|
1
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Yang L, Parkin DM, Ferlay J, Li L and Chen
Y: Estimates of cancer incidence in China for 2000 and projections
for 2005. Cancer Epidemiol Biomarkers Prev. 14:243–250.
2005.PubMed/NCBI
|
|
3
|
Chon HS and Lancaster JM: Microarray-based
gene expression studies in ovarian cancer. Cancer Control. 18:8–15.
2011.PubMed/NCBI
|
|
4
|
Vitucci M, Hayes DN and Miller CR: Gene
expression profiling of gliomas: merging genomic and
histopathological classification for personalised therapy. Br J
Cancer. 104:545–553. 2011. View Article : Google Scholar :
|
|
5
|
Nannini M, Pantaleo MA, Maleddu A, Astolfi
A, Formica S and Biasco G: Gene expression profiling in colorectal
cancer using microarray technologies: results and perspectives.
Cancer Treat Rev. 35:201–209. 2009. View Article : Google Scholar
|
|
6
|
Bartel DP: MicroRNAs: genomics,
biogenesis, mechanism and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kozomara A and Griffiths-Jones S: miRBase:
integrating microRNA annotation and deep-sequencing data. Nucleic
Acids Res. 39(Database Issue): 152–157. 2011. View Article : Google Scholar
|
|
9
|
Li X, Zhang J, Gao L, et al: MiR-181
