|
1
|
Maciotta S, Meregalli M and Torrente Y:
The involvement of microRNAs in neurodegenerative diseases. Front
Cell Neurosci. 7:2652013. View Article : Google Scholar
|
|
2
|
Rao AT, Degnan AJ and Levy LM: Genetics of
Alzheimer disease. AJNR Am J Neuroradiol. 35:457–458. 2014.
View Article : Google Scholar
|
|
3
|
Davinelli S, Calabrese V, Zella D and
Scapagnini G: Epigenetic nutraceutical diets in Alzheimer’s
disease. J Nutr Health Aging. 18:800–805. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wirth M, Villeneuve S, La Joie R, Marks SM
and Jagust WJ: Gene-environment interactions: Lifetime cognitive
activity, APOE genotype, and β-amyloid burden. J Neurosci.
34:8612–8617. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Hébert SS, Horré K, Nicolaï L, et al: Loss
of microRNA cluster miR-29a/b-1 in sporadic Alzheimer’s disease
correlates with increased BACE1/beta-secretase expression. Proc
Natl Acad Sci USA. 105:6415–6420. 2008. View Article : Google Scholar
|
|
6
|
Selbach M, Schwanhäusser B, Thierfelder N,
et al: Widespread changes in protein synthesis induced by
microRNAs. Nature. 455:58–63. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Fineberg SK, Kosik KS and Davidson BL:
MicroRNAs potentiate neural development. Neuron. 64:303–309. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hampel H, Shen Y, Walsh DM, et al:
Biological markers of amyloid beta-related mechanisms in
Alzheimer’s disease. Exp Neurol. 223:334–346. 2010. View Article : Google Scholar :
|
|
9
|
Wang WX, Rajeev BW, Stromberg AJ, et al:
The expression of microRNA miR-107 decreases early in Alzheimer’s
disease and may accelerate disease progression through regulation
of beta-site amyloid precursor protein-cleaving enzyme 1. J
Neurosci. 28:1213–1223. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kocerha J, Kauppinen S and Wahlestedt C:
microRNAs in CNS disorders. Neuromolecular Med. 11:162–172. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
National Research Council (US) Committee
for the Update of the Guide for the Care and Use of Laboratory
Animals. Guide for the Care and Use of Laboratory Animals. 8th
edition. National Academies Press; Washington DC, US: 2011
|
|
12
|
Livak KF and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
13
|
Decourt B, Walker A, Gonzales A, et al:
Can platelet BACE1 levels be used as a biomarker for Alzheimer’s
disease? Proof-of-concept study. Platelets. 24:235–238. 2013.
View Article : Google Scholar
|
|
14
|
Shaw LM, Vanderstichele H, Knapik-Czajka
M, et al: Alzheimer’s Disease Neuroimaging Initiative:
Cerebrospinal fluid biomarker signature in Alzheimer’s disease
neuroimaging initiative subjects. Ann Neurol. 65:403–413. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Bibl M, Esselmann H, Lewczuk P, et al:
Combined Analysis of CSF Tau, Aβ42, Aβ1-42% and AβCombined Analysis
of CSF Tau, Aβ42, Aβ1-42% and Aβ1-40% in Alzheimer’s Disease,
Dementia with Lewy Bodies and Parkinson’s Disease Dementia. Int J
Alzheimers Dis pii. 7615712010.
|
|
16
|
Müller M, Kuiperij HB, Claassen JA,
Küsters B and Verbeek MM: MicroRNAs in Alzheimer’s disease:
Differential expression in hippocampus and cell-free cerebrospinal
fluid. Neurobiol Aging. 35:152–158. 2014. View Article : Google Scholar
|
|
17
|
Bettens K, Brouwers N, Engelborghs S, et
al: APP and BACE1 miRNA genetic variability has no major role in
risk for Alzheimer disease. Hum Mutat. 30:1207–1213. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Marques SC, Lemos R, Ferreiro E, et al:
Epigenetic regulation of BACE1 in Alzheimer’s disease patients and
in transgenic mice. Neuroscience. 220:256–266. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Cattabeni F, Colciaghi F and Di Luca M:
Platelets provide human tissue to unravel pathogenic mechanisms of
Alzheimer disease. Prog Neuropsychopharmacol Biol Psychiatry.
