Introduction
Atherosclerosis, a chronic inflammatory disease of
blood vessels, is one of the main causes of cardiovascular disease,
which is the most common cause of mortality in industrialized
societies and is increasingly becoming the leading cause of
mortality worldwide (1).
Atherosclerosis is induced by multiple factors and regulated by a
number of genes (2). Macrophages,
which are known to reside within atherosclerotic plaques,
contribute to the pathology of atherosclerosis by internalizing
native or modified lipoproteins or lipoprotein remnants that have
invaded the vessel wall to form cholesterol-rich foam cells
(3). As macrophages are one of the
precursors of foam cells, their cholesterol counter transport
system (cholesterol efflux) is important to maintain the balance of
cholesterol in cells and influence the formation of foam cells
(4). The conversion of macrophages
into foam cells is orchestrated by disruption of the normal
cholesterol homeostatic mechanism that controls the uptake,
intracellular metabolism and efflux of cholesterol (5).
The peroxisome proliferator-activated receptor γ
(PPARγ), a member of a superfamily of ligand-dependent
transcription factors that regulate immunity and inflammation, is
one of the nuclear receptors expressed in macrophages (6,7).
Numerous studies have indicated that PPARγ and its ligands promote
cholesterol efflux from macrophages through the PPARγ-liver X
receptor α-ATP-binding cassette, sub-family A, member 1 signaling
pathway (8), and this process may
downregulate the expression of pro-inflammatory genes in
macrophages that may be associated with the transrepression of the
transcription factor nuclear factor κ-light-chain-enhancer of
activated B cells (NF-κB) (9,10). A
study concerning the anti-inflammatory effects of PPARγ has shown
that its agonists markedly inhibit the secretion of
pro-inflammatory mediators, including tumor necrosis factor-α and
interleukin-1 and -6 in activated macrophages (11). However, the effect of PPARγ on the
cholesterol efflux of macrophages in inflammation remains unclear.
Pretreating wild-type mice with PPARγ ligands may reduce the
expression of pro-inflammatory cytokines and alleviate injury of
local and distant tissues (12),
which has a therapeutic effect in numerous inflammatory diseases,
including acute myocarditis, autoimmune encephalitis and multiple
sclerosis (13).
Atherosclerosis has been acknowledged as a
consequence of lipid metabolism disorder and chronic inflammation
(14). Thus, with multiple
factors, including hypercholesterolemia and inflammation, promoting
atherosclerosis either individually or in combination, it is of
great significance to clarify how the mechanism of cholesterol
efflux from macrophages changes and the role of PPARγ in these
situations. This will help to explain the formation of foam cells
and provide novel methods of preventing and curing
atherosclerosis.
In view of the action of PPARγ on various key
transcriptional factors, we proposed the hypothesis that PPARγ is
the primary regulator of macrophage cholesterol efflux and
suppressor of the inflammatory response. The present study aimed to
provide evidence to elucidate the possible mechanism of PPARγ on
the cholesterol efflux of peritoneal macrophages in inflammation
and the role of PPARγ in maintaining the balance between the
cholesterol efflux and anti-inflammatory response.
Materials and methods
Reagents and kits
LPS (Escherichia coli, O111:B4) was purchased
from Sigma (St. Louis, MO, USA) and reconstituted in
phosphate-buffered saline (PBS). PPARγ antibody (rabbit anti-mouse)
and PPARγ antibody (goat anti-rabbit) were purchased from Sigma.
Phosphor-nuclear factor of κ light polypeptide gene enhancer in
B-cells inhibitor α (IκBα; Ser32) was purchased from Youyizhonglian
Bio-Corporation (Beijing, China). [H3] cholesterol and
apolipoprotein AI (ApoAI) were purchased from Sigma. Ciglitazone
was purchased from Sigma, and the final concentration of
ciglitazone dissolved in dimethylsulfoxide (DMSO) was 3
μmol/ml. The sequences of the PPARγ antisense and missense
oligonucleotides were 5′-CATGAGGCTTATTGTAGAGCTGA-3′ and
5′-GCCAGGTACCACTCACTCTGCAGT-3′, respectively. The procedure of
synthesis, purification and subpackage of the sequence was operated
by Shenggong Bio-Corporation (Shanghai, China).
Animals
Fifteen C57BL/6 mice (8–10 weeks old, males,
weighing 20–26 g) were obtained from the Laboratory Animal Centre
of Chongqing Medical University (Chongqing, China). These mice were
housed in an animal room and fed a standard diet. All experimental
protocols described in this study were approved by the Ethics
Review Committee for Animal Experimentation of Chongqing Medical
University.
Experimental protocol
The 15 mice were randomly divided into three groups.
Proceeding from isolation and culture of peritoneal macrophages
from the C57BL/6 mice, the cells were divided into three groups:
The control group, the ciglitazone group and the PPARγ antisense
oligonucleotide group. The expression of PPARγ and IκBα in each
group was observed through the levels of protein and mRNA, and then
the cholesterol efflux of each group was investigated. The
intraperitoneal injection of LPS into mice is a widely used method
of constructing inflammatory animal models (15). In addition, the same experiment was
repeated subsequent to stimulation of each group with LPS.
