Introduction
Telomeres are special structures at the ends of the
chromosomes consisting of ‘TTAGGG’ tandem repeat sequences along
with its associated protein complex called ‘shelterin’ (1). As a result of the inability of DNA
polymerases to duplicate the ends of the linear DNA molecules, the
lengths of the human telomeres get shortened to ~50 bp with each
cell division and the attrition of telomere length is counteracted
by telomerase reverse transcriptase (TERT), which is the catalytic
component of the enzyme telomerase (2). It has been reported that TERT is
reactivated in cancer cells and was found to be overexpressed in
tumors (3). Several mechanisms are
associated with TERT reactivation, including TERT promoter
mutations or rearrangements, and copy number amplification and
methylation (4). Cancer cells
survive by exploiting the telomere maintenance mechanism. The
details of cellular immortality through the telomere length
maintenance are poorly understood in many human cancer types,
including cervical cancer.
The prevalence of cervical cancer has increased
rapidly in rural areas and overall it ranks second in both
incidence and mortality rates in India, regardless of its
incidental difference between rural and urban areas (5). In total, 95% of cases are caused by
persistent infection with high-risk human papilloma virus (HPV)
(6). Persistent HPV infection and
viral oncogene expression results in the inactivation of tumor
suppressor genes, including tumor protein 53 and pRb, that
consequently leads to increased genomic instability and
accumulation of mutations, which often results in tumorigenesis
(7). Cervical cancer remains a
serious problem among women, especially in developing nations like
India even after several decades of cervical cancer research that
has identified various therapeutic regiments.
Recent advancement in next generation sequencing has
enabled the whole genome sequencing of tumors with their paired
normal controls. The Cancer Genome Atlas and International Cancer
Genome Consortium studies revealed significant non-coding mutations
and single nucleotide polymorphism (SNPs) in the regulatory regions
of genes associated with tumorigenesis. The pan-cancer analysis of
the whole genome aimed to analyze whole genomes of ~2,500 tumors
and matched normal controls, mainly to identify driver mutations
and to differentiate from passenger mutations (8). The previously identified driver
mutations (C228T and C250T) in the TERT promoter were found to
create a new binding site for the E-twenty six (ETS) family of
transcription factors, resulting in the increased expression of
TERT, which was initially observed in melanomas (9,10) and
later in other cancer types (11). A
previous study also observed the same functional TERT promoter
mutations with high frequency (21.4%) in South Indian cervical
cancer (12).
Previously, an SNP rs2853669 (A>G) in the TERT
promoter was identified in chr5:1,295,349 T>C (T349C) and was
shown to be associated with cancer risk in a different population
(13-15).
The presence of the SNP rs2853669 along with reactivating promoter
mutations has been reported to increase the risk and shown to be
associated with a poor survival rate in hepatocellular carcinoma
(16). At present, the role of this
particular SNP with TERT promoter driver mutations and its genetic
association with HPV in patients with cervical cancer has not been
determined. To understand the genetic association of the TERT SNP
with cervical cancer risk, the present study focused on analyzing
the SNP rs2853669 in cervical cancer samples from South Indian
women and healthy controls. Furthermore, a meta-analysis of the SNP
rs2853669 in various cancer types of world populations was
conducted to elucidate the distribution and risk association of
variant alleles with different types of cancer.
Materials and methods
Study design and subjects
The present study was conducted after obtaining
approval from the Institutional Ethics Committee (approval no.
04092010) of Madras Medical College and Hospital (Chennai, India).
The present study was conducted between January 2011 and June 2015.
The tissue/blood samples were collected at The Institute of Social
Obstetrics and Government Kasturba Gandhi Hospital for Women and
Children (Chennai, India) and Government Royapettah Hospital
(Chennai, India) after obtaining written informed consent from each
patient. The present study included tissue samples from 257
patients with cervical cancer confirmed by histologic examination
of biopsies and curettage specimens in pathology laboratory. For
histologic classification, a two-tiered cervical intraepithelial
neoplasia system was employed (17).
The samples with no reported malignancy in the pathology were
excluded. The biopsy samples were collected during the diagnosis of
the patients and transported to the laboratory in RNAlater solution
(Thermo Fisher Scientific, Inc.). The transported samples were
homogenized and stored in RNAlater solution at 4˚C overnight. On
the subsequent day, RNAlater solution was discarded and DNA was
isolated from the tissues after PBS washes following a standard
phenol-chloroform extraction method. In total, 295 healthy women
were recruited as controls. About 2 ml of blood was collected in
EDTA-coated vacutainers by venipuncture of the dorsal hand veins.
Blood samples were also collected from patients with cervical
cancer to sequence them to confirm the germline nature of the TERT
SNP. The mean age of the patients and controls were 51.2±11.3 and
39.41±10.6 years, respectively. DNA was isolated from all the blood
samples using a standard phenol-chloroform extraction and ethanol
precipitation method (18).
SNP genotyping
To analyze the frequency of the SNP rs2853669 A>G
in patients with cervical cancer and the controls, a hydrolysis
probe-based allelic discrimination assay was performed (Thermo
Fisher Scientific, Inc.). The PCR reaction was carried out in a
total of 5 µl comprised of 10 ng DNA and 2.5 µl 2X TaqMan Universal
PCR master mix (No UNG) and 0.125 µl 40X TaqMan SNP genotyping
assay mix (Assay ID: C-8773290_10; Thermo Fisher Scientific, Inc.).
The reaction was performed in QuantStudio 6 Flex Real-Time PCR
(Thermo Fisher Scientific, Inc.) using a standard protocol (2 min
at 50˚C, 10 min at 95˚C followed by 15 sec at 92˚C and 60 sec at
60˚C for 40 cycles) and the allelic discriminations were conducted
by detecting the fluorescence in the PCR reactions. A control with
no template was included in each plate. Genotype calls of 95%
quality were scored using Sequence Detection Software v2.4.1
(Thermo Fisher Scientific, Inc.).
DNA sequencing
The germline nature of the identified TERT SNP was
confirmed by sequencing 2% parallel blood DNA available from the
patients with cervical cancer and the controls. The human TERT
promoter region was amplified from cervical cancer tissue DNA
samples and sequencing was performed, as previously described
(12). The promoter-specific primers
TERT forward: 5'-TGTAAAACGACGGCCAGTGGCCGATTCGACCTCTCTC-3'
and reverse: 5'-CAGCGCTGCCTGAAACTCG-3' (the underlined sequence in
the TERT F is the M13 universal sequencing primer) were used to
amplify the TERT promoter. Briefly, the thermocycling conditions
for PCR were 5 min at 95˚C once followed by 30 sec at 95˚C, 45 sec
at 60˚C, and 45 sec at 72˚C for 10 cycles and 30 cycles of 30 sec
at 95˚C, 45 sec at 60˚C and 30 sec (with 5 sec increases in each
cycle) at 72˚C and final extension for 7 min at 72˚C. The PCR
products were purified using a commercial kit (Qiagen, Inc.) and
sequenced by the Sanger sequencing method (Macrogen, Inc.). The
representative sequence for all three genotypes is presented in
Fig. S1.
Statistical analysis
The genotype and allele frequency of the cervical
cancer and control groups were estimated using the gene count
method. Hardy-Weinberg Equilibrium (HWE) was assessed by the
goodness-of-fit χ2 test (19). The association between the
polymorphism and cancer was analyzed using a χ2 test.
