Introduction
Alzheimer's disease (AD) is a chronic
neurodegenerative disease characterized by a progressive decline in
cognitive and memory function (1).
The prevalence of AD was 35.6 million globally in 2010, and this
number is expected to double by 2030. AD has become one of the most
common forms of dementia in the elderly worldwide (2), exerting a huge burden on families and
society.
AD is a complex disease that is influenced by
environmental and genetic factors (3). Heritability studies have shown that
~70% of AD risk may be attributed to genetic factors (4). Epigenetic modifications are thought
to link environmental and genetic factors (5–7). DNA
methylation, a type of epigenetic modification, has been shown to
have a significant role in the etiology of several diseases,
including leukemia (8), type 2
diabetes (9,10), essential hypertension (9), coronary heart disease (11,12),
schizophrenia (13) and AD
(14,15). DNA methylation often occurs in a
CpG dinucleotide context (16).
The CpG islands of a promoter are CpG-rich regions that are
predominantly hypomethylated (17). Alterations in promoter methylation
often affect gene expression (7,18–20).
Dopamine receptor D4 (DRD4) encodes the D4
subtype of the dopamine receptor (21). An increasing amount of evidence
supports a link between DRD4 and AD (22). DRD4 polymorphism has been
observed to be significantly associated with AD (22) and DRD4 promoter methylation
(23). DRD4 gene
hypermethylation has been demonstrated to increase DRD4 gene
expression and the risk of schizophrenia in males (13). A hypermethylated DRD4
promoter has also been identified in patients with alcohol
addiction (24).
Although DNA methylation levels vary among tissues,
independent studies have revealed that the DNA methylation patterns
of multiple loci in peripheral blood were similar to those in brain
tissues (25–28). In light of these previous findings,
the goal of the present study was to evaluate the contribution of
DRD4 promoter methylation to AD using peripheral blood as a
surrogate of brain tissue.
Materials and methods
Sample collection
A total of 46 sporadic AD patients and 61 matched
controls were selected from Ningbo No. 1 Hospital (Ningbo, China)
and Ningbo Kangning Hospital (Ningbo, China). AD was diagnosed by
two professional neurological physicians (CZ and ZQ) according to
National Institute of Neurological and Communicative Disorders and
Stroke-Alzheimer's Disease and Related Disorders Association
criteria from ICD-10 (29), on the
basis of medical and family histories, neurological examination,
blood studies, brain imaging studies, neuropsychological testing
and cognitive screening tests. All controls were free of any type
of physical or mental disorder. In the current study, two kinds of
drugs (Exelon and Aricept) were used for AD patients. Exelon
(Novartis Farmaceutica S.A, Spain) was administered at 1.5 mg twice
per day with morning and evening meals. Following a minimum of two
weeks of treatment, if the initial dose was well-tolerated, it was
increased to 3 mg twice per day. Initial treatment with Aricept
(Eisai China Inc., Jiangsu, China) was 5 mg per day at bedtime. The
dose was increased to 10 mg per day after 4 to 6 weeks if the
response was not adequate. All individuals in the present study
were Han Chinese originating from Ningbo city in Eastern China.
Peripheral blood samples were collected in 3.2% sodium
citrate-treated tubes (Jiangsu Kailijian Medical Device Co., Ltd.,
Jiangsu, China) and then stored at −80°C. The study protocol was
approved by the Ethical Committee of Ningbo University (Ningbo,
China), Ningbo No.1 Hospital and Ningbo Kangning Hospital. Written
informed consent was obtained from all subjects or their
guardians.
Detection of biochemical factors
The serum concentrations of total protein (TP) and
albumin (ALB) were detected using the biuret (30) and bromocresol green methods
(31), respectively. Globulin
(GLB) was calculated as TP minus ALB. The concentrations of
glutamic-pyruvic transaminase, alkaline phosphatase and glutamic
oxalacetic transaminase were determined using the velocity method
(32,33). The levels of total bile acid and
homocysteine (Hcy) were measured using the cycling enzymatic method
(34,35). Plasma concentrations of blood
glucose, triglyceride, total cholesterol, carbamide (UREA),
creatinine (CRE) and uric acid were determined using the enzymatic
methods (36–41). The high-density lipoprotein
cholesterol level was determined using the one-step detection
method (42). The proportions of
apolipoprotein-A (ApoA) and apolipoprotein-B were measured by the
turbidimetry method (43,44). The concentrations of lipoprotein A
(Lp (a)) and C Reactive Protein (CRP) were detected using the
endpoint method (45) and latex
agglutination assay (46),
respectively. The apolipoprotein E levels were detected using the
immunoturbidimetric assay (47).
Bisulfite pyrosequencing assay
DNA extraction and consequent bisulfite
pyrosequencing assays were performed as described in our previous
studies (8,9,12,13,48,49).
PCR primers were designed using PyroMark Assay Design software
version 2 (Qiagen China Co., Ltd., Shanghai, China). Primer
sequences were 5′-biotin-GGGAGGTTTTGTTAGATATTAGGT-3′ for the
forward primer; 5′-CCACCCTAAACCCAATATTTACTCATCTTA-3′ for the
reverse primer; and 5′-ACCAAACCAAACCCT-3′ for the sequencing
primer.
Statistical analyses
Statistical analyses were performed using SPSS
software version 16.0 (SPSS, Inc., Chicago, IL, USA), and a
P<0.05 was considered to indicate a statistically significant
difference. The two independent samples t test was used to
compare the differences in the mean values of continuous variables
between AD patients and controls. Pearson's correlation test was
used to assess the associations between DRD4 methylation and
the metabolic characteristics of the AD subjects. Bonferroni
correction was used to adjust the results. Power analysis was
estimated with the Power and Sample Size Calculation software
version 3.043 (http://biostat.mc.vanderbilt.edu/wiki/Main/PowerSampleSize).
