Introduction
Retinopathy of prematurity (ROP) is a vascular
proliferative disease that primarily occurs in preterm infants,
causing disorganized growth of retinal blood vessels, which results
in scarring, retinal detachment and occasionally blindness
(1). ROP is associated with low
gestational age and birth weight; although advances in neonatal
care have resulted in improved survival rates for premature
infants, the incidence of ROP has increased (2,3).
Numerous studies have been undertaken to examine the
pathophysiological mechanisms of ROP (4,5);
however, these currently remain unclear.
Animal models of oxygen-induced retinopathy (OIR)
have previously been used to reveal mechanisms of ROP development,
including oxidant stress, the functions of growth factors including
insulin-like growth factor (IGF) and vascular endothelial growth
factor (VEGF), and structure-functional anomalies (6,7).
Bioinformatic strategies have also been widely used to investigate
ROP mechanisms; a study by Sato et al (8) investigated gene-expression profiles
in murine oxygen-induced retinopathy as early as 2009.
microRNAs (miRNAs) are small non-coding RNAs of
18–26 nucleotides in length that regulate gene expression through
sequence-specific base pairing with target mRNAs. Numerous miRNAs
have been identified through molecular cloning and bioinformatics
strategies (9,10). miRNAs generally inhibit mRNA
translation of their target genes (11), and have been reported to be
important in biological processes including vascular angiogenesis,
cell growth, embryonic development, cell proliferation, tissue
differentiation and apoptosis (12–15).
It is well documented that miRNAs regulate vascular
angiogenesis (16) and the
regulation of VEGF, IGF-2 and hypoxia-inducible factor 1α (HIF-1α)
by miRNA (miR)-126 is associated with retinal vascular angiogenesis
under conditions of OIR (17).
However, whether other miRNAs are involved in the pathological
process of angiogenesis in OIR conditions currently remains
unclear. Therefore, the present study aimed to screen for miRNA
candidates affecting retinal neovascularization using an OIR mouse
model.
Materials and methods
Establishment of oxygen-induced
retinopathy mouse model
A total of 80 (36 female; 44 male) 7-day-old healthy
C57BL/6 neonatal mice (weight, 3±0.3 g) were provided by the Animal
Center of Nanjing University (Nanjing, China). All procedures in
the study followed the protocols approved by the Animal Care and
Use Committees of Nanjing Medical University (Nanjing, China). The
mice were maintained in specific-pathogen-free conditions at the
Animal Center of Nanjing Medical University. The mice were reared
in a clean grade animal room, at 23±2°C and 50±10% humidity. The
light did not exceed 350 lux and the light/dark cycle was 12-h.
Food, water and bedding were replaced every two days. The oxygen
concentration was monitored with an oxygen concentration detector.
The mice were randomly divided into the OIR group (n=40) or the
control group (n=40). For OIR modeling, 7-day-old mice were
maintained in 75% oxygen condition for 5 days, then in normal room
air for a further 5 days. Mice in the control group were maintained
in normal room air for the duration of the experiment. The present
study was approved by the ethics review board of Nanjing Children's
Hospital of Nanjing Medical University.
Retinal angiography
On postnatal day 17 (P17), 50 mg high molecular
weight fluorescein isothiocyanate dextrans (catalog no. 75005;
Sigma-Aldrich; Merck Millipore, Darmstadt, Germany) were dissolved
in 1 ml phosphate buffer containing 4% polyformaldehyde and
directly perfused through the left ventricle of the heart following
an intraperitoneal injection of 10% chloral hydrate (3 ml/kg;
Sigma-Aldrich; Merck Millipore). The eyes were then enucleated and
fixed with 4% paraformaldehyde at 4°C for 2 h. The retina was
separated and mounted on a microscope slide. The vasculature was
examined under a fluorescent microscope (Nikon Corporation, Tokyo,
Japan) at an original magnification of ×40. In total, 30 sections
were counted, and 3 fields of view were assessed for each
slide.
Hematoxylin and eosin (HE)
staining
Paraformaldehyde-fixed retinal tissue was embedded
in paraffin and sliced into 6 µm sections. The sections were
stained with HE and examined by routine light microscopy (Olympus
Corporation, Tokyo, Japan). Briefly, sections were dried, followed
by xylene dewaxing and rehydration with decreasing concentrations
of alcohol. The slides were then stained with hematoxylin for 15
min and differentiated with hydrochloric acid alcohol for 30 sec.
Following staining with 1% ammonia, sections were stained with 1%
eosin for 2 min, followed by dehydration in alcohol and mounting.
The number of vascular endothelial cells with nuclei breaking
through the internal limiting membrane (ILM) was counted. In total,
30 sections were counted, and 3 fields of view were assessed for
each slide.
Extraction and quantification of total
RNA
For total RNA extraction, retinal tissue was frozen
at −80°C and crushed using a BioPulverizer (Aoran Technology Ltd.,
Shanghai, China). Homogenization was performed by adding 1 ml
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) to the tissue fragments, according to the
manufacturer's protocol. Extracted RNA was dissolved in RNAse-free
water and incubated at 60°C for 10 min.
Absorbance at 260, 280 and 230 nm was measured using
an ultraviolet spectrophotometer (NanoDrop Technologies; Thermo
Fisher Scientific, Inc., Wilmington, DE, USA). The purity of the
extracted RNAs was determined by calculating the A260/A280 and
A260/A230 ratios.
miRNA microarray
Following isolation of total RNA from retinal
tissue, microarray was performed by LC Sciences (Houston, TX, USA).
