Introduction
Breast cancer (BC) is the most prevalent malignant
tumor among women and the leading cause of cancer-related
mortality. Although medical advancements have significantly
improved clinical outcomes, incidence and mortality rates remain
persistently high (1). This
situation is mainly due to the complexity of BC, which involves
dysregulation across multiple molecular networks, including
accumulation of genetic mutations, abnormal epigenetic
modifications, and alterations in the tumor microenvironment (TME)
(2). Despite increased
understanding of these molecular mechanisms, BC's inherent
complexity and heterogeneity present substantial therapeutic
challenges, particularly limited treatment efficacy and drug
resistance (3). Therefore, deeper
investigation of the molecular mechanisms underlying BC
pathogenesis and identification of novel therapeutic targets
constitute critical research priorities requiring immediate
attention.
MicroRNAs (miRNAs or miRs) are a class of highly
conserved, non-coding RNA molecules of ~22 nucleotides in length.
They mediate mRNA degradation or translational repression by
specifically binding to the 3' untranslated region (3' UTR) of
target mRNAs, thus achieving fine-tuned regulation of gene
expression at the post-transcriptional level (4). Extensive research indicates that
dysregulated miRNA expression is closely linked to the initiation
and progression of various malignant tumors. By modulating
malignant cellular behaviors, such as proliferation, invasion,
migration and drug resistance, miRNAs play a crucial role in cancer
progression. For example, in BC, miR-182-3p can induce apoptosis of
tumor cells by targeting TRF2, significantly inhibiting tumor
growth (5); in lung cancer (LC),
miR-193b-3p downregulates prion protein expression by targeting
PRNP, thus diminishing the proliferation, migration and invasive
capabilities of LC cells (6).
Notably, systematic investigations into miRNA
families have emerged as novel approaches to elucidate core tumor
regulatory networks. Compared with individual miRNAs, members of
the same family often share sequence homology and functional
synergy, thus forming more stable regulatory networks across
various pathways, such as the miR-34 family (7) and the miR-200 family (8). Among these, the miR-200 family has
received considerable attention due to its pivotal role in
regulating key biological processes such as epithelial-mesenchymal
transition (EMT). Dysregulation of this family correlates with
tumor progression and poor prognosis in various cancers, including
hepatocellular carcinoma (9), LC
(10) and ovarian cancer (OC)
(11). This pan-cancer functional
conservation suggests that in-depth investigations of the miR-200
family may not only elucidate the pathogenesis of specific cancers
but also uncover shared molecular mechanisms underlying malignant
tumor development. From a preventive medicine perspective,
identifying and validating these critical regulatory factors, which
exhibit abnormalities at early stages of multiple cancers, holds
significant value for developing universal early screening
biomarkers and molecular intervention strategies targeting
precancerous lesions. In BC, the miR-200 family influences disease
progression by regulating multiple critical pathways, including
cell proliferation, EMT, invasion, migration, cellular stemness and
TME modulation. However, the precise mechanisms of action,
synergistic or antagonistic relationships between family members,
and dynamic changes across BC subtypes and stages remain
incompletely reviewed. The present study aims to systematically
review research progress concerning the miR-200 family in BC,
examine its molecular regulatory mechanisms and biological
functions, and evaluate its potential clinical utility as a
diagnostic biomarker and prognostic indicator for BC. This endeavor
seeks to provide novel theoretical foundations and therapeutic
strategies for managing BC.
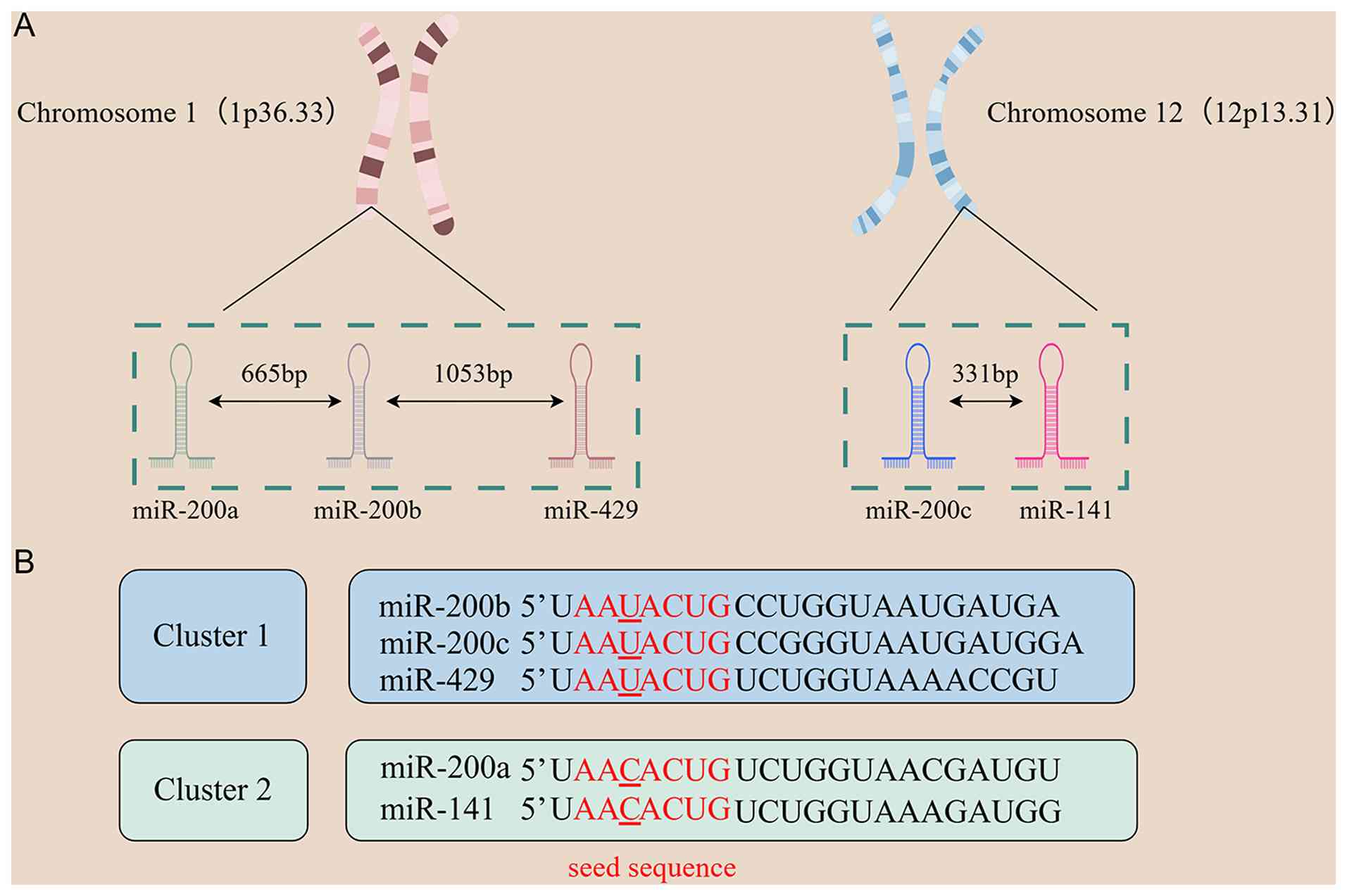
Overview of the miR-200 family
The miR-200 family consists of five members:
miR-200a, miR-200b, miR-200c, miR-429 and miR-141. Based on their
chromosomal locations, this family is divided into two gene
clusters (Fig. 1A): The
miR-200b/miR-200a/miR-429 cluster located at 1p36.33 (intergenic
spacing: 665 bp between miR-200b and miR-200a, 1053 bp between
miR-200a and miR-429) and the miR-141/miR-200c cluster at 12p13.31
(spacing: 331 bp). The miR-200 family is highly conserved across
vertebrates, suggesting a critical regulatory function throughout
evolution (12,13).
The biosynthesis of the miR-200 family follows the
classical miRNA maturation pathway. Its expression is finely
regulated by epigenetic modifications. In epithelial cells,
activating histone modifications (H3K4me3 and H3Ac) are enriched at
the promoters of miR-200 family members, maintaining their high
expression levels; in mesenchymal cells, these family members are
specifically suppressed by distinct epigenetic mechanisms. The
miR-200b/miR-200a/miR-429 cluster is suppressed by PRC2-mediated
H3K27me3, while the miR-200c/miR-141 cluster is silenced through
DNA methylation and H3K9me2 modifications (14,15). Although both clusters exhibit high
expression levels in epithelial cells via H3K4me3 and H3Ac
modifications, they show distinct epigenetic silencing patterns in
mesenchymal cells. This phenomenon reveals the complex regulatory
network of the miR-200 family during evolution and suggests its
essential role in maintaining epithelial phenotype and inhibiting
EMT. Furthermore, oxidative stress reportedly upregulates miR-200
family expression levels, though this response varies among
subclusters. Expression of the miR-200c/miR-141 cluster is
significantly enhanced, whereas the miR-200b/miR-200a/miR-429
cluster displays relatively modest upregulation (16). Additionally, maturation of the
miR-200 family is regulated by multiple transcription factors. For
example, p53 can directly bind to response elements RE1 and
RE2within the miR-200 promoter to activate transcription, thereby
increasing its expression (17).
The ZEB family also serves as a critical regulator. Specifically,
ZEB1/2 bind to the conserved E-box element (CACCTG) in the promoter
regions of miR-200 family members, inhibiting their transcription
and expression. Conversely, miR-200 family members target the 3'UTR
of ZEB1/2, reducing their expression. This reciprocal regulation
establishes a double-negative feedback loop between ZEB1/2 and the
miR-200 family (18,19).
Members of the miR-200 family exhibit high sequence
homology and can be divided into two clusters based on nucleotide
differences in their seed sequences: the miR-200b/200c/429 cluster
(seed sequence: AAUACUG) and the miR-200a/141 cluster (seed
sequence: AACACUG). The seed sequences of these clusters differ by
only one nucleotide (Fig. 1B).
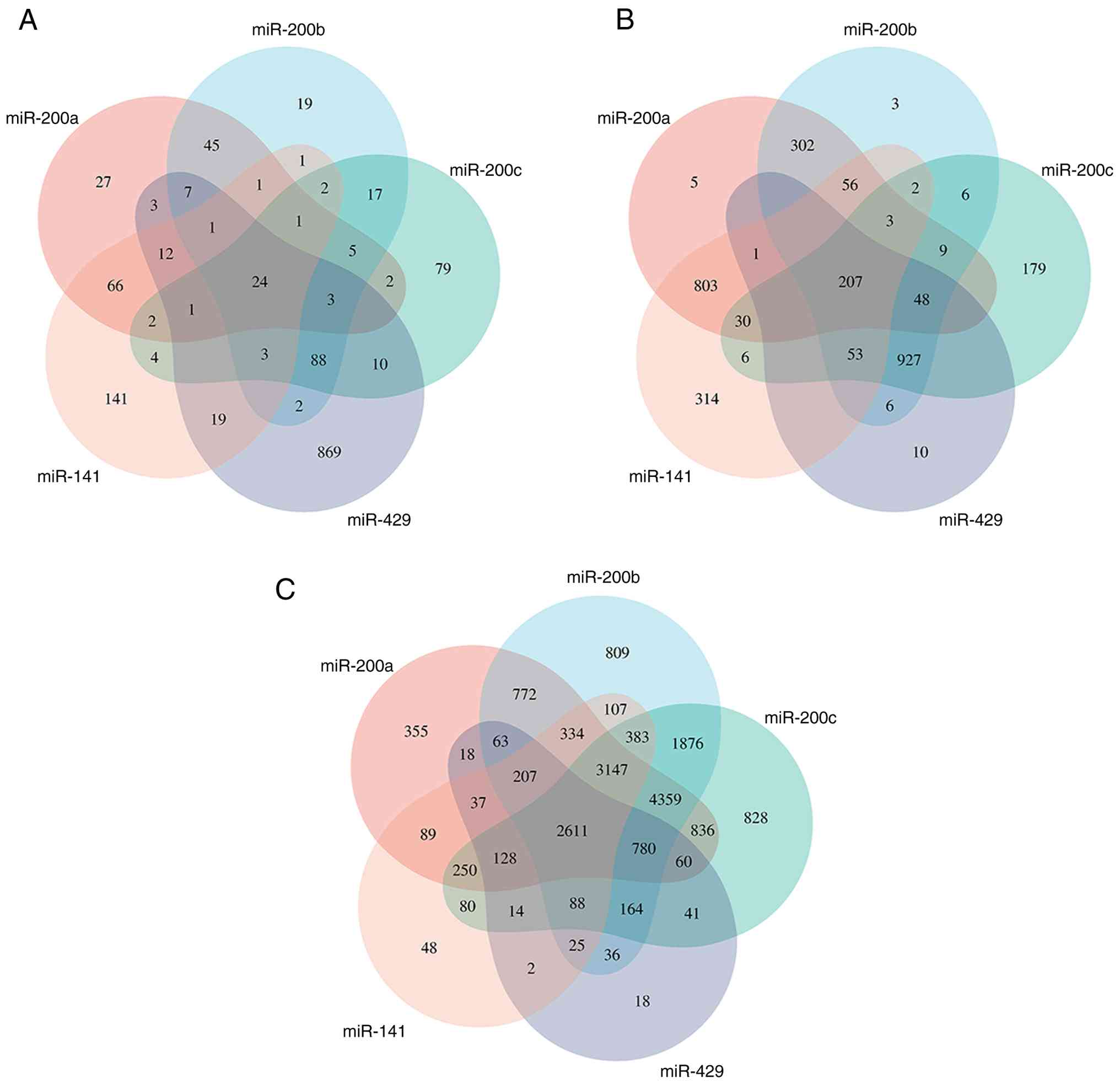
suggesting substantial overlap in their target genes. To validate
this hypothesis, potential target genes of miR-200 family members
were systematically analyzed using bioinformatics approaches.