Isolation and treatment of peritoneal
macrophages
Pre-cooled PBS (2 ml) was injected into the
abdominal cavity of the mice, whilst the abdomen was kneaded softly
for 2 min. The PBS was drawn out and collected, and then
centrifuged for 10 min at 2,000 × g. The supernatant liquid was
discarded and placed in RPMI-1640, which regulated the
concentration of the cells at 3–5×106cells/ml. The cells
were cultivated in 24-well plates at 37°C for 2 h until they had
adhered, then the cultivation holes were washed with pre-cooled
PBS. The adherent cells were peritoneal macrophages. The peritoneal
macrophages were cultivated for 24 h and then randomly divided into
three groups: The control group (RMPI-1640+25 μl DMSO), the
ciglitazone group (RMPI-1640+25 μl DMSO+ciglitazone; final
concentration, 10 μmol/l), and the PPARγ antisense oligonucleotide
group (RMPI-1640+25 μl DMSO+PPARγ antisense oligonucleotide; final
concentration, 400 nmol/l). The final concentration of LPS was 80
ng/ml.
Immunocytochemical staining analysis of
peritoneal macrophages
The peritoneal macrophages were cultured on a
chamber slide, which was washed with PBS and air-dried. The slide
was fixed with methanol for 30 min at −20°C and then stained with
PPARγ antibody (rabbit anti-mouse; Abcam, Cambridge, MA, USA) and
PPARγ antibody (goat anti-rabbit; Abcam) for 24 h at room
temperature. The cells that were stained purple were considered
positive.
Assessment of gene expression of PPARγ by
quantitative reverse transcription-polymerase chain reaction
(RT-PCR)
Total RNA samples of the peritoneal macrophages were
extracted using an RNA extraction kit Takara Bio Inc. (Shiga,
Japan) according to the manufacturer’s instructions. Total RNA was
quantified with the ratio of absorption values of RNA samples at
260 and 280 nm. Each total RNA sample was reversely transcribed to
complementary DNA using an RT-PCR kit and stored at −70°C. All PCR
products were electrophoresed on 2% agarose gels. The RT-PCR was
performed using the sense and antisense primers for PPARγ or
β-actin (Table I). The relative
expression of mRNAs were assessed by taking the ratio of the
intensity of the DNA bands of PPARγ to the β-actin band using the
Bio-Image analysis system (Gel Doc 2000; Bio-Rad, Hercules, CA,
USA) and expressed as arbitrary units.
 | Table IPrimer sequences for RT-PCR. |
Table I
Primer sequences for RT-PCR.
| DNA amplified | Primer sequence
(5′→3′) | Size (bp) |
|---|
| PPARγ | | 476 |
| Sense |
CAATCCGAATTTTTCAAGGGTGCCA | |
| Antisense |
GAGCACCTTGGCGAACAGCTGAGAG | |
| β-actin | | 355 |
| Sense |
GAGAAGAGCTATGAACTTCCTGACG | |
| Antisense |
TTTGCTGGAAGGTGGACAGAGAGGC | |
Western blotting analysis of PPARγ and
IκBα
Total protein of the peritoneal macrophages was
extracted by homogenizing the macrophages in a cell lysis buffer
(Beyotime Institute of Biotechnology, Shanghai, China), then by two
cycles of centrifugation at 12,000 × g for 15 min. Protein
concentration was determined using a Bradford assay kit (Beyotime
Institute of Biotechnology). The total protein was separated by 10%
SDS-PAGE and transferred to polyvinylidene fluoride membranes,
which were then incubated with rabbit anti-mouse PPARγ polyclonal
antibody (diluted 1:1000, IMG-441; Sigma) and horseradish
peroxidase-conjugated goat anti-rabbit IgG (diluted 1:2000;
Zhongshan Jinqiao, Beijing, China). The immune complexes were
developed with an Enhanced Chemiluminescence Detection kit (Pierce
Biotechnology, Inc., Rockford, IL, USA) and the membranes were then
immediately exposed to autoradiographic film (Kodak, Rochester, NY,
USA). The relative amount of PPARγ protein was quantified from the
optical density of the corresponding band by Bio-Image analysis
system (Gel Doc 2000; Bio-Rad).
Determination of peritoneal macrophage
cholesterol efflux
The concentration of macrophages was regulated at
3.0×109 cells/l and the macrophages were transferred
into RPMI-1640 containing fetal bovine serum, penicillin and
streptomycin, and [3H] cholesterol. After 24 h, the
cells were cultivated in new medium containing 50 μg/ml ApoAI for
12 h. The [3H]cholesterol in the culture solution and
cells was detected by liquid scintillation counting. The effluxion
of cholesterol was calculated using the following formula:
[3H] (culture solution)/[3H] (culture
solution and cells) × 100.
Statistical analysis
Data are reported as the mean ± standard deviation
and were analyzed using one-way analysis of variance with Tukey’s
multiple comparison test, and the statistical program SPSS, version
11.0 (SPSS, Inc., Chicago, IL, USA). P≤0.05 was considered to
indicate a statistically significant difference.