mediates cell differentiation by interrupting the Lin28 and let-7
feedback circuit. Cell Death Differ. 19:378–386. 2012. View Article : Google Scholar :
|
|
10
|
Krützfeldt J, Rajewsky N, Braich R, et al:
Silencing of microRNAs in vivo with ‘antagomirs’. Nature.
438:685–689. 2005. View Article : Google Scholar
|
|
11
|
Elmén J, Lindow M, Schütz S, et al:
LNA-mediated microRNA silencing in non-human primates. Nature.
452:896–899. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lee YS and Dutta A: MicroRNAs: small but
potent oncogenes or tumor suppressors. Curr Opin Investig Drugs.
7:560–564. 2006.PubMed/NCBI
|
|
13
|
Caldas C and Brenton JD: Sizing up miRNAs
as cancer genes. Nat Med. 11:712–714. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Mathé EA, Nguyen GH, Bowman ED, Zhao Y, et
al: MicroRNA expression in squamous cell carcinoma and
adenocarcinoma of the esophagus: associations with survival. Clin
Cancer Res. 15:6192–6200. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Li J, Chen Z, Tian L, Zhou C, et al:
LncRNA profile study reveals a three-lncRNA signature associated
with the survival of patients with oesophageal squamous cell
carcinoma. Gut. 63:1700–1710. PubMed/NCBI
|
|
16
|