Results revealed that the miRTarBase (https://mirtarbase.cuhk.edu.cn/) identified 24 shared
target genes (Fig. 2A), miRDB
(https://mirdb.org/) predicted 207 shared target genes
(Fig. 2B) and miRWalk (http://mirwalk.umm.uni-heidelberg.de/)
predicted 2,611 shared target genes (Fig. 2C).
Role of the miR-200 family in BC and its
molecular mechanisms
Studies indicate that the miR-200 family regulates
essential biological behaviors in BC cells, including
proliferation, migration, invasion and apoptosis, through diverse
molecular mechanisms. Currently identified regulatory mechanisms
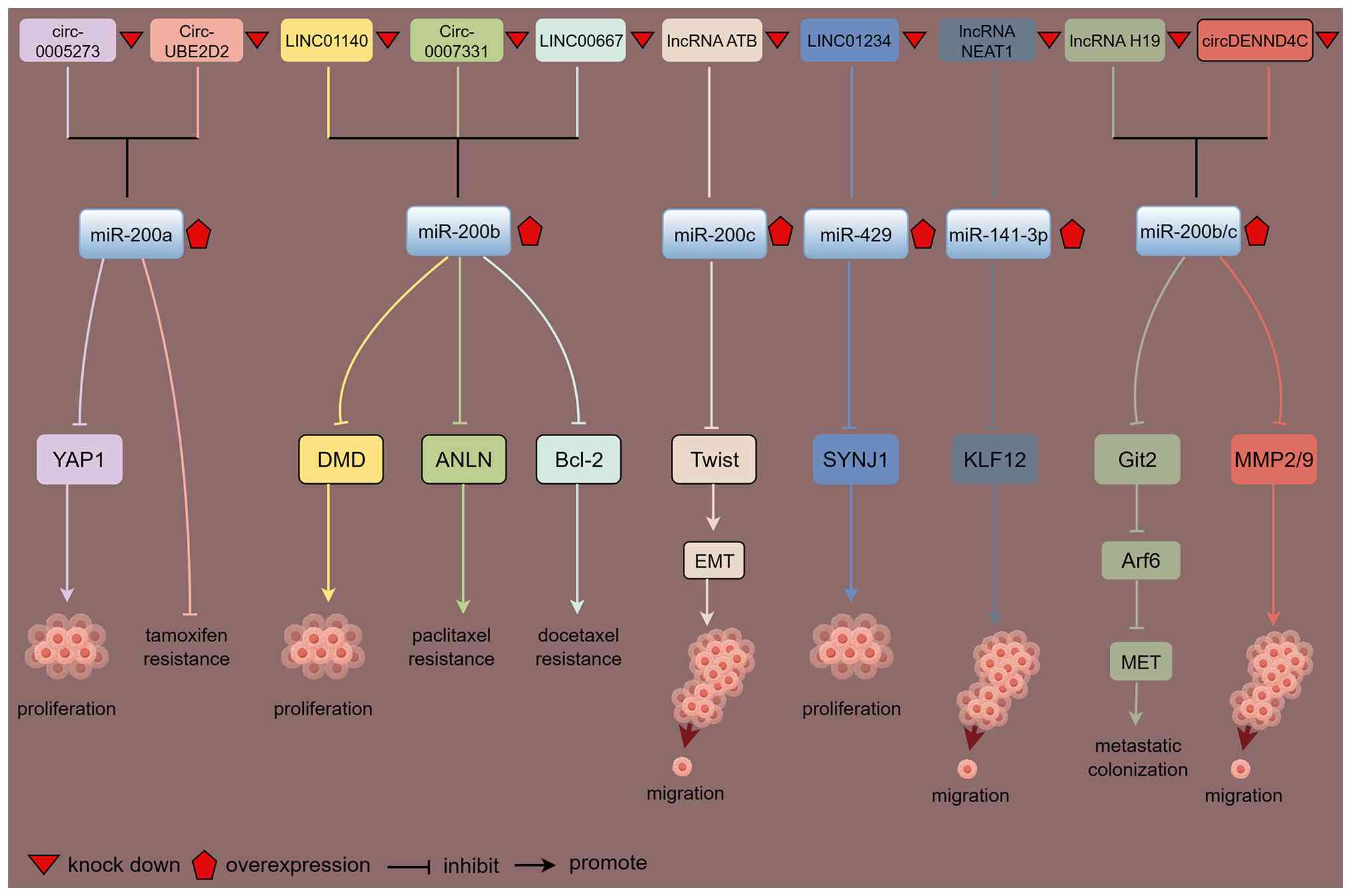
include (Fig. 3): (i) Members of
the miR-200 family directly bind to the 3'UTR regions of downstream
target genes, precisely modulating their expression by mediating
mRNA degradation or translational inhibition (20-24). This direct targeting constitutes
the fundamental mechanism behind their biological functions. (ii)
Multiple long non-coding RNAs (lncRNAs) and circular RNAs act as
'molecular sponges' (25-28) (Fig.
4). These molecules competitively bind miR-200 family members
via miR-200 response elements, preventing miR-200 from interacting
with its target mRNAs and thereby attenuating its inhibitory
effects. Additionally, certain cytokines dynamically regulate
miR-200 expression by interacting with its binding sites (29-32). (iii) The miR-200 family further
modulates downstream signaling pathways, such as PI3K/AKT, MAPK/ERK
and NF-κB, by regulating the expression of target genes, thereby
systematically influencing tumor progression (33-39). Thus, the miR-200 family constructs
a multi-tiered regulatory network in BC, spanning direct gene
regulation to complex signaling pathway modulation, forming a
precise molecular regulatory system. These findings not only
enhance understanding of BC pathogenesis but also provide a
theoretical foundation for developing targeted therapeutic
strategies.
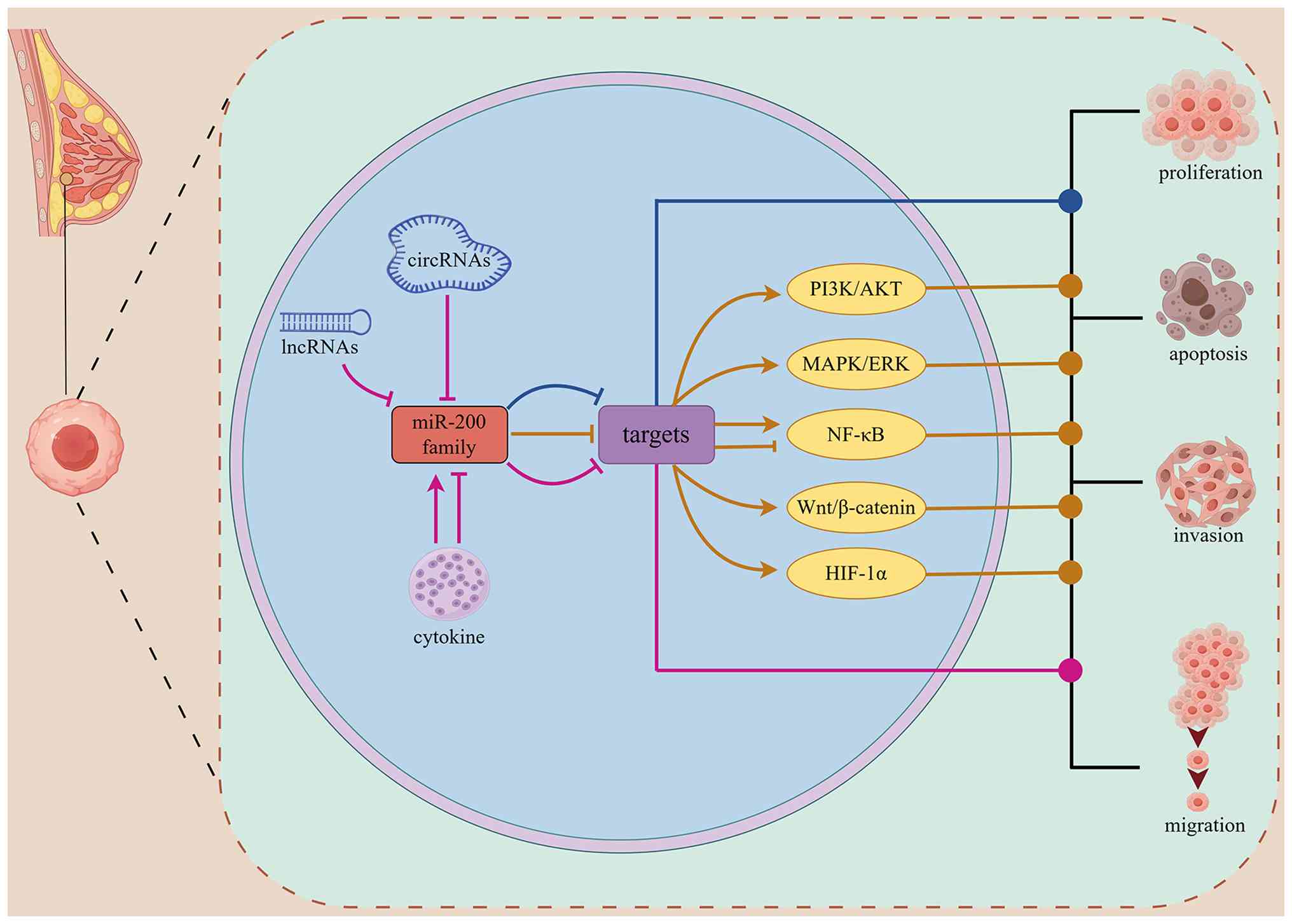 | Figure 3The molecular mechanisms by which the
miR-200 family regulates malignant behavior in breast cancer cells.
Blue pathway: the miR-200 family directly inhibits related target
genes, regulating tumor proliferation, invasion, migration, and
apoptosis. Pink pathway: lncRNAs, circRNAs, and cytokines regulate
tumor progression by upregulating/downregulating miR-200 levels,
thereby inhibiting/promoting target gene expression. Yellow
pathway: the miR-200 family regulates tumor progression by
modulating the expression of its target genes, thereby influencing
the activation status of downstream signaling pathways. lncRNA,
long non-coding RNA; circRNA, circular RNA; miR, microRNA; ⊣,
inhibition; →, promotion. The figure was created by www.figdraw.com. |
Invasion and migration
The primary threat to survival in patients with BC
arises from the highly invasive and migratory characteristics of
tumor cells. These malignant cells break through tissue barriers
via multiple pathological pathways and molecular mechanisms,
causing systemic dissemination and eventually forming lethal
metastatic lesions in distant organs (40). Numerous studies have recently
confirmed that differential expression of the miR-200 family
between epithelial and mesenchymal cells positions it as a key
molecular regulator of BC invasion and migration. Members of this
family significantly influence BC cell invasion and migration
through multi-pathway and multi-target regulatory networks
(Table I), providing critical
insights into the mechanisms underlying BC metastasis.
 | Table IRole of the miR-200 family in BC and
its target genes. |
Table I
Role of the miR-200 family in BC and
its target genes.
