Introduction
The insulin-like growth factor 1 receptor (IGF-1R)
is a tyrosine kinase (TK) receptor that plays a crucial role in
regulating various biological processes, such as cell growth,
differentiation and apoptosis (1-3).
Upon binding its ligand, IGF-1, IGF-1R activates downstream
signaling pathways that modulate cellular functions and contribute
to physiological processes, such as embryonic development, organ
formation and tissue regeneration (4-6).
Aberrant activation of the IGF-1R is closely associated with the
initiation and progression of various malignancies. For instance,
elevated IGF-1R expression has been strongly correlated with tumor
cell proliferation, invasion, metastasis and therapy resistance in
multiple solid tumors, including hepatocellular carcinoma (7), prostate cancer (8), ovarian cancer (9), laryngeal carcinoma (10) and breast cancer (11). Furthermore, IGF-1R promotes
malignant progression by modulating the tumor microenvironment,
specifically by positively correlating with the infiltration of
immune cells, including CD4+ T cells, dendritic cells
and macrophages (12) and
sustaining cancer stem cell properties (13).
Traditionally, IGF-1R has been viewed as a mediator
of signal transduction at the plasma membrane. However, studies
have suggested that after being internalized, IGF-1R can either be
recycled back to the cell surface, directed for degradation or
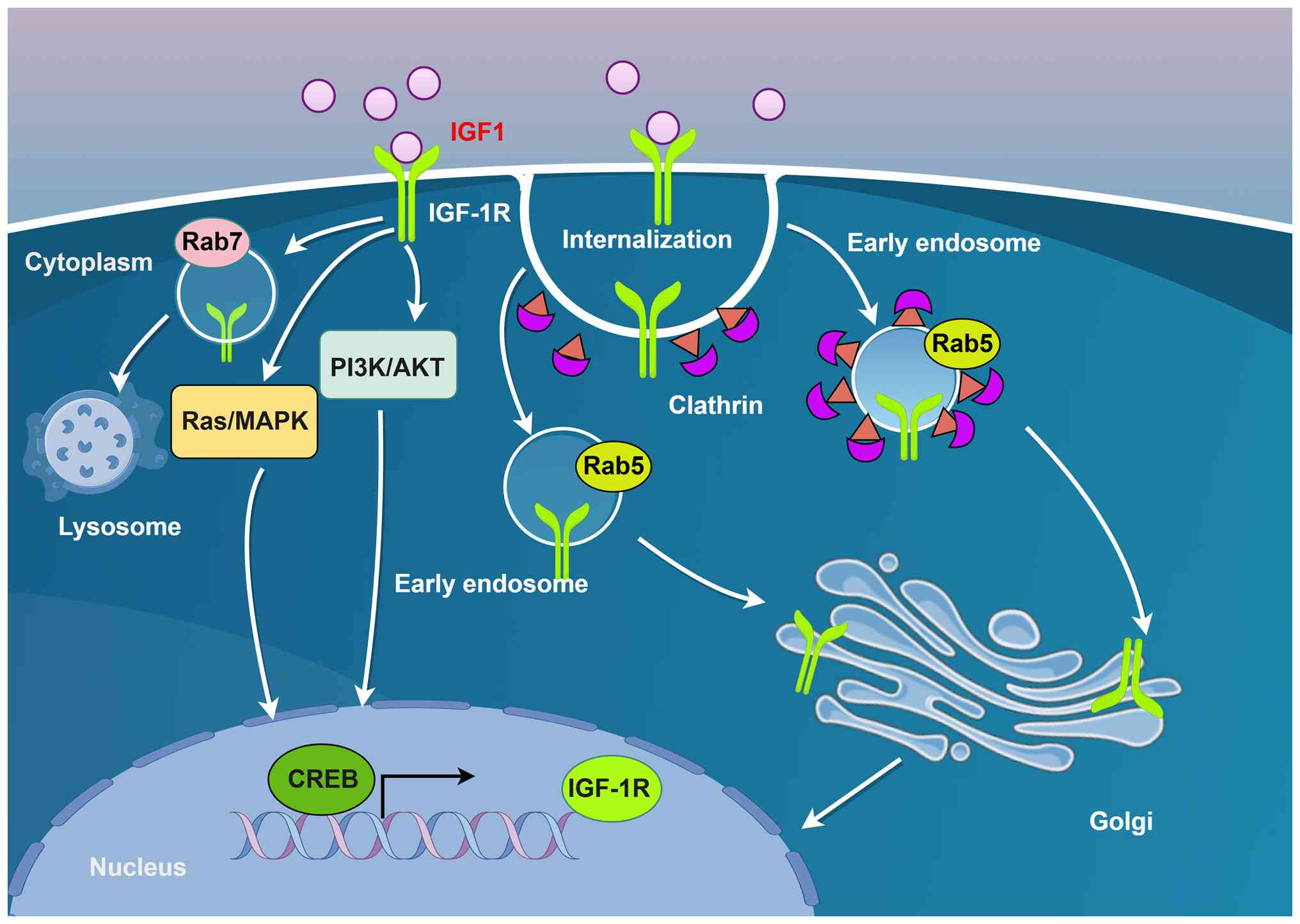
moved to specific intracellular compartments (14-18). The internalization and subsequent
localization of IGF-1R to various subcellular regions, such as the
Golgi apparatus, nucleus and mitochondria, exert distinct and
critical regulatory effects on tumor behavior (15,16). Notably, IGF-1R accumulation in the
Golgi apparatus is associated with enhanced tumor cell migration,
whereas its nuclear expression is strongly correlated with
increased invasiveness and poor clinical prognosis (17,18). Furthermore, compartmentalized
signaling of IGF-1R plays a role in regulating various cellular
processes, such as gene expression and mitochondrial function,
revealing new mechanisms that contribute to tumor progression and
resistance to therapy (19,20).
Emerging evidence underscores the subcellular
localization of IGF-1R as a novel predictive biomarker of
therapeutic response, providing a compelling rationale for its
integration into precision oncology frameworks to optimize targeted
therapy (16). Although initial
clinical trials of IGF-1R monotherapy have yielded limited
efficacy, recent insights into its compartmentalized signaling have
revitalized the therapeutic landscape, prompting the investigation
of synergistic combination strategies (17-19). Co-targeting IGF-1R with
immunotherapy, chemotherapy or other targeted agents has
demonstrated improved outcomes in preclinical and clinical settings
(1,21). In conclusion, IGF-1R contributes
crucially to tumor pathogenesis via canonical plasma membrane
signaling and through internalization and compartment-specific
signaling, which critically influences malignant progression and
treatment resistance. The present study provides a comprehensive
review of the molecular mechanisms that govern IGF-1R
internalization and compartmental signaling, explores their
implications in tumor progression and therapeutic resistance, and
integrates recent advancements in the field, thereby providing a
theoretical foundation and a novel perspective for precision cancer
therapy.
IGF-1R-related signaling pathway
Literature search strategy
A comprehensive and systematic literature search was
conducted to identify relevant studies on the internalization,
compartmentalization and signaling of IGF-1R in cancer cells. The
search was performed across multiple electronic databases,
including PubMed (https://pubmed.ncbi.nlm.nih.gov/), Web of Science
(https://www.webofscience.com/) and
ScienceDirect (https://www.sciencedirect.com/), from their inception
in September 2025. To ensure the inclusion of the most recent and
comprehensive evidence, Google Scholar (https://scholar.google.com/) was also used for
supplementary searches, minimizing the risk of publication bias.
The search strategy employed a combination of key words and Medical
Subject Headings related to the core themes. The primary search
terms included: ('Insulin-like growth factor 1 receptor' OR
'IGF-1R' OR 'IGF1R') AND ('Internalization' OR 'Subcellular
Localization' OR 'Compartmentalization') AND ('Cancer' OR 'Tumor')
AND ('Signaling' OR 'Pathway'). Boolean operators (AND and OR) were
used to combine these terms effectively. The following inclusion
criteria were applied: i) Studies focused on IGF-1R
internalization, subcellular localization, compartmentalized
signaling or related trafficking mechanisms in cancer contexts; ii)
articles providing mechanistic insights, functional data or
clinical correlations; and iii) publications in peer-reviewed
journals. Exclusion criteria included: i) Non-English articles; ii)
conference abstracts, editorials, case reports or studies without
full-text availability; and iii) studies not directly addressing
IGF-1R biology in malignancy. The initial search results were
screened based on titles and abstracts to select studies that
focused on the molecular mechanisms and biological functions of
IGF-1R internalization and compartmentalization in cancer. The full
texts of potentially eligible articles were retrieved and assessed
for inclusion. Reference lists of key reviews and eligible studies
were manually searched to identify additional relevant
publications. Only articles published in English were included.
Original research articles, authoritative reviews and significant
clinical trial reports were selected.
Molecular structure and functional
domains of IGF-1R
IGF-1R is a classic receptor TK (RTK), structurally
comprising two α subunits and two β subunits linked by disulfide
bonds to form a heterotetramer. Studies in structural biology have
shown that IGF-1R presents a symmetrical 'Λ' or 'U' conformation in
its unactivated state (22,23). The α subunit is entirely located
extracellularly and is primarily responsible for the recognition
and binding of ligands (such as IGF-1); its structure contains
multiple functional domains, including leucine-rich repeats (L1 and
L2), a cysteine-rich region (CR) and three fibronectin type III
(FnIII-1, FnIII-2 and FnIII-3) domains, which together constitute a
high-affinity binding site for IGF-1 (23). The ligand-binding sites of IGF-1R
are mainly distributed in the L1 and α-carboxyl terminal (CT) helix
regions of the α subunit, where these regions form multiple points
of contact with the IGF-1 ligand, ensuring high specificity of
binding (23,24). The CR domain and the FnIII-1
domain do not directly interact strongly with the ligand; they
assist in maintaining the spatial conformation of the binding
pocket and in transmitting conformational signals, thereby
enhancing the stability and specificity of ligand binding (14,23,24).
The β subunit spans the cell membrane and possesses
TK activity, capable of triggering downstream signaling pathways.
The extracellular portion is connected to the α subunit, while the
intracellular portion contains the juxtamembrane region, TK domain
and CT region (25). The TK
domain is the core of signal transduction, located in the
intracellular region of the β subunit, and contains a highly
conserved ATP-binding site and catalytic residues. Upon
receptor-ligand binding, conformational changes in the kinase
region lead to the phosphorylation of key tyrosine residues (such
as Y1131, Y1135 and Y1136), forming anchor sites for downstream
effector proteins (25,26). For instance, insulin receptor
substrate (IRS) proteins in the cytoplasm are recruited and bind to
the activated IGF-1R via their specific domains (such as the
pleckstrin homology and phosphotyrosine binding domains), which in
turn leads to the phosphorylation of multiple tyrosine residues on
IRS (26). Each phosphorylated
tyrosine site on phosphorylated IRS-1 can recruit downstream
signaling proteins with Src homology 2 (SH2) domains [such as PI3K
and growth factor receptor-bound protein 2/son of sevenless). Once
these proteins are recruited and bound, they initiate multiple
signaling cascades, including two major signaling pathways: The
PI3K-Akt and Shc-Ras-MAPK pathways. These signaling pathways play
critical roles in cell metabolism, proliferation and survival
(27,28) (Fig.
1). In summary, the structure and function of IGF-1R highlight
its important role in cell proliferation and survival, and the
activation of multiple signaling pathways indicates a critical role
of IGF-1R in tumor progression.