Diboun I, Wernisch L, Orengo CA and
Koltzenburg M: Microarray analysis after RNA amplification can
detect pronounced differences in gene expression using limma. BMC
Genomics. 7:2522006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
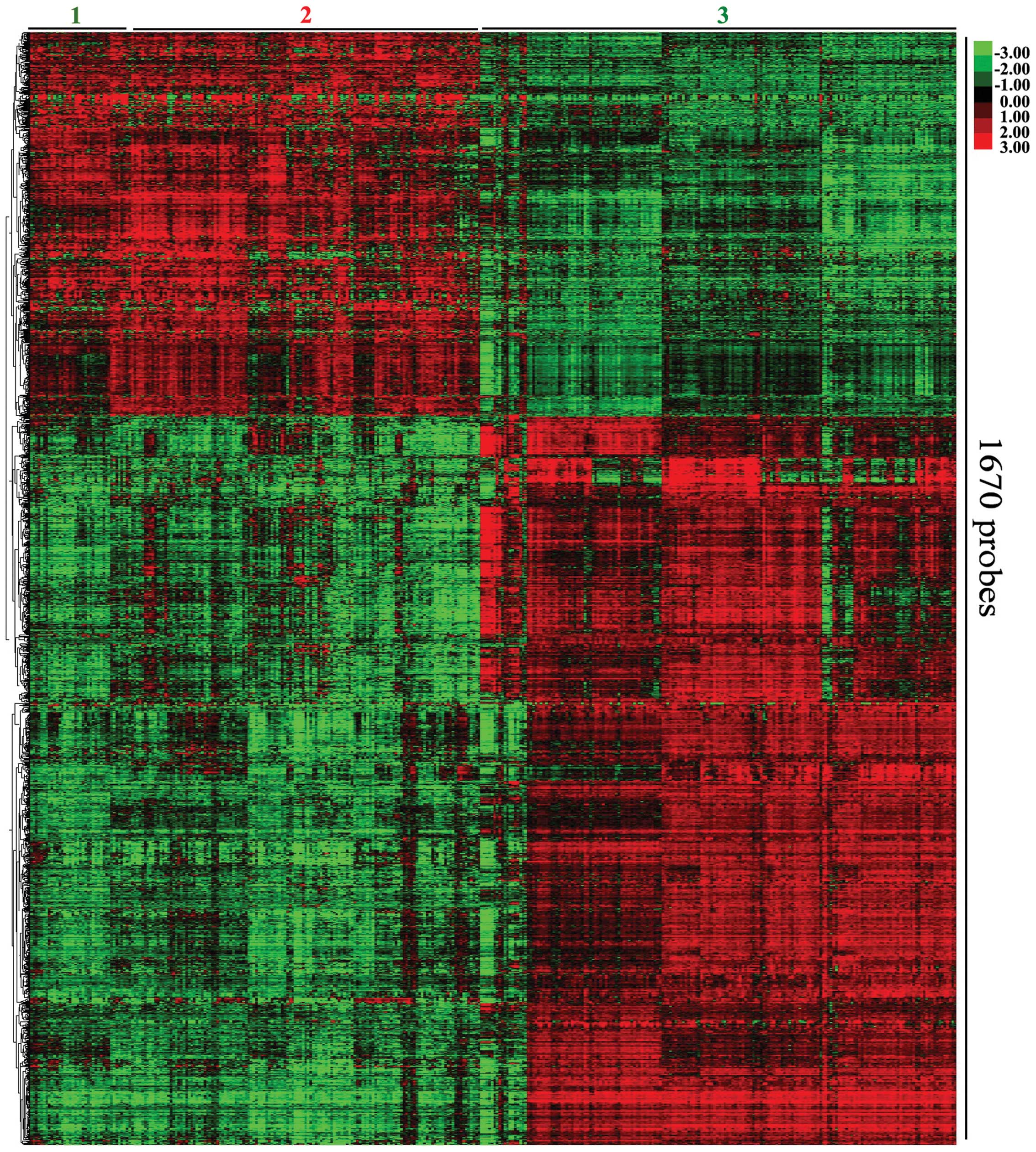
Eisen MB, Spellman PT, Brown PO and
Botstein D: Cluster analysis and display of genome-wide expression
patterns. Proc Natl Acad Sci USA. 95:14863–14868. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Dennis G Jr, Sherman BT, Hosack DA, et al:
DAVID: Database for Annotation, Visualization and Integrated
Discovery. Genome Biol. 4:P32003. View Article : Google Scholar
|
|
19
|
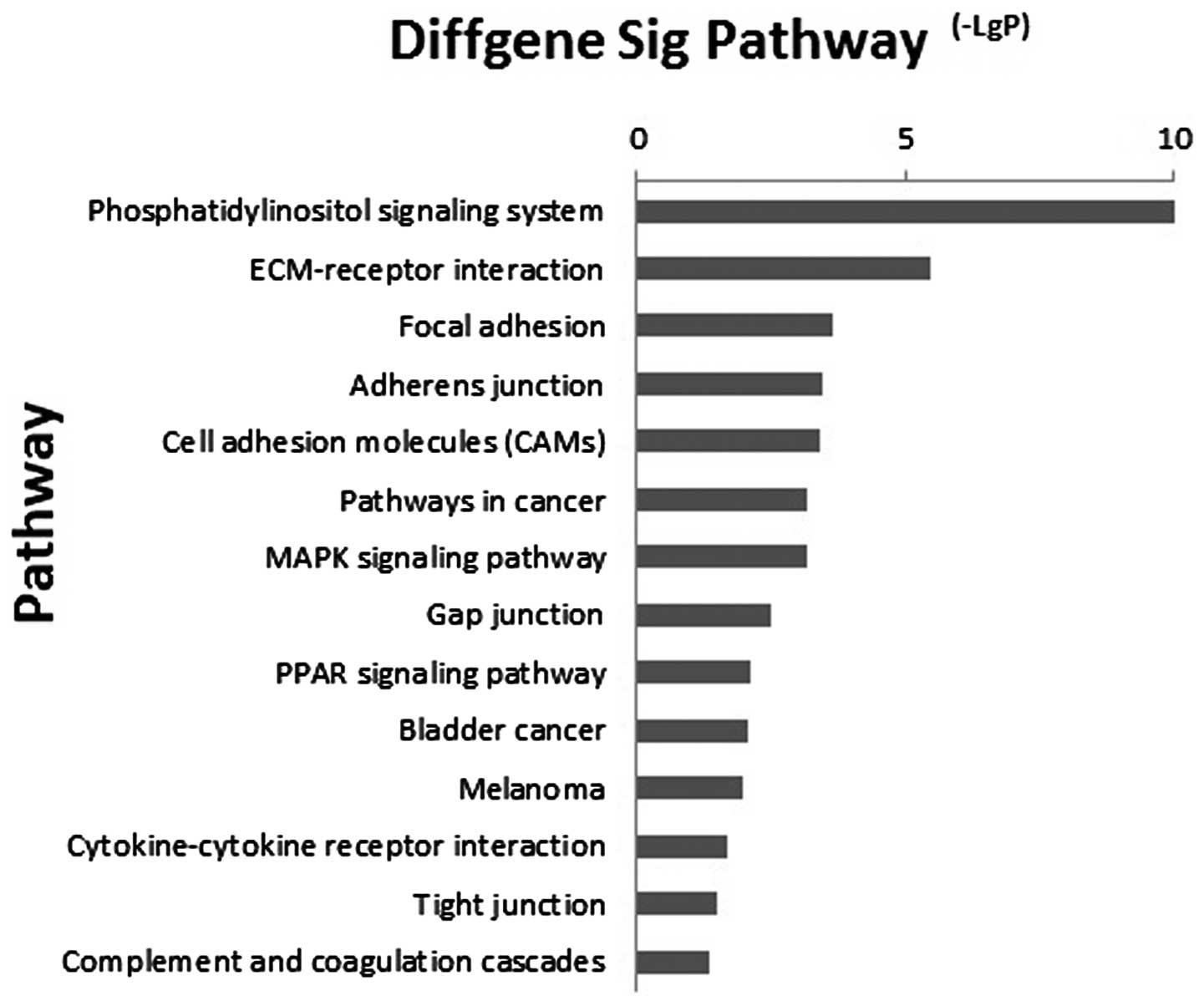
Kanehisa M, Goto S, Kawashima S, Okuno Y
and Hattori M: The KEGG resource for deciphering the genome.
Nucleic Acids Res. 32(Database Issue): 277–280. 2004. View Article : Google Scholar
|
|
20
|
Yi M, Horton JD, Cohen JC, Hobbs HH and
Stephens RM: WholePathwayScope: a comprehensive pathway-based
analysis tool for high-throughput data. BMC Bioinformatics.
7:302006. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Draghici S, Khatri P, Tarca AL, et al: A
systems biology approach for pathway level analysis. Genome Res.
