|
1
|
de Jong MME, Kellermayer Z, Papazian N,
Tahri S, Hofste Op Bruinink D, Hoogenboezem R, Sanders MA, van de
Woestijne PC, Bos PK, Khandanpour C, et al: The multiple myeloma
microenvironment is defined by an inflammatory stromal cell
landscape. Nat Immunol. 22:769–780. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ramberger E, Sapozhnikova V, Ng YLD,
Dolnik A, Ziehm M, Popp O, Sträng E, Kull M, Grünschläger F, Krüger
J, et al: The proteogenomic landscape of multiple myeloma reveals
insights into disease biology and therapeutic opportunities. Nat
Cancer. 5:1267–1284. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Guan L, Su W, Zhong J and Qiu L: M-protein
detection by mass spectrometry for minimal residual disease in
multiple myeloma. Clin Chim Acta. 552:1176232024. View Article : Google Scholar
|
|
4
|
Kumar SK, Rajkumar V, Kyle RA, van Duin M,
Sonneveld P, Mateos MV, Gay F and Anderson KC: Multiple myeloma.
Nat Rev Dis Prim. 3:170462017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Mikhael J, Bhutani M and Cole CE: Multiple
myeloma for the primary care provider: A practical review to
promote earlier diagnosis among diverse populations. Am J Med.
136:33–41. 2023. View Article : Google Scholar
|
|
6
|
Rajkumar SV: Multiple myeloma: 2022 update
on diagnosis, risk stratification, and management. Am J Hematol.
97:10862022. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Huang J, Chan SC, Lok V, Zhang L,
Lucero-Prisno DE III, Xu W, Zheng ZJ, Elcarte E, Withers M, Wong
MCS, et al: The epidemiological landscape of multiple myeloma: A
global cancer registry estimate of disease burden, risk factors,
and temporal trends. Lancet Haematol. 9:e670–e677. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Das S, Juliana N, Yazit NAA, Azmani S and
Abu IF: Multiple myeloma: Challenges encountered and future options
for better treatment. Int J Mol Sci. 23:16492022. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Mohty M, Facon T, Malard F and Harousseau
JL: A roadmap towards improving outcomes in multiple myeloma. Blood
Cancer J. 14:1352024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hu S, Xu J, Cui W, Jin H, Wang X and
Maimaitiyiming Y: Post-translational modifications in multiple
myeloma: Mechanisms of drug resistance and therapeutic
opportunities. Biomolecules. 15:7022025. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Rajkumar SV and Kumar S: Multiple myeloma
current treatment algorithms. Blood Cancer J. 10:942020. View Article : Google Scholar :
|
|
12
|
Bhatt P, Kloock C and Comenzo R:
Relapsed/refractory multiple myeloma: A review of available
therapies and clinical scenarios encountered in myeloma relapse.
Curr Oncol. 30:2322–2347. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zeng L, Huang H, Qirong C, Ruan C, Liu Y,
An W, Guo Q and Zhou J: Multiple myeloma patients undergoing
chemotherapy: Which symptom clusters impact quality of life? J Clin
Nurs. 32:7247–7259. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Goodman RS, Johnson DB and Balko JM:
Corticosteroids and cancer immunotherapy. Clin Cancer Res.
29:2580–2587. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Luo H, Feng Y, Wang F, Lin Z, Huang J, Li
Q, Wang X, Liu X, Zhai X, Gao Q, et al: Combinations of ivermectin
with proteasome inhibitors induce synergistic lethality in multiple
myeloma. Cancer Lett. 565:2162182023. View Article : Google Scholar : PubMed/NCBI
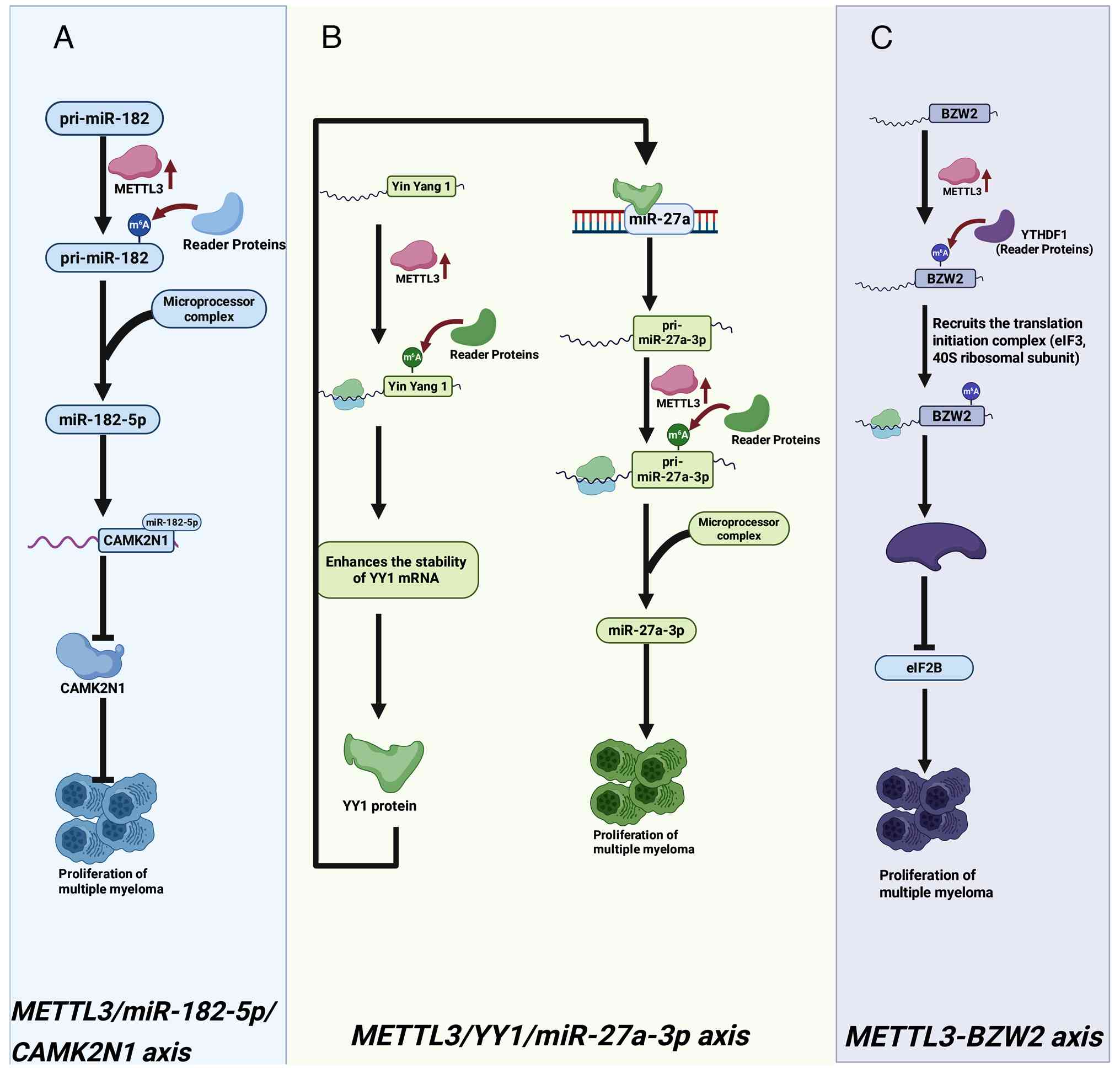
|
|
16
|
Minařík J and Ševčíková S:
Immunomodulatory agents for multiple myeloma. Cancers (Basel).