In the present study, according to the online power calculator, α
was the type I error probability for a two-sided test; n was the
sample number of AD patients; δ was the difference in population
means, which was equal to the mean methylation levels in AD
patients minus those in normal controls; σ was the within-group
standard deviation; and m was the ratio of control to experimental
patients.
Results
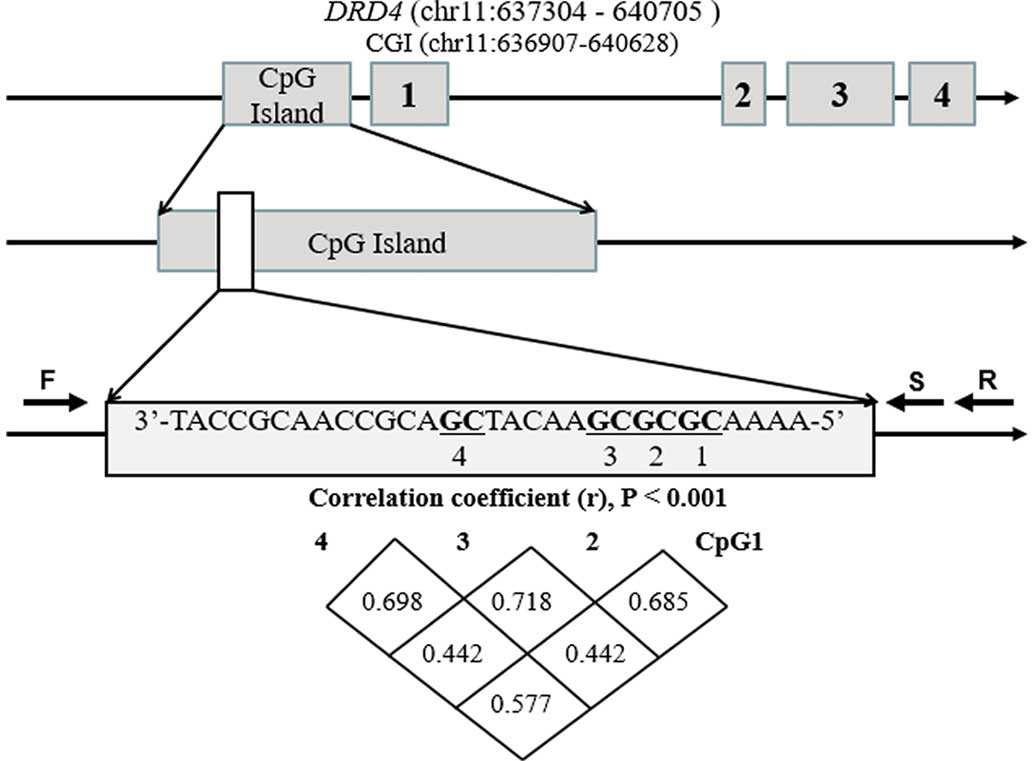
The selected promoter fragment in the
current methylation assay
The bisulfite pyrosequencing assay was performed on
the CpG island region (chr11:636877-637167) of the DRD4
promoter. As shown in Fig. 1, a
total of four CpG sites were measured. As there was a significant
correlation among the methylation levels of the four CpG sites
(Fig. 1; r=0.442, P<0.001), the
mean DNA methylation of the four CpGs was also evaluated in the
subsequent analyses.
Association tests between clinical
phenotypes with AD
The present study involved a total of 46 AD patients
and 61 controls. As shown in Table
I, among the 22 phenotypes, the plasma levels of ApoA, Lp (a),
Hcy and CRP were observed to significantly differ between AD
patients and controls (Table I;
ApoA, P=0.011; Lp (a), P<0.001; Hcy, P=0.046; CRP, P=0.016). A
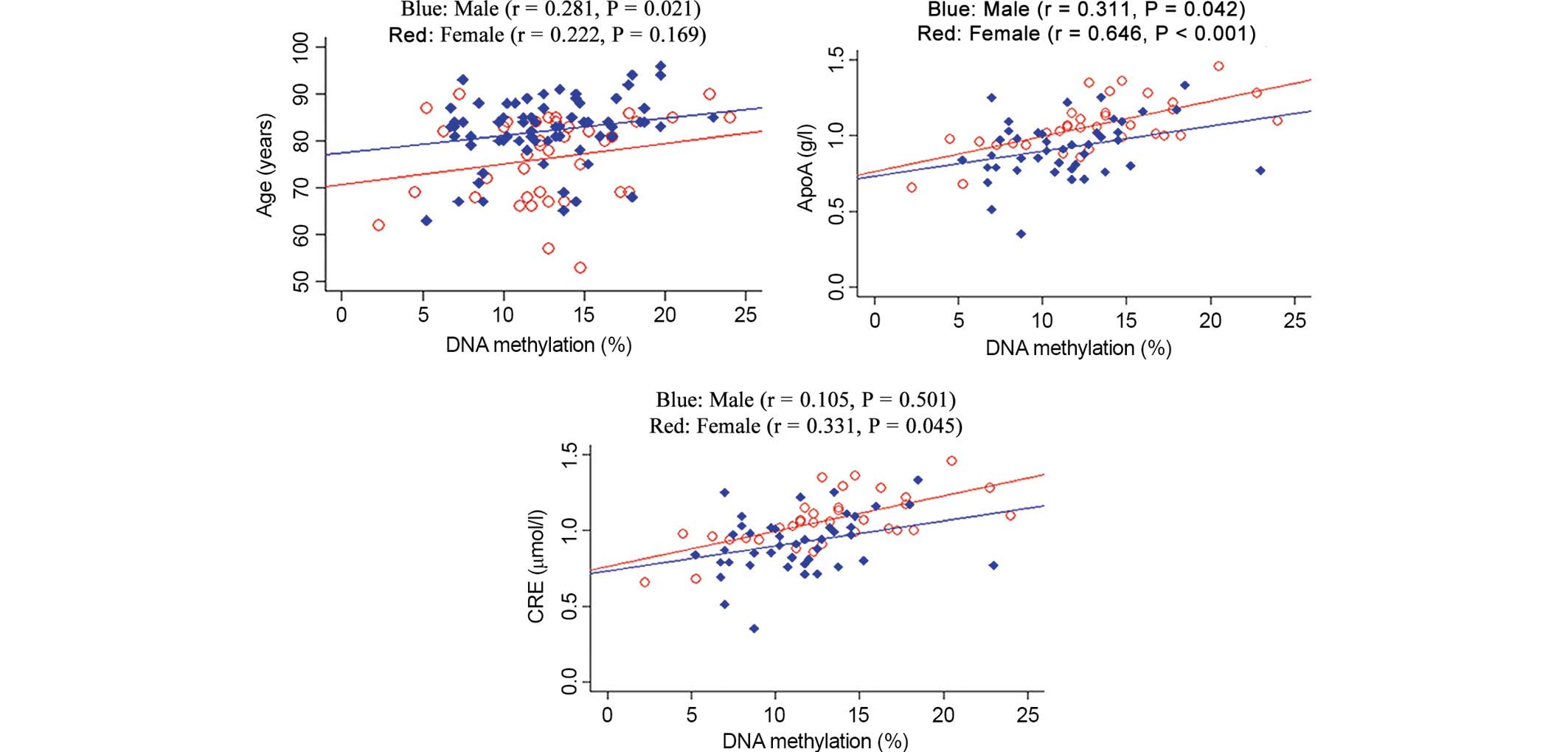
significant male-specific association was identified between the
mean DRD4 methylation and ApoA and level (Fig. 2; ApoA, P=0.042). A significant
female-specific association was observed between the average
DRD4 methylation and several phenotypes, including ApoA and
CRE levels (Fig. 2; ApoA,
P<0.001; CRE, P=0.045). Age is a well-known risk factor for AD,
therefore the association between age and DRD4 methylation
was tested. The results revealed an association between age and
DRD4 methylation (Fig. 2;
males: r=0.281, P=0.021; females: r=0.222, P=0.169).
 | Table ICharacteristics of the 107
subjects. |
Table I
Characteristics of the 107
subjects.
| Characteristic | Patients (n=46),
mean ± SD | Controls (n=61),
mean ± SD | P-value |
|---|
| Age, years | 80.67±9.20 | 79.54±7.87 | 0.495 |
| TP, g/l | 68.79±6.91 | 65.48±9.45 | 0.099 |
| ALB, g/l | 38.32±3.83 | 36.86±3.61 | 0.090 |
| GLB, g/l | 30.46±5.32 | 29.55±4.58 | 0.418 |
| A/G | 1.29±0.22 | 1.28±0.20 | 0.768 |
| ALT, U/l | 13.87±10.68 | 18.20±13.46 | 0.193a |