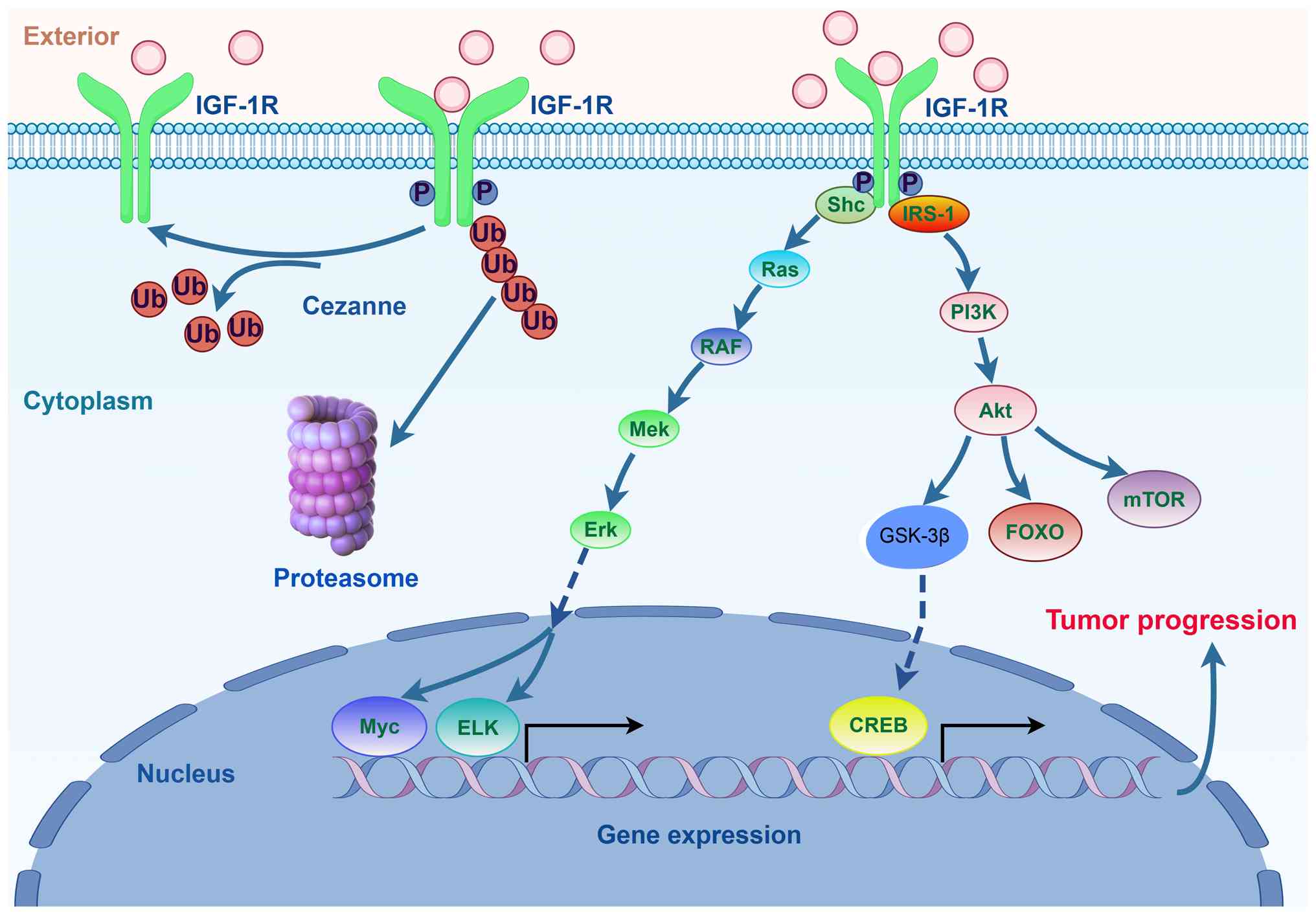 | Figure 1Schematic of the canonical IGF-1R
signaling pathway. Upon binding to its ligand, IGF-1, IGF-1R
activates major downstream signaling cascades, primarily the
PI3K-Akt and Shc-Ras-MAPK pathways. These pathways collectively
promote cell proliferation, survival and metabolic reprogramming,
while inhibiting apoptosis, thereby driving oncogenic processes
(created with Figdraw). IGF-1R, insulin-like growth factor 1
receptor; P, Phosphorylated; Ub, Ubiquitin; IRS-1, Insulin receptor
substrate 1; ELK, E twenty-six-like kinase 1. |
It has been reported that certain mutations in the
kinase domain of IGF-1R (such as K1055R) can significantly reduce
kinase activity but have limited effects on the subcellular
localization of the receptor and some non-classical signaling
functions, suggesting that the structural microenvironment of the
kinase activity center finely regulates receptor function (26). Additionally, the kinase region of
IGF-1R interacts with various regulatory proteins (such as Ras
GTPase-activating-like protein IQGAP1) and is associated with
glycosylation modifications that affect receptor stability and
signal output (29). These
structural characteristics determine the signaling capability of
IGF-1R and provide a theoretical basis for targeted drug design. In
conclusion, as a heterotetrameric RTK, the clearly defined
structure and functional domain division of its α and β subunits,
along with the precise regulation of ligand-binding sites and
kinase domains, collectively determine the activation and signaling
output characteristics of the receptor. A deeper understanding of
the structure and functional domains of IGF-1R provides a solid
molecular foundation for elucidating its signal transduction
mechanism and targeted interventions.
Expression of IGF-1R in various tumor
tissues
A comprehensive analysis of the IGF-1R expression
patterns across 33 tumor types using the University of California
Santa Cruz XENA tool (https://xenabrowser.net/datapages/) (30) was performed. Among the 33 tumor
types, the expression of IGF-1R was compared between pan-cancer
cohorts from The Cancer Genome Atlas (https://portal.gdc.cancer.gov) and matched normal
tissues from patients in the Genotype-Tissue Expression project
(https://gtexportal.org/home/) using the
Mann-Whitney U test. The R software and the R packages
ggplot2[3.4.4], stats[4.2.1] and car[3.1-0] (R Core Team; R
Foundation for Statistical Computing) were employed, with P<0.05
considered to indicate a statistically significant difference. The
results revealed that IGF-1R expression was significantly
downregulated in adrenocortical carcinoma, bladder urothelial
carcinoma, colon adenocarcinoma, lymphoid neoplasm diffuse large
B-cell lymphoma, glioblastoma multiforme, liver hepatocellular
carcinoma, lung adenocarcinoma, ovarian serous cystadenocarcinoma,
rectum adenocarcinoma, testicular germ cell tumor, thyroid
carcinoma, uterine corpus endometrial carcinoma and uterine
carcinosarcoma compared with that in normal tissues. Conversely,
significant upregulation was observed in breast invasive carcinoma,
cholangiocarcinoma, esophageal carcinoma, head and neck squamous
cell carcinoma, acute myeloid leukemia, brain lower grade glioma,
lung squamous cell carcinoma, pancreatic adenocarcinoma, prostate
adenocarcinoma, skin cutaneous melanoma and thymoma tissues
compared with that in corresponding normal controls (Fig. 2). These findings highlight the
cancer type-specific nature of IGF-1R expression and suggest that
its levels and activity are dynamically regulated across different
tissues and pathological contexts, thereby influencing diverse
biological functions. This variability was further supported by
evidence from specific malignancies, such as myxoid liposarcoma,
where elevated IGF-1R expression was associated with poor
metastasis-free survival, underscoring its role in tumor
pathogenesis (31).
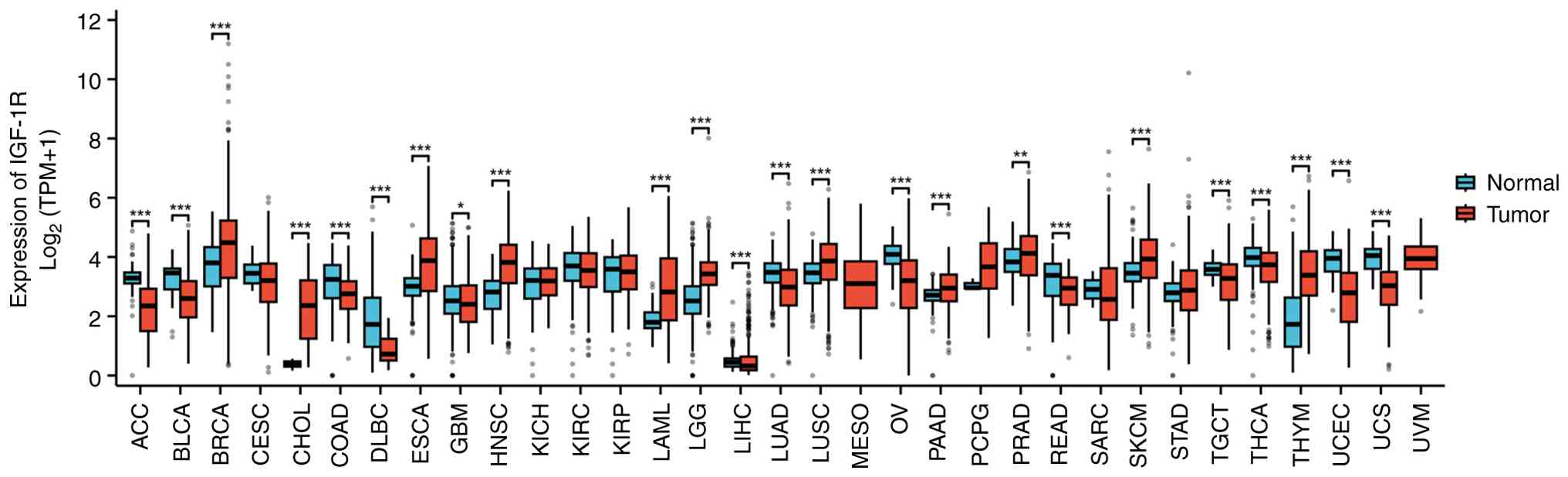 | Figure 2Pan-cancer analysis of IGF-1R
expression across tumor and normal tissues. The expression levels
of IGF-1R (in TPM) were compared between tumor samples from The
Cancer Genome Atlas pan-cancer cohorts and matched normal tissue
samples from the Genotype-Tissue Expression project. Data for 33
cancer types were uniformly processed and obtained from the
University of California Santa Cruz XENA database. Statistical
significances between tumor and adjacent normal groups for each
cancer type were assessed using the non-parametric Mann-Whitney U
test (Wilcoxon rank-sum test); *P<0.05,
**P<0.01 and ***P<0.001. CESC, cervical
squamous cell carcinoma and endocervical adenocarcinoma; IGF-1R,
insulin-like growth factor 1 receptor; KICH, kidney chromophobe;
KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal
papillary cell carcinoma; MESO, mesothelioma; PCPG,
pheochromocytoma and paraganglioma; SARC, sarcoma; STAD, stomach
adenocarcinoma; TPM, transcripts per million; UVM, uveal
melanoma. |
Multilayered regulatory network of IGF-1R
as a central oncogenic driver
Emerging evidence indicates that the functional role
of IGF-1R extends beyond classical cell membrane signaling, with
its nuclear localization garnering increasing attention. For
instance, IGF-1R can undergo nuclear translocation and participate
in the regulation of gene transcription, suggesting a potential
mechanism for tumor progression (27). Furthermore, the internalization,
intracellular trafficking and sublocalization of IGF-1R have been
shown to significantly influence signaling dynamics. These
processes may lead to distinct biological outcomes, thereby
impacting cellular behavior and oncogenesis (26,28). Therefore, elucidating the detailed
mechanisms of IGF-1R localization and trafficking is essential for
deepening our understanding of its biological functions and for
identifying novel therapeutic targets for cancer treatment.
IGF-1R signaling serves as a central oncogenic
driver in multiple malignancies, including colorectal, breast,
lung, ovarian and endometrial cancer; its aberrant activation,
often induced by ligand overexpression (e.g., IGF2), environmental
factors (e.g., nicotine) or cues from the tumor microenvironment
[e.g., tumor-associated macrophage (TAM)-derived C-X-C motif
chemokine ligand 1 (CXCL1) or collagen], orchestrates an extensive
downstream signaling network (32-44). Key pathways regulated by IGF-1R
include PI3K/AKT/mTOR (32-35), Elk1/AP1/Myc (36), STAT3/HMGB1 (37-39), NuMA/53BP1 (40) and forkhead box P3
(FOXP3)/β-catenin (41).
Collectively, these pathways facilitate critical oncogenic
processes, such as tumor proliferation (42), metastasis (43), stemness maintenance (44), metabolic reprogramming via
glycolysis (45), therapy
resistance (46-49) and immune evasion (50). For instance, TEA domain
transcription factor 4 transcriptionally upregulates RNA-binding
motif protein 8A (RBM8A), which partners with eukaryotic initiation
factor-4A3 to stabilize the IGF-1R and IRS-2 mRNAs, thereby
activating the PI3K/AKT signaling pathway and promoting breast
cancer progression. This RBM8A-mediated post-transcriptional
regulation represents a novel mechanism underlying IGF-1R-driven
oncogenesis (32). G-protein
coupled receptor-associated sorting protein 1 (GASP1) stabilizes
IGF-1R by competitively inhibiting MDM2-mediated ubiquitination and
activating the downstream NF-κB, PI3K/AKT and MAPK/ERK pathways to
drive breast cancer progression. This GASP1-IGF-1R signaling loop
creates a vicious cycle that enhances malignant phenotypes and
reduces paclitaxel sensitivity, suggesting the potential for dual
therapeutic targeting (51).
Metformin inhibits endometrial cancer cell viability in
vitro by inducing cytotoxicity and cell cycle arrest, while,
in vivo, it suppresses tumor growth and downregulates key
components of the IGF-1R and PI3K/AKT/mTOR signaling pathways. The
drug's antitumor effects involve the modulation of multiple
targets, including mTOR and MAPK3, suggesting a complex mechanism
of action beyond direct IGF-1 level reduction (52). Elevated FOXP3 expression promotes
breast cancer progression by directly binding to the β-catenin
promoter and enhancing its transcription. IGF-1R signaling
activates this FOXP3-β-catenin axis, suggesting a novel therapeutic
target that is effectively inhibited by the compound elesclomol
(41). Monensin suppresses
colorectal cancer proliferation and migration by elevating IGF1 to
inhibit IGF-1R signaling and concurrently targeting multiple
oncogenic pathways, including Elk1/AP1/Myc. This antibiotic
demonstrates repurposing potential as an IGF-1R-axis-targeting
agent in colorectal cancer therapy (36). Nicotine upregulates cholinergic
receptor nicotinic α9 subunit and IGF-1R expression to promote
cancer stemness, migration and metastasis in triple-negative breast
cancer (TNBC). Targeting IGF-1R signaling suppresses
nicotine-induced malignancy and represents a therapeutic strategy
for high-risk patients with TNBC (53). Lactate-induced lactylation
stabilizes the IGF-1R protein and enhances its binding to IGF1,
thereby promoting glycolysis and proliferation in lung cancer
cells. This metabolic reprogramming establishes a feed-forward loop
in which IGF-1R-driven lactate production further amplifies its own
oncogenic signaling (45).
