Introduction
Flavonoids are an important class of secondary
metabolites present in soybean. They have important physiological
functions, such as antifungal and antioxidation. They can inhibit
the growth of microorganisms and promote the growth of the soybean
(1). Soybean glycosides are one of
the components of isoflavones and are important physiologically
active substances (2). A study has
reported that chalcone reductase (CHR) is required for the
synthesis of precursors of soybean glycosides; CHR synthesizes
isoliquiritigenin, a precursor to daidzein (3). For the synthesis of daidzein,
cinnamic amide is first formed from phenylalanine under the action
of phenylalanine lyase and then catalyzed by 4-hydroxycinnamamidase
to coumaric acid. Coumaric acid coenzyme A ligase (4CL) effects the
conversion to the coumarin coenzyme A. Coumarin coenzyme A is a
common precursor of daidzein and genistein synthesis. In the
synthesis pathway of daidzein, CHR and chalcone synthase combine to
catalyze the synthesis of allyl coenzyme A. Glycyrrhizin is
catalyzed by chalcone isomerase to produce licorice and licorice is
catalyzed by isoflavone synthase (IFS) to synthesize daidzein.
Soybean glycosides have certain biological effects
that genistein does not. Soybean glycosides have various beneficial
effects, including preventing breast cancer (4), improving immunity (5), preventing skin burns caused by
ultraviolet radiation (6) and the
treatment of women with postpartum depression (7). They have an important role in
protecting against soybean Phytophthora root rot (8,9).
There are multiple forms of CHR gene in the
soybean genome (10). A previous
study report identified five CHR genes (11). Besides the CHR1 gene, the
function of the others has not been fully characterized (12,13).
Graham et al (14) verified
4 CHR gene fragments using RNAi technology, and demonstrated
that not all of the CHR proteins were involved in the synthesis of
soybean glycosides. In 2009, Liu (15) isolated a new CHR gene in
soybean. Based on the previous research, the type and quantity of
CHR genes present in soybean is inconclusive. Therefore,
CHR gene cloning and identification are also the important
for understanding the regulation of glycoside synthesis in
soybean.
In the current study, a CHR3 gene expression
vector was constructed, and transformed into soybean varieties to
overexpress CHR3. Subsequently, the effects of CHR3
on the synthesis of soybean glycosides were analyzed, the function
and efficiency of different CHR genes were identified and
the mechanism of CHR action in the synthesis of soybean glycosides
was determined, which lays a theoretical foundation for breeding
soybean resistant to Phytophthora root rot by using genetic
engineering technology to control the synthesis and metabolism of
soybean glycosides.
Materials and methods
Materials
Soybean varieties ‘Jinong 17’ and ‘Jilin 30’, E.
coli DH5α, Agrobacterium tumefaciens strain EHA105,
pMD18-T cloning vector, the recombinant prokaryotic expression
vector BL21-pET28a and expression vector pCAMBIA3300 were all
provided and maintained by the Plant Biotechnology Center of Jilin
Agricultural University (Changchun, China).
Obtaining the objective fragment
The BL21-pET28a-CHR3 recombinant expression plasmid,
which was cloned by Zhang et al (16) (GenBank accession no: KF927169), was
used as the template, and objective fragment was amplified using
specific primers CHR3 sense/CHR3 antisense (Table I) and sequenced by the NCBI
(National Institutes of Health, Bethesda, MD, USA).
 | Table I.Primers of different genes used for
cloning and reverse transcription-quantitative polymerase chain
reaction. |
Table I.