28:763–770. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
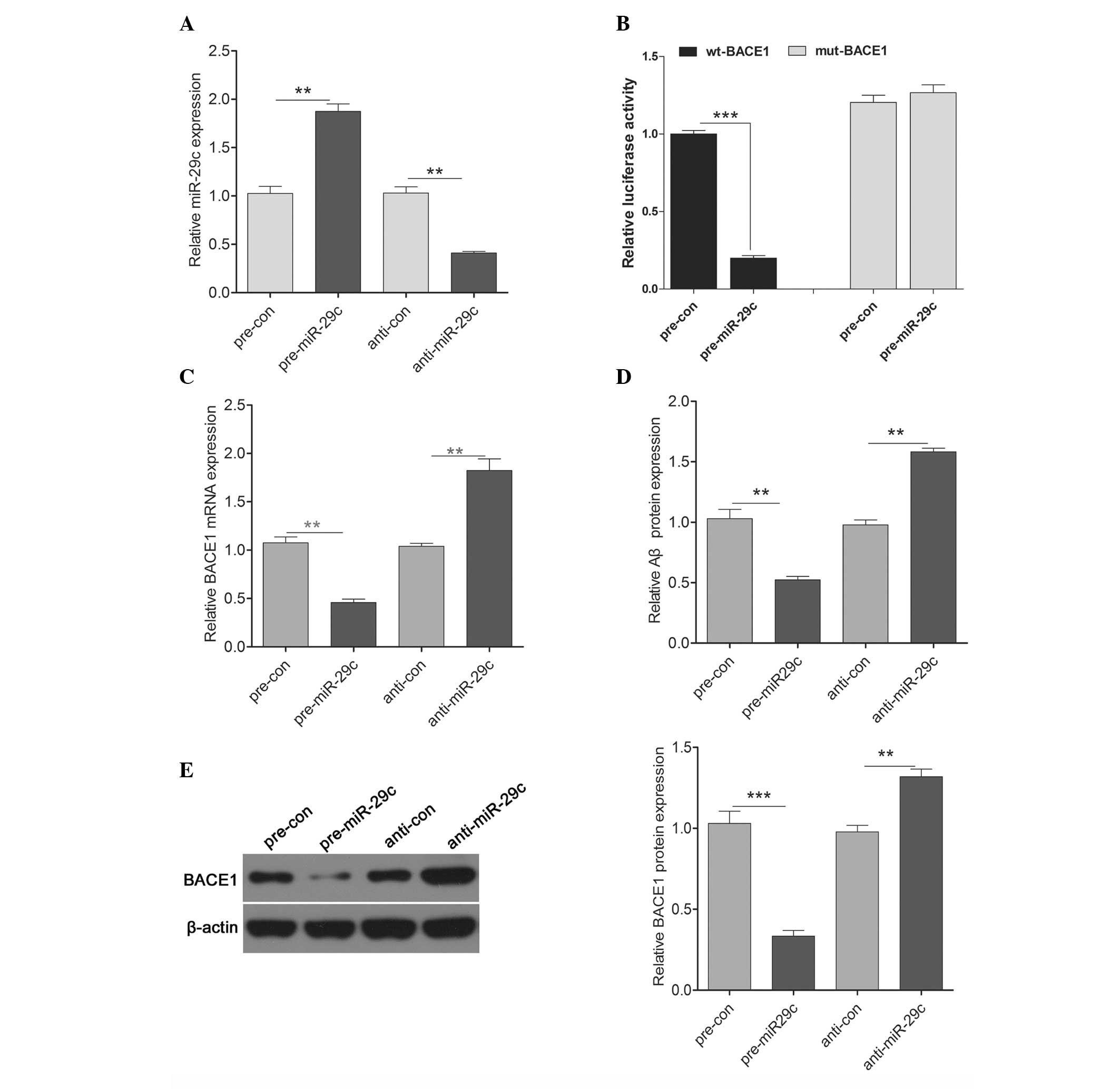
Zong Y, Wang H, Dong W, et al: miR-29c
regulates BACE1 protein expression. Brain Res. 1395:108–115. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cogswell JP, Ward J, Taylor IA, et al:
Identification of miRNA changes in Alzheimer’s disease brain and
CSF yields putative biomarkers and insights into disease pathways.