Results
Immunocytochemical staining analysis of
peritoneal macrophages
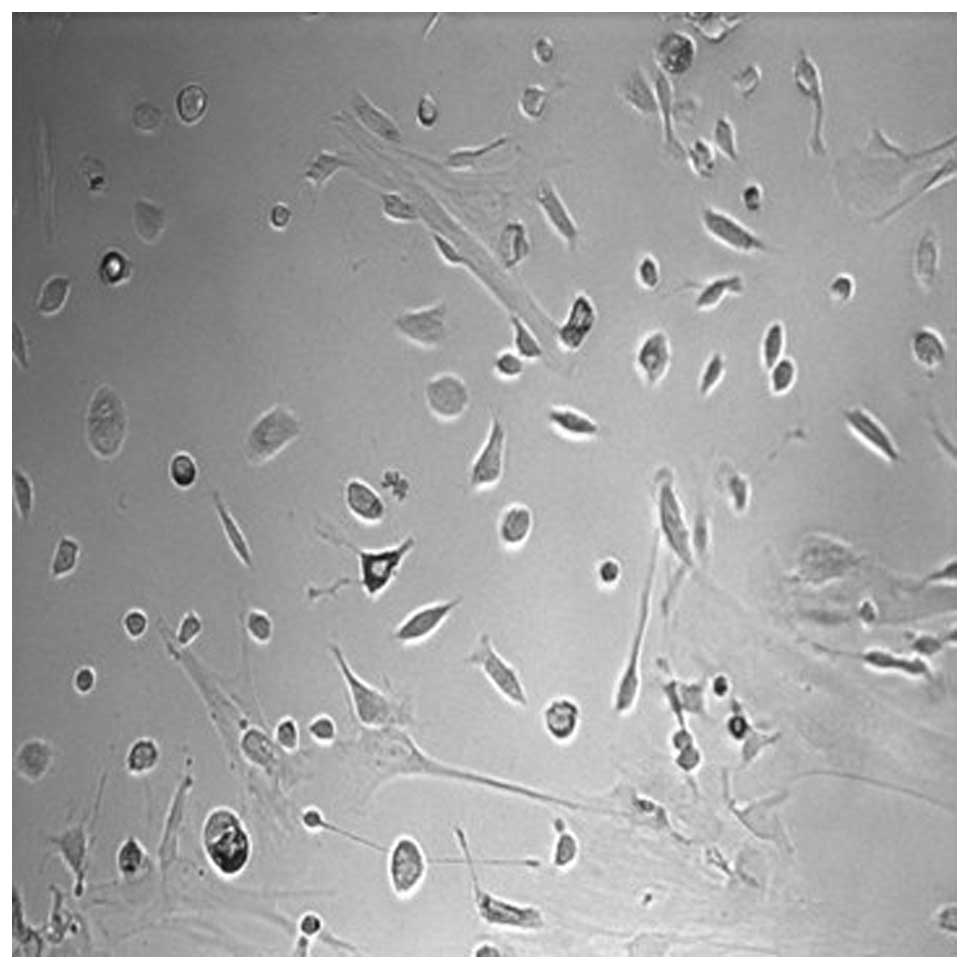
Peritoneal macrophages from C57BL/6 mice were
isolated and cultured (Fig. 1).
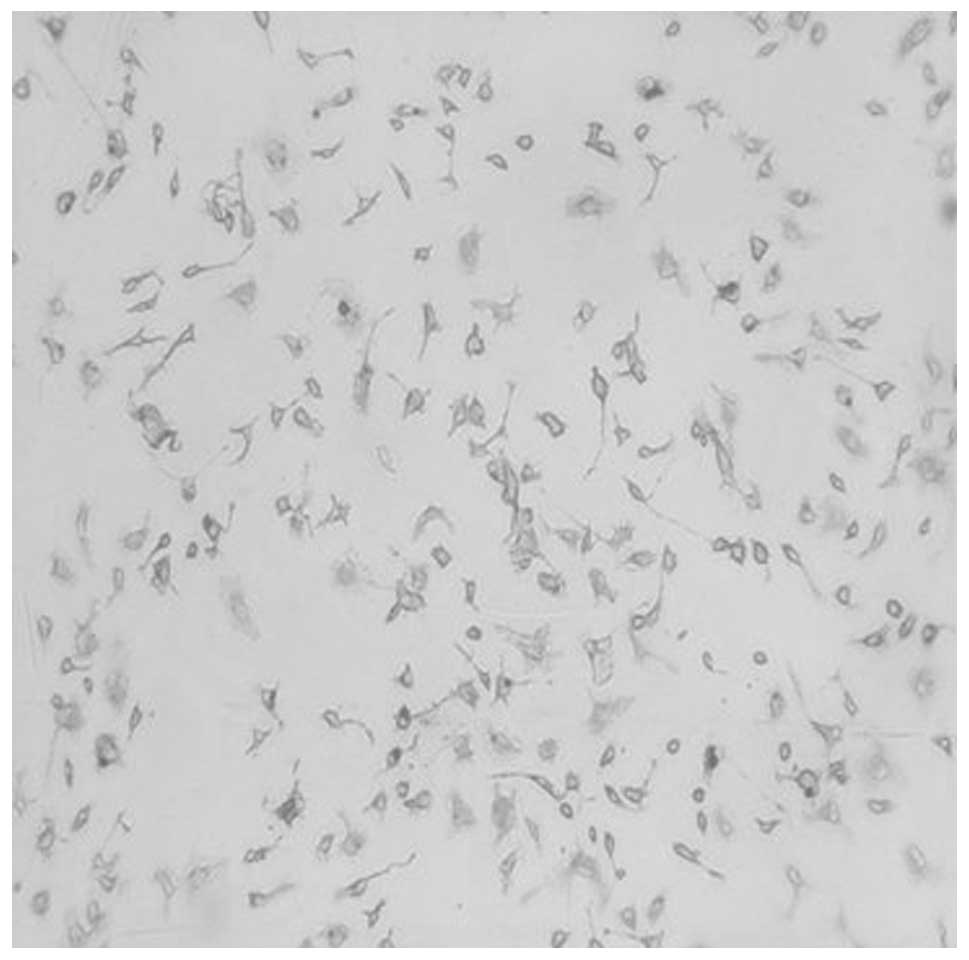
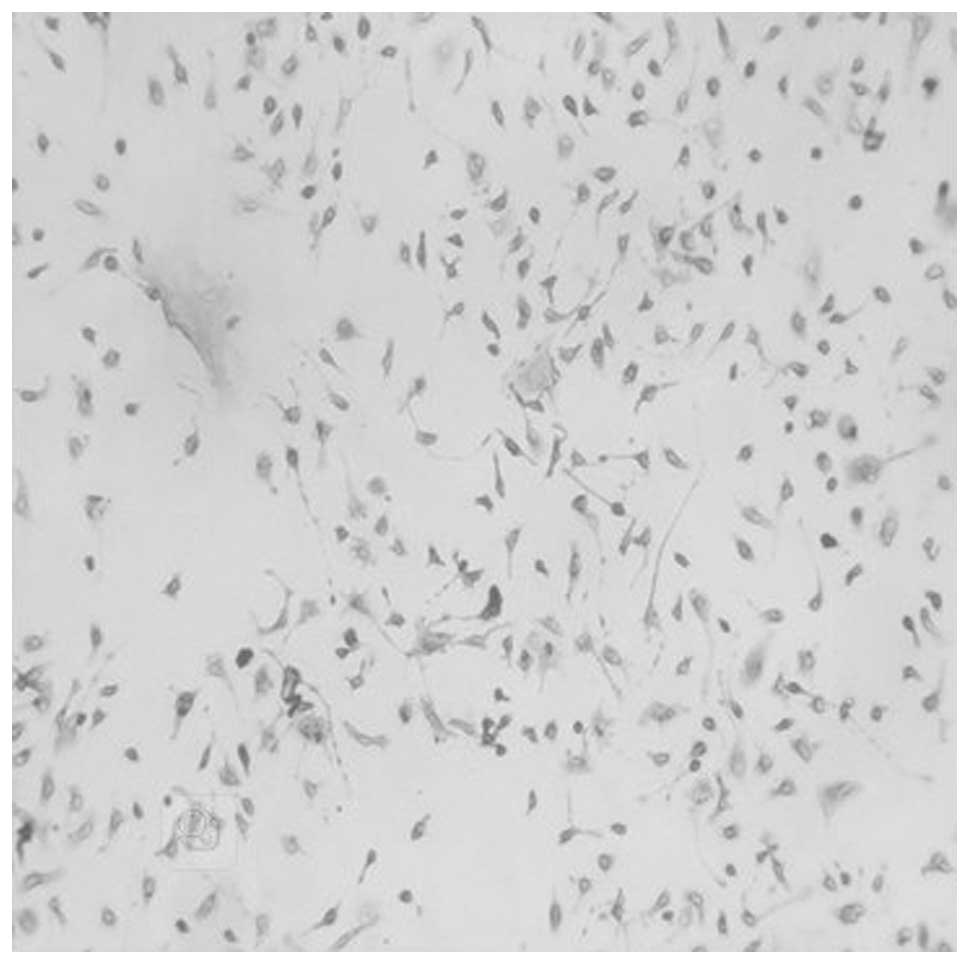
Subsequently, PPARγ in peritoneal macrophages prior to and
following stimulation by LPS was examined using immunocytochemistry
(Figs. 2 and 3). It was found that the number of
PPARγ-positive cells following stimulation by LPS was greater than
that prior to stimulation by LPS. The PPARγ-positive cells were
stained purple.
