Introduction
Hepatocellular carcinoma (HCC) is the predominant
histological subtype of hepatic cancer, comprising 90% of all
primary hepatic malignancies worldwide (1). Its high mortality rate and poor
prognosis render it a significant global public health threat. In
particular, the survival rate of patients with HCC remains among
the lowest across all cancer types, with studies indicating a
five-year survival rate of only 21%. The prevalence of HCC exhibits
notable geographic variation, with China bearing a substantial
proportion of the global burden, accounting for more than 50% of
all cases globally (2).
A major challenge in managing HCC is that the
disease often remains asymptomatic during its early phases, leading
to delayed diagnosis (3). As a
result, most patients are diagnosed at intermediate or advanced
stages, missing the optimal window for curative interventions such
as surgical resection, liver transplantation, or percutaneous
ablation for early-stage liver cancer (4). Although liver transplantation is
considered the most effective treatment for eligible patients, its
utility is often limited by high rates of recurrence and metastasis
(5-7).
Therefore, developing strategies for early precision diagnosis and
targeted therapy is critically important. In this end, there is an
urgent need to identify effective biomarkers to aid in early
screening, accurate diagnosis, treatment evaluation and prognosis
prediction for patients with HCC (8).
The pathogenesis of HCC involves dysregulation of
key cellular processes such as cell cycle control, apoptosis and
DNA repair (9). Among the key
molecular drivers is the ubiquitin-proteasome system (UPS), the
primary pathway for intracellular protein degradation. The UPS also
plays a critical role in regulating antigen processing, signal
transduction, and cell cycle progression (10). In HCC, aberrant UPS function, often
marked by the overexpression of specific proteasome subunits, has
been recognized as a hallmark of HCC, supported by the clinical
efficacy of proteasome inhibitors in certain patient subgroups
(11).
As cancer cells exhibit rapid proliferation, they
generate an increased load of misfolded and damaged proteins,
creating a highly promiscuous cellular environment (12), where proteasomes can combine with
highly promiscuous substrates selectively and induce the
degradation of proteins modified with ubiquitin chains (13). The 26S proteasome selectively
recognizes and degrades ubiquitin-tagged proteins, thereby
maintaining protein homeostasis. This complex includes PSMD
(proteasome 26S subunit, non-ATPase) proteins, which comprise 14
distinct subunits responsible for the recognition and degradation
of damaged, misfolded, or foreign proteins (14).
Increasing evidence suggests that several
PSMD components are overexpressed in various malignancies
and may serve as potential biomarkers and therapeutic targets in
HCC due to their distinct molecular functions (15,16).
For example, PSMD11 has been reported to promote HCC
progression through interactions with ATP7A, DLAT and
PDHA1 (17). Similarly, high expression of PSMD13 is
associated with sustained tumor cell activity,
epithelial-mesenchymal transition (EMT) and genomic instability
(18). However, the roles of
numerous other PSMD family members in hepatocarcinogenesis
remain poorly understood. Notably, 26S proteasome non-ATPase
regulatory subunit 6 (PSMD6, also known as Rpn7) has
been implicated in maintaining genomic stability; its knockdown has
been shown to induce DNA damage and apoptosis in experimental
models (19). Nevertheless, the
relationship between PSMD6 and HCC pathogenesis has not been
systematically investigated.
The present study aims to comprehensively analyze
the expression patterns of PSMD6 in HCC at both mRNA and
protein levels and to evaluate its potential as a biomarker for
early diagnosis and prognosis assessment. Through a combination of
bioinformatics analysis and experimental validation, it was aimed
to clarify the clinical significance of PSMD6 and its
underlying mechanisms in HCC progression.
Materials and methods
Data acquisition and processing
Transcriptome data (RNA-seq) and corresponding
clinical information of the Liver Hepatocellular Carcinoma (LIHC)
project were downloaded from The Cancer Genome Atlas (TCGA) portal
(https://portal.gdc.cancer.gov).
The initial dataset comprised 424 clinical samples,
including 371 tumor tissues and 53 paired adjacent normal tissues.
Samples with incomplete clinical information were excluded from
subsequent survival and clinical correlation analyses. To augment
the normal liver tissue sample size, transcriptome data from 110
normal liver samples were obtained from the Genotype-Tissue
Expression (GTEx) project database (https://commonfund.nih.gov/GTEx). All gene expression
data were normalized to Transcripts Per Million (TPM) format. For
downstream analyses, the TPM values were transformed using log2(TPM
+ 1) to approximate a normal distribution. All data processing was
performed using R software (version 4.2.1; https://www.R-project.org).
Open databases and bioinformatics
analysis. Xiantao Toolbox
The Xiantao Toolbox (https://xiantaozi.com/) platform provides a non-coding
suite of tools for multi-dimensional bioinformatics analysis
without requiring programming expertise. It was utilized in the
present study for initial data exploration and specific analytical
modules.
TISCH database. Tumor Immune Single-cell Hub
(TISCH, http://tisch.comp-genomics.org/) is a curated database
dedicated to the tumor microenvironment (TME), providing
single-cell RNA sequencing (scRNA-seq) data across various cancer
types. It was consulted to explore the cellular composition of the
TME in HCC at a single-cell resolution.
BioGRID database. Biological General
Repository for Interaction Datasets (BioGRID, https://thebiogrid.org) is a publicly accessible
repository of protein, genetic and chemical interactions. This
database provides over 2.8 million protein types and genetic
interactions, and more than 30.000 chemical interactions, and this
database was queried to identify known and predicted
protein-protein interactions (PPIs) involving PSMD6.
UALCAN database. UALCAN is a comprehensive
web-based bioinformatics toolbox platform (https://ualcan.path.uab.edu/) for analyzing cancer
OMICS data, particularly from TCGA. It enables in-depth gene
expression analysis, patient survival analysis, and promoter
methylation profiling across different tumor subgroups. This
database was used to validate PSMD6 expression patterns and
assess its correlation with key clinical parameters (for example,
cancer stage and tumor grade).
SANGERBOX database. SANGERBOX is an
integrated bioinformatics analysis platform (http://vip.sangerbox.com) offering a user-friendly
interface for various analyses, including differential expression,
pathway enrichment and Weighted Gene Co-expression Network Analysis
(WGCNA). It was employed for specific functional enrichment
analyses.
Kaplan-Meier Plotter. Kaplan-Meier Plotter
data repository (https://kmplot.com/analysis/) is designed to assess
the effect of gene expression on survival outcomes across multiple
cancer types using data from sources such as GEO and TCGA. It was
used specifically to evaluate the prognostic significance of PSMD6
expression in terms of overall survival (OS) and disease-specific
survival (DSS) in the TCGA-LIHC cohort. The platform employs a Cox
proportional hazards model, and hazard ratios with 95% confidence
intervals and log-rank P-values are reported.
STRING database. STRING is a database of
known and predicted PPIs (https://string-db.org). It was used to construct a PPI
network centered on PSMD6, with a confidence score threshold set to
ensure high-quality interactions.
TIMER database. TIMER (https://cistrome.shinyapps.io/timer/) is a web server
for comprehensive analysis of immune cell infiltrates across
diverse cancer types using TCGA data. It was utilized to
investigate the correlations between PSMD6 expression levels and
the abundance of six immune cell populations (B cells,
CD4+ T cells, CD8+ T cells, neutrophils,
macrophages and dendritic cells) within the HCC TME, with
adjustment for tumor purity.
Cell culture
The human immortalized hepatic epithelial cell line
(THLE-2) and three human liver cancer cell lines were procured from
Zhejiang Meisen Cell Technology Co., Ltd. (THLE-2, cat. no.
CTCC-004-0030; HepG2, cat. no. CTCC-DZ-0057; MHCC97-L, cat. no.
CTCC-400-0194; MHCC97-H, cat. no. CTCC-0395-Luc1). The 4 cell lines
was authenticated using short tandem repeat profiling. The obtained
STR profile was compared with reference databases (DSMZ) to confirm
identity. Cells were routinely tested for mycoplasma contamination
and used within 20 passages after thawing. All cell lines were
cultured in RPMI-1640 medium (Gibco; Thermo Fisher Scientific,
Inc.), supplemented with 10% heat-inactivated fetal bovine serum
(FBS; Gibco; Thermo Fisher Scientific, Inc.) and 1%
penicillin-streptomycin (Gibco; Thermo Fisher Scientific, Inc.).
Cells were maintained in a humidified incubator at 37˚C with 5%
CO2. Regular mycoplasma testing was performed to ensure
cell line authenticity and absence of contamination.
Western blot analysis
Total protein was extracted from cultured cells
using RIPA lysis buffer (Guangzhou Biolight Biotechnology Co.,
Ltd.) containing protease and phosphatase inhibitors (Tiandz,
Inc.). Protein concentration was determined using a bicinchoninic
acid (BCA) assay kit. Equal amounts of protein (typically 20-30 µg
per lane) were separated by 10% SDS-polyacrylamide gel
electrophoresis and subsequently transferred onto polyvinylidene
difluoride membranes (Seebio; https://www.seebio.cn/). After blocking with 5% (w/v)
bovine serum albumin (cat. no. BS114; Biosharp Life Sciences) in
Tris-buffered saline with 0.1% Tween-20 (TBST) for 1 h at room
temperature, the membranes were incubated overnight at 4˚C with the
primary antibody against PSMD6 (1:1,000; cat. no. Ag3251;
Proteintech Group, Inc.) and GAPDH as a loading control (1:50,000;
cat. no. 60004-1-Ig; Proteintech Group, Inc.). Following three
washes with TBST, the membranes were incubated with an appropriate
horseradish peroxidase (HRP)-conjugated secondary antibody
(1:50,000; cat. no. BL003A; Biosharp Life Sciences) for 1 h at room
temperature. Protein bands were visualized using an enhanced
chemiluminescence (ECL) detection kit (MilliporeSigma) and imaged
with a chemiluminescence imaging system. Densitometric analysis of
the bands was performed using ImageJ software (version 1.8.0.;
National Institutes of Health). All experiments were independently
repeated at least three times.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from cultured cells using
TriQuick Reagent (Beijing Solarbio Science & Technology Co.,
Ltd.) according to the manufacturer's instructions. RNA
concentration and purity (A260/A280 ≥1.8) were measured using a
NanoDrop spectrophotometer (Thermo Fisher Scientific, Inc.).