J Alzheimers Dis. 14:27–41. 2008.PubMed/NCBI
|
|
22
|
Moon M, Cha MY and Mook-Jung I: Impaired
hippocampal neurogenesis and its enhancement with ghrelin in 5XFAD
mice. J Alzheimers Dis. 41:233–241. 2014.PubMed/NCBI
|
|
23
|
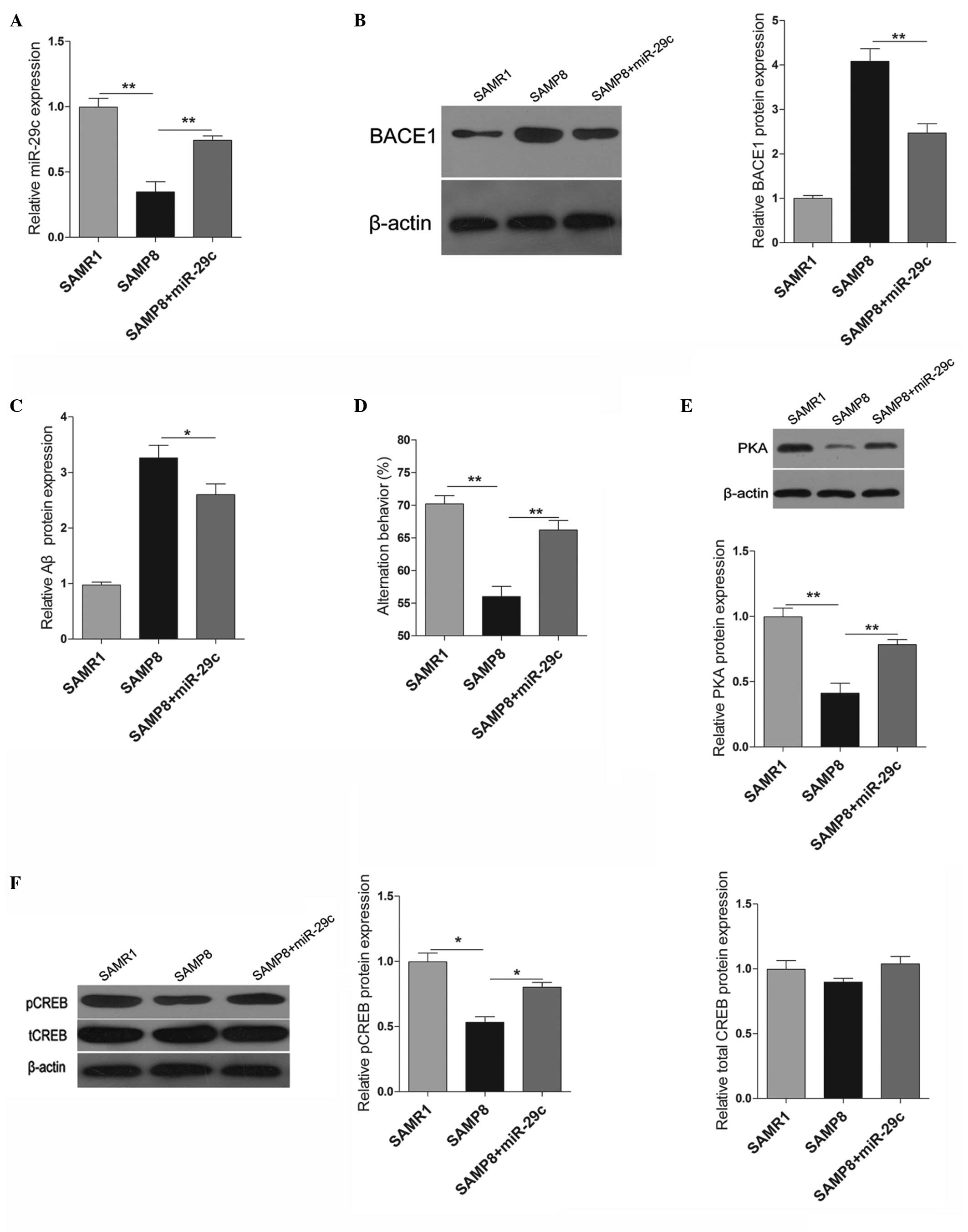
Shi YQ, Huang TW, Chen LM, et al:
Ginsenoside Rg1 attenuates amyloid-beta content, regulates PKA/CREB
activity and improves cognitive performance in SAMP8 mice. J
Alzheimers Dis. 19:977–989. 2010.
|
|
24
|
Valera E, Sánchez-Martín FJ,
Ferrer-Montiel AV, Messeguer A and Merino JM: NMDA-induced
neuroprotection in hippo-campal neurons is mediated through the
protein kinase A and CREB (cAMP-response element-binding protein)
pathway. Neurochem Int. 53:148–154. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zeitlin R, Patel S, Burgess S, Arendash GW
and Echeverria V: Caffeine induces beneficial changes in PKA
signaling and JNK and ERK activities in the striatum and cortex of
Alzheimer’s transgenic mice. Brain Res. 1417:127–136. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Sierksma AS, Rutten K, Sydlik S, et al:
Chronic phosphodiesterase type 2 inhibition improves memory in the
APPswe/PS1dE9 mouse model of Alzheimer’s disease.
Neuropharmacology. 64:124–136. 2013. View Article : Google Scholar
|
|
27
|
Chen Y, Huang X, Zhang YW, et al:
Alzheimer’s β-secretase (BACE1) regulates the cAMP/PKA/CREB pathway
independently of β-amyloid. J Neurosci. 32:11390–11395. 2012.
View Article : Google Scholar : PubMed/NCBI
|