Expression of PPARγ mRNA in peritoneal
macrophages pre- and post-LPS stimulation
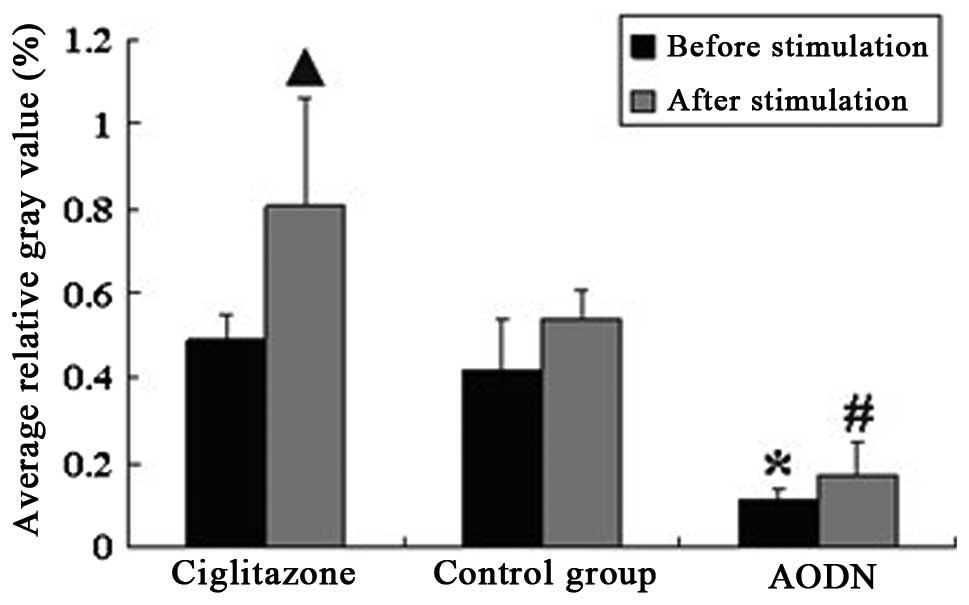
The results of the RT-PCR indicated that there was
no significant difference in the average relative gray value
between the control group and ciglitazone group. The average
relative gray value of the PPARγ antisense oligonucleotide group
was evidently lower than that of control group. Following
stimulation with LPS, the expression levels of PPARγ mRNA in the
ciglitazone group were higher than those in the control group,
while the PPARγ mRNA expression levels of the PPARγ antisense
oligonucleotide group were lower than those of the control group
(Figs. 4 and 5).
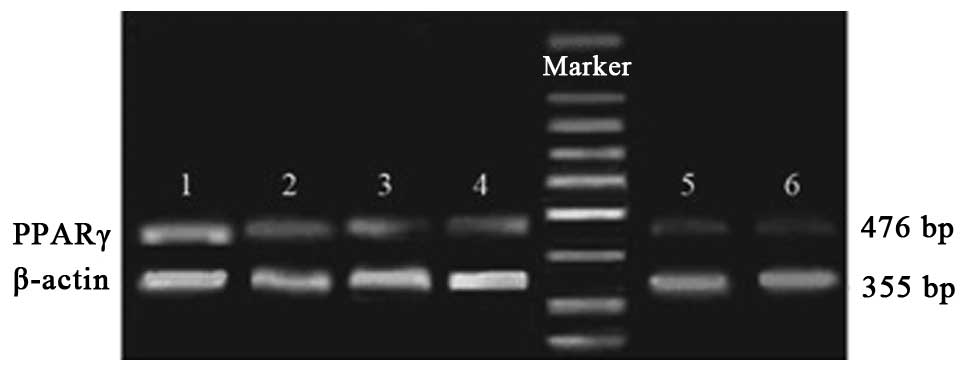 | Figure 4RT-PCR analysis of PPARγ mRNA
expression in each group prior to and following stimulation with
LPS. Group 1, stimulated ciglitazone group; group 2, unstimulated
ciglitazone group; group 3, stimulated control group; group 4,
unstimulated control group; group 5, stimulated AODN group; group
6, unstimulated AODN group. PPARγ, peroxisome
proliferator-activated receptor γ; RT-PCR, reverse
transcription-polymerase chain reaction; LPS, lipopolysaccharide;
AODN, antisense oligonucleotide group. |
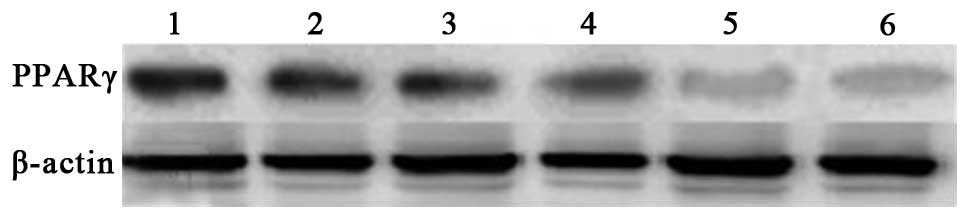
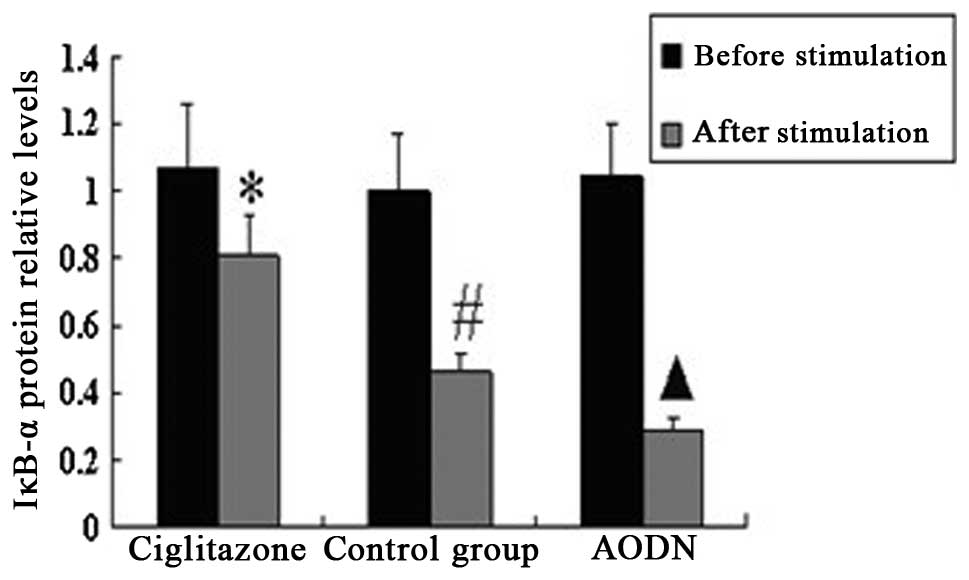
Expression of PPARγ and IκBα protein in
peritoneal macrophages pre- and post-LPS stimulation
The results of the western blotting suggested that
there was no significant difference in the expression of PPARγ
protein between the control group and ciglitazone group. The PPARγ
protein expression levels of the PPARγ antisense oligonucleotide
group were considerably lower than those of the control group.
Subsequent to stimulation with LPS, the expression levels of PPARγ
protein in the three groups were higher than those of each group