17:1537–1545. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
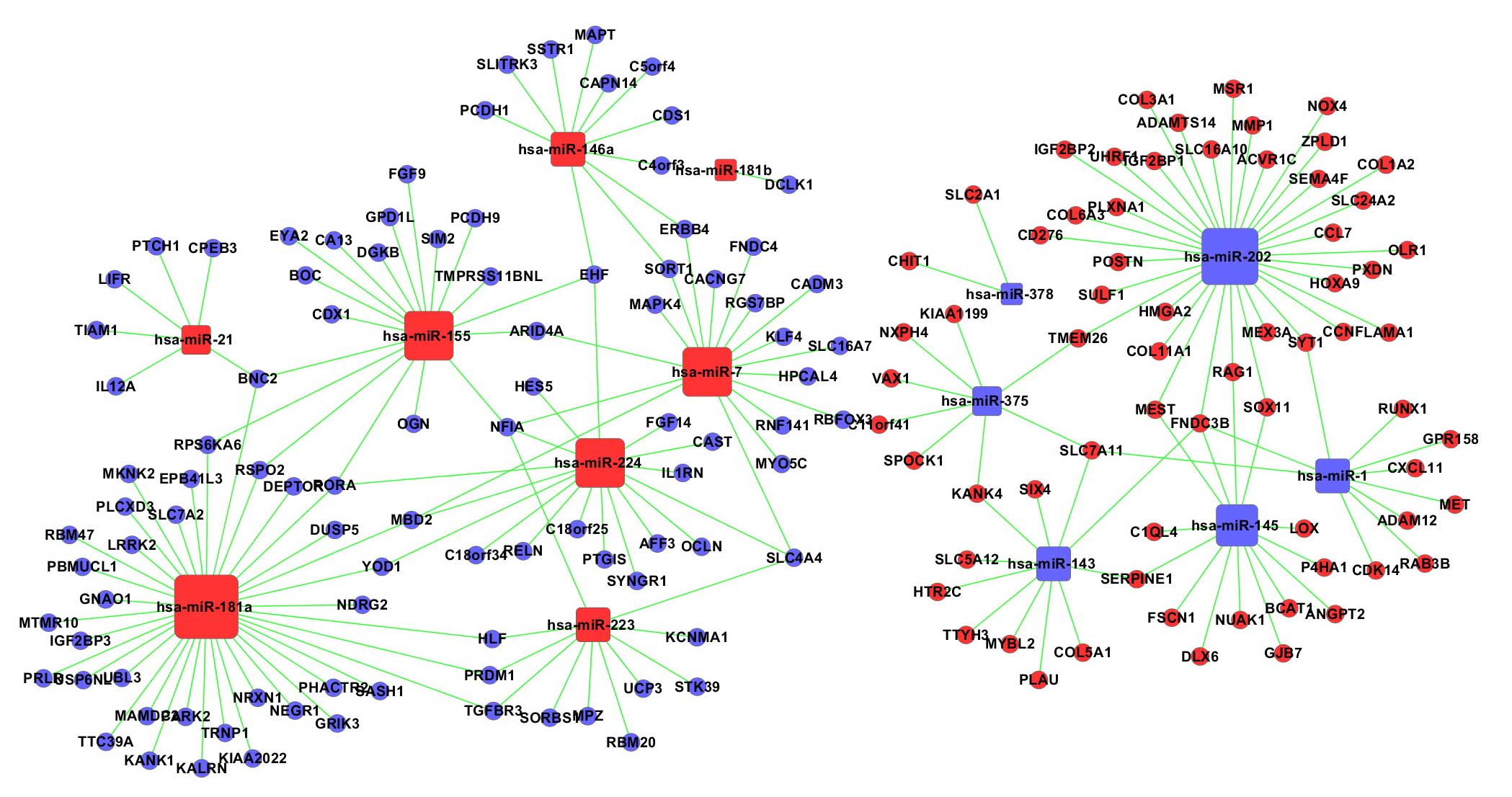
Friedman RC, Farh KK, Burge CB and Bartel
DP: Most mammalian mRNAs are conserved targets of microRNAs. Genome
Res. 19:92–105. 2009. View Article : Google Scholar :
|
|
23
|
Grimson A, Farh KK, Johnston WK,
Garrett-Engele P, Lim LP and Bartel DP: MicroRNA targeting
specificity in mammals: determinants beyond seed pairing. Mol Cell.
27:91–105. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Garcia DM, Baek D, Shin C, Bell GW,
Grimson A and Bartel DP: Weak seed-pairing stability and high
target-site abundance decrease the proficiency of lsy-6 and other
microRNAs. Nat Struct Mol Biol. 18:1139–1146. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kuhn RM, Haussler D and Kent WJ: The UCSC
genome browser and associated tools. Brief Bioinform. 14:144–161.
2013. View Article : Google Scholar :
|
|
26
|
Chen X and Wang L: Integrating biological
knowledge with gene expression profiles for survival prediction of
cancer. J Comput Biol. 16:265–278. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lu J, Getz G, Miska EA, et al: MicroRNA
expression profiles classify human cancers. Nature. 435:834–838.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Lovering RC, Camon EB, Blake JA and Diehl
AD: Access to immunology through the Gene Ontology. Immunology.
