Introduction
Hepatocellular carcinoma (HCC) is one of the most
prevalent types of cancer and represents a notable global public
health challenge. It is the sixth most common type of cancer and
the third leading cause of cancer-related deaths worldwide, with
~905,000 new cases and 830,000 deaths reported in 2020 (1,2).
The progression of liver cancer from a normal liver typically
involves three stages: Hepatitis, liver cirrhosis and liver cancer.
Key risk factors for these diseases include alcohol consumption,
viral infection, aflatoxin exposure and smoking. The treatment
options for HCC mainly include surgical resection, chemotherapy,
targeted therapy and local embolization (3,4).
Despite the availability of these treatments, high recurrence and
mortality rates persist. Specifically, the 5-year recurrence rate
is >70% following resection, and the overall 5-year survival
rate is <20%. These poor outcomes are primarily due to
late-stage diagnosis and limited effective treatment options
(1). Moreover, recurrence and
distant metastasis are notable contributors to treatment failure in
advanced liver cancer (5,6). Thus, understanding the molecular
mechanisms driving HCC development is key.
Small nucleolar RNAs (snoRNAs) serve a central role
in the post-transcriptional modification and maturation of
ribosomal RNAs (rRNAs) and other cellular RNAs (7). Rapid cancer cell proliferation
requires the production of large quantities of ribosomes to meet
the biosynthetic demands (8,9).
Elevated ribosome production is not a passive consequence of
proliferation, but actively drives cancer progression.
Consequently, key regulators of ribosome assembly, such as snoRNAs
and their associated proteins, are key players in oncogenesis
(10,11). snoRNAs are implicated in various
cancers, including breast and colorectal cancer, pancreatic ductal
carcinoma and HCC (12-18). Small nuclear RNAs, such as U1-U7,
are named for their high uridine content and primarily function in
pre-mRNA splicing. By contrast, snoRNAs, including U3 snoRNA, are
predominantly localized in the nucleolus and guide the modification
and maturation of 18S rRNA (19,20). rRNA processing 9 (RRP9), also
known as U3-55K, is a core subunit of the U3-snoRNP complex
(21). Previous studies have
demonstrated that RRP9 promotes the development of numerous types
of cancer, including pancreatic, colorectal and breast cancer
(22-24). The AKT signaling pathway is
frequently activated in cancer and RRP9 has been shown to activate
the AKT pathway in pancreatic and breast cancer (22,23). RRP9 may promote HCC proliferation
through AKT pathway activation (22-24). Given the oncogenic role of RRP9 in
other cancers and the key role of dysregulated ribosome biogenesis
in HCC, RRP9 may also play a pivotal role in HCC pathogenesis.
Thus, the present study aimed to systematically investigate the
biological functions of RRP9 in HCC and determine whether its
mechanisms align with or differ from those in other types of
cancer, thereby identifying potential tissue-specific therapeutic
targets.
Materials and methods
Collection and processing of HCC
specimens
A total of 216 patients were included in the present
study. These patients (176 male and 40 females; age range, 23-77
years) were recruited from Shouyi Campus and East Campus of the
Renmin Hospital of Wuhan University (Wuhan, China). To account for
potential demographic variations between the two campuses, they
were divided into two independent cohorts: Cohort 1 comprised 131
patients (110 males and 21 female; age range, 23-77 years), and
cohort 2 comprised 85 patients (66 males and 19 females; age range,
28-75 years). Paired tumorous and matched adjacent non-tumorous
tissues (located at least 3 cm away from the tumor margin) were
collected from these patients who underwent surgical resection at
the Renmin Hospital of Wuhan University (Wuhan, China) from January
2022 to December 2023. The inclusion criteria were as follows: i)
Pathologically confirmed primary HCC; and ii) availability of
complete clinicopathological and follow-up data. The exclusion
criteria were: i) Patients who had received preoperative
anti-cancer treatments (such as chemotherapy, radiotherapy,
targeted therapy, or local embolization); and ii) presence of other
primary malignancies. The present study was conducted in accordance
with the principles of the Declaration of Helsinki and was approved
by the Ethics Committee of Renmin Hospital of Wuhan University
(Wuhan, China; approval no. WDRY2022-K013). Written informed
consent was obtained from all patients prior to surgery and sample
collection.
Cell culture and transfection
Huh7 (cat. no. CL-0120) and Hep-3B (cat. no.
CL-0102) cell lines were purchased from Wuhan Pricella
Biotechnology Co. MHCC 97h (cat. no. CBP60227) and HLF (cat. no.
CBP60589) cell lines were purchased from Nanjing Kebai
Biotechnology Co., Ltd. Snu449 (cat. no. CC0105) cells were
purchased from Saiku Biotechnology Co. 97h is a highly metastatic
cell line derived from a patient with HCC lung metastasis. Huh7 is
a well-differentiated, non-metastatic cell line with mutant p53.
Hep-3B is a poorly differentiated cell line deficient in p53
expression and harboring an integrated hepatitis B virus genome.
HLF is a highly metastatic cell line with mutant p53, while Snu449
is derived from a primary tumor and also carries a mutant p53. 97h,
Huh7, Hep-3B and HLF cells were cultured in DMEM supplemented with
10% FBS, while Snu449 cells were maintained in RPMI-1640 medium
(all Gibco; Thermo Fisher Scientific, Inc.) with 10% FBS. All cell
lines were incubated under standard conditions (37°C, 5%
CO2).
For lentiviral production, 293T cells (Wuhan
Pricella Biotechnology; cat. no. CL-0001) were co-transfected with
10 μg 2nd-generation RRP9, CCNA2 or their respective
negative control lentiviral plasmids and helper plasmids (psPAX2
and pMD2.G; ratio 4:3:1; MiaoLing Plasmid) using Lipo8000™
(Beyotime Biotechnology; cat. no. C0533) at 37°C for 10 h. Viral
particles collected at 48 h were used to infect HCC cells at an MOI
of 10 for 24 h at 37°C with 5% CO2. Stable transductants
were selected with 10 μg/ml puromycin, maintained at 2
μg/ml and used for subsequent experiments 7 days
post-transduction.
740 Y-P is a cell-permeable peptide that mimics a
tyrosine-phosphorylated segment of the platelet-derived growth
factor receptor, thereby directly activating PI3K by binding its
SH2 domains (34,35). Cells were treated with 20
μM 740 Y-P or 10 μM PI3K/AKT/mTOR-IN-2 for 24 h at
37°C (25,26).
Immunohistochemistry
Tissue specimens, fixed in 4% paraformaldehyde at
room temperature for 24 h, were embedded in paraffin and cut into
1.5 μm thick sections. The sections were deparaffinized in
xylene and rehydrated through a descending alcohol series. Antigen
retrieval was performed by heating in Tris-EDTA antigen repair
solution (pH 9.0) at 95°C for 15 min. Endogenous peroxidase
activity was quenched with 3% hydrogen peroxide for 15 min,
followed by membrane permeabilization using 0.2% Triton X-100 for
45 min. Non-specific binding was blocked with 10% donkey serum
(Beijing Solarbio Biotechnology; cat. no. SL050) at room
temperature for 1 h. Subsequently, sections were incubated at 4°C
for 12-16 h with primary antibodies, including anti-RRP9 (Santa
Cruz Biotechnology; 1:50; cat. no. sc-100592) or Ki-67 (Wuhan
Pricella Biotechnology; 1:50; cat No. 27309-1-AP). Sections were
incubated with an HRP-conjugated secondary antibody (Thermo Fisher
Scientific Inc.; 1:500; cat. no. 31460) using the MaxVision
detection reagent at room temperature for 30 min. Immunoreactivity
was visualized using a DAB Chromogen detection kit (Abcam). Nuclei
were counterstained with hematoxylin at room temperature for 5 min.
