. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Yang J, Li C, Mudd A and Gu X: LncRNA PVT1
predicts prognosis and regulates tumor growth in prostate cancer.
Biosci Biotechnol Biochem. 81:2301–2306. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Yan YJ, Zhang L, Zhou JJ, Chen ZJ, Liao
YX, Zeng JM and Shen H: Comprehensive characterization of common
and cancer-specific differently expressed lncRNAs in urologic
cancers. Comput Math Methods Med. 2021:55152182021. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Lyko F and Brown R: DNA methyltransferase
inhibitors and the development of epigenetic cancer therapies. J
Natl Cancer Inst. 97:1498–1506. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Tuncel G and Kalkan R: Importance of m
N6-methyladenosine (m6A) RNA modification in
cancer. Med Oncol. 36:362019. View Article : Google Scholar
|
|
55
|
Sun HL, Zhu AC, Gao Y, Terajima H, Fei Q,
Liu S, Zhang L, Zhang Z, Harada BT, He YY, et al: Stabilization of
ERK-Phosphorylated METTL3 by USP5 Increases m6A
Methylation. Mol Cell. 80:633–647.e7. 2020. View Article : Google Scholar
|
|
56
|
Liu N, Parisien M, Dai Q, Zheng G, He C
and Pan T: Probing N6-methyladenosine RNA modification status at
single nucleotide resolution in mRNA and long noncoding RNA. RNA.
19:1848–1856. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Amort T, Rieder D, Wille A,
Khokhlova-Cubberley D, Riml C, Trixl L, Jia XY, Micura R and Lusser
A: Distinct 5-methylcytosine profiles in poly(A) RNA from mouse
embryonic stem cells and brain. Genome Biol. 18:12017. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han
D, Fu Y, Parisien M, Dai Q, Jia G, et al:
N6-methyladenosine-dependent regulation of messenger RNA stability.
Nature. 505:117–120. 2014. View Article : Google Scholar
|
|
59
|
Chan JJ and Tay Y: Noncoding RNA:RNA
regulatory networks in cancer. Int J Mol Sci. 19:13102018.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Sun F, Wu K, Yao Z, Mu X, Zheng Z, Sun M,
Wang Y, Liu Z and Zhu Y: Long Noncoding RNA PVT1 promotes prostate
cancer metastasis by increasing NOP2 expression via targeting tumor
suppressor MicroRNAs. Onco Targets Ther. 13:6755–6765. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Liu HT, Fang L, Cheng YX and Sun Q: LncRNA
PVT1 regulates prostate cancer cell growth by inducing the
methylation of miR-146a. Cancer Med. 5:3512–3519. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Chang Z, Cui J and Song Y: Long noncoding
RNA PVT1 promotes EMT via mediating microRNA-186 targeting of
Twist1 in prostate cancer. Gene. 654:36–42. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Lu J, Xu F and Lu H: LncRNA PVT1 regulates
ferroptosis through miR-214-mediated TFR1 and p53. Life Sci.
260:1183052020. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Chen J, Yu Y, Li H, Hu Q, Chen X, He Y,
Xue C, Ren F, Ren Z, Li J, et al: Long non-coding RNA PVT1 promotes
tumor progression by regulating the miR-143/HK2 axis in gallbladder
cancer. Mol Cancer. 18:332019. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Han M, Li N, Li F, Wang H and Ma L:
MiR-27b-3p exerts tumor suppressor effects in esophageal squamous
cell carcinoma by targeting Nrf2. Hum Cell. 33:641–651. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Sun Y, Xu T, Cao YW and Ding XQ: Antitumor
effect of miR-27b-3p on lung cancer cells via targeting Fzd7. Eur
Rev Med Pharmacol Sci. 21:4113–4123. 2017.PubMed/NCBI
|
|
67
|
Li S, Zhao X, Cheng S, Li J, Bai X and
Meng X: Downregulating long non-coding RNA PVT1 expression
inhibited the viability, migration and phenotypic switch of
PDGF-BB-treated human aortic smooth muscle cells via targeting
miR-27b-3p. Hum Cell. 34:335–348. 2021. View Article : Google Scholar
|
|
68
|
Lu X, Yu Y, Yin F, Yang C, Li B, Lin J and
Yu H: Knockdown of PVT1 inhibits IL-1β-induced injury in
chondrocytes by regulating miR-27b-3p/TRAF3 axis. Int
Immunopharmacol. 79:1060522020. View Article : Google Scholar
|
|
69
|
Ledet EM, Antonarakis ES, Isaacs WB, Lotan
TL, Pritchard C and Sartor AO: Germline BLM mutations and
metastatic prostate cancer. Prostate. 80:235–237. 2020. View Article : Google Scholar :
|
|
70
|
Ruan Y, Xu H, Ji X and Zhao J: BLM
interaction with EZH2 regulates MDM2 expression and is a poor
prognostic biomarker for prostate cancer. Am J Cancer Res.
11:1347–1368. 2021.PubMed/NCBI
|
|
71
|
Deng L, Meng T, Chen L, Wei W and Wang P:
The role of ubiquitination in tumorigenesis and targeted drug
discovery. Signal Transduct Target Ther. 5:112020. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Chen Y, Zhao J, Duan Z, Gong T, Chen W,
Wang S and Xu H: miR27b3p and miR607 cooperatively regulate BLM
gene expression by directly targeting the 3′UTR in PC3 cells. Mol
Med Rep. 19:4819–4831. 2019.PubMed/NCBI
|