|
1
|
Siegel RL, Giaquinto AN and Jemal A:
Cancer statistics, 2024. CA Cancer J Clin. 74:12–49.
2024.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Qu R, Ma Y, Zhang Z and Fu W: Increasing
burden of colorectal cancer in China. Lancet Gastroenterol Hepatol.
7(700)2022.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Dantas AAG, de Oliveira NPD, Costa GAB,
Martins LFL, Dos Santos JEM, Migowski A, de Camargo Cancela M and
de Souza DLB: Multilevel analysis of social determinants of
advanced stage colorectal cancer diagnosis. Sci Rep.
14(9667)2024.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Myers DJ and Arora K: Villous adenoma
(Archived). In: StatPearls. StatPearls Publishing, Treasure Island,
FL, 2025.
|
|
5
|
Nguyen LH, Goel A and Chung DC: Pathways
of colorectal carcinogenesis. Gastroenterology. 158:291–302.
2020.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Uno Y, Murayama N, Kunori M and Yamazaki
H: Characterization of microsomal glutathione S-transferases MGST1,
MGST2, and MGST3 in cynomolgus macaque. Drug Metab Dispos.
41:1621–1625. 2013.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Bolger AM, Lohse M and Usadel B:
Trimmomatic: A flexible trimmer for Illumina sequence data.
Bioinformatics. 30:2114–2120. 2014.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Dobin A, Davis CA, Schlesinger F, Drenkow
J, Zaleski C, Jha S, Batut P, Chaisson M and Gingeras TR: STAR:
Ultrafast universal RNA-seq aligner. Bioinformatics. 29:15–21.
2013.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Ghosh S and Chan CK: Analysis of RNA-seq
data using tophat and cufflinks. Methods Mol Biol. 1374:339–361.
2016.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Adan A, Kiraz Y and Baran Y: Cell
proliferation and cytotoxicity assays. Curr Pharm Biotechnol.
17:1213–1221. 2016.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Franken NA, Rodermond HM, Stap J, Haveman
J and van Bree C: Clonogenic assay of cells in vitro. Nat Protoc.
1:2315–2319. 2006.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Liang CC, Park AY and Guan JL: In vitro
scratch assay: A convenient and inexpensive method for analysis of
cell migration in vitro. Nat Protoc. 2:329–333. 2007.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Justus CR, Marie MA, Sanderlin EJ and Yang
LV: Transwell in vitro cell migration and invasion assays. Methods
Mol Biol. 2644:349–359. 2023.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Mahmood T and Yang PC: Western blot:
Technique, theory, and trouble shooting. N Am J Med Sci. 4:429–434.
2012.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Schubert M, Klinger B, Klünemann M, Sieber
A, Uhlitz F, Sauer S, Garnett MJ, Blüthgen N and Saez-Rodriguez J:
Perturbation-response genes reveal signaling footprints in cancer
gene expression. Nat Commun. 9(20)2018.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Szklarczyk D, Nastou K, Koutrouli M,
Kirsch R, Mehryary F, Hachilif R, Hu D, Peluso ME, Huang Q, Fang T,
et al: The STRING database in 2025: protein networks with
directionality of regulation. Nucleic Acids Res. 53:D730–D737.
2025.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Guo X, Liang X, Wang Y, Cheng A, Zhang H,
Qin C and Wang Z: Significance of tumor mutation burden combined
with immune infiltrates in the progression and prognosis of
advanced gastric cancer. Front Genet. 12(642608)2021.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Mayakonda A, Lin DC, Assenov Y, Plass C
and Koeffler HP: Maftools: Efficient and comprehensive analysis of
somatic variants in cancer. Genome Res. 28:1747–1756.
2018.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Tan TZ, Miow QH, Miki Y, Noda T, Mori S,
Huang RY and Thiery JP: Epithelial-mesenchymal transition spectrum
quantification and its efficacy in deciphering survival and drug
responses of cancer patients. EMBO Mol Med. 6:1279–1293.