Each detection probe on the microfluidic chips consisted of a
chemically modified nucleotide coding segment complementary to
target a total of 25,141 miRNAs (from miRBase version 19.0,
www.mirbase.org/) or other RNA (control or
customer-defined sequences) and a spacer segment of polyethylene
glycol to extend the coding segment away from the substrate. The
detection probes were made by in situ synthesis using
photogenerated reagent chemistry. The hybridization melting
temperatures were balanced by chemical modifications of the
detection probes. Hybridization used 100 µl 6X saline sodium
phosphate EDTA buffer (0.90 M NaCl, 60 mM
Na2HPO4, 6 mM ethylenediaminetetraacetic
acid, pH 6.8) containing 25% formamide at 34°C. Following RNA
hybridization, tag-conjugating Cyanine3 dye was circulated through
the microfluidic chip at 34°C overnight. Fluorescence images were
collected using a GenePix 4000B microarray scanner (Molecular
Devices LLC, Sunnyvale, CA, USA) and visualized using Array-Pro
image analysis software version 6.3 (Media Cybernetics, Inc.,
Rockville, MD, USA). Data were analyzed by subtracting the
background, and then normalizing the signals using a LOWESS filter
(locally-weighted regression).
Following normalization, differentially expressed
miRNAs were identified through fold-change filtering (fold change
≥2). Hierarchical clustering was performed using MeV software
version 4.6 (The Institute for Genomic Research, J. Craig Venter
Institute, Rockville, MA, USA).
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Following isolation of total RNA, 1 µg total RNA was
used for RT using the TaqMan® MicroRNA Reverse
Transcription kit (Applied Biosystems; Thermo Fisher Scientific,
Inc.) according to the manufacturer's protocol. The cDNA obtained
was used for qPCR, with probes for Mus musculus
(mmu)-miR-130a-3p and mmu-miR-5107-5p, listed in Table I. The hot start method was used for
fluorescent qPCR and the standard internal reference [U6
spliceosomal RNA (snRNA)] coupled with the control group was
amplified for each sample. qPCR was performed in a 20-µl reaction
mixture, with each tube containing 10 µl 2X TaqMan Universal Master
Mix II, 1 µl TaqMan MicroRNA assays (Applied Biosystems; Thermo
Fisher Scientific, Inc.), 1.33 µl cDNA template and 7.67 µl
RNase-free water. The cycling conditions were as follows: An
initial denaturation step at 50°C for 2 min and 95°C for 2 min,
followed by 40 cycles of denaturation at 95°C for 15 sec, and
annealing/extension at 60°C for 1 min. The U6 snRNA was used as an
internal control for normalization. The relative quantitative (RQ)
value, obtained using the 2−ΔΔCq method (18), was used for statistical analysis.
Briefly, the cycle threshold (Cq) values were calculated using
automatic baseline settings. The ΔCq value between internal control
U6 and the corresponding Cq value for the target gene was used for
the relative quantification of miRNA expression.
 | Table I.Context sequences of the probes of
mmu-miR-130a-3p, mmu-miR-5107-5p and the internal control U6 snRNA
used in the present study. |
Table I.
Context sequences of the probes of
mmu-miR-130a-3p, mmu-miR-5107-5p and the internal control U6 snRNA
used in the present study.
| Assay name | Context sequence
(5′-3′) |
|---|
| mmu-miR-130a |
CAGUGCAAUGUUAAAAGGGCAU |
| mmu-miR-5107 |
UGGGCAGAGGAGGCAGGGACA |
| U6 snRNA |
GTGCTCGCTTCGGCAGCACATATACTAAAATTGGAACGATACAGAGAAGATTAGCATGGCCCCTGCGCAAGGATGACACGCAAATTCGTGAAGCGTTCCATATTTT |
Statistical analysis
Statistical analyses were performed using SPSS
version 13.0 (SPSS, Inc., Chicago, IL, USA). Data were expressed as
the mean ± standard deviation. Comparisons of retinal endothelial
cell nucleus between the OIR and control groups were performed
using the means t-test. RT-qPCR data were evaluated by one-way
analysis of variance and independent sample t-test. P<0.05 was
considered to indicate a statistically significant difference. To
evaluate miRNA expression profiles, t-tests were performed, and
P<0.01 was considered to indicate a statistically significant
difference.