| First author/s,
year | miR-200 family
members | Biological
processes | Targets | Roles | (Refs.) |
|---|
| Ahmad et al,
2011; Gregory et al, 2008; Lorenzo-Martin et al,
2019; Lu et al, 2015; Roy et al, 2014; Ye et
al, 2015; Zhang et al, 2019; ZOU et al, 2018 | All members | Invasion and
migration | ZEB1/2 | Inhibit EMT,
thereby suppressing invasion and migration | (43-50) |
| Choi et al,
2016 | | | - | Promotes the
phosphorylation of FAK and AKT, thereby enhancing cell migration
and invasion | (72) |
| Lim et al,
2013 | | Cell stemness | - | Suppressing stem
cell properties | (14) |
| Eades et al,
2011; Kim et al, 2020; Kim et al, 2018 | miR-200a | Invasion and
migration | SIRT1, ELK3,
IMP2/3 | Inhibit EMT,
thereby suppressing invasion and migration | (20-22) |
| Yu et al,
2013 | | | YAP1 | Induce cells to
resist anoikis, thereby promoting metastasis | (61) |
| Tsouko et
al, 2015; Ming et al, 2015 | | | EPHA2, Cx43 | Inhibit cell
invasion and migration | (65,66) |
| Zeng et al,
2019; Yao et al, 2014 | | Proliferation and
apoptosis | TFAM, MET,
EGFR | Inhibit cell
proliferation | (31,80) |
| Wang et al,
2021 | | | YAP1 | Inhibit the cell
cycle, thereby suppressing cell proliferation | (89) |
| Sossey et
al, 2018; Xu et al, 2018 | miR-200b | Invasion and
migration | Suz12,
Kindlin-2 | Inhibit EMT,
thereby suppressing invasion and migration | (23,29) |
| Hong et al,
2016; Li et al, 2014; Yuan et al, 2020 | | | ERM | Disrupts
cytoskeletal remodeling and inhibits invasion | (68-70) |
| Humphries et
al, 2014; Humphries et al, 2017; Zheng et al,
2017 | | | Protein kinase Cα,
ARHGAP18, FUT4 | Inhibit cell
invasion and migration | (74,75,78) |
| Wang et al,
2021 | | Proliferation and
apoptosis | JAZF1 | Promoting
pyroptosis and apoptosis | (34) |
| Peng et al,
2023 | | | DMD | Inhibit cell
proliferation | (83) |
| Yao et al,
2015 | | | Sp1 | Promote apoptosis
and inhibit cell proliferation | (90) |
| Iliopoulos et
al, 2010 | | Cell stemness | Suz12 | Inhibit stem cell
growth | (100) |
| Peng et al,
2021; Li et al, 2018 | miR-200c | Invasion and
migration | c-JUN, Twist | Inhibit EMT,
thereby suppressing invasion and migration | (24,25) |
| Howe et al,
2011 | | | TrkB | Restore anoikis,
thereby inhibiting cell migration | (62) |
| Sigloch et
al, 2015 | | | PAK | Inhibit cell
invasion and migration | (76) |
| Song et al,
2015; Zhang et al, 2019 | | Proliferation and
apoptosis | KRAS, PDE7B | Inhibit cell
proliferation | (35,92) |
| Lin et al,
2024 | | TME | - | Reprogramming CAF
states to promote carcinogenesis | (95) |
| Meng et al,
2020 | | | - | Upregulating PAI-2
promotes M2-type TAM polarization, thereby accelerating
triple-negative BC metastasis | (96) |
| Raue et al,
2022 | | | - | Reduce macrophage
recruitment to tumor sites | (97) |
| Tang et al,
2019; Wu et al, 2017 | | Cell stemness | Jagged1, ZEB1,
Bmi1 | Inhibit cell
stemness | (101,102) |
| Zhang et al,
2020 | miR-429 | Invasion and
migration | CRKL, MMP9 | Inhibiting bone
metastasis in BC | (71) |
| Zhang et al,
2020; Bi et al, 2021 | | Proliferation and
apoptosis | Fibronectin 1,
SYNJ1 | Inhibit cell
proliferation | (36,86) |
| Wang et al,
2015 | | | XIAP | Promote
apoptosis | (85) |
| Li et al,
2023 | | | DLC1 | Promote tumor
proliferation | (87) |
| Zhou et al,
2021; Sun et al, 2020; Li et al, 2017 | miR-141 | Invasion and
migration | High-mobility group
box 1, ANP32E, KLF12 | Inhibit cell
invasion and migration | (28,33,77) |
| Xu et al,
2023 | | Proliferation and
apoptosis | Malat 1 | Inhibit cell
proliferation | (37) |
| Dong et al,
2021 | | | RBMS3 | Inhibit apoptosis
and promote cell proliferation | (88) |
| Tang et al,
2019 | | TME | TCF12 | Inhibiting CAFs
from secreting CXCL12 thereby suppresses BC progression | (32) |
| Zhou et al,
2017 | miR-200b/c | Invasion and
migration | Git2 | Inhibit MET to
suppress the colonization of metastatic lesions | (26) |
| Ren et al,
2019 | | | MMP2/9 | Inhibit cell
invasion | (27) |
| Sun et al,
2018 | | TME | IKKβ | Weakening the
ability of CAFs to promote epithelial-mesenchymal transition and
invasion in tumor cells | (38) |
| Jin et al,
2017 | miR-200c/141 | Invasion and
Migration | - | Increased SerpinB2
promotes cell migration | (73) |
| Liu et al,
2018 | | Cell stemness | HIPK1 | Overexpression of
miR-200c/141 promotes the formation of epithelial-like
ALDH+ BC stem cells, while its underexpression favors
maintaining the characteristics of mesenchymal-like
CD24−CD44+ BC stem cells | (39) |
| Li et al,
2017 |
miR-200b-5p/429-5p | Proliferation and
apoptosis | LIMK1 | Inhibit the cell
cycle | (93) |
| Tang et al,
2019 | miR-200s | TME | DNMT3B | Inhibit CAF
activation | (32) |
EMT
Under normal physiological conditions, epithelial
cells maintain tissue integrity through tight junctions, adherens
junctions and other cellular structures. These features provide
strong adhesion and limit migratory capability. Mesenchymal cells,
by contrast, show reduced cell-cell adhesion and reorganized
cytoskeletal structures, resulting in increased motility and
invasiveness. EMT refers to the transition from epithelial to
mesenchymal phenotypes, characterized by the downregulation of
epithelial markers (for example, E-cadherin) and the upregulation
of mesenchymal markers (for example, N-cadherin and vimentin). This
transformation allows tumor cells to gain migratory and invasive
properties, significantly enhancing their metastatic potential
(41). EMT is primarily mediated
by core transcription factors, such as ZEB1, ZEB2, Snail and Twist,
which repress epithelial gene expression by binding to E-box motifs
in their promoter regions (42).
Notably, a precise double-negative feedback loop exists between the
miR-200 family and the ZEB family (ZEB1/ZEB2). On one hand, miR-200
family members directly target and suppress the expression of ZEB1
and ZEB2 (43-50), blocking EMT and inhibiting the
invasive and migratory capacities of BC cells. Conversely, the ZEB
family transcriptionally suppresses miR-200 family expression
(51-53), thereby promoting EMT and
increasing tumor metastatic potential. This reciprocal regulatory
mechanism establishes a dynamic equilibrium that critically governs
BC invasion and metastasis. Notably, this classical miR-200/ZEB
feedback loop has also been identified as a central mechanism
driving invasion and metastasis in various epithelial tumors,
including OC (54) and colorectal
cancer (CRC) (55), highlighting
its conserved role in maintaining epithelial phenotypes and
suppressing malignant transformation. In addition to the ZEB
family, miR-200 family members further regulate tumor metastasis by
targeting other drivers of EMT. For instance, the lncRNA ATB acts
as a molecular sponge to competitively bind miR-200c, thereby
alleviating its inhibitory effect on Twist. This interaction
promotes EMT and enhances the invasive and migratory capabilities
of BC cells (25). SIRT1 induces
the epigenetic silencing of E-cadherin in BC (56). Eades et al (20) demonstrated that miR-200a targets
and suppresses SIRT1 expression, thereby inhibiting EMT and
reducing tumor invasion.
In addition, the miR-200 family regulates tumor EMT
through several other mechanisms, thereby influencing tumor
invasion and migration. Studies indicate that ELK3 acts
synergistically with ZEB1 to suppress E-cadherin expression
(57). Kim et al (21) demonstrated that miR-200a
specifically binds to the 3'UTR of ELK3 mRNA, significantly
reducing ELK3 protein levels, restoring E-cadherin expression, and
suppressing BC cell invasion and migration. Kim et al
(22) further showed that
miR-200a targets and downregulates IMP2 and IMP3 expression,
thereby blocking EMT and inhibiting cell migration and invasion.
Another study revealed that the p38 MAPK subtype p38γ negatively
regulates miR-200b expression by inhibiting its transcriptional
activator GATA3. Conversely, miR-200b overexpression reduces Suz12
expression, reversing p38γ-induced EMT process (29). Additionally, miR-200b suppresses
EMT by directly inhibiting Kindlin-2 (23). Peng et al (24) demonstrated that miR-200c blocks
EMT and inhibits migration by directly targeting c-JUN. Tumor
metastasis is a dynamic, reversible process. Tumor cells entering
circulation must activate EMT to enhance invasive capacity;
however, during implantation in distant organs, they must revert to
epithelial characteristics via mesenchymal-epithelial transition
(MET) to establish metastatic colonies (58). Zhou et al (26) found that in metastatic BC (MBC)
lesions, the lncRNA H19 sequesters miR-200b/c, thereby relieving
suppression of its target gene Git2. This results in Arf6
inactivation and maintains E-cadherin expression, thus inhibiting
EMT and promoting MET, facilitating the colonization of metastatic
sites (26). Collectively, these
studies indicate that the miR-200 family influences EMT via a
multi-level regulatory network, playing a pivotal role in BC
metastasis.
Anoikis
Anoikis is a form of programmed cell death triggered
when cells detach from the extracellular matrix or loss
intercellular adhesion. It effectively limits the survival and
migration of detached tumor cells. However, malignant tumor cells
can acquire resistance to anoikis, promoting metastasis (59). The role of the miR-200 family in
this process appears complex. It has been reported that the loss of
YAP1 function can protect cells from anoikis (60). Yu et al (61) showed that miR-200a promotes
metastasis by inhibiting YAP1, thereby inducing tumor cell
resistance to anoikis. By contrast, Howe et al (62) reported that miR-200c restores
apoptosis sensitivity by downregulating TrkB, thus suppressing cell
migration. This functional discrepancy suggests that miR-200 family
members may exhibit distinct roles depending on specific
microenvironmental contexts. The underlying mechanisms require
further exploration. Additionally, miR-200-mediated regulation of
anoikis through the Orphanage pathway is not exclusive to BC. In
endometrial carcinoma, miR-200 family members (miR-141 and
miR-200b/200c/429) directly suppress Sestrin proteins (SESN2 and
SESN3), weakening resistance to anoikis following loss of
anchorage. This observation suggests that the miR-200/Sestrin axis
may also play a critical role in endometrial carcinoma metastasis
(63).
Genes associated with invasion and
migration
The miR-200 family also exerts its biological
effects by regulating multiple genes involved in invasion and
migration. Studies indicate that EPHA2 binds its ligand, Ephrin-A1,
resulting in EPHA2 degradation and suppression of tumor invasion
and migration. In the absence of Ephrin-A1, EPHA2 accumulates
excessively, promoting invasion and migration (64). Tsouko et al (65) reported that Ephrin-A1 expression
is nearly absent in triple-negative BC (TNBC), leading to elevated
EPHA2 expression. miR-200a directly targets EPHA2, reducing its
expression, thereby increasing AMPK activity and ultimately
suppressing tumor cell invasion and migration (65). Additionally, miR-200a targets and
downregulates connexin 43 (cx43) expression, thus inhibiting
cellular metastasis (66). The
ezrin-radixin-moesin (ERM) protein family is critical for
maintaining cytoskeletal structure, facilitating cell movement, and
contributing to BC invasion and migration (67). Studies have demonstrated that
miR-200b downregulates ERM family proteins (Ezrin, Radixin and
Moesin), disrupting cytoskeletal remodeling and inhibiting invasion
(68-70). Additionally, Ren et al
(27) showed that knockout of
circDENND4C upregulates miR-200b/c, which suppresses glycolysis and
MMP2/9 expression, consequently decreasing cell invasion. Other
studies reported that miR-141-3p downregulates high-mobility group
box 1 protein expression, suppressing hypoxia-induced HIF-1α
pathway activity and cell migration (33). miR-429 can reduce BC bone
metastasis by targeting and downregulating CRKL and MMP9 expression
(71). However, a comprehensive
study on the miR-200 family reported conflicting results. Stable
overexpression of the miR-200b/200a/429 or miR-141/200c clusters
inhibited MDA-MB-231 cell proliferation but significantly enhanced
migration and invasion. Compared with miR-200b/200a/429, the
miR-141/200c cluster showed stronger suppression of cell
proliferation and greater promotion of migration and invasion.
Further analysis revealed that overexpressing either
miR-200b/200a/429 or miR-141/200c increased phosphorylation of FAK
and AKT, thus enhancing cell migration and invasion. Compared with
miR-200b/200a/429-overexpressing cells, those expressing
miR-141/200c secreted higher levels of vascular endothelial growth
factor A and exhibited increased integrin-αV expression, further
augmenting migration and invasion (72). Additionally, Jin et al
(73) suggested that the
miR-200c/141 cluster promotes BC metastasis by upregulating
SerpinB2 (PAI-2); however, the precise molecular mechanisms
involved require further clarification. These discrepancies
highlight the functional heterogeneity of the miR-200 family across
different BC subtypes and TME.
Other mechanisms
Besides the aforementioned pathways, the miR-200
family can inhibit cell invasion and migration by targeting
molecules such as protein kinase Cα (74), ARHGAP18 (75), PKA subunits (PRKAR1A and PRKACB)
(76), ANP32E (77) and FUT4 (78). Furthermore, lncRNA NEAT1 acts as a
sponge for miR-141-3p. Upon interference with NEAT1 expression,
miR-141-3p levels significantly increase, thereby reducing KLF12
expression, inhibiting cell invasion and migration, and decreasing
chemotherapy resistance (28).
Conversely, FOXP3, as an endogenous tumor suppressor, inhibits BC
metastasis by promoting the expression of miR-200c and miR-141
(30). Liu et al (79) notably reported that BC cells
specifically package miR-200c into exosomes through a
Rab1A-mediated sorting mechanism. The cells then actively secrete
these exosomes, thereby evading the tumor suppressive effects of
miR-200c, promoting metastasis, and reshaping the immune
microenvironment. This finding reveals a novel mechanism underlying
BC metastasis and suggests new therapeutic opportunities (79).
In summary, the miR-200 family plays a complex and
critical role in BC invasion and metastasis through multiple
pathways, including the regulation of EMT, anoikis, and invasion-
and migration-related genes. Although most studies support its
tumor-suppressive role, some findings suggest pro-metastatic
effects. These discrepancies indicate that future research should
clarify its regulatory mechanisms across BC subtypes and
microenvironments. More broadly, pathways regulated by the miR-200
family, such as EMT and anoikis, represent core biological
processes that enable cells to acquire migratory capabilities.
Therefore, deepening the understanding of miR-200 family-regulated
networks in cancer not only provides potential therapeutic targets
for advanced metastatic disease but also suggests that monitoring
these pathway activities in precancerous lesions or early-stage
cancers may enable earlier intervention by predicting invasive
potential.
Proliferation and apoptosis
The miR-200 family also plays an essential role in
regulating BC cell proliferation and apoptosis. Its members
influence tumor biological behaviors by targeting diverse signaling
molecules (Table I). Current
studies suggest that miR-200a, miR-200b and miR-200c predominantly
function as tumor suppressors in BC by inhibiting cell
proliferation and promoting apoptosis. Yao et al (80) demonstrated that miR-200a directly
targets TFAM, inhibiting its protein expression and thereby
consequently BC cell proliferation. Additionally, FEN1, a critical
DNA metabolism regulator involved in DNA replication, damage
repair, and telomere maintenance, is closely associated with tumor
proliferation (81,82). Studies indicated that FEN1 forms a
FEN1/proliferating cell nuclear antigen (PCNA)/DNA
methyltransferase 3a (DNMT3a) complex with PCNA and DNMT3a,
suppressing miR-200a expression via DNMT3a-mediated methylation.