14:57592022. View Article : Google Scholar
|
|
17
|
Koniarczyk HL, Ferraro C and Miceli T:
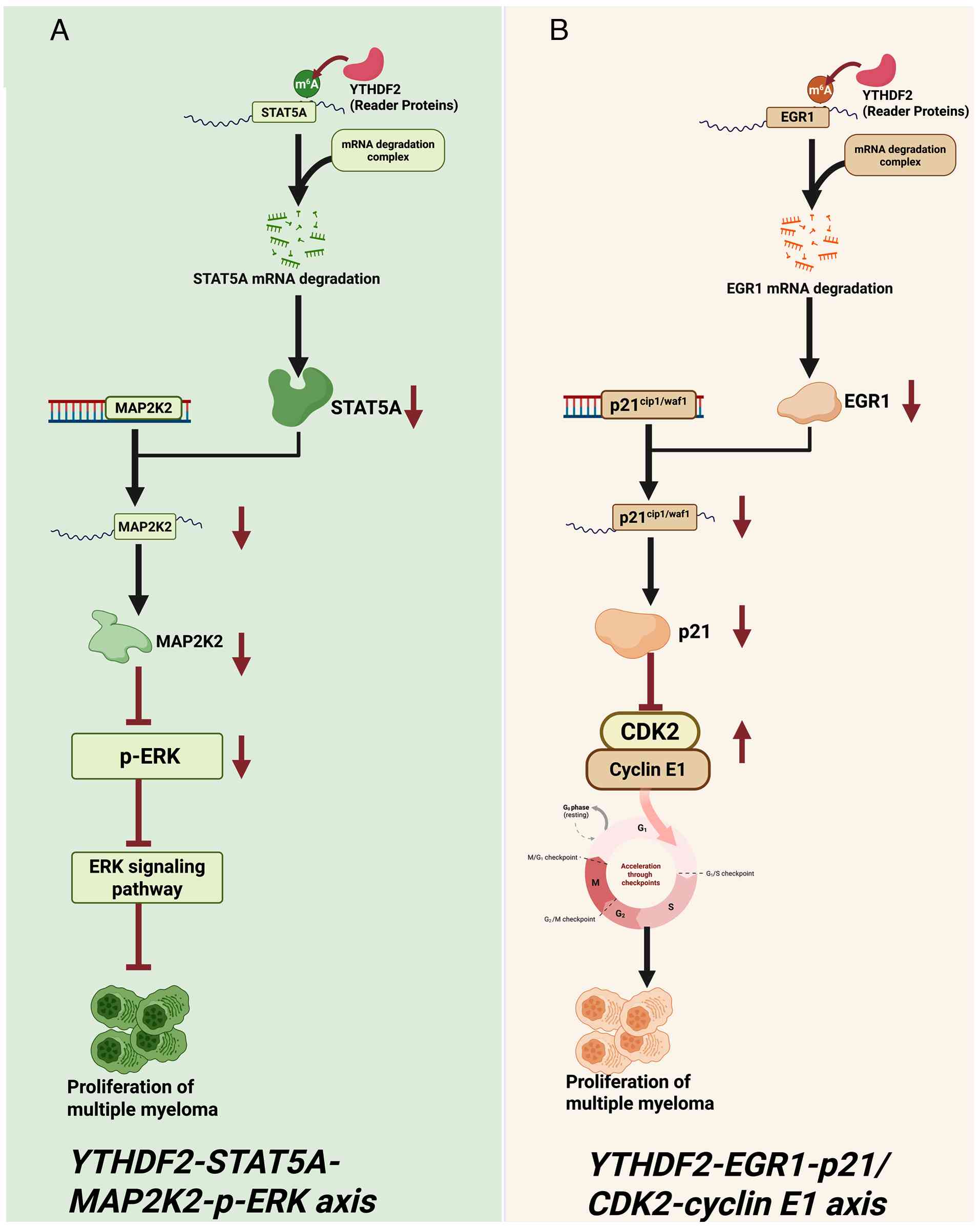
Hematopoietic stem cell transplantation for multiple myeloma. Semin
Oncol Nurs. 33:265–278. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Dima D, Jiang D, Singh DJ, Hasipek M, Shah
HS, Ullah F, Khouri J, Maciejewski JP and Jha BK: Multiple myeloma
therapy: Emerging trends and challenges. Cancers (Basel).
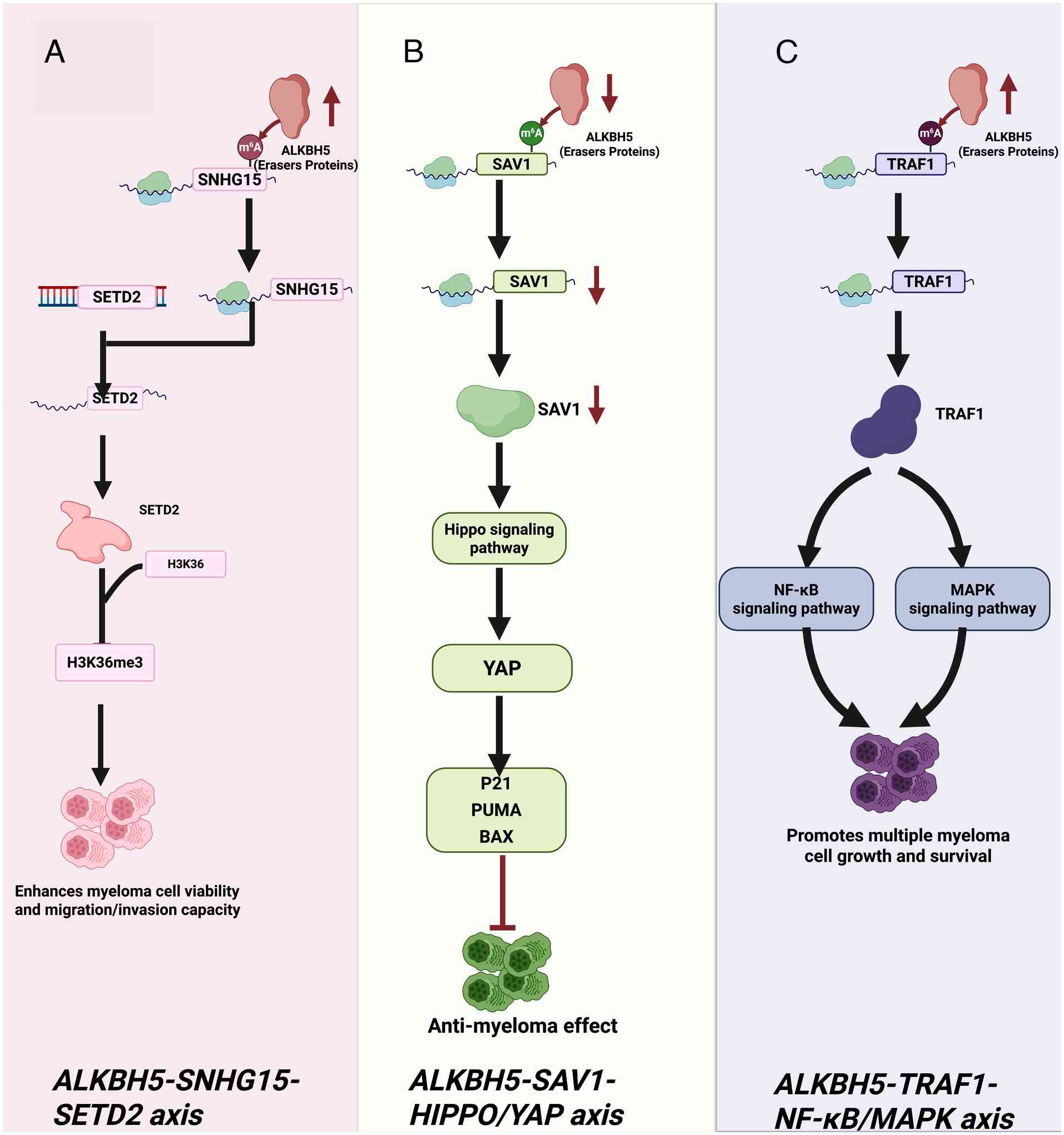
14:40822022. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sheykhhasan M, Ahmadieh-Yazdi A,
Vicidomini R, Poondla N, Tanzadehpanah H, Dirbaziyan A, Mahaki H,
Manoochehri H, Kalhor N and Dama P: CAR T therapies in multiple
myeloma: Unleashing the future. Cancer Gene Ther. 31:667–686. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Neri P, Leblay N, Lee H, Gulla A, Bahlis
NJ and Anderson KC: Just scratching the surface: novel treatment
approaches for multiple myeloma targeting cell membrane proteins.
Nat Rev Clin Oncol. 21:590–609. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Tam T, Smith E, Lozoya E, Heers H and
Andrew Allred P: Roadmap for new practitioners to navigate the
multiple myeloma landscape. Heliyon. 8:e105862022. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhu Y, Jian X, Chen S, An G, Jiang D, Yang
Q, Zhang J, Hu J, Qiu Y, Feng X, et al: Targeting gut microbial
nitrogen recycling and cellular uptake of ammonium to improve
bortezomib resistance in multiple myeloma. Cell Metab.
36:159–175.e8. 2024. View Article : Google Scholar
|
|
23
|
Neri P, Barwick BG, Jung D, Patton JC,
Maity R, Tagoug I, Stein CK, Tilmont R, Leblay N, Ahn S, et al:
ETV4-Dependent transcriptional plasticity maintains MYC expression
and results in IMiD resistance in multiple myeloma. Blood Cancer
Discov. 5:56–73. 2024. View Article : Google Scholar :
|
|
24
|
Zhang L, Peng X, Ma T, Liu J, Yi Z, Bai J,
Li Y, Li L and Zhang L: Natural killer cells affect the natural
course, drug resistance, and prognosis of multiple myeloma. Front
Cell Dev Biol. 12:13590842024. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Malard F, Neri P, Bahlis NJ, Terpos E,
Moukalled N, Hungria VTM, Manier S and Mohty M: Multiple myeloma.
Nat Rev Dis Prim. 10:452024. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Negrete-Rodríguez P, Gallardo-Pérez MM,
Lira-Lara O, Melgar-de-la-Paz M, Hamilton-Avilés LE, Ocaña-Ramm G,
Robles-Nasta M, Sánchez-Bonilla D, Olivares-Gazca JC, Mateos MV, et
al: Prevalence and consequences of a delayed diagnosis in multiple
myeloma: A single institution experience. Clin Lymphoma Myeloma
Leuk. 24:478–483. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yang P, Qu Y, Wang M, Chu B, Chen W, Zheng
Y, Niu T and Qian Z: Pathogenesis and treatment of multiple
myeloma. MedComm (2020). 3:e1462022. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Dimopoulos K, Gimsing P and Grønbæk K: The
role of epigenetics in the biology of multiple myeloma. Blood
Cancer J. 4:e2072014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Song H, Feng X, Zhang H, Luo Y, Huang J,
Lin M, Jin J, Ding X, Wu S, Huang H, et al: METTL3 and ALKBH5
oppositely regulate m6A modification of TFEB mRNA, which dictates
the fate of hypoxia/reoxygenation-treated cardiomyocytes.