125:154–160. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Matsushima K, Isomoto H, Yamaguchi N, et
al: MiRNA-205 modulates cellular invasion and migration via
regulating zinc finger E-box binding homeobox 2 expression in
esophageal squamous cell carcinoma cells. J Transl Med. 9:302011.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Omoto S and Fujii YR: Regulation of human
immunodeficiency virus 1 transcription by nef microRNA. J Gen
Virol. 86:751–755. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang N, Li Y, Zhou RM, et al:
Hsa-miR-196a2 functional SNP is associated with the risk of ESCC in
individuals under 60 years old. Biomarkers. 19:43–48. 2014.
View Article : Google Scholar
|
|
32
|
Yang M, Liu R, Li X, et al: miRNA-183
suppresses apoptosis and promotes proliferation in esophageal
cancer by targeting PDCD4. Mol Cells. 37:873–880. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Yuan JM, Mao WM, Luo J, Peng BF, Zheng ZG
and Ling ZQ: Effect of miRNA-106a expression on the prognosis of
patients with esophageal squamous cell carcinoma. Zhonghua Zhong
Liu Za Zhi. 35:590–594. 2013.In Chinese. PubMed/NCBI
|
|
34
|
Wang XC, Zhang ZB, Wang YY, et al:
Increased miRNA-22 expression sensitizes esophageal squamous cell
carcinoma to irradiation. J Radiat Res. 54:401–408. 2013.
View Article : Google Scholar :
|
|
35
|
Watanabe N, Takaoka M, Sakurama K, et al:
Dual tyrosine kinase inhibitor for focal adhesion kinase and
insulin-like growth factor-I receptor exhibits anticancer effect in
esophageal adeno-carcinoma in vitro and in vivo. Clin Cancer Res.
14:4631–4639. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Ying GX, Wen Sheng LI, Xia ZL and Tao WH:
CagA+H. pylori filtrate induces cytokine IL-8 secretion by
esophageal squamous carcinoma EC 109 cells via a p38 pathway.
Indian J Pathol Microbiol. 57:13–18. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Chan SH, Yee Ko JM, Chan KW, et al: The
ECM protein LTBP-2 is a suppressor of esophageal squamous cell
carcinoma tumor formation but higher tumor expression associates
with poor patient outcome. Int J Cancer. 129:565–573. 2011.
View Article : Google Scholar
|
|
38
|
Zhang X, Nie Y, Li X, et al: MicroRNA-181a
Functions as an Oncomir in Gastric Cancer by Targeting the Tumour
Suppressor Gene ATM. Pathol Oncol Res. 2014. View Article : Google Scholar
|
|
39
|
Zhang Y, Zheng D, Xiong Y, et al: miR-202
suppresses cell proliferation in human hepatocellular carcinoma by
downregu-lating LRP6 post-transcriptionally. FEBS Lett.
588:1913–1920. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Zhang J, Cheng C, Yuan X, He JT, Pan QH
and Sun FY: microRNA-155 acts as an oncogene by targeting the tumor
protein 53-induced nuclear protein 1 in esophageal squamous cell
carcinoma. Int J Clin Exp Pathol. 7:602–610. 2014.PubMed/NCBI
|
|
41
|
Cai C, Rajaram M, Zhou X, et al:
Activation of multiple cancer pathways and tumor maintenance
function of the 3q amplified oncogene FNDC3B. Cell Cycle.
11:1773–1781. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
42
|
Akagi T, Ito T, Kato M, et al: Chromosomal
abnormalities and novel disease-related regions in progression from
Barrett’s esophagus to esophageal adenocarcinoma. Int J Cancer.
125:2349–2359. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Kasap E, Boyacioglu SO, Korkmaz M, et al:
Aurora kinase A (AURKA) and never in mitosis gene A-related kinase
6 (NEK6) genes are upregulated in erosive esophagitis and
esophageal adenocarcinoma. Exp Ther Med. 4:33–42. 2012.PubMed/NCBI
|
|
44
|
Berger J and Bird A: Role of MBD2 in gene
regulation and tumorigenesis. Biochem Soc Trans. 33:1537–1540.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Yuan K, Xie K, Fox J, et al: Decreased
levels of miR-224 and the passenger strand of miR-221 increase
MBD2, suppressing maspin and promoting colorectal tumor growth and
metastasis in mice. Gastroenterology. 145:853–864. e8592013.
View Article : Google Scholar : PubMed/NCBI
|