1. Introduction
The latest statistics from the Global Burden of
Cancer Data reveal that malignant tumors of the digestive system
account for 26% of all new cancer cases yet cause 35% of
cancer-related deaths, exhibiting the prominent characteristics of
‘high incidence and even higher mortality’ (1). Among these, gastric cancer,
colorectal cancer, esophageal cancer, liver cancer, and pancreatic
cancer are the primary pathological types. Ranked by mortality,
colorectal cancer, liver cancer, and gastric cancer occupy the top
three positions, accounting for 9.3, 7.8, and 6.8% of global cancer
deaths, respectively (2). Tumor
progression is a multistage process involving the gradual
accumulation of genetic mutations and epigenetic alterations, with
the tumor microenvironment playing a decisive role. Recent advances
in mechanistic research reveal that the circulating microorganisms
(CM) are associated with alterations in the tumor microenvironment
by regulating inflammatory signaling and immune evasion pathways,
potentially serving as a critical factor in the development and
progression of gastrointestinal malignancies (3). Therefore, systematically deciphering
the dynamic characteristics and functional mechanisms of CM in
gastrointestinal malignancies not only deepens our understanding of
their pathogenesis but also provides potential targets for
developing microbiome-targeted intervention strategies. In light of
this, this review summarizes the latest research advances on the
relationship between CM and the occurrence, progression, and
outcomes of gastrointestinal tumors, aiming to provide
evidence-based guidance and directional references for subsequent
basic and clinical translational research.
In the context of this review, the term ‘circulating
microorganisms (CM)’ refers broadly to microbial signatures
detectable in blood and body fluids, including circulating
microbial DNA (cmDNA), cell-free microbial RNA, circulating
pathogen-associated molecular patterns (PAMPs), microbial
metabolites (e.g., short-chain fatty acids, bile acids), and
microbial extracellular vesicles (mEVs). It is critical to
distinguish these molecular components from viable, culturable
microbial cells. While translocation of intact bacteria into
circulation may occur under pathological conditions (e.g., severe
leaky gut), current metagenomic evidence predominantly detects
microbial nucleic acids and products rather than living,
replicating microbial communities in the bloodstream. Therefore, CM
discussed herein primarily function as liquid biopsy biomarkers and
signaling molecules rather than as active, colonizing agents.
2. Sources and composition of recirculating
microorganisms
For a long time, the academic community has
generally regarded blood as an ‘absolutely sterile’ closed system,
where the detection of any microorganisms was equated with severe
infection. However, with the rapid advancement of sensitive
technologies such as Next-Generation Sequencing (NGS), this
classical notion is gradually being rewritten. In 2001, Nikkari
et al (4) first amplified
bacterial DNA in the blood of healthy subjects. Subsequently,
Sciarra et al (5) and
others further confirmed that nucleic acids from archaea, fungi,
and viruses could also be stably detected in healthy individuals. A
series of findings reveal the existence of a circulating microbial
ecosystem within the human body that far exceeds traditional
understanding. This ecosystem not only colonizes open cavities like
the oral cavity and intestines but may also enter the bloodstream
through various pathways, forming a dynamic equilibrium and
interacting with the host. This discovery lays a solid foundation
for subsequent research into the ‘blood microbiome’.
Origin: Gut microbiota, oral cavity,
and skin-blood transport pathway
Currently, the presence of CM is largely attributed
to microbial transfer from other body sites rich in microbiota,
primarily the gastrointestinal tract, oral cavity, and skin. This
process is often triggered by compromised structural integrity or
functional imbalance of epithelial barriers: for instance, in
periodontitis, widened gingival crevicular spaces allow oral
microbiota to directly enter microvessels; whereas ‘leaky gut’
occurs due to downregulation of tight junction proteins and
thinning of the mucus layer, enabling trans-epithelial migration of
intestinal commensals and their metabolites (6). Once in the bloodstream, bacterial
components, including cell wall fragments or cell wall components
(lipopolysaccharides, peptidoglycans, flagellar proteins) along
with secondary metabolites (short-chain fatty acids and bile acids)
circulate systemically, creating a low-grade systemic inflammatory
response that provides sustained signaling for microenvironmental
remodeling in distant organs (7).
Composition: Reflecting but distinct
from gut microbiota
While CM composition mirrors the gut microbiota to
some extent, significant alterations occur during translocation.
The total human gut microbiota population can reach 4x10¹³ CFU,
with 97% colonizing the colon. Archaea and eukaryotes (including
fungi) constitute only 0.1-1% (8).
However, CM represent only a highly selected fraction of these
organisms that successfully translocate and are detectable in
blood. Notably, the blood microbiome is dominated by the
Proteobacteria phylum, often reaching relative abundances of
85-90%, exhibiting marked enrichment compared to their relative
abundance in the gut, while Firmicutes, and
Bacteroidetes are underrepresented, suggesting selective
pressure during translocation (6).
However, it is critical to acknowledge that Proteobacterial
dominance is also the hallmark signature of reagent contamination
(‘kitome’) in low-biomass samples, particularly from
Pseudomonas and Acinetobacter species commonly found
in DNA extraction kits and laboratory water. Therefore, whether
this dominance reflects true biological translocation or technical
artifact remains an active subject of debate and rigorous
contamination control. Goraya et al (9) further noted that the composition and
abundance of the CM exhibit significant heterogeneity across
different diseases and across stages of the same disease,
reflecting not merely gut dysbiosis but also disease-specific
selective pressures in the bloodstream. This provides a new
perspective for elucidating the causal role of the CM in disease
progression.