prior to stimulation, and the IκBα protein expression levels of the
three groups were lower than those of each group prior to
stimulation (Figs. 6–9).
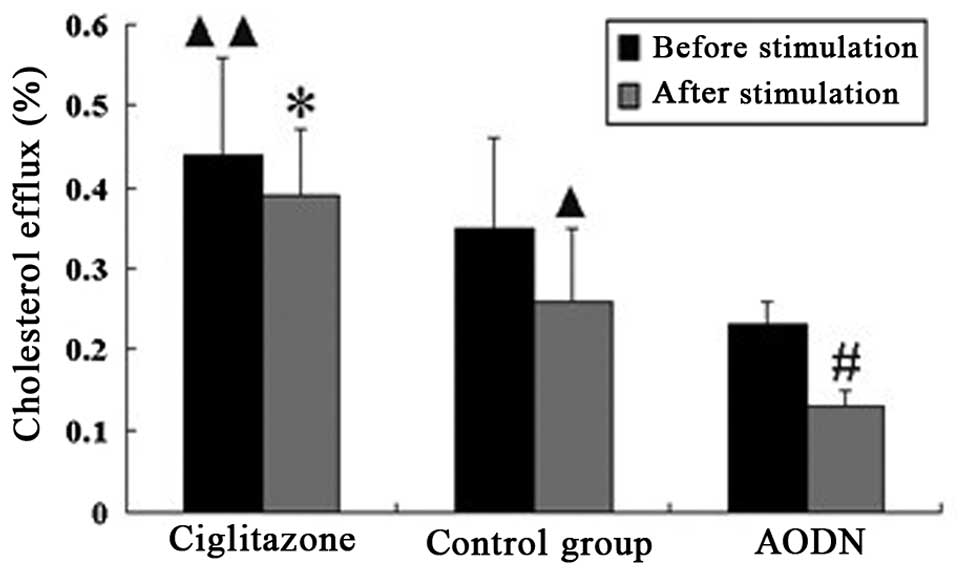
Cholesterol efflux of peritoneal
macrophages in each group pre- and post-LPS stimulation
The cholesterol efflux of the ciglitazone group was
suppressed following stimulation with LPS, and the suppression
ratio was lower than that of the control group. However, the
cholesterol efflux of the PPARγ antisense oligonucleotide group was
greatly suppressed following stimulation with LPS, and the
suppression ratio was higher than that of the control group
(Table II, Fig. 10).
 | Table IICholesterol efflux of the peritoneal
macrophages in each group prior to and following stimulation with
LPS (mean ± standard deviation, %). |
Table II
Cholesterol efflux of the peritoneal
macrophages in each group prior to and following stimulation with
LPS (mean ± standard deviation, %).
| Measurement | Ciglitazone | Control | AODN |
|---|
| Cholesterol
efflux |
| Prior to
stimulation | 0.44±0.12d | 0.35±0.11 | 0.23±0.03 |
| Following
stimulation | 0.39±0.08a | 0.26±0.09b | 0.13±0.02c |
| Suppression
ratio | 11.37 | 25.72 | 43.48 |
Discussion
It has been universally acknowledged that
atherosclerosis is a disease associated with lipid metabolic
disturbance and chronic inflammation (16). Foam cells form from macrophages, in
which the cholesterol accumulation is the significant pathological
characteristic of atherosclerotic lesions (17). The balance of cholesterol in
macrophages depends on coordinated regulation of cholesterol,
including intake, storage, de novo synthesis and efflux
(18). As a thiazolidinedione,
ciglitazone is a high-affinity ligand for PPARγ (19,20),
and is able to activate PPARγ to suppress the activation of
inflammatory cells and delivery of inflammatory mediators (21).