Primers of different genes used for
cloning and reverse transcription-quantitative polymerase chain
reaction.
| Name | Sequence (5′-3′) | Product (bp) |
|---|
| CHR3 sense | CCCGAGCTCTTCAGGACACAAATGCCA | CHR3 (932) |
| CHR3 antisense | TTTGGATCCCTTAAACGTCTCCATCCC |
|
| 35S sense |
TAGAGGACCTAACAGAAC | 35S (500) |
| 35S antisense |
CCGTGTTCTCTCCAAATG |
|
| Bar sense |
TCAAATCTCGGTGACGGGC | Bar (552) |
| Bar antisense |
ATGAGCCCAGAACGACGC |
|
| Q-CHR3 sense |
GGTGGGTTACCGTCATTTTG |
|
| Q-CHR3
antisense |
TCATGTCTCAGCCTCACTGG |
|
| Q-ACT forward |
ATCTTGACTGAGCGTGGTTATTCC |
|
| Q-ACT reverse |
GCTGGTCCTGGCTGTCTCC |
|
The polymerase chain reaction (PCR) amplification
system was as follows: 2.5 µl MgCl2, 2.5 µl 10X Taq
buffer, 1 µl specific primer, 1 µl template, 0.5 µl dNTP, 0.2 µl
Taq polymerase (reagents from Takara Biotechnology Co., Ltd.,
Dalian, China), sterile water was added up to 25 µl. Amplification
conditions were as follows: 94°C predenaturation for 10 min, then
94°C denaturation for 30 sec, 55°C renaturation for 40 sec and 72°C
extension for 50 sec for 35 cycles; a final 72°C extension for 10
min was performed and then maintained at 4°C. PCR products were
separated by 1% agarose gel electrophoresis, and then ligated into
the pMD18-T vector following recovery from the gel (DNA gel
extraction kit; Takara Biotechnology Co., Ltd.). The mixture was: 5
µl CHR3, 1 µl solution I and 4 µl pMD18-T at 16°C overnight.
Recombinant cloning vector pMD18-T-CHR3 was transformed into
competent cells E. coli DH5α, and then monoclonal colonies
were selected and sequenced by Comate Bioscience Co., Ltd.,
(Changchun, China).
Construction of overexpression vector
of CHR3 gene
The CHR3 gene fragment was amplified by PCR as
described above and its products were separated by 1% agarose gel
electrophoresis, and obtaining purified target fragments. Following
electrophoresis, the gel was placed under UV light and the target
gel cut out with a knife and placed in tubes and the AxyPrep DNA
Gel Extraction kit (Corning Life Sciences Limited. Wujiang, China)
employed. Fragment and the objective based expression vector
pCAMBIA3300 were digested with SacI and BamHI (restriction
endonucleases from Takara Biotechnology Co., Ltd.), and the enzyme
digestion system was as follows: 2 µl SacI, 1 µl BamHI, 8 µl
carrier (fragment), 2 µl 10X BamHI buffer and 7 µl
DDH2O; these were incubated at 37°C for 2 h, and
inactivated at 80°C. Enzyme digestion products were separated by 1%
agarose gel electrophoresis, and then the vector and the target
fragment were collected and ligated in the following system: 4.5 µl
CHR3 DNA fragment, 2.5 µl pCAMBIA3300 vector, 1 µl T4 ligase, 2 µl
T4 buffer and 10 µl DDH2O, incubated at 22°C overnight.
The expression vector pCAMBIA3300-CHR3 was produced and
anti-herbicide Bar gene used as a screening marker. The recombinant
expression vector was identified by PCR and double enzymic
digestion.
Genetic transformation of soybean
In this experiment, the pCAMBIA3300-CHR3 DNA was
transferred into the receptor soybean cultivar Jinong 17 and Jilin
30 by Agrobacterium infection (17,18),
thus obtaining positive plants. At the time of Agrobacterium
infection, only the T-DNA region was transferred into the recipient
soybean, with the Bar gene as the marker gene, thereby obtaining a
positive plant with herbicide resistance.
Progeny analysis of transgenic
plants
PCR detection
The pCAMBIA3300 plasmid vector contained
constitutive promoters 35S and marker gene Bar. Primers for the
resistance gene Bar (552 bp) and promoter 35S (500 bp) sequences
(Bar sense/Bar antisense and 35S sense/35S antisense) were designed
by Primer software version 5.00 (Premier Biosoft International,
Palo Alto, CA, USA; Table I).