TAM-secreted CXCL1 promotes autophagic chemoresistance in breast
cancer by stabilizing the IGF-1R through suppressed von
Hippel-Lindau (VHL)-mediated ubiquitination, subsequently
activating STAT3/HMGB1 signaling. Targeting the CXCL1/IGF-1R axis
reverses paclitaxel resistance and correlates with improved
clinical outcomes (37).
Additionally, p52-zinc-finger estrogen receptor interaction clone 6
(ZER6) maintains cancer stem cell populations by transcriptionally
upregulating IGF-1R to enhance pro-survival mitophagy,
distinguishing its function from that of the p71-ZER6 isoform.
Targeting this axis represents a promising therapeutic strategy to
exhaust cancer stem cells (CSCs) and inhibit tumor progression
(44).
Furthermore, IGF-1R activity is fine-tuned through
multi-faceted regulatory mechanisms, including transcriptional
control (e.g., by SOX4), post-translational modifications (e.g.,
VHL-mediated ubiquitination) and non-coding RNA networks (e.g.,
circular (circ)RNA/microRNA (miR/miRNA)-mediated competing
endogenous RNA mechanisms), underscoring its role as a dynamic
signaling hub (Fig. 3) (54). Specifically, some miRNAs (Table SI), including miR-770-5p,
miR-376a-3p, miR-133b, miR-99b-5p, miR-144, miR-98-5p, miR-520b,
miR-99a, miR-15b and miR-194-5p, have also been reported to
regulate corresponding downstream signaling pathways by interfering
with IGF-1R expression, thereby exhibiting anticancer effects or
promoting chemotherapy sensitivity (35,46,47,55-61). In addition, long non-coding RNAs
that regulate IGF-1R (Table SII)
include MLETA1, LINC02381, HULC, OIP5-AS1, MIAT, LINC01287, SNHG11,
AFAP1-AS1, CASC11, LINC00324, Linc00210, LINC00958, LINC00319 and
LINC00339 (11,62-74), and circRNAs (Table SIII) including Circ_0006174,
Circ_PPAPDC1A, Circ-IGF-1R, CircVAPA, Circ_0067835, CircRNF13,
Circ_0002577, CircGNB1 and Circ_0006089 (75-83) promote the expression or stability
of IGF-1R by inhibiting miRNAs.
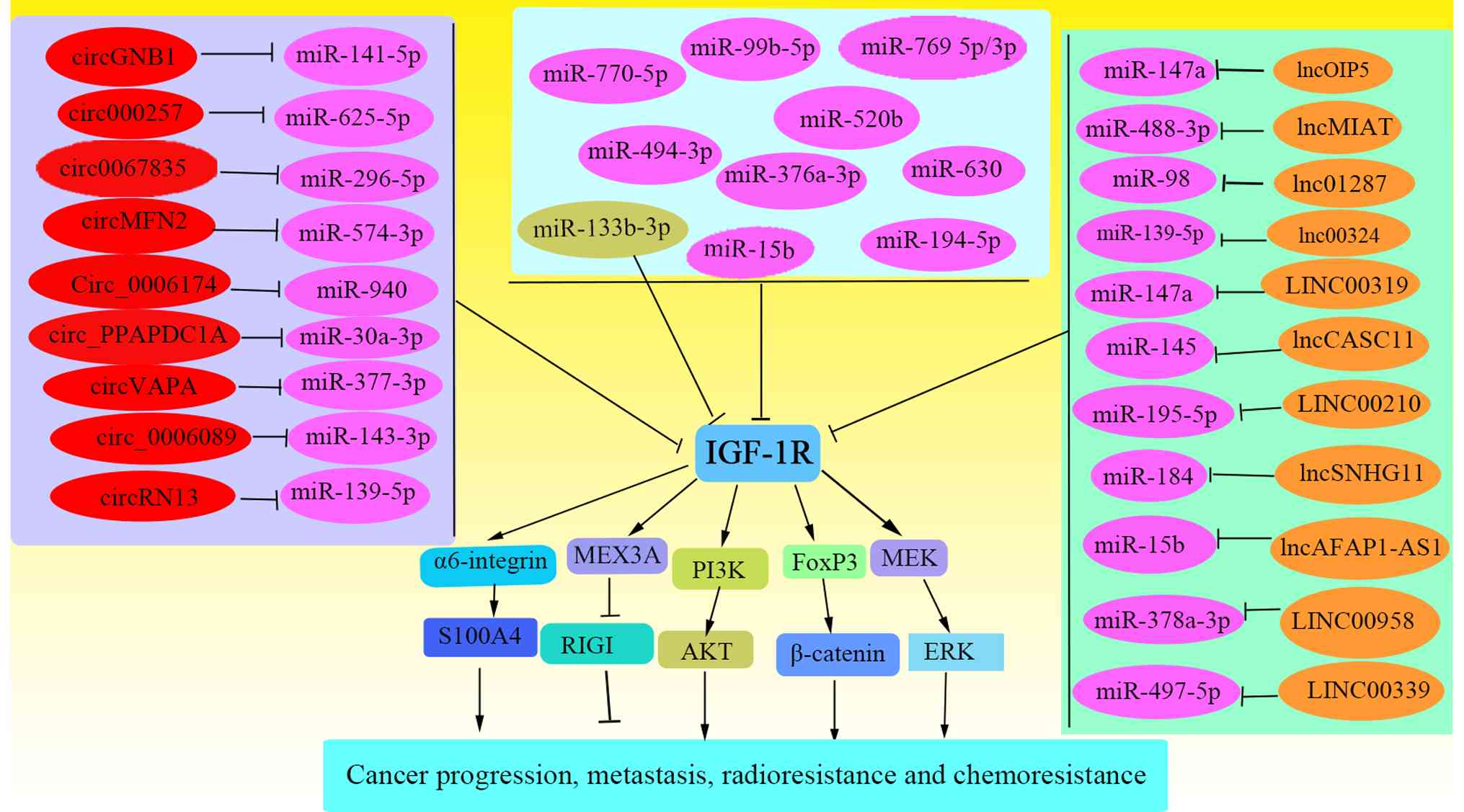 | Figure 3ceRNA network regulating
IGF-1R-mediated cancer phenotypes. A conceptual model illustrating
how ncRNAs, including lncRNAs and circRNAs, can function as
molecular sponges to sequester miRNAs that target IGF-1R
mRNA. Through this ceRNA mechanism, these ncRNAs indirectly
upregulate IGF-1R expression and activity, thereby influencing key
cancer hallmarks, such as tumor progression, metastasis and
therapeutic resistance to radiotherapy and chemotherapy. ceRNA,
competing endogenous RNA; IGF-1R, insulin-like growth factor 1
receptor; ncRNAs, non-coding RNAs; lncRNAs, long non-coding RNAs;
circRNA, circular RNAs; miRs/miRNAs, microRNAs; MEX3A, Mex-3 RNA
binding family member A; RIGI, antiviral innate immune response
receptor RIG-I. |
Molecular mechanisms of IGF-1R
internalization and compartmentalized signaling
Emerging research has fundamentally shifted the
traditional paradigm, revealing that IGF-1R internalization is not
an endpoint, but a critical regulatory node that diversifies
signaling outputs through subsequent compartmentalization (15-18). This review section delineates the
molecular journey from endocytosis to functional engagement within
distinct subcellular locations, providing a mechanistic basis for
context-dependent signaling.
Mechanism for the initiation of
ligand-dependent endocytosis
The ligand-dependent endocytosis of IGF-1R begins
with the binding of IGF-1 to the receptor, which induces a
conformational change in the receptor's structure. This triggers a
spatial rearrangement of the extracellular domain of IGF-1R and
promotes receptor dimerization and exposure of its
autophosphorylation active sites, thereby activating its TK
function (14,84). This conformational change marks
the initiation of signal transduction and is a crucial prerequisite
for endocytosis. Receptor endocytosis and subsequent subcellular
localization are critical for determining specific signaling
responses. For example, IGF-1R is regulated by cell adhesion
receptors and endocytosis-related proteins in different cell types,
indicating that endocytosis has a high degree of cell specificity
(14).
Receptor phosphorylation is the second most
important event in ligand-dependent endocytosis. After ligand
binding, the intracellular tyrosine residues of IGF-1R are rapidly
phosphorylated. This phosphorylation activates downstream signaling
pathways and provides molecular recognition sites for the
recruitment of endocytosis-related proteins. Specifically,
phosphorylated IGF-1R can attract various endocytic adaptor
proteins, among which the adaptor-related protein complex 2 subunit
α1/2 (AP2A1/2) complex and flotillin-1 (Flot-1) are core adaptor
molecules that mediate the classical clathrin-dependent and
noncaveolar lipid raft-associated endocytic pathways, respectively
(84). It has been reported that
AP2A1/2 dominates receptor endocytosis at high IGF-1
concentrations, whereas Flot-1 is more sensitive at low IGF-1
concentrations. Dual knockdown of both proteins significantly
inhibits endocytosis, indicating their functional synergy (84). Additionally, the palmitoylation
modification of Flot-1 dynamically regulates its function, with
acyl protein thioesterases-1 (APT-1) and palmitoyltransferase
ZDHHC19 (ZDHHC19) responsible for its depalmitoylation and
repalmitoylation, maintaining its cycling and function, thereby
preventing IGF-1R desensitization and degradation while promoting
sustained receptor signaling output (85). The cooperation of these proteins
determines the efficiency of IGF-1R endocytosis and affects
subsequent signal partitioning and downstream biological effects of
the receptor. Thus, the dynamic regulation of endocytosis-related
proteins provides a molecular basis for the adaptive response of
cells to IGF-1 signaling, and the selectivity of different
endocytic pathways may be closely linked to changes in the cell
type and microenvironment.
Internalization: Pathways, trafficking
and fate decisions
IGF-1R initiates signal transduction at the plasma
membrane under ligand stimulation and undergoes endocytosis,
entering different compartments within the cell. Studies have found
that IGF-1R forms specific signaling complexes in endosomes,
providing spatial specificity to signal transduction (14,86). Using methods such as confocal
microscopy and immunoprecipitation, it has been confirmed that
IGF-1R can enter Rab5-positive early endosomes via
clathrin-mediated endocytosis, where IGF-1R can form active
complexes with downstream signaling proteins (such as IRS and AKT),
maintaining continuous signal transmission. Notably, the
endocytosis and transport of IGF-1R in different cell types are
regulated by cell adhesion receptors and related proteins,
affecting the specificity of the signals (14). In intestinal epithelial cell
models, IGF-1R, after clathrin-dependent endocytosis, enters
Rab5-positive endosomes and is transported to the Golgi apparatus
with the help of the dynein-microtubule system, laying the
groundwork for subsequent signal compartmentalization and nuclear
transport (86) (Fig. 4). This process reveals that
endosomes are sites for signal termination and key nodes for the
continuous regulation and compartmentalization of signals.
Furthermore, the signaling activity of IGF-1R in
endosomes is closely related to its subsequent sorting. Some
IGF-1Rs in early endosomes are recycled to the plasma membrane,
maintaining the cell's sensitivity to IGF-1, while another portion
is transported to the lysosomes for degradation, terminating the
signal (14). Thus, once
internalized, IGF-1R may undergo one of three main outcomes:
Dephosphorylation, degradation through ubiquitination or recycling
(87). The internalization and
subsequent dephosphorylation of IGF-1R play critical roles in
signal termination and prevention of aberrant activation (88). The precise regulation of these
processes has emerged as an important area of research in cancer
and metabolic diseases. These mechanisms suggest that receptor
signaling activation in endosomes results from a balance between
receptor degradation and recycling, and this balance underlies the
cell's finely tuned response to external stimuli.
i) Clathrin-mediated endocytosis mechanism. The
endocytosis of IGF-1R is mainly mediated by the clathrin-mediated
endocytic pathway, a process that is far from a simple signal
termination mechanism, but rather initiates a precisely regulated
intracellular journey and becomes a critical hub determining
receptor signal partitioning and downstream biological effects
(86,87). It has been reported that, in
various cell types, after IGF-1 binds to the receptor, the
activated IGF-1R interacts with clathrin and its core adaptor
protein complex (such as AP2A1/2) and is encapsulated into
clathrin-coated vesicles, thus completing endocytosis. This pathway
is particularly pronounced under stimulation at high concentrations
of IGF-1 (84).
After entering the cell, the receptor is sorted into
the Rab5-positive early endosome, which is the first decision point
in the intracellular transport pathway. The key transport
regulatory factors determine the subsequent destination.
Specifically, the transport system relies on the microtubule
network, particularly for retrograde transport, mediated by the
dynein activator complex subunit p150Glued (89). This system transports receptors
back to the perinuclear region. A critical step for the receptor's
entry into the nucleus depends on the classical nuclear transport
mechanism: The receptor interacts with the nuclear transport
receptor importin β at the nuclear pore and ultimately crosses the
pore under the regulation of the nuclear pore protein NUP358
(RanBP2) (86). Notably, small
ubiquitin-related modifier 1 (SUMO1) modification of IGF-1R is a
key driving factor in this process, which is catalyzed by SUMO E3
ligases, such as RanBP2, and is crucial for stabilizing the
receptor and promoting its interaction with the nuclear pore
complex (89). Specifically,
IGF-1 stimulation can induce SUMO1 modification on three conserved
lysine residues (Lys1025, Lys1100 and Lys1120) of the IGF-1R β
subunit. This modification is a prerequisite for nuclear
translocation, and mutations at these sites completely block
nuclear entry without affecting the classical kinase signaling
pathway (90). In a number of
cancer types (such as breast and colorectal cancer), IGF-1R has
been observed to accumulate in the nucleus. This is closely related
to increased tumor malignancy and a poor prognosis, suggesting that
inhibiting its nuclear translocation may be a potential therapeutic
strategy (14).