The odds ratios (ORs) with 95% CIs were calculated to evaluate the
association between the rs2853669 polymorphism and cervical cancer
risk, including variant heterozygous vs. wild-type homozygous
model, and variant homozygous vs. wild-type homozygous model and
dominant model. All tests were two-tailed and P<0.05 was
considered to indicate a statistically significant difference. All
the analyses were performed using SPSS, version 20.0. (IBM,
Corp.).
Meta-analysis
Meta-analysis was performed to identify the
distribution of rs2583669 variant allele across various types of
cancers among different populations. An extensive literature search
was conducted using PubMed (https://www.ncbi.nlm.nih.gov/pubmed/), Embase
(https://www.embase.com/) and Web of Science
(https://clarivate.com/products/web-of-science/)
databases (last search date January 10, 2019). The key words used
in the search included ‘TERT or telomerase reverse transcriptase’,
‘cancer or carcinoma’ and ‘rs2583669 polymorphism’. After the
initial screening, the full text of all relevant papers was
obtained and further filtered in order to fit into the inclusion
criteria: i) Studies examining the association of TERT rs2583669
polymorphisms with cancer; ii) prospective case-control studies;
and iii) studies with information to calculate ORs. Then, two
authors independently extracted the genotype frequencies of the
TERT rs2583669 polymorphism. This meta-analysis evaluated the
association of TERT rs2583669 polymorphism with cancer risk
according to the guidelines issued in the Preferred Reporting Items
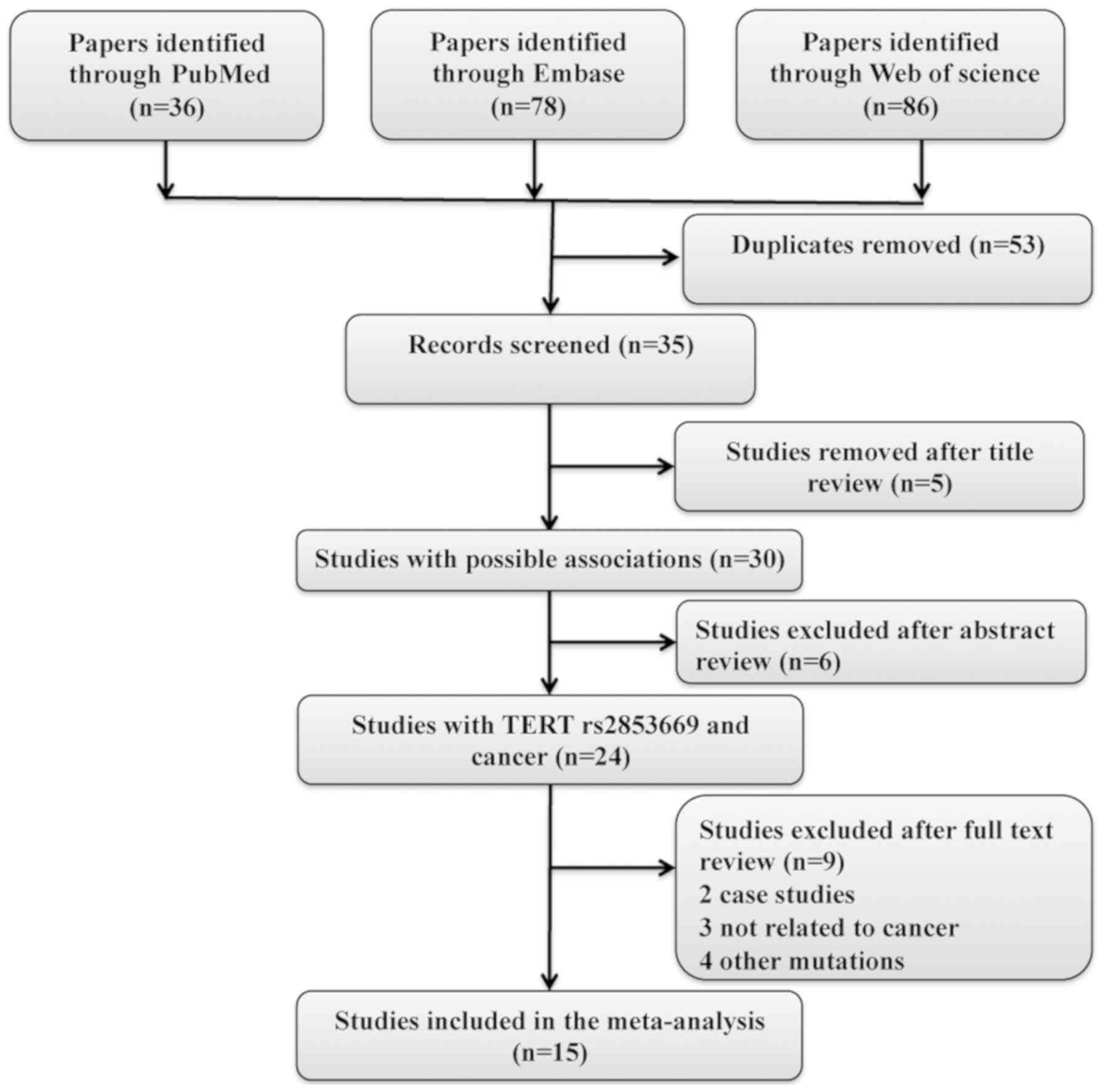
for Systematic Reviews and Meta-Analyses statement (20). A flow-chart describing the present
study and the selection criteria for the meta-analysis is presented
in Fig. 1. All meta-analyses were
conducted using OpenMetaAnalyst software (21). All the previous studies included in
the meta-analysis were evaluated for the HWE using a χ2
test in the control groups and P>0.05 was considered as the
sample neutrality. The carrier risk associations for rs2853669 were
calculated in the allelic, dominant and recessive models using R
(version 3.2.0) statistical software (https://cran.r-project.org/bin/windows/base/old/3.2.0/).
The OR and 95% CIs were illustrated using Forest plot graphical
representation. Furthermore, a sensitivity analysis was conducted
by excluding one study in each analysis to examine the robustness
of the method used for the meta-analysis. The potential publication
bias was assessed by Begg's funnel plot and Egger's test (22).
Results
Frequency of rs2853669 in South Indian
cervical cancer
In the present study, the promoter SNP rs2853669
(-245A>G) was genotyped in 552 samples, consisting of 257
cervical cancer samples and 295 controls from South India. The
distribution of alleles along with various clinical variables is
presented in Table I. The
proportions of genotypes were 23.3% AA, 35.4% AG and 41.3% GG in
the cervical cancer cases, and 21.4% AA, 49.5% AG and 29.1% GG in
the controls (Table II). A higher
incidence [41.25% (106/257)] of homozygous variant ‘GG’ was
identified in the cervical cancer samples compared with the
controls [29.15% (86/295)] in the study population, although
without any statistical significance (P=0.265). The TERT rs2853669
polymorphism showed significant association with cervical cancer in
the recessive model (GG vs. AA+AG; OR=1.71; 95% CI=1.20-2.43;
P=0.003; Table II). The
distribution of TERT rs2853669 polymorphism among TERT activating
promoter mutations in our previous study (C228T and C250T) is
documented in Table III (12). The individuals carrying rs2853669
variant allele (CT+CC) was found in 66.7% (20/30) of TERT promoter
mutation-positive patients and this interaction did not increase
the risk (OR=1.54; 95% CI=0.62-3.67; P=0.33). Furthermore, the
distribution of the TERT rs2853669 polymorphism among patients with
high-risk HPV also did not increase the risk of cervical cancer
(OR=1.54; 95% CI=0.79-3.01; P=0.203; Table IV).