2014.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Subramanian A, Tamayo P, Mootha VK,
Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub
TR, Lander ES and Mesirov JP: Gene set enrichment analysis: A
knowledge-based approach for interpreting genome-wide expression
profiles. Proc Natl Acad Sci USA. 102:15545–15550. 2005.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Chen B, Khodadoust MS, Liu CL, Newman AM
and Alizadeh AA: Profiling tumor infiltrating immune cells with
CIBERSORT. Methods Mol Biol. 1711:243–259. 2018.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Yoshihara K, Shahmoradgoli M, Martínez E,
Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW,
Levine DA, et al: Inferring tumour purity and stromal and immune
cell admixture from expression data. Nat Commun.
4(2612)2013.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Yan H, Talty R and Johnson CH: Targeting
ferroptosis to treat colorectal cancer. Trends Cell Biol.
33:185–188. 2023.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Jardim BV, Moschetta MG, Leonel C,
Gelaleti GB, Regiani VR, Ferreira LC, Lopes JR and Zuccari DAPDC:
Glutathione and glutathione peroxidase expression in breast cancer:
An immunohistochemical and molecular study. Oncol Rep.
30:1119–1128. 2013.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Inoue T, Ishida T, Sugio K, Maehara Y and
Sugimachi K: Glutathione S transferase Pi is a powerful indicator
in chemotherapy of human lung squamous-cell carcinoma. Respiration.
62:223–227. 1995.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Xu H, Hu C, Wang Y, Shi Y, Yuan L, Xu J,
Zhang Y, Chen J, Wei Q, Qin J, et al: Glutathione peroxidase 2
knockdown suppresses gastric cancer progression and metastasis via
regulation of kynurenine metabolism. Oncogene. 42:1994–2006.
2023.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Xiao Y and Meierhofer D: Glutathione
metabolism in renal cell carcinoma progression and implications for
therapies. Int J Mol Sci. 20(3672)2019.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Sobhakumari A, Love-Homan L, Fletcher EV,
Martin SM, Parsons AD, Spitz DR, Knudson CM and Simons AL:
Susceptibility of human head and neck cancer cells to combined
inhibition of glutathione and thioredoxin metabolism. PLoS One.
7(e48175)2012.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Schmitt M and Greten FR: The inflammatory
pathogenesis of colorectal cancer. Nat Rev Immunol. 21:653–667.
2021.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Kalluri R and Weinberg RA: The basics of
epithelial-mesenchymal transition. J Clin Invest. 119:1420–1428.
2009.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Bethmann D, Feng Z and Fox BA:
Immunoprofiling as a predictor of patient's response to cancer
therapy-promises and challenges. Curr Opin Immunol. 45:60–72.
2017.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Tiwari N, Gheldof A, Tatari M and
Christofori G: EMT as the ultimate survival mechanism of cancer
cells. Semin Cancer Biol. 22:194–207. 2012.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Wellenstein MD and de Visser KE:
Cancer-cell-intrinsic mechanisms shaping the tumor immune
landscape. Immunity. 48:399–416. 2018.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Mukherjee AG, Wanjari UR, Namachivayam A,
Murali R, Prabakaran DS, Ganesan R, Renu K, Dey A, Vellingiri B,
Ramanathan G, et al: Role of immune cells and receptors in cancer
treatment: An immunotherapeutic approach. Vaccines (Basel).
10(1493)2022.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Mazari AMA, Zhang L, Ye ZW, Zhang J, Tew
KD and Townsend DM: The multifaceted role of glutathione
S-transferases in health and disease. Biomolecules.
13(688)2023.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Moyer MP, Manzano LA, Merriman RL,
Stauffer JS and Tanzer LR: NCM460, a normal human colon mucosal
epithelial cell line. In Vitro Cell Dev Biol Anim. 32:315–317.