A Nuclean Plant Genomic DNA kit (CW Biotech,
Beijing, China) was used to extract genomic DNA from young soybean
leaves, and the soybean leaves genome of untransformed plants were
used as a negative control. The PCR reaction volume for Bar
was 25 µl: 2.5 µl MgCl2, 2.5 µl Buffer, 1 µl BarS, 1 µl
BarAS, 1 µl genome DNA, 0.8 µl dNTP, 0.2 µl Taq and
DDH2O to 25 µl. The PCR reaction conditions were as
follows: 94°C predenaturation for 5 min, 94°C denaturation for 40
sec, 60°C renaturation for 40 sec and 72°C extension for 40 sec for
30 cycles; the last extension step was at 72°C for 8 min and then
maintained at 4°C. The PCR reaction volume for 35S was 25 µl: 2.5
µl MgCl2, 2.5 µl Buffer, 1 µl 35S, 1 µl 35AS, 1 µl
genome DNA, 0.8 µl dNTP, 0.2 µl Taq and DDH2O to 25 µl.
The reaction conditions of PCR were: 94°C predenaturation for 5
min, 94°C denaturation for 30 sec, 55°C renaturation for 30 sec and
72°C extension for 30 sec for 40 cycles; a final extension step was
performed at 72°C for 8 min, and then maintained at 4°C. PCR
products were separated by 1% agarose gel electrophoresis and
sequenced after recovery using a AxyPrep DNA Gel Extraction kit
(Corning Life Sciences, Wujiang, China).
Southern blotting detection of transgenic
plants
PCR was performed for the initial detection of the
target gene integration and to identify positive plants for
Southern blotting to further verify the integration of the target
gene at the genome level. This enabled the number of copies of the
gene of interest in the genome to be detected.
The genomic DNA of positive T1 generation transgenic
plants was extracted using a Nuclean Plant Genomic DNA kit (CW
Biotech, Beijing, China) and then digested using BamHI.
Southern blotting was conducted with probe labeling, sample
preparation, transfer of DNA to membrane, hybridization, washing
the membrane and signal development performed according to the
manufacturer's protocols. Purified 35S DNA (https://www.ncbi.nlm.nih.gov/nucleotide/1050047859?report=genbank&log$=nuclalign&blast_rank=1&RID=NXN567FN014)
was used as the DNA probe and the DIG High Primer DNA Labeling and
Detection Starter kit I (Roche Diagnostics, Basel, Switzerland) was
used.
Reverse transcription-quantitative PCR (RT-qPCR)
detection of transgenic plants
Total RNA was extracted from leaves of transgenic
plants which were detected by Southern blotting. These plants were
tested by RT-qPCR to verify the integration of the target gene at
the mRNA level. A Total RNA Extraction kit (Omega Bio-Tek, Inc.,
Norcross, GA, USA) was used and then reverse transcribed into cDNA
using the All-in-One™ First-Strand cDNA Synthesis kit
(GeneCopoeia, Inc., Rockville, MD, USA). The reaction volume was 25
µl; 1 µg 250 µM Total RNA, 1 µl 60 µM Random Primer, 1 µl
Oligo(dT)18 and DDH2O to 13 µl, heated to
65°C for 10 min and put in an ice bath. Then was added 5 µl 25 mM
5*RT Reaction Buffer, 1 µl 25 U dNTP, 1 µl 200 U RNase Inhibitor, 1
µl M-MLV RTase and DDH2O to 25 µl. This was diluted
5-fold for subsequent use. The RT-qPCR primer sequences for CHR3
(Q-CHR3) are presented in Table I.