Complementary DNA (cDNA) was synthesized from 1 µg of total RNA
using a 5X RT SuperMix for qPCR kit (Vazyme Biotech Co., Ltd.).
qPCR was performed using a 2X SYBR Green qPCR Master Mix (Vazyme
Biotech Co., Ltd.) on an iQ5 Multicolor Real-Time PCR Detection
System (Bio-Rad Laboratories, Inc.). The PCR cycling conditions
were as follows: Initial denaturation at 95˚C for 30 sec, followed
by 40 cycles of 95˚C for 5 sec and 60˚C for 30 sec. The relative
mRNA expression level of PSMD6 was calculated using the
2-ΔΔCq method (20), with GAPDH as the internal reference
gene. The specific primer sequences used are listed in Table I. Each experiment was performed in
triplicate wells and repeated independently three times.
 | Table IPrimer sequences. |
Table I
Primer sequences.
| Gene name | Primer sequence
(5'-3') |
|---|
| GAPDH | F:
CATCATCCCTGCCTCTACTGG |
| | R:
GTGGGTGTCGCTGTTGAAGTC |
| Proteasome 26s
subunit, non-ATPase 6 | F:
AAGAACCCCGACTTGCGTATC |
| | R:
GGCTTCATAGTAAGGAGCCATGT |
Statistical analysis
Bioinformatics data analysis was primarily conducted
using R software (version 4.2.1 or 3.6.4 as specified for specific
packages). Differential expression of PSMD6 between HCC tumors and
normal tissues was assessed using the Wilcoxon rank-sum test.
Correlations between PSMD6 expression and immune cell infiltration
levels were evaluated using Pearson's or Spearman's correlation
coefficients via the corr.test function in the Psych R package
(v2.1.6; CRAN.R-project.org/package=psych). Survival analysis
was performed using the Kaplan-Meier method, and differences in
survival curves were compared using the log-rank test via the
Survival R package. For in vitro experiments, data are
presented as the mean ± standard deviation (x̄±s) from at least
three independent experiments. Statistical comparisons between two
groups were analyzed using independent two-tailed unpaired
Student's t-test. Comparisons among multiple groups were performed
using one-way analysis of variance (ANOVA) followed by Dunnett's
multiple comparison test. P<0.05 was considered to indicate a
statistically significant difference.
Results
PSMD6 is significantly overexpressed
in HCC and correlates with adverse clinicopathological
features
Investigation was initiated by assessing the
pan-cancer expression profile of PSMD6 to determine its potential
relevance in oncogenesis. To tackle this, the pan-cancer expression
of PSMD6 was assessed utilizing SANGERBOX database, and it
was revealed that PSMD6 was significantly dysregulated
across multiple cancer types, being upregulated in 15 and
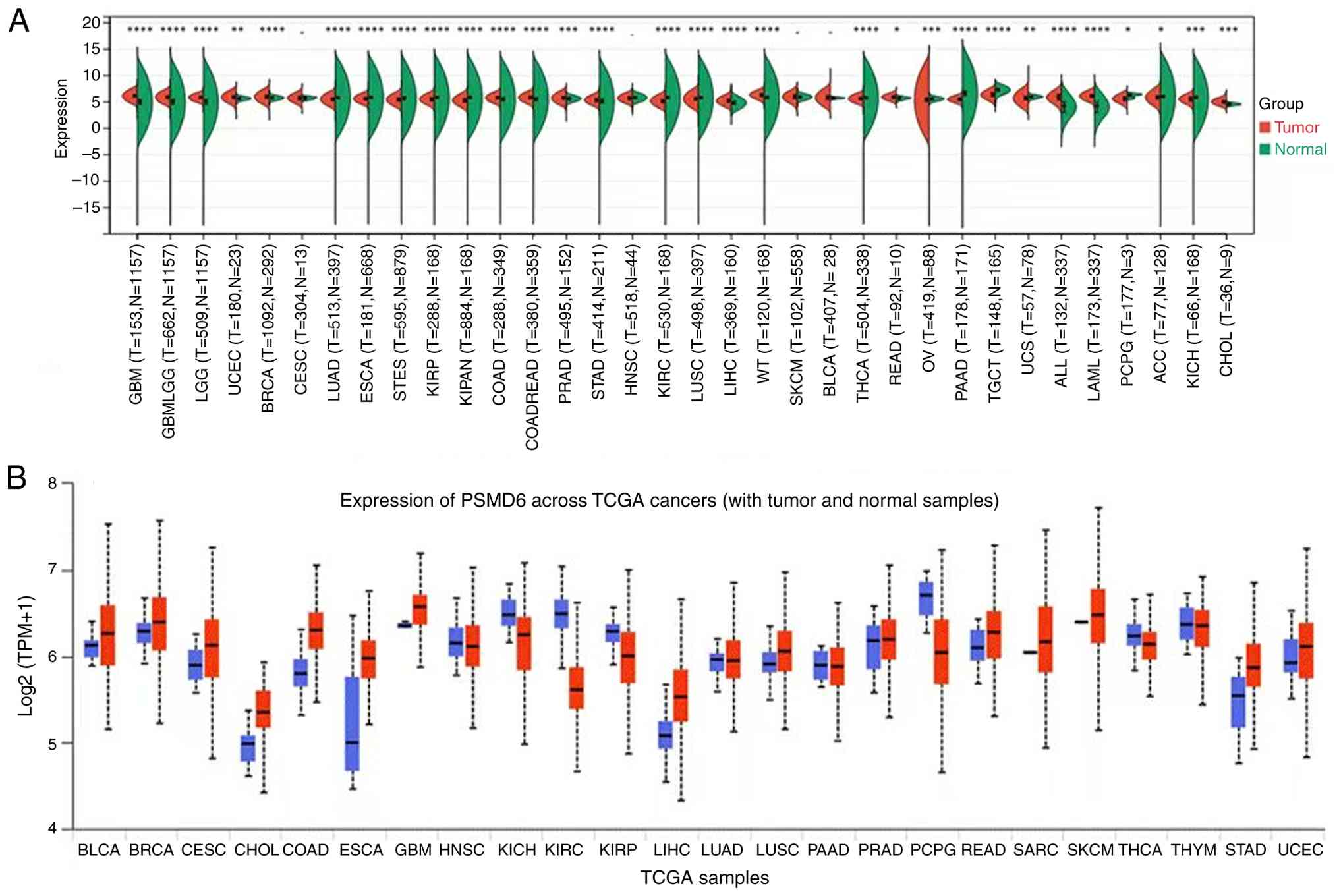
downregulated in 15 other malignancies (Table II, Fig. 1A). Consistent findings were gained
through analyzing with UALCAN data repository (Fig. 1B). This pan-cancer aberration
highlighted PSMD6 as a gene of general interest in cancer
research.
 | Table IIComparison of proteasome 26s subunit,
non-ATPase 6 expression in multiple tumor and non-tumor
tissues. |
Table II
Comparison of proteasome 26s subunit,
non-ATPase 6 expression in multiple tumor and non-tumor
tissues.