| ALP, U/l | 78.00±24.30 | 96.82±63.34 | 0.113a |
| TBA,
µmol/l | 6.91±3.84 | 6.07±5.86 | 0.503 |
| AST, U/l | 20.52±7.22 | 23.58±11.70 | 0.258a |
| Glu, mmol/l | 5.23±1.58 | 5.53±2.71 | 0.444b |
| TG, mmol/l | 1.35±0.76 | 1.42±0.98 | 0.896b |
| TC, mmol/l | 4.48±1.04 | 4.28±1.22 | 0.378 |
| HDL-C, mmol/l | 1.12±0.27 | 1.04±0.30 | 0.118 |
| ApoA, g/l | 1.06±0.21 | 0.94±0.18 | 0.011c |
| ApoB, g/l | 0.66±0.19 | 0.73±0.25 | 0.194 |
| Lp(a), g/l | 184.86±233.63 | 34.86±27.32 | <0.001a,c |
| ApoE, mg/l | 37.73±17.44 | 36.69±10.37 | 0.800 |
| UREA, mmol/l | 7.77±10.00 | 6.45±3.45 | 0.804b |
| CRE,
µmol/l | 82.73±47.25 | 78.52±30.04 | 0.626 |
| UA,
µmol/l | 309.93±106.30 | 308.88±112.75 | 0.967 |
| Hcy,
µmol/l | 19.76±10.82 | 17.67±20.84 | 0.046a,c |
| CRP, mg/l | 6.20±11.72 | 15.00±26.21 | 0.016a,c |
Association tests of DRD4 methylation
with AD
Significantly increased DRD4 methylation
levels were observed in AD patients compared with controls
(Table II; CpG1, P=0.001; CpG2,
P=0.013; CpG3, P=0.001; CpG4, P<0.001; mean CpG1-4 methylation,
P<0.001). Among 23 male patients and 45 male controls, elevated
methylation levels of four CpG sites of the DRD4 promoter
were observed following breakdown analysis by gender (Table II; CpG1, P=0.013; CpG2, P=0.012;
CpG3, P<0.001; CpG4, P<0.001; mean CpG1-4 methylation,
P<0.001). As shown in Table
II, the power was sufficient in the overall (power=0.978) and
male-based subgroup (power=0.976) case-control comparisons;
however, the power was only 0.405 in the female-based subgroup
analysis, suggesting that the negative association in the female
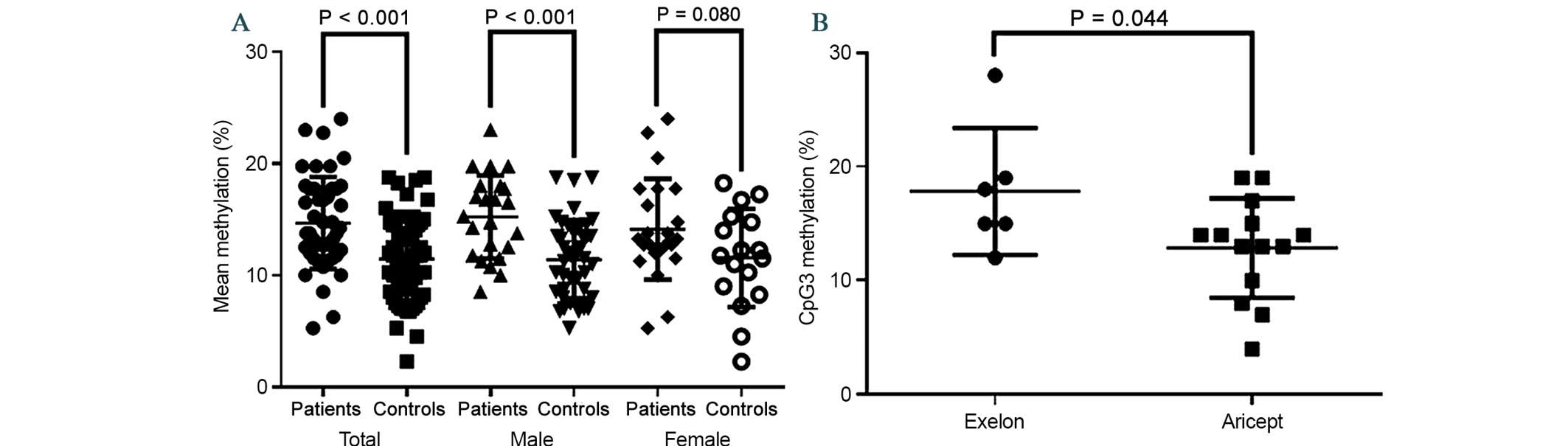
subgroup may be due to a lack of power. A consequent breakdown
analysis by gender revealed that the above significant association
of DRD4 methylation with AD existed only in males (Fig. 3A; males, P<0.001; females,
P=0.080).
 | Table IIComparison of DRD4 DNA
methylation levels between AD patients and controls. |
Table II
Comparison of DRD4 DNA
methylation levels between AD patients and controls.
A, All patients
|
|---|
| Characteristic,
% | Patients, mean ± SD
(n=46) | Controls, mean ± SD
(n=62) | P-value | Power |
|---|
| CpG1 | 16.50±5.14 | 13.13±4.67 |
0.001a,b | 0.916 |
| CpG2 | 12.26±4.84 | 10.13±3.92 |
0.013a,b | 0.610 |
| CpG3 | 14.98±5.27 | 11.59±4.59 |