Slides were examined and imaged using a light microscope (Olympus
Corporation), with quantitative analysis performed using ImageJ
software (version 1.8.0; National Institutes of Health).
Tissue microarray
To evaluate RRP9 expression in the clinical cohorts,
a tissue microarray was constructed using the collected
formalin-fixed, paraffin-embedded (FFPE) HCC and matched adjacent
non-tumor tissues. Due to the large sample size exceeding the
maximum capacity of a single recipient block, the cohort samples
were distributed across three independent TMA blocks to preserve
tissue integrity and ensure high-quality sectioning. To minimize
technical variations and ensure direct comparability of the data,
all three TMA blocks were prepared, sectioned, and
immunohistochemically stained simultaneously using an identical
protocol.
TUNEL assay
To evaluate apoptosis in the xenograft tumor
tissues, TUNEL) assay was performed using a TUNEL Apoptosis Assay
Kit (Beyotime Biotechnology; cat. no. C1086) according to the
manufacturer's instructions.
Western blotting
Total protein was extracted from cultured cells and
HCC tissues using RIPA lysis buffer (Beyotime Biotechnology)
supplemented with phosphatase and protease inhibitors (including
PMSF). Protein concentrations were determined using a BCA protein
assay kit (Thermo Fisher Scientific). Equal amounts of protein (20
μg per lane) were separated via 10% SDS-PAGE. Proteins were
then electrotransferred onto PVDF membranes. The membranes were
blocked with 5% non-fat milk for 1 h at room temperature, and
incubated overnight at 4°C with primary antibodies against the
target proteins and the internal reference, RRP9 (Santa Cruz
Biotechnology; 1:50; cat. no. sc-100592); E-cadherin (Proteintech
Group, Inc.; 1:5,000, cat. no. 22018-1-AP), N-cadherin (Proteintech
Group, Inc.; 1:5,000, cat. no. 20874-1-AP), Vimentin (Proteintech
Group, Inc.; 1:5,000, cat. no. 10366-1-AP), Snail-1 (Proteintech
Group, Inc.; 1:5,000, cat. no. 13099-1-AP), Bax (Proteintech Group,
Inc.; 1:5,000, cat. no. 50599-2-Ig), Bcl-2 (Proteintech Group,
Inc.; 1:5,000, cat. no. 12789-1-AP), PI3K (Proteintech Group, Inc.;
1:5,000, cat. no. 20584-1-AP), AKT (Proteintech Group, Inc.;
1:5,000, cat. no. 10176-2-AP), mTOR (1:5,000, cat. no. 20657-1-AP)
and GAPDH (1:5,000; cat. no. 60004-1-Ig; all Proteintech Group,
Inc). Following three washes with TBST containing 0.1% Tween-20,
the membranes were incubated with HRP-conjugated secondary antibody
(1:5,000; Proteintech Group, Inc.), HRP-Goat Anti-Mouse Secondary
Antibody (Proteintech Group, Inc.; 1:5,000, cat. no. RGAR001) and
HRP-Goat Anti-Rabbit Secondary Antibody (Proteintech Group, Inc.;
1:5,000, cat. no. RGAM001) for 1 h at room temperature. Protein
bands were visualized using an enhanced chemiluminescence kit
(Proteintech Group, Inc.). Densitometric analysis was performed
using ImageJ software (version 1.8.0).
Co-immunoprecipitation
Cells were lysed in cold IP buffer (Beyotime
Biotechnology; cat. no. P0013) containing protease inhibitors.
Total cell lysates were incubated with primary antibodies against
RRP9, CCNA2, or normal IgG (as a negative control) at 4°C
overnight. Protein A/G magnetic (Beyotime Biotechnology; cat. no.
P2108; 25 μl) beads were added to capture the immune
complexes. After washing three times with lysis buffer, the
precipitated proteins were eluted by boiling in SDS loading buffer
and analyzed by western blotting.
Cell Counting Kit (CCK)-8 assay
After establishing stable RRP9-overexpression and
RRP9-knockdown models, the cells were seeded into 96-well culture
plates at an initial density of 8×103 cells/well in
quadruplicate. The cells were cultured in DMEM supplemented with
10% FBS and incubated under standard conditions (37°C, 5%
CO2). CCK-8 solution (Biosharp Life Sciences) was added
at 0, 24, 48 and 72 h according to the manufacturer's protocol.
Following 120 min incubation, optical density was measured at 450
nm using a multi-mode microplate reader.
Colony formation assay
Stable RRP9-overexpressing and -knockdown and
control HCC cells were plated in 6-well culture dishes at a density
of 1,000 cells/well and maintained for 14 days under standard
conditions. Cells were fixed with 4% paraformaldehyde for 15 min at
room temperature. The colonies were stained with 2% crystal violet
for 20 min at room temperature, followed by rinsing with distilled
water. Images were captured using a digital imaging system. The
number of colonies (clusters of >50 cells) was quantified using
ImageJ software (version 1.8.0).
Transwell assay
To evaluate cell migration and invasion, Transwell
chambers with 8.0 μm porous membranes (Corning, Inc.) were
used. For the migration assay, 4×105 cells in serum-free
medium [DMEM (Thermo Fisher Scientific Inc.; cat. no. 11965092) for
HLF, 97h, and Huh7 cells; RPMI-1640 (Thermo Fisher Scientific Inc.;
cat. no. 11875119) for Snu449 cells] was added to the upper
chamber, while the lower chamber was filled with culture medium
supplemented with 10% FBS. In the invasion assay, the upper chamber
was pre-coated with Matrigel at 37°C for 2 h prior to seeding
5×105 cells in the aforementioned serum-free media; the
lower chamber contained medium with 10% FBS as a chemoattractant.
After 24-48 h incubation at 37°C, cells that had migrated through
the membrane were fixed with 4% paraformaldehyde at room
temperature for 20 min. The cells were stained with 0.1% crystal
violet solution at room temperature for 20 min and quantified using
a light microscope.
Wound healing assay
Migration of Snu449, HLF, 97h and Huh7 cells was
evaluated using a wound healing assay. A uniform scratch was made
in a cell monolayer at >90% confluence using a sterile pipette
tip. Following three washes with PBS, the cells were maintained in
serum-free medium [HLF, 97h and Huh7 cell cultivated in DMEM,
Snu449 cultivated in RPMI-1640 (Thermo Fisher Scientific Inc.; cat.
no. 11875119)] for 48 h at 37°C. Images were captured at 0 and 48 h
using a light microscope (Olympus). The relative wound closure rate
was determined by measuring the wound width at both time points and
calculating the percentage reduction using Adobe Photoshop software
(version 26.8.0; Adobe Systems, Inc.).
RNA sequencing
Total RNA was extracted from HLF cells of the
wild-type (WT) and RRP9-overexpressing (RRP9-OE) groups using
TRIzol. Three independent biological replicates were performed for
each group to ensure statistical reliability. RNA quality and
quantity were assessed using a Fragment Analyzer, Agilent 2100
Bioanalyzer (Agilent Technologies, Inc.), or Qseq-400 (BiOptic,
Inc.). The DESeq2 software package (v1.34.0) was employed to
identify differentially expressed genes (DEGs) for subsequent
investigations. Detailed RNA sequencing procedures, including
extraction, library preparation, quality control, and data
analysis, are provided in Supplementary Material 1. Venn diagrams
were generated using the VennDiagram R package (version_1.8;
github.com/) and Gene Ontology (GO; geneontology.org/) and Kyoto Encyclopedia of Genes and
Genomes (KEGG; kegg.jp/) enrichment analyses were
performed using the cluster-Profiler R package (version 4.18.4;
https://github.com/).
Protein-protein interaction (PPI) network
construction
Differentially expressed genes (DEGs) [adjusted
P-value <0.05 and |log2(fold-change)|>1] identified by RNA
sequencing were imported into the STRING database (cn.string-db.org/) with a minimum interaction score of
0.9. The data were processed in Cytoscape 3.10.3 (cytoscape.org/), and core targets of the PPI network
were identified by calculating the degree values.