In the present study, measurement of the cholesterol
efflux of peritoneal macrophages in each group indicated that
pretreating macrophages with ciglitazone increases the cholesterol
efflux. However, the cholesterol efflux was weaker in the PPARγ
antisense oligonucleotide group compared with that in the control
group. This demonstrates that PPARγ, when activated by its ligand
ciglitazone, greatly reinforces the cholesterol efflux of
peritoneal macrophages. In inflammation, the cholesterol efflux of
the three groups was suppressed, but the suppression ratio varied.
The cholesterol efflux of the PPARγ antisense oligonucleotide group
was evidently suppressed following stimulation with LPS, and the
suppression ratio was higher than that of the other two groups. The
technique of knockdown using antisense nucleic acids was selected
due to its benefits, which include strong specificity to target
site, few side-effects and a precise depression effect (22). The results indicate that PPARγ is
associated with the suppression of the cholesterol efflux resulting
from LPS stimulation. This study demonstrated that stimulation of
peritoneal macrophages with LPS suppresses the cholesterol efflux,
even when the expression of PPARγ is upregulated, so it was
presumed that the activation of PPARγ was affected due to its
anti-inflammatory characteristic. When pro-atherosclerotic factors,
including inflammation and hypercholesteremia coexist, the
anti-inflammatory effect of PPARγ is of great significance.
The activation of NF-κB is an important signal
transmission pathway that produces various pro-inflammatory factors
(23). NF-κB consists mainly of
the heterodimer p50/p65, which is generally bound to IκB,
maintaining a state of inactivation in the cytoplasm (24,25).
LPS binds to the corresponding receptor in the cytomembrane and
leads to phosphorylation and degradation of IκB, and then NF-κB is
released into the nucleus to promote the transcription of target
genes (26). Certain studies have
indicated that PPARγ may suppress several inflammation-correlated
signaling pathways, including Janus kinase-signal transducer and
activator of transcription, NF-κB, nuclear factor of activated T
cell and activator protein 1, to express the anti-inflammatory
effect (27–30). In order to explore the association
between PPARγ and NF-κB in peritoneal macrophages in inflammation,
the three groups in the present study were stimulated with LPS. The
results indicated that the expression of IκBα was downregulated in
each group by LPS, and the downregulation of IκBα in the PPARγ
antisense oligonucleotide group was more significant than that in
the other two groups. We consider PPARγ to be closely connected
with NF-κB in peritoneal macrophages in inflammation, and PPARγ may
produce anti-inflammatory effects by protecting IκBα from being
phosphorylated and degraded in order to influence the activation
and nuclear translocation of NF-κB.
In conclusion, the present study demonstrates that
PPARγ performs a role in anti-inflammation by means of protecting
IκBα from being phosphorylated and degraded and promoting
cholesterol efflux from peritoneal macrophages in inflammation. As
the understanding of the complex association between PPARγ and the
cholesterol efflux from peritoneal macrophages in inflammation
increases, there will undoubtedly be an increasing number of
opportunities to apply knowledge to the management and ultimately
the prevention of atherosclerosis.
Acknowledgements
This study was granted financial support from the
National Natural Science Foundation of China (grant 30772098) and
Chongqing Science Technology Commission (No.s cstc2010bb5386 and
cstc2012jjA10090).
References
|
1
|
Klingenberg R and Hansson GK: Treating
inflammation in atherosclerotic cardiovascular disease: emerging
therapies. Eur Heart J. 30:2838–2844. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Moore KJ and Tabas I: Macrophages in the
pathogenesis of atherosclerosis. Cell. 145:341–355. 2011.
View Article : Google Scholar
|
|
3
|
Hansson GK and Hermansson A: The immune
system in atherosclerosis. Nat Immunol. 12:204–212. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Reiss AB and Cronstein BN: Regulation of
foam cells by adenosine. Arterioscler Thromb Vasc Biol. 32:879–886.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
McLaren JE, Michael DR, Ashlin TG and
Ramji DP: Cytokines, macrophage lipid metabolism and foam cells:
implications for cardiovascular disease therapy. Prog Lipid Res.
50:331–347. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Glass CK and Saijo K: Nuclear receptor
transrepression pathways that regulate inflammation in macrophages
and T cells. Nat Rev Immunol. 10:365–376. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Villacorta L, Schopfer FJ, Zhang J, et al:
PPARgamma and its ligands: therapeutic implications in
cardiovascular disease. Clin Sci (Lond). 116:205–218. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Bouhlel MA, Staels B and Chinetti-Gbaguidi
G: Peroxisome proliferator-activated receptors - from active
regulators of macrophage biology to pharmacological targets in the
treatment of cardiovascular disease. J Intern Med. 263:28–42.