1996.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Xue X, Wang M, Cui J, Yang M, Ma L, Kang
R, Tang D and Wang J: Glutathione metabolism in ferroptosis and
cancer therapy. Cancer Lett. 621(217697)2025.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Ma T, Du J, Zhang Y, Wang Y, Wang B and
Zhang T: GPX4-independent ferroptosis-a new strategy in disease's
therapy. Cell Death Discov. 8(434)2022.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Sun X, Ou Z, Xie M, Kang R, Fan Y, Niu X,
Wang H, Cao L and Tang D: HSPB1 as a novel regulator of ferroptotic
cancer cell death. Oncogene. 34:5617–5625. 2015.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Zhang Y, Zou L, Li X, Guo L, Hu B, Ye H
and Liu Y: SLC40A1 in iron metabolism, ferroptosis, and disease: A
review. WIREs Mech Dis. 16(e1644)2024.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Chaudhary N, Choudhary BS, Shah SG,
Khapare N, Dwivedi N, Gaikwad A, Joshi N, Raichanna J, Basu S,
Gurjar M, et al: Lipocalin 2 expression promotes tumor progression
and therapy resistance by inhibiting ferroptosis in colorectal
cancer. Int J Cancer. 149:1495–1511. 2021.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Klusek J, Głuszek S and Klusek J: GST gene
polymorphisms and the risk of colorectal cancer development.
Contemp Oncol (Pozn). 18:219–221. 2014.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Xie Z, Kawasaki T, Zhou H, Okuzaki D,
Okada N and Tachibana M: Targeting GGT1 eliminates the
tumor-promoting effect and enhanced immunosuppressive function of
myeloid-derived suppressor cells caused by G-CSF. Front Pharmacol.
13(873792)2022.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Chen J, Wu Y, Zhou Q, Song Y, Zhuang J, Lu
K and Yang X: GPX3 is a key cholesterol-related gene associated
with prognosis and tumor-infiltrating T cells in colorectal cancer.
Neoplasma. 70(230704N348)2023.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Lagal DJ, Montes-Osuna AM, Ortiz-Olivencia
A, Arribas-Parejas C, Ortiz-Alcántara Á, Pescuezo-Castillo C,
Bárcena JA, Padilla CA and Requejo-Aguilar R: Tumoral malignancy
decreases coupled with higher ROS and lipid peroxidation in HCT116
Colon cancer cells upon loss of PRDX6. Antioxidants (Basel).
13(881)2024.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Merino DM, McShane LM, Fabrizio D, Funari
V, Chen SJ, White JR, Wenz P, Baden J, Barrett JC, Chaudhary R, et
al: Establishing guidelines to harmonize tumor mutational burden
(TMB): In silico assessment of variation in TMB quantification
across diagnostic platforms: Phase I of the friends of cancer
research TMB harmonization project. J ImmunoTher Cancer.
8(e000147)2020.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Sha D, Jin Z, Budczies J, Kluck K,
Stenzinger A and Sinicrope FA: Tumor mutational burden as a
predictive biomarker in solid tumors. Cancer Discov. 10:1808–1825.
2020.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Zgura A, Chipuc S, Bacalbasa N, Haineala
B, Rodica A and Sebastian V: Evaluating tumour mutational burden as
a key biomarker in personalized cancer immunotherapy: A Pan-cancer
systematic review. Cancers (Basel). 17(480)2025.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Kim ES, Velcheti V, Mekhail T, Yun C,
Shagan SM, Hu S, Chae YK, Leal TA, Dowell JE, Tsai ML, et al:
Blood-based tumor mutational burden as a biomarker for atezolizumab
in non-small cell lung cancer: The phase 2 B-F1RST trial. Nat Med.