Xenograft tumor model in nude mice
Female immunocompromised BALB/c nude mice (age, 4
weeks; weight, 16-18 g) were obtained from Bainter Biotechnology
(Wuhan, China). The mice were housed under specific pathogen-free
conditions at 22±2°C and 50±10% humidity, with a 12-h light/dark
cycle and ad libitum access to food and water. The animals
(n=18) were randomly divided into three experimental groups (n=6):
WT, OE and OE + 740 Y-P. Tumor xenografts were generated by
subcutaneously implanting 5×106 HLF cells suspended in
100 μl PBS into the right flank region. Tumor growth was
monitored weekly, with specimens collected after 4 weeks. Tumor
dimensions were recorded and volumes were calculated using the
standard ellipsoid formula: V=(L × W2)/2, where L is the
longest diameter and W is the perpendicular width. The humane
endpoints included a maximum tumor volume of 1,500 mm3,
severe tumor ulceration or >20% loss of initial body weight.
During the 4-week experimental period, none of the mice reached
these humane endpoints, and no animals were prematurely euthanized.
All procedures involving laboratory animals were conducted in
accordance with protocols approved by the Animal Research Ethics
Committee of Wuhan University People's Hospital (Wuhan, China;
approval no. WDRM20250303).
Statistical analysis
The experimental data were processed and visualized
using SPSS software (version 22.0; IBM Corp.) and GraphPad Prism
(version 9.0; Dotmatics). Comparisons between two groups were
performed using paired or unpaired t-test. For comparisons
involving >2 groups, a one-way ANOVA followed by Tukey's
honestly significant difference post hoc test was used.
χ2 test was used to analyze the association between
target gene expression and clinicopathological characteristics.
Overall survival curves were plotted using the Kaplan-Meier method,
and the differences between the survival curves were evaluated
using the log-rank test. All data are presented as the mean ±
standard deviation of ≥3 independent experiments. P<0.05 was
considered to indicate a statistically significant difference.
Results
alidation of RRP9 expression in clinical
samples
The expression of RRP9 in the samples was assessed
using western blotting, revealing lower RRP9 expression in cancer
compared with adjacent non-tumor tissues (Fig. 1A). Tissue microarray analysis of
both cohorts showed reduced RRP9 expression in HCC tissues compared
with adjacent non-tumor tissue (Figs.
1B and S1), consistent with
western blot results. Based on RRP9 expression in HCC tissue, the
patients were categorized into high- and low-expression groups
using the median score of 133 as the cut-off value. The demographic
and clinicopathological characteristics of these groups are
summarized in Table I. Lower RRP9
expression in HCC was associated with more tumor nodules, Barcelona
Clinic Liver Cancer stage, tumor size and Tumor-Node-Metastasis
stage (27,28) (Table
I). Kaplan-Meier survival analysis revealed that patients with
higher RRP9 expression had significantly longer overall survival
(Fig. 1C).
 | Table IClinicopathological characteristics
of patients with low- and high-RRP9 expression in primary liver
cancer tissue. |
Table I
Clinicopathological characteristics
of patients with low- and high-RRP9 expression in primary liver
cancer tissue.
| Characteristic | Cohort 1 (n=131)
| Cohort 2 (n=85)
| P-value
|
|---|
| Low | High | Low | High | Group 1 | Group 2 |
|---|
| Cases | 61 | 70 | 40 | 45 | | |
| Age, years | | | | | 0.7982 | 0.5352 |
| ≤60 | 45 | 53 | 25 | 31 | | |
| >60 | 16 | 17 | 15 | 14 | | |
| Sex | | | | | 0.2889 | 0.9755 |
| Male | 49 | 61 | 31 | 35 | | |
| Female | 12 | 9 | 9 | 10 | | |
| Diagnosis | | | | | 0.8398 | 0.3429 |
| HCC | 58 | 66 | 40 | 44 | | |
| ICC | 3 | 4 | 0 | 1 | | |
| Child-Pugh
score | | | | | 0.0333 | 0.0001 |
| A | 49 | 65 | 11 | 31 | | |
| B | 12 | 5 | 29 | 14 | | |
| BCLC stage | | | | | 0.0092 | 0.0010 |
| A | 33 | 55 | 9 | 25 | | |
| B | 8 | 6 | 14 | 15 | | |
| C | 20 | 9 | 17 | 5 | | |
| Number of
tumors | | | | | 0.0013 | 0.0131 |
| ≤3 | 22 | 45 | 12 | 33 | | |
| >3 | 39 | 25 | 28 | 12 | | |
| Tumor diameter,
cm | | | | | 0.0139 | 0.0005 |
| ≤5 | 13 | 29 | 9 | 27 | | |
| >5 | 48 | 41 | 31 | 18 | | |
| TNM stage | | | | | 0.0144 | 0.0089 |
| I | 31 | 54 | 10 | 18 | | |
| II | 1 | 1 | 5 | 15 | | |
| III | 25 | 14 | 15 | 7 | | |
| IV | 4 | 1 | 10 | 5 | | |
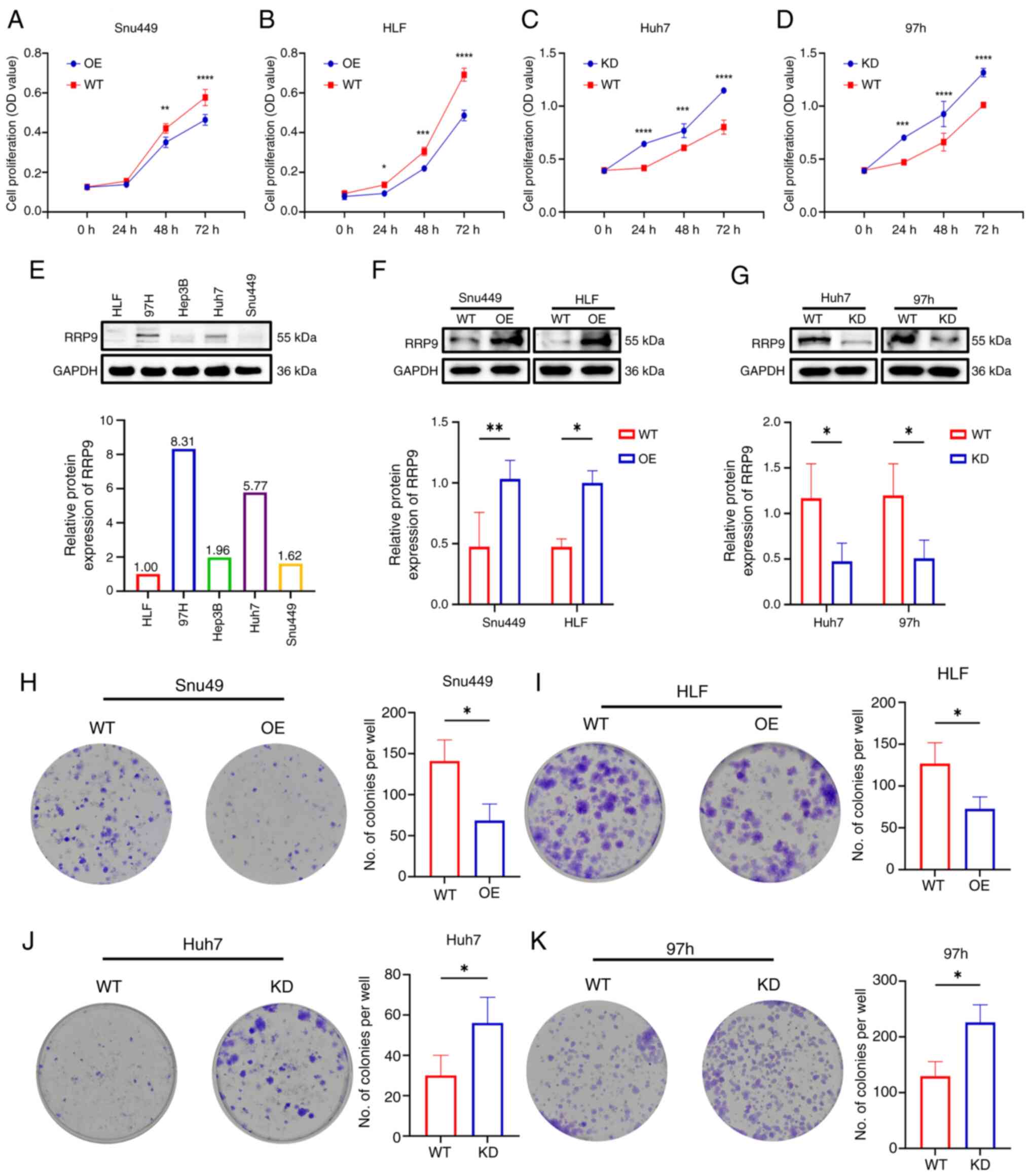
RRP9 inhibits HCC cell proliferation,
migration and invasion
Western blot analysis was performed to assess RRP9
expression in HCC cell lines. RRP9 levels were lower in HLF and
Snu449 cells, while higher levels were observed in Huh7 and 97h
cells (Fig. 2E). RRP9 was
overexpressed in cell lines with low basal expression (HLF and
Snu449) to evaluate its tumor-suppressive potential, and knocked
down in high-expressing lines (Huh7 and 97H) to assess its
tumor-promoting effects. Western blotting confirmed the successful
construction of these cell lines (Figs. 2F and G, S3A-D). The CCK-8 assay demonstrated
RRP9-OE reduced short-term proliferative activity in Snu449 and HLF
cells (Fig. 2A and B), while RRP9
knockdown increased short-term proliferative activity in Huh7 and
97h cells (Fig. 2C and D).