0.001a,b | 0.906 |
| CpG4 | 14.96±4.22 | 10.92±4.90 | <0.001a,b | 0.987 |
| Mean
methylation | 14.67±4.12 | 11.44±3.72 | <0.001a,b | 0.978 |
B, Male patients
|
|---|
| Characteristic,
% | Patients, mean ± SD
(n=46) | Controls, mean ± SD
(n=62) | P-value | Power |
|---|
| CpG1 | 16.74±4.92 | 13.68±4.55 |
0.013a,b | 0.666 |
| CpG2 | 12.87±3.94 | 10.39±3.62 |
0.012a,b | 0.676 |
| CpG3 | 15.61±4.35 | 11.16±4.27 | <0.001a,b | 0.975 |
| CpG4 | 15.65±4.60 | 10.36±4.42 | <0.001a,b | 0.992 |
| Mean
methylation | 15.22±3.72 | 11.40±3.47 | <0.001a,b | 0.976 |
C, Female patients
|
|---|
| Characteristic,
% | Patients, mean ± SD
(n=46) | Controls, mean ± SD
(n=62) | P-value | Power |
|---|
| CpG1 | 16.26±5.45 | 11.71±4.82 |
0.009a,b | 0.720 |
| CpG2 | 11.65±5.63 |
9.47±4.67 | 0.201 | 0.212 |
| CpG3 | 14.35±6.10 | 12.71±5.31 | 0.380 | 0.125 |
| CpG4 | 14.26±3.78 | 12.35±5.88 | 0.220 | 0.162 |
| Mean
methylation | 14.13±4.51 | 11.56±4.40 | 0.080 | 0.405 |
DRD4 methylation levels in AD patients
with various treatments
Exelon and Aricept are commonly used
acetylcholinesterase inhibitors in the treatment of AD (48). Both drugs aim to enhance
cholinergic neurotransmission in specific parts of the brain and to
improve the clinical symptoms of AD (50). The results of the present study
revealed that the patients using Exelon-treated patients had
significantly higher CpG3 methylation levels compared with
Aricept-treated patients Aricept (Fig.
3B; P=0.044).
Discussion
The present study evaluated the levels of
DRD4 promoter methylation in AD patients and matched
controls to clarify the contribution of DRD4 promoter
methylation to AD risk. It was observed that the DRD4
promoter methylation levels in AD patients were significantly
higher than those in controls. In a breakdown analysis by gender,
there were varying associations of methylation status in males and
females. Positive results were identified for all four CpG sites
observed in males. Positive correlations between DRD4
methylation and age, as well as DRD4 methylation and ApoA
level, were also observed in males. In addition, clear positive
correlations were observed between DRD4 methylation and ApoA
and between DRD4 methylation and CRE in females.
Furthermore, varying methylation levels were observed in patients
who used Exelon and those who used Aricept.