Autophagy. 15:1419–1437. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Liu R, Gao Q, Foltz SM, Fowles JS, Yao L,
Wang JT, Cao S, Sun H, Wendl MC, Sethuraman S, et al: Co-evolution
of tumor and immune cells during progression of multiple myeloma.
Nat Commun. 12:25592021. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Vo JN, Wu YM, Mishler J, Hall S, Mannan R,
Wang L, Ning Y, Zhou J, Hopkins AC, Estill JC, et al: The genetic
heterogeneity and drug resistance mechanisms of relapsed refractory
multiple myeloma. Nat Commun. 13:37502022. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Xu L, Wen C, Xia J, Zhang H, Liang Y and
Xu X: Targeted immunotherapy: Harnessing the immune system to
battle multiple myeloma. Cell Death Discov. 10:552024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Skerget S, Penaherrera D, Chari A,
Jagannath S, Siegel DS, Vij R, Orloff G, Jakubowiak A, Niesvizky R,
Liles D, et al: Comprehensive molecular profiling of multiple
myeloma identifies refined copy number and expression subtypes. Nat
Genet. 56:1878–1889. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Rosselló-Tortella M, Ferrer G and Esteller
M: Epitranscriptomics in hematopoiesis and hematologic
malignancies. Blood Cancer Discov. 1:26–31. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Frye M, Jaffrey SR, Pan T, Rechavi G and
Suzuki T: RNA modifications: What have we learned and where are we
headed? Nat Rev Genet. 17:365–372. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Yang C, Han H and Lin S: RNA
epitranscriptomics: A promising new avenue for cancer therapy. Mol
Ther. 30:2–3. 2022. View Article : Google Scholar
|
|
37
|
Yang L, Tang L, Min Q, Tian H, Li L, Zhao
Y, Wu X, Li M, Du F, Chen Y, et al: Emerging role of RNA
modification and long noncoding RNA interaction in cancer. Cancer
Gene Ther. 31:816–830. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Pham CT, Rangan L and Schlenner S: RNA
modifications-a regulatory dimension yet to be deciphered in
immunity. Genes Immun. 24:281–282. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Xue C, Chu Q, Zheng Q, Jiang S, Bao Z, Su
Y, Lu J and Li L: Role of main RNA modifications in cancer:
N6-methyladenosine, 5-methylcytosine, and pseudouridine. Signal
Transduct Target Ther. 7:1422022. View Article : Google Scholar
|
|
40
|
Deng X, Qing Y, Horne D, Huang H and Chen
J: The roles and implications of RNA m6A modification in cancer.
Nat Rev Clin Oncol. 20:507–526. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Cui L, Ma R, Cai J, Guo C, Chen Z, Yao L,
Wang Y, Fan R, Wang X and Shi Y: RNA modifications: importance in
immune cell biology and related diseases. Signal Transduct Target
Ther. 7:3342022. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Yao L, Yin H, Hong M, Wang Y, Yu T, Teng
Y, Li T and Wu Q: RNA methylation in hematological malignancies and
its interactions with other epigenetic modifications. Leukemia.
35:1243–1257. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Qing Y, Su R and Chen J: RNA modifications
in hematopoietic malignancies: A new research frontier. Blood.
138:637–648. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Chao CT, Kuo FC and Lin SH: Epigenetically
regulated inflammation in vascular senescence and renal progression
of chronic kidney disease. Semin Cell Dev Biol. 154:305–315. 2024.
View Article : Google Scholar
|
|
45
|
Kan RL, Chen J and Sallam T: Crosstalk
between epitranscriptomic and epigenetic mechanisms in gene
regulation. Trends Genet. 38:182–193. 2022. View Article : Google Scholar :
|
|
46
|
Ye Z, Mayila M, Bu N, Hao W and
Maimaitiyiming Y: Epigenetic and epitranscriptomic landscape of
phthalate toxicity: Implications for human health and disease.
Environ Pollut. 391:1275592026. View Article : Google Scholar
|
|
47
|
Frye M, Harada BT, Behm M and He C: RNA
modifications modulate gene expression during development. Science.
361:1346–1349. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Qiu L, Jing Q, Li Y and Han J: RNA
modification: Mechanisms and therapeutic targets. Mol Biomed.
4:252023. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Sun H, Li K, Liu C and Yi C: Regulation
and functions of non-m6A mRNA modifications. Nat Rev Mol Cell Biol.
24:714–731. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Martinez De La Cruz B, Darsinou M and
Riccio A: From form to function: m6A methylation links mRNA
structure to metabolism. Adv Biol Regul. 87:1009262023. View Article : Google Scholar
|
|
51
|
Deng LJ, Deng WQ, Fan SR, Chen MF, Qi M,
Lyu WY, Qi Q, Tiwari AK, Chen JX, Zhang DM and Chen ZS: m6A
modification: Recent advances, anticancer targeted drug discovery
and beyond. Mol Cancer. 21:522022. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Su Y, Maimaitiyiming Y, Wang L, Cheng X
and Hsu CH: Modulation of phase separation by RNA: A glimpse on
N6-Methyladenosine modification. Front Cell Dev Biol. 9:7864542021.
View Article : Google Scholar
|
|
53
|
Mendel M, Delaney K, Pandey RR, Chen KM,
Wenda JM, Vågbø CB, Steiner FA, Homolka D and Pillai RS: Splice
site m6A methylation prevents binding of U2AF35 to inhibit RNA
splicing. Cell. 184:3125–3142.e25. 2021. View Article : Google Scholar
|
|
54
|
Aufgebauer CJ, Bland KM and Horner SM:
Modifying the antiviral innate immune response by selective
writing, erasing, and reading of m6A on viral and cellular RNA.
Cell Chem Biol. 31:100–109. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Lee SY, Kim JJ and Miller KM: Emerging
roles of RNA modifications in genome integrity. Brief Funct
Genomics. 20:106–112. 2021. View Article : Google Scholar :
|
|
56
|
Zaccara S, Ries RJ and Jaffrey SR:
Reading, writing and erasing mRNA methylation. Nat Rev Mol Cell
Biol. 20:608–624. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Esteva-Socias M and Aguilo F: METTL3 as a
master regulator of translation in cancer: Mechanisms and
implications. NAR Cancer. 6:zcae0092024. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Yan X, Liu F, Yan J, Hou M, Sun M, Zhang
D, Gong Z, Dong X, Tang C and Yin P: WTAP-VIRMA counteracts dsDNA
binding of the m(6)A writer METTL3-METTL14 complex and maintains
N(6)-adenosine methylation activity. Cell Discov. 9:1002023.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Yang Z, Zhang S, Xiong J, Xia T, Zhu R,
Miao M, Li K, Chen W, Zhang L, You Y and You B: The m6A
demethylases FTO and ALKBH5 aggravate the malignant progression of
nasopharyngeal carcinoma by coregulating ARHGAP35. Cell Death
Discov. 10:432024. View Article : Google Scholar
|
|
60
|
Zou Z, Sepich-Poore C, Zhou X, Wei J and
He C: The mechanism underlying redundant functions of the YTHDF
proteins. Genome Biol. 24:172023. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Xiao W, Adhikari S, Dahal U, Chen YS, Hao
YJ, Sun BF, Sun HY, Li A, Ping XL, Lai WY, et al: Nuclear m(6)A
Reader YTHDC1 Regulates mRNA splicing. Mol Cell. 61:507–519. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Huang H, Weng H, Sun W, Qin X, Shi H, Wu
H, Zhao BS, Mesquita A, Liu C, Yuan CL, et al: Recognition of RNA N
6 -methyladenosine by IGF2BP proteins enhances mRNA stability and
translation. Nat Cell Biol. 20:285–295. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Alarcón CR, Goodarzi H, Lee H, Liu X,
Tavazoie S and Tavazoie SF: HNRNPA2B1 is a mediator of
m6A-dependent nuclear RNA processing events. Cell. 162:1299–1308.
2015. View Article : Google Scholar
|
|
64
|
Xu W, Huang Z, Xiao Y, Li W, Xu M, Zhao Q
and Yi P: HNRNPC promotes estrogen receptor-positive breast cancer
cell cycle by stabilizing WDR77 mRNA in an m6A-dependent manner.