Results
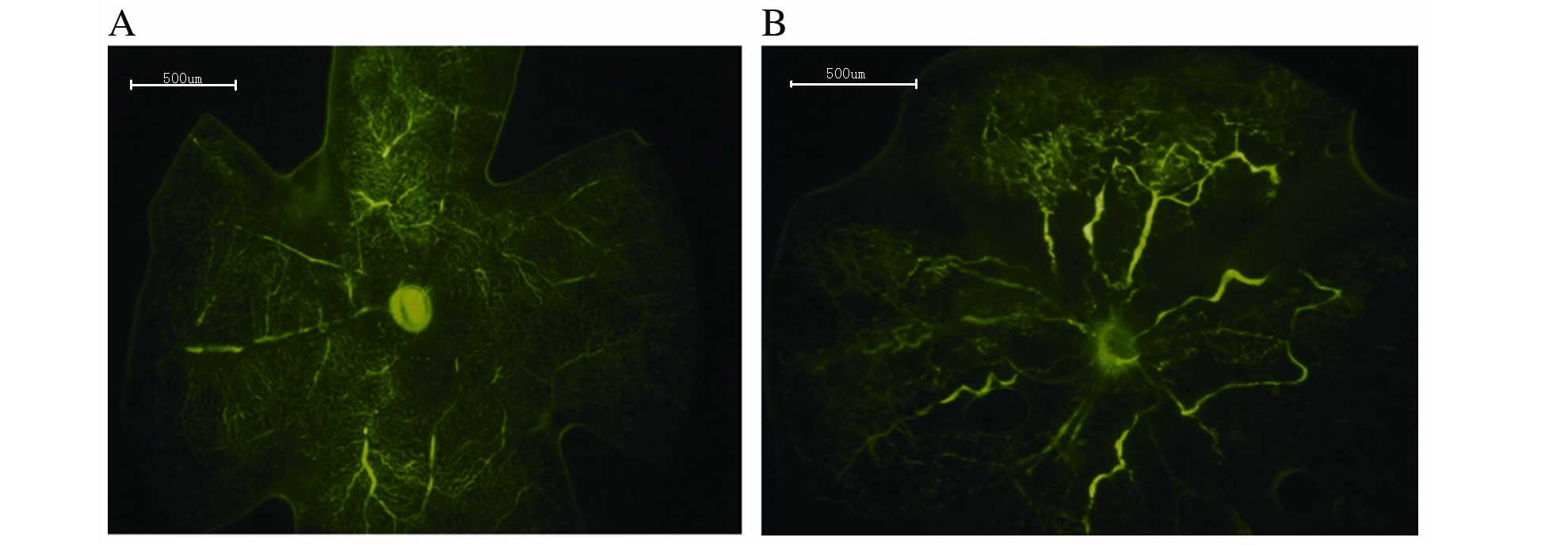
Result of retinal angiography
Retinal angiography was performed to detect
differences between the control and OIR groups, and thus determine
whether the murine model was successfully established. The retinal
vasculature of the control group extended from the optic nerve to
the periphery, with the vessels forming a fine radial branching
pattern (Fig. 1A). The retinal
vascular pattern in the OIR group was characterized by decreased
central perfusion, which was particularly notable in the central
retina (Fig. 1B). The murine OIR
model was, therefore, successfully established.
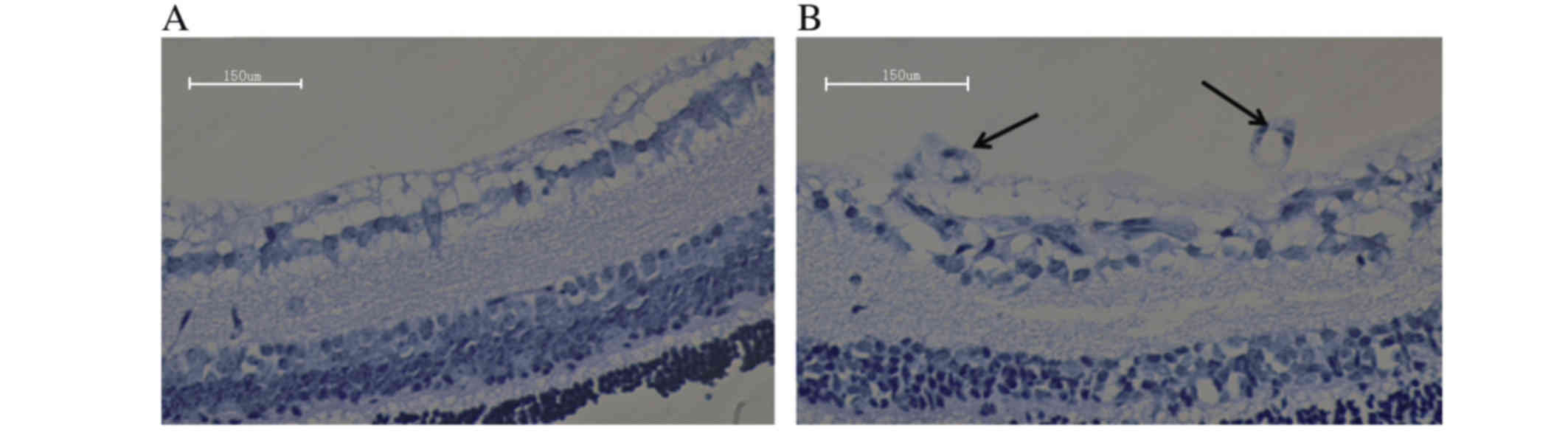
Histological changes
Histology was conducted to examine differences in
retinal tissue between the control and OIR groups. Vascular cells
in the control group did not extend from the internal limiting
membrane into the vitreous (Fig.