Anomalies in dopaminergic transmission may lead to
the disturbance of synaptic plasticity and advanced cognitive
behavior (22). A male-specific
association between DRD4 methylation and schizophrenia has
been reported in Han Chinese (13). As a significant receptor of
dopamine, DRD4 is pivotal to the development of AD (51). In the present study, significantly
hypermethylated DRD4 promoters were observed in AD patients
compared with controls. These results indicated that DRD4
may be involved in the progression of AD. Understanding the DNA
methylation changes in the DRD4 promoter may aid in
understanding the pathological mechanisms of AD. This information
may also provide insight into the function of AD-associated genes,
including DRD4, and help identify novel targets for
therapeutic strategies to reverse the promoter methylation of
DRD4.
Gender differences in AD have been widely
documented. Females have a higher risk of AD at all ages, and the
age-adjusted odds ratio for females has been shown to be 3.1
between AD patients and controls (52). Gender-specific DNA methylation
exists in mice (53) and humans
(12,54). DRD4 methylation research in
another nervous system disorder, schizophrenia, has also identified
a male-specific significant association (13). Significant differences were
reported in all CpG sites observed in the present study in males,
but a significant difference was observed in only CpG1 in females.
These phenomena of varying methylation statuses in the DRD4
promoter also provide insight into gender differences in AD. The
results of the present study support the idea that gender
differences should be considered when establishing a clinical
treatment plan for AD.
Furthermore, a significant association between DNA
methylation and age in males but not in females was observed in the
present study. Age is considered a major risk factor for AD
(54). A previous study reported
that DNA methylation is dynamically regulated in the human cerebral
cortex throughout life, involves differentiated neurons and affects
a substantial proportion of genes predominantly by an
age-associated increase (55).
Detailed descriptions of associations between DNA methylation and
age in various gender subgroups require additional study.
A total of 22 phenotypes were analyzed in the
present study. It is well-known that ApoA1, an increase in which
leads to an increased risk of AD, is the major apolipoprotein
constituent of high-density lipoprotein, and Apo A1 has been
observed to affect brain cholesterol metabolism and angiogenesis
(56). In an earlier study, serum
ApoA concentration was shown to have a high correlation with the
severity of AD (57).
Significantly increased levels of ApoA1 were observed in AD
patients compared with controls in the present study. Based on
previous findings, it was speculated that Lp (a) may participate in
the progression of dementia (58)
and AD (59). The present study
showed that AD patients had higher levels of Lp (a), which was
similar to the results of a previous cross-sectional study
(59). Hcy levels, another AD
factor that may induce amyloid β accumulation, synaptic dysfunction
and memory impairment, were significantly different between AD
patients and matched controls in the present study (60–63).
It has been reported that CRP participates in the systemic response
to inflammation (64). Additional
neuropathological studies have shown that CRP is associated with
neurofibrillary tangles (65) and
senile plaques (66) in AD brain
tissue. The subjects with AD had significantly lower levels of
plasma CRP than control subjects in the present study, which was
consistent with the results of a previous study (64).
Aricept is an acetylcholinesterase inhibitor used
for the symptomatic treatment of AD (67). Furthermore, Aricept is a
high-affinity sigma-1 receptor antagonist, which has been
investigated as a potential disease-modifying agent for several CNS
disorders (64). Exelon is an oral
drug approved by the US Food and Drug Administration for the
treatment of AD (67).
Significantly higher levels of DNA methylation were reported at the
CpG3 site in the patients who were treated with Exelon compared
with those treated with Aricept. This finding indicates that
varying DNA methylation effects were produced by Exelon and
Aricept. However, additional research is required to uncover the
detailed association between DNA methylation, Exelon and
Aricept.
The current study has a number of limitations.
Firstly, the sample size was small, which may have influenced the
results, particularly for the gender-stratified association test of
DRD4 methylation with AD. In addition, the levels of DNA
methylation of the four CpGs that were tested cannot represent the
entire influence of DRD4. Additional studies investigating
the DRD4 promoter or gene body regions are required.
Furthermore, samples were mainly from the elderly, who may have
underlying diseases. In other words, there may be certain unknown
or potential risk factors for AD present in this study. Also, the
DNA methylation level of DRD4 was measured in peripheral
blood only, which may not be an accurate reflection of the
situation in the brain tissue. Additional comprehensive studies on
the association of DRD4 methylation within brain tissues and
peripheral blood are required. Finally, four CpG positions were
assessed per pyrosequencing read. Certain P-values may not retain
their significance after being corrected for this number of CpG
sites. The possibility that the present positive findings arose by
chance cannot be excluded.
In conclusion, the present study supports
DRD4 promoter hypermethylation as a risk factor for AD in
males. Additionally, positive associations between DRD4
methylation and age, as well as DRD4 methylation and ApoA
levels, were observed in males. Disparate DRD4 methylation
levels were reported for patients taking various drugs: Patients
taking Exelon appear to have higher levels than those taking
Aricept. The results of the present study may contribute to an
improved understanding of the molecular mechanisms underlying the
pathophysiology of AD.
Acknowledgments
The present research was supported by grants from
the National Natural Science Foundation of China (grant nos.
81371469, 81471938 and U1503223), the Natural Science Foundation of
Zhejiang Province (grant no. Y15H090032), Ningbo International
Science and Technology Cooperation Project (grant no. 2014D10019),
the Public Technology Research and Social Development Project of
Zhejiang Province (grant no. 2015C33155), the Disciplinary Project
of Ningbo University (grant no. B01350104900) and K. C. Wong Magna
Fund in Ningbo University.
References
|
1
|
Wilson RS, Barnes LL, Mendes de Leon CF,
Aggarwal NT, Schneider JS, Bach J, Pilat J, Beckett LA, Arnold SE,
Evans DA and Bennett DA: Depressive symptoms, cognitive decline,
and risk of AD in older persons. Neurology. 59:364–370. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Querfurth HW and LaFerla FM: Alzheimer's
disease. N Engl J Med. 362:329–344. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kubota T: Epigenome: What we learned from
Rett syndrome, a neurological disease caused by mutation of a
methyl-CpG binding protein. Rinsho Shinkeigaku. 53:1339–1341.