3. Circulating microbial detection
Compared to the vast genetic background of the host,
the biomass of commensal microorganisms is extremely low, making
their precise capture and qualitative/quantitative analysis highly
challenging. With the advancement of highly sensitive genetic
technologies, methods such as 16S rRNA sequencing, metagenomic
sequencing, and intergenic spacer analysis (IS-pro) have become key
analytical tools for studying prokaryotes, viruses, fungi, and
other microorganisms (Table
I).
 | Table IComparison of methods for detecting
CMS. |
Table I
Comparison of methods for detecting
CMS.
| | Testing method |
|---|
| Characteristic | 16S rRNA
sequencing | Metagenomic
sequencing | IS-pro
technology |
|---|
| Resolution | Genus/Species
level | Strain level | Species level |
|
Primer-dependent | Yes (conservative
zone + variable zone) | No | Yes
(phylum-specific primers) |
| Testing scope | Bacteria and
archaea | Bacteria, archaea,
fungi and viruses | Bacteria |
| Cost | Low | High | Moderate |
| Turnaround
time | 1-2 days | 4-7 days | 3-6 h |
| Contamination
risk | High (PCR
amplification preference) | Moderate (host DNA
interference) | Moderate |
| Primary
contamination categories | Pseudomonas,
Acinetobacter, Bacillus. |
Cutibacterium, Pseudomonas
(kit residue); host DNA (>90% of reads) |
Staphylococcus,
Streptococcus (skin microbiota) |
| Sources of
contamination | DNA extraction kit
(kitome), PCR reagents, laboratory environment, index hopping | DNA extraction kit,
host DNA contamination, environmental microorganisms, batch
effects | PCR reagents,
laboratory environment, operator skin microbiota |
| Recommended
controls | Negative extraction
control, PCR negative control, mock community, batch control | Negative extraction
control, positive control, batch control, internal reference
control | Negative control,
standardized strain control, operator control |
16S rRNA sequencing is a widely adopted method for
analyzing bacteria and archaea (10). This molecular marker combines
conserved regions (reflecting species affinity) with highly
variable regions (defining species differences), enabling rapid and
cost-effective characterization of circulating microbial DNA
composition in low-biomass samples like tissues and blood. De
Oliveira et al (11)
utilized high-throughput sequencing of the V3-V4 region to identify
an enrichment of colibactin-producing Escherichia coli in
right-sided colorectal cancer lesions. The study hypothesizes that
such bacteria may suppress antigen presentation by reshaping the
local glycerophospholipid microenvironment, thereby reducing tumor
immunogenicity. However, this conclusion remains a preliminary
hypothesis: On one hand, colibactin production is believed to be
confined to the tumor microenvironment, lacking direct functional
evidence establishing a causal link to immune evasion in
vivo (or its presence in the circulating microbial fraction).
On the other hand, 16S rRNA sequencing itself is constrained by
primer specificity, making it difficult to distinguish closely
related species and prone to PCR amplification bias that may
overestimate diversity. Nevertheless, this study provides important
clues for exploring the potential regulatory mechanisms of CM
components on the host microenvironment, while also highlighting
the ongoing technical challenges associated with current CM
detection methods based on 16S rRNA sequencing (12).
Metagenomic sequencing (also known as shotgun
sequencing) involves off-target sequencing of all genomes, enabling
simultaneous analysis of both the species composition and
functional gene profiles of the CM (13). Chen et al (14) combined fecal metagenomics with
metabolomics, revealing that Fusobacterium nucleatum and
Bifidobacterium longum accelerate or inhibit colorectal
cancer progression via pro-inflammatory metabolites and short-chain
fatty acid pathways, respectively, visually demonstrating the
causal chain of the ‘microbe-metabolite-tumor’ axis. Compared to
16S rRNA sequencing, metagenomics is not limited by primers and can
capture non-bacterial members such as fungi, viruses, and
mycoplasmas. It achieves strain-level resolution, yielding more
accurate results. However, this approach requires detecting all
gene sequences (including those from normal host cells and tumor
cells), making it relatively time-consuming, complex, and costly
(15).
IS-pro technology targets the 16S-23S rDNA
intergenic spacer region across bacterial species. This fragment
exhibits high variability between species while maintaining high
intra-species conservation, serving as a bacterial ‘fingerprint’
(16). Research indicates that
IS-pro enables rapid species-level identification and relative
quantification in a single-tube reaction through gate-specific PCR
primers coupled with fluorescent labeling. Furthermore, compared to
16S rRNA sequencing, IS-pro accelerates analysis, reduces costs,
and maintains equivalent analytical performance, offering a
scalable, lightweight solution for rapid CM research (17).
Although the emergence of novel detection
technologies has facilitated the detection of circulating microbial
signatures (CMS), CMS analysis still faces significant technical
challenges due to the extremely low microbial DNA biomass in blood.
This low biomass characteristic makes CMS research highly
susceptible to multiple sources of contamination, including DNA
extraction kits (‘kit sets’), PCR reagents, laboratory
environments, and sequencing tag jumping (18). Studies indicate that contaminant
DNA is prevalent in commonly used DNA extraction kits, with
significant variation between batches, potentially obscuring
genuine biological signals (19).
To ensure reliable interpretation of CMS data,
multiple stringent controls must be implemented: i) Negative
extraction controls and PCR negative controls; ii) batch
randomization and calibration; iii) computational decontamination
(removing contaminants like Pseudomonas and Bacillus
detected in negative controls); iv) quantitative PCR measurement of
bacterial DNA load to distinguish true signals from background
contamination (18).
These limitations significantly impact the
interpretation of CMS studies. Research lacking adequate controls
or decontamination steps may misidentify kit contaminants as
disease biomarkers. Furthermore, variations in blood collection
tube types, storage conditions, and extraction protocols introduce
batch effects that interfere with cross-study comparisons.
Therefore, caution is warranted when interpreting CMS findings, and
standardized contamination control protocols are urgently needed in
this field.
4. The link between circulating
microorganisms and gastrointestinal tumors
In recent years, as the aforementioned detection
technologies have transformed CM into quantifiable liquid biopsy
markers, it has been confirmed that CM may be associated with
synergistically regulating host immunity, inflammatory signaling,
and tumor microenvironment remodeling through bacterial structures,
nucleic acid fragments, and metabolic small molecules. The
following sections will systematically examine the pivotal role of
CM in gastrointestinal tumorigenesis, progression, and outcomes
from two dimensions: ‘Diversity disparities’ and ‘oncogenic
mechanisms’. This analysis aims to provide novel strategic
footholds for early diagnosis, precision therapy, and prognostic
assessment.
Diversity differences: From gut
dysbiosis to circulating signals
The composition and diversity of the gut microbiota
in gastrointestinal cancer patients undergo significant remodeling,
contrasting sharply with healthy individuals. Colorectal cancer
(CRC) is a malignant tumor originating in the colon or rectum,
classified as colon cancer or rectal cancer based on its site of
origin (20). Compared to control
patients, colorectal cancer patients frequently exhibit increased
levels of pro-inflammatory bacterial species, including
Clostridium difficile and Bacteroides fragilis;
conversely, samples from control patients typically show greater
abundance of beneficial bacteria such as Bifidobacterium and
Bacteroides species (21).