2A), while vascular cells in the OIR group extended from the
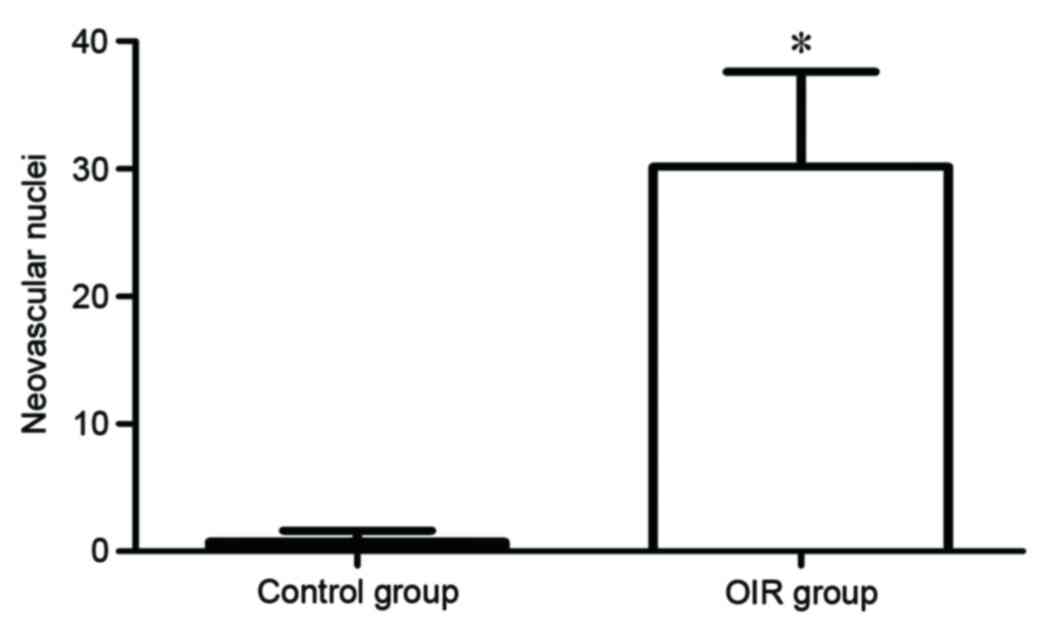
internal limiting membrane into the vitreous (Fig. 2B). The degree of hyperoxia-induced
neovascularization was quantified by counting the number of
vascular cell nuclei on the vitreal side of the internal limiting
membrane of the retina. There was an average of 0.89±0.71 nuclei
extending into the internal limiting membrane in the control group,
compared with 30.17±7.43 nuclei in the OIR group (P=0.00006;
Fig. 3). The retinal tissue in the
OIR model was, therefore demonstrated to be damaged, further
confirming the successful establishment of the OIR model.
Quality assessment of total retinal
RNA
In order to determine the quality of the extracted
retinal tissue RNA, total RNA quality assessment was performed. RNA
with A260/A280 ratio close to 2.0 is regarded as pure RNA:
A260/A280 ratio <1.8 indicates sample contamination, while a
ratio >2.0 suggests RNA hydrolysis. An A260/A280 ratio in the
range of 1.8–2.1 is acceptable. In addition, for pure RNA samples,
the A260/A230 ratio should be >1.8. As presented in Table II, the A260/A280 ratios of the
extracted RNA were ~2.0 and the 260/A230 ratios were all >2.2.
The quality of the extracted RNA was therefore good.
 | Table II.RNA purity from retinal tissue RNA
extractions. |
Table II.
RNA purity from retinal tissue RNA
extractions.
| Sample group | Concentration
(µg/µl) | UV A260/A280 | UV A260/A230 | Quantity (µg) |
28S/18Sa | Bioanalyzer
result |
|---|
| Oxygen-induced
retinopathy | 0.25 | 2.00 | 2.21 | 10.72 | 2.5 | Pass |
| Control | 0.34 | 2.03 | 2.28 | 14.59 | 2.4 | Pass |
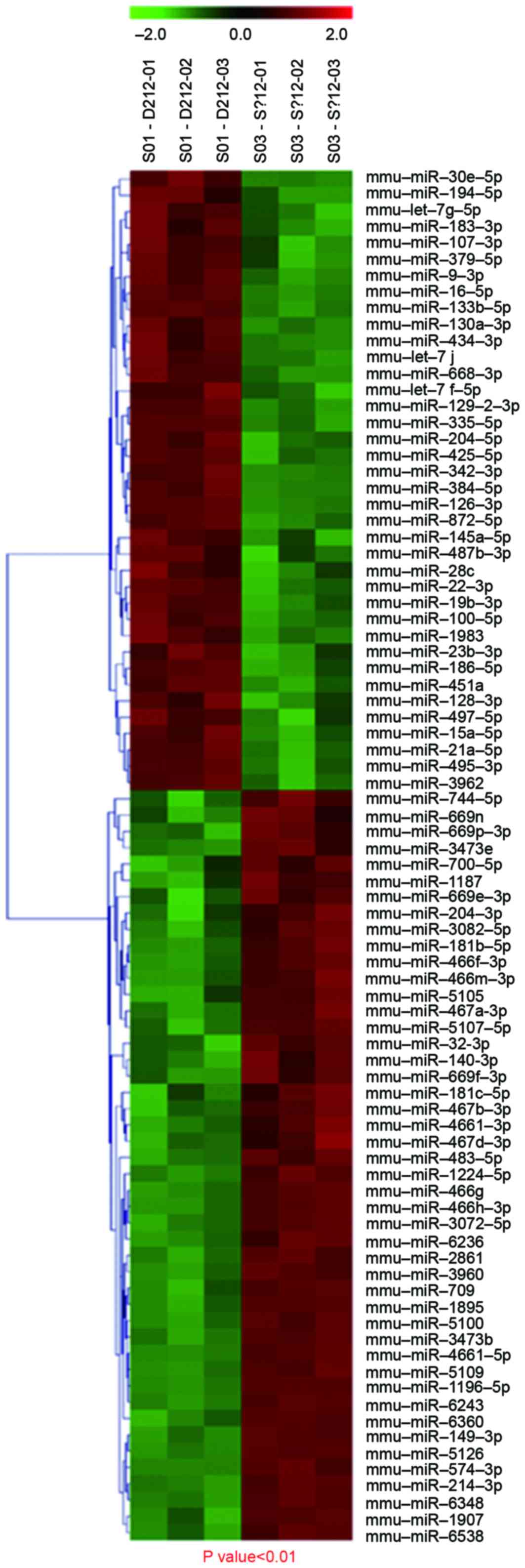
miRNA expression profile
To examine changes in miRNA expression levels in
mice in the OIR group, a microarray analysis was performed to
evaluate miRNA expression profiles. There were 67 differentially
expressed miRNAs in the OIR group compared with the control group,
of which, 34 were upregulated and 33 were downregulated. There were
32 differentially expressed miRNAs with a fold change ≥2.0, of
which, 21 were upregulated (Table
III) and 11 were downregulated (Table IV). Of these miRNAs, 6 miRNAs that
were upregulated in the OIR group compared with the control group
(miR-1907, miR-3970, miR-5107-5p, miR-133b-5p, miR-3962 and
miR-145b; Table III) and 11
miRNAs that were downregulated in the OIR group compared with the
control group (miR-495-3p, miR-677-3p, miR-let-7d-3p, miR-328-3p,
miR-301a-3p, miR-128-3p, miR-872-5p, miR-181c-5p, miR-744-5p,
miR-130a-3p and miR-377-3p; Table
IV) were statistically significantly different (P<0.05),
with a ≥2-fold change. Of the upregulated miRNAs in the OIR group,
mmu-miR-1907 had the greatest fold-change compared with the control
group, and of the downregulated miRNAs, mmu-miR-495-3p had the
greatest difference compared with the control group. Hierarchical
clustering revealed distinguishable miRNA expression profiles
between the two groups, with S01 as the control group and S03 as
the experimental group (Fig. 4).