RRP9-OE inhibited colony formation (Fig. 2H and I), while RRP9 knockdown
promoted colony formation (Fig. 2J
and K). These results suggested that RRP9 can inhibit
short-term and long-term proliferation of HCC cells.
The effect of RRP9 on the migration and invasion of
HCC cells was evaluated using wound healing and Transwell assay.
Wound healing assays showed that RRP9-OE inhibited migration in
Snu449 and HLF cells, while RRP9 knockdown enhanced migration in
Huh7 and 97h cells (Fig. 3A-D).
Transwell assay confirmed RRP9-OE inhibited migration and invasion
in Snu449 and HLF cells, whereas RRP9 knockdown decreased migration
and invasion in Huh7 and 97h cells (Fig. 3E-H).
RRP9 promotes apoptosis and inhibits
epithelial-mesenchymal transition (EMT) in HCC
EMT marker expression was analyzed by western
blotting, revealing that RRP9-OE decreased the levels of Snail-1,
N-cadherin and vimentin in Snu449 and HLF cell lines, while
increasing E-cadherin expression (Fig. 4A). By contrast, RRP9 knockdown in
Huh7 and 97H cells significantly increased the expression of
Snail-1, N-cadherin, and vimentin, while decreasing E-cadherin
levels (Fig. 4B). To investigate
the impact of RRP9 on apoptosis, the expression of Bax and Bcl-2
was measured. The results demonstrated a decrease in the expression
of anti-apoptotic factor Bcl-2 and an increase in the pro-apoptotic
factor Bax in Snu449 and HLF cells following RRP9 OE (Fig. 4C). Conversely, RRP9 knockdown in
Huh7 and 97H cells significantly increased the expression of the
anti-apoptotic factor Bcl-2 and decreased the expression of the
pro-apoptotic factor Bax (Fig.
4D). These results suggested that RRP9 suppressed HCC cell
invasion and migration, promoted apoptosis and inhibited EMT.
 | Figure 4Effect of RRP9 on apoptosis- and
EMT-associated protein in HCC. (A) Western blot analysis of
EMT-associated proteins (E-cad, N-cad, vimentin, and Snail-1) in
Snu449 and HLF cells following RRP9 overexpression and (B) Western
blot analysis of EMT-associated proteins (E-cad, N-cad, vimentin,
and Snail-1) in Huh7 and 97h cells following RRP9 knockdown and (C)
apoptosis-related proteins (Bax and Bcl-2) in Snu449 and HLF cells
following RRP9 overexpression and (D) Western blot analysis of
apoptosis-related proteins (Bax and Bcl-2) in Huh7 and 97h cells
following RRP9 knockdown. *P<0.05,
**P<0.01. EMT, epithelial-mesenchymal transition;
HCC, hepatocellular carcinoma; cad, cadherin; WT, wild-type; OE,
overexpression. |
RRP9 regulates HCC progression via the
PI3K/AKT/mTOR signaling pathway
To elucidate the mechanism underlying the action of
RRP9 action in HCC, transcriptome sequencing was performed on WT
and RRP9-OE HLF cells. The sequencing results revealed a number of
DEGs following RRP9 OE (Fig. 5A).
The Venn diagram indicated that 15,926 genes were shared between
the WT and OE groups (Fig. 5B).
Gene Ontology enrichment analysis highlighted significant
enrichment of DEGs in processes associated with the cell cycle,
'sites of DNA damage' and 'extracellular exosome' (Fig. 5C). Kyoto Encyclopedia of Genes and
Genomes enrichment analysis showed that RRP9-OE influenced pathways
such as 'PI3K-Akt signaling', 'pathways in cancer', 'AMPK signaling
pathway', 'mTOR signaling pathway', 'MAPK signaling pathway' and
'Ras signaling pathway' (Fig.
5D). To identify core regulatory nodes within the RRP9-mediated
transcriptomic changes, 465 DEGs were selected for PPI network
analysis. These DEGs were imported into the STRING database and a
core target network containing 25 targets and 96 edges was
constructed using Cytoscape 3.10.3. Cyclin A2 (CCNA2; degree=10)
was associated with RRP9 (Fig.
5E). Given that CCNA2 is a well-established key regulator of
the cell cycle and its dysregulation is frequently implicated in
cancer progression (29-31), it was selected for experimental
validation to explore its functional connection to RRP9 in HCC.
Given the key role of the PI3K/AKT/mTOR signaling
pathway in cancer, the expression of PI3K, AKT, mTOR,
phosphorylated (p-)PI3K, p-AKT and p-mTOR was assessed by western
blotting. The ratios of p-PI3K/PI3K, p-AKT/AKT and p-mTOR/mTOR were
decreased in RRP9-OE Snu449 and HLF cell lines, suggesting
inhibition of the PI3K/AKT/mTOR pathway (Fig. 6A). Conversely, RRP9 knockdown in
Huh7 and 97H cells significantly increased the ratios of
p-PI3K/PI3K, p-AKT/AKT and p-mTOR/mTOR (Fig. 6B). To investigate the association
between RRP9 and CCNA2, proteins were extracted from RRP9-OE,
RRP9-knockdown, and wild-type cells. Western blot analysis revealed
that RRP9-OE decreased CCNA2 protein levels (Fig. 6C), whereas RRP9 knockdown led to
increased CCNA2 expression (Fig.
6D). Co-immunoprecipitation (Co-IP) experiments confirmed that
RRP9 interacted with CCNA2 in both 293T and 97h cell lines
(Fig. S2). Following the
validation of CCNA2 knockdown and overexpression via western
blotting (Fig. S3E and F), we
observed that CCNA2 overexpression increased the ratios of
p-PI3K/PI3K, p-AKT/AKT, and p-mTOR/mTOR, whereas CCNA2 knockdown
exerted the opposite inhibitory effect, demonstrating that CCNA2
positively regulates the PI3K/AKT/mTOR signaling pathway (Fig. S2D and E).