CM as liquid biopsy markers
Critically, these gut dysbiotic signatures translate
into detectable CM alterations. Giacconi et al (19) conducted a case-control study
analyzing plasma samples from 50 CRC patients and 40 healthy
controls, demonstrating significantly elevated bacterial DNA loads
in the circulation of CRC patients compared to controls
(P<0.001). Their analysis revealed distinct microbial
signatures, including increased abundance of oral-originating taxa
such as Cutibacterium and Sphingomonas, that could
discriminate early-stage CRC from healthy individuals with high
accuracy (AUC=0.92). These findings suggest that translocated
microbial DNA in circulation reflects underlying gut dysbiosis and
tumor-associated microbial shifts, supporting the theoretical
potential of CM as liquid biopsy biomarkers for CRC detection,
pending standardized validation.
Gastric cancer exemplifies the interaction between
CM dysbiosis and host epithelial cells. Helicobacter pylori
(Hp) is the most extensively documented risk factor for gastric
cancer, accounting for approximately 65-80% of all cases. It may be
associated with driving patients' progression from atrophic
gastritis to intestinal metaplasia (22). It can also enter the bloodstream
through mucosal lesions to form CM, whose DNA and antigenic
components can be detected in the patient's peripheral blood.
Beyond Hp, the gut microbiota of gastric cancer patients exhibits
enrichment of pro-inflammatory bacteria such as
Proteobacteria and Spirochaetes, alongside reduced
levels of beneficial bacteria like Firmicutes (butyrate
producers) and Bacteroidetes (bile acid degraders) (23). Hu et al (24) demonstrated that bovine anti-H.
pylori antibodies effectively clear H. pylori infections
in individuals with blood type O. They also reported that other
spirochetes (e.g., Helicobacter heilmannii) can colonize the
human stomach alongside H. pylori. This dysbiosis is
associated with intestinal barrier disruption, inflammatory
responses, and tumor-promoting microenvironmental changes.
Microbiome dysbiosis-leaky
gut-inflammation axis: Pathogenic mechanisms of CM disruption
Dysbiosis may compromise the intestinal barrier
through mechanisms such as downregulating tight junction proteins,
disrupting the mucus layer, and causing immune dysregulation,
potentially facilitating bacterial translocation into the
bloodstream. Notably, barrier disruption permits the translocation
of diverse, as-yet-uncharacterized microbial species and their
products; consequently, while barrier dysfunction and inflammation
are plausible contributors to colorectal carcinogenesis, ascribing
tumor-promoting effects to specific taxa or discrete mechanisms
remains challenging at this stage. Zhang et al's circadian
rhythm disruption (CRD) mouse model demonstrated that 21 days of
continuous light exposure increased intestinal permeability by
114.7%, significantly elevated plasma FITC-dextran levels, and
induced a reduction in Prevotellaceae alongside an increase
in Bacteroidaceae, forming a vicious cycle of
‘dysbiosis-leaky gut-circulating microbial dysregulation’ (25). Lipopolysaccharide (LPS) derived
from intestinal Gram-negative bacteria enters the liver via the
portal vein and is normally cleared by Kupffer cells. When
clearance efficiency declines, LPS translocate into the
bloodstream, triggering metabolic endotoxemia. This activates the
TLR4-NF-κB pathway, promoting the release of pro-inflammatory
factors such as IL-1β, IL-6, and TNF-α, leading to low-grade
chronic inflammation and significantly increasing disease risk
(26).
5. Mechanistic studies of CM and
gastrointestinal tumors
The mechanisms by which circulating microorganisms
contribute to gastrointestinal tumorigenesis involve complex
interactions between microbial components and host signaling
pathways (Fig. 1). Proposed
mechanisms implicating CM in chronic inflammation and immune
evasion lies in gut microbiota dysbiosis but manifests systemically
through translocated components. PAMPs, namely LPS and flagellar
proteins from gram-negative gut bacteria, first breach the
intestinal barrier via paracellular or transcellular routes,
entering the bloodstream as CM components. Once in circulation,
these PAMPs bind with high affinity to Toll-like receptor 4 (TLR4)
on distant immune cells and endothelial cells, activating systemic
rather than localized inflammatory responses (27).