 | Table IClinicopathological characteristics
among the rs2853669 genotypes in patients with cervical cancer. |
Table I
Clinicopathological characteristics
among the rs2853669 genotypes in patients with cervical cancer.
| | | SNP rs2853669 |
|---|
| Clinicopathological
characteristics | Total (%) | AA (%) | AG+GG (%) |
|---|
| No. of cases | 257 | 60 (23.3) | 197 (76.7) |
| Age in years (mean
± SD) | 51.2 ±11.3 | | |
|
<51 | 138 (53.7) | 29 (21.1) | 109 (78.9) |
|
≥51 | 116 (45.1) | 31 (26.7) | 85 (73.3) |
|
Unknown | 3 (1.2) | - | 3(100) |
| Tumor cell
differentiation | | | |
|
Well
differentiated | 54(21) | 11 (20.4) | 43 (79.6) |
|
Moderately
differentiated | 116 (45.1) | 34 (29.4) | 82 (70.6) |
|
Poorly
differentiated | 52 (20.2) | 10 (19.3) | 42 (80.7) |
|
Unknown | 35 (13.6) | 5 (14.7) | 30 (85.7) |
| CIN grade | | | |
|
IB1 or
IB2 | 17 (6.6) | 6 (35.3) | 11 (64.7) |
|
II or IIA2
and IIB | 86 (33.5) | 18 (20.9) | 68 (79.1) |
|
III or
IIIB | 58 (22.6) | 17 (29.3) | 41 (70.7) |
|
IV or
IVB | 3 (1.2) | - | 3(100) |
|
Unknown | 93 (36.2) | 19 (20.4) | 74 (79.6) |
| Infiltration | | | |
|
Infiltrnfiltrated | 208 (80.9) | 50 (24.1) | 158 (75.9) |
 | Table IIAssociation between TERT rs2853669
polymorphism and cervical cancer. |
Table II
Association between TERT rs2853669
polymorphism and cervical cancer.
| TERT SNP rs2853669
A>G | Cervical cancer
(n=257) (Percentage frequency) | Control (n=295)
(Percentage frequency) | OR | 95% CI | P-value |
|---|
| Genotype | | | | | |
|
AA | 60 (23.3) | 63 (21.4) | Reference |
|
AG | 91 (35.4) | 146 (49.5) | 0.66 | 0.42-1.02 | 0.058 |
|
GG | 106 (41.3) | 86 (29.1) | 1.29 | 0.82-2.04 | 0.265 |
| Dominant model
AG+GG vs. AA | 197 (76.7) | 232 (78.6) | 0.89 | 0.60-1.33 | 0.575 |
| Recessive model GG
vs. AA+AG | 106 (41.3) | 86 (29.1) | 1.71 | 1.20-2.43 | 0.003 |
| Allelic model | | | | | |
|
A | 211 (41.1) | 272 (46.1) | Reference |
|
G | 303 (58.9) | 318 (53.9) | 1.23 | 0.97-1.56 | 0.091 |
| HWE
χ2 | 18.51 | 0.005 | | | |
| HWE P-value | <0.001 | 0.944 | | | |
 | Table IIIAssociation of TERT promoter
mutations (C228T and C250T) with rs2853669 (-245T/C) in cervical
cancer from our previous study (12). |
Table III
Association of TERT promoter
mutations (C228T and C250T) with rs2853669 (-245T/C) in cervical
cancer from our previous study (12).
| | TERT mutation
status | | |
|---|
| SNP rs2853669T>C
(n=140) | Wild-type | Mutated | OR (95% CI) | P-value |
|---|
| TT (37) | 27 | 10 | Reference |
| CT+CC (103) | 83 | 20 | 1.54
(0.62-3.67) | 0.333 |
 | Table IVAssociation of HPV and telomerase
reverse transcriptase rs2853669 (-245T/C) in cervical cancer. |
Table IV
Association of HPV and telomerase
reverse transcriptase rs2853669 (-245T/C) in cervical cancer.
| | HPV status | | |
|---|
| SNP rs2853669T>C
(n=257) | Positive (%) | Negative (%) | OR (95% CI) | P-value |
|---|
| TT (60) | 46 (76.7) | 14 (23.3) | Reference |
| CT+CC (197) | 134 (68.0) | 63 (32.0) | 1.54
(0.79-3.01) | 0.203 |
Meta-analysis results
Based on the inclusion criteria, a total of 17
studies from 14 previously published articles (15,23-35)
along with the present study results were analyzed for the
association between the TERT rs2583669 polymorphism and cancer
susceptibility; this included 9,537 cancer cases and 12,370
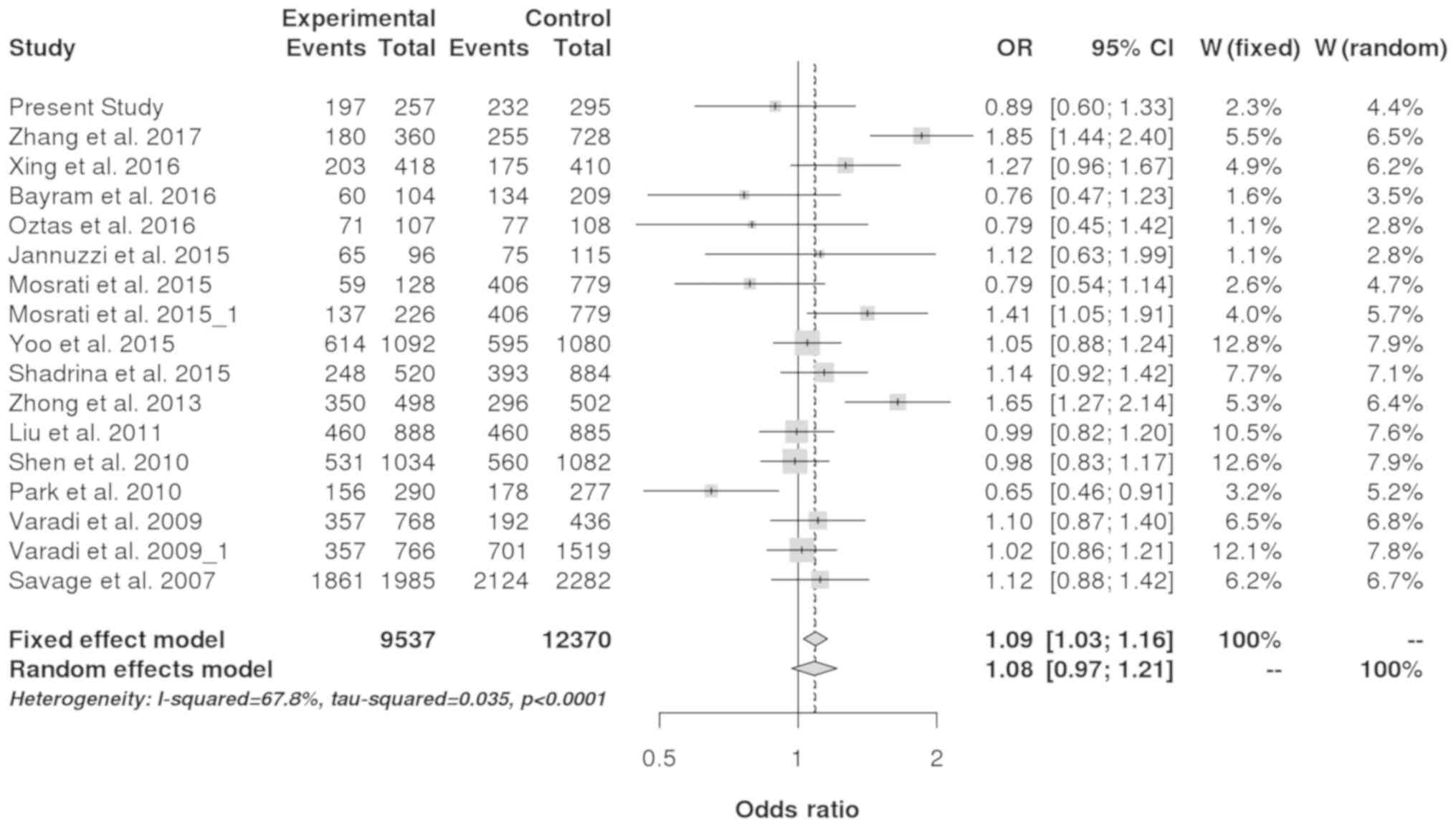
controls (Table V). By pooling all
the previous studies, a statistically significant association
between the TERT rs2583669 polymorphism and cancer risk was
identified (dominant model: Pooled OR=1.09; 95% CI, 1.03-1.16;
P=0.004; Fig. 2). Further,
stratification analyses were performed to assess the risk by type
of cancer and ethnicity (Table SI).