Mol Carcinog. 63:859–873. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Meyer KD, Patil DP, Zhou J, Zinoviev A,
Skabkin MA, Elemento O, Pestova TV, Qian SB and Jaffrey SR: 5' UTR
m6A promotes cap-independent translation. Cell. 163:999–1010. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Zhang F, Kang Y, Wang M, Li Y, Xu T, Yang
W, Song H, Wu H, Shu Q and Jin P: Fragile X mental retardation
protein modulates the stability of its m6A-marked messenger RNA
targets. Hum Mol Genet. 27:3936–3950. 2018.PubMed/NCBI
|
|
67
|
Mao-Mao, Zhang JJ, Xu YP, Shao MM and Wang
MC: Regulatory effects of natural products on N6-methyladenosine
modification: A novel therapeutic strategy for cancer. Drug Discov
Today. 29:1038752024. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Zhu ZM, Huo FC, Zhang J, Shan HJ and Pei
DS: Crosstalk between m6A modification and alternative splicing
during cancer progression. Clin Transl Med. 13:e14602023.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Jain S, Koziej L, Poulis P, Kaczmarczyk I,
Gaik M, Rawski M, Ranjan N, Glatt S and Rodnina MV: Modulation of
translational decoding by m6A modification of mRNA. Nat Commun.
14:47842023. View Article : Google Scholar
|
|
70
|
Qiao Y, Sun Q, Chen X, He L, Wang D, Su R,
Xue Y, Sun H and Wang H: Nuclear m6A Reader YTHDC1 promotes muscle
stem cell activation/proliferation by regulating mRNA splicing and
nuclear export. Elife. 12:e827032023. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Boulias K and Greer EL: Biological roles
of adenine methylation in RNA. Nat Rev Genet. 24:143–160. 2023.
View Article : Google Scholar
|
|
72
|
Wang Y, Li Y, Skuland T, Zhou C, Li A,
Hashim A, Jermstad I, Khan S, Dalen KT, Greggains GD, et al: The
RNA m6A landscape of mouse oocytes and preimplantation embryos. Nat
Struct Mol Biol. 30:703–709. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Wen J, Lv R, Ma H, Shen H, He C, Wang J,
Jiao F, Liu H, Yang P, Tan L, et al: Zc3h13 regulates nuclear RNA
m6A methylation and mouse embryonic stem cell self-renewal. Mol
Cell. 69:1028–1038.e6. 2018. View Article : Google Scholar
|
|
74
|
Yoon KJ, Ringeling FR, Vissers C, Jacob F,
Pokrass M, Jimenez-Cyrus D, Su Y, Kim NS, Zhu Y, Zheng L, et al:
Temporal control of mammalian cortical neurogenesis by m6A
methylation. Cell. 171:877–889.e17. 2017. View Article : Google Scholar
|
|
75
|
Wang L, Maimaitiyiming Y, Su K and Hsu CH:
RNA m6A Modification: The Mediator Between Cellular Stresses and
Biological Effects. RNA Technologies. 12:353–390. 2021. View Article : Google Scholar
|
|
76
|
Xiang Y, Laurent B, Hsu CH, Nachtergaele
S, Lu Z, Sheng W, Xu C, Chen H, Ouyang J, Wang S, et al: RNA m(6)A
methylation regulates the ultraviolet-induced DNA damage response.
Nature. 543:573–576. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Wang L, Zhan G, Maimaitiyiming Y, Su Y,
Lin S, Liu J, Su K, Lin J, Shen S, He W, et al: m6A modification
confers thermal vulnerability to HPV E7 oncotranscripts via reverse
regulation of its reader protein IGF2BP1 upon heat stress. Cell
Rep. 41:1115462022. View Article : Google Scholar
|
|
78
|
Engel M, Eggert C, Kaplick PM, Eder M, Röh
S, Tietze L, Namendorf C, Arloth J, Weber P, Rex-Haffner M, et al:
The role of m6A/m-RNA methylation in stress response regulation.
Neuron. 99:389–403.e9. 2018. View Article : Google Scholar
|
|
79
|
Chuong NN, Doan PPT, Wang L, Kim JH and
Kim J: Current insights into m6A RNA methylation and its emerging
role in plant circadian clock. Plants (Basel). 12:6242023.
|
|
80
|
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang
Y, Jin Y, Yang Y, Chen LL, Wang Y, et al: Extensive translation of
circular RNAs driven by N 6 -methyladenosine. Cell Res. 27:626–641.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Alarcón CR, Lee H, Goodarzi H, Halberg N
and Tavazoie SF: N6-methyladenosine marks primary microRNAs for
processing. Nature. 519:482–485. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Ma JZ, Yang F, Zhou CC, Liu F, Yuan JH,
Wang F, Wang TT, Xu QG, Zhou WP and Sun SH: METTL14 suppresses the
metastatic potential of hepatocellular carcinoma by modulating
N6-methyladenosine-dependent primary MicroRNA processing.
Hepatology. 65:529–543. 2017. View Article : Google Scholar
|
|
83
|
Deng S, Zhang J, Su J, Zuo Z, Zeng L, Liu
K, Zheng Y, Huang X, Bai R, Zhuang L, et al: RNA m6A regulates
transcription via DNA demethylation and chromatin accessibility.
Nat Genet. 54:1427–1437. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Höfler S and Duss O: Interconnections
between m6 A RNA modification, RNA structure, and protein-RNA
complex assembly. Life Sci Alliance. 7:e2023022402024. View Article : Google Scholar
|
|
85
|
Vaid R, Thombare K, Mendez A,
Burgos-Panadero R, Djos A, Jachimowicz D, Lundberg KI, Bartenhagen
C, Kumar N, Tümmler C, et al: MET TL3 drives telomere targ eting of
TERRA lncRNA through m 6 A-dependent R-loop formation: A
therapeutic target for ALT-positive neuroblastoma. Nucleic Acids
Res. 52:2648–2671. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Lee SY, Lee SH, Kwak MJ, Kim JY, Perren
JO, Miller KM and Kim JJ: Depletion of BRD9-mediated R-loop
accumulation inhibits leukemia cell growth via
transcription-replication conflict. Nucleic Acids Res.
53:gkaf6132025. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Verghese M, Wilkinson E and He YY: Role of
RNA modifications in carcinogenesis and carcinogen damage response.
Mol Carcinog. 62:24–37. 2023. View Article : Google Scholar :
|
|
88
|
Yang J, Xu J, Wang W, Zhang B, Yu X and
Shi S: Epigenetic regulation in the tumor microenvironment:
Molecular mechanisms and therapeutic targets. Signal Transduct
Target Ther. 8:2102023. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Lin S and Kuang M: RNA
modification-mediated mRNA translation regulation in liver cancer:
Mechanisms and clinical perspectives. Nat Rev Gastroenterol
Hepatol. 21:267–281. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Dong L, Cao Y, Hou Y and Liu G:
N6-methyladenosine RNA methylation: A novel regulator of the
development and function of immune cells. J Cell Physiol.
237:329–345. 2022. View Article : Google Scholar
|
|
91
|
Delaunay S, Helm M and Frye M: RNA
modifications in physiology and disease: Towards clinical
applications. Nat Rev Genet. 25:104–122. 2024. View Article : Google Scholar
|
|
92
|
Berdasco M and Esteller M: Towards a
druggable epitranscriptome: Compounds that target RNA modifications
in cancer. Br J Pharmacol. 179:2868–2889. 2022. View Article : Google Scholar
|
|
93
|
Zheng J, Lu Y, Lin Y, Si S, Guo B, Zhao X
and Cui L: Epitranscriptomic modifications in mesenchymal stem cell
differentiation: Advances, mechanistic insights, and beyond. Cell
Death Differ. 31:9–27. 2024. View Article : Google Scholar
|
|
94
|
Delaunay S and Frye M: RNA modifications
regulating cell fate in cancer. Nat Cell Biol. 21:552–559. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Shi J, Zhang Q, Yin X, Ye J, Gao S, Chen
C, Yang Y, Wu B, Fu Y, Zhang H, et al: Stabilization of IGF2BP1 by
USP10 promotes breast cancer metastasis via CPT1A in an
m6A-dependent manner. Int J Biol Sci. 19:449–464. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Lv D, Gimple RC, Zhong C, Zhong C, Wu Q,
Yang K, Prager BC, Godugu B, Qiu Z, Zhao L, et al: PDGF signaling
inhibits mitophagy in glioblastoma stem cells through
N6-methyladenosine. Dev Cell. 57:1466–1481.e6. 2022. View Article : Google Scholar
|
|
97
|
Feng Y, Yuan P, Guo H, Gu L, Yang Z, Wang
J, Zhu W, Zhang Q, Cao J, Wang L and Jiao Y: METTL3 mediates
epithelial-mesenchymal transition by modulating FOXO1 mRNA
N6-Methyladenosine-Dependent YTHDF2 Binding: A novel mechanism of
radiation-induced lung injury. Adv Sci (Weinh). 10:e22047842023.
View Article : Google Scholar
|
|
98
|
Geula S, Moshitch-Moshkovitz S,
Dominissini D, Mansour AA, Kol N, Salmon-Divon M, Hershkovitz V,
Peer E, Mor N, Manor YS, et al: Stem Cells m6A mRNA methylation
facilitates resolution of naïve pluripotency toward
differentiation. Science. 347:1002–1006. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Chen Y, Peng C, Chen J, Chen D, Yang B, He
B, Hu W, Zhang Y, Liu H, Dai L, et al: WTAP facilitates progression
of hepatocellular carcinoma via m6A-HuR-dependent epigenetic
silencing of ETS1. Mol Cancer. 18:1272019. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Zhang S, Zhao BS, Zhou A, Lin K, Zheng S,
Lu Z, Chen Y, Sulman EP, Xie K, Bögler O, et al: m6A demethylase
ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by
sustaining FOXM1 expression and cell proliferation program. Cancer
Cell. 31:591–606.e6. 2017. View Article : Google Scholar
|
|
101
|
Xiao S, Ma S, Sun B, Pu W, Duan S, Han J,
Hong Y, Zhang J, Peng Y, He C, et al: The tumor-intrinsic role of
the m6A reader YTHDF2 in regulating immune evasion. Sci Immunol.