28:939–945. 2022.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Addeo A, Friedlaender A, Banna GL and
Weiss GJ: TMB or not TMB as a biomarker: That is the question. Crit
Rev Oncol/Hematol. 163(103374)2021.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Brabletz T, Kalluri R, Nieto MA and
Weinberg RA: EMT in cancer. Nat Rev Cancer. 18:128–134.
2018.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Kim DH, Xing T, Yang Z, Dudek R, Lu Q and
Chen YH: Epithelial mesenchymal transition in embryonic
development, tissue repair and cancer: A comprehensive overview. J
Clin Med. 7(1)2017.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Atreya I and Neurath MF: Immune cells in
colorectal cancer: Prognostic relevance and therapeutic strategies.
Expert Rev Anticancer Ther. 8:561–572. 2008.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Ye L, Zhang T, Kang Z, Guo G, Sun Y, Lin
K, Huang Q, Shi X, Ni Z, Ding N, et al: Tumor-infiltrating immune
cells act as a marker for prognosis in colorectal cancer. Front
Immunol. 10(2368)2019.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Nguyen A, Loo JM, Mital R, Weinberg EM,
Man FY, Zeng Z, Paty PB, Saltz L, Janjigian YY, de Stanchina E and
Tavazoie SF: PKLR promotes colorectal cancer liver colonization
through induction of glutathione synthesis. J Clin Investig.
126:681–694. 2016.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Endale HT, Tesfaye W and Mengstie TA: ROS
induced lipid peroxidation and their role in ferroptosis. Front
Cell Dev Biol. 11(1226044)2023.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Li S, Tao K, Yun H, Yang J, Meng Y, Zhang
F and Ma X: Ferroptosis is a protective factor for the prognosis of
cancer patients: A systematic review and meta-analysis. BMC Cancer.
24(604)2024.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Liebl MC and Hofmann TG: The role of p53
signaling in colorectal cancer. Cancers (Basel).