TERT rs2583669 polymorphism and cancer risk was not statistically
significant in different ethnicities. Stratification analyses by
type of cancer showed significant association of TERT rs2583669
polymorphism with acute myeloid leukemia, hepatocellular carcinoma
and lung cancer (Table SI).
Significant heterogeneity was observed in all genetic models tested
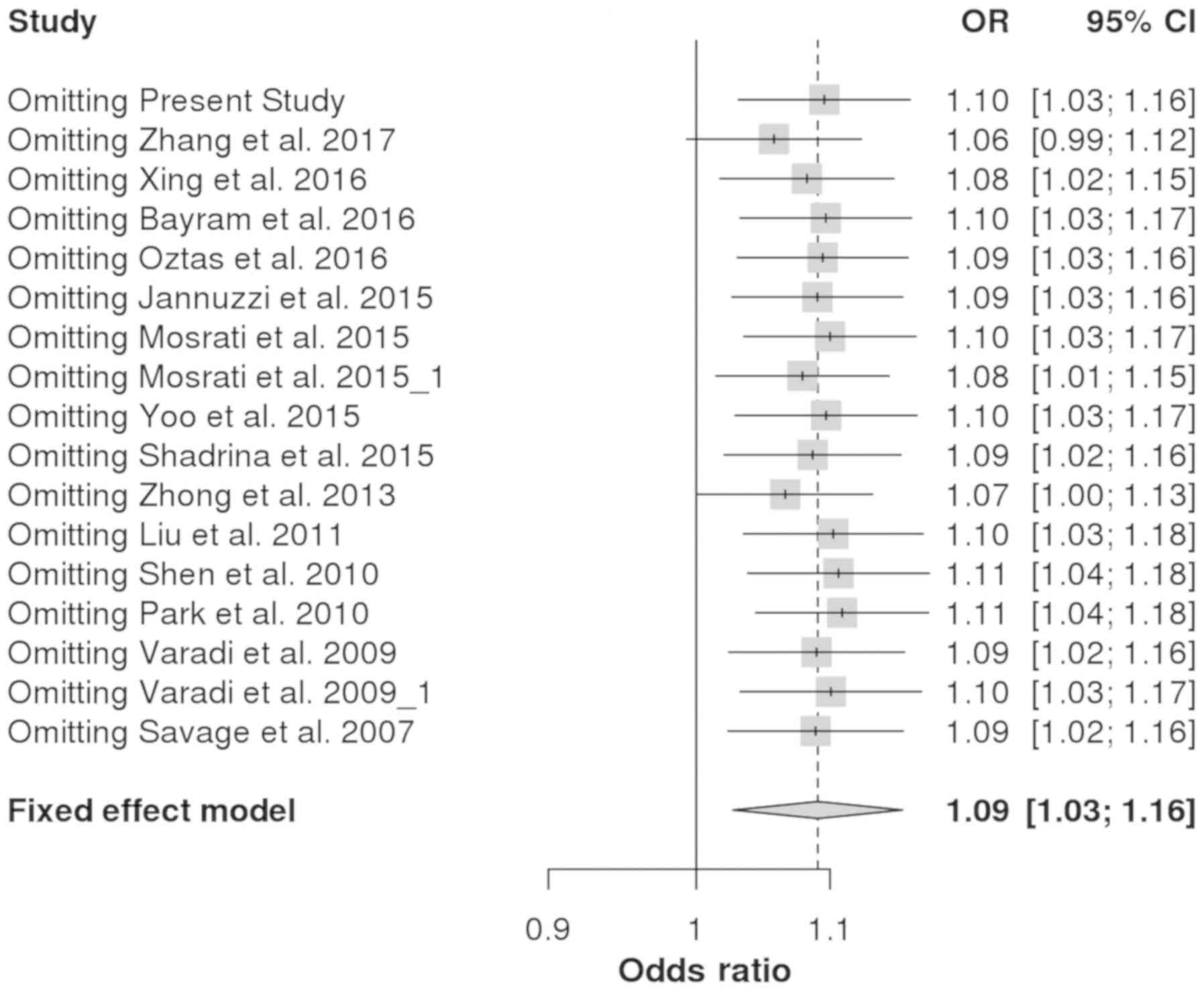
(Table SI). Sensitivity analysis
performed by omitting individual studies revealed that there was no
change in the pooled ORs (Fig. 3).