FEN1 knockout significantly upregulated miR-200a, which then
targeted and downregulated hepatocyte growth factor (MET) and EGFR,
thereby inhibiting the PI3K/AKT and MAPK/ERK signaling pathways and
ultimately reducing BC cell proliferation (31). Chrysophanol activated the NF-κB
signaling pathway by upregulating miR-200b and downregulating
JAZF1, inducing pyroptosis and apoptosis, thus significantly
reducing BC cell viability (34).
Additionally, LINC01140 promotes BC progression by competitively
inhibiting miR-200b through sponge adsorption. Knockdown of
LINC01140 increased miR-200b levels, reducing downstream DMD
expression and suppressing cell proliferation (83). miR-200c inhibits tumor
proliferation both in vitro and in vivo by directly
targeting KRAS and subsequently suppressing the AKT and ERK
signaling pathways (35).
Notably, Jones et al (84)
showed that miR-200c expression is downregulated in Claudin-low BC,
a rare subtype of TNBC, and restoring its expression significantly
inhibited tumor cell proliferation, colony formation and tumor
growth in vivo (84).
It is noteworthy that miR-429 exhibits a dual role
in BC. Zhang et al (36)
reported that miR-429 suppresses cell proliferation by targeting
fibronectin 1, thereby inhibiting the Wnt/β-catenin signaling
pathway. Other studies demonstrated that vitamin E δ-tocotrienol
exerts anticancer effects both in vitro and in vivo
by upregulating miR-429, suppressing its target gene XIAP, and
inducing apoptosis (85).
Additionally, Bi et al (86) showed that LINC01234 is highly
expressed in TNBC, and its knockdown upregulates miR-429, decreases
SYNJ1 expression, inhibits cell proliferation, and promotes
apoptosis. By contrast, Li et al (87) found that miR-429 is overexpressed
in the TNBC cell line MDA-MB-468 and promotes tumor proliferation
by degrading the tumor suppressor factor DLC1. The regulatory role
of miR-141 is also context dependent in BC. Xu et al
(37) reported that MALAT1 is
overexpressed as an oncogene in TNBC and is further induced under
hypoxic conditions. Overexpression of miR-141-3p suppresses the
post-transcriptional MALAT1/HIF-1α signaling pathway, thereby
inhibiting autophagy initiation and cell proliferation (37). However, another study identified
that miR-141-3p promotes proliferation and inhibits apoptosis by
targeting and downregulating RBMS3 (88).
In addition, members of the miR-200 family influence
BC cell proliferation and by regulating cell cycle progression.
Dysregulation of the Hippo signaling pathway has been reported to
promote proliferation and inhibit apoptosis, with YAP1 acting as a
key downstream effector. Wang et al (89) demonstrated that miR-200a directly
targets YAP1, induces G0/G1 cell cycle arrest, and reverses the
pro-proliferative effects of circ-0005273. Furthermore, Yao et
al (90) found that miR-200b
targets Sp1, leading to G2/M phase arrest, a reduction in S-phase
cells, inhibition of proliferation, and induction of apoptosis.
Elevated intracellular cAMP levels are known to induce cell cycle
arrest and apoptosis (91). Zhang
et al (92) demonstrated
that miR-200c downregulates PDE7B, increases intracellular cAMP
levels, elevates the proportion of cells in the G1 phase, and
suppresses cell proliferation. Moreover, Li et al (93) demonstrated that miR-200b-5p and
miR-429-5p downregulate LIMK1 and its substrate CFL1, inhibit
cyclin D1/CDK4/CDK6 and cyclin E1/CDK2, and ultimately impair cell
cycle progression. Nevertheless, studies focusing on miR-200
family-mediated cell cycle regulation in BC remain limited. Whether
this family regulates the cell cycle through additional molecular
mechanisms to promote BC progression requires further
investigation.
TME
It has been demonstrated that the TME, composed of
cellular and non-cellular components, plays a pivotal regulatory
role in BC progression (94).
Among these components, the miR-200 family acts as an essential
regulator, influencing TME remodeling through multiple targets and
pathways (Table I). Sun et
al (38) showed significantly
reduced miR-200b/c expression in cancer-associated fibroblasts
(CAFs), closely linked to their pro-tumorigenic properties.
Restoration of miR-200b/c expression directly targets and decreases
IKKβ, inhibiting NF-κB signaling. This process subsequently reduces
expression and secretion of downstream effector PAI-1, diminishing
the ability of CAFs to facilitate tumor cell EMT and invasion
(38). Tang et al
(32) revealed a
TGF-β1-miR-200-DNMT3B regulatory axis that promotes tumor
progression: TGF-β1 suppresses miR-200 (miR-200a/b/c and miR-141)
expression in CAFs, alleviating their post-transcriptional
inhibition of DNMT3B. Elevated DNMT3B promotes methylation of CpG
islands in the miR-200 promoters, leading to stable epigenetic
silencing. This self-sustaining positive feedback loop enables CAFs
to maintain a persistently activated phenotype. Additionally,
reduced miR-141 expression elevates its target gene TCF12,
promoting CXCL12 secretion by CAFs. CXCL12 subsequently activates
the c-Myc/Cyclin D1 pathway in tumor cells, enhancing BC
progression (32). Conversely,
Lin et al (95) reported
that oxidative stress induces DNA demethylation of the miR-200c
promoter in CAFs, significantly upregulating its expression. Highly
expressed miR-200c reprograms CAFs toward a MET state via the
miR-205-COMMD1-NFκB-HIF axis, inducing a senescent phenotype. These
reprogrammed CAFs enhance cancer cell proliferation, apoptosis
resistance and immune suppression through paracrine secretion of
factors such as TGFB2, CCL5, PDGFA/B and lactate, ultimately
promoting tumor progression (95).
In another crucial TME component, tumor-associated
macrophages (TAMs), the miR-200 family similarly exerts significant
regulatory effects. TAMs, abundant immune cells within the TME, can
be classified based on activation status into antitumor M1 and
pro-tumor M2 subtypes. Meng et al (96) demonstrated that miR-200c promotes
M2-type TAM polarization by upregulating PAI-2, thereby
accelerating TNBC metastasis; however, the detailed molecular
mechanisms remain to be clarified. Raue et al (97) described another aspect of miR-200c
function: Apoptotic tumor cells release miR-200c, which macrophages
then internalize. Uptake of miR-200c suppresses migration-related
gene expression in macrophages, reducing their recruitment to tumor
sites (97). These findings
illustrate the complex mechanisms by which the miR-200 family
regulates BC progression by modulating various cellular components
of the TME.
Cell stemness
A major cause of BC treatment failure is the
persistence of cancer stem cells (CSCs). CSCs drive tumor
initiation, progression and resistance to treatment. Due to their
unique self-renewal capacity and multipotent differentiation
potential, CSCs not only directly contribute to tumor progression
and metastasis but also mediate chemotherapy resistance through
various molecular mechanisms (98). Previous studies demonstrated that
the miR-200 family, as key epigenetic regulators, plays a crucial
role in controlling the stemness and phenotypic plasticity of BC
stem cells (BCSCs) (Table I). For
example, Hsu et al (99)
found that inhibition of the miRNA processing enzyme Dicer
significantly reduces miR-200b expression, whereas transfection
with miR-200b mimics decreases the expression of
stemness-associated transcription factors (Oct-4, Nanog, SOX-2 and
KLF4), thus inhibiting CSC properties. Moreover, miR-200b
suppresses mammary spheroid formation and maintenance by directly
targeting Suz12, ultimately impairing CSC proliferation (100). Similarly, Tang et al
(101) confirmed that miR-200c
expression is generally reduced in BC cells and BCSCs.
Overexpression of miR-200c directly targets and suppresses Jagged1,
ZEB1 and Bmi1, thereby inhibiting tumor-sphere formation in
HER2+ BC cells and decreasing the proportion of
CD44+CD24− cells. Importantly,
miR-200c-mediated inhibition of stemness significantly restores
sensitivity to trastuzumab, suppressing tumor growth and metastasis
(101). Additional studies
indicate that miR-200c inhibits BCSC proliferation and
tumorigenicity by targeting BMI1 (102). Notably, BCSCs exist in two
plastic states: epithelial-like (ALDH+) and
mesenchymal-like (CD24−CD44+). These states
interconvert and possess distinct biological behaviors:
Epithelial-like BCSCs exhibit higher proliferation potential,
whereas mesenchymal-like BCSCs show stronger migratory and invasive
abilities (103). Liu et
al (39) clarified the
molecular basis of this phenomenon, demonstrating that miR-200c/141
inhibits HIPK1 expression by directly binding its 3'UTR. HIPK1
downregulation reduces phosphorylation at β-catenin Ser552,
suppressing Wnt/β-catenin signaling. This regulatory axis
determines BCSC state transitions: High miR-200c/141 expression
promotes epithelial-like (ALDH+) BCSCs, while low
expression favors mesenchymal-like
(CD24−CD44+) BCSCs. These results provide a
theoretical basis for novel therapeutic strategies targeting tumor
growth and metastasis, suggesting the miR-200c/141-HIPK1-β-catenin
axis as a potential therapeutic target (39). Additionally, Lim et al
(14) reported that immortalized
human mammary epithelial cells can spontaneously transition from a
non-stem cell to a stem-like phenotype, which strongly correlates
with the absence of miR-200 family expression. Restoration of
miR-200 family expression reverses stem-like properties and
promotes differentiation toward an epithelial phenotype,
highlighting the crucial role of that the miR-200 family in stem
cell and non-stem cell state transitions (14).
Biomarker
Currently, BC diagnosis, staging and prognosis
primarily depend on tissue biopsy. However, this method is
invasive, causes significant patient discomfort, and is difficult
to repeat. Moreover, tissue biopsy has limited effectiveness in
early diagnosis and dynamic monitoring, often delaying detection
until advanced stages. Therefore, developing non-invasive,
effective biomarkers is crucial for early screening and precise
management of BC. Although CA125 and CA153 are widely used clinical
markers for BC monitoring, they face substantial limitations,
including low sensitivity, poor specificity and limited value in
early diagnosis, prognosis assessment and treatment guidance
(104,105). By contrast, miRNAs possess
unique advantages such as high stability, ease of quantitative
detection, and abundant presence in bodily fluids, enabling
non-invasive and dynamic monitoring (106). Previous studies (107,108) demonstrate that specific miRNAs,
particularly members of the miR-200 family, show promise as novel
biomarkers for early BC detection, offering opportunities to
overcome existing diagnostic limitations.
Diagnostic value
Numerous studies highlight the significant
diagnostic and classification roles of the miR-200 family in BC
(Table II). However, expression
patterns vary across research contexts. Xu et al (107) quantified expression levels of
all the miR-200 family members in 99 pairs of BC tissues and
adjacent normal tissues using quantitative PCR (qPCR). They
observed significant downregulation of all miR-200 members in
cancer tissues and found that miR-200 could effectively distinguish
patients with and without lymph node metastasis (107). These findings support the
miR-200 family as potential diagnostic biomarkers for BC.
Inflammatory BC (IBC) is a rare yet highly aggressive subtype often
misdiagnosed due to the lack of reliable molecular biomarkers.
Fahim et al (108)
analyzed miRNA expression in primary tumors of patients with IBC
and non-IBC using miRNA PCR arrays. They found significant
downregulation of miR-200b-3p, miR-200c-3p and miR-203a-3p, with
significant upregulation of miR-181b-5p in IBC. Receiver Operating
Characteristic (ROC) curves demonstrated that each miRNA
individually distinguished patients with IBC from patients with
non-IBC. Notably, combining miR-181b-5p, miR-200b-3p and
miR-200c-3p significantly improved diagnostic accuracy [area under
the curve (AUC)=0.897], indicating potential molecular diagnostic
markers for this challenging subtype (108). Qiao et al (109) reported significantly decreased
expression of miR-200c in serum exosomes from patients with BC. ROC
analysis indicated that miR-200c effectively discriminated patients
with BC from healthy controls (AUC=0.854). Combining miR-200c with
CEA, CA125 and CA153 increased diagnostic accuracy to an AUC of
0.914 (109). However,
discrepancies exist regarding miR-200 expression patterns in
circulation. Khalil et al (110) used qPCR to analyze blood samples
from 54 patients with non-metastatic BC (NMBC), 46 patients with
MBC and 50 healthy controls. They observed significant
overexpression of miR-200a in the serum of patients with BC, with
higher levels in MBC compared with patients with NMBC. Further
analysis showed that miR-200a distinguished between MBC and NMBC,
achieving an AUC of 0.708 (110). Similarly, Papadaki et al
(111) found significantly
higher plasma expression of miR-200b and miR-200c in patients with
MBC compared with early-stage patients. The diagnostic AUC for
miR-200b alone was 0.720, improving to 0.797 when combined with
miR-21, miR-190 and miR-200c, suggesting that circulating miRNAs
reflect distinct tumor biological characteristics (111). Additionally, Braicu et al
(112) analyzed The Cancer
Genome Atlas data and found significant overexpression of miR-200b
was in TNBC and DPBC (ER+, PR+ and
HER2−) compared with normal breast tissue, consistent
with clinical plasma measurements. miR-200b effectively
discriminated patients with BC from healthy controls, achieving an
AUC of 0.8772. Notably, the present study independently validated
miR-200b expression consistency between tumor tissues and plasma
samples (21 TNBC, 47 DPBC and 19 normal samples), providing robust
evidence supporting miR-200b as a reliable liquid biopsy biomarker
(112).