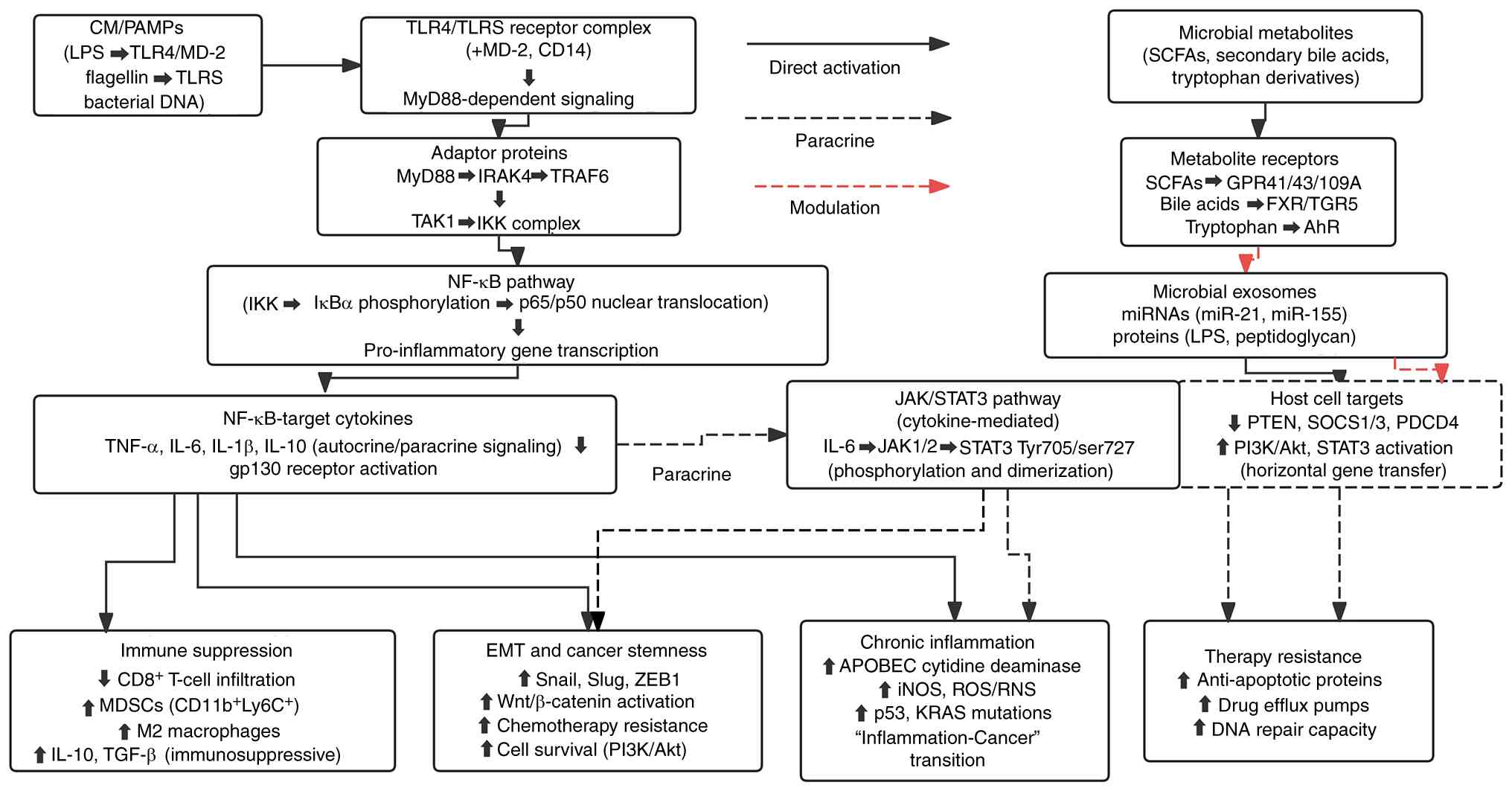 | Figure 1Mechanistic pathways of CMS in
gastrointestinal tumorigenesis. CMS, circulating microbial
signatures. CMS, circulating microbial signatures; CM, circulating
microorganisms; PAMPs, pathogen-associated molecular patterns; LPS,
lipopolysaccharide; TLR, Toll-like receptor; IRAK4, IL-1
receptor-associated kinase 4; TRAF6, TNF receptor-associated factor
6; TAK1, TGF-β-activated kinase 1; IKK, IκB kinase; STAT3, signal
transducer and activator of transcription 3; APOBEC, apolipoprotein
B mRNA-editing enzyme catalytic; iNOS, inducible nitric oxide
synthase; ROS, reactive oxygen species; RNS, reactive nitrogen
species; MDSCs, myeloid-derived suppressor cells; EMT,
epithelial-mesenchymal transition; ZEB1, zinc finger E-box binding
homeobox 1; SCFAs, short-chain fatty acids; FXR, farnesoid X
receptor; TGR5, Takeda G protein-coupled receptor 5; AhR, aryl
hydrocarbon receptor; miR, microRNA; PDCD4, programmed cell death
4. |
Upon binding to TLR4 on intestinal epithelial or
immune cells, LPS rapidly activates dual signaling pathways, NF-κB
and MAPK, via the MyD88 adaptor protein, continuously amplifying
proinflammatory signals such as IL-6, IL-1β, and TNF-α. This
systemic inflammation, driven by circulating microbial components
(such as PAMPs and metabolites) rather than localized gut bacteria,
leads to recruiting CD11b+ Ly6C+ myeloid-derived suppressor cells
myeloid-derived suppressor cells, inducing macrophage polarization
toward the M2 phenotype, and secreting IL-10 and TGF-β. This may
directly inhibit CD8+ T cells infiltration, potentially weakening
the first line of immune defense (28). Furthermore, the chronic
inflammatory environment upregulates apolipoprotein B mRNA editing
enzyme catalyzing polypeptide (APOBEC) cytidine deaminase and
nitric oxide synthase, driving the accumulation of mutations in key
genes like p53. This may promote the ‘inflammation-to-cancer’
transition and early immune evasion (27).
Moreover, PAMPs can trigger signal transducer and
activator of transcription 3 (STAT3) serine phosphorylation via the
TLR4/TLR5-JAK2 axis, driving epithelial-mesenchymal transition,
maintenance of cellular stemness, and the formation of
drug-resistant phenotypes. Balic et al (29) confirmed that this non-canonical
STAT3 activation is a core node in TLR4-mediated glycolytic
reprogramming and macrophage polarization. Extensive crosstalk
exists between the Wnt and STAT3 pathways; microbe-mediated STAT3
activation indirectly enhances Wnt signaling, further boosting
chemotherapy tolerance (30).
Concurrently, microbial metabolites or LPS-induced inflammatory
factors like TNF-α inhibit apoptosis by activating the PI3K/Akt
pathway, thereby enhancing tumor cell survival and resistance to
chemotherapeutic agents (31).
Beyond the bacteria themselves, secondary bile
acids, short-chain fatty acids, and microbial exosomes in CM serve
as critical chemical messengers in the ‘inflammation-to-cancer’
evolution. Gao et al (32)
demonstrated that high concentrations of deoxycholic acid (DCA),
generated from hepatic primary bile acids via 7α-dehydroxylation,
act as DNA-breaking agents and epigenetic disruptors. DCA inhibits
DNA repair enzymes in intestinal epithelial cells, increases
alkylating agent-induced mutation rates, and amplifies NF-κB
inflammatory signaling, significantly elevating gastrointestinal
tumor risk. Meanwhile, reduced butyrate-producing bacteria in
vivo cause insufficient β-oxidation energy supply in intestinal
epithelium, while abnormal increases in propionic acid and
isovaleric acid enhance tumor cell migration and metastasis via the
GPR43-PI3K axis (33).
Furthermore, circulating microbial components, particularly PAMPs,
induce host cells to release exosomes rich in specific miRNAs and
proteins. Ponton-Almodovar et al (34) propose that these exosomes, once
internalized by tumor cells, sustain activation of PI3K/Akt and
STAT3 pathways by downregulating tumor suppressors like PTEN and
SOCS6, thereby synergistically driving metastasis and chemotherapy
resistance.
In summary, CM have been implicated in contributing
to a self-reinforcing oncogenic network through
TLR-inflammation-mutation-pathway activation-metabolic
reprogramming-exosome remote regulation, potentially spanning the
initiation, progression, and resistance of gastrointestinal
tumors.