9:eadl21712024. View Article : Google Scholar
|
|
102
|
Wang X, Zhao BS, Roundtree IA, Lu Z, Han
D, Ma H, Weng X, Chen K, Shi H and He C: N6-methyladenosine
modulates messenger RNA translation efficiency. Cell.
161:1388–1399. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Zeng L, Huang X, Zhang J, Lin D and Zheng
J: Roles and implications of mRNA N6-methyladenosine in cancer.
Cancer Commun (Lond). 43:729–748. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Zhuang H, Yu B, Tao D, Xu X, Xu Y, Wang J,
Jiao Y and Wang L: The role of m6A methylation in therapy
resistance in cancer. Mol Cancer. 22:912023. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Jin Z, MacPherson K, Liu Z and Vu LP: RNA
modifications in hematological malignancies. Int J Hematol.
117:807–820. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Lv J, Zhang Y, Gao S, Zhang C, Chen Y, Li
W, Yang YG, Zhou Q and Liu F: Endothelial-specific m6A modulates
mouse hematopoietic stem and progenitor cell development via Notch
signaling. Cell Res. 28:249–252. 2018. View Article : Google Scholar
|
|
107
|
Li Z, Weng H, Su R, Weng X, Zuo Z, Li C,
Huang H, Nachtergaele S, Dong L, Hu C, et al: FTO plays an
oncogenic role in acute myeloid leukemia as a N6-Methyladenosine
RNA demethylase. Cancer Cell. 31:127–141. 2017. View Article : Google Scholar
|
|
108
|
Vu LP, Pickering BF, Cheng Y, Zaccara S,
Nguyen D, Minuesa G, Chou T, Chow A, Saletore Y, MacKay M, et al:
The N 6 -methyl-adenosine (m 6 A)-forming enzyme METTL3 controls
myeloid differentiation of normal hematopoietic and leukemia cells.
Nat Med. 23:1369–1376. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Barbieri I, Tzelepis K, Pandolfini L, Shi
J, Millán-Zambrano G, Robson SC, Aspris D, Migliori V, Bannister
AJ, Han N, et al: Promoter-bound METTL3 maintains myeloid leukaemia
by m6A-dependent translation control. Nature. 552:126–131. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Zhang N, Shen Y, Li H, Chen Y, Zhang P,
Lou S and Deng J: The m6A reader IGF2BP3 promotes acute myeloid
leukemia progression by enhancing RCC2 stability. Exp Mol Med.
54:194–205. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Wilkinson E, Cui YH and He YY:
Context-dependent roles of RNA modifications in stress responses
and diseases. Int J Mol Sci. 22:19492021. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Wang J, Zuo Y, Lv C, Zhou M and Wan Y:
N6-methyladenosine regulators are potential prognostic biomarkers
for multiple myeloma. IUBMB Life. 75:137–148. 2023. View Article : Google Scholar :
|
|
113
|
Deng Y, Zhu H and Peng H: Enhancing
staging in multiple myeloma using an m6A regulatory gene-pairing
model. Clin Exp Med. 25:402025. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Bao J, Xu T, Wang W, Xu H, Chen X and Xia
R: N6-methyladenosine-induced miR-182-5p promotes multiple myeloma
tumorigenesis by regulating CAMK2N1. Mol Cell Biochem.
479:3077–3089. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Che F, Ye X, Wang Y, Wang X, Ma S, Tan Y,
Mao Y and Luo Z: METTL3 facilitates multiple myeloma tumorigenesis
by enhancing YY1 stability and pri-microRNA-27 maturation in
m6A-dependent manner. Cell Biol Toxicol. 39:2033–2050. 2023.
View Article : Google Scholar
|
|
116
|
Huang X, Yang Z, Li Y and Long X: m6A
methyltransferase METTL3 facilitates multiple myeloma cell growth
through the m6A modification of BZW2. Ann Hematol. 102:1801–1810.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Lu X, Li Y, Li R, Zhang J, Peng J and
Zhang Y: Regulatory role of the METTL3/MALAT1 axis in multiple
myeloma progression. J Bone Oncol. 53:1006952025. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Zhao Y, Zhang E, Lv N, Ma L, Yao S, Yan M,
Zi F, Deng G, Liu X, He J, et al: Metformin and FTY720
synergistically induce apoptosis in multiple myeloma cells. Cell
Physiol Biochem. 48:785–800. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Gao L, Li L, Hu J, Li G, Zhang Y, Dai X,
De Z and Xu F: Metformin inhibits multiple myeloma serum-induced
endothelial cell thrombosis by down-regulating miR-532. Ann Vasc
Surg. 85:347–357.e2. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Chen CJ, Huang JY, Huang JQ, Deng JY,
Shangguan XH, Chen AZ, Chen LT and Wu WH: Metformin attenuates
multiple myeloma cell proliferation and encourages apoptosis by
suppressing METTL3-mediated m6A methylation of THRAP3, RBM25, and
USP4. Cell Cycle. 22:986–1004. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Jiang J, Zhong F, Xiao Z, Yao F, Liu J, Li
M, Zeng H, Qiu Y, Zhang J, Zhang H, et al: METTL5 regulates
SEPHS2-mediated selenoprotein synthesis to promote multiple myeloma
survival and progression. Cell Death Dis. 16:5852025. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Wu Y, Luo Y, Yao X, Shi X, Xu Z, Re J, Shi
M, Li M, Liu J, He Y and Du X: KIAA1429 increases FOXM1 expression
through YTHDF1-mediated m6A modification to promote aerobic
glycolysis and tumorigenesis in multiple myeloma. Cell Biol
Toxicol. 40:582024. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Su Q, Liu W, Wang P and Wang M: Long
non-coding RNA FEZF1-AS1 suppresses ferroptosis in multiple myeloma
cells through KIAA1429-mediated m6A modification. Hum Cell.
38:1782025. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Xu H, Xu M, Ding J and Bao J: WTAP
promotes the proliferation of multiple myeloma by regulating the
hippo pathway through m(6)A modification of MAP6D1. Leuk Lymphoma.
67:148–163. 2026. View Article : Google Scholar
|
|
125
|
Jia Y, Yu X, Liu R, Shi L, Jin H, Yang D,
Zhang X, Shen Y, Feng Y, Zhang P, et al: PRMT1 methylation of WTAP
promotes multiple myeloma tumorigenesis by activating oxidative
phosphorylation via m6A modification of NDUFS6. Cell Death Dis.
14:5122023. View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Hua Z, Wei R, Guo M, Lin Z, Yu X, Li X, Gu
C and Yang Y: YTHDF2 promotes multiple myeloma cell proliferation
via STAT5A/MAP2K2/p-ERK axis. Oncogene. 41:1482–1491. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Liu R, Miao J, Jia Y, Kong G, Hong F, Li
F, Zhai M, Zhang R, Liu J, Xu X, et al: N6-methyladenosine reader
YTHDF2 promotes multiple myeloma cell proliferation through
EGR1/p21cip1/waf1/CDK2-Cyclin E1 axis-mediated cell cycle
transition. Oncogene. 42:1607–1619. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Xu J, Wang Y, Ren L, Li P and Liu P:
IGF2BP1 promotes multiple myeloma with chromosome 1q gain via
increasing CDC5L expression in an m6A-dependent manner. Genes Dis.
12:1012142024. View Article : Google Scholar
|
|
129
|
Bernstein ZS, Kim EB and Raje N: Bone
disease in multiple myeloma: Biologic and clinical implications.
Cells. 11:23082022. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Terpos E, Ntanasis-Stathopoulos I,
Gavriatopoulou M and Dimopoulos MA: Pathogenesis of bone disease in
multiple myeloma: From bench to bedside. Blood Cancer J. 8:72018.
View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Liu R, Zhong Y, Chen R, Chu C, Liu G, Zhou
Y, Huang Y, Fang Z and Liu H: m6A reader hnRNPA2B1 drives multiple
myeloma osteolytic bone disease. Theranostics. 12:7760–7774. 2022.
View Article : Google Scholar
|
|
132
|
Jiang F, Tang X, Tang C, Hua Z, Ke M, Wang
C, Zhao J, Gao S, Jurczyszyn A, Janz S, et al: HNRNPA2B1 promotes
multiple myeloma progression by increasing AKT3 expression via
m6A-dependent stabilization of ILF3 mRNA. J Hematol Oncol.
14:542021. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Guo Y, Jia C, Wang X, Luo K, Chi L, Xu Q,
Gong T and Quan L: HNRNPA2B1 promotes the progression of multiple
myeloma via endoplasmic reticulum stress and autophagy mediated by
CK2 Kinase. J Proteome Res. 24:5921–5931. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Tang J, Li J, Qin S, Xiao Y, Liu J, Chen X
and Zhang Y: Identification and validation of the m6A-binding
protein LRPPRC to promote tumorigenesis in multiple myeloma.