 | Table IIPotential utility of miR-200 family
in cancer diagnosis. |
Table II
Potential utility of miR-200 family
in cancer diagnosis.
| First author/s,
year | miR-200 family
members | Cases | Sample type | Testing
technology | Expression | AUC | (Refs.) |
|---|
| Xu et al,
2016 | All members | 99 BC vs. 99
NC | Tissue | qPCR | Downward | - | (107) |
| Fahim et al,
2020 |
miR-200b-3p/200c-3p | 17 IBC vs.18
non-IBC | Tissue | qPCR | Downward | 0.713/0.743 | (108) |
| Qiao et al,
2024 | miR-200c | 51 BC vs. 47
NC | Serum exosomal | qPCR | Downward | 0.854 | (109) |
| Khalil et
al, 2024 | miR-200a | 46 MBC vs. 54 NMBC
vs. 50 NC | Serum | qPCR | Highly expressed in
BC, and MBC is higher than NMBC | Distinguishing
between MBC and NMBC: 0.708 | (110) |
| Papadaki et
al, 2019 | miR-200b | 110 MBC vs. 133
Early BC | Plasma | qPCR | Upward | 0.72 | (111) |
| Braicu et
al, 2018 | | 48 BC vs. 28
NC | Plasma | qPCR | Upward | 0.8772 | (112) |
Prognosis assessment value
The miR-200 family also exhibits significant
prognostic value in BC (Table
III). Yao et al (90)
analyzed miR-200b expression levels in 278 paired BC and adjacent
normal tissues by qPCR, showing significant downregulation in BC
tissues. Further analysis indicated that low miR-200b expression
was closely associated with aggressive clinicopathological features
(advanced TNM stage, ER-negative status and HER-2-positive status)
and predicted poorer clinical outcomes (90). Similarly, Ye et al
(113) found reduced miR-200b
expression in BC tissues. Patients with lower miR-200b levels
exhibited significantly decreased overall survival (OS) and
disease-free survival (DFS) compared with those with higher
expression (113). A total of
~70% of BCs are estrogen receptor-positive (luminal subtype), and
endocrine therapy remains one of the most effective adjuvant
treatments. However, recurrence occurs in up to 40% of patients.
Amorim et al (114)
demonstrated that low expression of miR-200b-3p correlated with
shorter endocrine resistance-free survival and DFS. Combining
miR-200b-3p with miR-182-5p further improved prognostic accuracy
(114). However, when examining
circulating miRNAs, the clinical significance of miR-200 family
expression patterns differs. Two studies by Fischer et al
(115,116) revealed that high baseline plasma
miR-200 expression in MBC correlated positively with circulating
tumor cell (CTC) positivity and predicted shorter OS and
progression-free survival (PFS). Moreover, high baseline miR-429
levels specifically indicated an increased risk of early
recurrence. Notably, dynamic miR-200 expression changes during
treatment also held prognostic importance: Effective therapy
significantly reduced miR-200a/b/141 levels, which rebounded upon
disease progression. Patients maintaining high miR-200 expression
after treatment experienced a 3-5-fold increase in early recurrence
risk. These findings suggest that miR-200 family members could
serve as liquid biopsy biomarkers for clinical prognosis and
treatment monitoring, particularly when combined with CTC detection
for identifying high-risk patients (115,116). Similarly, Madhavan et al
(117) performed a multicohort
study demonstrating that elevated plasma miR-200 levels correlated
with poorer OS and PFS, surpassing the predictive power of CTC
detection alone. Notably, high miR-200a/b/c expression predicted
metastasis risk up to two years before clinical diagnosis (AUC=
0.82), highlighting its potential as an early surveillance
biomarker (117). MBC typically
carries a poor prognosis and primarily relies on chemotherapy;
however, drug resistance frequently leads to treatment failure and
disease progression. Shao et al (118) observed significantly higher
plasma miR-200a levels were in patients with chemotherapy-resistant
MBC compared with sensitive patients, with high expression
correlating with advanced disease stage. ROC analysis confirmed the
high accuracy (AUC=0.881) of plasma miR-200a in differentiating
chemotherapy-resistant from sensitive cases, indicating its
potential as a predictive biomarker for chemotherapy resistance in
MBC (118). Navarro-Manzano
et al (119) measured
plasma miR-200c-3p levels in 28 healthy women, 42 patients with MBC
and 171 patients with locally advanced BC (LABC) using qPCR. They
found significantly elevated miR-200c-3p levels in LABC and MBC
groups compared with healthy controls, with high expression
associated with reduced OS (119).
 | Table IIIPotential utility of miR-200 family
in cancer prognostic assessment. |
Table III
Potential utility of miR-200 family
in cancer prognostic assessment.
| First author/s,
year | miR-200 family
members | Sample type | Expression |
Characteristics | (Refs.) |
|---|
| Yao et al,
2015 | miR-200b | Tissue | Downward | Low expression
indicates greater invasiveness | (90) |
| Ye et al,
2014 | | Tissue | Downward | Low expression is
associated with shorter OS and DFS | (113) |
| Amorim et
al, 2019 | | Tissue | Downward | Low expression is
associated with shorter ERFS and DFS | (114) |
| Fischer et
al, 2022; Fischer et al, 2022 | All members | Plasma | Upward | High expression is
associated with shorter OS and PFS; the risk of recurrence is
significantly increased by 3-5 times if miR-200a/b/141 remains
highly expressed after treatment | (115,116) |
| Madhavan et
al, 2016 | | Plasma | Upward | High expression is
associated with shorter OS and PFS | (117) |
| Shao et al,
2019 | miR-200a | Plasma | Upward | High expression is
associated with chemotherapy resistance | (118) |
| Navarro-Manzano
et al, 2022 | miR-200c-3p | Plasma | Upward | High expression is
associated with shorter OS | (119) |
Discussion on miR-200 expression
differences between tissues and circulation
Although the miR-200 family shows significant
diagnostic and prognostic value in BC tissues and circulating
samples, noteworthy differences in expression patterns exist
between these two contexts. Most tissue-based studies indicate that
miR-200 members are downregulated in BC tissues, exerting
tumor-suppressive effects (90,107,108,113). Conversely, circulating miR-200
levels, particularly in plasma or serum from patients with MBC, are
frequently upregulated, and high levels correlate with poorer
prognosis (110,111,115-117,119). Several mechanisms may explain
this discrepancy: (i) Active secretion and selective miRNA
packaging: Tumor cells actively release miRNAs into extracellular
vesicles, such as exosomes, influencing the TME or evading
intracellular tumor-suppressive roles. For example, Liu et
al (79) demonstrated that BC
cells selectively load miR-200c into exosomes through a
Rab1A-mediated mechanism, thus reducing intracellular miR-200c
levels while increasing circulating miR-200c. This 'secretory
escape' could explain elevated circulating miR-200 despite
suppressed tissue expression (79). (ii) Tumor heterogeneity and
molecular subtype variability: BC is highly heterogeneous, and
miRNA expression and functions differ across molecular subtypes
(for example, luminal, HER2-positive, TNBC) (120). (iii) Sample type and detection
methodology differences: Tissue samples reflect localized miRNA
expression, while circulating samples represent systemic average
levels. Additionally, differences in detection platforms, RNA
extraction methods and normalization strategies may contribute to
observed discrepancies (121).
Despite expression inconsistencies, these
differences carry valuable biological and clinical insights.
Elevated circulating miR-200 may indicate active secretion by tumor
cells, closely linked to metastatic behavior. Therefore, clinical
applications should clearly differentiate between tissue-derived
and circulating miRNAs. Tissue miR-200 may be more appropriate for
assessing local tumor characteristics and identifying therapeutic
targets. By contrast, circulating miR-200 serves as a non-invasive
biomarker for dynamic monitoring, metastasis prediction and
treatment efficacy evaluation. Future studies should further
explore miR-200 expression dynamics across sample types and its
specific relationships with tumor biology to advance precision
clinical applications.
Drug resistance
Paclitaxel (PTX) is a first-line chemotherapy agent
for BC treatment; however, tumor cell resistance significantly
limits its clinical efficacy (122). It has been revealed that miR-200
family members play key roles in PTX resistance through multiple
molecular mechanisms. Yu et al (123) reported that miR-200a promotes
resistance of BC cells to paclitaxel, cisplatin and gemcitabine by
targeting TP53INP1 and YAP1. Mechanistically, miR-200a directly
inhibits TP53INP1, weakening the pro-apoptotic p73/p53 pathway, and
simultaneously downregulates YAP1, reducing p73 protein stability.
These coordinated effects suppress transcription of pro-apoptotic
genes (for example, PUMA, Bax, Bim and Noxa), ultimately reducing
chemotherapy-induced apoptosis sensitivity (123). Additionally, Yang et al
(124) demonstrated that
circ-0007331 promotes PTX resistance. Knockdown of circ-0007331
upregulates miR-200b-3p, targeting and reducing ANLN expression,
thus restoring PTX sensitivity (124). Similarly, Li et al
(125) observed significantly
increased LINC00667 expression in exosomes derived from docetaxel
(TXT)-resistant TNBC cells. These exosomes transfer LINC00667 to
recipient TNBC cells, downregulating miR-200b-3p, elevating Bcl-2
expression, and promoting TXT resistance (125). Chen et al (126) demonstrated that miR-200c-3p
restores PTX sensitivity by directly targeting and reducing SOX2
expression. Notably, miR-200 family members may exhibit opposing
regulatory effects under specific conditions. Duan et al
(127) reported that miR-141-3p
inhibits Keap1 expression, activates the Nrf2/SLC7A11-GSH-GPX4
signaling pathway, suppresses ferroptosis, and thus promotes
paclitaxel resistance. Conversely, Tao et al (128) found that miR-141-3p enhances
tumor cell sensitivity to PTX by suppressing autophagy through
downregulation of RAB10.
The miR-200 family also plays a critical role in
regulating resistance to endocrine therapies. Regarding tamoxifen
(TAM) resistance, Hu et al (129) reported that the
circ-UBE2D2/miR-200a-3p axis significantly reduces sensitivity of
ER-positive BC cells to TAM (129). Gao et al (130) revealed that miR-200b/c targets
and suppresses c-MYB, inhibiting EMT and restoring TAM sensitivity.
In relation to other chemotherapies, several studies indicated that
miR-200c restores sensitivity to doxorubicin by inhibiting
molecules such as MDR1 (131),
ANLN (132) and Moesin (133). Moreover, miR-141-3p enhances
trastuzumab sensitivity by targeting and reducing CDK8 expression
(134). In summary, miR-200
family members mediate BC chemotherapy resistance via multiple
target genes and signaling pathways (Table IV), providing potential
intervention targets to reverse resistance.
 | Table IVRegulation of drug resistance by
members of the miR-200 family. |
Table IV
Regulation of drug resistance by
members of the miR-200 family.
| First author/s,
year | miR-200 family
members | Chemotherapy
drugs | Targets | Drug
resistance | (Refs.) |
|---|
| Yu et al,
2018 | miR-200a | PTX | TP53INP1, YAP1 | ↑ | (123) |
| Hu et al,
2020 | | TAM | - | ↓ | (129) |
| Yang et al,
2022 | miR-200b | PTX | ANLN | ↓ | (124) |
| Li et al,
2022 | | TXT | Bcl-2 | ↓ | (125) |
| Chen et al,
2018 | miR-200c | PTX | SOX2 | ↓ | (126) |
| Safaei et
al, 2022 | | DOX | MDR1 | ↓ | (131) |
| Zhang et al,
2020 | | DOX | ANLN | ↓ | (132) |
| Alam et al,
2017 | | DOX | Moesin | ↓ | (133) |
| Duan et al,
2024 | miR-141-3p | PTX | Keap1 | ↑ | (127) |
| Tao et al,
2025 | | PTX | RAB10 | ↓ | (128) |
| Song et al,
2019 | | Trastuzumab | CDK8 | ↓ | (134) |
| Gao et al,
2019 | miR-200b/c | TAM | c-MYB | ↓ | (130) |
Conclusion
In summary, the miR-200 family acts as a critical
regulator in BC initiation and progression. Through diverse
molecular mechanisms, this family controls key biological
processes, including cell proliferation, apoptosis, cell cycle
regulation, invasion, migration and maintenance of stemness. These
findings highlight the significant therapeutic potential of miR-200
and provide novel insights for molecularly targeted BC treatments.
Importantly, the functional impact of the miR-200 family extends
beyond BC. Extensive research demonstrates its conserved role in
regulating EMT and metastasis across multiple cancer types. Such
pan-cancer functional conservation suggests that research on the
miR-200 family may uncover shared tumor metastasis mechanisms,
offering biological insights for early prevention of cancer spread.
From a clinical translation perspective, miR-200 family members
show potential as stable biomarkers for early diagnosis, dynamic
monitoring, and prognosis assessment in BC, allowing integration
into comprehensive screening-diagnosis-monitoring frameworks.
Critically, aberrant miR-200 expression in other cancers, such as
CRC (135) and
cholangiocarcinoma (136),
mirrors its diagnostic and prognostic significance in BC,
positioning miR-200 as a potential pan-cancer early-risk marker.