2013.In Japanese. View Article : Google Scholar
|
|
4
|
Alonso Vilatela ME, Lopez-Lopez M and
Yescas-Gomez P: Genetics of Alzheimer's disease. Genetics of
Alzheimer's disease. 43:622–631. 2012.
|
|
5
|
Tang LL, Liu Q, Bu SZ, Xu LT, Wang QW, Mai
YF and Duan SW: The effect of environmental factors and DNA
methylation on type 2 diabetes mellitus. Yi Chuan. 35:1143–1152.
2013.In Chinese. View Article : Google Scholar
|
|
6
|
Zhang W, Zheng Y and Hou L:
Pharmacogenomic discovery delineating the genetic basis of drug
response. Curr Genet Med Rep. 1:143–149. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wang T, Garcia JG and Zhang W: Epigenetic
regulation in particulate matter-mediated cardiopulmonary
toxicities: A systems biology perspective. Curr Pharmacogenomics
Person Med. 10:314–321. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Jiang D, Hong Q, Shen Y, Xu Y, Zhu H, Li
Y, Xu C, Ouyang G and Duan S: The diagnostic value of DNA
methylation in leukemia: A systematic review and meta-analysis.
PloS One. 9:e968222014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cheng J, Wang L, Xu L, Wang H, Liu P, Bu
S, Ye M, Zhang L, Wang Q and Duan S: Gender-dependent miR-375
promoter methylation and the risk of type 2 diabetes. Exp Ther Med.
5:1687–1692. 2013.PubMed/NCBI
|
|
10
|
Tang LL, Liu Q, Bu SZ, Xu LT, Wang QW, Mai
YF and Duan SW: The effect of environmental factors and DNA
methylation on type 2 diabetes mellitus. Yi Chuan. 35:1143–1152.
2013.In Chinese. View Article : Google Scholar
|
|
11
|
Xu L, Zheng D, Wang L, Jiang D, Liu H, Xu
L, Liao Q, Zhang L, Liu P, Shi X, et al: GCK gene-body
hypomethylation is associated with the risk of coronary heart
disease. Biomed Res Int. 2014:1517232014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jiang D, Zheng D, Wang L, Huang Y, Liu H,
Xu L, Liao Q, Liu P, Shi X, Wang Z, et al: Elevated PLA2G7 gene
promoter methylation as a gender-specific marker of aging increases
the risk of coronary heart disease in females. PloS One.
8:e597522013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Cheng J, Wang Y, Zhou K, Wang L, Li J,
Zhuang Q, Xu X, Xu L, Zhang K, Dai D, et al: Male-specific
association between dopamine receptor D4 gene methylation and
schizophrenia. PloS One. 9:e891282014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Hodgson N, Trivedi M, Muratore C, Li S and
Deth R: Soluble oligomers of amyloid-b cause changes in redox
state, DNA methylation and gene transcription by inhibiting EAAT3
mediated cysteine uptake. J Alzheimers Dis. 36:197–209. 2013.
|
|
15
|
Ji Huihui, Zhou Xiaohui, Chen Zhongming,
Li Ying, Zhou Dongsheng, Zhuo Renjie, Duan Shiwei and Wang Qinwen:
Research progress of DNA methylation in Alzheimer's disease. Chi J
Cell Biol. 36:1551–1559. 2014.
|
|
16
|
Al Akeel R: Role of epigenetic
reprogramming of host genes in bacterial pathogenesis. Saudi J Biol
Sci. 20:305–309. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Fatemi M, Pao MM, Jeong S, Gal-Yam EN,
Egger G, Weisenberger DJ and Jones PA: Footprinting of mammalian
promoters: Use of a CpG DNA methyltransferase revealing nucleosome
positions at a single molecule level. Nucleic Acids Res.
33:e1762005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yu Z, Kong Q and Kone BC: Aldosterone
reprograms promoter methylation to regulate αENaC transcription in
the collecting duct. Am J Physiol Renal Physiol. 305:F1006–F1013.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Moen EL, Zhang X, Mu W, Delaney SM, Wing
C, McQuade J, Myers J, Godley LA, Dolan ME and Zhang W: Genome-wide
variation of cytosine modifications between European and African
populations and the implications for complex traits. Genetics.
194:987–996. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Moen EL, Stark AL, Zhang W, Dolan ME and
Godley LA: The role of gene body cytosine modifications in MGMT
expression and sensitivity to temozolomide. Mol Cancer Ther.
13:1334–1344. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kirley A, Hawi Z, Daly G, McCarron M,
Mullins C, Millar N, Waldman I, Fitzgerald M and Gill M:
Dopaminergic system genes in ADHD: Toward a biological hypothesis.
Neuropsychopharmacology. 27:607–619. 2002.PubMed/NCBI
|
|
22
|
Lin WY, Wu BT, Lee CC, Sheu JJ, Liu SH,
Wang WF, Tsai CH, Liu HP and Tsai FJ: Association analysis of
dopaminergic gene variants (Comt, Drd4 And Dat1) with Alzheimer s
disease. J Biol Regul Homeost Agents. 26:401–410. 2012.PubMed/NCBI
|
|
23
|
Docherty SJ, Davis OS, Haworth CM, Plomin
R, D'Souza U and Mill J: A genetic association study of DNA
methylation levels in the DRD4 gene region finds associations with
nearby SNPs. Behav Brain Funct. 8:312012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang H, Herman AI, Kranzler HR, Anton RF,
Zhao H, Zheng W and Gelernter J: Array-based profiling of DNA
methylation changes associated with alcohol dependence. Alcohol
Clin Exp Res. 37(Suppl 1): E108–E115. 2013. View Article : Google Scholar
|
|
25
|
Mill J and Heijmans BT: From promises to
practical strategies in epigenetic epidemiology. Nat Rev Genet.