It is important to note that the mechanistic
pathways discussed above, including TLR-NF-κB, STAT3, Wnt, and
PI3K/Akt signaling, have been primarily established through studies
of gut-resident or intratumoral microbiota rather than circulating
microbial components per se. The extrapolation of these mechanisms
to CM is based on the shared molecular structures (e.g., LPS,
flagellin) between tissue-resident and circulating microbes.
However, whether CM activate identical signaling nodes with
comparable kinetics and potency as their tissue-bound counterparts
requires direct experimental validation in future studies.
6. Circulating microbes shape epigenetic
memory: The long-term legacy of inflammation
Prolonged exposure to circulating microbial
components can induce epigenetic reprogramming, driving persistent
alterations in gene expression. This process encompasses abnormal
changes in DNA methylation patterns, chromatin structural
remodeling, and sustained activation of inflammatory transcription
programs. The resulting epigenetic imprints persist long after the
initial stimulus has completely subsided, driving fundamental
shifts in cellular function.
At the DNA methylation level, chronic activation of
the TLR4 and NF-κB signaling pathways can recruit DNA
methyltransferases to specific genomic regions, leading to
hypermethylation of tumor suppressor gene promoters (8). For instance, prolonged exposure to
LPS in the human microenvironment induces significantly elevated
methylation levels in promoter regions such as p16, thereby causing
gene silencing and impairing DNA damage repair capacity (35).
Histone modifications also represent a core
component in inflammation-mediated epigenetic alterations. Key
transcription factors like NF-κB and signal transducer and
activator of STAT3 recruit histone acetyltransferases to gene
enhancer regions, maintaining sustained activity of
proliferation-related genes and establishing a stable pro-tumor
transcriptional environment. Concurrently, this ‘open’ chromatin
conformation possesses epigenetic memory properties, allowing the
state to persist long-term even after pathogenic microorganisms are
completely cleared, without requiring continuous external
stimulation (36).
Microbial metabolites in the circulation can also
participate in gene expression reprogramming by directly regulating
the activity of epigenetic enzymes. As a natural inhibitor of
histone deacetylases (HDACs), reduced circulating levels of
butyrate, commonly observed in patients with advanced
gastrointestinal tumors, lead to dysregulated HDAC hyperactivation.
This triggers abnormal chromatin condensation and silences key DNA
repair genes (37).
These multi-layered epigenetic alterations provide a
molecular explanation for a key clinical observation: even after
successful early eradication of pathogenic bacteria (such as
Helicobacter pylori), the host's cancer risk does not
immediately decline (38). Because
inflammation-induced epigenetic ‘memory’ is firmly established in
target tissue cells, this intrinsic driver of malignant
transformation persists and operates autonomously. Even after the
original circulating microbial components have completely
disappeared, the organism inevitably continues progressing along
the inflammation-to-cancer conversion trajectory.
7. Clinical translation of CMS and
validation requirements
Currently, multiple studies have explored the
potential of CMS as non-invasive liquid biopsies for
gastrointestinal tumors. In the field of CRC, circulating bacterial
DNA levels in CRC patients are significantly higher than in healthy
controls and positively correlate with tumor burden (19). Microbiome-based cfDNA classifiers
achieve 90% accuracy in distinguishing primary hepatocellular
carcinoma from metastatic CRC (39). In gastric cancer, detection of
Helicobacter pylori DNA and antigen components in peripheral
blood provides direct evidence of bacterial translocation triggered
by mucosal lesions, effectively supplementing diagnostic
information (6). Furthermore, the
association between CMS and postoperative recurrence risk in lung
cancer suggests broad prospects for this approach across cancer
types (7).
However, its clinical translation still faces
significant challenges. Existing studies are predominantly
single-center designs with limited sample sizes and lack
prospective validation in independent cohorts. Additionally, the
absence of universally accepted negative control standards and
protocols for calculating decontamination processes complicates
cross-study comparisons. More importantly, the precise origin of
circulating microbial DNA (intestinal, oral, or other sites) and
its functional relationship with the live microbial community
remain to be elucidated. Future large-scale, multicenter
prospective studies are urgently needed. These should establish the
clinical utility of circulating microbiome as a liquid biopsy tool
through rigorous contamination control, standardized methodologies,
and integration with established clinical biomarkers.
8. Application of circulating microorganisms
in gastrointestinal tumor therapy
Currently, surgery combined with chemoradiotherapy
remains the standard treatment regimen for gastrointestinal tumors.
Recent studies on circulating microorganisms have revealed its dual
‘oncogenic-suppressive’ role in tumor initiation and progression.
By interacting with host immunity and metabolism, CM can either
enhance or diminish treatment response, positioning it as a
potential key target for precision interventions. Furthermore,
emerging strategies such as engineered strain immunotherapy,
probiotic interventions, and fecal microbiota transplantation
further highlight the role of CM in exploring novel cancer
treatments.
CM and chemotherapy
Chemotherapy remains one of the core treatment
modalities for advanced gastrointestinal tumors, yet the emergence
of chemotherapy resistance significantly shortens patient survival
(40). Recent evidence indicates
that microbial enzymes detected in circulation (likely originating
from translocated bacteria or circulating microbial fragments)
alter the activity of chemotherapeutic agents, positioning them as
both ‘accomplices’ and ‘assistants’ in tumor treatment (41). On one hand, enzymes associated with
certain CMs inactivate drugs by altering their structures; for
example, Vibrio gamma-deoxythymidine deaminase inactivates
gemcitabine, inducing resistance in pancreatic ductal
adenocarcinoma, an effect reversible by ciprofloxacin (42). Similarly, bacterial β-glucuronidase
converts the inert metabolite of irinotecan into its active form,
triggering dose-limiting diarrhea (43).
Conversely, multi-omics experiments by the Iida team
confirmed that probiotics like Lactobacillus acidophilus
activate tumor-associated macrophages and dendritic cells via the
TLR-MyD88-NF-κB axis. This induces sustained release of high
concentrations of reactive oxygen species (ROS), enhancing DNA
damage in tumor cells while blocking repair and transcription
pathways, thereby boosting the efficacy of the anticancer drug
oxaliplatin (44). The dual nature
of CM, both antagonizing and enhancing chemotherapy effects,
underscores the necessity for further investigation. Only by
thoroughly understanding the specific drug-microbiome interactions
can precise microbial intervention strategies be designed to
optimize therapeutic outcomes.