Thus, modulation of oxygen conditions resulted in differential
miRNA expression.
 | Table III.Differentially expressed microRNAs
upregulated (>2-fold) in the oxygen-induced retinopathy group
compared with the control group, listed and arranged according to
magnitude of change. |
Table III.
Differentially expressed microRNAs
upregulated (>2-fold) in the oxygen-induced retinopathy group
compared with the control group, listed and arranged according to
magnitude of change.
| Name | Fold-change | P-value |
|---|
| mmu-miR-1907 | 14.389 | 0.000108 |
| mmu-miR-3970 | 12.300 | 0.000069 |
|
mmu-miR-5107-5p |
9.714 | 0.000632 |
|
mmu-miR-133b-5p |
7.753 | 0.000973 |
| mmu-miR-3962 |
7.068 | 0.000009 |
| mmu-miR-145b |
5.077 | 0.000924 |
|
mmu-miR-3072-5p |
3.690 | 0.002920 |
|
mmu-miR-129-2-3p |
3.074 | 0.001340 |
| mmu-miR-714 |
3.000 | 0.004800 |
| mmu-miR-29c-3p |
2.833 | 0.001420 |
|
mmu-miR-145a-5p |
2.500 | 0.003340 |
| mmu-miR-100-5p |
2.425 | 0.000081 |
|
mmu-miR-3107-5p |
2.385 | 0.005480 |
|
mmu-miR-30c-1-3p |
2.172 | 0.004510 |
| mmu-miR-96-5p |
2.157 | 0.009780 |
| mmu-miR-6236 |
2.148 | 0.006500 |
| mmu-miR-3473b |
2.122 | 0.005810 |
| mmu-miR-214-3p |
2.093 | 0.006740 |
| mmu-miR-9-5p |
2.089 | 0.000624 |
|
mmu-miR-181d-5p |
2.070 | 0.002950 |
| mmu-miR-3473e |
2.045 | 0.005690 |
 | Table IV.Differentially expressed microRNAs
downregulated (>2-fold) in the oxygen-induced retinopathy group
compared with the control group, listed and arranged according to
magnitude of change. |
Table IV.
Differentially expressed microRNAs
downregulated (>2-fold) in the oxygen-induced retinopathy group
compared with the control group, listed and arranged according to
magnitude of change.
| Name | Fold-change | P-value |
|---|
| mmu-miR-495-3p | 3.354 | 0.000002 |
| mmu-miR-677-3p | 3.255 | 0.000206 |
|
mmu-miR-let-7d-3p | 2.606 | 0.002650 |
| mmu-miR-328-3p | 2.465 | 0.001420 |
|
mmu-miR-301a-3p | 2.462 | 0.003470 |
| mmu-miR-128-3p | 2.149 | 0.002440 |
| mmu-miR-872-5p | 2.076 | 0.008250 |
|
mmu-miR-181c-5p | 2.070 | 0.006170 |
| mmu-miR-744-5p | 2.065 | 0.004720 |
|
mmu-miR-130a-3p | 2.000 | 0.000412 |
| mmu-miR-377-3p | 2.000 | 0.006950 |
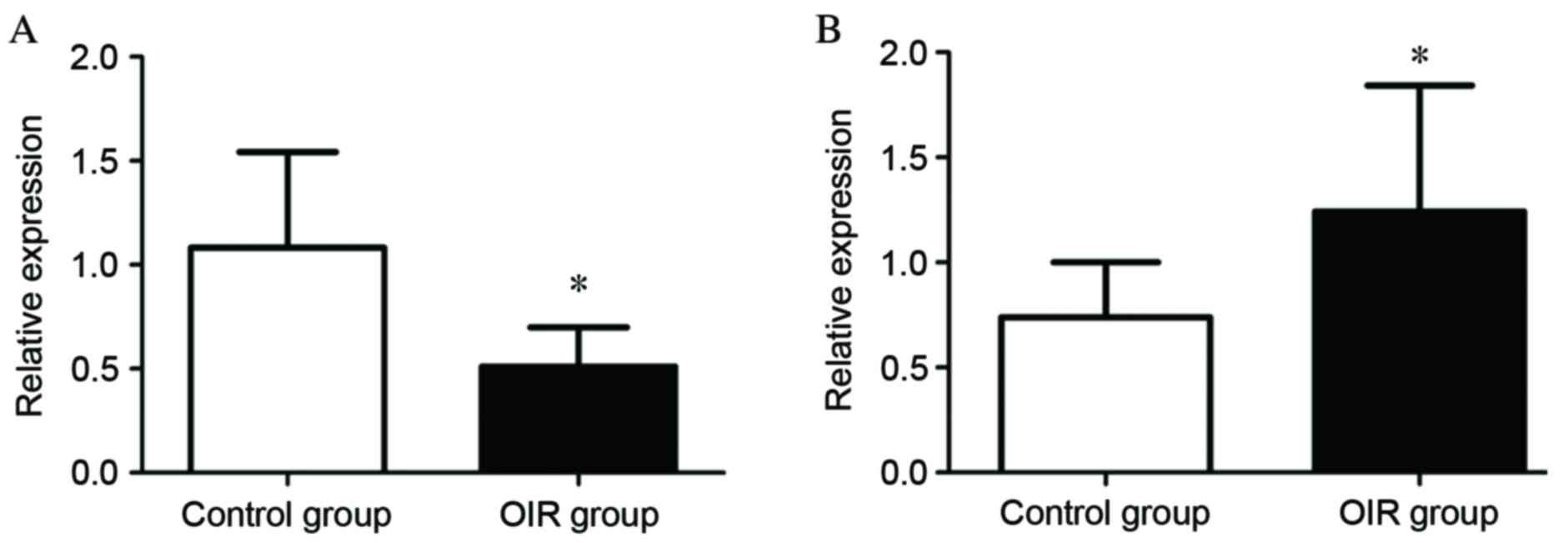
Confirmation of differentially
expressed miRNAs by RT-qPCR
To validate the differential expression profiles of
miRNAs indicated by microarray analysis, RT-qPCR analysis of miRNAs
was performed for mmu-miR-130a-3p and mmu-miR-5107-5p. These miRNAs
were selected for confirmatory testing by RT-qPCR as they exhibited
large fold-changes, large mean values and small P-values. The
relative expression level of mmu-miR-130a-3p was significantly
lower in the OIR group compared with the control group (P=0.00239;
Fig. 5A), while that of
mmu-miR-5107-5p was significantly higher in the OIR group compared
with the control group (P=0.0352; Fig.
5B). The data from this confirmatory RT-qPCR analysis was,
therefore, consistent with that of the microarray analysis.
Discussion
In the present study, a mouse model of OIR was
constructed and subsequently used to identify differentially