| Cancer | Tumor | Normal | P-value | Result |
|---|
| GBM | 6.11±0.42 | 4.86±1.30 |
4.7x10-72 | Increase |
| GBMLGG | 5.89±0.35 | 4.86±1.30 |
1.1x10-187 | Increase |
| LGG | 5.83±0.30 | 4.86±1.30 |
4.5x10-150 | Increase |
| UCEC | 5.85±0.56 | 5.64±0.27 |
2.1x10-3 | Increase |
| BRCA | 5.96±0.47 | 5.77±0.23 |
1.2x10-16 | Increase |
| COAD | 5.77±0.42 | 5.36±1.70 |
4.4x10-13 | Increase |
| COADREAD | 5.80±0.42 | 5.37±1.68 |
2.2x10-18 | Increase |
| PRAD | 5.73±0.42 | 5.60±0.35 |
1.5x10-4 | Increase |
| STAD | 5.37±0.46 | 4.99±1.56 |
6.0x10-8 | Increase |
| LIHC | 5.21±0.46 | 4.79±0.46 |
3.6x10-20 | Increase |
| WT | 6.32±0.46 | 5.65±1.35 |
3.2x10-21 | Increase |
| READ | 5.92±0.39 | 5.68±0.26 | 0.04 | Increase |
| ALL | 5.83±0.76 | 4.00±1.34 |
1.8x10-39 | Increase |
| LAML | 6.08±0.42 | 4.00±1.34 |
5.6x10-64 | Increase |
| CHOL | 5.06±0.44 | 4.56±0.21 |
2.8x10-4 | Increase |
| LUAD | 5.53±0.41 | 5.76±0.87 |
1.7x10-30 | Decrease |
| ESCA | 5.62±0.40 | 5.71±1.27 |
3.0x10-13 | Decrease |
| STES | 5.45±0.46 | 5.53±1.38 |
1.1x10-22 | Decrease |
| KIRP | 5.50±0.52 | 5.65±1.35 |
6.7x10-10 | Decrease |
| KIPAN | 5.31±0.51 | 5.65±1.35 |
1.8x10-28 | Decrease |
| KIRC | 5.18±0.46 | 5.65±1.35 |
2.5x10-39 | Decrease |
| LUSC | 5.59±0.40 | 5.76±0.87 |
1.2x10-20 | Decrease |
| THCA | 5.63±0.32 | 5.71±0.90 |
2.4x10-8 | Decrease |
| OV | 5.36±0.91 | 5.58±0.29 |
3.2x10-4 | Decrease |
| PAAD | 5.50±0.36 | 6.48±1.90 |
4.6x10-44 | Decrease |
| TGCT | 6.34±0.52 | 7.22±0.48 |
6.1x10-34 | Decrease |
| UCS | 5.76±0.69 | 5.95±0.24 |
2.7x10-3 | Decrease |
| PCPG | 5.59±0.53 | 6.20±0.43 | 0.05 | Decrease |
| ACC | 5.80±0.69 | 5.93±1.47 | 0.01 | Decrease |
| KICH | 5.49±0.57 | 5.65±1.35 |
2.8x10-4 | Decrease |
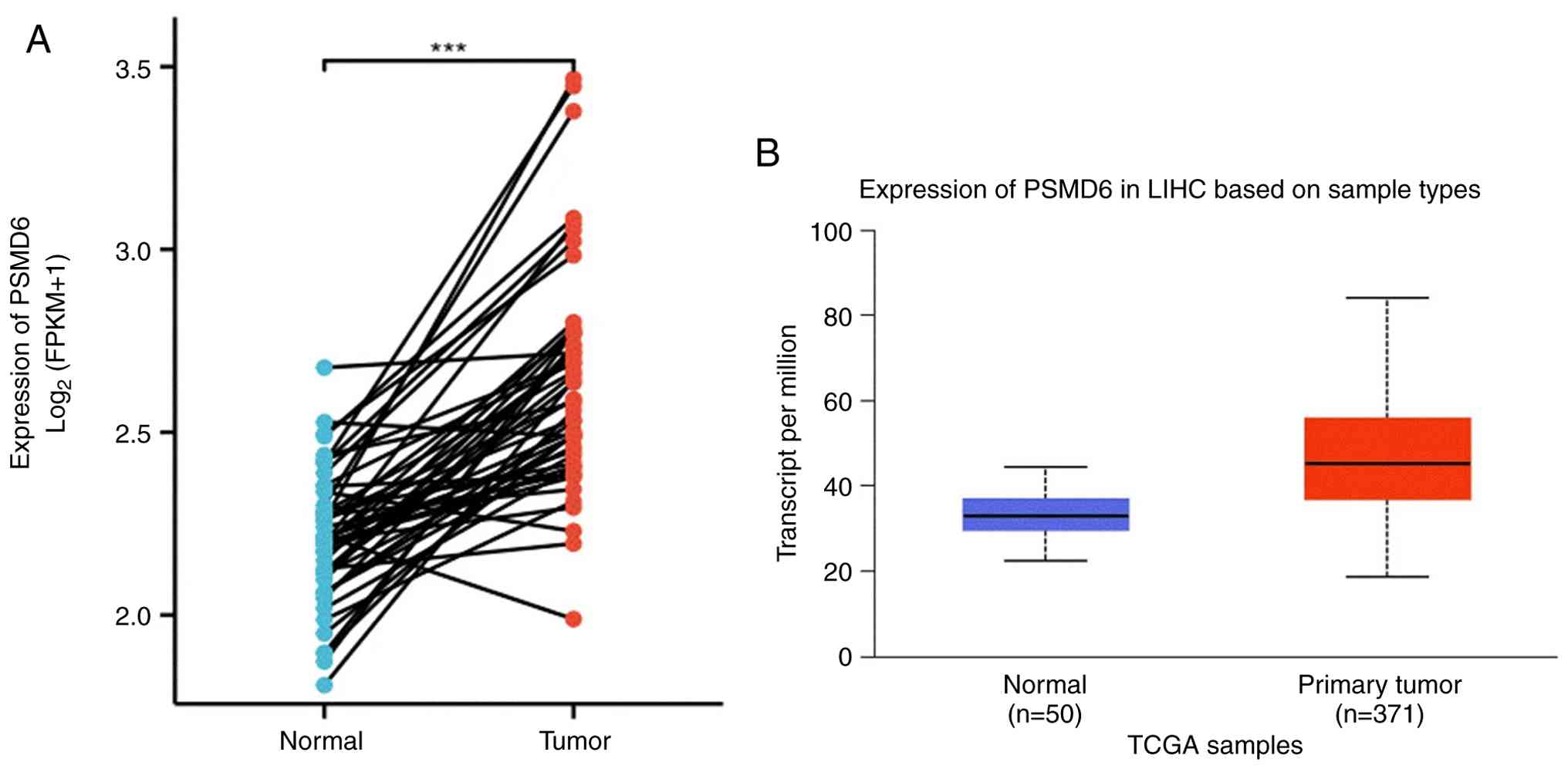
Subsequent focused analysis on LIHC from the TCGA
cohort demonstrated a pronounced overexpression of PSMD6 in tumor
specimens (n=371) compared with adjacent normal tissues (n=50;
P<0.001; Fig. 2A and B).
To evaluate the clinical relevance of PSMD6
overexpression, its correlation with key clinicopathological
parameters was examined using Xiantao tools. PSMD6 levels were
significantly elevated in HCC tissues across all pathology grades
(G1-G4), T stages (T1-T4), nodal involvement (N0/N1), metastasis
status (M0/M1), as well as irrespective of patient weight, sex and
ethnicity (all P<0.05) (Fig.
3A-G). This consistent upregulation across diverse clinical
subgroups underscores the potential of PSMD6 as a robust biomarker
associated with HCC progression and aggressiveness.
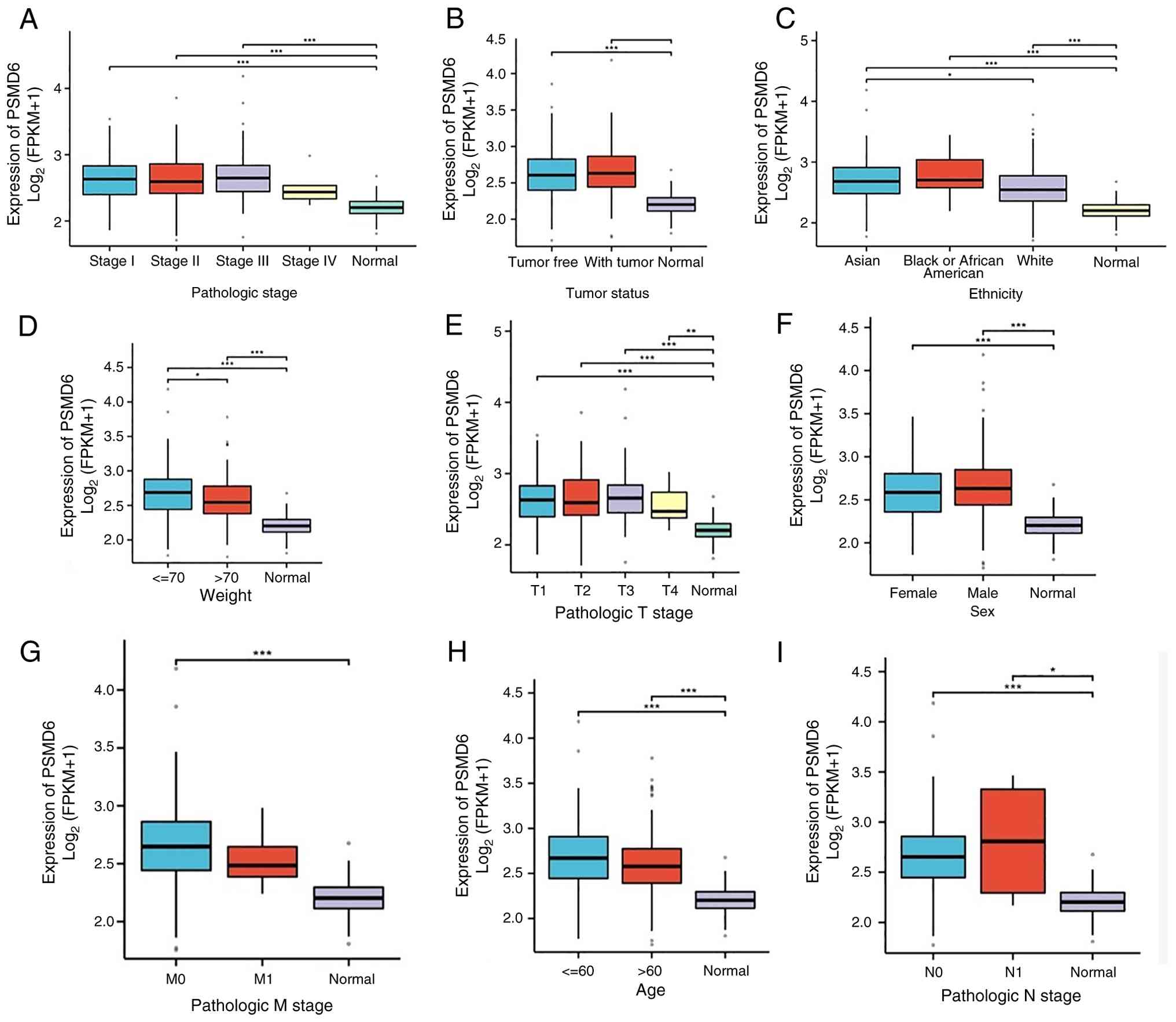 | Figure 3Association between PSMD6 expression
and clinicopathological characteristics in LIHC. (A-I) Box plots
showing PSMD6 expression levels stratified by (A) pathologic stage,
(B) tumor status, (C) ethnicity, (D) patient weight, (E) pathologic
T stage, (F) sex, (G) pathologic M stage, (H) age and (I)
pathologic N stage. Analyses were performed using data from The
Cancer Genome Atlas-LIHC cohort via XIANTAO tools.