CM and radiation therapy
Radiotherapy relies on ionizing radiation to exert
cytotoxic effects on tumor cells, serving as another treatment
modality for mid-to-late stage tumors. Recent studies indicate that
commensal microorganisms such as CM significantly influence
radiotherapy efficacy and prognosis (45). Selective elimination of
Gram-positive bacteria via oral vancomycin enhances antitumor
activity during radiotherapy by facilitating cross-presentation of
tumor-associated antigens to CD8+ T cells and promoting IFN-γ
secretion (46). Conversely,
broad-spectrum antibiotic mixtures, while eliminating symbiotic
bacteria, promote the expansion of symbiotic fungi, thereby
diminishing radiosensitivity (47). In the same year, Dong et al
(45) demonstrated significantly
improved radiotherapy outcomes after using metronidazole to
specifically eliminate Clostridium difficile. Furthermore,
researchers discovered that tumor lactic acid bacteria
producing L-lactic acid can induce radiation resistance in tumor
cells through metabolic remodeling (48). Collectively, these findings suggest
that optimizing disease-specific and personalized probiotic
combinations and applications will be an essential future research
direction, building upon improvements in radiotherapy efficacy and
reduction of radiation-induced damage.
CM and novel engineering
therapies
In recent years, fecal microbiota transplantation
(FMT) has emerged as a novel approach for reshaping the gut
microbiome, demonstrating significant advantages in treating
gastrointestinal diseases. FMT involves transferring the fecal
microbiota from a healthy donor into a patient's gut to restore
microbial balance, with demonstrated efficacy in treating
conditions such as recurrent Clostridioides difficile
infection and ulcerative colitis (49,50).
In oncology, FMT can recruit antigen-presenting cells like
dendritic cells into the tumor microenvironment, reshape innate and
adaptive immunity, and exert antitumor effects through microbial
metabolism (51). Davar et
al (52) demonstrated that FMT
from responder donors reversed immune therapy efficacy in some of
15 advanced melanoma patients resistant to anti-PD-1 therapy,
confirming FMT's potential for cancer prevention and treatment.
However, the mechanisms of action, ethical guidelines, and optimal
protocols for FMT require further clarification through
high-quality randomized controlled trials.
Probiotics and prebiotics represent another hotspot
in microbiome interventions. Probiotics are live beneficial
microorganisms that activate macrophages and induce antitumor
factors such as TNF-α, IFN-γ, IL-12, and NO. Prebiotics,
conversely, are substrates utilized by these beneficial bacteria,
selectively promoting the proliferation of beneficial commensal
flora (53). Zheng et al
(54) demonstrated in a rat
gastric cancer postoperative model that composite probiotics reduce
inflammation, downregulate intestinal permeability signals, and
restore microbial community structure. However, clinical
translation of probiotics faces challenges including short shelf
life and quality variability due to production and storage
differences. Standardized protocols and long-term follow-up data
are urgently needed to support their integration into frontline
cancer treatment.
Overall, FMT and probiotic interventions restore
barrier function by reshaping the gut microbiome, reducing
bacterial translocation and the entry of pro-tumor factors into the
bloodstream. Simultaneously, they competitively inhibit pathogenic
bacteria, block carcinogenic components from entering the
bloodstream, promote the systemic release of beneficial
metabolites, stabilize the circulating microbiome, indirectly
regulate cell membrane composition and function, and drive the body
toward restoring a healthy steady state.
9. Conclusion
In recent years, with the iterative development of
high-sensitivity genetic technologies, CM has evolved from a
‘detection noise’ to a core liquid biomarker potentially associated
with driving the occurrence, progression, and efficacy regulation
of gastrointestinal tumors. This paper examines CM research
progress in gastrointestinal tumors across four dimensions,
revealing a ‘gut dysbiosis-CM translocation-systemic tumor
microenvironment remodeling’ axis that offers novel perspectives
for non-invasive diagnosis.
However, as reviewed herein, current evidence
remains predominantly preclinical or derived from small-scale
retrospective cohorts. Future research should focus on the
following pathways: First, integrate individual patient data on
metagenomics, metabolomics, and immune phenotypes; second, employ
machine learning algorithms to eliminate contamination signals from
kits and laboratories, thereby identifying microbial biomarkers
with diagnostic value; finally, validate through multicenter
prospective cohort studies whether these biomarkers exhibit a
direct causal relationship with tumor development, rather than
merely being associated phenomena. Should standardized protocols
and rigorous clinical validation be achieved, real-time monitoring
of CM dynamics could eventually contribute to the development of
‘liquid microbiome precision medicine’. Currently, however,
CM-based diagnostics remain in the proof-of-concept phase,
requiring substantial methodological refinement before clinical
implementation.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
Not applicable.
Authors' contributions
YY and YD conceptualized the study. YY and YR
conducted the investigation. YY wrote the original draft. TM, SJ,
YR, JA and YD reviewed and edited the manuscript. SJ and JA
prepared the figures. YD supervised the study. Data authentication
is not applicable. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Rauth S, Malafa M, Ponnusamy MP and Batra
SK: Emerging trends in gastrointestinal cancer targeted therapies:
Harnessing tumor microenvironment, immune factors, and metabolomics
insights. Gastroenterology. 167:867–884. 2024.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Asawa S, Nüesch M, Gvozdenovic A and Aceto
N: Circulating tumour cells in gastrointestinal cancers: Food for
thought? Br J Cancer. 128:1981–1990. 2023.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Nikkari S, McLaughlin IJ, Bi W, Dodge DE
and Relman DA: Does blood of healthy subjects contain bacterial
ribosomal DNA? J Clin Microbiol. 39:1956–1959. 2001.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Sciarra F, Franceschini E, Campolo F and
Venneri MA: The diagnostic potential of the human blood microbiome:
Are we dreaming or awake? Int J Mol Sci. 24(10422)2023.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Lee YY, An J, Han J, Moon Y and Lee SI:
Plasma-based digital PCR assay for early detection of gastric
cancer using multiple methylation biomarkers. Sci Rep.
16(1727)2025.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Law HKW and Yim HCH: Early diagnosis of
cancer using circulating microbial DNA. Cell Rep Med.
5(101502)2024.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Zhang C, Geng H, Tan Y and Wang L:
Multidimensional regulation of the microbe-TLR4 signaling axis in
colorectal cancer: From molecular mechanisms to microbe-targeted
therapies. Biochim Biophys Acta Rev Cancer.
1880(189397)2025.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Goraya MU, Li R, Mannan A, Gu L, Deng H
and Wang G: Human circulating bacteria and dysbiosis in
non-infectious diseases. Front Cell Infect Microbiol.