expressed miRNAs via microarray. Two of the differentially
expressed miRNAs identified by microarray were confirmed by
RT-qPCR. Hyperoxia was, therefore, demonstrated to result in the
dysregulation of miRNAs in murine retinal tissue, suggesting that
miRNAs are important in the development and progression of
oxygen-induced ROP.
The OIR mouse model was first developed in 1994
(7) and is commonly used for
studies of retinal neovascularization in ROP and other
vasculopathologies. Subsequent studies have demonstrated its
utility in investigating the pathogenesis of retinal
neovascularization and medical interventions for ROP and other
retinal angiopathies (8,19). The model has also proven to be
reproducible and quantifiable, making it a favorable model for
microarray analysis.
In the present study, a miRNA microarray was used to
analyze miRNA expression profiles and to identify differentially
expressed miRNAs in the retinal tissues of mice subjected to OIR:
67 differentially expressed miRNAs were identified in the OIR group
compared with the control group. The functions of the majority of
the upregulated miRNAs identified in the present study, including
miR-1907, miR-3970, miR-5107-5p, miR-3962 and miR-145b, have not
previously been reported. As a result, the effects of these miRNAs
and potential applications of their functions require further
investigations.
In the present study, the expression of miR-133b-5p
was revealed to be up to 7.753-fold higher in the OIR group
compared with the control group, indicating a potential role for
this miRNA in the development of ROP. miR-133b-5p regulates the
expression of heat shock protein 70 (Hsp70) during gp120 V3 loop
peptide-induced rat neuronal cell apoptosis (20). Hsp70 is a chaperone protein that
inhibits key effectors of the apoptotic machinery at the pre- and
postmitochondrial level (21). In
addition, miR-133b, the precursor of miR-133b-5p, is downregulated
in human osteosarcoma cells, resulting in increased apoptosis, and
potentially acting as a tumor suppressor gene (22). The expression of miR-133b-5p may,
therefore, regulate programmed cell apoptosis pathways and be
involved in the development of ROP.
In contrast to the largely uncharacterized
upregulated miRNAs, some of the downregulated miRNAs identified in
the present study have been demonstrated to be involved in numerous
diseases. The miR-1et-7 family is widely distributed in vertebrates
and invertebrates, and participates in numerous biological
processes, including reticulocytes (23), T-cell generation, angiogenesis
(24) and the Fas-mediated
apoptotic process in peripheral blood mononuclear cells (25,26).
It is, therefore, unsurprising that the expression of miR-let-7d-3p
is decreased in a disease such as ROP, which is caused by abnormal
blood vessel development. miR-328 expression has been demonstrated
to be elevated in numerous lung adenocarcinoma subtypes, which may
affect the degree of malignancy of tumor progression by inhibiting
the proliferation of GBM cells in gliomas (27–30).