Soybean β-actin gene (GenBank accession number: NM_001252731.2) was
selected as the reference gene and appropriate primers were
designed (Q-ACT; Table I). The
total cDNA of the soybean leaf tissue was analyzed by 3000P Mx
fluorescence Real-time RT-qPCR instrument (Agilent StrataGene
Mx3000P) (19,20) according to the protocols of the
SYBR Premix Ex Taq™ kit (Takara Biotechnology Co.,
Ltd.). The PCR amplification system was as follows: 12.5 µl 2X SYBR
Premix Ex Taq polymerase, 1 µl Q-CHR3 sense primer, 1 µl Q-CHR3
reverse primer, 2 µl cDNA and sterile water to 25 µl. PCR
amplification conditions were as follows: 95°C predenaturation for
3 min, followed by 40 cycles of 95°C denaturation for 10 sec and
60°C reaction for 35 sec. Analysis of relative gene expression data
was performed by the 2−ΔΔCq method (21,22).
Determination of isoliquiritigenin
production
The content of isoliquiritigenin was measured using
high-performance liquid chromatography (HPLC). Soybean leaves were
treated at 70°C to dry them, and untransformed soybean leaves were
used as the control. Leaves (0.5 g dry weight) were ground to a
fine powder in liquid nitrogen and dissolved in methanol (methanol:
sample 4:1 v/v), and then exposed to ultrasonic treatment at 40°C
for 50 min following soaking overnight in methanol. Ethyl acetate
was used to extract the distribution of the isoliquiritigenin in
the enzyme hydrolysate, and then the ethyl acetate was extracted.
The sample was dissolved in methanol solution, and filtered using a
0.22 µm organic membrane. A 20 µl sample volume was analyzed using
HPLC (23,24). A Shimadzu LC-20AT HPLC system
(Shimadzu Corporation, Kyoto, Japan) was used. Detection was