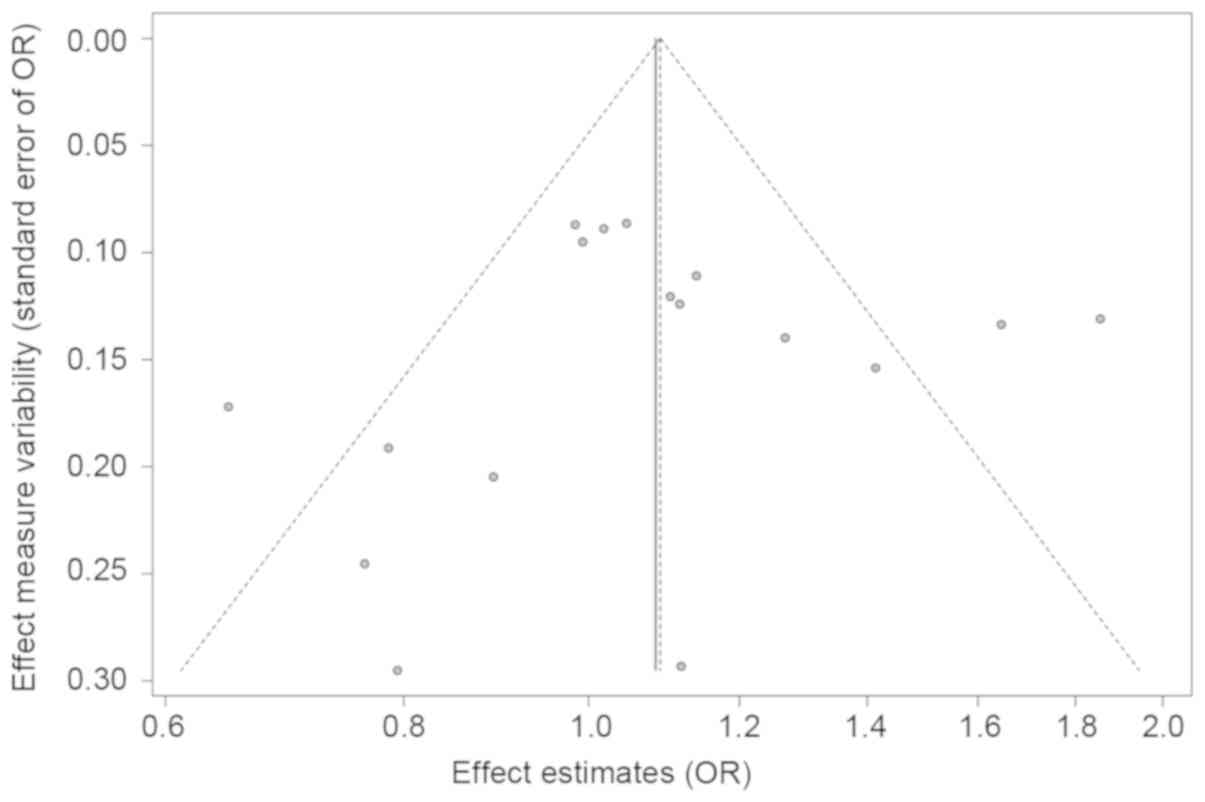
The shape of the funnel plot (Fig.
4) and Egger's tests did not reveal any evidence for asymmetry
in all three genetic models (Table
SI).
 | Table VTelomerase reverse transcriptase
rs2853669 and cancer association studies included in the
meta-analysis. |
Table V
Telomerase reverse transcriptase
rs2853669 and cancer association studies included in the
meta-analysis.
| | | | | | Cases | Control | HWE | |
|---|
| Author, year | Serial no. | Disease | Country | Ethnicity | CC | CT | TT | CC | CT | TT | P-value | (Refs.) |
|---|
| Present Study | 1 | Cervical
cancer | India | Asian | 106 | 91 | 60 | 86 | 146 | 63 | 0.944 | - |
| Zhang et al,
2017 | 2 | Gastric cancer | China | Asian | 44 | 136 | 180 | 36 | 219 | 473 | 0.109 | (23) |
| Xing et al,
2016 | 3 | Lung cancer | China | Asian | 41 | 162 | 215 | 30 | 145 | 235 | 0.249 | (24) |
| Bayram et
al, 2016 | 4 | Gastric cancer | Turkey | Caucasian | 13 | 47 | 44 | 35 | 99 | 75 | 0.810 | (25) |
| Oztas et al,
2016 | 5 | Breast cancer | Turkey | Caucasian | 24 | 47 | 36 | 25 | 52 | 31 | 0.723 | (26) |
| Jannuzzi et
al, 2015 | 6 | Colorectal
cancer | Turkey | Caucasian | 15 | 50 | 31 | 17 | 58 | 40 | 0.587 | (27) |
| Mosrati et
al, 2015 | 7 | Glioblastoma | Sweden | Caucasian | 11 | 48 | 69 | 65 | 341 | 373 | 0.293 | (15) |
| Mosrati et
al, 2015 | 8 | AML | Sweden | Caucasian | 38 | 99 | 89 | 65 | 341 | 373 | 0.293 | (15) |
| Yoo et al,
2015 | 9 | Lung cancer | Korea | Asian | 137 | 477 | 478 | 105 | 490 | 485 | 0.242 | (28) |
| Shadrina et
al, 2015 | 10 | NHL | Russia | Caucasian | 35 | 213 | 272 | 71 | 322 | 491 | 0.079 | (29) |
| Zhong et al,
2013 | 11 | Lung cancer | China | Asian | 108 | 242 | 148 | 72 | 224 | 206 | 0.381 | (30) |
| Liu et al,
2011 | 12 | SCCHN | USA | Caucasian | 79 | 381 | 428 | 85 | 375 | 425 | 0.863 | (31) |
| Shen et al,
2010 | 13 | Breast cancer | USA | Caucasian | 86 | 445 | 503 | 128 | 432 | 522 | 0.009 | (32) |
| Park et al,
2010 | 14 | HCC | Korea | Asian | 35 | 121 | 134 | 68 | 110 | 99 | 0.001 | (33) |
| Varadi et
al, 2009 | 15 | Breast cancer | Poland | Caucasian | 58 | 299 | 411 | 38 | 154 | 244 | 0.059 | (34) |
| Varadi et
al, 2009 | 16 | Breast cancer | Sweden | Caucasian | 47 | 310 | 409 | 143 | 558 | 818 | 0.001 | (34) |
| Savage et
al, 2007 | 17 | Breast cancer | Poland | Caucasian | 1,095 | 766 | 124 | 1,224 | 900 | 158 | 0.669 | (35) |
Discussion
TERT has been found to be overexpressed in 90% of
human cancer (36). Furthermore,
genetic alterations in the proximal promoter of TERT were shown to
be significantly associated with a range of clinical stages of
different cancer types (37).
Recently, one of the mechanisms of TERT regulation through the
non-coding driver mutations (C228T and C250T) in the TERT promoter
has been reported in several cancer types with different
frequencies (38-40).
The mutations created a new binding site for the ETS family of
transcription factors, resulting in the overexpression of TERT
(9,10). However, the mechanism by which it
overexpresses and the length of the telomere were reported to be
different in different types of tumor. TERT has been shown to
co-operate with activated oncogenes and inactivated tumor
suppressor genes in tumorigenesis (41). In addition, viral oncogene expression
and other genetic alterations, including mutations and SNPs in both
coding and non-coding regions, were shown to play a major role in
carcinogenesis (42). Advances in
next generation sequencing resulted in the identification of
several SNPs and mutations in non-coding regulatory regions,
highlighting the role of genetic variants in the regulatory region
of genes in tumorigenesis (8).
The TERT promoter SNP rs2853669 and its association
with cancer risk has been reported in various cancer types in
different populations (13-15).
However, the role of this particular SNP with the TERT promoter
driver mutations and its genetic association with HPV in patients
with cervical cancer had not yet been studied, to the best of our
knowledge. In the present study, to understand the genetic
association of the TERT SNP with cervical cancer risk, the SNP
rs2853669 was genotyped in 257 cervical cancer samples of South
Indian origin and 295 control samples, and a higher frequency of
the rs2853669 variant allele (G) was observed in the cervical
cancer samples compared with the control samples. The SNP rs2853669
in the TERT promoter has been shown to have a functional impact on
TERT regulation and telomere length in various cancer types
(34,35,43,44).