It was hypothesized that RRP9 regulates EMT in HCC
cells via the PI3K/AKT/mTOR pathway. Therefore, Snu449 and HLF
cells were treated with the PI3K activator 740 Y-P and Huh7 and 97h
cells with the PI3K/AKT/mTOR pathway inhibitor PI3K/AKT/mTOR-IN-2.
Western blotting showed that 740 Y-P activated the PI3K/AKT/mTOR
pathway and the inhibitory effect of RRP9-OE on EMT was reversed
following treatment with 740 Y-P (Fig. 7A). To confirm that RRP9 knockdown
regulated EMT via the PI3K/AKT/mTOR pathway, Huh7 and 97h cells
were treated with PI3K/AKT/mTOR-IN-2. Western blotting demonstrated
that PI3K/AKT/mTOR-IN-2 inhibited the promotion of EMT induced by
RRP9 KD (Fig. 7B). These results
suggested that RRP9 regulates EMT in HCC cells through the
PI3K-AKT/mTOR signaling pathway.
 | Figure 7Rescue experiments in cells with RRP9
OE or KD with PI3K/AKT/mTOR pathway activator and inhibitor
treatment. (A) Western blot analysis of PI3K, AKT, mTOR, p-PI3K,
p-AKT, p-mTOR, E-cad, N-cad, vimentin and Snail-1 protein
expression in Snu449 and HLF cells following RRP9 overexpression
and (B) Huh7 and 97h cells following RRP9 knockdown.
*P<0.05, **P<0.01,
***P<0.001. RRP9, ribosomal RNA processing 9; OE,
overexpression; KD, knockdown; p-, phosphorylated-; E-cad,
epithelial cadherin; N-cad, neural cadherin; WT, wild-type; IN-2,
PI3K/Akt/mTOR-IN-2. |
It was hypothesized that RRP9 affected the migration
and invasion of HCC cells via the PI3K/AKT/mTOR signaling pathway.
Snu449 and HLF cells overexpressing RRP9 were treated with 740 Y-P.
The wound healing and Transwell assays revealed that 740 Y-P
reversed the inhibitory effect of RRP9-OE on the migration and
invasion of HCC cells (Fig.
8A-D).
 | Figure 8Rescue experiments using 740 Y-P in
RRP9-OE Snu449 and HLF cells. (A) Wound healing assay evaluating
the migration of Snu449 and (B) Wound healing assay evaluating the
migration of HLF cells. (C) Transwell assays assessing the
migration and invasion of Snu449 cells in the WT, OE, and OE + 740
Y-P groups. (D) Transwell assays assessing the migration and
invasion of HLF cells in the WT, OE, and OE + 740 Y-P groups. Scale
bar, 100 μm. *P<0.05, **P<0.01,
***P<0.001, ****P<0.0001. RRP9,
ribosomal RNA processing 9; OE, overexpression; WT, wild-type. |
Huh7 and 97h cells with RRP9 knockdown were treated
with PI3K/AKT/mTOR-IN-2; wound healing and Transwell assays
indicated that PI3K/AKT/mTOR-IN-2 inhibited the enhanced migratory
and invasive capacity induced by RRP9 knockdown (Fig. 9A-D). In summary, RRP9 regulated
HCC cell migration, invasion and EMT by regulating the
PI3K/AKT/mTOR signaling pathway.
 | Figure 9Rescue experiments using ribosomal
RNA processing 9 KD 97h and Huh7 cells treated with PI3K/AKT/mTOR
pathway inhibitor. (A) Wound healing assay evaluating the migration
of Huh7 and (B) 97h cells in the WT, KD, and KD + IN-2 groups. (C)
Transwell assays assessing the migration and invasion of Huh7 cells
in the WT, KD, and KD + IN-2 groups. (D) Transwell assay of
migration and invasion of 97h cells in the WT, KD, and KD + IN-2
groups. Scale bar, 100 μm. *P<0.05,
**P<0.01. KD, knockdown; IN-2, PI3K/Akt/mTOR-IN-2;
WT, wild-type. |
RRP9 suppresses tumor growth in vivo by
inhibiting the PI3K/AKT/mTOR signaling pathway
To investigate the role of RRP9 in tumor
progression, an in vivo model was constructed. Nude mice
were divided into three groups: WT, OE and OE + 740 Y-P. HLF cells,
which exhibited one of the lowest basal levels of RRP9 protein,
were selected for xenograft experiments to maximize the phenotypic
difference between the OE and WT groups. RRP9-OE significantly
inhibited tumor growth in a subcutaneous xenograft model, with
marked decreases in both tumor weight and volume. However, 740 Y-P
reversed the inhibitory effect of RRP9-OE on tumor growth (Fig. 10A-C, E), suggesting that RRP9
suppressed tumor growth in vivo via the PI3K/AKT/mTOR
signaling pathway. TUNEL staining showed that RRP9 promoted
apoptosis in subcutaneous tumors (Fig. 10D). Western blot analysis
confirmed effective OE of RRP9 in xenograft tumors, with increased
E-cadherin and decreased N-cadherin expression in the OE group
compared with the WT group (Fig.
10F). Immunohistochemical analysis revealed that the OE group
exhibited decreased expression of the proliferation marker Ki-67
and the mesenchymal marker N-cadherin, alongside increased
expression of the epithelial marker E-cadherin, compared with the
WT group (Fig. 10G). These
results indicated that RRP9 effectively suppressed tumor growth
in vivo.
 | Figure 10In vivo validation of RRP9
role in tumor progression. (A) Mice on day 14 post-injection. (B)
Xenograft tumors excised from nude mice. Xenograft (C) tumor weight
and (D) volume. (E) TUNEL staining. (F) Western blot analysis of
RRP9, E-cad and N-cad protein expression in WT and OE groups. (G)
Immunohistochemistry analysis of Ki67, E-cad and N-cad expression
in xenograft tumor tissues. Scale bar, 100 μm.
*P<0.05, **P<0.01,
***P<0.001 ****P<0.0001. RRP9,
ribosomal RNA processing 9; E-cad, epithelial cadherin; N-cad,
neural cadherin; WT, wild-type; OE, overexpression. |
Discussion
Liver cancer, one of the most common and deadly
types of malignancy, is characterized by aggressive biology, rapid
progression and a tendency for early metastasis. Conventional
therapies provide limited benefits, highlighting the need for new
molecular targets and effective treatments (32,33). EMT is a cell reprogramming process
in which epithelial cells lose apical-basal polarity and tight
intercellular junctions, acquiring mesenchymal characteristics
(34). This conversion enhances
cell migratory and invasive ability, confers resistance to
therapeutic agents and allows evasion of immune surveillance
(34). Apoptosis, a highly
regulated form of programmed cell death, serves a key
tumor-suppressive role (35). EMT
and apoptosis are key for cancer promotion and suppression,
respectively, making it crucial to understand the association
between RRP9 and these processes in HCC. snoRNAs are involved in
the post-transcriptional modification and maturation of rRNA, which
is key for ribosome biogenesis. Increasing evidence has implicated
dysregulated snoRNAs in the development of multiple types of
cancer, including HCC, colorectal cancer and pancreatic ductal
adenocarcinoma (18,36,37). snoRNAs are classified into seven
families, U1-U7, based on their high U content (20,38). The present study demonstrated that
RRP9/U3-55K, a core subunit of the U3-snoRNP complex (19), exerts tumor-suppressive activity
in HCC. This is in contrast with its established oncogenic roles in
pancreatic, breast, and colorectal cancer, where it drives
malignant progression by constitutively activating AKT signaling,
promoting chemoresistance and interacting with proteins such as JUN
and DExD-box helicase 21 (22,23,39,40). RRP9 interacts with distinct
signaling molecules or proteins in different cancer contexts,
yielding opposing regulatory outcomes; this highlights the
context-dependent nature of RRP9 functionality and provides a
direction for future research.