14:585–594. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Klengel T, Mehta D, Anacker C, Rex-Haffner
M, Pruessner JC, Pariante CM, Pace TW, Mercer KB, Mayberg HS,
Bradley B, et al: Allele-specific FKBP5 DNA demethylation mediates
gene-childhood trauma interactions. Nature Neurosci. 16:33–41.
2013. View Article : Google Scholar
|
|
27
|
Provencal N, Suderman MJ, Guillemin C,
Massart R, Ruggiero A, Wang D, Bennett AJ, Pierre PJ, Friedman DP,
Côté SM, et al: The signature of maternal rearing in the methylome
in rhesus macaque prefrontal cortex and T cells. J Neurosci.
32:15626–15642. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Liu X, Zheng Y, Zhang W, Zhang X,
Lioyd-Jones DM, Baccarelli AA, Ning H, Fornage M, He K, Liu K and
Hou L: Blood methylomics in response to arsenic exposure in a
low-exposed US population. J Expo Sci Environ Epidemiol.
24:145–149. 2014. View Article : Google Scholar
|
|
29
|
Fiedler U, Wiltfang J, Peters N and
Benninghoff J: Advances in the diagnostics of Alzheimer's disease.
Nervenarzt. 83:661–673. 2012.In German. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Doumas BT: Standards for total serum
protein assays-a collaborative study. Clin Chem. 21:1159–1166.
1975.PubMed/NCBI
|
|
31
|
McPherson IG and Everard DW: Serum albumin
estimation: Modification of the bromcresol green method. Clinic
Chim Acta. 37:117–121. 1972. View Article : Google Scholar
|
|
32
|
Yang X, Liu B, Sang Y, Yuan Y, Pu J, Liu
Y, Li Z, Feng J, Xie Y, Tang R, et al: Kinetic analysis of the
lactate-dehydrogenase-coupled reaction process and measurement of
alanine transaminase by an integration strategy. Anal Sci.
26:1193–1198. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Nath RL and Ghosh NK: A preliminary report
on the determination of the normal values of serum alkaline
phosphatase activity by velocity constant method. Bull Calcutta Sch
Trop Med. 10:71–72. 1962.PubMed/NCBI
|
|
34
|
Zhang GH, Cong AR, Xu GB, Li CB, Yang RF
and Xia TA: An enzymatic cycling method for the determination of
serum total bile acids with recombinant 3alpha-hydroxysteroid
dehydrogenase. Biochem Biophys Res Commun. 326:87–92. 2005.
View Article : Google Scholar
|
|
35
|
Roberts RF and Roberts WL: Performance
characteristics of a recombinant enzymatic cycling assay for
quantification of total homocysteine in serum or plasma. Clin Chim
Acta. 344:95–99. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Jaynes PK, Feld RD and Johnson GF: An
enzymic, reaction-rate assay for serum creatinine with a
centrifugal analyzer. Clin Chem. 28:114–117. 1982.PubMed/NCBI
|
|
37
|
Tabacco A, Meiattini F, Moda E and Tarli
P: Simplified enzymic/colorimetric serum urea nitrogen
determination. Clin Chem. 25:336–337. 1979.PubMed/NCBI
|
|
38
|
Whitlow K and Gochman N: Continuous-flow
enzymic method evaluated for measurement of serum triglycerides
with use of an improved lipase reagent. Clin Chem. 24:2018–2019.
1978.PubMed/NCBI
|
|
39
|
Knob M and Rosenmund H: Enzymic
determination of total serum cholesterol with centrifugal analyzers
(author's transl). Z Klin Chem Klin Biochem. 13:493–498. 1975.In
German. PubMed/NCBI
|
|
40
|
Hunziker P and Keller H: A mechanized
enzymic method for the determination of uric acid (author's
transl). Z Klin Chem Klin Biochem. 13:89–96. 1975.In German.
PubMed/NCBI
|
|
41
|
Asrow G: Semiautomated enzymic micro
methods for blood glucose and lactic acid on a single filtrate.
Anal Biochem. 28:130–138. 1969. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Egloff M, Léglise D, Duvillard L,
Steinmetz J, Boyer MJ, Ruelland A, Agher R and Beucler I:
Multicenter evaluation on different analyzers of three methods for
direct HDL-cholesterol assay. Ann Biol Clin (Paris). 57:561–572.
1999.
|
|
43
|
DaCol P and Kostner GM:
Immunoquantification of total apolipoprotein B in serum by
nephelometry: Influence of lipase treatment and detergents. Clin
Chem. 29:1045–1050. 1983.PubMed/NCBI
|
|
44
|
Girault A, Loiseau D and Girault M:
Quantitative determination of apolipoprotein A in human serum by
laser nephelometry. Ric Clin Lab. 11(Suppl 2): S19–S29. 1981.
|
|
45
|
Cazzolato G, Prakasch G, Green S and
Kostner GM: The determination of lipoprotein Lp(a) by rate and
endpoint nephelometry. Clin Chim Acta. 135:203–208. 1983.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Deyo RA, Pope RM and Persellin RH:
Interference by rheumatoid factor with the detection of C-reactive
protein by the latex agglutination method. J Rheumatol. 7:279–287.
1980.PubMed/NCBI
|
|
47
|
Rifai N and Silverman LM: A simple
immunotechnique for the determination of serum concentration of
apolipoprotein E. Clin Chim Acta. 163:207–213. 1987. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Ma J, Cheng J, Wang L, Wang H, Xu L, Liu
P, Bu S, Zhang L, Le Y, Ye M, et al: No association between IRS-1
promoter methylation and type 2 diabetes. Mol Med Rep. 8:949–953.