Notably, differential miR-200 expression between drug-sensitive and
resistant cells underscores its central role in chemotherapy
resistance mechanisms, providing a therapeutic target for
overcoming clinical resistance. However, current research on the
miR-200 family in BC faces limitations: (i) Studies primarily focus
on individual miRNA functions, with insufficient exploration of
synergistic or antagonistic interactions among family members. (ii)
Controversy persists regarding miR-200 expression patterns and
functions across BC subtypes and bodily fluids. Additionally,
efficient utilization of miR-200 for early BC diagnosis and
accurate prognostic assessment remain challenging. Critically,
circulating miR-200 expression used for dynamic monitoring may be
influenced by non-tumor factors, including treatment-related
stress, inflammation, or comorbidities, impeding its specificity
and reliability as a clinical biomarker. (iii) Current research
remains predominantly experimental, lacking substantial integration
into clinical practice. Therefore, future research should
emphasize: First, mechanistically, emerging technologies such as
single-cell and spatial transcriptomics should be leveraged to
elucidate miR-200 expression heterogeneity among different cell
types (for example, cancer cells, CAFs and TAMs) within the BC
microenvironment and to clarify their intercellular communication
roles, enabling a comprehensive understanding of functional
complexity. Second, translational research should urgently advance
three areas: (i) Developing miR-200-based interventions, such as
targeted delivery systems using exosomes or lipid nanoparticles for
miR-200 mimics or inhibitors; (ii) establishing standardized
detection and validation systems through multicenter, multicohort
studies to define absolute quantification standards and dynamic
thresholds for circulating miR-200 across different BC subtypes and
treatment stages; (iii) advancing precision detection technologies
and constructing multifactorial dynamic monitoring models to fully
harness clinical application potential. Finally, prospective
clinical studies should systematically evaluate miR-200 biomarker
utility throughout BC management, from prevention to diagnosis,
treatment and monitoring, while exploring combined application with
imaging, pathology and existing serum markers. Comprehensive and
systematic investigation promises to establish novel theoretical
foundations and therapeutic targets for precision BC management,
ultimately improving patient outcomes.
Availability of data and materials
Not applicable.
Authors' contributions
JL and HD were responsible for manuscript writing,
conceived and designed the study. YS and JL were responsible for
the collection and assembly of data. YS and HD were responsible for
data analysis and interpretation. All authors read and approved the
final version of the manuscript. Data authentication is not
applicable.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declate that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
The present study was supported by the Inner Mongolia Science
and Technology Research Project (grant no. 2024MS08069), the
Science and Technology Program of the Joint Fund of Scientific
Research for the Public Hospitals of Inner Mongolia Academy of
Medical Sciences (grant no. 2024GLLH0323), the Key Technologies
Research and development program of Inner Mongolia (grant no.
2021GG0170), the general Program of Inner Mongolia Medical
University (grant no. YKD2021MS006), the 14th Five-Year Plan of
Science and Technology Innovation in Inner Mongolia Autonomous
Region (grant no. 2022YFSH0078), the Key project of Inner Mongolia
Medical University (grant no. YKD2021ZD007), the Zhiyuan Talent
Program of Inner Mongolia Medical University (grant nos. ZY0202020
and ZY20242107), the Doctoral Start-up Foundation Project of Inner
Mongolia Medical University (grant no. YKD2024BSQD026), the
Undergraduate Teaching Reform Research and Practice Project of
Inner Mongolia Medical University in 2024 (grant no.
NYJXGGSJ20244046), the Inner Mongolia Medical University 2024 Maker
Cultivation'Project (grant no. 101322024038) and the Inner Mongolia
Medical University 2025 Cultivation of Excellence Programme in
Science and Technology Innovation for Undergraduates (grant no.
YCPY2025057).
References
|
1
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI
|
|
2
|
Xiong X, Zheng LW, Ding Y, Chen YF, Cai
YW, Wang LP, Huang L, Liu CC, Shao ZM and Yu KD: Breast cancer:
Pathogenesis and treatments. Signal Transduct Target Ther.
10:492025. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Khan MM, Yalamarty SSK, Rajmalani BA,
Filipczak N and Torchilin VP: Recent strategies to overcome breast
cancer resistance. Crit Rev Oncol Hematol. 197:1043512024.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Diener C, Keller A and Meese E: Emerging
concepts of miRNA therapeutics: from cells to clinic. Trends Genet.
38:613–626. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Dinami R, Pompili L, Petti E, Porru M,
D'Angelo C, Di Vito S, Rizzo A, Campani V, De Rosa G, Bruna A, et
al: MiR-182-3p targets TRF2 and impairs tumor growth of
triple-negative breast cancer. EMBO Mol Med. 15:e160332023.
View Article : Google Scholar :
|
|
6
|
Ho HL, Lin SC, Chiang CW, Lin C, Liu CW,
Yeh YC, Chen MY and Chou TY: miR-193b-3p suppresses lung cancer
cell migration and invasion through PRNP targeting. J Biomed Sci.
32:282025. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhang T, Hu Y, Yang N, Yu S and Pu X: The
microRNA-34 family and its functional role in lung cancer. Am J
Clin Oncol. 47:448–457. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Klicka K, Grzywa TM, Mielniczuk A, Klinke
A and Włodarski PK: The role of miR-200 family in the regulation of
hallmarks of cancer. Front Oncol. 12:9652312022. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Mao Y, Chen W, Wu H, Liu C, Zhang J and
Chen S: Mechanisms and functions of MiR-200 family in
hepatocellular carcinoma. Onco Targets Ther. 13:13479–13490. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Liu C, Hu W, Li LL, Zhou Q, Zhang F,
Song-Yang YY, Zhu W, Sun CC and Li DJ: Roles of miR-200 family
members in lung cancer: more than tumor suppressors. Future Oncol.
14:2875–2886. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Choi PW and Ng SW: The Functions of
MicroRNA-200 Family in ovarian cancer: Beyond
epithelial-mesenchymal transition. Int J Mol Sci. 18:12072017.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Choi PS, Zakhary L, Choi WY, Caron S,
Alvarez-Saavedra E, Miska EA, McManus M, Harfe B, Giraldez AJ,
Horvitz HR, et al: Members of the miRNA-200 family regulate
olfactory neurogenesis. Neuron. 57:41–55. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Trümbach D and Prakash N: The conserved
miR-8/miR-200 microRNA family and their role in invertebrate and
vertebrate neurogenesis. Cell Tissue Res. 359:161–177. 2015.
View Article : Google Scholar
|
|
14
|
Lim YY, Wright JA, Attema JL, Gregory PA,
Bert AG, Smith E, Thomas D, Lopez AF, Drew PA, Khew-Goodall Y and
Goodall GJ: Epigenetic modulation of the miR-200 family is
associated with transition to a breast cancer stem-cell-like state.
J Cell Sci. 126(Pt 10): 2256–2266. 2013.PubMed/NCBI
|
|
15
|
Vrba L, Garbe JC, Stampfer MR and Futscher
BW: Epigenetic regulation of normal human mammary cell
type-specific miRNAs. Genome Res. 21:2026–2037. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Magenta A, Cencioni C, Fasanaro P,
Zaccagnini G, Greco S, Sarra-Ferraris G, Antonini A, Martelli F and
Capogrossi MC: miR-200c is upregulated by oxidative stress and
induces endothelial cell apoptosis and senescence via ZEB1
inhibition. Cell Death Differ. 18:1628–1639. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Jing J, Xiong S, Li Z, Wu J, Zhou L, Gui
JF and Mei J: A feedback regulatory loop involving p53/miR-200 and
growth hormone endocrine axis controls embryo size of zebrafish.
Sci Rep. 5:159062015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Bracken CP, Gregory PA, Kolesnikoff N,
Bert AG, Wang J, Shannon MF and Goodall GJ: A double-negative
feedback loop between ZEB1-SIP1 and the microRNA-200 family
regulates epithelial-mesenchymal transition. Cancer Res.
68:7846–7854. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Burk U, Schubert J, Wellner U, Schmalhofer
O, Vincan E, Spaderna S and Brabletz T: A reciprocal repression
between ZEB1 and members of the miR-200 family promotes EMT and
invasion in cancer cells. EMBO Rep. 9:582–589. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Eades G, Yao Y, Yang M, Zhang Y, Chumsri S
and Zhou Q: miR-200a regulates SIRT1 expression and epithelial to
mesenchymal transition (EMT)-like transformation in mammary
epithelial cells. J Biol Chem. 286:25992–6002. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kim HK, Park JD, Choi SH, Shin DJ, Hwang
S, Jung HY and Park KS: Functional link between miR-200a and ELK3
regulates the metastatic nature of breast cancer. Cancers (Basel).
12:12252020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kim HY, Ha Thi HT and Hong S: IMP2 and
IMP3 cooperate to promote the metastasis of triple-negative breast
cancer through destabilization of progesterone receptor. Cancer
Lett. 415:30–39. 2018. View Article : Google Scholar
|
|
23
|
Sossey-Alaoui K, Pluskota E, Szpak D,
Schiemann WP and Plow EF: The Kindlin-2 regulation of
epithelial-to-mesenchymal transition in breast cancer metastasis is
mediated through miR-200b. Sci Rep. 8:73602018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Peng F, Tang H, Du J, Chen J and Peng C:
Isoliquiritigenin suppresses EMT-Induced metastasis in
triple-negative breast cancer through miR-200c/C-JUN/β-Catenin. Am
J Chin Med. 49:505–523. 2021. View Article : Google Scholar
|
|
25
|
Li RH, Chen M, Liu J, Shao CC, Guo CP, Wei
XL, Li YC, Huang WH and Zhang GJ: Long noncoding RNA ATB promotes
the epithelial-mesenchymal transition by upregulating the
miR-200c/Twist1 axe and predicts poor prognosis in breast cancer.
Cell Death Dis. 9:11712018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhou W, Ye XL, Xu J, Cao MG, Fang ZY, Li
LY, Guan GH, Liu Q, Qian YH and Xie D: The lncRNA H19 mediates
breast cancer cell plasticity during EMT and MET plasticity by
differentially sponging miR-200b/c and let-7b. Sci Signal.
10:eaak95572017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ren S, Liu J, Feng Y, Li Z, He L, Li L,
Cao X, Wang Z and Zhang Y: Knockdown of circDENND4C inhibits
glycolysis, migration and invasion by up-regulating miR-200b/c in
breast cancer under hypoxia. J Exp Clin Cancer Res. 38:3882019.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zhou D, Gu J, Wang Y, Wu H, Cheng W, Wang
Q, Zheng G and Wang X: Long non-coding RNA NEAT1 transported by
extracellular vesicles contributes to breast cancer development by
sponging microRNA-141-3p and regulating KLF12. Cell Biosci.
11:682021. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Xu M, Wang S, Wang Y, Wu H, Frank JA,
Zhang Z and Luo J: Role of p38γ MAPK in regulation of EMT and
cancer stem cells. Biochim Biophys Acta Mol Basis Dis.
1864:3605–3617. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zhang G, Zhang W, Li B, Stringer-Reasor E,
Chu C, Sun L, Bae S, Chen D, Wei S, Jiao K, et al: MicroRNA-200c
and microRNA-141 are regulated by a FOXP3-KAT2B axis and associated
with tumor metastasis in breast cancer. Breast Cancer Res.
19:732017. View Article : Google Scholar
|
|
31
|
Zeng X, Qu X, Zhao C, Xu L, Hou K, Liu Y,
Zhang N, Feng J, Shi S, Zhang L, et al: FEN1 mediates miR-200a
methylation and promotes breast cancer cell growth via MET and EGFR
signaling. FASEB J. 33:10717–10730. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tang X, Tu G, Yang G, Wang X, Kang L, Yang
L, Zeng H, Wan X, Qiao Y, Cui X, et al: Autocrine
TGF-β1/miR-200s/miR-221/DNMT3B regulatory loop maintains CAF status
to fuel breast cancer cell proliferation. Cancer Lett. 452:79–89.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Sun S, Ma J, Xie P, Wu Z and Tian X:
Hypoxia-responsive miR-141-3p is involved in the progression of
breast cancer via mediating the HMGB1/HIF-1α signaling pathway. J
Gene Med. 22:e32302020. View Article : Google Scholar
|
|
34
|
Wang JG, Jian WJ, Li Y and Zhang J:
Nobiletin promotes the pyroptosis of breast cancer via regulation
of miR-200b/JAZF1 axis. Kaohsiung J Med Sci. 37:572–582. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Song C, Liu LZ, Pei XQ, Liu X, Yang L, Ye
F and Xie X, Chen J, Tang H and Xie X: miR-200c inhibits breast
cancer proliferation by targeting KRAS. Oncotarget. 6:34968–34978.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhang L, Liu Q, Mu Q, Zhou D, Li H, Zhang
B and Yin C: MiR-429 suppresses proliferation and invasion of
breast cancer via inhibiting the Wnt/β-catenin signaling pathway.
Thorac Cancer. 11:3126–3138. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Xu F, Hu Y, Gao J, Wang J, Xie Y, Sun F,
Wang L, Miyamoto A, Xia O and Zhang C: HIF-1α/Malat1/miR-141 axis
activates autophagy to increase proliferation, migration, and
invasion in triple-negative breast cancer. Curr Cancer Drug
Targets. 23:363–378. 2023. View Article : Google Scholar
|
|
38
|
Sun Y, Yang D, Xi L, Chen Y, Fu L, Sun K,
Yin J, Li X, Liu S, Qin Y, et al: Primed atypical ductal
hyperplasia-associated fibroblasts promote cell growth and polarity
changes of transformed epithelium-like breast cancer MCF-7 cells
via miR-200b/c-IKKβ signaling. Cell Death Dis. 9:1222018.
View Article : Google Scholar
|
|
39
|
Liu B, Du R, Zhou L, Xu J, Chen S, Chen J,
Yang X, Liu DX, Shao ZM, Zhang L, et al: miR-200c/141 regulates
breast cancer stem cell heterogeneity via targeting HIPK1/β-Catenin
axis. Theranostics. 8:5801–5813. 2018. View Article : Google Scholar
|
|
40
|
Saha T, Solomon J, Samson AO and Gil-Henn
H: Invasion and metastasis as a central hallmark of breast cancer.