Hematology. 30:25230822025. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Yao L, Li T, Teng Y, Guo J, Zhang H, Xia L
and Wu Q: ALKHB5-demethylated lncRNA SNHG15 promotes myeloma
tumorigenicity by increasing chromatin accessibility and recruiting
H3K36me3 modifier SETD2. Am J Physiol Cell Physiol. 326:C684–C697.
2024. View Article : Google Scholar :
|
|
136
|
Yu T, Yao L, Yin H, Teng Y, Hong M and Wu
Q: ALKBH5 promotes multiple myeloma tumorigenicity through inducing
m6A-demethylation of SAV1 mRNA and myeloma stem cell phenotype. Int
J Biol Sci. 18:2235–2248. 2022. View Article : Google Scholar
|
|
137
|
Qu J, Hou Y, Chen Q, Chen J, Li Y, Zhang
E, Gu H, Xu R, Liu Y, Cao W, et al: RNA demethylase ALKBH5 promotes
tumorigenesis in multiple myeloma via TRAF1-mediated activation of
NF-κB and MAPK signaling pathways. Oncogene. 41:400–413. 2022.
View Article : Google Scholar :
|
|
138
|
Badraldin SQ, Alfarttoosi KH, Sameer HN,
Bishoyi AK, Ganesan S, Shankhyan A, Ray S, Nathiya D, Yaseen A,
Athab ZH and Adil M: Mechanistic role of FTO in cancer
pathogenesis, immune evasion, chemotherapy resistance, and
immunotherapy response. Semin Oncol. 52:1523682025. View Article : Google Scholar : PubMed/NCBI
|
|
139
|
Xu A, Zhang J, Zuo L, Yan H, Chen L, Zhao
F, Fan F, Xu J, Zhang B, Zhang Y, et al: FTO promotes multiple
myeloma progression by posttranscriptional activation of HSF1 in an
m6A-YTHDF2-dependent manner. Mol Ther. 30:1104–1118. 2022.
View Article : Google Scholar
|
|
140
|
Li JJ, Yu T, Zeng P, Tian J, Liu P, Qiao
S, Wen S, Hu Y, Liu Q, Lu W, et al: Wild-type IDH2 is a therapeutic
target for triple-negative breast cancer. Nat Commun. 15:34452024.
View Article : Google Scholar : PubMed/NCBI
|
|
141
|
Song S, Fan G, Li Q, Su Q, Zhang X, Xue X,
Wang Z, Qian C, Jin Z, Li B and Zhuang W: IDH2 contributes to
tumorigenesis and poor prognosis by regulating m6A RNA methylation
in multiple myeloma. Oncogene. 40:5393–5402. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
142
|
Zhang Y, Deng Z, Sun S, Xie S, Jiang M,
Chen B, Gu C and Yang Y: NAT10 acetylates BCL-XL mRNA to promote
the proliferation of multiple myeloma cells through PI3K-AKT
pathway. Front Oncol. 12:9678112022. View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Liu H, Zhang X, Lu Q and Zhang H: NAT10
contributes to the progression of multiple myeloma through ac4C
modification of GPR37. Hematology. 30:25557792025. View Article : Google Scholar : PubMed/NCBI
|
|
144
|
Ren H, Liu C, Wu H, Wang Z, Chen S, Zhang
X, Ren J, Qiu H and Zhou L: m5C Regulator-mediated methylation
modification clusters contribute to the immune microenvironment
regulation of multiple myeloma. Front Genet. 13:9201642022.
View Article : Google Scholar
|
|
145
|
Jiang Y, Sun J, Chen Y, Cheng L, Feng S,
Wang Y and Sun C: NSUN2-mediated RNA m(5)C modification drives
multiple myeloma progression by enhancing the stability of HIP1
mRNA. Sci Rep. 15:278882025. View Article : Google Scholar : PubMed/NCBI
|
|
146
|
Fu J, Han X, Gao W, Yu M and Cui X: m1A
regulator-mediated methylation modifications and gene signatures
and their prognostic value in multiple myeloma. Exp Ther Med.
29:182025. View Article : Google Scholar
|
|
147
|
Cohen YC, Zada M, Wang SY, Bornstein C,
David E, Moshe A, Li B, Shlomi-Loubaton S, Gatt ME, Gur C, et al:
Identification of resistance pathways and therapeutic targets in
relapsed multiple myeloma patients through single-cell sequencing.
Nat Med. 27:491–503. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
148
|
Ferguson ID, Patiño-Escobar B, Tuomivaara
ST, Lin YT, Nix MA, Leung KK, Kasap C, Ramos E, Nieves Vasquez W,
Talbot A, et al: The surfaceome of multiple myeloma cells suggests
potential immunotherapeutic strategies and protein markers of drug
resistance. Nat Commun. 13:41212022. View Article : Google Scholar : PubMed/NCBI
|
|
149
|
Sun J, Corradini S, Azab F, Shokeen M, Muz
B, Miari KE, Maksimos M, Diedrich C, Asare O, Alhallak K, et al:
IL-10R inhibition reprograms tumor-associated macrophages and
reverses drug resistance in multiple myeloma. Leukemia.
38:2355–2365. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
150
|
Bird S and Pawlyn C: IMiD resistance in
multiple myeloma: Current understanding of the underpinning biology
and clinical impact. Blood. 142:131–140. 2023.PubMed/NCBI
|
|
151
|
Tzelepis K, Rausch O and Kouzarides T:
RNA-modifying enzymes and their function in a chromatin context.
Nat Struct Mol Biol. 26:858–862. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
152
|
Chen H, Jia B, Zhang Q and Zhang Y:
Meclofenamic acid restores gefinitib sensitivity by downregulating
breast cancer resistance protein and multidrug resistance protein 7
via FTO/m6A-Demethylation/c-Myc in non-small cell lung cancer.
Front Oncol. 12:8706362022. View Article : Google Scholar : PubMed/NCBI
|
|
153
|
Yuan J, Guan W, Li X, Wang F, Liu H and Xu
G: RBM15-mediating MDR1 mRNA m6A methylation regulated by the TGF-β
signaling pathway in paclitaxel-resistant ovarian cancer. Int J
Oncol. 63:1122023. View Article : Google Scholar
|
|
154
|
Liu R, Shen Y, Hu J, Wang X, Wu D, Zhai M,
Bai J and He A: Comprehensive Analysis of m6A RNA methylation
regulators in the prognosis and immune microenvironment of multiple
myeloma. Front Oncol. 11:7319572021. View Article : Google Scholar : PubMed/NCBI
|
|
155
|
Wang C, Li L, Li M, Wang W and Jiang Z:
FTO promotes Bortezomib resistance via m6A-dependent
destabilization of SOD2 expression in multiple myeloma. Cancer Gene
Ther. 30:622–628. 2023. View Article : Google Scholar
|
|
156
|
Prabhu KS, Ahmad F, Kuttikrishnan S, Leo
R, Ali TA, Izadi M, Mateo JM, Alam M, Ahmad A, Al-Shabeeb Akil AS,
et al: Bortezomib exerts its anti-cancer activity through the
regulation of Skp2/p53 axis in non-melanoma skin cancer cells and
C. elegans. Cell Death Discov. 10:2252024. View Article : Google Scholar : PubMed/NCBI
|
|
157
|
Sogbein O, Paul P, Umar M, Chaari A,
Batuman V and Upadhyay R: Bortezomib in cancer therapy: Mechanisms,
side effects, and future proteasome inhibitors. Life Sci.
358:1231252024. View Article : Google Scholar : PubMed/NCBI
|
|
158
|
Hurt EM, Thomas SB, Peng B and Farrar WL:
Integrated molecular profiling of SOD2 expression in multiple
myeloma. Blood. 109:3953–3962. 2007. View Article : Google Scholar
|
|
159
|
Hodge DR, Peng B, Pompeia C, Thomas S, Cho
E, Clausen PA, Marquez VE and Farrar WL: Epigenetic silencing of
manganese superoxide dismutase (SOD-2) in KAS 6/1 human multiple
myeloma cells increases cell proliferation. Cancer Biol Ther.
4:585–592. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
160
|
Song IS, Kim HK, Lee SR, Jeong SH, Kim N,
Ko KS, Rhee BD and Han J: Mitochondrial modulation decreases the
bortezomib-resistance in multiple myeloma cells. Int J Cancer.
133:1357–1367. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
161
|
Jomova K, Alomar SY, Alwasel SH,
Nepovimova E, Kuca K and Valko M: Several lines of antioxidant
defense against oxidative stress: antioxidant enzymes,
nanomaterials with multiple enzyme-mimicking activities, and
low-molecular-weight antioxidants. Arch Toxicol. 98:1323–1367.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
162
|
Huang R, Chen H, Liang J, Li Y, Yang J,
Luo C, Tang Y, Ding Y, Liu X, Yuan Q, et al: Dual role of reactive
oxygen species and their application in cancer therapy. J Cancer.