2013.PubMed/NCBI
|
|
49
|
Zhang LN, Liu PP, Wang L, Yuan F, Xu L,
Xin Y, Fei LJ, Zhong QL, Huang Y, Xu L, et al: Lower ADD1 gene
promoter DNA methylation increases the risk of essential
hypertension. PloS One. 8:e634552013. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Zemek F, Drtinova L, Nepovimova E, Sepsova
V, Korabecny J, Klimes J and Kuca K: Outcomes of Alzheimer's
disease therapy with acetylcholinesterase inhibitors and memantine.
Expert Opin Drug Saf. 13:759–774. 2014.PubMed/NCBI
|
|
51
|
Pritchard AL, Ratcliffe L, Sorour E, Haque
S, Holder R, Bentham P and Lendon CL: Investigation of dopamine
receptors in susceptibility to behavioural and psychological
symptoms in Alzheimer's disease. Int J Geriatr Psychiatry.
24:1020–1025. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Fratiglioni L, Viitanen M, von Strauss E,
Tontodonati V, Herlitz A and Winblad B: Very old women at highest
risk of dementia and Alzheimer's disease: Incidence data from the
Kungsholmen Project, Stockholm. Neurology. 48:132–138. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Penaloza CG, Estevez B, Han DM, Norouzi M,
Lockshin RA and Zakeri Z: Sex-dependent regulation of cytochrome
P450 family members Cyp1a1, Cyp2e1, and Cyp7b1 by methylation of
DNA. FASEB J. 28:966–977. 2014. View Article : Google Scholar :
|
|
54
|
Piferrer F: Epigenetics of sex
determination and gonadogenesis. Dev Dyn. 242:360–370. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Siegmund KD, Connor CM, Campan M, Long TI,
Weisenberger DJ, Biniszkiewicz D, Jaenisch R, Laird PW and Akbarian
S: DNA methylation in the human cerebral cortex is dynamically
regulated throughout the life span and involves differentiated
neurons. PloS One. 2:e8952007. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Paula-Lima AC, Tricerri MA, Brito-Moreira
J, Bomfim TR, Oliveira FF, Magdesian MH, Grinberg LT, Panizzutti R
and Ferreira ST: Human apolipoprotein A-I binds amyloid-beta and
prevents Abeta-induced neurotoxicity. Int J Biochem Cell Biol.
41:1361–1370. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Fagan AM, Younkin LH, Morris JC, Fryer JD,
Cole TG, Younkin SG and Holtzman DM: Differences in the
Abeta40/Abeta42 ratio associated with cerebrospinal fluid
lipoproteins as a function of apolipoprotein E genotype. Ann
Neurol. 48:201–210. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Emanuele E, Peros E, Tomaino C, Feudatari
E, Bernardi L, Binetti G, Maletta R, Micieli G, Bruni AC and
Geroldi D: Relation of apolipoprotein(a) size to alzheimer's
disease and vascular dementia. Dement Geriatr Cogn Disord.
18:189–196. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Solfrizzi V, Panza F, D'Introno A,
Colacicco AM, Capurso C, Basile AM and Capurso A: Lipoprotein(a),
apolipoprotein E genotype, and risk of Alzheimer's disease. J
Neurol Neurosurg Psychiatry. 72:732–736. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Suszynska-Zajczyk J, Luczak M, Marczak L
and Jakubowski H: Inactivation of the paraoxonase 1 gene affects
the expression of mouse brain proteins involved in
neurodegeneration. J Alzheimers Dis. 42:247–260. 2014.PubMed/NCBI
|
|
61
|
Nazef K, Khelil M, Chelouti H, Kacimi G,
Bendini M, Tazir M, Belarbi S, El Hadi Cherifi M and Djerdjouri B:
Hyperhomocysteinemia is a risk factor for Alzheimer's disease in an
Algerian population. Arch Med Res. 45:247–250. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Hosseinzadeh S, Dabidi Roshan V and
Pourasghar M: Effects of intermittent aerobic training on passive
avoidance test (shuttle box) and stress markers in the dorsal
hippocampus of wistar rats exposed to administration of
homocysteine. Iran J Psychiatry Behav Sci. 7:37–44. 2013.
|
|
63
|
Lu YQ, Luo Y, He ZF, Chen J, Yan BL, Wang
Y and Yu Q: Hydroxysafflor yellow A ameliorates
homocysteine-induced Alzheimer-like pathologic dysfunction and
memory/synaptic disorder. Rejuvenation Res. 16:446–452. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Yarchoan M, Louneva N, Xie SX, Swenson FJ,
Hu W, Soares H, Trojanowski JQ, Lee VM, Kling MA, Shaw LM, et al:
Association of plasma C-reactive protein levels with the diagnosis
of Alzheimer's disease. J Neurol Sci. 333:9–12. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Duong T, Nikolaeva M and Acton PJ:
C-reactive protein-like immunoreactivity in the neurofibrillary
tangles of Alzheimer's disease. Brain Res. 749:152–156. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Iwamoto N, Nishiyama E, Ohwada J and Arai
H: Demonstration of CRP immunoreactivity in brains of Alzheimer's
disease: Immunohistochemical study using formic acid pretreatment
of tissue sections. Neurosci Lett. 177:23–26. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Ponnayyan Sulochana S, Sharma K, Mullangi
R and Sukumaran SK: Review of the validated HPLC and LC-MS/MS
methods for determination of drugs used in clinical practice for
Alzheimer's disease. Biomed Chromatogr. 28:1431–1490. 2014.
View Article : Google Scholar : PubMed/NCBI
|