A known target of miR-328 is RTP801, which is also a target of
miR-181b-5p, which had an effect on the cancer stem cells fraction
and neurosphere formation as surrogate marker (31). RTP801 expression is induced in the
retina following hyperoxia treatment, while the development of
retinopathy is significantly attenuated in an RTP801-knockout mouse
model of ROP (32). These findings
suggest important roles of miR-328 and miR-181b-5p in the
pathogenesis of ROP by regulating the expression of RTP801.
The potential targets of other miRNAs are also
related to ROP. One of the miR-872-5p targets is insulin-like
growth factor 1, which acts upstream of retinal VEGF and is
important in ROP (33).
Angiopoietin 2, a potential target of miR-128-3p, induces
endothelial cell apoptosis and is involved in ROP (31). miR-301a and miR-130a are miRNAs
that have been suggested to bind to conserved sites of the
peroxisome proliferator-activated receptor γ (PPARγ) 3′
untranslated region (34). PPARγ
is widely expressed in the eye, is involved in the pathogenesis of
OIR (35), and provides a
pharmacological target for treating ocular angiogenesis (36). Furthermore, miR-130a contributes to
angiogenesis by inhibiting the expression of two antiangiogenic
homeobox transcription factors, homeobox A5 (HOXA5) and mesenchyme
homeobox 2 (MEOX2; also termed GAX) (11). However, further research is
required to establish whether these miRNAs (miR-872-5p, miR-128-3p,
miR-301a and miR-130a) regulate ROP development.
In summary, the present study has identified the
differential expression of miRNAs in a murine model of OIR some of
whose functions, or those of their known targets, are associated
with the development of ROP. These findings are useful to
understand the molecular basis of ROP and suggest that miRNAs are
involved in the progression of ROP. However, further studies are
required to confirm the involvement of the miRNAs identified in the
present study in the development of ROP, and their molecular
mechanisms.
Acknowledgements
The present study was supported by the National
Natural Science Foundation of China (grant no. 81260106).
References
|
1
|
McGregor ML, Bremer DL, Cole C, McClead
RE, Phelps DL, Fellows RR and Oden N: HOPE-ROP Multicenter Group.
High Oxygen Percentage in Retinopathy of Prematurity study:
Retinopathy of prematurity outcome in infants with prethreshold
retinopathy of prematurity and oxygen saturation >94% in room
air: The high oxygen percentage in retinopathy of prematurity
study. Pediatrics. 110:540–544. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Sola A, Chow L and Rogido M: Retinopathy
of prematurity and oxygen therapy: A changing relationship. An
Pediatr (Barc). 62:48–63. 2005.(In spanish). PubMed/NCBI
|
|
3
|
Chamney S, McGrory L, McCall E, Twaij S,
Napier M, Rollins R, Marshall AH, Craig S and McLoone E: Treatment
of retinopathy of prematurity in Northern Ireland, 2000–2011: A
population-based study. J AAPOS. 19:223–227. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Levesque BM, Kalish LA, Winston AB, Parad
RB, Hernandez-Diaz S, Phillips M, Zolit A, Morey J, Gupta M,
Mammoto A, et al: Low urine vascular endothelial growth factor
levels are associated with mechanical ventilation, bronchopulmonary
dysplasia and retinopathy of prematurity. Neonatology. 104:56–64.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Patel S Jivabhai, Bany-Mohammed F, McNally
L, Valencia GB, Lazzaro DR, Aranda JV and Beharry KD: Exogenous
superoxide dismutase mimetic without scavenging
H2O2 causes photoreceptor damage in a rat
model for oxygen-induced retinopathy. Invest Ophthalmol Vis Sci.
56:1665–1677. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Penn JS, Tolman BL and Henry MM:
Oxygen-induced retinopathy in the rat: Relationship of retinal
nonperfusion to subsequent neovascularization. Invest Ophthalmol
Vis Sci. 35:3429–3435. 1994.PubMed/NCBI
|
|
7
|
Smith LE, Wesolowski E, McLellan A, Kostyk
SK, D'Amato R, Sullivan R and D'Amore PA: Oxygen-induced
retinopathy in the mouse. Invest Ophthalmol Vis Sci. 35:101–111.
1994.PubMed/NCBI
|
|
8
|
Sato T, Kusaka S, Hashida N, Saishin Y,
Fujikado T and Tano Y: Comprehensive gene-expression profile in
murine oxygen-induced retinopathy. Br J Ophthalmol. 93:96–103.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bentwich I, Avniel A, Karov Y, Aharonov R,
Gilad S, Barad O, Barzilai A, Einat P, Einav U, Meiri E, et al:
Identification of hundreds of conserved and nonconserved human
microRNAs. Nat Genet. 37:766–770. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Xie M, Zhang S and Yu B: microRNA
biogenesis, degradation and activity in plants. Cell Mol Life Sci.
72:87–99. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Doench JG and Sharp PA: Specificity of
microRNA target selection in translational repression. Genes Dev.
18:504–511. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kuhnert F, Mancuso MR, Hampton J,
Stankunas K, Asano T, Chen CZ and Kuo CJ: Attribution of vascular
phenotypes of the murine Egfl7 locus to the microRNA miR-126.
Development. 135:3989–3993. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen Y and Gorski DH: Regulation of
angiogenesis through a microRNA (miR-130a) that down-regulates
antiangiogenichomeobox genes GAX and HOXA5. Blood. 111:1217–1226.