Furthermore, the genetic association of the TERT SNP rs2853669 has
been reported in many cancer types of different populations where
the disease association was different between the population and
the cancer types. The variant rs2853669 is located in the binding
site of ETS-2, another transcription factor of the ETS family,
which regulates various genes involved in cellular senescence and
tumorigenesis (45,46). The results of either the SNP of TERT
(rs2853669) alone or the modifying effect of rs2853669 on TERT
hotspot promoter mutations have been shown to be highly
controversial with varying results across different previous
studies (15,31,34,35,44,47,48).
The combination of the rs2853669 variant allele (G)
and TERT promoter hotspot mutations was also high compared with
hotspot mutations and the wild-type allele. Almost all the patients
with the rs2853669 and hotspot mutations were observed in poorly
differentiated squamous cell carcinoma tumors except one patient
where it was a moderately differentiated tumor (49). It has been reported that the TERT
hotspot mutation-positive cases showed poor survival in the absence
of the variant allele (47). On the
contrary, previous studies on glioblastoma showed a shortest mean
overall survival, which was mainly detected in patients harboring
both an activating TERT promoter mutation and the rs2853669 variant
homozygous allele (14,15). Moreover, the role of rs2853669 with
or without TERT hotspot mutations on TERT regulation and activation
has been demonstrated through functional experiments in
hepatocellular carcinoma (HCC) cell lines. E2F1 binding to the TERT
promoter can enhance interaction with other epigenetic modifiers
like DNMT1 and HDAC1 to enable promoter methylation-mediated
regulation of TERT transcription, which suggested that E2F1 could
potentially function as a transcriptional repressor of TERT
(16). The SNP (rs2853669) may
interfere with the binding of E2F1 and influence promoter
methylation associated with the TERT transcription as the SNP
resides at 2 bp downstream of E2F1 binding site on the TERT
promoter (16). Recently, the
combination of -124 C>T (C228T) mutation and rs2853669
(-245T>C) variant was reported to be correlated with increased
TERT transcription activity in HCC cases that frequently resulted
in higher risk of recurrence (16).
The increased mortality in HCC has been observed in cases with the
co-existence of the two hotspot TERT promoter mutations and SNPs
(16). The expression level of TERT
is increased by the inhibition of the transcriptional repressor
E2F1 in the presence of the variant allele -245T>C, together
with the activation of the ETS2 transcription factor due to the
-124C>T mutation. A recent study reported that the -245T region
located on an ETS2 binding site is not a native ETS2 binding site,
suggesting that the TERT promoter mutation (-124C>T and
-146C>T) cooperates with its native ETS binding sites to form
high-order structures such as G-quadruplexes, contributing to the
recruitment rs2853669 with the non-coding driver hotspot mutation
in the TERT promoter has been shown to be associated with poor
prognosis in HCC (16). Furthermore,
the present results suggested that the hotspot TERT promoter
mutations and the SNP combinations might play an important role in
TERT regulation. In addition, the expression of TERT was shown to
be regulated by HPV E6 oncoprotein (50,51). The
combination of the rs2853669 variant allele and TERT promoter
hotspot mutation (C228T and C250T) along with HPV could be an
additional risk to patients with cervical cancer. However, the
present study did not identify any significant association between
the TERT SNP (rs2853669) and the presence of HPV.
In addition, to understand and compare the genotype
frequency and the risk associated with the SNP rs2853669 among
various cancer types and different populations, a meta-analysis was
conducted. Due to the lack of previous studies on the association
of rs2853669 and cervical cancer risk in different populations, a
meta-analysis using different cancer types of the world population
was conducted. In the present study, the variant allele ‘G’ showed
a significantly increased association with cancer risk in the
dominant model. This result was consistent with previous studies
reported from other human cancer types and this result was also
obtained in the present meta-analysis (14,28,29,46).
Overall, the present meta-analysis showed a significant risk
association of the SNP rs2853669 with various cancer types of the
different ethnic population. In contrast, Shen et al
(52) reported that the SNP had no
association with cancer risk and prognosis. The number and ethnic
origin of the samples could influence the outcome of the results.
However, when the SNP was combined with TERT promoter mutations, a
modifying effect of rs2853669 among patients with cancer with TERT
promoter mutations was observed and only those patients carrying
the TT genotype had a poor survival (52). Although the effect of both the SNP
and the TERT promoter mutations were analyzed in the present study,
it was not possible to identify an association between the TERT SNP
with the clinicopathological features of the patients with cervical
cancer, as the majority of the patient history showed poor follow
up and/or drop out from the treatments. Moreover, all previous
genome-wide association studies (GWAS) showed an association of
this SNP with telomere length (53).
To the best of our knowledge, no previous GWAS study has reported
the association of SNP with cancer risk/clinical features, although
many individual previous studies have reported the association of
this SNP with cancer risk across human cancer but not in cervical
cancer (14-16,33).
In conclusion, the present study suggests that the
TERT rs2853669 variant ‘GG’ may play a role in the progression of
cervical cancer in South India as well in different cancer types of
world populations. The present study enrolled female patients from
a relatively homogenous population of South India; however, the
limitations of the present study should be considered when
interpreting the results. A major limitation of the present study
is the collection of tissue from cases and blood samples from
controls for DNA extraction and genotyping. Furthermore, a number
of demographic and clinicopathological characteristics limited the
present study. Owing to inadequate data, the impact of demographic
and clinicopathological characteristics was not considered to
perform multivariate analysis. Furthermore, non-availability of the
data on Helicobacter pylori infection, alcohol consumption
and smoking, limited the evaluation of the potential interactions
between these risk factors and TERT rs2853669 polymorphism. In
conclusion, within the limitations, the present study provides an
insight on the significance of genetic variants present in the
non-coding regions of genes and their association with the hotspot
mutations. However, functional studies are warranted to establish
the role of rs2853669 in cervical carcinogenesis.
Supplementary Material
Representative sequences for
telomerase reverse transcriptase rs2853669 genotypes. (A)
Homozygous variant, (B) homozygous wild-type and (C)
heterozygous.
Subgroup analysis by type of cancer
and ethnicity in the meta-analysis on the association between
telomerase reverse transcriptase rs2853669 polymorphism and cancer
risk.
Acknowledgements
Not applicable.
Funding
The present study was supported by The University
Grants Commission (UGC-SAP) and Department of Science and
Technology (DST-FIST) grants and the Department of Health Research,
DHR-MRU grant, Government of India for the research infrastructural
facilities. Research fellowships were awarded to VV and KA from
Council of Scientific and Industrial Research, and SR and GA from
University Grants Commission, New Delhi, Government of India. The
present study was supported by research grants from Department of
Atomic Energy, Board of Research in Nuclear Sciences, Mumbai, India
(grant no. 35/14/10/2014-BRNS/0210) and Department of
Biotechnology, New Delhi, India (grant no.
BT/PR10023/AGR/36/27/2007) to AKM.
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
AKMun conceived the study, designed the experiments
and supervised the study. VV and SR contributed to the study
design, performed the experiments, analyzed the data and drafted
the manuscript. KA and GA helped conduct the experiments and
drafted the manuscript. RR provided tumor samples and clinical
data. AKMur, LVKSB and AKMun analyzed the data and critically
reviewed the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was conducted after obtaining
approval from The Institutional Ethics Committee (approval no.
04092010) of Madras Medical College and Hospital (Chennai, China).