12(932702)2022.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Dass M, Singh Y and Ghai M: A review on
microbial species for forensic body fluid identification in healthy
and diseased humans. Curr Microbiol. 80(299)2023.PubMed/NCBI View Article : Google Scholar
|
|
11
|
De Oliveira Alves N, Dalmasso G, Nikitina
D, Vaysse A, Ruez R, Ledoux L, Pedron T, Bergsten E, Boulard O,
Autier L, et al: The colibactin-producing Escherichia coli
alters the tumor microenvironment to immunosuppressive lipid
overload facilitating colorectal cancer progression and
chemoresistance. Gut Microbes. 16(2320291)2024.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Yu J, Zhang L, Gao D, Wang J, Li Y and Sun
N: Comparison of metagenomic next-generation sequencing and blood
culture for diagnosis of bloodstream infections. Front Cell Infect
Microbiol. 14(1338861)2024.PubMed/NCBI View Article : Google Scholar
|
|
13
|
El Tekle G and Garrett WS: Bacteria in
cancer initiation, promotion and progression. Nat Rev Cancer.
23:600–618. 2023.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Chen F, Dai X, Zhou CC, Li KX, Zhang YJ,
Lou XY, Zhu YM, Sun YL, Peng BX and Cui W: Integrated analysis of
the faecal metagenome and serum metabolome reveals the role of gut
microbiome-associated metabolites in the detection of colorectal
cancer and adenoma. Gut. 71:1315–1325. 2022.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Zhou X, Kandalai S, Hossain F and Zheng Q:
Tumor microbiome metabolism: A game changer in cancer development
and therapy. Front Oncol. 12(933407)2022.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Budding AE, Grasman ME, Lin F, Bogaards
JA, Soeltan-Kaersenhout DJ, Vandenbroucke-Grauls CM, van Bodegraven
AA and Savelkoul PH: IS-pro: High-throughput molecular
fingerprinting of the intestinal microbiota. FASEB J. 24:4556–4564.
2010.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Singer M, Koedooder R, Bos MP, Poort L,
Schoenmakers S, Savelkoul PHM, Laven JSE, de Jonge JD, Morré SA and
Budding AE: The profiling of microbiota in vaginal swab samples
using 16S rRNA gene sequencing and IS-pro analysis. BMC Microbiol.
21(100)2021.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Fierer N, Leung PM, Lappan R, Eisenhofer
R, Ricci F, Holland SI, Dragone N, Blackall LL, Dong X, Dorador C,
et al: Guidelines for preventing and reporting contamination in
low-biomass microbiome studies. Nat Microbiol. 10:1570–1580.
2025.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Giacconi R, Donghia R, Arborea G, Savino
MT, Provinciali M, Lattanzio F, Caponio GR, Coletta S, Bianco A,
Notarnicola M, et al: Plasma bacterial DNA load as a potential
biomarker for the early detection of colorectal cancer: A
case-control study. Microorganisms. 11(2360)2023.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Siegel RL, Wagle NS, Cercek A, Smith RA
and Jemal A: Colorectal cancer statistics, 2023. Ca Cancer J Clin.
73:233–254. 2023.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Zhan ZS, Zheng ZS, Shi J, Chen J, Wu SY
and Zhang SY: Unraveling colorectal cancer prevention: The vitamin
D-gut flora-immune system nexus. World J Gastrointest Oncol.
16:2394–2403. 2024.PubMed/NCBI View Article : Google Scholar
|
|
22
|
López MJ, Carbajal J, Alfaro AL, Saravia
LG, Zanabria D, Araujo JM, Quispe L, Zevallos A, Buleje JL, Cho CE,
et al: Characteristics of gastric cancer around the world. Crit Rev
Oncol Hematol. 181(103841)2023.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Fakharian F, Asgari B, Nabavi-Rad A,
Sadeghi A, Soleimani N, Yadegar A and Zali MR: The interplay
between Helicobacter pylori and the gut microbiota: An
emerging driver influencing the immune system homeostasis and
gastric carcinogenesis. Front Cell Infect Microbiol.
12(953718)2022.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Hu D, Zhang F, Zhou J, Xu B, Zhang H,
Qiang H, Ren S, Shan B, Yin C, Zhang Z, et al: The clearance effect
of bovine anti-Helicobacter pylori antibody-containing milk
in O blood group Helicobacter pylori-infected patients: A
randomized double-blind clinical trial. J Transl Med.
13(205)2015.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Zhang TW, Song JC, Hao NB, Qu MY, Guo BS
and Li CZ: Continuous light-induced circadian rhythm disruption
impairs intestinal barrier integrity in male C57BL/6 mice through
gut microbiota dysbiosis and the apoptosis-inflammation-oxidative
stress cascade. bioRxiv: doi: https://doi.org/10.1101/2025.11.11.687914.
|
|
26
|
Lu J, Zhang W, He Y, Jiang M, Liu Z, Zhang
J, Zheng L, Zhou B, Luo J, He C, et al: Multi-omics decodes
host-specific and environmental microbiome interactions in sepsis.
Front Microbiol. 16(1618177)2025.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Fu Y, Kim H, Lee DS, Han A, Heine H,
Zamyatina A and Kim HM: Structural insight into TLR4/MD-2
activation by synthetic LPS mimetics with distinct binding modes.
Nat Commun. 16(4164)2025.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Li J, Qin Y, Chen Y, Zhao P, Liu X, Dong
H, Zheng W, Feng S, Mao X and Li C: Mechanisms of the
lipopolysaccharide-induced inflammatory response in alveolar
epithelial cell/macrophage co-culture. Exp Ther Med.
20(76)2020.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Balic JJ, Albargy H, Luu K, Kirby FJ,
Jayasekara WSN, Mansell F, Garama DJ, De Nardo D, Baschuk N, Louis
C, et al: STAT3 serine phosphorylation is required for TLR4
metabolic reprogramming and IL-1β expression. Nat Commun.
11(3816)2020.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Parsons MJ, Tammela T and Dow LE: WNT as a
driver and dependency in cancer. Cancer Discov. 11:2413–2429.