*P<0.05, **P<0.01 and
***P<0.001. PSMD6, proteasome 26s subunit, non-ATPase
6; LIHC, liver hepatocellular carcinoma. |
High PSMD6 expression predicts poor
prognosis in patients with HCC
Given the strong association of PSMD6 with advanced
disease stages, the prognostic value of PSMD6 expression in HCC was
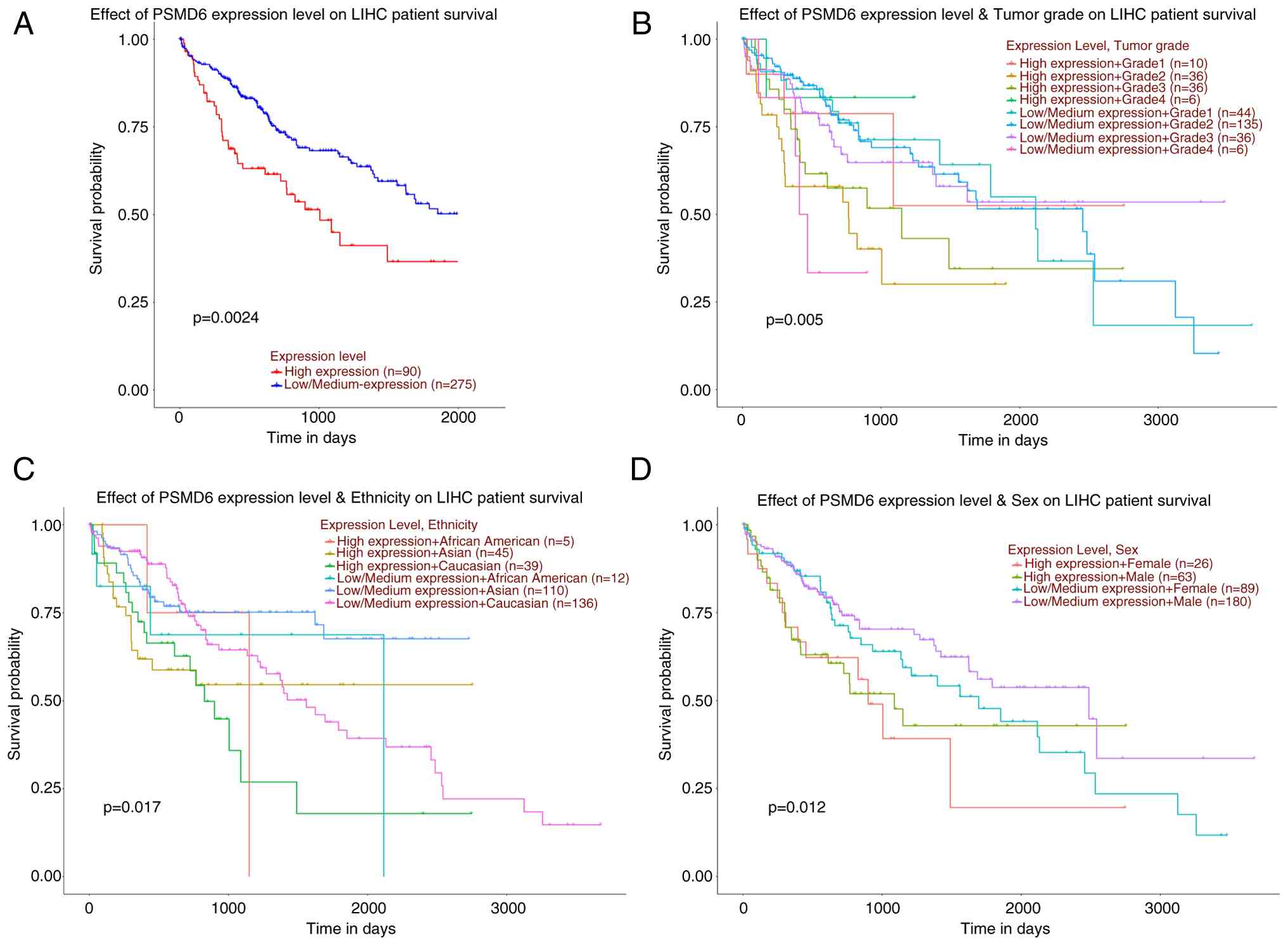
next evaluated. A 5-year survival analysis was conducted on
patients using the UALCAN database, and the results showed that
patients with high PSMD6 expression (n=90) had a significantly
lower probability of overall survival compared with those with low
expression (n=275; P<0.05; Fig.
4A).
In the subgroup analysis stratified by pathology
grade, the best prognosis was found in grade 1 patients with a low
expression level of PSMD6, and the worst prognosis was found
in grade 4 patients highly expressing PSMD6. In the subgroup
analysis stratified by ethnicity, African Americans with high
PSMD6 expression had the worst prognosis, while Caucasians
with low PSMD6 expression had the best prognosis. In the
subgroup analysis stratified by sex, female group with high
expression of PSMD6 had the most unfavorable prognosis, and
female group with low expression of PSMD6 had the best
prognosis. All comparisons exhibited statistical significance
(Fig. 4B-D).
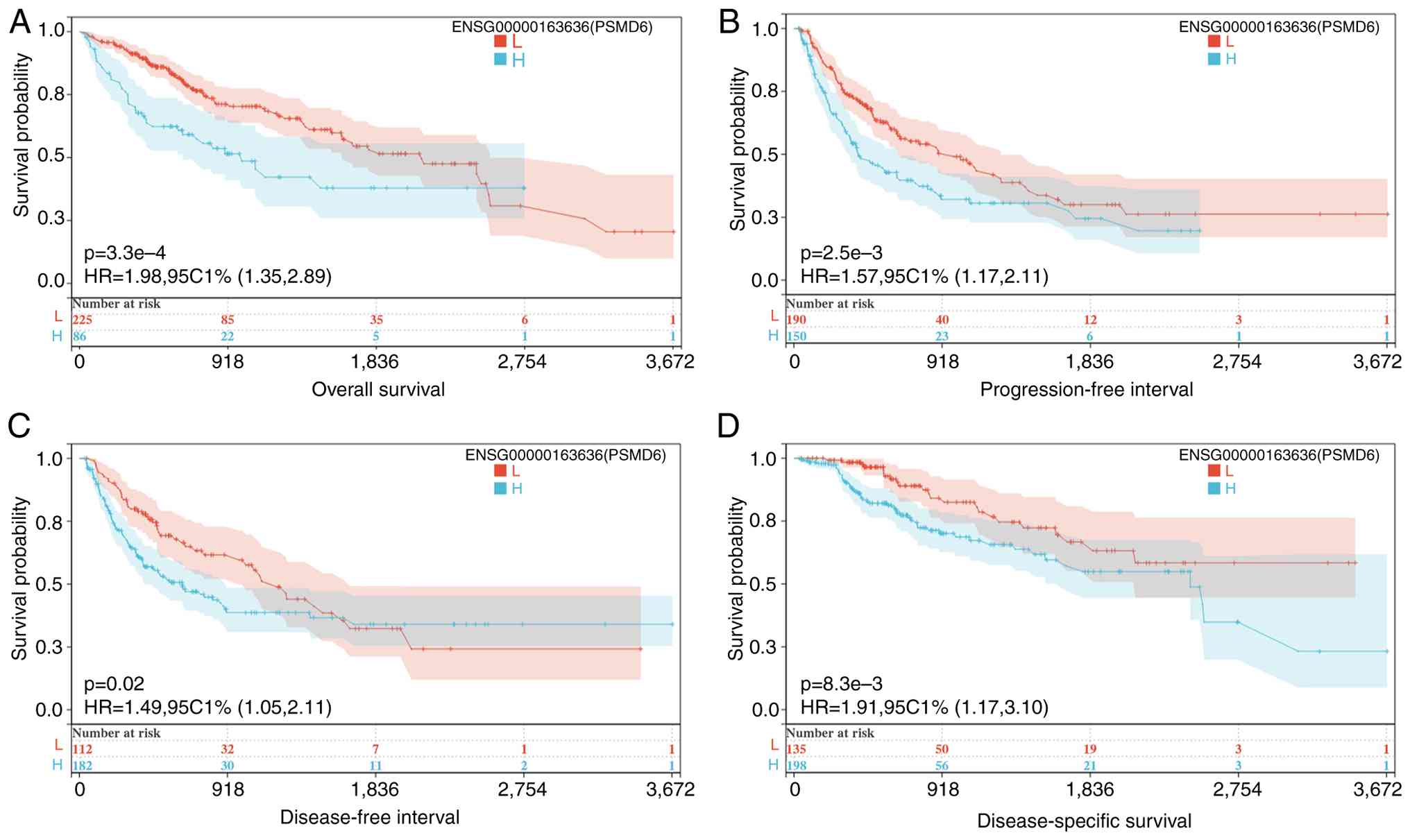
Further validation using the SANGERBOX database
corroborated these findings across multiple survival endpoints.
High PSMD6 expression was significantly associated with poorer OS,
DSS, disease-free interval and progression-free interval (all
P<0.05; Fig. 5).