By contrast, the pathway directing receptor
degradation is controlled by the GTPase Rab7a. Rab7a guides the
maturation of early endosomes to late endosomes/lysosomes,
ultimately leading to receptor degradation (91,92). Inhibiting the function of Rab7a
significantly enhances IGF-1R stability and prolongs its downstream
signaling, highlighting the core role of this molecule in the
precise regulation of receptor turnover (93).
ii) Non-clathrin-mediated endocytic pathways and
their molecular basis. In addition to the classical
clathrin-mediated mechanism, IGF-1R can also mediate intracellular
transport through non-clathrin-mediated endocytic pathways (such as
caveolae- and macropinocytosis-mediated pathways), where Flot-1
plays a central role. It has been reported that the dynamic
regulation of Flot-1 palmitoylation affects IGF-1R sensitivity and
degradation, preventing receptor desensitization and lysosomal
degradation (85). Specifically,
the palmitoylation of Flot-1 is co-regulated by APT-1
depalmitoylating enzyme and ZDHHC19 palmitoyl transferase,
modulating the membrane localization and functional activity of
Flot-1. The dynamic changes in this modification can prevent
excessive degradation of IGF-1R after endocytosis, maintain the
receptor's continuous activation state, and particularly promote
malignant phenotypes such as epithelial-mesenchymal transition,
migration and invasion in tumor cells (85). Furthermore, the endocytosis of
IGF-1R is also regulated by cell adhesion molecules, with β1
integrin (ITGB1) enhancing receptor endocytosis and Golgi apparatus
accumulation by promoting the phosphorylation of the C-terminal
Tyr1250/1251 sites of IGF-1R, thereby initiating specific
intracellular signaling distinct from the insulin receptor
(18,94). In migratory cancer cells, IGF-1R
can localize to the Golgi apparatus and continuously activate
signaling pathways, enhancing cell migration ability (15-18). Considering the high expression of
Flot-1 and its associated enzymes, which are closely related to
cancer progression (85),
interventions targeting this pathway are expected to provide new
strategies for antitumor therapy. In summary, the endocytic
mechanisms of IGF-1R exhibit diversity and high plasticity, and the
synergy and complementarity between different endocytic pathways
assure the fine regulation of receptor signals.
iii) Post-endocytic trafficking and fate
determination. The fate of the internalized IGF-1R represents a
critical branch point in signal regulation, culminating in three
primary outcomes: Recycling back to the membrane following
dephosphorylation by protein tyrosine phosphatases (PTPs)
(including PTP1B and SHP2), targeted destruction via
ubiquitination-mediated degradation and compartmental sequestration
of organelles, such as the Golgi apparatus or nucleus. Regulatory
inputs, including ITGB1-mediated phosphorylation at Tyr1250/1251,
significantly bias this fate decision by modulating receptor
accumulation in specific subcellular compartments, a process
distinct from that involving the insulin receptor (18,94). Nuclear translocation requires
interaction with nuclear pore complex components (e.g., NUP358) to
facilitate receptor transport into the nucleus (95,96).
Dysregulation of these finely balanced fate
decisions, a common hallmark of cancer, results in sustained IGF-1R
signaling and ultimately promotes malignant progression. This can
manifest as failed degradation and as aberrant receptor
accumulation on the plasma membrane, as observed with IGFBP7 in
acute lymphoblastic leukemia, which prolongs surface residency to
enhance pro-survival signaling (97).
Compartmentalized signaling and its
functional outputs
The spatial segregation of internalized IGF-1R
confers distinct signaling capabilities, profoundly influencing
tumor cell behavior and therapeutic response. This
compartmentalized signaling is a key mechanism behind the
functional pleiotropy of IGF-1R.
Golgi apparatus. IGF-1R localized to the Golgi
complex is actively involved in promoting cell migration and
invasion (3,98). This accumulation is a hallmark of
migratory cancer cells and is regulated by phosphorylation at
Tyr1250/1251, often downstream of integrin signaling (18,94). From this perinuclear location,
IGF-1R activates signaling pathways that facilitate cytoskeletal
reorganization and support invasive behavior (18). New cell adhesion events can
dynamically shift Golgi complex-accumulated IGF-1R back to the
plasma membrane, illustrating a sophisticated feedback loop between
migration and adhesion (18,94).
Nucleus. Emerging research solidifies the direct
involvement of nuclear IGF-1R in orchestrating the DNA damage
response (DDR), a key mechanism underlying therapy resistance
(99). IGF-1R modulates DDR
through both cytoplasmic signaling and direct nuclear actions.
Cytoplasmically, IGF-1R-activated PI3K/AKT and MAPK/ERK pathways
can upregulate the expression of DNA repair scaffold proteins, such
as X-ray repair cross-complement group 1, which is crucial for base
excision repair (100,101). Consequently, inhibition of these
pathways can attenuate repair capacity and enhance
chemosensitivity. More directly, IGF-1R translocates to the nucleus
via mechanisms involving SUMOylation (102). Once inside, it influences DDR in
two primary ways: First, by binding to enhancer regions and
regulating the transcription of genes involved in repair and
survival; and second, by physically facilitating the repair process
itself. Functional studies in colorectal cancer models provide
direct evidence, as depletion or inhibition of IGF-1R markedly
sensitizes cells to radiation. This sensitization is
mechanistically linked to a significant reduction in the formation
of RAD51 and 53BP1 foci, key markers of homologous recombination
and non-homologous end joining repair, respectively, following
irradiation. This impairment in the repair complex assembly leads
to an accumulation of unrepaired DNA damage and enhanced
G2/M-phase cell cycle arrest (99). Therefore, nuclear IGF-1R acts both
as a transcriptional regulator and as a direct coordinator of DNA
repair machinery, making it a central player in conferring
resistance to radiotherapy and genotoxic chemotherapies.
Endosomes/lysosomes. Signaling from endosomal
compartments can prolong pathway activation, while lysosomal
degradation represents a definitive signal termination mechanism
(103). The balance between
these outcomes is critically regulated by Rab GTPases (e.g., Rab5
and Rab7a) and the ubiquitination status (104,105).
Mitochondrial crosstalk. Through the PI3K-AKT axis,
IGF-1R signaling indirectly promotes mitophagic clearance and
reduces reactive oxygen species, supporting metabolic adaptation
and cell survival (106-108).
This compartmentalized signaling network ensures
that IGF-1R can elicit appropriate biological responses, such as
proliferation, migration, DNA repair or metabolic adaptation,
depending on its subcellular localization and the cellular context
(109,110). In summary, the internalization
and compartmentalization of IGF-1R constitute a sophisticated
spatial regulatory system. By directing the receptor to specific
organelles, tumor cells harness distinct signaling outputs to drive
migration, enhance survival, repair DNA damage and adapt
metabolically, thereby fueling progression and evading therapies.
This refined understanding moves beyond the membrane-centric view
and reveals novel vulnerabilities for therapeutic intervention,
such as targeting specific endocytic adaptors (Flot-1 and Cezanne)
or disrupting the nuclear translocation machinery.
Immune regulation by IGF-1R and the tumor
microenvironment
The internalization and compartmentalized signaling
of IGF-1R influence the biological behavior of tumor cells and
profoundly shape the adaptability and immune-modulatory capacity of
the tumor microenvironment (TME) (111,112). Following internalization, IGF-1R
can sustain the activation of downstream signaling pathways, such
as PI3K/AKT/mTOR and MAPK/ERK, from subcellular compartments,
including endosomes and lysosomes (113). This enhances the adaptive
responses of tumor cells to stressors and therapeutic pressures
within the TME. Compartmentalized IGF-1R signaling modulates how
tumor cells respond to classic TME features, such as metabolic
stress, hypoxia and acidosis, while also influencing the
recruitment and differentiation of immune cells. For example,
aberrant activation of mTOR signaling promotes the metabolic
reprogramming of tumor cells and regulates immune cell metabolism
and function, thereby fostering an immunosuppressive
microenvironment (111). For
instance, IGF-1R activation promotes colorectal tumor growth
through a non-canonical β-arrestin-2/Mex-3 RNA binding family
member A pathway that degrades antiviral innate immune response
receptor RIG-I and suppresses type I interferon responses in the
tumor microenvironment. Targeting this immune evasion mechanism
restores sensitivity to checkpoint inhibitors, revealing a novel
therapeutic strategy for IGF-1R-driven cancers (50). Inhibition of IGF2-IGF-1R signaling
with PQ401 delays the growth of IGF2-high colorectal cancer cells
by modulating myeloid-derived suppressor cells (MDSCs), which
enhance T cell-mediated antitumor immunity. PQ401 treatment reduces
the suppressive function and recruitment of MDSCs, leading to
increased infiltration and activity of CD4(+) and CD8(+) T cells,
thereby inhibiting tumor growth in IGF2-high colorectal cancer
(114). Collagen-rich tumor
microenvironments in TNBC upregulate the IGF-1R via SOX4 and DDR1,
promoting immunosuppression and T-cell exhaustion. Targeting IGF-1R
reverses this cold phenotype and synergizes with anti-programmed
cell death protein 1 (PD-1) therapy to enhance antitumor immunity
(115). Combined IGF-1R
inhibition and PD-1 blockade synergistically enhance dendritic cell
maturation and CD8(+) T cell-mediated antitumor immunity in
epithelial ovarian cancer. This dual targeting strategy reduces the
tumor burden more effectively than monotherapy by remodeling the
immunosuppressive tumor microenvironment (116). Glucosidase 2 subunit β (PRKCSH)
stabilizes IGF-1R and enhances its oncogenic signaling, thereby
conferring resistance to tumor necrosis factor superfamily
(TNFSF)-mediated apoptosis in lung cancer. Targeting PRKCSH
sensitizes tumors to natural killer (NK) cell-mediated
cytotoxicity, suggesting a combined therapeutic approach for
TNFSF-resistant malignancies (49). Furthermore, tumor cells facilitate
the recruitment and functional enhancement of immunosuppressive
cells through exosome-mediated transfer of signaling molecules,
while simultaneously inhibiting the antitumor response of effector
immune cells. This process ultimately promotes tumor progression
and immune escape (117). IGF-1R
promotes lung metastatic implantation and progression by remodeling
the tumor microenvironment through enhanced vascularization,
inflammation and immunosuppression. Host IGF-1R deficiency
significantly attenuates the metastatic burden and alters multiple
stromal components, positioning IGF-1R as both a therapeutic target
and potential biomarker in non-small cell lung cancer metastasis
(118). In summary, the
internalization and compartmentalized signaling of IGF-1R represent
a key adaptive mechanism through which tumor cells respond to
microenvironmental changes and serve as a crucial determinant of
immune regulation and treatment response within the TME. These
insights provide a strong theoretical foundation for the
development of novel therapeutic strategies that target TME
adaptation and immunomodulation.
For the present review, a correlation analysis was
performed between IGF-1R expression and immune infiltration
profiles across pan-cancer datasets from The Cancer Genome Atlas
(TCGA) database. Immune cell infiltration levels were quantified
via the single-sample Gene Set Enrichment Analysis algorithm
implemented in the R package GSVA (v1.46.0) (119), based on a curated panel of 24
immune cell-specific gene sets described by Bindea et al
(120). Spearman's rank
correlation test was applied between IGF-1R expression
(ENSG00000140443.15) and the infiltration scores of each immune
subset. The analysis was conducted using R software (v4.2.1; R
Foundation for Statistical Computing) with the ggplot2 package
(v3.4.4) for visualization. Correlation coefficients and
corresponding P-values were calculated for all immune cell types
across TCGA cancer cohorts. Data are presented as Spearman's rho
(ρ) in the heatmap (Fig. 5;
Tables SIV and SV), with statistical significance
defined as a two-sided P-value of <0.05. The results, visualized
by a heatmap (Fig. 5; Tables SIV and SV), revealed a positive correlation
between the IGF-1R and T central memory (Tcm) cells and significant
negative correlations with plasmacytoid dendritic cells (pDCs), T
cells, NK CD56bright cells, NK CD56dim cells, and T helper 1 (Th1)
cells. These findings indicate that the IGF-1R signaling pathway
plays a key regulatory role in the TME. Although other immune
subsets (e.g., general T helper cells) also show correlations in
the heatmap, the selected populations exhibited the most robust and
reproducible associations with IGF-1R expression, supporting their
potential mechanistic and translational relevance. The effect of
IGF-1R signaling on immune cells is primarily reflected in the
regulation of their differentiation, recruitment and functional
states. For instance, IGF-1R activation can promote the
accumulation of immunosuppressive cells, such as regulatory T cells
and MDSCs, while inhibiting the antitumor activity of effector T
cells and NK cells, thereby facilitating immune evasion by tumor
cells. Furthermore, various signaling molecules within the TME,
including growth factors, lipid metabolites and metal ions,
interact with the IGF-1R pathway to modulate the functional state
of tumor-associated immune cells. For example, the PI3K/AKT/mTOR
signaling axis, a key downstream pathway of IGF-1R, regulates
immune cell differentiation, metabolism and effector functions; its
aberrant activation often creates an immunosuppressive
microenvironment that can compromise the efficacy of immunotherapy
(111). IGF-1 and IGF-1R
demonstrate distinct pan-cancer expression patterns with
significant prognostic implications, with elevated levels
correlating with poor survival outcomes in specific malignancies
and altered tumor immune microenvironments. Their association with
immune checkpoint markers and genomic instability features
positions IGF-1R signaling as both a prognostic biomarker and a
potential immunotherapeutic target across multiple cancer types
(12).