The tissue/blood samples were collected from The Institute of
Social Obstetrics and Government Kasturba Gandhi Hospital for Women
and Children, Triplicane, Chennai and Government Royapettah
Hospital after obtaining written informed consent from each
patient.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Low KC and Tergaonkar V: Telomerase:
Central regulator of all of the hallmarks of cancer. Trends Biochem
Sci. 38:426–434. 2013.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Maciejowski J and de Lange T: Telomeres in
cancer: Tumour suppression and genome instability. Nat Rev Mol Cell
Biol. 18:175–186. 2017.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Pestana A, Vinagre J, Sobrinho-Simões M
and Soares P: TERT biology and function in cancer: Besyond
immortalisation. J Mol Endocrinol. 58:R129–R146. 2017.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Barthel FP, Wei W, Tang M,
Martinez-Ledesma E, Hu X, Amin SB, Akdemir KC, Seth S, Song X, Wang
Q, et al: Systematic analysis of telomere length and somatic
alterations in 31 cancer types. Nat Genet. 49:349–357.
2017.PubMed/NCBI View
Article : Google Scholar
|
|
5
|
Sreedevi A, Javed R and Dinesh A:
Epidemiology of cervical cancer with special focus on India. Int J
Womens Health. 7:405–414. 2015.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Schiffman M, Wentzensen N, Wacholder S,
Kinney W, Gage JC and Castle PE: Human papillomavirus testing in
the prevention of cervical cancer. J Natl Cancer Inst. 103:368–383.
2011.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Cancer Genome Atlas Research Network;
Albert Einstein College of Medicine; Analytical Biological
Services; Barretos Cancer Hospital; Baylor College of Medicine;
Beckman Research Institute of City of Hope; Buck Institute for
Research on Aging; Canada's Michael Smith Genome Sciences Centre;
Harvard Medical School; Helen F. Graham Cancer Center &
Research Institute at Christiana Care Health Services, et al:
Integrated genomic and molecular characterization of cervical
cancer. Nature 543: 378.384, 2017.
|
|
8
|
Khurana E, Fu Y, Chakravarty D, Demichelis
F, Rubin MA and Gerstein M: Role of non-coding sequence variants in
cancer. Nat Rev Genet. 17:93–108. 2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Horn S, Figl A, Rachakonda PS, Fischer C,
Sucker A, Gast A, Kadel S, Moll I, Nagore E, Hemminki K, et al:
TERT promoter mutations in familial and sporadic melanoma. Science.
339:959–961. 2013.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Liu X, Bishop J, Shan Y, Pai S, Liu D,
Murugan AK, Sun H, El-Naggar AK and Xing M: Highly prevalent TERT
promoter mutations in aggressive thyroid cancers. Endocr Relat
Cancer. 20:603–610. 2013.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Killela PJ, Reitman ZJ, Jiao Y, Bettegowda
C, Agrawal N, Diaz LA Jr, Friedman AH, Friedman H, Gallia GL,
Giovanella BC, et al: TERT promoter mutations occur frequently in
gliomas and a subset of tumors derived from cells with low rates of
self-renewal. Proc Natl Acad Sci USA. 110:6021–6026.
2013.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Vinothkumar V, Arunkumar G, Revathidevi S,
Arun K, Manikandan M, Rao AK, Rajkumar KS, Ajay C, Rajaraman R,
Ramani R, et al: TERT promoter hot spot mutations are frequent in
Indian cervical and oral squamous cell carcinomas. Tumour Biol.
37:7907–7913. 2016.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Park CK, Lee SH, Kim JY, Kim JE, Kim TM,
Lee ST, Choi SH, Park SH and Kim IH: Expression level of hTERT is
regulated by somatic mutation and common single nucleotide
polymorphism at promoter region in glioblastoma. Oncotarget.
5:3399–3407. 2014.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Spiegl-Kreinecker S, Lotsch D, Ghanim B,
Pirker C, Mohr T, Laaber M, Weis S, Olschowski A, Webersinke G,
Pichler J and Berger W: Prognostic quality of activating TERT
promoter mutations in glioblastoma: Interaction with the rs2853669
polymorphism and patient age at diagnosis. Neuro Oncol.
17:1231–1240. 2015.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Mosrati MA, Malmström A, Lysiak M,
Krysztofiak A, Hallbeck M, Milos P, Hallbeck AL, Bratthäll C,
Strandéus M, Stenmark-Askmalm M and Söderkvist P: TERT promoter
mutations and polymorphisms as prognostic factors in primary
glioblastoma. Oncotarget. 6:16663–16673. 2015.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Ko E, Seo HW, Jung ES, Kim BH and Jung G:
The TERT promoter SNP rs2853669 decreases E2F1 transcription factor
binding and increases mortality and recurrence risks in liver
cancer. Oncotarget. 7:684–699. 2016.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Ostör AG: Natural history of cervical
intraepithelial neoplasia: A critical review. Int J Gynecol Pathol.
12:186–192. 1993.PubMed/NCBI
|
|
18
|
Green MR: Molecular Cloning: A Laboratory
Manual. Green MR and Sambrook J (eds). Cold Spring Harbor
Laboratory Press, New York, NY, 2012.
|
|
19
|
Chen B, Cole JW and Grond-Ginsbach C:
Departure from Hardy Weinberg equilibrium and genotyping error.
Front Genet. 8(167)2017.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Moher D, Liberati A, Tetzlaff J and Altman
DG: Preferred reporting items for systematic reviews and
meta-analyses: The PRISMA statement. PLoS Med.
6(e1000097)2009.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Wallace BC, Dahabreh IJ, Trikalinos TA,
Lau J, Trow P and Schmid CH: Closing the gap between methodologists
and end-users: R as a computational back-end. J Stat Software.
49(15)2012.
|
|
22
|
Egger M, Davey Smith G, Schneider M and
Minder C: Bias in meta-analysis detected by a simple, graphical
test. BMJ. 315:629–634. 1997.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Zhang J, Ju H, Gao JR, Jiao XL and Lu Y:
Polymorphisms in human telomerase reverse transcriptase (hTERT)
gene, gene-gene and gene-smoking interaction with susceptibility to
gastric cancer in Chinese Han population. Oncotarget.
8:20235–20243. 2017.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Xing YL, Liu F, Li JF, Lin JC, Zhu GD, Li
M, Zhang CR and Niu YY: Case-control study on impact of the
telomerase reverse transcriptase gene polymorphism and additional
single nucleotide polymorphism (SNP)-SNP interaction on non-small
cell lung cancers risk in Chinese Han population. J Clin Lab Anal.
30:1071–1077. 2016.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Bayram S, Ülger Y, Sümbül AT, Kaya BY,
Genç A, Rencüzoğullari E and Dadaş E: Polymorphisms in human
telomerase reverse transcriptase (hTERT) gene and susceptibility to
gastric cancer in a Turkish population: Hospital-based case-control
study. Gene. 585:84–92. 2016.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Oztas E, Kara H, Kara ZP, Aydogan MU, Uras
C and Ozhan G: Association between human telomerase reverse
transcriptase gene variations and risk of developing breast cancer.
Genet Test Mol Biomarkers. 20:459–464. 2016.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Jannuzzi AT, Karaman E, Oztas E, Yanar HT
and Özhan G: Telomerase reverse transcriptase (TERT) gene
variations and susceptibility of colorectal cancer. Genet Test Mol
Biomarkers. 19:692–697. 2015.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Yoo SS, Do SK, Choi JE, Lee SY, Lee J, Cha
SI, Kim CH and Park JY: TERT polymorphism rs2853669 influences on
lung cancer risk in the Korean population. J Korean Med Sci.