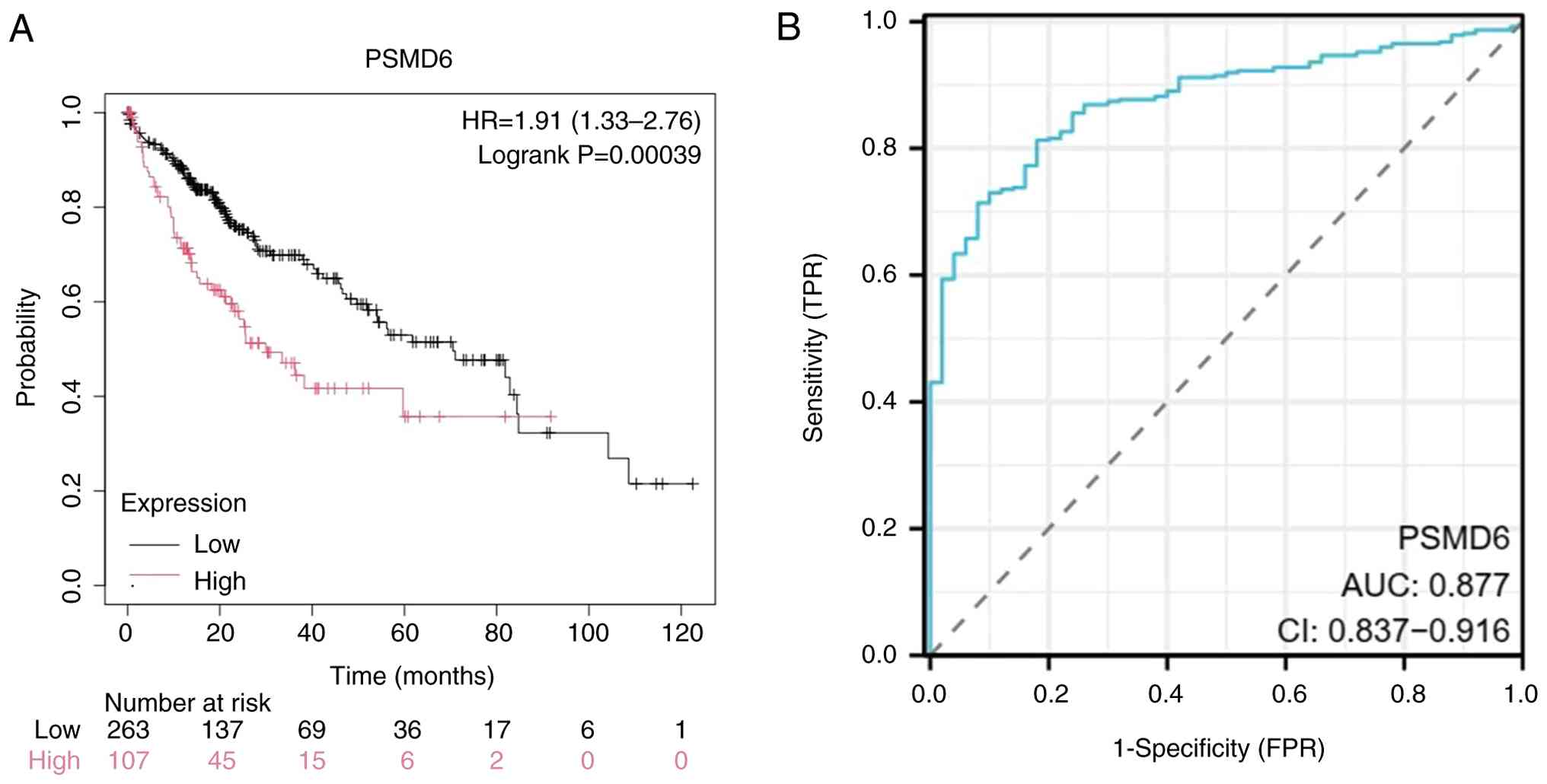
The robustness of PSMD6 as a prognostic indicator
was confirmed by Receiver Operating Characteristic (ROC) curve
analysis utilizing the Kaplan-Meier Plotter data repository. Cases
were classified into high-expression groups and low-expression
groups relying on the mean expression at different quantiles. The
analysis revealed that PSMD6 gene expression had a favorable
prognostic value (P=0.00039; Fig.
6A), which yielded an area under the curve of 0.877 (95%
confidence interval: 0.837-0.916; Fig.
6B), indicating high diagnostic accuracy.
Experimental validation confirms
elevated PSMD6 expression in HCC cell lines
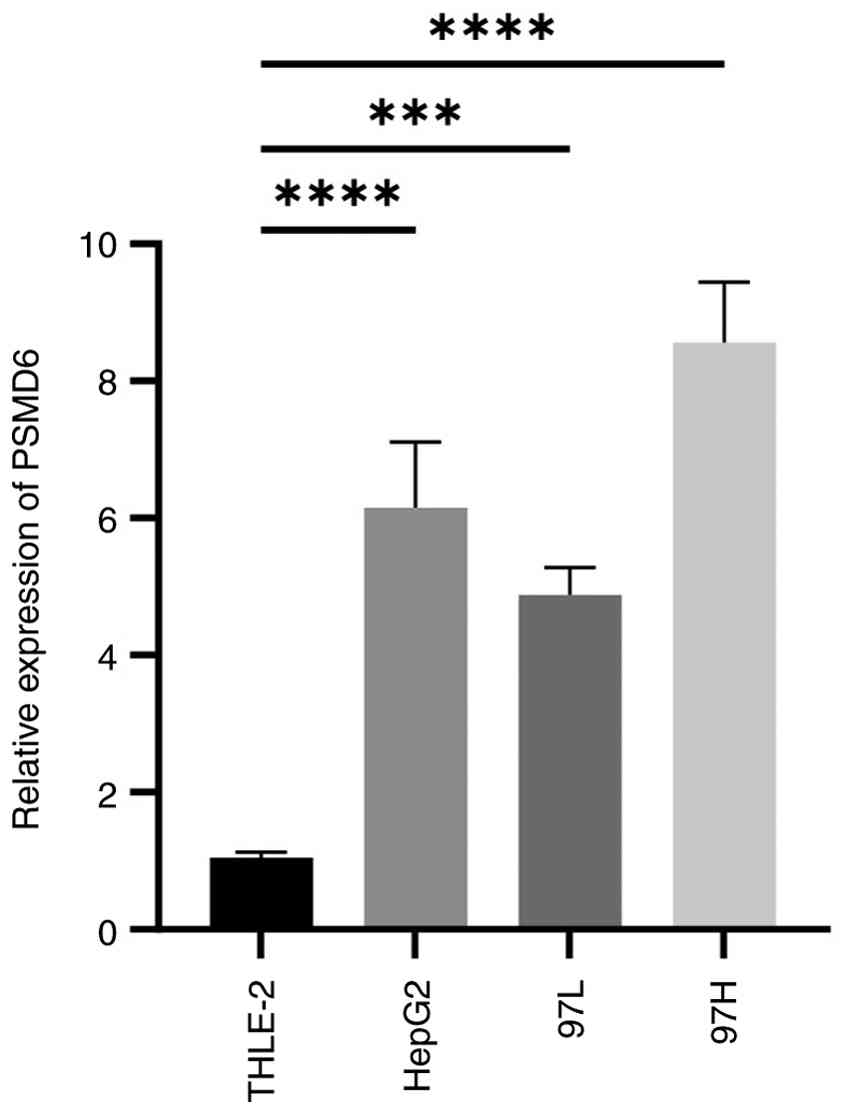
To further validate bioinformatic predictions, in
vitro experiments were performed using a normal human hepatic
cell line (THLE-2) and three distinct HCC cell lines (HepG2,
MHCC97-L and MHCC97-H). RT-qPCR analysis demonstrated that
PSMD6 mRNA levels were significantly elevated in all three
HCC cell lines compared with the normal hepatic cell line
(P<0.0001; Fig. 7).
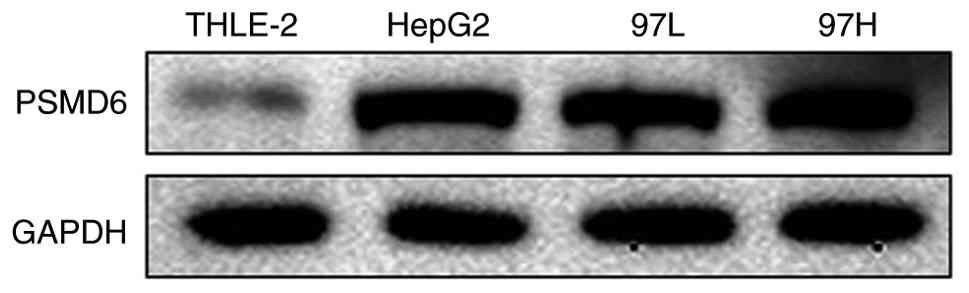
This transcriptional upregulation was further
confirmed at the protein level by western blot analysis, which
revealed consistently higher PSMD6 protein expression in the HCC
cell lines (Fig. 8). These
findings from mRNA and protein levels provide strong evidence for
the overexpression of PSMD6 in HCC.
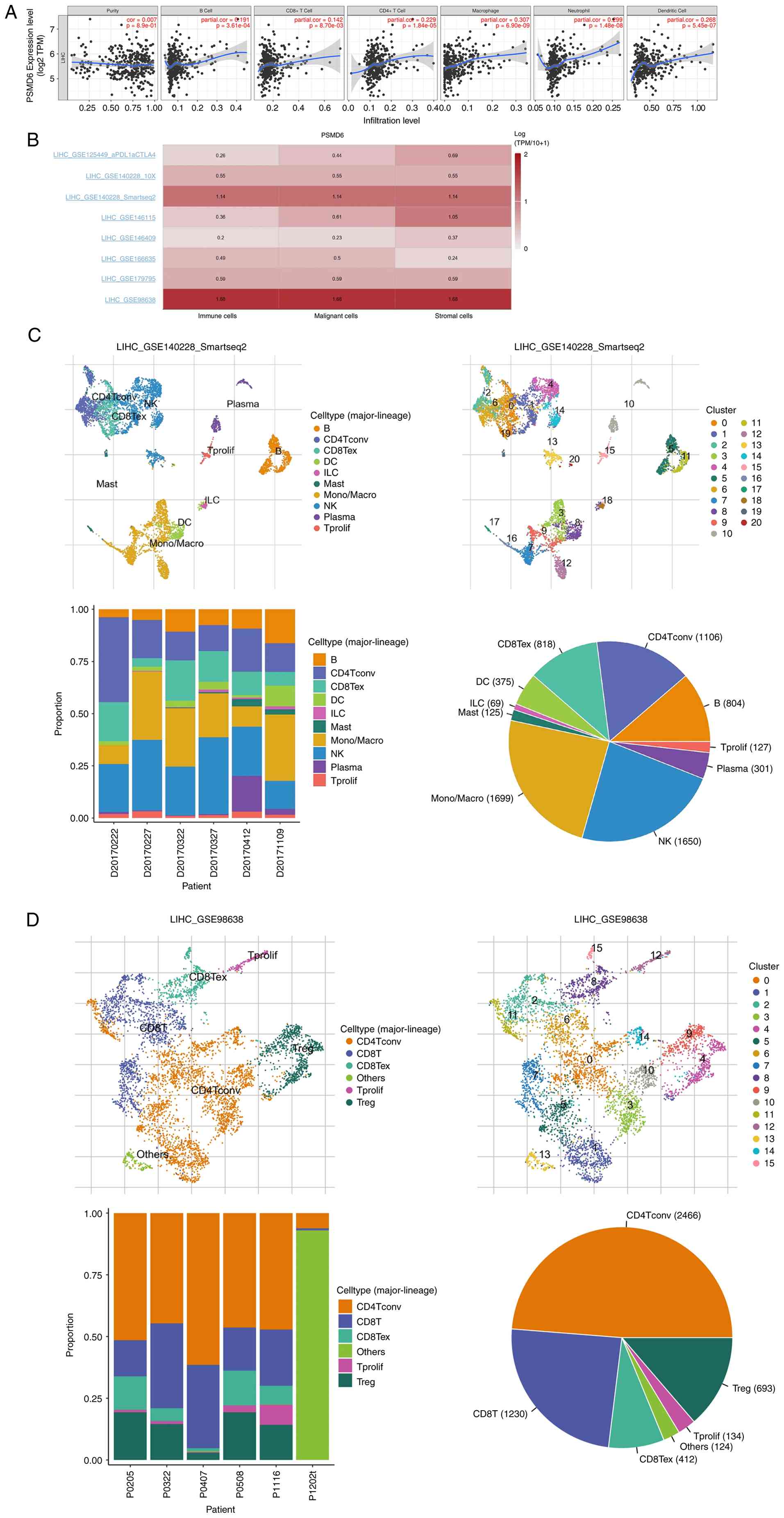
PSMD6 expression correlates with immune cell
infiltration in the TME. Tumorigenesis and development usually
correlate to infiltration of immune cells including B cell and
macrophage. In this context, the TIMER database was utilized to
investigate the relationship between PSMD6 expression and
the abundance of various immune cell populations within the HCC
TME. PSMD6 expression was found to be independent of tumor
purity (r=0.007, P=0.89). However, it exhibited significant
positive correlations with the infiltration levels of B cells
(r=0.191, P=0.00003), CD8+ T cells (r=0.1416, P=0.0087),
CD4+ T cells (r=0.2287, P=1.84x10-5),
neutrophils (r=0.299, P=1.48x10-8), macrophages
(r=0.307, P=6.9x10-9) and dendritic cells (r=0.268,
P=5.45x10-7) (Table
III, Fig. 9A).