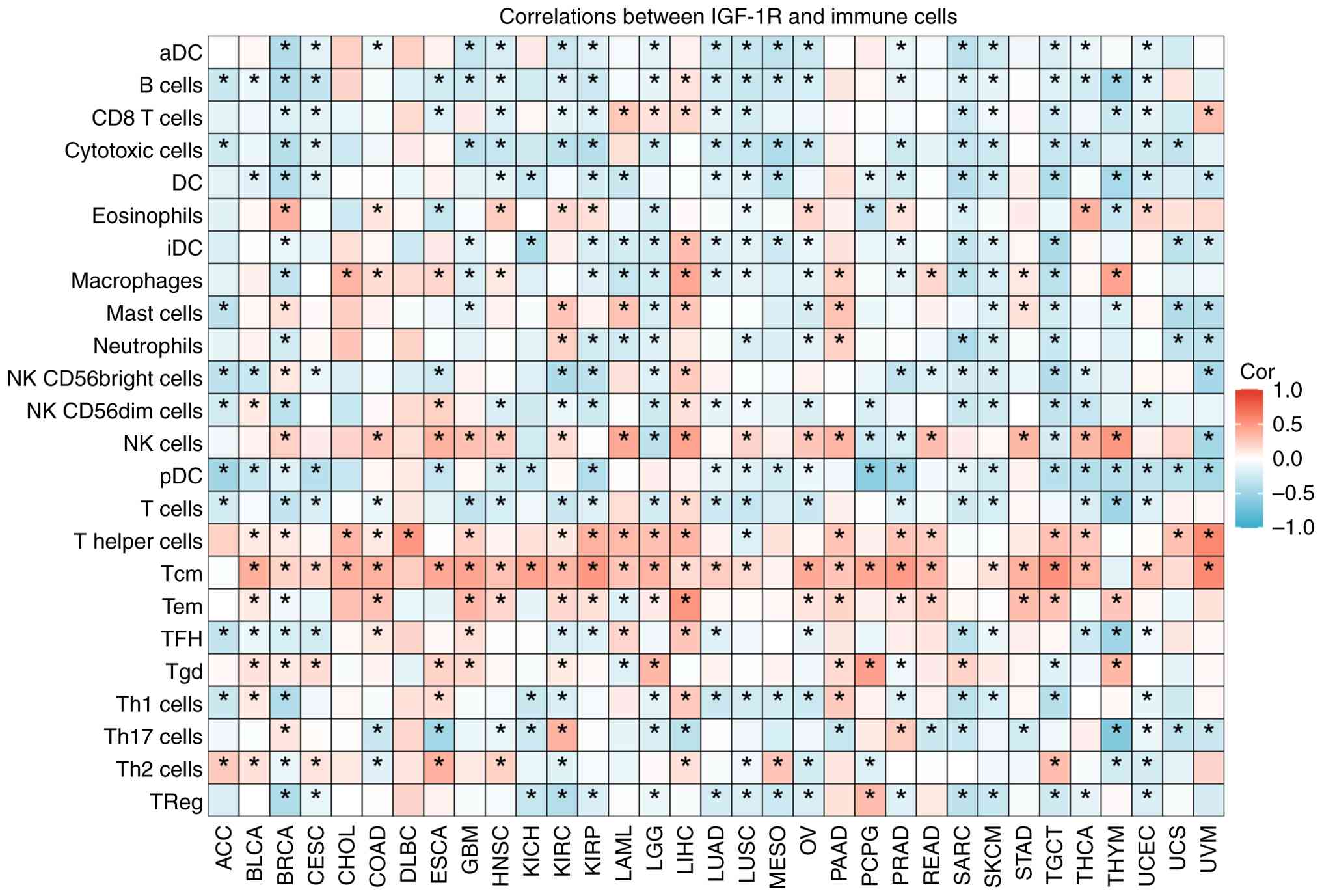 | Figure 5Correlation between IGF-1R expression
and immune cell infiltration in the tumor microenvironment across
cancers. A heatmap depicting Spearman's correlation coefficients
between IGF-1R expression levels (transcripts per million) and the
estimated abundance of various immune cell populations across 33
cancer types from The Cancer Genome Atlas (*P<0.05).
Red indicates positive correlations and blue indicates negative
correlations. This pan-cancer analysis suggests the potential role
of IGF-1R in modulating the tumor-immune microenvironment. IGF-1R,
insulin-like growth factor 1 receptor; cor, correlation. ACC,
adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA,
breast invasive carcinoma; CESC, cervical squamous cell carcinoma
and endocervical adenocarcinoma; CHOL, cholangiocarcinoma; COAD,
colon adenocarcinoma; DLBC, diffuse large B-cell lymphoma; ESCA,
esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and
neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC,
kidney renal clear cell carcinoma; KIRP, kidney renal papillary
cell carcinoma; LAML, acute myeloid leukemia; LGG, brain lower
grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung
adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO,
mesothelioma; OV, ovarian serous cystadenocarcinoma; PAAD,
pancreatic adenocarcinoma; PCPG, pheochromocytoma and
paraganglioma; PRAD, prostate adenocarcinoma; READ, rectum
adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD,
stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA,
thyroid carcinoma; THYM, thymoma; UCEC, uterine corpus endometrial
carcinoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma; iDC,
immature dendritic cells; NK, natural killer cells; Tcm, T central
memory cells; Tem, T effector memory cells; Tgd, γδ T cells; TFH, T
follicular helper cells; Th1, T helper 1 cells. |
Role of IGF-1R internalization and
compartmentalized signaling in cancer therapy
Mechanisms of resistance to
IGF-1R-targeted therapy
Targeted therapy against IGF-1R has demonstrated
considerable therapeutic potential in multiple cancer types;
however, drug resistance remains a major limitation to its clinical
efficacy. Receptor recycling and sustained signaling activation are
among the key mechanisms that confer resistance to IGF-1R
inhibitors (112). Tumor cells
enhance IGF-1R recycling, enabling continuous signal transduction
under therapeutic conditions, thereby diminishing drug inhibition.
Furthermore, internalized IGF-1R can engage in crosstalk with other
signaling pathways, such as by forming heterodimers with EGFR or
HER2, leading to activation of downstream survival and
proliferation pathways, including PI3K/AKT and Ras/RAF/ERK. This
results in redundant signaling networks that further exacerbate
resistance (121). Additionally,
certain tumor cells upregulate IGF-1R ligands (e.g., IGF-1 and
IGF-2) or downstream signaling molecules to enhance pathway
activity and counteract monotherapeutic inhibition (122,123). IGF-1R signaling also interacts
with DNA damage repair mechanisms and resistance-associated
proteins such as polycomb complex protein BMI-1. Co-inhibition of
these pathways can increase sensitivity to radiotherapy,
chemotherapy and targeted agents, thereby delaying the emergence of
resistance (99,124). Therefore, combination
therapeutic strategies that simultaneously target IGF-1R
internalization/compartmentalization and its synergistic pathways
may enable multilevel, multitarget interventions in tumor biology,
offering a rational approach to overcome mono-drug resistance and
improve treatment outcomes.
Progress in drug development and
preclinical research
Targeting of the IGF-1R signaling axis has led to
the development of diverse therapeutic agents, including monoclonal
antibodies, small-molecule TK inhibitors and soluble receptor
mutants (125-128). In preclinical studies, a
recombinant adenovirus-expressing dominant-negative mutant of
IGF-1R effectively suppressed tumor cell proliferation, migration
and tumorigenesis, demonstrating broad inhibition of downstream
pathways such as PI3K/AKT and Ras/RAF/MAPK (125). Moreover, IGF-1R-specific
inhibitors (e.g., NVP-AEW541 and picropodophyllin) have shown
promising antitumor activity in multiple tumor models. Combination
therapies involving mTOR inhibitors (e.g., rapamycin), immune
checkpoint inhibitors, CDK4/6 inhibitors, BRAF/MEK inhibitors or
DNA damage repair inhibitors significantly enhance treatment
efficacy and help overcome resistance and tumor heterogeneity
(1,126). Preclinical evidence indicates
that IGF-1R inhibitors can synergize with radiotherapy and
chemotherapy by increasing tumor cell sensitivity to DNA damage and
promoting cell cycle arrest and apoptosis (99). In the context of CSC-targeted
therapies, IGF-1R signaling inhibitors effectively reduce stemness
and suppress tumor recurrence and metastasis (7,13).
Novel tools, such as molecular imaging probes and fluorescently
labeled antibodies against IGF-1R, are also under development,
providing a technical foundation for precise diagnosis and
treatment response monitoring (127,128).
Biomarkers and prediction of
therapeutic response
Recent advances have highlighted the potential of
IGF-1R internalization- and compartmentalization-associated
molecules as predictive biomarkers of tumor therapeutic responses.
The subcellular localization of internalized IGF-1R, such as within
the Golgi apparatus or the nucleus, has been closely linked to
specific biological behaviors and clinical outcomes (15-18). For instance, IGF-1R accumulation
in the Golgi apparatus of migratory cancer cells promotes invasive
behaviors, such as cell migration, while nuclear localization of
IGF-1R is associated with poorer clinical outcomes (14), suggesting that the intracellular
distribution of IGF-1R may serve as a potential biomarker for tumor
progression and prognosis (14).
In osteosarcoma and other malignancies, elevated levels of
phosphorylated IGF-1R in the nucleus negatively correlate with
patient survival, further supporting its prognostic utility
(129). Moreover, changes in
IGF-1R expression in circulating tumor cells (CTCs) are closely
related to treatment outcomes. These findings reveal that the loss
of IGF-1R expression in CTCs is significantly associated with
unfavorable outcomes in patients with metastatic breast cancer,
suggesting that the abundance of IGF-1R-negative CTCs may serve as
an independent prognostic indicator to guide patient stratification
and personalized treatment strategies (130). Collectively, these findings
underscore the clinical value of IGF-1R and its
compartmentalization-related molecules for predicting therapeutic
responses and evaluating prognosis in cancer treatment.
Limitations and future perspectives
Despite the efforts of the present review to provide
a comprehensive overview of the non-canonical roles of IGF-1R,
several limitations of both it and the current body of literature
must be acknowledged. First, the review was constrained by the
availability and quality of published literature. Despite the
systematic search, relevant studies may have been missed, leading
to a potential selection bias. Second, a significant portion of the
evidence supporting the direct role of IGF-1R in DNA repair comes
from observational studies (e.g., co-localization and
knockdown/knockout experiments showing reduced repair focus
formation). While these associations are highly suggestive, more
direct biochemical evidence (e.g., detailed structural
interactions) in vivo is needed to unequivocally establish
causality. Finally, although preclinical data on targeting IGF-1R
are promising, the transition to clinical success has been limited.
This highlights the complexity of human biology and the challenges
in overcoming compensatory pathways and on-target toxicity.
However, our understanding of the crosstalk between IGF-1R and
other resistance mechanisms remains incomplete.
Future research should therefore prioritize several
avenues: i) Utilizing advanced techniques (e.g.,
cryo-electron microscopy, live-cell imaging and proximity labeling)
to map the precise molecular interactions of nuclear IGF-1R; ii)
developing more sophisticated in vivo models to validate
these mechanisms and test novel therapeutic strategies, such as
inhibiting IGF-1R nuclear translocation rather than its kinase
activity; and iii) designing biomarker-driven clinical trials to
identify patient subpopulations most likely to benefit from
IGF-1R-targeted therapies, potentially in combination with other
agents to overcome resistance.
Conclusion
In recent years, internalization and
compartmentalization of IGF-1R have been demonstrated to play
critical regulatory roles in tumor initiation, progression and
therapeutic response. Advances in molecular and cellular biology
techniques have significantly advanced our understanding of the
dynamic membrane trafficking, subcellular localization and
signaling networks of the IGF-1R. Studies have revealed that IGF-1R
internalization influences ligand binding and signal transduction
efficiency, and triggers compartment-specific downstream effects,
thereby regulating key oncogenic processes, such as proliferation,
apoptosis, migration and drug resistance.
Despite substantial progress in elucidating the
mechanisms and functional consequences of IGF-1R internalization,
several aspects remain controversial, including the detailed
molecular networks, cross-compartmental signal integration and
interactions with the TME. While some studies have emphasized the
role of classical endocytic pathways in the spatiotemporal
regulation of IGF-1R signaling, others have suggested that
non-canonical internalization routes and interactions with the
cytoskeleton and vesicular transport systems critically contribute
to tumor heterogeneity and drug resistance. Therefore, further
experimental validation is required to clarify how IGF-1R
internalization and intercompartmental signaling regulate tumor
progression and treatment resistance.