30:1423–1428. 2015.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Shadrina AS, Boyarskikh UA, Oskina NA,
Sinkina TV, Lazarev AF, Petrova VD and Filipenko ML: TERT
polymorphisms rs2853669 and rs7726159 influence on prostate cancer
risk in Russian population. Tumour Biol. 36:841–847.
2015.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Zhong R, Liu L, Zou L, Zhu Y, Chen W, Zhu
B, Shen N, Rui R, Long L, Ke J, et al: Genetic variations in
TERT-CLPTM1L locus are associated with risk of lung cancer in
Chinese population. Mol Carcinog. 52((Suppl 1)): E118–E126.
2013.PubMed/NCBI View
Article : Google Scholar
|
|
31
|
Liu Z, Ma H, Wei S, Li G, Sturgis EM and
Wei Q: Telomere length and TERT functional polymorphisms are not
associated with risk of squamous cell carcinoma of the head and
neck. Cancer Epidemiol Biomarkers Prev. 20:2642–2645.
2011.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Shen J, Gammon MD, Wu HC, Terry MB, Wang
Q, Bradshaw PT, Teitelbaum SL, Neugut AI and Santella RM: Multiple
genetic variants in telomere pathway genes and breast cancer risk.
Cancer Epidemiol Biomarkers Prev. 19:219–228. 2010.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Park J, Chang H, Ahn S, Lee J, Kim D, Lee
K, Chon C, Moon Y and Han K: hTERT promoter gene polymorphism and
the risk of hepatocellular carcinoma (HCC) in patients with chronic
hepatitis. B. J Hepatol. 52:(Suppl 1): S345,. 2010.
|
|
34
|
Varadi V, Brendle A, Grzybowska E,
Johansson R, Enquist K, Butkiewicz D, Pamula-Pilat J, Pekala W,
Hemminki K, Lenner P and Försti A: A functional promoter
polymorphism in the TERT gene does not affect inherited
susceptibility to breast cancer. Cancer Genet Cytogenet. 190:71–74.
2009.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Savage SA, Chanock SJ, Lissowska J,
Brinton LA, Richesson D, Peplonska B, Bardin-Mikolajczak A,
Zatonski W, Szeszenia-Dabrowska N and Garcia-Closas M: Genetic
variation in five genes important in telomere biology and risk for
breast cancer. Br J Cancer. 97:832–836. 2007.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Jiang Y, Chen C, Chen SM, Wang YQ, Xu Y,
Wang Y, Chen Z, Xiao BK and Tao ZZ: Telomerase reverse
transcriptase promotes the proliferation of human laryngeal
carcinoma cells through activation of the activator protein 1.
Oncol Lett. 6:75–80. 2013.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Shay JW: Role of telomeres and telomerase
in aging and cancer. Cancer Discov. 6:584–593. 2016.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Liu R and Xing M: TERT promoter mutations
in thyroid cancer. Endocr Relat Cancer. 23:R143–R155.
2016.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Shimoi T, Yoshida M, Kitamura Y, Yoshino
T, Kawachi A, Shimomura A, Noguchi E, Yunokawa M, Yonemori K,
Shimizu C, et al: TERT promoter hotspot mutations in breast cancer.
Breast Cancer. 25:292–296. 2018.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Thiem S, Herold T, Krafft U, Bremmer F,
Tolkach Y, Szász AM, Kriegsmann J, Gaisa NT, Niedworok C, Szarvas T
and Reis H: Telomerase reverse transcriptase (TERT) promoter
mutations are rare in urachal cancer. Pathol Int. 67:597–601.
2017.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Liu X, Dakic A, Chen R, Disbrow GL, Zhang
Y, Dai Y and Schlegel R: Cell-restricted immortalization by human
papillomavirus correlates with telomerase activation and engagement
of the hTERT promoter by Myc. J Virol. 82:11568–11576.
2008.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Chen Y, Williams V, Filippova M, Filippov
V and Duerksen-Hughes P: Viral carcinogenesis: Factors inducing DNA
damage and virus integration. Cancers (Basel). 6:2155–2186.
2014.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Hsu CP, Hsu NY, Lee LW and Ko JL: Ets2
binding site single nucleotide polymorphism at the hTERT gene
promoter-effect on telomerase expression and telomere length
maintenance in non-small cell lung cancer. Eur J Cancer.
42:1466–1474. 2006.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Bojesen SE, Pooley KA, Johnatty SE,
Beesley J, Michailidou K, Tyrer JP, Edwards SL, Pickett HA, Shen
HC, Smart CE, et al: Multiple independent variants at the TERT
locus are associated with telomere length and risks of breast and
ovarian cancer. Nat Genet. 45:371–384, 384e1-2,. 2013.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Horikawa I, Cable PL, Afshari C and
Barrett JC: Cloning and characterization of the promoter region of
human telomerase reverse transcriptase gene. Cancer Res.
59:826–830. 1999.PubMed/NCBI
|
|
46
|
Helbig S, Wockner L, Bouendeu A,
Hille-Betz U, McCue K, French JD, Edwards SL, Pickett HA, Reddel
RR, Chenevix-Trench G, et al: Functional dissection of breast
cancer risk-associated TERT promoter variants. Oncotarget.
8:67203–67217. 2017.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Rachakonda PS, Hosen I, de Verdier PJ,
Fallah M, Heidenreich B, Ryk C, Wiklund NP, Steineck G, Schadendorf
D, Hemminki K and Kumar R: TERT promoter mutations in bladder
cancer affect patient survival and disease recurrence through
modification by a common polymorphism. Proc Natl Acad Sci USA.
110:17426–17431. 2013.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Nagore E, Heidenreich B, Requena C,
García-Casado Z, Martorell-Calatayud A, Pont-Sanjuan V,
Jimenez-Sanchez AI and Kumar R: TERT promoter mutations associate
with fast-growing melanoma. Pigment Cell Melanoma Res. 29:236–238.
2016.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Bell RJA, Rube HT, Xavier-Magalhães A,
Costa BM, Mancini A, Song JS and Costello JF: Understanding TERT
promoter mutations: A common path to immortality. Mol Cancer Res.
14:315–323. 2016.PubMed/NCBI View Article : Google Scholar
|
|
50
|
McMurray HR and McCance DJ: Human
papillomavirus type 16 E6 activates TERT gene transcription through
induction of c-Myc and release of USF-mediated repression. J Virol.
77:9852–9861. 2003.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Liu X, Dakic A, Zhang Y, Dai Y, Chen R and
Schlegel R: HPV E6 protein interacts physically and functionally
with the cellular telomerase complex. Proc Natl Acad Sci USA.
106:18780–18785. 2009.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Shen N, Lu Y, Wang X, Peng J, Zhu Y and
Cheng L: Association between rs2853669 in TERT gene and the risk
and prognosis of human cancer: A systematic review and
meta-analysis. Oncotarget. 8:50864–50872. 2017.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Soerensen M, Thinggaard M, Nygaard M, Dato
S, Tan Q, Hjelmborg J, Andersen-Ranberg K, Stevnsner T, Bohr VA,
Kimura M, et al: Genetic variation in TERT and TERC and human
leukocyte telomere length and longevity: A cross-sectional and
longitudinal analysis. Aging Cell. 11:223–227. 2012.PubMed/NCBI View Article : Google Scholar
|