13(2125)2021.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Xie Y, Zhu S, Song X, Sun X, Fan Y, Liu J,
Zhong M, Yuan H, Zhang L, Billiar TR, et al: The tumor suppressor
p53 limits ferroptosis by blocking DPP4 activity. Cell Rep.
20:1692–1704. 2017.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Yang L, WenTao T, ZhiYuan Z, Qi L, YuXiang
L, Peng Z, Ke L, XiaoNa J, YuZhi P, MeiLing J, et al: Cullin-9/p53
mediates HNRNPC degradation to inhibit erastin-induced ferroptosis
and is blocked by MDM2 inhibition in colorectal cancer. Oncogene.
41:3210–3221. 2022.PubMed/NCBI View Article : Google Scholar
|















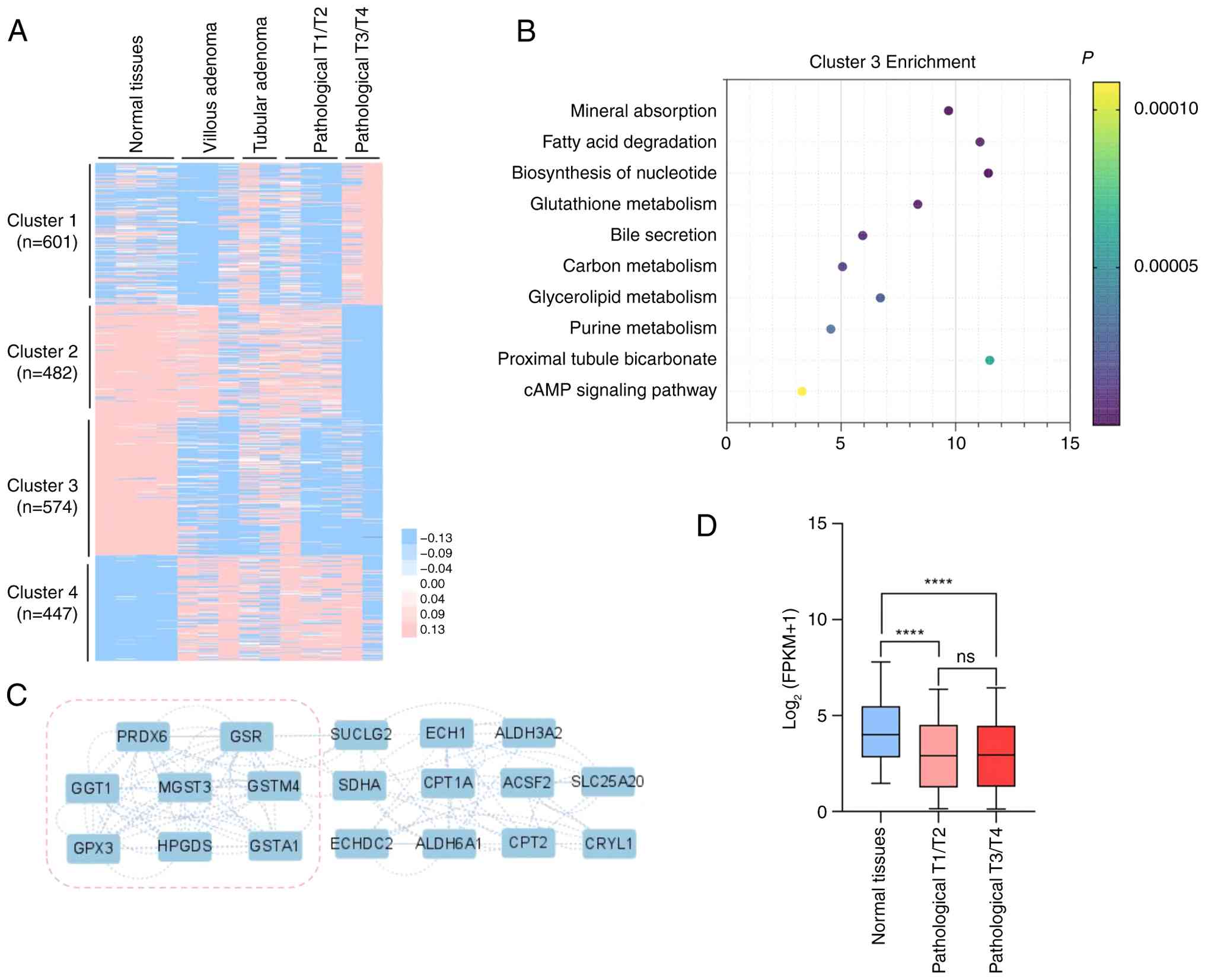
![Differences in tumor characteristics
and the immune microenvironment in glutathione gene set expression
profiles in The Cancer Genome Atlas-colon adenocarcinoma cohort.
(A) Differences in TMB between the high (n=178) and low (n=179)
glutathione gene set expression samples (two-tailed unpaired
t-test; *P<0.05). (B) Distribution of high and low
glutathione gene set expression across lymphocyte, macrophage,
dendritic cell, mast cell, neutrophil and eosinophil populations
(two-tailed unpaired t-test; ns, not significant;
**P<0.01, ****P<0.0001). (C) Difference
in the EMT scores (gene set: EMTome) between the high (n=236) and
low (n=235) glutathione gene set expression samples (two-tailed
unpaired t-test; ***P<0.001). (D) EMT scores [gene
set: Tan et al (20)]
between high (n=236) and low (n=235) glutathione gene set
expression samples (two-tailed unpaired t-test; ns, not
significant; **P<0.01, ****P<0.0001).
(E) Differences in immune score, estimate comprehensive score and
stromal score between the high (n=236) and low (n=235) glutathione
gene set expression samples. (two-tailed unpaired t-test;
*P<0.05, ***P<0.001). EMT,
epithelial-mesenchymal transition; TMB, tumor mutation burden.](/article_images/mco/24/5/mco-24-05-02938-g02.jpg)