From a clinical perspective, a deeper understanding
of IGF-1R internalization and compartmentalization provides a
theoretical foundation for the development of novel therapeutic
strategies. The limited efficacy of traditional IGF-1R inhibitors
may be attributed, in part, to signal adaptation via
internalization and spatial reorganization. Future targeting
strategies should aim to precisely intercept compartment-specific
IGF-1R signaling or combine it with agents that prevent adaptive
resistance, thereby improving antitumor efficacy. Furthermore,
emerging biomarkers that capture the dynamic subcellular behavior
of IGF-1R hold promise for tumor classification, prognostic
assessment and personalized therapy. Advances in liquid biopsy and
single-cell multiomics will likely accelerate the application of
IGF-1R dynamic signaling features in precision oncology.
In summary, research on IGF-1R internalization and
compartmentalization represents a cutting-edge area in tumor
biology that extends our understanding of oncogenesis and offers
new avenues for targeted therapy. Future studies should focus on
the molecular complexity of IGF-1R trafficking and signaling,
strengthen interdisciplinary collaboration, and promote the
development and clinical translation of novel biomarkers. These
efforts will ultimately contribute to more precise and effective
therapeutic strategies that improve the survival outcomes and
quality of life for patients with cancer.
Supplementary Data
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
TZ, LL and CD guarantee the integrity of the entire
study. TZ designed the paper. TZ, LL and CD performed the
literature research, data acquisition and data analysis. LL and CD
prepared the manuscript, and TZ wrote the manuscript. TZ and LL
confirm the authenticity of all the raw data. All authors have read
and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
No funding was received.
References
|
1
|
Guven DC, Ahmed J, Stephen B and Naing A:
IGF-1R inhibitors in cancer: A review of available evidence and
future outlook. Crit Rev Oncol Hematol. 214:1048092025. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Gui R, Li W, Li Z, Wang H, Wu Y, Jiao W,
Zhao G, Shen Y, Wang L, Zhang J, et al: Effects and potential
mechanisms of IGF1/IGF1R in the liver fibrosis: A review. Int J
Biol Macromol. 251:1262632023. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Robertson DM, Zhu M and Wu YC: Cellular
distribution of the IGF-1R in corneal epithelial cells. Exp Eye
Res. 94:179–186. 2012. View Article : Google Scholar
|
|
4
|
Martin A, Fernandez MC, Miraglia S, Venara
M, Clement F, Papendieck P, De Matteo E and Pennisi PA: Brachyury
and IGF1R: Potential opposing roles in pediatric thyroid nodular
pathology. J Endocrinol Invest. 48:2643–2655. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Truong T and Silkiss RZ: The role of
insulin-like growth factor-1 and its receptor in the eye: A review
and implications for IGF-1R inhibition. Ophthalmic Plast Reconstr
Surg. 39:4–12. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Nieto Guil AF, Oksdath M, Weiss LA, Grassi
DJ, Sosa LJ, Nieto M and Quiroga S: IGF-1 receptor regulates
dynamic changes in neuronal polarity during cerebral cortical
migration. Sci Rep. 7:77032017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ngo MT, Jeng HY, Kuo YC, Diony Nanda J,
Brahmadhi A, Ling TY, Chang TS and Huang YH: The role of IGF/IGF-1R
signaling in hepatocellular carcinomas: Stemness-related properties
and drug resistance. Int J Mol Sci. 22:19312021. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liu G, Zhu M, Zhang M and Pan F: Emerging
role of IGF-1 in prostate cancer: A promising biomarker and
therapeutic target. Cancers (Basel). 15:12872023. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Weiwei Z, Ya X, Wenwen W, Jia J, Jing B,
Ruitao Z, Chunfang W and Ruixia G: IGF-1R anti-idiotypic antibody
antagonist exhibited anti-ovarian cancer bioactivity and reduced
cisplatin resistance. Hum Cell. 34:1197–1214. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ben Elhadj M, Goucha A, Fourati A, Adouni
O, Dhambri S, Hsairi M, El May MV and Mokni Baizig N: The
prognostic significance of IGF-1R and the predictive risk value of
circulating IGF-1 in tunisian patients with laryngeal carcinoma.
Cancer Invest. 38:289–299. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Guo C, Zhang M, Qian J, Li P and Guo L:
Oncogenic long noncoding RNA Linc01287 promotes IGF1R expression by
sponging miR-98 in breast cancer. Crit Rev Eukaryot Gene Expr.
32:31–44. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhang Y, Gao C, Cao F, Wu Y, Chen S, Han
X, Mo J, Qiu Z, Fan W, Zhou P and Shen L: Pan-cancer analysis of
IGF-1 and IGF-1R as potential prognostic biomarkers and
immunotherapy targets. Front Oncol. 11:7553412021. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Liu F, Ye S, Zhao L and Niu Q: The role of
IGF/IGF-1R signaling in the regulation of cancer stem cells. Clin
Transl Oncol. 26:2924–2934. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Rieger L and O'Connor R: Controlled
signaling-insulin-like growth factor receptor endocytosis and
presence at intracellular compartments. Front Endocrinol
(Lausanne). 11:6200132021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Peloso Maia ML, Albuquerque RM, do Carmo
Silva SD, Lima CX, Costa Diniz PH and Vieira Teixeira Vidigal P:
Immunohistochemical expression of insulin-like growth factor-1
receptor and its association with clinicopathological parameters in
hepatocellular carcinoma. Oncology. 102:494–502. 2024. View Article : Google Scholar :
|
|
16
|
Soni UK, Jenny L and Hegde RS: IGF-1R
targeting in cancer-does sub-cellular localization matter? J Exp
Clin Cancer Res. 42:2732023. View Article : Google Scholar
|
|
17
|
Grice DM, Vetter I, Faddy HM, Kenny PA,
Roberts-Thomson SJ and Monteith GR: Golgi calcium pump secretory
pathway calcium ATPase 1 (SPCA1) is a key regulator of insulin-like
growth factor receptor (IGF1R) processing in the basal-like breast
cancer cell line MDA-MB-231. J Biol Chem. 285:37458–37466. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Rieger L, O'Shea S, Godsmark G, Stanicka
J, Kelly G and O'Connor R: IGF-1 receptor activity in the Golgi of
migratory cancer cells depends on adhesion-dependent
phosphorylation of Tyr(1250) and Tyr(1251). Sci Signal.
13:eaba31762020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Crudden C and Girnita L: The tale of a
tail: The secret behind IGF-1R's oncogenic power. Sci Signal.
13:eabb78872020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Werner H, Sarfstein R and Laron Z: The
role of nuclear insulin and IGF1 receptors in metabolism and
cancer. Biomolecules. 11:5312021. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Amin HM, Morani AC, Daw NC,
Lamhamedi-Cherradi SE, Subbiah V, Menegaz BA, Vishwamitra D,
Eskandari G, George B, Benjamin RS, et al: IGF-1R/mTOR targeted
therapy for ewing sarcoma: A meta-analysis of five IGF-1R-related
trials matched to proteomic and radiologic predictive biomarkers.
Cancers (Basel). 12:17682020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Batishchev OV, Kuzmina NV, Mozhaev AA,
Goryashchenko AS, Mileshina ED, Orsa AN, Bocharov EV, Deyev IE and
Petrenko AG: Activity-dependent conformational transitions of the
insulin receptor-related receptor. J Biol Chem. 296:1005342021.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Ferguson KM, Hu C and Lemmon MA: Insulin
and epidermal growth factor receptor family members share parallel
activation mechanisms. Protein Sci. 29:1331–1344. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Sheff J, Wang P, Xu P, Arbour M, Masson L,
van Faassen H, Hussack G, Kemmerich K, Brunette E, Stanimirovic D,
et al: Defining the epitope of a blood-brain barrier crossing
single domain antibody specific for the type 1 insulin-like growth
factor receptor. Sci Rep. 11:42842021. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang C, Liu S, Wu Q, Cheng Y, Feng T, Song
J, Yang R, Geng H, Lu G, Wang S and Hao L: Porcine IGF-1R
synonymous mutations in the intracellular domain affect cell
proliferation and alter kinase activity. Int J Biol Macromol.
152:147–153. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Bano N, Hossain MM, Bhat AQ, Ayaz MO,
Kumari M, Sandhu P, Akhter Y and Dar MJ: Analyzing structural
differences between insulin receptor (IR) and IGF1R for designing
small molecule allosteric inhibitors of IGF1R as novel anti-cancer
agents. Growth Horm IGF Res. 55:1013432020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Mills JV, Osher E, Rieunier G, Mills IG
and Macaulay VM: IGF-1R nuclear import and recruitment to chromatin
involves both alpha and beta subunits. Discov Oncol. 12:132021.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Khan N, Althobiti M, Chinnadurai RK,
Alharbi S and Kumar R: Evolutionary sequences and structural
information-driven reconstruction of new insulin-like growth
factor-I peptide variants. Curr Mol Med. 25:652–661. 2025.
View Article : Google Scholar
|
|
29
|
Salvi R, Kumar C, Brahmbhatt K, Subedi R,
Idicula-Thomas S, Madan T and Biswas B: N-Linked glycosylation in
chinese hamster ovary cells is critical for insulin-like growth
factor 1 signaling. Int J Mol Sci. 23:149522022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Vivian J, Rao AA, Nothaft FA, Ketchum C,
Armstrong J, Novak A, Pfeil J, Narkizian J, Deran AD,
Musselman-Brown A, et al: Toil enables reproducible, open source,
big biomedical data analyses. Nat Biotechnol. 35:314–316. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Cheng H, Dodge J, Mehl E, Liu S, Poulin N,
van de Rijn M and Nielsen TO: Validation of immature adipogenic
status and identification of prognostic biomarkers in myxoid
liposarcoma using tissue microarrays. Hum Pathol. 40:1244–1251.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li F, Wang X, Zhang J, Zhang J, Jing X,
Jiang Q, Zhou J, Cao L, Peng H, Tong D and Huang C: RBM8A, a new
target of TEAD4, promotes breast cancer progression by regulating
IGF1R and IRS-2. J Transl Med. 22:8232024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang J, Chen B, Li H, Wang Y, Liu X, Wong
KY, Chan WN, Chan AK, Cheung AH, Leung KT, et al: Cancer-associated
fibroblasts potentiate colorectal cancer progression by crosstalk
of the IGF2-IGF1R and Hippo-YAP1 signaling pathways. J Pathol.
259:205–219. 2023. View Article : Google Scholar
|
|
34
|
Huang YK, Kang WM, Ma ZQ, Liu YQ, Zhou L
and Yu JC: NUCKS1 promotes gastric cancer cell aggressiveness by
upregulating IGF-1R and subsequently activating the PI3K/Akt/mTOR
signaling pathway. Carcinogenesis. 40:370–379. 2019. View Article : Google Scholar
|
|
35
|
Jassi C, Kuo WW, Chang YC, Wang TF, Li CC,
Ho TJ, Hsieh DJ, Kuo CH, Chen MC and Huang CY: Aloin and CPT-11
combination activates miRNA-133b and downregulates
IGF1R-PI3K/AKT/mTOR and MEK/ERK pathways to inhibit colorectal
cancer progression. Biomed Pharmacother. 169:1159112023. View Article : Google Scholar
|
|
36
|
Zhou Y, Deng Y, Wang J, Yan Z, Wei Q, Ye
J, Zhang J, He TC and Qiao M: Effect of antibiotic monensin on cell
proliferation and IGF1R signaling pathway in human colorectal
cancer cells. Ann Med. 55:954–964. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yang B, Li G, Wang S, Zheng Y, Zhang J,
Pan B, Wang N and Wang Z: Tumor-associated macrophages/C-X-C motif
chemokine ligand 1 promotes breast cancer autophagy-mediated
chemoresistance via IGF1R/STAT3/HMGB1 signaling. Cell Death Dis.
15:7432024. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Zhu S, Soutto M, Chen Z, Blanca Piazuelo
M, Kay Washington M, Belkhiri A, Zaika A, Peng D and El-Rifai W:
Activation of IGF1R by DARPP-32 promotes STAT3 signaling in gastric
cancer cells. Oncogene. 38:5805–5816. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Chen C, Gupta P, Parashar D, Nair GG,
George J, Geethadevi A, Wang W, Tsaih SW, Bradley W, Ramchandran R,
et al: ERBB3-induced furin promotes the progression and metastasis
of ovarian cancer via the IGF1R/STAT3 signaling axis. Oncogene.
39:2921–2933. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Yang C, Zhang Y, Segar N, Huang C, Zeng P,
Tan X, Mao L, Chen Z, Haglund F, Larsson O, et al: Nuclear IGF1R
interacts with NuMA and regulates 53BP1-dependent DNA double-strand
break repair in colorectal cancer. Oncol Rep. 46:1682021.