To elucidate the mechanism of action of RRP9 in HCC,
HCC cell lines were constructed with either RRP9 overexpression or
knockdown. RRP9 significantly inhibited tumor cell proliferation,
invasion and migration, while simultaneously promoting apoptosis
and suppressing EMT, thereby exerting a notable tumor-suppressive
effect. The PI3K/AKT/mTOR signaling pathway is key for HCC
metabolism and malignant biological behavior such as proliferation,
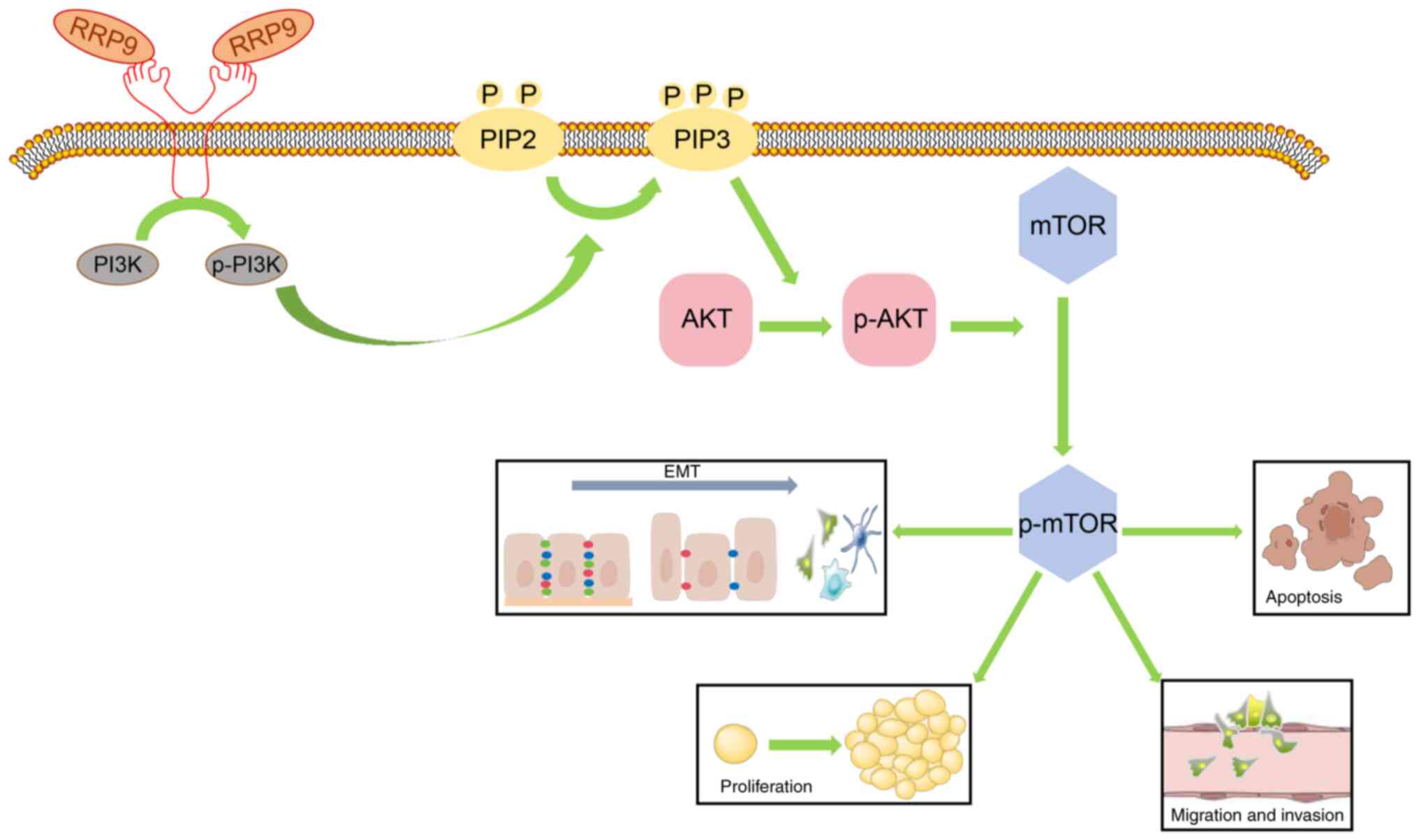
invasion and migration (41).
PI3K activation catalyzes the conversion of phosphatidylinositol
diphosphate to phosphatidylinositol triphosphate on the cell
membrane, which activates AKT. Activated AKT triggers biological
effects by phosphorylating downstream substrates, including mTOR
and glycogen synthase kinase-3β (42) (Fig.
11). Inactivation of this pathway inhibits HCC development
(43,44). RRP9 modulates the invasion,
migration and EMT of HCC cells via the PI3K/AKT/mTOR signaling
pathway. Furthermore, RRP9 was shown to impact HCC progression by
inhibiting this pathway in vivo. Although previous studies
have suggested that RRP9 activates the PI3K/AKT/mTOR pathway
(22,23), the present study observed the
opposite effect. To explore the association between RRP9 and CCNA2,
co-IP was performed, which confirmed that RRP9 interacted with
CCNA2. Furthermore, the present study demonstrated that CCNA2
regulated the PI3K/AKT/mTOR pathway, linking RRP9-mediated effects
to this signaling cascade.
While the present PPI network and co-IP data
suggested an interaction between RRP9 and CCNA2 and western blot
analysis confirmed that RRP9 modulates CCNA2 protein levels, the
precise nature of this regulation requires further investigation.
Future studies should determine whether RRP9 directly regulates
CCNA2 at the transcriptional level by influencing its promoter
activity or at the post-transcriptional level by affecting mRNA
stability. Moreover, the specific mechanism by which CCNA2, a key
regulator of the cell cycle, influences the PI3K/AKT/mTOR signaling
pathway in HCC warrants further exploration. Future studies should
use techniques such as luciferase reporter assays to assess
transcriptional regulation and RNA immunoprecipitation to examine
potential binding to CCNA2 mRNA to determine whether the effects of
RRP9 on the PI3K/AKT/mTOR pathway and HCC progression are
functionally dependent on CCNA2. Clarifying the precise
upstream-to-downstream regulatory sequence and the functional
dependencies within the RRP9/CCNA2/PI3K/AKT/mTOR pathway may
provide more comprehensive understanding of tumor suppression in
HCC.
Given the contrasting oncogenic roles of RRP9
reported in pancreatic, breast and colorectal cancer, its
tumor-suppressive activity in HCC may be shaped by a broader
regulatory landscape (22-24).
In addition to the CCNA2-mediated regulatory axis, other mechanisms
may underlie the context-dependent role of RRP9 in modulating
PI3K/AKT/mTOR signaling in HCC. First, post-translational
modifications of RRP9 affect its function. For example, RRP9 has
been reported to undergo neddylation by the ubiquitin-like modifier
Nedd8 via the E3 ligase Smurf1, which enhances its activity in
tumorigenesis in colorectal cancer models; this demonstrates that
RRP9 function is dynamically regulated by covalent modification and
changes in such modifications may alter downstream signaling
outcomes (45). Moreover,
acetylation and deacetylation of RRP9 modulate its binding to U3
snoRNA and influence ribosome biogenesis, suggesting that
additional post-translational modifications such as
phosphorylation, methylation or ubiquitination may similarly
regulate RRP9 interactions and downstream effects in a cell
type-specific manner (46,47).