performed with a fluorescence spectrometer (Beijing Jitian
Instrument Co., Ltd., Beijing, China) with excitation at 366 nm and
emission at 417 nm. The C18 column (GL Sciences Inc., Tokyo, Japan;
5 µm, 4.6×150 mm) was used at room temperature. The mobile phase
was methanol: H2O 80:20 (0–5 min, 30% methanol; 5–20
min, 45% methanol; 20–30 min, 45% methanol; 30–35 min, 30%
methanol; and, 35–40 min, 30% methanol). The flow rate was 0.8
ml/min.
Statistical analysis
Significant differences of CHR3 gene expression
between the transgenic plants and the non-transformed plants
determined by RT-qPCR were analyzed by one-way analysis of variance
followed by LSD post hoc test using SPSS version 19.0 (IBM Corp.,
Armonk, NY, USA). P<0.05 was considered to indicate a
statistically significant difference.
Results
Cloning of soybean chalcone reductase
gene CHR3 fragment and construction of over expression vector
Cloned BL21-pET28a-CHR3 plasmid was used as the
template, and the objective fragment was amplified using specific
primers (CHR3 sense/CHR3 antisense). The amplified fragment length
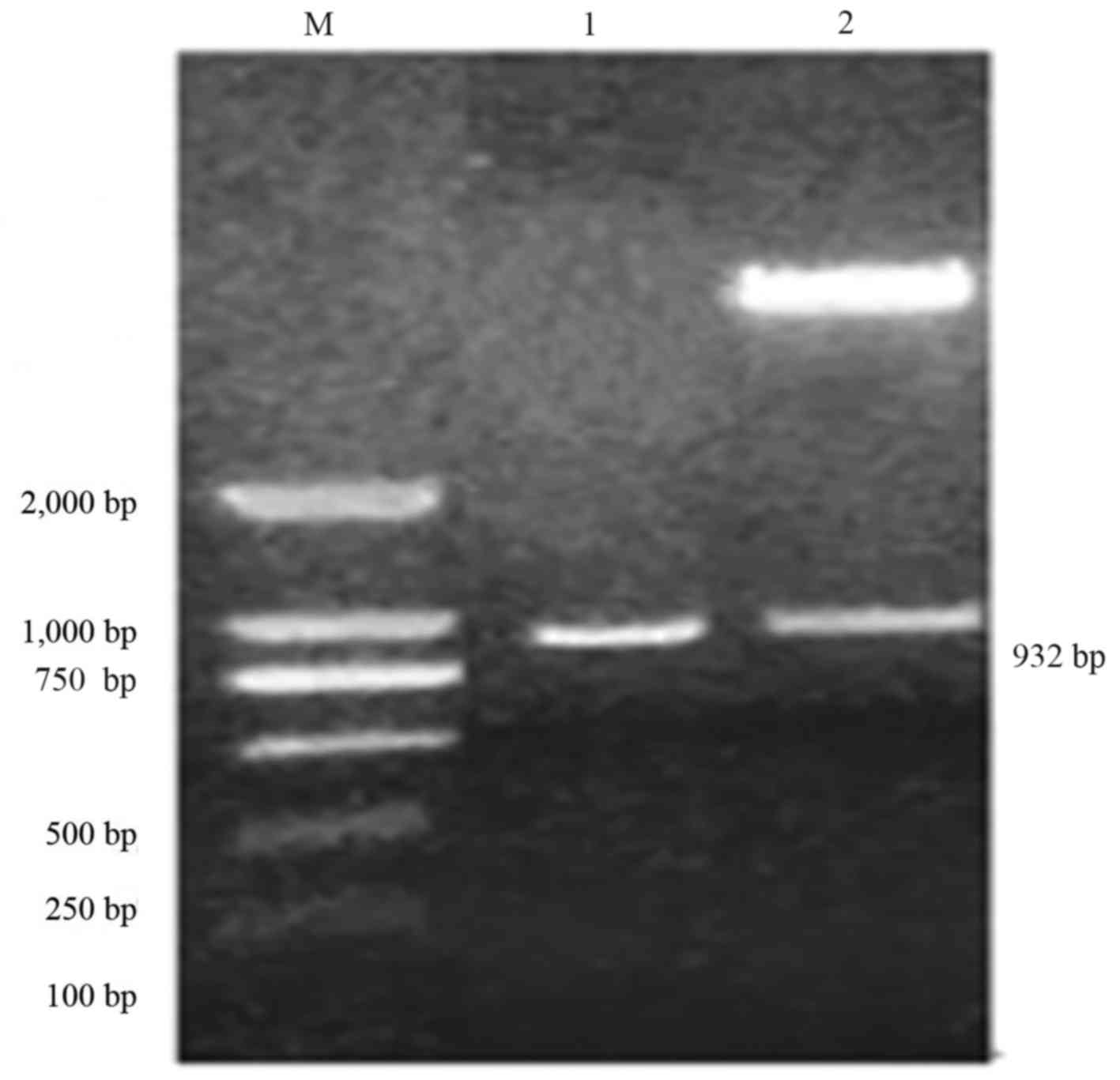
was 932 bp. The amplified fragment was cloned into pCAMBIA3300 to
obtain an overexpression vector, pCAMBIA3300-CHR3. The target
fragment was 932 bp was identified using SacI/BamHI double enzyme
digestion. As presented in Fig. 1,
results of PCR and double enzyme digestion were consistent with the
expected size, demonstrating the success of vector
construction.
Creation and detection of T1
generation transgenic plants
PCR analysis of T1 generation plants
The pCAMBIA3300-CHR3 overexpression vector was
transferred into the soybean varieties Jinong 17 and Jilin 30.