2021.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Liu L, Yan M, Yang R, Qin X, Chen L, Li L,
Si J, Li X and Ma K: Adiponectin attenuates
lipopolysaccharide-induced apoptosis by regulating the
Cx43/PI3K/AKT pathway. Front Pharmacol. 12(644225)2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Gao Y, Lin J, Ye C, Guo S and Jiang C:
Microbial transformations of bile acids and their receptors in the
regulation of metabolic dysfunction-associated steatotic liver
disease. Liver Res. 7:165–176. 2023.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Kong L, Hoshi N, Sui Y, Yamada Y, Yoshida
R, Ooi M, Tian Z, Kimura I and Kodama Y: GPR43 suppresses
intestinal tumor growth by modification of the mammalian target of
rapamycin complex 1 activity in ApcMin/+ mice. Med Princ Pract.
31:39–46. 2022.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Ponton-Almodovar A, Sanderson S, Rattan R,
Bernard JJ and Horibata S: Ovarian tumor microenvironment
contributes to tumor progression and chemoresistance. Cancer Drug
Resist. 7(53)2024.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Chen S, Tan Y, Xiao X, Xiao XC, Li Q, Wu
Q, Peng YY, Ren J and Dong ML: Deletion of TLR4 attenuates
lipopolysaccharide-induced acute liver injury by inhibiting
inflammation and apoptosis. Acta Pharmacol Sin. 42:1610–1619.
2021.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Netea MG, Domínguez-Andrés J, Barreiro LB,
Chavakis T, Divangahi M, Fuchs E, Joosten LAB, van der Meer JWM,
Mhlanga MM, Mulder WJM, et al: Defining trained immunity and its
role in health and disease. Nat Rev Immunol. 20:375–388.
2020.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Vinolo MAR, Rodrigues HG, Nachbar RT and
Curi R: Regulation of inflammation by short chain fatty acids.
Nutrients. 3:858–876. 2011.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Tanaka I, Ono S, Watanabe Y, Yamamoto H,
Oikawa R, Matsumoto S, Kubo M, Nishimura Y, Shimoda Y, Ono M, et
al: Long-term changes in aberrant DNA methylation and gastritis
after Helicobacter pylori eradication focused on
metachronous gastric cancer. Helicobacter.
27(e12915)2022.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Guccione C, Dantas Machado AC, Youssef F,
Angeli-Pahim I, Duarte S, Warren C, Farmer S, Humphrey G, Richter
RA, McDonald D, et al: Blood microbial DNA signature differentiates
hepatocellular carcinoma from metastatic lesions.
eGastroenterology. 3(e100193)2025.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Ramos A, Sadeghi S and Tabatabaeian H:
Battling chemoresistance in cancer: Root causes and strategies to
uproot them. Int J Mol Sci. 22(9451)2021.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Herrera-Quintana L, Vázquez-Lorente H,
Lopez-Garzon M, Cortés-Martín A and Plaza-Diaz J: Cancer and the
microbiome of the human body. Nutrients. 16(2790)2024.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Zhang H, Fu L, Leiliang X, Qu C, Wu W, Wen
R, Huang N, He Q, Cheng Q, Liu G and Cheng Y: Beyond the gut: The
intratumoral microbiome's influence on tumorigenesis and treatment
response. Cancer Commun (Lond). 44:1130–1167. 2024.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Sharma S, Hegde P, Panda S, Orimoloye MO
and Aldrich CC: Drugging the microbiome: Targeting small microbiome
molecules. Curr Opin Microbiol. 71(102234)2023.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Iida N, Dzutsev A, Stewart CA, Smith L,
Bouladoux N, Weingarten RA, Molina DA, Salcedo R, Back T, Cramer S,
et al: Commensal bacteria control cancer response to therapy by
modulating the tumor microenvironment. Science. 342:967–970.
2013.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Dong J, Li Y, Xiao H, Cui M and Fan S:
Commensal microbiota in the digestive tract: A review of its roles
in carcinogenesis and radiotherapy. Cancer Biol Med. 18:43–55.
2021.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Uribe-Herranz M, Rafail S, Beghi S,
Gil-de-Gómez L, Verginadis I, Bittinger K, Pustylnikov S, Pierini
S, Perales-Linares R, Blair IA, et al: Gut microbiota modulate
dendritic cell antigen presentation and radiotherapy-induced
antitumor immune response. J Clin Invest. 130:466–479.
2020.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Shiao SL, Kershaw KM, Limon JJ, You S,
Yoon J, Ko EY, Guarnerio J, Potdar AA, McGovern DPB, Bose S, et al:
Commensal bacteria and fungi differentially regulate tumor
responses to radiation therapy. Cancer Cell. 39:1202–1213.e6.
2021.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Colbert LE, El Alam MB, Wang R, Karpinets
T, Lo D, Lynn EJ, Harris TA, Elnaggar JH, Yoshida-Court K, Tomasic
K, et al: Tumor-resident Lactobacillus iners confer
chemoradiation resistance through lactate-induced metabolic
rewiring. Cancer Cell. 41:1945–1962.e11. 2023.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Alam MZ, Maslanka JR and Abt MC:
Immunological consequences of microbiome-based therapeutics. Front
Immunol. 13(1046472)2023.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Hizay A, Dag K, Oz N, Comak-Gocer EM,
Ozbey-Unlu O, Ucak M and Keles-Celik N: Lactobacillus
acidophilus regulates abnormal serotonin availability in
experimental ulcerative colitis. Anaerobe.
80(102710)2023.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Zheng L, Ji YY, Wen XL and Duan SL: Fecal
microbiota transplantation in the metabolic diseases: Current
status and perspectives. World J Gastroenterol. 28:2546–2560.
2022.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Davar D, Dzutsev AK, McCulloch JA,
Rodrigues RR, Chauvin JM, Morrison RM, Deblasio RN, Menna C, Ding
Q, Pagliano O, et al: Fecal microbiota transplant overcomes
resistance to anti-PD-1 therapy in melanoma patients. Science.
371:595–602. 2021.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Sun J, Song S, Liu J, Chen F, Li X and Wu
G: Gut microbiota as a new target for anticancer therapy: From
mechanism to means of regulation. NPJ Biofilms Microbiomes.
11(43)2025.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Zheng C, Chen T, Lu J, Wei K, Tian H, Liu
W, Xu T, Wang X, Wang S, Yang R, et al: Adjuvant treatment and
molecular mechanism of probiotic compounds in patients with gastric
cancer after gastrectomy. Food Funct. 12:6294–6308. 2021.PubMed/NCBI View Article : Google Scholar
|