2008.PubMed/NCBI
|
|
14
|
Kuehbacher A, Urbich C, Zeiher AM and
Dimmeler S: Role of Dicer and Drosha for endothelial microRNA
expression and angiogenesis. Circ Res. 101:59–68. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Johnson CD, Esquela-Kerscher A, Stefani G,
Byrom M, Kelnar K, Ovcharenko D, Wilson M, Wang X, Shelton J,
Shingara J, et al: The let-7 microRNA represses cell proliferation
pathways in human cells. Cancer Res. 67:7713–7722. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Shen J, Yang X, Xie B, Chen Y, Swaim M,
Hackett SF and Campochiaro PA: MicroRNAs regulate ocular
neovascularization. Mol Ther. 16:1208–1216. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Bai Y, Bai X, Wang Z, Zhang X, Ruan C and
Miao J: MicroRNA-126 inhibits ischemia-induced retinal
neovascularization via regulating angiogenic growth factors. Exp
Mol Pathol. 91:471–477. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C (T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wang W, Li Z, Sato T and Oshima Y:
Tenomodulin inhibits retinal neovascularization in a mouse model of
oxygen-induced retinopathy. Int J Mol Sci. 13:15373–15386. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Xia C, Cai Y, Lin Y, Guan R, Xiao G and
Yang J: MiR-133b-5p regulates the expression of the heat shock
protein 70 during rat neuronal cell apoptosis induced by the gp120
V3 loop peptide. J Med Virol. 88:437–447. 2015.PubMed/NCBI
|
|
21
|
Rérole AL, Jego G and Garrido C: Hsp70:
Anti-apoptotic and tumorigenic protein. Methods. 787:205–230.
2011.
|
|
22
|
Zhao H, Li M, Li L, Yang X, Lan G and
Zhang Y: MiR-133b is down-regulated in human osteosarcoma and
inhibits osteosarcoma cells proliferation, migration, and invasion
and promotes apoptosis. PLoS One. 8:e835712013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Noh SJ, Miller SH, Lee YT, Goh SH,
Marincola FM, Stroncek DF, Reed C, Wang E and Miller JL: Let-7
microRNAs are developmentally regulated in circulating human
erythroid cells. J Transl Med. 7:982009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Landskroner-Eiger S, Moneke I and Sessa
WC: miRNAs as modulators of angiogenesis. Cold Spring Harb Perspect
Med. 3:a0066432013.PubMed/NCBI
|
|
25
|
Vaz F, Hanenberg H, Schuster B, Barker K,
Wiek C, Erven V, Neveling K, Endt D, Kesterton I, Autore F, et al:
Mutation of the RAD51C gene in a Fanconi anemia-like disorder. Nat
Genet. 42:406–409. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang S, Tang Y, Cui H, Zhao X, Luo X, Pan
W, Huang X and Shen N: Let-7/miR-98 regulate Fas and Fas-mediated
apoptosis. Genes Immun. 12:149–154. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ulivi P, Foschi G, Mengozzi M, Scarpi E,
Silvestrini R, Amadori D and Zoli W: Peripheral Blood mir-328
expression as a potential biomarker for the early diagnosis of
NSCLC. Int J Mol Sci. 14:10332–10342. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Arora S, Ranade AR, Tran NL, Nasser S,
Sridhar S, Korn RL, Ross JT, Dhruv H, Foss KM, Sibenaller Z, et al:
MicroRNA-328 is associated with (non-small) cell lung cancer
(NSCLC) brain metastasis and mediates NSCLC migration. Int J
Cancer. 129:2621–2631. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Okumus NO, Gursel B, Meydan D, Ozdemir O,
Odabas E and Gonullu G: Prognostic significance of concomitant
radiotherapy in newly diagnosed glioblastomamultiforme: A
multivariate analysis of 116 patients. Ann Saudi Med. 32:250–255.
2012.PubMed/NCBI
|
|
30
|
Delic S, Lottmann N, Stelzl A, Liesenberg
F, Wolter M, Götze S, Zapatka M, Shiio Y, Sabel MC, Felsberg J, et
al: MiR-328 promotes glioma cell invasion via SFRP1-dependent
Wnt-signaling activation. Neuro Oncol. 16:179–190. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kozomara A and Griffiths-Jones S: miRBase:
Annotating high confidence microRNAs using deep sequencing data.
Nucleic Acids Res. 42:D68–D73. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Brafman A, Mett I, Shafir M, Gottlieb H,
Damari G, Gozlan-Kelner S, Vishnevskia-Dai V, Skaliter R, Einat P,
Faerman A, et al: Inhibition of oxygen-induced retinopathy in
RTP801-deficient mice. Invest Ophthalmol Vis Sci. 45:3796–3805.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Nidadavolu LS, Niedernhofer LJ and Khan
SA: Identification of microRNAs dysregulated in cellular senescence
driven by endogenous genotoxicstress. Aging (Albany NY). 5:460–473.
2013.PubMed/NCBI
|
|
34
|
Huang JY, Chou SF, Lee JW, Chen HL, Chen
CM, Tao MH and Shih C: MicroRNA-130a can inhibit hepatitis B virus
replication via targeting PGC1α and PPARγ. RNA. 21:385–400. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Tawfik A, Sanders T, Kahook K, Akeel S,
Elmarakby A and Al-Shabrawey M: Suppression of retinal peroxisome
proliferator-activated receptor gamma in experimental diabetes and
oxygen-induced retinopathy: Role of NADPH oxidase. Invest
Ophthalmol Vis Sci. 50:878–884. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Simpson-Haidaris PJ, Pollock SJ, Ramon S,
Guo N, Woeller CF, Feldon SE and Phipps RP: Anticancer role of
PPARgamma agonists in hematological malignancies found in the
vasculature, marrow and eyes. PPAR Res. 2010:8146092010.PubMed/NCBI
|