Second, tumor- or tissue-specific cofactors and interacting
partners may direct RRP9 toward distinct signaling programs. In
pancreatic cancer, for example, RRP9 interacts with insulin-like
growth factor 2 mRNA-binding protein 1 to activate AKT signaling
and promote chemoresistance, whereas in breast cancer RRP9
interacts with JUN to regulate AKT pathway activity and tumor
progression (23). These
interactions demonstrate how RRP9 engages different RNA-binding
proteins or transcriptional regulators depending on the cell
context, thereby altering the downstream impact on PI3K/AKT/mTOR
and associated pathways (22,23).
Additionally, differences in upstream regulatory
signaling environments across cancer types may shape how RRP9
influences PI3K/AKT/mTOR signaling. The repertoire of activated
receptor tyrosine kinases, metabolic state or viral infection
status (hepatitis B or C in HCC) may change the balance of
interacting signaling nodes available to RRP9 compared with cancer
types lacking these stimuli (48,49). Given that snoRNAs and their
associated proteins are influenced by broader epigenetic and
transcriptional networks in cancer, including other non-coding RNAs
that regulate PI3K/AKT activity, these upstream differences may
contribute to divergent functional outcomes (50,51).
To the best of our knowledge, the present study is
the first to report the tumor-suppressive effect of RRP9, which
contrasts with previous findings (23,52). Additionally, the present study
demonstrated that RRP9 exerted a tumor-suppressive effect by
inhibiting the PI3K/AKT/mTOR pathway. The present study provided
key insights into the role of RRP9 in HCC progression and
highlights its potential as a therapeutic target.
Supplementary Data
Availability of data and materials
The data generated in the present study may be found
in the National Center for Biotechnology Information under
accession number PRJNA1330788 or at the following URL: ncbi.nlm.nih.gov.
Authors' contributions
ZF, WW, JF, KD and ML conceived and designed the
study. ZF, ML and WW confirm the authenticity of all the raw data.
WW and JF provided administrative support. All authors wrote the
manuscript. All authors have read and approved the final
manuscript.
Ethics approval and consent to
participate
The human sample experiments were approved by the
Ethics Committee of Wuhan University People's Hospital (Wuhan,
China; approval no. WDRY2022-K013) and written informed consent was
obtained from all patients prior to participation. The animal
experiments were approved by the Animal Research Ethics Committee
of Wuhan University People's Hospital (approval no.
WDRM20250303).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Acknowledgements
Not applicable.
Funding
The present study was supported by Discipline Construction Funds
of Hubei Province (grant no. CZ2025020003-5), Chen Xiao-ping
Foundation for the Development of Science and Technology of Hubei
Province (grant no. CXPJJH123003-094) and Natural Science
Foundation of Hubei Province (grant no. 2023AFB197).
References
|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249.
2021.PubMed/NCBI
|
|
2
|
Rumgay H, Arnold M, Ferlay J, Lesi O,
Cabasag CJ, Vignat J, Laversanne M, McGlynn KA and Soerjomataram I:
Global burden of primary liver cancer in 2020 and predictions to
2040. J Hepatol. 77:1598–606. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hwang SY, Danpanichkul P, Agopian V, Mehta
N, Parikh ND, Abou-Alfa GK, Singal AG and Yang JD: Hepatocellular
carcinoma: Updates on epidemiology, surveillance, diagnosis and
treatment. Clin Mol Hepatol. 31(Suppl): S228–S254. 2025. View Article : Google Scholar :
|
|
4
|
Toh MR, Wong EYT, Wong SH, Ng AWT, Loo LH,
Chow PK and Ngeow J: Global epidemiology and genetics of
hepatocellular carcinoma. Gastroenterology. 164:766–782. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Anwanwan D, Singh SK, Singh S, Saikam V
and Singh R: Challenges in liver cancer and possible treatment
approaches. Biochim Biophys Acta Rev Cancer. 1873:1883142020.
View Article : Google Scholar
|
|
6
|
Galicia-Moreno M, Silva-Gomez JA,
Lucano-Landeros S, Santos A, Monroy-Ramirez HC and
Armendariz-Borunda J: Liver cancer: Therapeutic challenges and the
importance of experimental models. Can J Gastroenterol Hepatol.
2021:88378112021. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Liang J, Wen J, Huang Z, Chen XP, Zhang BX
and Chu L: Small nucleolar RNAs: Insight into their function in
cancer. Front Oncol. 9:5872019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Pecoraro A, Pagano M, Russo G and Russo A:
Ribosome biogenesis and cancer: Overview on ribosomal proteins. Int
J Mol Sci. 22:54962021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Gaviraghi M, Vivori C and Tonon G: How
cancer exploits ribosomal RNA biogenesis: A journey beyond the
boundaries of rRNA transcription. Cells. 8:10982019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Elhamamsy AR, Metge BJ, Alsheikh HA,
Shevde LA and Samant RS: Ribosome biogenesis: A central player in
cancer metastasis and therapeutic resistance. Cancer Res.
82:2344–2353. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lorenzo HK: Small nucleolar RNAs as
emerging players in cancer biology and precision medicine. Cancers
(Basel). 17:38472025. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gee HE, Buffa FM, Camps C, Ramachandran A,
Leek R, Taylor M, Patil M, Sheldon H, Betts G, Homer J, et al: The
small-nucleolar RNAs commonly used for microRNA normalisation
correlate with tumour pathology and prognosis. Br J Cancer.
104:1168–1177. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Blenkiron C, Hurley DG, Fitzgerald S,
Print CG and Lasham A: Links between the oncoprotein YB-1 and small
non-coding RNAs in breast cancer. PLoS One. 8:e801712013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yang X, Li Y, Li L, Liu J, Wu M and Ye M:
SnoRNAs are involved in the progression of ulcerative colitis and
colorectal cancer. Dig Liver Dis. 49:545–551. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yoshida K, Toden S, Weng W, Shigeyasu K,
Miyoshi J, Turner J, Nagasaka T, Ma Y, Takayama T, Fujiwara T and
Goel A: SNORA21-an oncogenic small nucleolar RNA, with a prognostic
biomarker potential in human colorectal cancer. EBioMedicine.
22:68–77. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wu L, Zheng J, Chen P, Liu Q and Yuan Y:
Small nucleolar RNA ACA11 promotes proliferation, migration and
invasion in hepatocellular carcinoma by targeting the PI3K/AKT
signaling pathway. Biomed Pharmacother. 90:705–712. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li G, He Y, Liu X, Zheng Z, Zhang M, Qin F
and Lan X: Small nucleolar RNA 47 promotes tumorigenesis by
regulating EMT markers in hepatocellular carcinoma. Minerva Med.
108:396–404. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Cui L, Nakano K, Obchoei S, Setoguchi K,
Matsumoto M, Yamamoto T, Obika S, Shimada K and Hiraoka N: Small
nucleolar noncoding RNA SNORA23, up-regulated in human pancreatic
ductal adenocarcinoma, regulates expression of spectrin
repeat-containing nuclear envelope 2 to promote growth and
metastasis of xenograft tumors in mice. Gastroenterology.