 | Table IIIRelationship between proteasome 26s
subunit, non-ATPase 6 and immune cells in LIHC. |
Table III
Relationship between proteasome 26s
subunit, non-ATPase 6 and immune cells in LIHC.
| Cancer type | Variable | partial.cor | P-value |
|---|
| LIHC | Purity | 0.007451 | 0.890161 |
| LIHC | B cell | 0.191259 | 0.000361 |
| LIHC | CD8+ T
cell | 0.141669 | 0.008701 |
| LIHC | CD4+ T
cell | 0.2287 |
1.84x10-5 |
| LIHC | Macrophage | 0.307234 |
6.90x10-9 |
| LIHC | Neutrophil | 0.299014 |
1.48x10-8 |
| LIHC | Dendritic cell | 0.267707 |
5.45x10-7 |
To gain deeper, single-cell resolution insights, the
TISCH database was used to analyze the enrichment of PSMD6
gene in tumor and its immune microenvironment at the single-cell
level. Analysis of the GSE140228_SmartSeq2 and GSE98638 datasets
revealed that PSMD6 is predominantly enriched within
specific immune cell subsets, including monocytes/macrophages
(mono/macro) and natural killer (NK) cells in GSE140228_ smartseq2
single-cell cohort, and in conventional CD4+ T cells
(cd4tconv) and CD8+ T cells in GSE 98638 single-cell
cohort (Fig. 9B-D). This pattern
suggests that PSMD6 may influence the immune contexture of HCC,
potentially contributing to the immunosuppressive microenvironment
often observed in advanced tumors.
Functional enrichment and protein
interaction networks implicate PSMD6 in key oncogenic
processes
To elucidate the potential biological functions and
pathways through which PSMD6 contributes to hepatocarcinogenesis,
comprehensive functional enrichment analysis was performed.
Currently, Gene Ontology (GO) functional and Kyoto Encyclopedia of
Genes and Genomes (KEGG) pathway enrichment evaluations are the
methods that are used most frequently.
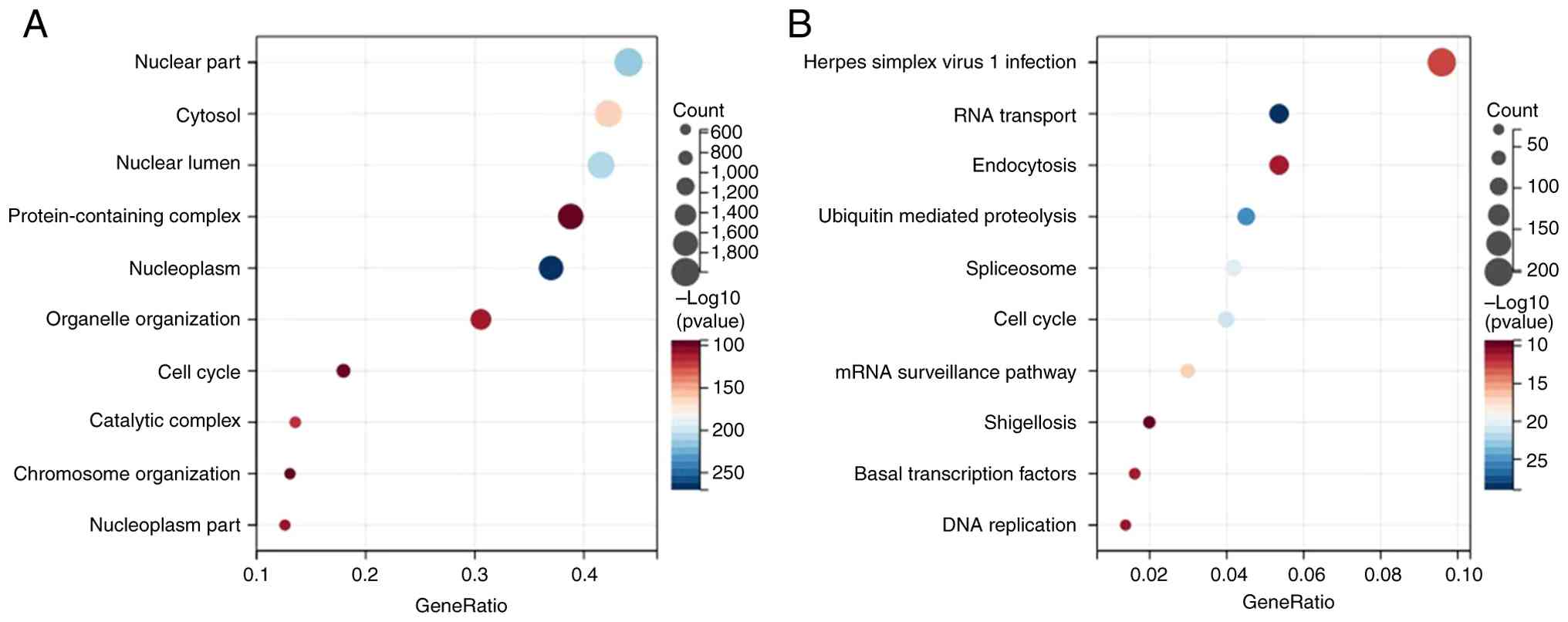
KEGG pathway analysis indicated that
PSMD6-coexpressed genes were significantly enriched in crucial
cancer-related pathways, including RNA transmit, Ubiquitin mediated
proteolysis, Cell cycle, mRNA surveillance pathway, Spliceosome,
Herpes simplex virus 1 infection, basal transcription factors,
shigellosis, endocytosis and DNA replication (Fig. 10). Concurrently, GO functional
analysis highlighted terms such as nucleoplasm, chromosome
organization, protein-containing complex and catalytic complex
(Fig. 10). The consistent
emergence of cell cycle and protein degradation pathways aligns
with the known role of the 26S proteasome, of which PSMD6 is
a regulatory subunit in controlling cell cycle progression and
protein turnover.