2008.
|
|
9
|
Rigamonti E, Chinetti-Gbaguidi G and
Staels B: Regulation of macrophage functions by PPAR-alpha,
PPAR-gamma, and LXRs in mice and men. Arterioscler Thromb Vasc
Biol. 28:1050–1059. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Tabas I: Macrophage death and defective
inflammation resolution in atherosclerosis. Nat Rev Immunol.
10:36–46. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Martinez FO, Helming L and Gordon S:
Alternative activation of macrophages: an immunologic functional
perspective. Annu Rev Immunol. 27:451–483. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Takano H and Komuro I: Peroxisome
proliferator-activated receptor gamma and cardiovascular diseases.
Circ J. 73:214–220. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lamers C, Schubert-Zsilavecz M and Merk D:
Therapeutic modulators of peroxisome proliferator-activated
receptors (PPAR): a patent review (2008-present). Expert Opin Ther
Pat. 22:803–841. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kajinami K and Kawai Y: Beyond C-reactive
protein; new evidence for another inflammatory biomarker predicting
cardiovascular disease risk. Atherosclerosis. 214:39–40. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lawrence CB, Brough D and Knight EM: Obese
mice exhibit an altered behavioural and inflammatory response to
lipopolysaccharide. Dis Model Mech. 5:649–659. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lloyd-Jones D, Adams RJ, Brown TM, et al:
American Heart Association Statistics Committee and Stroke
Statistics Subcommittee: Executive summary: heart disease and
stroke statistics - 2010 update: a report from the American Heart
Association. Circulation. 121:948–954. 2010. View Article : Google Scholar
|
|
17
|
Zhao Y, Pennings M, Vrins CL, et al:
Hypocholesterolemia, foam cell accumulation, but no atherosclerosis
in mice lacking ABC-transporter A1 and scavenger receptor BI.
Atherosclerosis. 218:314–322. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Uto-Kondo H, Ayaori M, Ogura M, et al:
Coffee consumption enhances high-density lipoprotein-mediated
cholesterol efflux in macrophages. Circ Res. 106:779–787. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ogawa Y, Yoneda M, Tomeno W, et al:
Peroxisome proliferator-activated receptor gamma exacerbates
Concanavalin A-induced liver injury via suppressing the
translocation of NF-κB into the nucleus. PPAR Res.
2012:9403842012.PubMed/NCBI
|
|
20
|
Norazmi MN, Mohamed R, Nurul AA and Jaacob
NS: The modulation of PPARγ1 and PPARγ2 mRNA expression by
ciglitazone in CD3/CD28-activated naïve and memory CD4+
T cells. Clin Dev Immunol. 2012:8491952012.
|
|
21
|
Tobiasova Z, Zhang L, Yi T, et al:
Peroxisome proliferator-activated receptor-γ agonists prevent in
vivo remodeling of human artery induced by alloreactive T
cells. Circulation. 124:196–205. 2011.
|
|
22
|
Visser ME, Witztum JL, Stroes ES and
Kastelein JJ: Antisense oligonucleotides for the treatment of
dyslipidaemia. Eur Heart J. 33:1451–1458. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Robinson SM and Mann DA: Role of nuclear
factor kappaB in liver health and disease. Clin Sci (Lond).
118:691–705. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tornatore L, Thotakura AK, Bennett J, et
al: The nuclear factor kappa B signaling pathway: integrating
metabolism with inflammation. Trends Cell Biol. 22:557–566. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kwak JH, Jung JK and Lee H: Nuclear
factor-kappa B inhibitors; a patent review (2006–2010). Expert Opin
Ther Pat. 21:1897–1910. 2011.
|
|
26
|
Kim IT, Ryu S, Shin JS, et al: Euscaphic
acid isolated from roots of Rosa rugosa inhibits LPS-induced
inflammatory responses via TLR4-mediated NF-κB inactivation in RAW
264.7 macrophages. J Cell Biochem. 113:1936–1946. 2012.PubMed/NCBI
|
|
27
|
Bright JJ, Kanakasabai S, Chearwae W and
Chakraborty S: PPAR regulation of inflammatory signaling in CNS
diseases. PPAR Res. 2008:6585202008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Hirsch J, Johnson CL, Nelius T, et al:
PEDF inhibits IL8 production in prostate cancer cells through PEDF
receptor/phospholipase A2 and regulation of NFκB and PPARγ.
Cytokine. 55:202–210. 2011.PubMed/NCBI
|
|
29
|
Simone RE, Russo M, Catalano A, et al:
Lycopene inhibits NF-κB-mediated IL-8 expression and changes redox
and PPARγ signalling in cigarette smoke-stimulated macrophages.
PLoS One. 6:e196522011.
|
|
30
|
Fan B, Ikuyama S, Gu JQ, et al: Oleic
acid-induced ADRP expression requires both AP-1 and PPAR response
elements, and is reduced by Pycnogenol through mRNA degradation in
NMuLi liver cells. Am J Physiol Endocrinol Metab. 297:E112–123.
2009. View Article : Google Scholar : PubMed/NCBI
|