J Clin Med. 10:34982021. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Pastushenko I and Blanpain C: EMT
transition states during tumor progression and metastasis. Trends
Cell Biol. 29:212–226. 2019. View Article : Google Scholar
|
|
42
|
Brabletz S, Schuhwerk H, Brabletz T and
Stemmler MP: Dynamic EMT: A multi-tool for tumor progression. EMBO
J. 40:e1086472021. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Ahmad A, Aboukameel A, Kong D, Wang Z,
Sethi S, Chen W, Sarkar FH and Raz A: Phosphoglucose
isomerase/autocrine motility factor mediates epithelial-mesenchymal
transition regulated by miR-200 in breast cancer cells. Cancer Res.
71:3400–3409. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Gregory PA, Bert AG, Paterson EL, Barry
SC, Tsykin A, Farshid G, Vadas MA, Khew-Goodall Y and Goodall GJ:
The miR-200 family and miR-205 regulate epithelial to mesenchymal
transition by targeting ZEB1 and SIP1. Nat Cell Biol. 10:593–601.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Lorenzo-Martín LF, Citterio C,
Menacho-Márquez M, Conde J, Larive RM, Rodríguez-Fdez S,
García-Escudero R, Robles-Valero J, Cuadrado M, Fernández-Pisonero
I, et al: Vav proteins maintain epithelial traits in breast cancer
cells using miR-200c-dependent and independent mechanisms.
Oncogene. 38:209–227. 2019. View Article : Google Scholar :
|
|
46
|
Lu Z, Jiao D, Qiao J, Yang S, Yan M, Cui S
and Liu Z: Restin suppressed epithelial-mesenchymal transition and
tumor metastasis in breast cancer cells through upregulating
mir-200a/b expression via association with p73. Mol Cancer.
14:1022015. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Roy SS, Gonugunta VK, Bandyopadhyay A, Rao
MK, Goodall GJ, Sun LZ, Tekmal RR and Vadlamudi RK: Significance of
PELP1/HDAC2/miR-200 regulatory network in EMT and metastasis of
breast cancer. Oncogene. 33:3707–3716. 2014. View Article : Google Scholar
|
|
48
|
Ye ZB, Ma G, Zhao YH, Xiao Y, Zhan Y, Jing
C, Gao K, Liu ZH and Yu SJ: miR-429 inhibits migration and invasion
of breast cancer cells in vitro. Int J Oncol. 46:531–538. 2015.
View Article : Google Scholar
|
|
49
|
Zhang Y, Li J, Jia S, Wang Y, Kang Y and
Zhang W: Down-regulation of lncRNA-ATB inhibits
epithelial-mesenchymal transition of breast cancer cells by
increasing miR-141-3p expression. Biochem Cell Biol. 97:193–200.
2019. View Article : Google Scholar
|
|
50
|
Zou Q, Zhou E, Xu F, Zhang D, Yi W and Yao
J: A TP73-AS1/miR-200a/ZEB1 regulating loop promotes breast cancer
cell invasion and migration. J Cell Biochem. 119:2189–2199. 2018.
View Article : Google Scholar
|
|
51
|
Damiano V, Brisotto G, Borgna S, di
Gennaro A, Armellin M, Perin T, Guardascione M, Maestro R and
Santarosa M: Epigenetic silencing of miR-200c in breast cancer is
associated with aggressiveness and is modulated by ZEB1. Genes
Chromosomes Cancer. 56:147–158. 2017. View Article : Google Scholar
|
|
52
|
Sundararajan V, Gengenbacher N, Stemmler
MP, Kleemann JA, Brabletz T and Brabletz S: The ZEB1/miR-200c
feedback loop regulates invasion via actin interacting proteins
MYLK and TKS5. Oncotarget. 6:27083–27096. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Teng Y, Mei Y, Hawthorn L and Cowell JK:
WASF3 regulates miR-200 inactivation by ZEB1 through suppression of
KISS1 leading to increased invasiveness in breast cancer cells.
Oncogene. 33:203–211. 2014. View Article : Google Scholar :
|
|
54
|
Bendoraite A, Knouf EC, Garg KS, Parkin
RK, Kroh EM, O'Briant KC, Ventura AP, Godwin AK, Karlan BY,
Drescher CW, et al: Regulation of miR-200 family microRNAs and ZEB
transcription factors in ovarian cancer: Evidence supporting a
mesothelial-to-epithelial transition. Gynecol Oncol. 116:117–125.
2010. View Article : Google Scholar
|
|
55
|
Wang Y, Wu Z and Hu L: The regulatory
effects of metformin on the [SNAIL/miR-34]:[ZEB/miR-200] system in
the epithelial-mesenchymal transition(EMT) for colorectal
cancer(CRC). Eur J Pharmacol. 834:45–53. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Pruitt K, Zinn RL, Ohm JE, McGarvey KM,
Kang SH, Watkins DN, Herman JG and Baylin SB: Inhibition of SIRT1
reactivates silenced cancer genes without loss of promoter DNA
hypermethylation. PLoS Genet. 2:e402006. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Cho HJ, Oh N, Park JH, Kim KS, Kim HK, Lee
E, Hwang S, Kim SJ and Park KS: ZEB1 Collaborates with ELK3 to
Repress E-Cadherin expression in triple-negative breast cancer
cells. Mol Cancer Res. 17:2257–2266. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Tsai JH, Donaher JL, Murphy DA, Chau S and
Yang J: Spatiotemporal regulation of epithelial-mesenchymal
transition is essential for squamous cell carcinoma metastasis.
Cancer Cell. 22:725–736. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Gilmore AP: Anoikis. Cell Death Differ.
12(Suppl 2): S1473–S1477. 2005. View Article : Google Scholar
|
|
60
|
Yuan M, Tomlinson V, Lara R, Holliday D,
Chelala C, Harada T, Gangeswaran R, Manson-Bishop C, Smith P,
Danovi SA, et al: Yes-associated protein (YAP) functions as a tumor
suppressor in breast. Cell Death Differ. 15:1752–1759. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Yu SJ, Hu JY, Kuang XY, Luo JM, Hou YF, Di
GH, Wu J, Shen ZZ, Song HY and Shao ZM: MicroRNA-200a promotes
anoikis resistance and metastasis by targeting YAP1 in human breast
cancer. Clin Cancer Res. 19:1389–1399. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Howe EN, Cochrane DR and Richer JK:
Targets of miR-200c mediate suppression of cell motility and
anoikis resistance. Breast Cancer Res. 13:R452011. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Kozak J, Wdowiak P, Maciejewski R and
Torres A: Interactions between microRNA-200 family and Sestrin
proteins in endometrial cancer cell lines and their significance to
anoikis. Mol Cell Biochem. 459:21–34. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Miao H, Li DQ, Mukherjee A, Guo H, Petty
A, Cutter J, Basilion JP, Sedor J, Wu J, Danielpour D, et al: EphA2
mediates ligand-dependent inhibition and ligand-independent
promotion of cell migration and invasion via a reciprocal
regulatory loop with Akt. Cancer Cell. 16:9–20. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Tsouko E, Wang J, Frigo DE, Aydoğdu E and
Williams C: miR-200a inhibits migration of triple-negative breast
cancer cells through direct repression of the EPHA2 oncogene.
Carcinogenesis. 36:1051–1060. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Ming J, Zhou Y, Du J, Fan S, Pan B, Wang
Y, Fan L and Jiang J: Identification of miR-200a as a novel
suppressor of connexin 43 in breast cancer cells. Biosci Rep.
35:e002512015. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Clucas J and Valderrama F: ERM proteins in
cancer progression. J Cell Sci. 127(Pt 2): 267–275. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Hong H, Yu H, Yuan J, Guo C, Cao H, Li W
and Xiao C: MicroRNA-200b impacts breast cancer cell migration and
invasion by regulating ezrin-radixin-moesin. Med Sci Monit.
22:1946–1952. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Li X, Roslan S, Johnstone CN, Wright JA,
Bracken CP, Anderson M, Bert AG, Selth LA, Anderson RL, Goodall GJ,
et al: MiR-200 can repress breast cancer metastasis through
ZEB1-independent but moesin-dependent pathways. Oncogene.
33:4077–4088. 2014. View Article : Google Scholar
|
|
70
|
Yuan J, Xiao C, Lu H, Yu H, Hong H, Guo C
and Wu Z: miR-200b regulates breast cancer cell proliferation and
invasion by targeting radixin. Exp Ther Med. 19:2741–2750.
2020.PubMed/NCBI
|
|
71
|
Zhang X, Yu X, Zhao Z, Yuan Z, Ma P, Ye Z,
Guo L, Xu S, Xu L, Liu T, et al: MicroRNA-429 inhibits bone
metastasis in breast cancer by regulating CrkL and MMP-9. Bone.
130:1151392020. View Article : Google Scholar
|
|
72
|
Choi SK, Kim HS, Jin T, Hwang EH, Jung M
and Moon WK: Overexpression of the miR-141/200c cluster promotes
the migratory and invasive ability of triple-negative breast cancer
cells through the activation of the FAK and PI3K/AKT signaling
pathways by secreting VEGF-A. BMC Cancer. 16:5702016. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Jin T, Suk Kim H, Ki Choi S, Hye Hwang E,
Woo J, Suk Ryu H, Kim K, Moon A and Kyung Moon W: microRNA-200c/141
upregulates SerpinB2 to promote breast cancer cell metastasis and
reduce patient survival. Oncotarget. 8:32769–32782. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Humphries B, Wang Z, Oom AL, Fisher T, Tan
D, Cui Y, Jiang Y and Yang C: MicroRNA-200b targets protein kinase
Cα and suppresses triple-negative breast cancer metastasis.
Carcinogenesis. 35:2254–2263. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Humphries B, Wang Z, Li Y, Jhan JR, Jiang
Y and Yang C: ARHGAP18 downregulation by miR-200b suppresses
metastasis of triple-negative breast cancer by enhancing activation
of RhoA. Cancer Res. 77:4051–4064. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Sigloch FC, Burk UC, Biniossek ML,
Brabletz T and Schilling O: miR-200c dampens cancer cell migration
via regulation of protein kinase A subunits. Oncotarget.
6:23874–23889. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Li P, Xu T, Zhou X, Liao L, Pang G, Luo W,
Han L, Zhang J, Luo X, Xie X and Zhu K: Downregulation of miRNA-141
in breast cancer cells is associated with cell migration and
invasion: involvement of ANP32E targeting. Cancer Med. 6:662–672.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Zheng Q, Cui X, Zhang D, Yang Y, Yan X,
Liu M, Niang B, Aziz F, Liu S, Yan Q and Liu J: miR-200b inhibits
proliferation and metastasis of breast cancer by targeting
fucosyltransferase IV and α1,3-fucosylated glycans. Oncogenesis.
6:e3582017. View Article : Google Scholar
|
|
79
|
Liu Y, Tang J, Qiu X, Teng LA, Sriwastva
MK, Han X, Li Z, Liu M, Liu S, Da D, et al: Rab1A-mediated exosomal
sorting of miR-200c enhances breast cancer lung metastasis. Breast
Cancer (Dove Med Press). 15:403–419. 2023.PubMed/NCBI
|
|
80
|
Yao J, Zhou E, Wang Y, Xu F, Zhang D and
Zhong D: microRNA-200a inhibits cell proliferation by targeting
mitochondrial transcription factor A in breast cancer. DNA Cell
Biol. 33:291–300. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Liu Y, Kao HI and Bambara RA: Flap
endonuclease 1: A central component of DNA metabolism. Annu Rev
Biochem. 73:589–615. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Zheng L, Dai H, Zhou M, Li M, Singh P, Qiu
J, Tsark W, Huang Q, Kernstine K, Zhang X, et al: Fen1 mutations
result in autoimmunity, chronic inflammation and cancers. Nat Med.
13:812–819. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
83
|
Peng G, Yan J, Shi P and Li H: LINC01140
hinders the development of breast cancer through targeting
miR-200b-3p to downregulate DMD. Cell Transplant.
32:96368972312112022023. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Jones R, Watson K, Bruce A, Nersesian S,
Kitz J and Moorehead R: Re-expression of miR-200c suppresses
proliferation, colony formation and in vivo tumor growth of murine
claudin-low mammary tumor cells. Oncotarget. 8:23727–23749. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Wang C, Ju H, Shen C and Tong Z: miR-429
mediates δ-tocotrienol-induced apoptosis in triple-negative breast
cancer cells by targeting XIAP. Int J Clin Exp Med. 8:15648–15656.
2015.
|
|
86
|
Bi M, Zheng L, Chen L, He J, Yuan C, Ma P,
Zhao Y, Hu F, Tang W and Sheng M: ln RNA LINC01234 promotes
triple-negative breast cancer progression through regulating the
miR-429/SYNJ1 axis. Am J Transl Res. 13:11399–11412.
2021.PubMed/NCBI
|
|
87
|
Li Y, Meng X, Luo Y, Luo S, Li J, Zeng J,
Huang X and Wang J: The oncogenic miR-429 promotes triple-negative
breast cancer progression by degrading DLC1. Aging (Albany NY).
15:9809–9821. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Dong S, Ma M, Li M, Guo Y, Zuo X, Gu X,
Zhang M and Shi Y: LncRNA MEG3 regulates breast cancer
proliferation and apoptosis through miR-141-3p/RBMS3 axis.