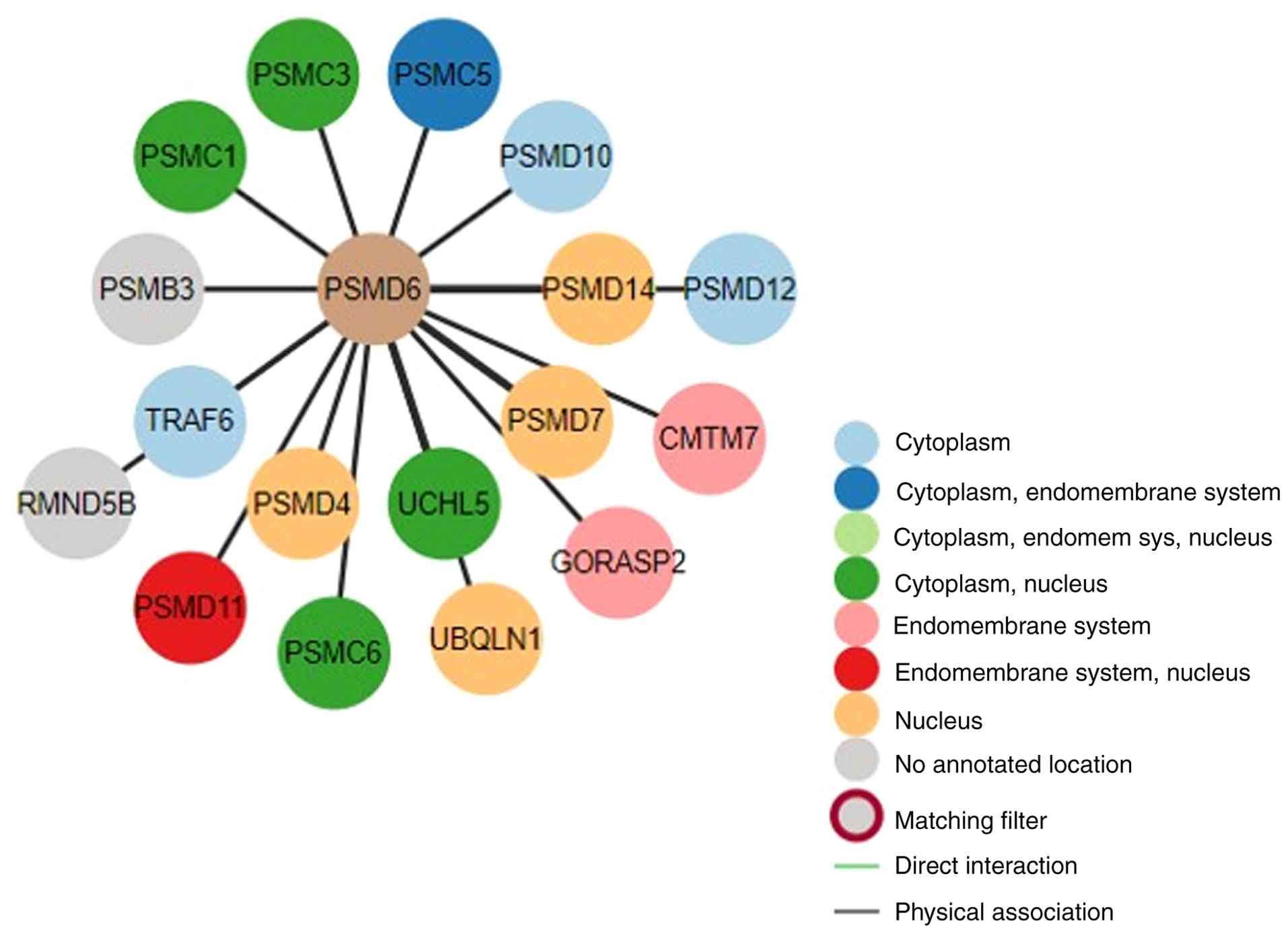
Furthermore, a PPI network was constructed to
identify potential functional partners of PSMD6. Queries using the
BioGRID and STRING databases revealed that PSMD6 interacts with
several core proteasomal subunits, including PSMC3, PSMD7, UCHL5,
PSMA2 and PSMB7 (P<0.001, Fig.
11).
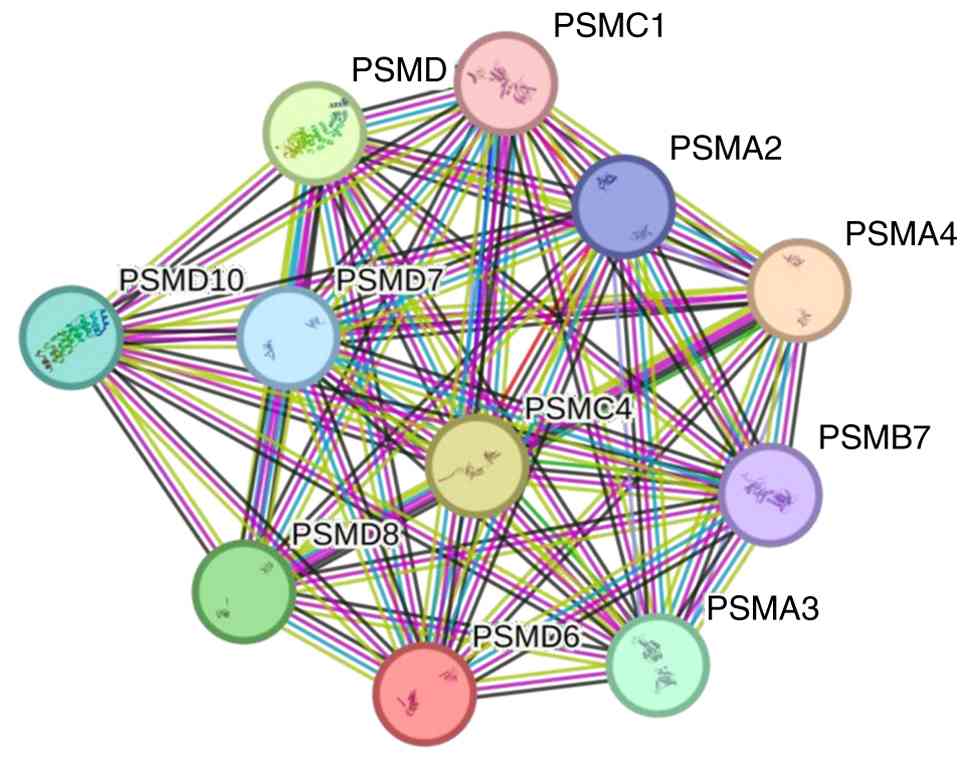
Moreover, as a PPI network can facilitate an
improved understanding of intracellular protein functions and
signal delivery, a PSMD6-centered PPI network was constructed using
STRING database. Multiple PSMD6-interacting proteins were
identified, such as PSMC1, PSMA2, PSMA4 and PSMB7 (Fig. 12), suggesting that its oncogenic
effects may be mediated through the regulation of proteasome
activity and subsequent impacts on critical cellular processes.
Discussion
Primary findings
The present multi-faceted analysis demonstrates that
PSMD6 is significantly overexpressed in HCC and is strongly
associated with malignant progression and poor patient survival.
Its influence likely extends to shaping the tumor immune
microenvironment. These findings nominate PSMD6 as a promising
diagnostic biomarker and a potential therapeutic target worthy of
further investigation in HCC.
Clinical significance and oncogenic
role of PSMD6 in HCC
Primary liver cancer is the seventh-most frequently
occurring cancer in the world and the second-most common cause of
cancer mortality (21). China has
the greatest number of primary liver cases, attributable to both an
elevated rate (18.3 per 100,000) and the world's largest population
(1.4 billion persons) (22).
Specifically, HCC is the dominant type of liver cancer, accounting
for ~75% of all liver cancers (23).
With the development of surgical intervention,
chemotherapy, targeted treatment and immunotherapy, the survival
rate of HCC cases has benefited more from multidisciplinary
treatment approaches. However, the survival rate remains low,
highlighting a pretty grim situation (24). Therefore, HCC remains a formidable
global health challenge, characterized by high mortality and
limited therapeutic options for advanced-stage patient.
The quest for reliable biomarkers for early
detection and prognosis prediction is therefore paramount. To
identify HCC early, current screening and surveillance strategies
depend on reliable and convenient serum biomarkers, but only
alpha-fetoprotein (AFP) has been extensively applied in clinical
settings (25). However, AFP has
poor sensitivity and specificity. The present study identified
PSMD6 as a potential key player in HCC pathogenesis. Bioinformatics
analysis across multiple databases (for example, TCGA and GTEx)
consistently revealed significant upregulation of PSMD6 in HCC
tissues compared with normal liver tissues. This finding was
robustly validated in vitro, where both mRNA and protein
levels of PSMD6 were markedly elevated in HCC cell lines (HepG2,
MHCC97-L, MHCC97-H) compared with the normal hepatic THLE-2 cell
line.
These findings align with and extend previous
bioinformatics studies that also identified PSMD6 as a
differentially methylated gene with significant prognostic value in
HCC (26,27). The consistency of PSMD6
overexpression across diverse cohorts and its strong association
with clinical outcomes underscore its potential as a robust
diagnostic and prognostic biomarker.
Potential mechanisms underlying PSMD6
in HCC
Proteasomes rapidly catalyze a diverse range of
biological reactions and then regulate the bioactivity of multiple
cells (28). In a previous study,
a proteasome system in Caenorhabditis elegans germline was
constructed and it was found that PSMD6 downregulation led
to reduced proteasome proteolytic activity and cell cycle defect
(29). PSMD family members
may exhibit an increasing trend in tumors due to the
characteristics of extremely high protein metabolic level and very
short cell division cycle in tumor cells. The PPI network analysis
revealed that PSMD6 interacts with core proteasomal subunits (for
example, PSMC3, PSMD7 and PSMA2), underscoring its integral role
within the 26S proteasome complex. This interaction network
suggests that the oncogenic effects of PSMD6 are likely mediated
through the regulation of proteasome activity, impacting a wide
array of downstream cellular processes.
Dysregulation of UPS components, including various
PSMD family members, is increasingly recognized as a hallmark of
cancer, contributing to uncontrolled proliferation and evasion of
apoptosis. The functional enrichment analyses (GO and KEGG) of the
present study shed light on the potential mechanisms by which PSMD6
contributes to hepatocarcinogenesis. PSMD6 was significantly
enriched in essential biological processes such as the cell cycle,
ubiquitin-mediated proteolysis and DNA replication.
A pivotal finding of the present study is the
significant correlation between PSMD6 expression and immune cell
infiltration. PSMD6 levels positively correlated with the abundance
of various immune cells, including B cells, CD4+ T
cells, CD8+ T cells, macrophages and dendritic cells.
scRNA-seq data from the TISCH database further indicated that PSMD6
is enriched in specific immune subsets including
monocytes/macrophages, NK cells and T cells within the HCC
microenvironment. This suggests that PSMD6 may influence the
functional state of the immune landscape, potentially fostering an
immunosuppressive niche that facilitates immune evasion. The role
of the immune microenvironment in determining HCC prognosis is
well-established, and the current findings position PSMD6 as a
novel modulator within this context.
Immunotherapy is important for patients with HCC
(30,31). Given the significant connection
amid the expression of PSMD6 and immune cell infiltration,
patients with HCC with poor prognosis and survival may benefit from
immunotherapy. However, whether PSMD6 gene expression affects the
effectiveness of immunotherapy warrants further validation.