12:5543–5561. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
163
|
Jahankhani K, Taghipour N, Nikoonezhad M,
Behboudi H, Mehdizadeh M, Kadkhoda D, Hajifathali A and Mosaffa N:
Adjuvant therapy with zinc supplementation; anti-inflammatory and
anti-oxidative role in multiple myeloma patients receiving
autologous hematopoietic stem cell transplantation: A randomized
controlled clinical trial. Biometals. 37:1609–1627. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
164
|
Yu W, Cao D, Zhou H, Hu Y and Guo T:
PGC-1α is responsible for survival of multiple myeloma cells under
hyperglycemia and chemotherapy. Oncol Rep. 33:2086–2092. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
165
|
Zhang Y, Cao X, Li W, Cui Z, Mao J, Yao R
and Liu L: ALKBH5 reverses romidepsin-mediated anti-multiple
myeloma activity via regulation of m6A modification of FOXM1.
Biochem Pharmacol. 239:1169982025. View Article : Google Scholar : PubMed/NCBI
|
|
166
|
Quan L, Jia C, Guo Y, Chen Y, Wang X, Xu Q
and Zhang Y: HNRNPA2B1-mediated m6A modification of TLR4 mRNA
promotes progression of multiple myeloma. J Transl Med. 20:5372022.
View Article : Google Scholar : PubMed/NCBI
|
|
167
|
Giallongo C, Tibullo D, Puglisi F, Barbato
A, Vicario N, Cambria D, Parrinello NL, Romano A, Conticello C,
Forte S, et al: Inhibition of TLR4 signaling affects mitochondrial
fitness and overcomes bortezomib resistance in myeloma plasma
cells. Cancers (Basel). 12:19992020. View Article : Google Scholar : PubMed/NCBI
|
|
168
|
Bagratuni T, Sklirou AD, Kastritis E,
Liacos CI, Spilioti C, Eleutherakis-Papaiakovou E, Kanellias N,
Gavriatopoulou M, Terpos E, Trougakos IP and Dimopoulos MA:
Toll-like receptor 4 activation promotes multiple myeloma cell
growth and survival via suppression of the endoplasmic reticulum
stress factor chop. Sci Rep. 9:32452019. View Article : Google Scholar : PubMed/NCBI
|
|
169
|
Jiang S, Gao L, Li J, Zhang F, Zhang Y and
Liu J: N6-methyladenosine-modified circ_0000337 sustains bortezomib
resistance in multiple myeloma by regulating DNA repair. Front Cell
Dev Biol. 12:13832322024. View Article : Google Scholar : PubMed/NCBI
|
|
170
|
Wang G and Wu W, He D, Wang J, Kong H and
Wu W: N6-methyladenosine-mediated upregulation of H19 promotes
resistance to bortezomib by modulating the miR-184/CARM1 axis in
multiple myeloma. Clin Exp Med. 25:1022025. View Article : Google Scholar : PubMed/NCBI
|
|
171
|
Wang Z, He J, Bach DH, Huang YH, Li Z, Liu
H, Lin P and Yang J: Induction of m6A methylation in adipocyte
exosomal LncRNAs mediates myeloma drug resistance. J Exp Clin
Cancer Res. 41:42022. View Article : Google Scholar
|
|
172
|
Sun X, Zhou Y, Zhu W and Chen H: Research
progress on N6-methyladenosine and non-coding RNA in multiple
myeloma. Discov Oncol. 16:6152025. View Article : Google Scholar : PubMed/NCBI
|
|
173
|
Huang Y, Xia W, Dong Z and Yang CG:
Chemical inhibitors targeting the oncogenic m6A Modifying Proteins.
Acc Chem Res. 56:3010–3022. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
174
|
He B, Hu Y, Wu Y, Wang C, Gao L, Gong C,
Li Z, Gao N, Yang H, Xiao Y and Yang S: Helicobacter pylori CagA
elevates FTO to induce gastric cancer progression via a
'hit-and-run' paradigm. Cancer Commun (Lond). 45:608–631. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
175
|
Xiao L, Li X, Mu Z, Zhou J, Zhou P, Xie C
and Jiang S: FTO inhibition enhances the antitumor effect of
temozolomide by targeting MYC-miR-155/23a cluster-MXI1 feedback
circuit in glioma. Cancer Res. 80:3945–3958. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
176
|
Xu Y, Zhou J, Li L, Yang W, Zhang Z, Zhang
K, Ma K, Xie H, Zhang Z, Cai L, et al: FTO-mediated autophagy
promotes progression of clear cell renal cell carcinoma via
regulating SIK2 mRNA stability. Int J Biol Sci. 18:5943–5962. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
177
|
Jiang L, Liang R, Luo Q, Chen Z and Song
G: Targeting FTO suppresses hepatocellular carcinoma by inhibiting
ERBB3 and TUBB4A expression. Biochem Pharmacol. 226:1163752024.
View Article : Google Scholar : PubMed/NCBI
|
|
178
|
Zhang J, Li G, Wu R, Shi L, Tian C, Jiang
H, Che H, Jiang Y, Jin Z, Yu R, et al: The m6A RNA demethylase FTO
promotes radioresistance and stemness maintenance of glioma stem
cells. Cell Signal. 132:1117822025. View Article : Google Scholar : PubMed/NCBI
|
|
179
|
Yang Q and Al-Hendy A: The functional role
and regulatory mechanism of FTO m6A RNA demethylase in human
uterine leiomyosarcoma. Int J Mol Sci. 24:79572023. View Article : Google Scholar
|
|
180
|
Su R, Dong L, Li Y, Gao M, Han L,
Wunderlich M, Deng X, Li H, Huang Y, Gao L, et al: Targeting FTO
suppresses cancer stem cell maintenance and immune evasion. Cancer
Cell. 38:79–96.e11. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
181
|
Huff S, Kummetha IR, Zhang L, Wang L, Bray
W, Yin J, Kelley V, Wang Y and Rana TM: Rational design and
optimization of m6A-RNA Demethylase FTO inhibitors as anticancer
agents. J Med Chem. 65:10920–10937. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
182
|
Peng S, Xiao W, Ju D, Sun B, Hou N, Liu Q,
Wang Y, Zhao H, Gao C, Zhang S, et al: Identification of entacapone
as a chemical inhibitor of FTO mediating metabolic regulation
through FOXO1. Sci Transl Med. 11:eaau71162019. View Article : Google Scholar : PubMed/NCBI
|
|
183
|
Ramedani F, Jafari SM, Saghaeian Jazi M,
Mohammadi Z and Asadi J: Anti-cancer effect of entacaponeon
esophageal cancer cells via apoptosis induction and cell cycle
modulation. Cancer Rep (Hoboken). 6:e17592023.
|
|
184
|
Yankova E, Blackaby W, Albertella M, Rak
J, De Braekeleer E, Tsagkogeorga G, Pilka ES, Aspris D, Leggate D,
Hendrick AG, et al: Small-molecule inhibition of METTL3 as a
strategy against myeloid leukaemia. Nature. 593:597–601. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
185
|
Sun Y, Shen W, Hu S, Lyu Q, Wang Q, Wei T,
Zhu W and Zhang J: METTL3 promotes chemoresistance in small cell
lung cancer by inducing mitophagy. J Exp Clin Cancer Res.
42:652023. View Article : Google Scholar : PubMed/NCBI
|
|
186
|
Jin X, Lv Y, Bie F, Duan J, Ma C, Dai M,
Chen J, Lu L, Xu S, Zhou J, et al: METTL3 confers oxaliplatin
resistance through the activation of G6PD-enhanced pentose
phosphate pathway in hepatocellular carcinoma. Cell Death Differ.
32:466–479. 2025. View Article : Google Scholar :
|
|
187
|
Hao S, Sun H, Sun H, Zhang B, Ji K, Liu P,
Nie F and Han W: STM2457 Inhibits the invasion and metastasis of
pancreatic cancer by down-regulating BRAF-Activated Noncoding RNA
N6-Methyladenosine modification. Curr Issues Mol Biol.
45:8852–8863. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
188
|
Tang H, Zhang R and Zhang A:
Small-molecule inhibitors targeting RNA m(6)A modifiers for cancer
therapeutics : Latest advances and future perspectives. J Med Chem.
68:18114–18142. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
189
|
Du Y, Yuan Y, Xu L, Zhao F, Wang W, Xu Y
and Tian X: Discovery of METTL3 small molecule inhibitors by
virtual screening of natural products. Front Pharmacol.
13:8781352022. View Article : Google Scholar : PubMed/NCBI
|
|
190
|
Dolbois A, Bedi RK, Bochenkova E, Müller
A, Moroz-Omori EV, Huang D and Caflisch A:
1,4,9-Triazaspiro[5.5]undecan-2-one derivatives as potent and
selective METTL3 Inhibitors. J Med Chem. 64:127382021. View Article : Google Scholar : PubMed/NCBI
|
|
191
|
Li J and Gregory RI: Mining for METTL3
inhibitors to suppress cancer. Nat Struct Mol Biol. 28:460–462.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
192
|
Malacrida A, Di Domizio A, Bentivegna A,
Cislaghi G, Messuti E, Tabano SM, Giussani C, Zuliani V, Rivara M
and Nicolini G: MV1035 overcomes temozolomide resistance in
patient-derived glioblastoma stem cell lines. Biology (Basel).