Genomics. 113:1689–1704. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Wang X, Ji C, Hu J, Deng X, Zheng W, Yu Y,
Hua K, Zhou X and Fang L: Hsa_circ_0005273 facilitates breast
cancer tumorigenesis by regulating YAP1-hippo signaling pathway. J
Exp Clin Cancer Res. 40:292021. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Yao Y, Hu J, Shen Z, Yao R, Liu S, Li Y,
Cong H, Wang X, Qiu W and Yue L: MiR-200b expression in breast
cancer: A prognostic marker and act on cell proliferation and
apoptosis by targeting Sp1. J Cell Mol Med. 19:760–769. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Wang W, Li Y, Zhu JY, Fang D, Ding HF,
Dong Z, Jing Q, Su SB and Huang S: Triple negative breast cancer
development can be selectively suppressed by sustaining an elevated
level of cellular cyclic AMP through simultaneously blocking its
efflux and decomposition. Oncotarget. 7:87232–87245. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Zhang DD, Li Y, Xu Y, Kim J and Huang S:
Phosphodiesterase 7B/microRNA-200c relationship regulates
triple-negative breast cancer cell growth. Oncogene. 38:1106–1120.
2019. View Article : Google Scholar
|
|
93
|
Li D, Wang H, Song H, Xu H, Zhao B, Wu C,
Hu J, Wu T, Xie D, Zhao J, et al: The microRNAs miR-200b-3p and
miR-429-5p target the LIMK1/CFL1 pathway to inhibit growth and
motility of breast cancer cells. Oncotarget. 8:85276–85289. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Bahrami A, Hassanian SM, Khazaei M,
Hasanzadeh M, Shahidsales S, Maftouh M, Ferns GA and Avan A: The
therapeutic potential of targeting tumor microenvironment in breast
cancer: rational strategies and recent progress. J Cell Biochem.
119:111–122. 2018. View Article : Google Scholar
|
|
95
|
Lin Z, Roche ME, Díaz-Barros V,
Domingo-Vidal M, Whitaker-Menezes D, Tuluc M, Uppal G, Caro J,
Curry JM and Martinez-Outschoorn U: MiR-200c reprograms fibroblasts
to recapitulate the phenotype of CAFs in breast cancer progression.
Cell Stress. 8:1–20. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Meng Z, Zhang R, Wang Y, Zhu G, Jin T, Li
C and Zhang S: miR-200c/PAI-2 promotes the progression of triple
negative breast cancer via M1/M2 polarization induction of
macrophage. Int Immunopharmacol. 81:1060282020. View Article : Google Scholar
|
|
97
|
Raue R, Frank AC, Fuhrmann DC, de la
Cruz-Ojeda P, Rösser S, Bauer R, Cardamone G, Weigert A, Syed SN,
Schmid T and Brüne B: MicroRNA-200c attenuates the
tumor-infiltrating capacity of macrophages. Biology (Basel).
11:3492022.PubMed/NCBI
|
|
98
|
Yang F, Xu J, Tang L and Guan X: Breast
cancer stem cell: the roles and therapeutic implications. Cell Mol
Life Sci. 74:951–966. 2017. View Article : Google Scholar
|
|
99
|
Hsu TW, Chen HA, Liao PH, Su YH, Chiu CF,
Huang CY, Lin YJ, Hung CC, Yeh MH, Sung SY and Su CM:
Dicer-mediated miR-200b expression contributes to cell
migratory/invasive abilities and cancer stem cells properties of
breast cancer cells. Aging (Albany NY). 14:6520–6536. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Iliopoulos D, Lindahl-Allen M, Polytarchou
C, Hirsch HA, Tsichlis PN and Struhl K: Loss of miR-200 inhibition
of Suz12 leads to polycomb-mediated repression required for the
formation and maintenance of cancer stem cells. Mol Cell.
39:761–772. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Tang H, Song C, Ye F, Gao G, Ou X, Zhang L
and Xie X and Xie X: miR-200c suppresses stemness and increases
cellular sensitivity to trastuzumab in HER2+ breast cancer. J Cell
Mol Med. 23:8114–8127. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Wu D, Ji N and Zhang L and Zhang L:
Differential expression of miR-200c in breast cancer stem cells.
Int J Clin Exp Pathol. 10:10085–10091. 2017.PubMed/NCBI
|
|
103
|
Liu S, Cong Y, Wang D, Deng L, Liu Y,
Martin-Trevino R, Shang L, McDermott SP, Landis MD, Hong S, et al:
Breast cancer stem cells transition between epithelial and
mesenchymal states reflective of their normal counterparts. Stem
Cell Reports. 2:78–91. 2013. View Article : Google Scholar
|
|
104
|
Lee Y, Ni J, Beretov J, Wasinger VC,
Graham P and Li Y: Recent advances of small extracellular vesicle
biomarkers in breast cancer diagnosis and prognosis. Mol Cancer.
22:332023. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Yang T, Li W, Huang T and Zhou J: Genetic
testing enhances the precision diagnosis and treatment of breast
cancer. Int J Mol Sci. 24:166072023. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Cardinali B, Tasso R, Piccioli P, Ciferri
MC, Quarto R and Del Mastro L: Circulating miRNAs in breast cancer
diagnosis and prognosis. Cancers (Basel). 14:23172022. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Xu F, He H, Huang W, Lin Y, Luo S, Du Q
and Duan R: Decreased expression of MicroRNA-200 family in human
breast cancer is associated with lymph node metastasis. Clin Transl
Oncol. 18:283–288. 2016. View Article : Google Scholar
|
|
108
|
Fahim SA, Abdullah MS, Espinoza-Sánchez
NA, Hassan H, Ibrahim AM, Ahmed SH, Shakir G, Badawy MA, Zakhary
NI, Greve B, et al: Inflammatory breast carcinoma: Elevated
microRNA miR-181b-5p and Reduced miR-200b-3p, miR-200c-3p, and
miR-203a-3p expression as potential biomarkers with diagnostic
value. Biomolecules. 10:10592020. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Qiao P, Du H, Guo X, Yu M, Zhang C and Shi
Y: Serum exosomal miR-200c is a potential diagnostic biomarker for
breast cancer. Biomarkers. 29:419–426. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Khalil M, Desouky EM, Khaliefa AK, Hozyen
WG, Mohamed SS and Hasona NA: Insights into the crosstalk between
miR-200a/lncRNA H-19 and IL-6/SIRT-1 axis in breast cancer. J
Interferon Cytokine Res. 44:191–197. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Papadaki C, Stoupis G, Tsalikis L,
Monastirioti A, Papadaki M, Maliotis N, Stratigos M, Mastrostamatis
G, Mavroudis D and Agelaki S: Circulating miRNAs as a marker of
metastatic disease and prognostic factor in metastatic breast
cancer. Oncotarget. 10:966–981. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Braicu C, Raduly L, Morar-Bolba G,
Cojocneanu R, Jurj A, Pop LA, Pileczki V, Ciocan C, Moldovan A,
Irimie A, et al: Aberrant miRNAs expressed in HER-2 negative breast
cancers patient. J Exp Clin Cancer Res. 37:2572018. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Ye F, Tang H, Liu Q and Xie X, Wu M, Liu
X, Chen B and Xie X: miR-200b as a prognostic factor in breast
cancer targets multiple members of RAB family. J Transl Med.
12:172014. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Amorim M, Lobo J, Fontes-Sousa M,
Estevão-Pereira H, Salta S, Lopes P, Coimbra N, Antunes L, Palma de
Sousa S, Henrique R and Jerónimo C: Predictive and prognostic value
of selected MicroRNAs in luminal breast cancer. Front Genet.
10:8152019. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Fischer C, Deutsch TM, Feisst M, Rippinger
N, Riedel F, Hartkopf AD, Brucker SY, Domschke C, Fremd C, Michel
L, et al: Circulating miR-200 family as predictive markers during
systemic therapy of metastatic breast cancer. Arch Gynecol Obstet.
306:875–885. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Fischer C, Turchinovich A, Feisst M,
Riedel F, Haßdenteufel K, Scharli P, Hartkopf AD, Brucker SY,
Michel L, Burwinkel B, et al: Circulating miR-200 family and CTCs
in metastatic breast cancer before, during, and after a new line of
systemic treatment. Int J Mol Sci. 23:95352022. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Madhavan D, Peng C, Wallwiener M, Zucknick
M, Nees J, Schott S, Rudolph A, Riethdorf S, Trumpp A, Pantel K, et
al: Circulating miRNAs with prognostic value in metastatic breast
cancer and for early detection of metastasis. Carcinogenesis.
37:461–470. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Shao B, Wang X, Zhang L, Li D, Liu X, Song
G, Cao H, Zhu J and Li H: Plasma microRNAs predict chemoresistance
in patients with metastatic breast cancer. Technol Cancer Res
Treat. 18:15330338198287092019. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Navarro-Manzano E, Luengo-Gil G,
González-Conejero R, García-Garre E, García-Martínez E,
García-Torralba E, Chaves-Benito A, Vicente V and Ayala de la Peña
F: Prognostic and predictive effects of tumor and plasma
miR-200c-3p in locally advanced and metastatic breast cancer.
Cancers (Basel). 14:23902022. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Hamam R, Hamam D, Alsaleh KA, Kassem M,
Zaher W, Alfayez M, Aldahmash A and Alajez NM: Circulating
microRNAs in breast cancer: novel diagnostic and prognostic
biomarkers. Cell Death Dis. 8:e30452017. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Moldovan L, Batte KE, Trgovcich J, Wisler
J, Marsh CB and Piper M: Methodological challenges in utilizing
miRNAs as circulating biomarkers. J Cell Mol Med. 18:371–390. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Abu Samaan TM, Samec M, Liskova A, Kubatka
P and Büsselberg D: Paclitaxel's mechanistic and clinical effects
on breast cancer. Biomolecules. 9:7892019. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Yu SJ, Yang L, Hong Q, Kuang XY, Di GH and
Shao ZM: MicroRNA-200a confers chemoresistance by antagonizing
TP53INP1 and YAP1 in human breast cancer. BMC Cancer. 18:742018.
View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Yang C, Liu L, Gao C, Zhang G, Zhang Y,
Zhang S, Li J and Liu Y: Circ_0,007,331 Promotes the PTX Resistance
and Progression of Breast Cancer via miR-200b-3p/ANLN. J Surg Res.
279:619–632. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Li J, Kang J, Liu W, Liu J, Pan G, Mao A,
Zhang Q, Lu J, Ding J and Li H: Docetaxel-resistant triple-negative
breast cancer cell-derived exosomal lncRNA LINC00667 reduces the
chemo-sensitivity of breast cancer cells to docetaxel
<em>via</em> targeting miR-200b-3p/Bcl-2 axis. Eur J
Histochem. 66:35292022. View Article : Google Scholar
|
|
126
|
Chen J, Tian W, He H, Chen F, Huang J,
Wang X and Chen Z: Downregulation of miR-200c-3p contributes to the
resistance of breast cancer cells to paclitaxel by targeting SOX2.
Oncol Rep. 40:3821–3829. 2018.PubMed/NCBI
|
|
127
|
Duan WL, Wang XJ, Guo A, Gu LH, Sheng ZM,
Luo H, Yang LX, Wang WH and Zhang BG: miR-141-3p promotes
paclitaxel resistance by attenuating ferroptosis via the Keap1-Nrf2
signaling pathway in breast cancer. J Cancer. 15:5622–5635. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Tao S, Ji Y, Li R, Xiao Y, Wu H, Ye R, Shi
J, Geng C, Tang G, Ran R, et al: Layered double hydroxide
LDH-Loaded miR-141-3p Targets RAB10 suppressing cellular autophagy
to reverse paclitaxel resistance in breast cancer. ACS Omega.
10:5886–5899. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Hu K, Liu X, Li Y, Li Q, Xu Y, Zeng W,
Zhong G and Yu C: Exosomes mediated transfer of Circ_UBE2D2
enhances the resistance of breast cancer to tamoxifen by binding to
MiR-200a-3p. Med Sci Monit. 26:e9222532020. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Gao Y, Zhang W, Liu C and Li G: miR-200
affects tamoxifen resistance in breast cancer cells through
regulation of MYB. Sci Rep. 9:188442019. View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Safaei S, Amini M, Najjary S, Mokhtarzadeh
A, Bolandi N, Saeedi H, Alizadeh N, Javadrashid D and Baradaran B:
miR-200c increases the sensitivity of breast cancer cells to
Doxorubicin through downregulating MDR1 gene. Exp Mol Pathol.
125:1047532022. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Zhang M, Wang F, Xiang Z, Huang T and Zhou
WB: LncRNA XIST promotes chemoresistance of breast cancer cells to
doxorubicin by sponging miR-200c-3p to upregulate ANLN. Clin Exp
Pharmacol Physiol. 47:1464–1472. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Alam F, Mezhal F, El Hasasna H, Nair VA,
Aravind SR, Saber Ayad M, El-Serafi A and bdel-Rahman WM: The role
of p53-microRNA 200-Moesin axis in invasion and drug resistance of
breast cancer cells. Tumour Biol. 39:10104283177146342017.
View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Song W, Wu S, Wu Q, Zhou L, Yu L, Zhu B
and Gong X: The microRNA-141-3p/CDK8 pathway regulates the
chemosensitivity of breast cancer cells to trastuzumab. J Cell
Biochem. 120:14095–14106. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Maierthaler M, Benner A, Hoffmeister M,
Surowy H, Jansen L, Knebel P, Chang-Claude J, Brenner H and
Burwinkel B: Plasma miR-122 and miR-200 family are prognostic
markers in colorectal cancer. Int J Cancer. 140:176–187. 2017.
View Article : Google Scholar
|
|
136
|
Prinz C, Frese R, Grams M and Fehring L:
Emerging role of microRNA dysregulation in diagnosis and prognosis
of extrahepatic cholangiocarcinoma. Genes (Basel). 13:14792022.
View Article : Google Scholar : PubMed/NCBI
|