The present study also noted that Zhou et al
(27) previously proposed in their
research that PSMD6 and its homolog PSMD11 may participate in liver
cancer progression and treatment response by regulating proteasome
dependent protein degradation pathways, thereby affecting the
stability of key cancer proteins such as p53 and Cyclin. This
hypothesis provides an important mechanistic perspective for the
findings of the present study. The experimental results further
support this viewpoint: PSMD6 expression is significantly
upregulated in liver cancer tissues, and its high expression is
closely related to poor prognosis in patients. Based on previous
studies, it was hypothesized that PSMD6 may promote the development
of liver cancer through the following pathways: Enhancing
proteasome activity and accelerating the degradation of tumor
suppressor proteins; regulating the stability of cell cycle and
apoptosis related proteins through the ubiquitin proteasome system;
synergistic effect with subunits such as PSMD11 affects the
proliferation and invasion ability of liver cancer cells. Future
research can further validate the specific regulatory mechanism of
PSMD6 in liver cancer by knocking down/overexpressing it and
detecting downstream target protein levels, providing new ideas for
targeted proteasome subunits in liver cancer therapy.
While the integrated bioinformatics and preliminary
experimental data highlight the significance of PSMD6 in HCC, the
present study has several limitations. First, the bioinformatic
findings, though validated across multiple databases, require
further confirmation in larger, prospectively collected clinical
cohorts to solidify their clinical translatability. Differences in
sample sources, sequencing platforms and batches in public
databases may lead to biased results, affecting the comparability
of PSMD6 expression. Second, the precise molecular
mechanisms by which PSMD6 regulates the cell cycle, immune
infiltration and other cancer-related pathways remain to be fully
elucidated through detailed functional experiments. Only partial
in vitro experimental studies were conducted without tissue
validation. No stratified analysis was conducted in the present
study based on liver cancer subtypes, staging, or treatment
response, which may mask potential heterogeneity and therefore lead
to weak clinical correlation analysis Although the co-expression
network or pathway enrichment of PSMD6 can be predicted, its
upstream regulation (such as methylation, miRNA) and downstream
effects in liver cancer still require experimental exploration.
Prospective cohorts and animal models are needed to further
validate its prognostic value and therapeutic potential.
In conclusion, PSMD6 was found to be significantly
overexpressed in HCC and is strongly associated with malignant
progression and poor patient survival. Its influence likely extends
to shaping the tumor immune microenvironment. These findings
nominate PSMD6 as a promising diagnostic biomarker and a
potential therapeutic target worthy of further investigation in
HCC. Further experiments and clinical samples are needed for
verification.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
JL proposed the initial research concept, was
responsible for collecting the data from the database, actively
participated in conducting the experiments, and wrote the first
complete version of the manuscript. TW contributed to the study
design, performed the PCR and Western blot experiments, and
critically revised the manuscript for important intellectual
content in response to the editor's comments. ZZ contributed to the
study design and methodology, provided critical supervision and
guidance throughout the experimental phase, rigorously validated
all collected data, and meticulously reviewed, critiqued, and
revised multiple drafts of the manuscript to ensure its academic
rigor and clarity. All authors read and approved the final version
of the manuscript. JL and TW confirm the authenticity of all the
raw data.
Ethics approval and consent to
participate
The present study was conducted in accordance with
the declaration of Helsinki and was approved by the Ethics
Committee of The First Hospital of Hunan University of Chinese
Medicine (approval no. HN-LL-LW-2024-062; Changsha, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Calderaro J, Seraphin TP, Luedde T and
Simon TG: Artificial intelligence for the prevention and clinical
management of hepatocellular carcinoma. J Hepatol. 76:1348–1361.
2022.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Zheng RS, Chen R, Han BF, Wang SM, Li L,
Sun KX, Zeng HM, Wei WW and He J: Cancer incidence and mortality in
China, 2022. Zhonghua Zhong Liu Za Zhi (Chinese). 46:221–231.
2024.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Qiu F, Qiu F, Liu L, Liu J, Xu J and Huang
X: The role of dermcidin in the diagnosis and staging of
hepatocellular carcinoma. Genet Test Mol Biomarkers. 22:218–223.
2018.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Urquijo-Ponce JJ, Alventosa-Mateu C,
Latorre-Sánchez M, Castelló-Miralles I and Diago M: Present and
future of new systemic therapies for early and intermediate stages
of hepatocellular carcinoma. World J Gastroenterol. 30:2512–2522.
2024.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Hoti E and Adam R: Liver transplantation
for primary and metastatic liver cancers. Transpl Int.
21:1107–1117. 2008.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Sapisochin G and Bruix J: Liver
transplantation for hepatocellular carcinoma: outcomes and novel
surgical approaches. Nat Rev Gastroenterol Hepatol. 14:203–217.
2017.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Neuberger J: An update on liver
transplantation: A critical review. J Autoimmun. 66:51–59.
2016.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Gao S, Gang J, Yu M, Xin G and Tan H:
Computational analysis for identification of early diagnostic
biomarkers and prognostic biomarkers of liver cancer based on GEO
and TCGA databases and studies on pathways and biological functions
affecting the survival time of liver cancer. BMC Cancer.
21(791)2021.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Fabregat I: Dysregulation of apoptosis in
hepatocellular carcinoma cells. World J Gastroenterol. 15:513–520.
2009.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Li X, Li X, Hu Y, Liu O, Wang Y, Li S,
Yang Q and Lin B: PSMD8 can serve as potential biomarker and
therapeutic target of the PSMD family in ovarian cancer: based on
bioinformatics analysis and in vitro validation. BMC Cancer.
23(573)2023.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Jiang L, Qi X, Lai M, Zhou J, Yuan M, You
J, Liu Q, Pan J, Zhao L, Ying M, et al: WDR20 prevents
hepatocellular carcinoma senescence by orchestrating the
simultaneous USP12/46-mediated deubiquitination of c-Myc. Proc Natl
Acad Sci USA. 121(e2407904121)2024.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Wang M, Yu F and Li P: Intratumor
microbiota in cancer pathogenesis and immunity: From mechanisms of
action to therapeutic opportunities. Front Immunol.
14(1269054)2023.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Li H, Ji Z, Paulo JA, Gygi SP and Rapoport
TA: Bidirectional substrate shuttling between the 26S proteasome
and the Cdc48 ATPase promotes protein degradation. Mol Cell.
84:1290–1303.e7. 2024.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Coll-Martínez B and Crosas B: How the 26S
proteasome degrades ubiquitinated proteins in the cell.
biomolecules. 9(395)2019.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Xuan DTM, Wu CC, Kao TJ, Ta HDK, Anuraga
G, Andriani V, Athoillah M, Chiao CC, Wu YF, Lee KH, et al:
Prognostic and immune infiltration signatures of proteasome 26S
subunit, non-ATPase (PSMD) family genes in breast cancer patients.
Aging (Albany NY). 13:24882–24913. 2021.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Li Y, Liu X, Zhao F, Zhao Z, Li X, Wang J,
Huang B and Chen A: Comprehensive analysis of PSMD family members
and validation of PSMD9 as a potential therapeutic target in human
glioblastoma. CNS Neurosci Ther. 30(e14366)2024.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Zhang C, Xu T, Ji K, Cao S, Cao Y, Ai J,
Jing L and Sun JH: An integrative analysis reveals the prognostic
value and potential functions of PSMD11 in hepatocellular
carcinoma. Mol Carcinog. 62:1355–1368. 2023.PubMed/NCBI View
Article : Google Scholar
|
|
18
|
Huang W, Mei J, Liu YJ, Li JP, Zou X, Qian
XP and Zhang Y: An analysis regarding the association between
proteasome (PSM) and hepatocellular carcinoma (HCC). J Hepatocell
Carcinoma. 10:497–515. 2023.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Tsolou A, Nelson G, Trachana V,
Chondrogianni N, Saretzki G, von Zglinicki T and Gonos ES: The 19S
proteasome subunit Rpn7 stabilizes DNA damage foci upon genotoxic
insult. IUBMB Life. 64:432–442. 2012.PubMed/NCBI View
Article : Google Scholar
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2-ΔΔCT method. Methods. 25:402–408. 2001.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Han B, Zheng R, Zeng H, Wang S, Sun K,
Chen R, Li L, Wei W and He J: Cancer incidence and mortality in
China, 2022. J Natl Cancer Cent. 4:47–53. 2024.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Valery PC, Laversanne M, Clark PJ, Petrick
JL, McGlynn KA and Bray F: Projections of primary liver cancer to
2030 in 30 countries worldwide. Hepatology. 67:600–611.
2018.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Singal AG, Kanwal F and Llovet JM: Global
trends in hepatocellular carcinoma epidemiology: Implications for
screening, prevention and therapy. Nat Rev Clin Oncol. 20:864–884.
2023.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Zhao K, Zhou X, Xiao Y, Wang Y and Wen L:
Research progress in alpha-fetoprotein-induced immunosuppression of
liver cancer. Mini Rev Med Chem. 22:2237–2243. 2022.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Zhou SS, Ye YP, Chen Y, Zeng DT, Zheng GC,
He RQ, Chi BT, Wang L, Lin Q, Su QY, et al: Overexpression pattern,
function, and clinical value of proteasome 26S subunit non-ATPase 6
in hepatocellular carcinoma. World J Clin Oncol.
16(99839)2025.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Zhou C, Li H, Han X, Pang H, Wu M, Tang Y
and Luo X: Prognostic value and molecular mechanisms of proteasome
26S subunit, non-ATPase family genes for pancreatic ductal
adenocarcinoma patients after pancreaticoduodenectomy. J Invest
Surg. 35:330–346. 2022.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Guedes RA, Serra P, Salvador JA and Guedes
RC: Computational approaches for the discovery of human proteasome
inhibitors: An overview. Molecules. 21(927)2016.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Zong Q, Mao B, Zhang HB, Wang B, Yu WJ,
Wang ZW and Wang YF: Comparative Ubiquitome analysis reveals
deubiquitinating effects induced by Wolbachia infection in
drosophila melanogaster. Int J Mol Sci. 23(9459)2022.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Donne R and Lujambio A: The liver cancer
immune microenvironment: Therapeutic implications for
hepatocellular carcinoma. Hepatology. 77:1773–1796. 2023.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Ruff SM, Shannon AH, Beane JD and Pawlik
TM: Highlighting novel targets in immunotherapy for liver cancer.
Expert Rev Gastroenterol Hepatol. 16:1029–1041. 2022.PubMed/NCBI View Article : Google Scholar
|