11:702022.PubMed/NCBI
|
|
193
|
Li N, Kang Y, Wang L, Huff S, Tang R, Hui
H, Agrawal K, Gonzalez GM, Wang Y, Patel SP and Rana TM: ALKBH5
regulates anti-PD-1 therapy response by modulating lactate and
suppressive immune cell accumulation in tumor microenvironment.
Proc Natl Acad Sci USA. 117:20159–20170. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
194
|
Tang W, Xu N, Zhou J, He Z, Lenahan C,
Wang C, Ji H, Liu B, Zou Y, Zeng H and Guo H: ALKBH5 promotes
PD-L1-mediated immune escape through m6A modification of ZDHHC3 in
glioma. Cell Death Discov. 8:4972022. View Article : Google Scholar : PubMed/NCBI
|
|
195
|
Schott A, Simon T, Müller S, Rausch A,
Busch B, Glaß M, Misiak D, Dipto M, Elrewany H, Peters LM, et al:
The IGF2BP1 oncogene is a druggable m6A-dependent enhancer of
YAP1-driven gene expression in ovarian cancer. NAR Cancer.
7:zcaf0062025. View Article : Google Scholar
|
|
196
|
Singh A, Singh V, Wallis N, Abis G,
Oberman F, Wood T, Dhamdhere M, Gershon T, Ramos A, Yisraeli J, et
al: Development of a specific and potent IGF2BP1 inhibitor: A
promising therapeutic agent for IGF2BP1-expressing cancers. Eur J
Med Chem. 263:1159402024. View Article : Google Scholar
|
|
197
|
Feng P, Chen D, Wang X, Li Y, Li Z, Li B,
Zhang Y, Li W, Zhang J, Ye J, et al: Inhibition of the m6A reader
IGF2BP2 as a strategy against T-cell acute lymphoblastic leukemia.
Leukemia. 36:2180–2188. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
198
|
Qian L, Ji Z, Mei L and Zhao J: IGF2BP2
promotes lung adenocarcinoma progression by regulating LOX1 and
tumor-associated neutrophils. Immunol Res. 73:162024. View Article : Google Scholar : PubMed/NCBI
|
|
199
|
Du W, Huang Y, Chen X, Deng Y, Sun Y, Yang
H, Shi Q, Wu F, Liu G, Huang H, et al: Discovery of a PROTAC
degrader for METTL3-METTL14 complex. Cell Chem Biol.
31:177–183.e17. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
200
|
Rauff R, Abedeera SM, Schmocker S, Xie J
and Abeysirigunawardena SC: Peptides targeting RNA m6A methylations
influence the viability of cancer cells. ChemMedChem.
18:e2022005492023. View Article : Google Scholar
|
|
201
|
Li Z, Feng Y, Han H, Jiang X, Chen W, Ma
X, Mei Y, Yuan D, Zhang D and Shi J: A stapled peptide inhibitor
targeting the binding interface of N6-Adenosine-Methyltransferase
Subunits METTL3 and METTL14 for cancer therapy. Angew Chem Int Ed
Engl. 63:e2024026112024. View Article : Google Scholar : PubMed/NCBI
|
|
202
|
Huang CS, Zhu YQ, Xu QC, Chen S, Huang Y,
Zhao G, Ni X, Liu B, Zhao W and Yin XY: YTHDF2 promotes
intrahepatic cholangiocarcinoma progression and desensitises
cisplatin treatment by increasing CDKN1B mRNA degradation. Clin
Transl Med. 12:e8482022. View Article : Google Scholar : PubMed/NCBI
|
|
203
|
Hua Z, Gong B and Li Z: Silencing YTHDF2
induces apoptosis of neuroblastoma cells in a cell line-dependent
manner via regulating the expression of DLK1. Mol Neurobiol.
62:8121–8134. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
204
|
Paris J, Morgan M, Campos J, Spencer GJ,
Shmakova A, Ivanova I, Mapperley C, Lawson H, Wotherspoon DA,
Sepulveda C, et al: Targeting the RNA m6A Reader YTHDF2 selectively
compromises cancer stem cells in acute myeloid leukemia. Cell Stem
Cell. 25:137–148.e6. 2019. View Article : Google Scholar
|
|
205
|
Bao Y, Zhai J, Chen H, Wong CC, Liang C,
Ding Y, Huang D, Gou H, Chen D, Pan Y, et al: Targeting m 6 A
reader YTHDF1 augments antitumour immunity and boosts anti-PD-1
efficacy in colorectal cancer. Gut. 72:1497–1509. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
206
|
Wang L, Zhu L, Liang C, Huang X, Liu Z,
Huo J, Zhang Y, Zhang Y, Chen L, Xu H, et al: Targeting
N6-methyladenosine reader YTHDF1 with siRNA boosts antitumor
immunity in NASH-HCC by inhibiting EZH2-IL-6 axis. J Hepatol.
79:1185–1200. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
207
|
Xie LJ, Yang XT, Wang RL, Cheng HP, Li ZY,
Liu L, Mao L, Wang M and Cheng L: Identification of flavin
mononucleotide as a cell-active artificial N6-Methyladenosine RNA
Demethylase. Angew Chem Int Ed Engl. 58:5028–5032. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
208
|
Wilson C, Chen PJ, Miao Z and Liu DR:
Programmable m6A modification of cellular RNAs with a
Cas13-directed methyltransferase. Nat Biotechnol. 38:1431–1440.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
209
|
Li J, Chen Z, Chen F, Xie G, Ling Y, Peng
Y, Lin Y, Luo N, Chiang CM and Wang H: Targeted mRNA demethylation
using an engineered dCas13b-ALKBH5 fusion protein. Nucleic Acids
Res. 48:5684–5694. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
210
|
Sun Y, Gong W and Zhang S: METTL3 promotes
colorectal cancer progression through activating JAK1/STAT3
signaling pathway. Cell Death Dis. 14:7652023. View Article : Google Scholar : PubMed/NCBI
|
|
211
|
Sun X, Wang DO and Wang J: Targeted
manipulation of m6A RNA modification through CRISPR-Cas-based
strategies. Methods. 203:56–61. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
212
|
Nar R, Wu Z, Li Y, Smith A, Zhang Y, Wang
J, Yu F, Gao S, Yu C, Huo Z, et al: Targeting METTL3 protein by
proteolysis-targeting chimeras: A novel therapeutic approach for
acute myeloid leukemia. Genes Dis. 12:1014522024. View Article : Google Scholar
|
|
213
|
Kim S, Hwang I, Kim YK, Kim DS, Choi YJ
and Jeung EB: Treatment of dexamethasone and lenalidomide-resistant
multiple myeloma via RAD51 degradation using PROTAC and synergistic
effects with chemotherapy. J Physiol Pharmacol. 75:2024.
|
|
214
|
Li Y, Zhu S, Chen Y, Ma Q, Kan D, Yu W,
Zhang B, Chen X, Wei W, Shao Y, et al: Post-transcriptional
modification of m6A methylase METTL3 regulates ERK-induced
androgen-deprived treatment resistance prostate cancer. Cell Death
Dis. 14:2892023. View Article : Google Scholar
|
|
215
|
Kamiya T, Oshima M, Koide S,
Nakajima-Takagi Y, Aoyama K, Itokawa N, Yamashita M, Doki N,
Kataoka K and Iwama A: Unraveling the heterogeneity of multiple
myeloma cells by single-cell RNA sequencing analysis. Blood.
140:9939–9940. 2022. View Article : Google Scholar
|
|
216
|
Wang Y, Peng Y, Yang C, Xiong D, Wang Z,
Peng H, Wu X, Xiao X and Liu J: Single-cell sequencing analysis of
multiple myeloma heterogeneity and identification of new
theranostic targets. Cell Death Dis. 15:6722024. View Article : Google Scholar : PubMed/NCBI
|
|
217
|
Liu C, Liang H, Wan AH, Xiao M, Sun L, Yu
Y, Yan S, Deng Y, Liu R, Fang J, et al: Decoding the m6A
epitranscriptomic landscape for biotechnological applications using
a direct RNA sequencing approach. Nat Commun. 16:7982025.
View Article : Google Scholar
|
|
218
|
Jin R, Zou Q and Luo X: From detection to
prediction: Advances in m6A methylation analysis through machine
learning and deep learning with implications in cancer. Int J Mol
Sci. 26:67012025. View Article : Google Scholar : PubMed/NCBI
|
|
219
|
Li Q, Liu J, Guo L, Zhang Y, Chen Y, Liu
H, Cheng H, Deng L, Qiu J, Zhang K, et al: Decoding the interplay
between m6A modification and stress granule stability by live-cell
imaging. Sci Adv. 10:eadp56892024. View Article : Google Scholar
|
|
220
|
Xi JF, Liu BD, Tang GR, Ren ZH, Chen HX,
Lan YL, Yin F, Li Z, Cheng WS, Wang J, et al: m6A modification
regulates cell proliferation via reprogramming the balance between
glycolysis and pentose phosphate pathway. Commun Biol. 8:4962025.
View Article : Google Scholar
|
|
221
|
Zhang L, Wei J, Zou Z and He C: RNA
modification systems as therapeutic targets. Nat Rev Drug Discov.
25:59–78. 2026. View Article : Google Scholar
|