153:292–306.e2. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Clerget G, Bourguignon-Igel V,
Marmier-Gourrier N, Rolland N, Wacheul L, Manival X, Charron C,
Kufel J, Méreau A, Senty-Ségault V, et al: Synergistic defects in
pre-rRNA processing from mutations in the U3-specific protein Rrp9
and U3 snoRNA. Nucleic Acids Res. 48:3848–3868. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ojha S, Malla S and Lyons SM: snoRNPs:
Functions in ribosome biogenesis. Biomolecules. 10:7832020.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Beltrame M and Tollervey D: Base pairing
between U3 and the pre-ribosomal RNA is required for 18S rRNA
synthesis. EMBO J. 14:4350–4356. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang Z, Yu H, Yao W, Zhu N, Miao R, Liu
Z, Song X, Xue C, Cai C, Cheng M, et al: RRP9 promotes gemcitabine
resistance in pancreatic cancer via activating AKT signaling
pathway. Cell Commun Signal. 20:1882022. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Huan J, Liu X, Wang N, Mu Y, Li L and Du
Y: The RRP9-JUN axis promotes breast cancer progression via the AKT
signalling pathway. Biol Direct. 19:1312024. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu H, Chi X, Yang N, Shan M, Xiao Y,
Zhang M, Hao Y, Hou S, Liu Y and Wang Y: Joint effect of RRP9 and
DDX21 on development of colorectal cancer and keloid. Aging (Albany
NY). 15:14703–14719. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Qin J, Sun X, Ma Y, Cheng Y, Ma Q, Jing W,
Qu S and Liu L: Design, synthesis and biological evaluation of
novel 1,3,4,9-tetrahydropyrano[3,4-b]indoles as potential treatment
of triple negative breast cancer by suppressing PI3K/AKT/mTOR
pathway. Bioorg Med Chem. 55:1165942022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
An Y, Cao C, Sun S, Wu H, Zhang J, Li R
and Zhao Y: SHP1 and its downstream p38/SP1/PI3K/YAP/Notch-1
signaling in trophoblast cells suppressed the progression of
Preeclampsia via inhibiting proliferation of SMCs. Sci Rep.
15:162052025. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Amin MB, Greene FL, Edge SB, Compton CC,
Gershenwald JE, Brookland RK, Meyer L, Gress DM, Byrd DR and
Winchester DP: The eighth edition AJCC cancer staging manual:
Continuing to build a bridge from a population-based to a more
'personalized' approach to cancer staging. CA Cancer J Clin.
67:93–99. 2017.PubMed/NCBI
|
|
28
|
Reig M, Forner A, Rimola J, Ferrer-Fàbrega
J, Burrel M, Garcia-Criado Á, Kelley RK, Galle PR, Mazzaferro V,
Salem R, et al: BCLC strategy for prognosis prediction and
treatment recommendation: The 2022 update. J Hepatol. 76:681–693.
2022. View Article : Google Scholar
|
|
29
|
Liu T, Shi Q, Yang L, Wang S, Song H, Wang
Z, Xu X, Liu H, Zheng H and Shen Z: Long non-coding RNAs HERH-1 and
HERH-4 facilitate cyclin A2 expression and accelerate cell cycle
progression in advanced hepatocellular carcinoma. BMC Cancer.
21:9572021. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Gan Y, Li Y, Li T, Shu G and Yin G: CCNA2
acts as a novel biomarker in regulating the growth and apoptosis of
colorectal cancer. Cancer Manag Res. 10:5113–5124. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Suski JM, Braun M, Strmiska V and Sicinski
P: Targeting cell-cycle machinery in cancer. Cancer Cell.
39:759–778. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Llovet JM, Kelley RK, Villanueva A, Singal
AG, Pikarsky E, Roayaie S, Lencioni R, Koike K, Zucman-Rossi J and
Finn RS: Hepatocellular carcinoma. Nat Rev Dis Primers. 7:62021.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI
|
|
34
|
Zhang J, Hu Z, Horta CA and Yang J:
Regulation of epithelial-mesenchymal transition by tumor
microenvironmental signals and its implication in cancer
therapeutics. Semin Cancer Biol. 88:46–66. 2023. View Article : Google Scholar :
|
|
35
|
Morana O, Wood W and Gregory CD: The
apoptosis paradox in cancer. Int J Mol Sci. 23:13282022. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Ding Y, Sun Z, Zhang S, Zhou L, Xu Q, Zhou
D, Li Y, Han X, Xu H, Bai Y, et al: Identification of snoRNA
SNORA71A as a novel biomarker in prognosis of hepatocellular
carcinoma. Dis Markers. 2020:88799442020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhang Z, Tao Y, Hua Q, Cai J, Ye X and Li
H: SNORA71A promotes colorectal cancer cell proliferation,
migration, and invasion. Biomed Res Int. 2020:82845762020.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Dragon F, Gallagher JE, Compagnone-Post
PA, Mitchell BM, Porwancher KA, Wehner KA, Wormsley S, Settlage RE,
Shabanowitz J, Osheim Y, et al: A large nucleolar U3
ribonucleoprotein required for 18S ribosomal RNA biogenesis.
Nature. 417:967–970. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Chi X, Yang N and Liu Y: RRP9 and DDX21 as
new biomarkers of colorectal cancer. Medicine (Baltimore).
102:e343842023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Li N, Jing Y, Xu L and Wang M: METTL1
enhances RRP9 mRNA stability through m7G modification to drive
colorectal tumorigenesis. Mol Carcinog. 64:858–869. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Alzahrani AS: PI3K/Akt/mTOR inhibitors in
cancer: At the bench and bedside. Semin Cancer Biol. 59:125–132.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Li Y, Liu Z, Yan H, Zhou T, Zheng L, Wen
F, Guo G and Zhang Z: Polygonatum sibiricum polysaccharide
ameliorates skeletal muscle aging and mitochondrial dysfunction via
PI3K/Akt/mTOR signaling pathway. Phytomedicine. 136:1563162025.
View Article : Google Scholar
|
|
43
|
Chen J, Chen J, Huang J, Li Z, Gong Y, Zou
B, Liu X, Ding L, Li P, Zhu Z, et al: HIF-2α upregulation mediated
by hypoxia promotes NAFLD-HCC progression by activating lipid
synthesis via the PI3K-AKT-mTOR pathway. Aging (Albany NY).
11:10839–10860. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Yang J, Pi C and Wang G: Inhibition of
PI3K/Akt/mTOR pathway by apigenin induces apoptosis and autophagy
in hepatocellular carcinoma cells. Biomed Pharmacother.
103:699–707. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Du MG, Liu F, Chang Y, Tong S, Liu W, Chen
YJ and Xie P: Neddylation modification of the U3 snoRNA-binding
protein RRP9 by Smurf1 promotes tumorigenesis. J Biol Chem.
297:1013072021. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Chen S, Blank MF, Iyer A, Huang B, Wang L,
Grummt I and Voit R: SIRT7-dependent deacetylation of the U3-55k
protein controls pre-rRNA processing. Nat Commun. 7:107342016.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Modenini G, Abondio P and Boattini A: The
coevolution between APOBEC3 and retrotransposons in primates. Mob
DNA. 13:272022. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Chen X, Wang X, Zhu F, Qian C, Xu F, Huang
X, Zhang W and Sun B: HBV infection-related PDZK1 plays an
oncogenic role by regulating the PI3K-Akt pathway and fatty acid
metabolism and enhances immunosuppression. J Immunol Res.
2022:87855672022. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Qu H, Xie Y, Hu S, Sun S, Yuan Y, Xia Y,
Liu M and Zhang XL: HBV upregulates TNNT1 expression through
PI3K/AKT/mTOR-c-Myc axis, which in turn induces EMT and liver
fibrosis in mice. Cell Signal. 134:1118992025. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Hussain MS, Moglad E, Afzal M, Gupta G,
Hassan Almalki W, Kazmi I, Alzarea SI, Kukreti N, Gupta S, Kumar D,
et al: Non-coding RNA mediated regulation of PI3K/Akt pathway in
hepatocellular carcinoma: Therapeutic perspectives. Pathol Res
Pract. 258:1553032024. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Zacchini F, Barozzi C, Venturi G and
Montanaro L: How snoRNAs can contribute to cancer at multiple
levels. NAR Cancer. 6:zcae0052024. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Wang Z, Zhao C, Li M, Zhang L, Diao J, Wu
Y, Yang T, Shi M, Lei Y, Wang Y, et al: Tuina therapy alleviates
knee osteoarthritis by modulating PI3K/AKT/mTOR-mediated autophagy:
An integrated machine learning and in vivo rat study. Front
Immunol. 16:16358182025. View Article : Google Scholar : PubMed/NCBI
|