View Article : Google Scholar
|
|
41
|
Li L, Zhang Z, Huang N, Ren J, Qin Y and
Luo Y: IGF1R activates FOXP3-β-catenin signaling to promote breast
cancer development. Breast Cancer Res Treat. 211:467–478. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Guo C, Chu H, Gong Z, Zhang B, Li C, Chen
J and Huang L: HOXB13 promotes gastric cancer cell migration and
invasion via IGF-1R upregulation and subsequent activation of
PI3K/AKT/mTOR signaling pathway. Life Sci. 278:1195222021.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Chen WJ, Tsai JH, Hsu LS, Lin CL, Hong HM
and Pan MH: Quercetin blocks the aggressive phenotype of
triple-negative breast cancer by inhibiting IGF1/IGF1R-mediated EMT
program. J Food Drug Anal. 29:98–112. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Li W, Huang C, Qiu L, Tang Y, Zhang X,
Zhang L, Zhao H, Miyagishi M, Kasim V and Wu S: p52-ZER6/IGF1R axis
maintains cancer stem cell population to promote cancer progression
by enhancing pro-survival mitophagy. Oncogene. 43:2115–2131. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang R, Li L and Yu J: Lactate-induced
IGF1R protein lactylation promotes proliferation and metabolic
reprogramming of lung cancer cells. Open Life Sci. 19:202208742024.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Zhang M, Li Z and Liu X: MiR-98-5p/IGF2
axis influence herceptin sensitivity through IGF1R/HER2 heterodimer
formation and AKT/mTOR signal pathway in HER2 positive breast
cancer. Asian Pac J Cancer Prev. 22:3693–3703. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Jassi C, Kuo WW, Chang YC, Wang TF, Ho TJ,
Hsieh DJ, Kuo CH, Chen MC, Li CC and Huang CY: MicroRNA-376a-3p
sensitizes CPT-11-resistant colorectal cancer by enhancing
apoptosis and reversing the epithelial-to-mesenchymal transition
(EMT) through the IGF1R/PI3K/AKT pathway. Transl Oncol.
50:1021252024. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Catalano R, Giardino E, Treppiedi D,
Mangili F, Morelli V, Elli FM, Serban AL, Luconi M, Mannelli M,
Spada A, et al: The cytoskeleton actin binding protein filamin A
impairs both IGF2 mitogenic effects and the efficacy of IGF1R
inhibitors in adrenocortical cancer cells. Cancer Lett. 497:77–88.
2021. View Article : Google Scholar
|
|
49
|
Shin GC, Lee HM, Kim N, Seo SU, Kim KP and
Kim KH: PRKCSH contributes to TNFSF resistance by extending IGF1R
half-life and activation in lung cancer. Exp Mol Med. 56:192–209.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Xie Q, Chu Y, Yuan W, Li Y, Li K, Wu X,
Liu X, Xu R, Cui S and Qu X: Activation of insulin-like growth
factor-1 receptor (IGF-1R) promotes growth of colorectal cancer
through triggering the MEX3A-mediated degradation of RIG-I. Acta
Pharm Sin B. 13:2963–2975. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Liu Z, Meng D, Wang J, Cao H, Feng P, Wu
S, Wang N, Dang C, Hou P and Xia P: GASP1 enhances malignant
phenotypes of breast cancer cells and decreases their response to
paclitaxel by forming a vicious cycle with IGF1/IGF1R signaling
pathway. Cell Death Dis. 13:7512022. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Santos Fortes Dos Reis VM, Ramos FM, de
Oliveira HL, Machado FD, Hartke S, Machado-Weber A, Germeyer A,
Strowitzki T, Kliemann LM, von Eye Corleta H, et al: Effects of
metformin treatment against endometrial cancer cells cultured in
vitro or grafted into female balb/C nude mice: Insights into cell
response and IGF-1R and PI3K/AKT/mTOR signaling pathways. Cell
Biochem Biophys. 83:5227–5245. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Kuo YC, Chen CL, Lee KL, Wang HF, Drew VJ,
Lan PC, Ho YS and Huang YH: Nicotine-driven enhancement of tumor
malignancy in triple-negative breast cancer via additive regulation
of CHRNA9 and IGF1R. J Pathol. 266:230–245. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Deo AN, Thorat R, Dhadve AC, De A, Rekhi B
and Ray P: IGF1R-α6 integrin-S100A4 network governs the
organ-specific metastasis of chemoresistant epithelial ovarian
cancer cells. Biochim Biophys Acta Mol Basis Dis. 1868:1662822022.
View Article : Google Scholar
|
|
55
|
Noyan S and Gur Dedeoglu B:
miR-770-5p-induced cellular switch to sensitize trastuzumab
resistant breast cancer cells targeting HER2/EGFR/IGF1R
bidirectional crosstalk. Turk J Biol. 48:153–162. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Bustamante-Marin X, Devlin KL, McDonell
SB, Dave O, Merlino JL, Grindstaff EJ, Ho AN, Rezeli ET, Coleman MF
and Hursting SD: Regulation of IGF1R by MicroRNA-15b contributes to
the anticancer effects of calorie restriction in a murine C3-TAg
model of triple-negative breast cancer. Cancers (Basel).
15:43202023. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Jiang S, Chen H, He K and Wang J: Human
bone marrow mesenchymal stem cells-derived exosomes attenuated
prostate cancer progression via the miR-99b-5p/IGF1R axis.
Bioengineered. 13:2004–2016. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Han B, Xu K, Feng D, Bai Y, Liu Y, Zhang Y
and Zhou L: miR-144 inhibits the IGF1R-ERK1/2 signaling pathway via
NUDCD1 to suppress the proliferation and metastasis of colorectal
cancer cells: A study based on bioinformatics and in vitro and in
vivo verification. J Cancer Res Clin Oncol. 148:1903–1918. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Zhang H, Zheng XD, Zeng XH, Li L and Zhou
Q: miR-520b Inhibits IGF-1R to increase doxorubicin sensitivity and
promote cell apoptosis in breast cancer. Yakugaku Zasshi.
141:415–426. 2021. View Article : Google Scholar
|
|
60
|
Han L: miR-99a inhibits proliferation and
migration of cervical cancer cells by targeting IGF1R. J BUON 2021.
26:1782–1788. 2021.
|
|
61
|
Bai R, Dou K, Wu Y, Ma Y and Sun J: The
NF-ĸB modulated miR-194-5p/IGF1R/PPFIBP axis is crucial for the
tumorigenesis of ovarian cancer. J Cancer. 11:3433–3445. 2020.
View Article : Google Scholar
|
|
62
|
Hsu XR, Wu JE, Wu YY, Hsiao SY, Liang JL,
Wu YJ, Tung CH, Huang MF, Lin MS, Yang PC, et al: Exosomal long
noncoding RNA MLETA1 promotes tumor progression and metastasis by
regulating the miR-186-5p/EGFR and miR-497-5p/IGF1R axes in
non-small cell lung cancer. J Exp Clin Cancer Res. 42:2832023.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Golestan S, Soltani BM, Jafarzadeh M,
Ghaemi Z and Nafisi N: LINC02381 suppresses cell proliferation and
promotes apoptosis via attenuating IGF1R/PI3K/AKT signaling pathway
in breast cancer. Funct Integr Genomics. 23:402023. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Zhou L, Li H, Sun T, Wen X, Niu C, Li M,
Li W, Hoffman AR, Hu JF and Cui J: HULC targets the IGF1R-PI3K-AKT
axis in trans to promote breast cancer metastasis and cisplatin
resistance. Cancer Lett. 548:2158612022. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Zhang L, Cai Y, Tian C, Li Y, Ma K, Gao X,
Liu L, Jiang Y, Wen W and Ma Z: LncRNA Opa interacting protein
5-antisense RNA 1 (OIP5-AS1) promotes the migration, invasion and
epithelial-mesenchymal transition (EMT) through targeting
miR-147a/insulin-like growth factor 1 receptor (IGF1R) pathway in
cervical cancer tissues and cell model. J Obstet Gynaecol Res.
48:1222–1232. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Liu Y, Peng H, Shen Y, Da R, Tian A and
Guo X: Downregulation of long noncoding RNA myocardial infarction
associated transcript suppresses cell proliferation, migration,
invasion, and glycolysis by regulation of miR-488-3p/IGF1R pathway
in colorectal cancer. Cancer Biother Radiopharm. 37:927–938.
2022.
|
|
67
|
Xie Q, Zhao S, Kang R and Wang X: lncRNA
SNHG11 facilitates prostate cancer progression through the
upregulation of IGF-1R expression and by sponging miR-184. Int J
Mol Med. 48:1822021. View Article : Google Scholar
|
|
68
|
Liu B, Jiang HY, Yuan T, Zhou WD, Xiang
ZD, Jiang QQ and Wu DL: Long Non-coding RNA AFAP1-AS1 facilitates
prostate cancer progression by regulating miR-15b/IGF1R axis. Curr
Pharm Des. 27:4261–4269. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Capik O, Sanli F, Kurt A, Ceylan O, Suer
I, Kaya M, Ittmann M and Karatas OF: CASC11 promotes aggressiveness
of prostate cancer cells through miR-145/IGF1R axis. Prostate
Cancer Prostatic Dis. 24:891–902. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Zhang M, Lin B, Liu Y, Huang T, Chen M,
Lian D, Deng S and Zhuang C: LINC00324 affects non-small cell lung
cancer cell proliferation and invasion through regulation of the
miR-139-5p/IGF1R axis. Mol Cell Biochem. 473:193–202. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Ma Z, Cai Y, Zhang L, Tian C and Lyu L:
LINC00319 promotes cervical cancer progression via targeting
miR-147a/IGF1R pathway. Cancer Biother Radiopharm. Jul 9–2020.Epub
ahead of print.
|
|
72
|
Du P, Liu F, Liu Y, Shao M, Li X and Qin
G: Linc00210 enhances the malignancy of thyroid cancer cells by
modulating miR-195-5p/IGF1R/Akt axis. J Cell Physiol.
235:1001–1012. 2020. View Article : Google Scholar
|
|
73
|
Cui Y, Xie M and Zhang Z: LINC00958
involves in bladder cancer through sponging miR-378a-3p to Elevate
IGF1R. Cancer Biother Radiopharm. 35:776–788. 2020.PubMed/NCBI
|
|
74
|
Zhang R, Hao S, Yang L, Xie J, Chen S and
Gu G: LINC00339 promotes cell proliferation and metastasis in
pancreatic cancer via miR-497-5p/IGF1R axis. J BUON. 24:729–738.
2019.PubMed/NCBI
|
|
75
|
Zhang X, Fang F, Zhang J, Zhang S, Li H,
Li B, Zhong Y and Zhen P: Circ_0006174 upregulates IGF1R to enhance
radioresistance and tumorigenesis in colorectal cancer via miR-940
suppression. Appl Biochem Biotechnol. 197:497–517. 2025. View Article : Google Scholar
|
|
76
|
Tang YF, Liu ZH, Zhang LY, Shi SH, Xu S,
Ma JA, Hu CH and Zou FW: circ_PPAPDC1A promotes Osimertinib
resistance by sponging the miR-30a-3p/IGF1R pathway in non-small
cell lung cancer (NSCLC). Mol Cancer. 23:912024. View Article : Google Scholar
|
|
77
|
Gao S, Zhang X, Bai W, Wang J and Jiang B:
Circ-IGF1R affects the progression of colorectal cancer by
activating the miR-362-5p/HMGB3-Mediated Wnt/β-catenin signal
pathway. Biochem Genet. 61:1210–1229. 2023. View Article : Google Scholar
|
|
78
|
Hua J, Wang X, Ma L, Li J, Cao G, Zhang S
and Lin W: CircVAPA promotes small cell lung cancer progression by
modulating the miR-377-3p and miR-494-3p/IGF1R/AKT axis. Mol
Cancer. 21:1232022. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Wang P, Sun Y, Yang Y, Chen Y and Liu H:
Circ_0067835 knockdown enhances the radiosensitivity of colorectal
cancer by miR-296-5p/IGF1R axis. Onco Targets Ther. 14:491–502.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Liu X, Zhou L, Chen Y, Jiang X and Jiang
J: CircRNF13 promotes the malignant progression of pancreatic
cancer through targeting miR-139-5p/IGF1R axis. J Oncol.
2021:69450462021. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Wang Y, Yin L and Sun X: CircRNA
I_circ_0002577 accelerates endometrial cancer progression through
activating IGF1R/PI3K/Akt pathway. J Exp Clin Cancer Res.
39:1692020. View Article : Google Scholar
|
|
82
|
Liu P, Zou Y, Li X, Yang A, Ye F, Zhang J,
Wei W and Kong Y: circGNB1 facilitates triple-negative breast
cancer progression by regulating miR-141-5p-IGF1R axis. Front
Genet. 11:1932020. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Wang X, Song Z, Meng Q, Xia S, Wang C and
Huang X: Circular RNA circ_0006089 regulates the IGF1R expression
by targeting miR-143-3p to promote gastric cancer proliferation,
migration and invasion. Cell Cycle. 1–14. 2022.Epub ahead of print.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Dam DHM, Jelsma SA, Yu JM, Liu H, Kong B
and Paller AS: Flotillin and AP2A1/2 Promote IGF-1 receptor
association with clathrin and internalization in primary human
keratinocytes. J Invest Dermatol. 140:1743–1752 e4. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Kwon H, Choi M, Ahn Y, Jang D and Pak Y:
Flotillin-1 palmitoylation turnover by APT-1 and ZDHHC-19 promotes
cervical cancer progression by suppressing IGF-1 receptor
desensitization and proteostasis. Cancer Gene Ther. 30:302–312.
2023. View Article : Google Scholar
|
|
86
|
Xiu M, Huan X, Ou Y, Ying S and Wang J:
The basic route of nuclear-targeted transport of IGF-1/IGF-1R and
potential biological functions in intestinal epithelial cells. Cell
Prolif. 54:e130302021. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Crudden C, Song D, Cismas S, Trocme E,
Pasca S, Calin GA, Girnita A and Girnita L: Below the surface:
IGF-1R therapeutic targeting and its endocytic journey. Cells.
8:12232019. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Chen X, Pang Z, Wang Y, Zhu L, Liu J and
Du J: Cezanne contributes to cancer progression by playing a key
role in the deubiquitination of IGF-1R. Am J Cancer Res.
10:4342–4356. 2020.
|
|
89
|
Packham S, Warsito D, Lin Y, Sadi S,
Karlsson R, Sehat B and Larsson O: Nuclear translocation of IGF-1R
via p150(Glued) and an importin-β/RanBP2-dependent pathway in
cancer cells. Oncogene. 34:2227–2238. 2015. View Article : Google Scholar
|
|
90
|
Sehat B, Tofigh A, Lin Y, Trocme E,
Liljedahl U, Lagergren J and Larsson O: SUMOylation mediates the
nuclear translocation and signaling of the IGF-1 receptor. Sci
Signal. 3:ra102010. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Gulec Taskiran AE, Husnugil HH, Soltani
ZE, Oral G, Menemenli NS, Hampel C, Huebner K, Erlenbach-Wuensch K,
Sheraj I, Schneider-Stock R, et al: Post-transcriptional regulation
of Rab7a in lysosomal positioning and drug resistance in
nutrient-limited cancer cells. Traffic. 25:e129562024. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Wong WH, Liu SZ, Li ASR, Liu X, Manolson
MF and Zirngibl RA: Evidence for Rab7b and its splice isoforms
having distinct biological functions from Rab7a. Int J Mol Sci.
26:26102025. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Hewawasam NV, Lhaf F, Taylor HA, Viloria
K, Austin A, King A, Jones P, Jones L, Turner MD and Hill NJ:
Modulation of Rab7a-mediated growth factor receptor trafficking
inhibits islet beta cell apoptosis and autophagy under conditions
of metabolic stress. Sci Rep. 10:157412020. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
McDermott N, O'Shea S, Rieger L, Cox OT
and O'Connor R: beta(1)-integrin controls IGF-1R internalization
and intracellular signaling. J Biol Chem. 301:1080212025.
View Article : Google Scholar
|
|
95
|
Guglielmi V, Lam D and D'Angelo MA:
Nucleoporin Nup358 drives the differentiation of myeloid-biased
multipotent progenitors by modulating HDAC3 nuclear translocation.
Sci Adv. 10:eadn89632024. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Guo Y, Tao T, Wu T, Hou J and Lin W:
Nucleoporin Nup98 is an essential factor for ipo4 dependent protein
import. J Cell Biochem. 125:e305732024. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Artico LL, Laranjeira ABA, Campos LW,
Correa JR, Zenatti PP, Carvalheira JBC, Brambilla SR, Nowill AE,
Brandalise SR and Yunes JA: Physiologic IGFBP7 levels prolong IGF1R
activation in acute lymphoblastic leukemia. Blood Adv. 5:3633–3646.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Chakraborty S, Bhat AM, Mushtaq I, Luan H,
Kalluchi A, Mirza S, Storck MD, Chaturvedi N, Lopez-Guerrero JA,
Llombart-Bosch A, et al: EHD1-dependent traffic of IGF-1 receptor
to the cell surface is essential for Ewing sarcoma tumorigenesis
and metastasis. Commun Biol. 6:7582023. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Zong R, Chen X, Feng J and Xu S: IGF-1R
depletion sensitizes colon cancer cell lines to radiotherapy.
Cancer Biomark. 32:199–206. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Ko JC, Syu JJ, Chen JC, Wang TJ, Chang PY,
Chen CY, Jian YT, Jian YJ and Lin YW: Resveratrol enhances
etoposide-induced cytotoxicity through down-regulating ERK1/2 and
AKT-Mediated X-ray repair cross-complement group 1 (XRCC1) protein
expression in human non-small-cell lung cancer cells. Basic Clin
Pharmacol Toxicol. 117:383–391. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Toulany M, Dittmann K, Fehrenbacher B,
Schaller M, Baumann M and Rodemann HP: PI3K-Akt signaling regulates
basal, but MAP-kinase signaling regulates radiation-induced XRCC1
expression in human tumor cells in vitro. DNA Repair (Amst).
7:1746–1756. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Deng H, Lin Y, Badin M, Vasilcanu D,
Stromberg T, Jernberg-Wiklund H, Sehat B and Larsson O:
Over-accumulation of nuclear IGF-1 receptor in tumor cells requires
elevated expression of the receptor and the SUMO-conjugating enzyme
Ubc9. Biochem Biophys Res Commun. 404:667–671. 2011. View Article : Google Scholar
|
|
103
|
Li J, Yin Q, Xuan N, Gan Q, Liu C, Zhang
Q, Yang M and Yang C: LYSMD proteins promote activation of
Rab32-family GTPases for lysosome-related organelle biogenesis. J
Cell Biol. 223:e2024020162024. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Haga K and Fukuda M: Comprehensive
knockout analysis of the RAB family small GTPases reveals an
overlapping role of RAB2 and RAB14 in autophagosome maturation.
Autophagy. 21:21–36. 2025. View Article : Google Scholar :
|
|
105
|
Matsumoto N, Sekiya M, Sun-Wada GH, Wada Y
and Nakanishi-Matsui M: The lysosomal V-ATPase a3 subunit is
involved in localization of Mon1-Ccz1, the GEF for Rab7, to
secretory lysosomes in osteoclasts. Sci Rep. 12:84552022.
View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Wang X, Wang Y, Lei P, Qu X, Qi R, Chen D
and Chang Y: IGFBP5 regulates fibrocartilage differentiation and
cartilage injury induced by T-2 toxin via blocking IGF-1/IGF-1R
signalling. Rheumatology (Oxford). 64:4051–4060. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Peng SW, Ngo MT, Kuo YC, Teng MH, Guo CL,
Lai HC, Chang TS and Huang YH: Niclosamide revitalizes sorafenib
through insulin-like growth factor 1 receptor (IGF-1R)/stemness and
metabolic changes in hepatocellular carcinoma. Cancers (Basel).
15:9312023. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Stuard WL, Titone R and Robertson DM: The
IGF/Insulin-IGFBP axis in corneal development, wound healing, and
disease. Front Endocrinol (Lausanne). 11:242020. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Zeng B, Liu L, Liao X and Zhang C:
Cardiomyocyte protective effects of thyroid hormone during
hypoxia/reoxygenation injury through activating of IGF-1-mediated
PI3K/Akt signalling. J Cell Mol Med. 25:3205–3215. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Icard P, Coquerel A, Wu Z, Gligorov J,
Fuks D, Fournel L, Lincet H and Simula L: Understanding the central
role of citrate in the metabolism of cancer cells and tumors: An
update. Int J Mol Sci. 22:65872021. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Mafi S, Mansoori B, Taeb S, Sadeghi H,
Abbasi R, Cho WC and Rostamzadeh D: mTOR-Mediated Regulation of
Immune Responses in Cancer and Tumor Microenvironment. Front
Immunol. 12:7741032022. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Nwabo Kamdje AH, Seke Etet PF, Kipanyula
MJ, Vecchio L, Tagne Simo R, Njamnshi AK, Lukong KE and Mimche PN:
Insulin-like growth factor-1 signaling in the tumor
microenvironment: Carcinogenesis, cancer drug resistance, and
therapeutic potential. Front Endocrinol (Lausanne). 13:9273902022.
View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Stefani C, Miricescu D, Stanescu-Spinu II,
Nica RI, Greabu M, Totan AR and Jinga M: Growth factors,
PI3K/AKT/mTOR and MAPK signaling pathways in colorectal cancer
pathogenesis: Where are we now? Int J Mol Sci. 22:102602021.
View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Zhu E, Liu Y, Xie S, Hou J, Yang X, Xu M,
Yang F, Li Z, Zhu B and Zha H: IGF2-IGF1R signaling inhibition
delays the growth of IGF2-high colorectal cancer by modulating
MDSCs. Biochem Biophys Res Commun. 746:1512302025. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Wan M, Mei J, Cai Y, Zhou J, Xue N, Jiang
Y, Zhang Y, Huang J and Zhu Y: Targeting IGF1R overcomes armored
and cold tumor microenvironment and boosts immune checkpoint
blockade in triple-negative breast cancer. Adv Sci (Weinh).
12:e013412025. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Somri-Gannam L, Meisel-Sharon S,
Hantisteanu S, Bar-Noy T, Sigal E, Groisman G, Hallak M, Werner H
and Bruchim I: IGF1R inhibition and PD-1 blockade improve
anti-tumor immune response in epithelial ovarian cancer. Front
Oncol. 14:14104472024. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Heitzeneder S, Sotillo E, Shern JF,
Sindiri S, Xu P, Jones R, Pollak M, Noer PR, Lorette J, Fazli L, et
al: Pregnancy-associated plasma protein-A (PAPP-A) in ewing
sarcoma: Role in tumor growth and immune evasion. J Natl Cancer
Inst. 111:970–982. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Alfaro-Arnedo E, Lopez IP, Pineiro-Hermida
S, Canalejo M, Gotera C, Sola JJ, Roncero A, Peces-Barba G,
Ruiz-Martinez C and Pichel JG: IGF1R acts as a cancer-promoting
factor in the tumor microenvironment facilitating lung metastasis
implantation and progression. Oncogene. 41:3625–3639. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Hanzelmann S, Castelo R and Guinney J:
GSVA: Gene set variation analysis for microarray and RNA-seq data.
BMC Bioinformatics. 14:72013. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Bindea G, Mlecnik B, Tosolini M,
Kirilovsky A, Waldner M, Obenauf AC, Angell H, Fredriksen T,
Lafontaine L, Berger A, et al: Spatiotemporal dynamics of
intratumoral immune cells reveal the immune landscape in human
cancer. Immunity. 39:782–795. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Kang J, Guo Z, Zhang H, Guo R, Zhu X and
Guo X: Dual inhibition of EGFR and IGF-1R signaling leads to
enhanced antitumor efficacy against esophageal squamous cancer. Int
J Mol Sci. 23:103822022. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Kumar S and Chaudhri S: Recent update on
IGF-1/IGF-1R signaling axis as a promising therapeutic target for
triple-negative breast cancer. Pathol Res Pract. 263:1556202024.
View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Zhao Q and Shen J: Insulin-like growth
factor-neutralizing antibodies for cancer therapy. J Leukoc Biol.
117:qiaf1222025. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Shen HT, Chien PJ, Sheu GT, Wang BY and
Chang WW: Reciprocal regulation between B lymphoma Mo-MLV insertion
region 1 homolog and type I insulin-like growth factor receptor in
pemetrexed-resistant lung cancer cells. Tzu Chi Med J. 37:285–292.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Cao D, Lei Y, Ye Z, Zhao L, Wang H, Zhang
J, He F, Huang L, Shi D, Liu Q, et al: Blockade of IGF/IGF-1R
signaling axis with soluble IGF-1R mutants suppresses the cell
proliferation and tumor growth of human osteosarcoma. Am J Cancer
Res. 10:3248–3266. 2020.PubMed/NCBI
|
|
126
|
Yang K, Hu Y, Feng Y, Li K, Zhu Z, Liu S,
Lin Y and Yu B: IGF-1R mediates crosstalk between nasopharyngeal
carcinoma cells and osteoclasts and promotes tumor bone metastasis.
J Exp Clin Cancer Res. 43:462024. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Liu Y, Yu S, Xu T, Bodenko V, Orlova A,
Oroujeni M, Rinne SS, Tolmachev V, Vorobyeva A and Graslund T:
Preclinical evaluation of a new format of (68)Ga- and
(111)In-labeled affibody molecule Z(IGF-1R:4551) for the
Visualization of IGF-1R expression in malignant tumors using PET
and SPECT. Pharmaceutics. 14:14752022. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Choi JH and Park JY: Insulin-like growth
factor-1 receptor targeted fluorescent imaging for gallbladder
cancer in orthotopic mouse models. Gut Liver. 16:606–612. 2022.
View Article : Google Scholar :
|
|
129
|
Molina ER, Chim LK, Lamhamedi-Cherradi SE,
Mohiuddin S, McCall D, Cuglievan B, Krishnan S, Porter RW, Ingram
DR, Wang WL, et al: Correlation of nuclear pIGF-1R/IGF-1R and
YAP/TAZ in a tissue microarray with outcomes in osteosarcoma
patients. Oncotarget. 13:521–533. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Gennari A, Foca F, Zamarchi R, Rocca A,
Amadori D, De Censi A, Bologna A, Cavanna L, Gianni L, Scaltriti L,
et al: Insulin-like growth factor-1 receptor (IGF-1R) expression on
circulating tumor cells (CTCs) and metastatic breast cancer
outcome: Results from the TransMYME trial. Breast Cancer Res Treat.
181:61–68. 2020. View Article : Google Scholar : PubMed/NCBI
|