There were four positive Jinong 17 plants in the T0 generation and
two positive Jilin 30 plants in the T0 generation, as detected by
PCR. From the T0 generation, 45 Jinong 17 seed grains were
harvested and18 Jilin 30 18 grains were harvested. Genomic DNA was
extracted from T1 generation plants, and 35S promoter sequences and
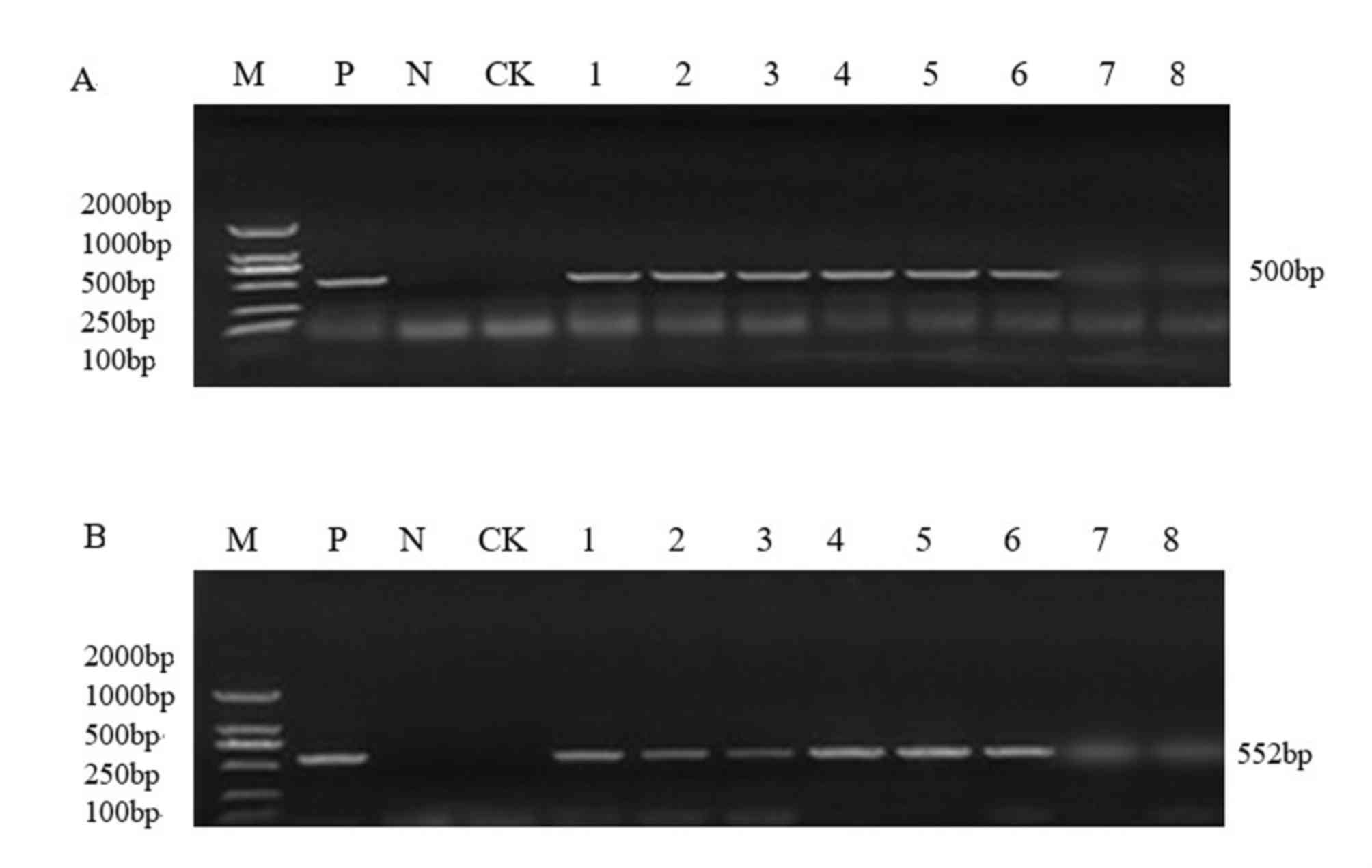
the Bar gene were detected by PCR using specific primers (Figs. 2 and 3). Recombinant expression plasmid DNA
vector pCAMBIA3300-CHR3 was the positive control and the
untransformed receptor soybean plants were a negative control.
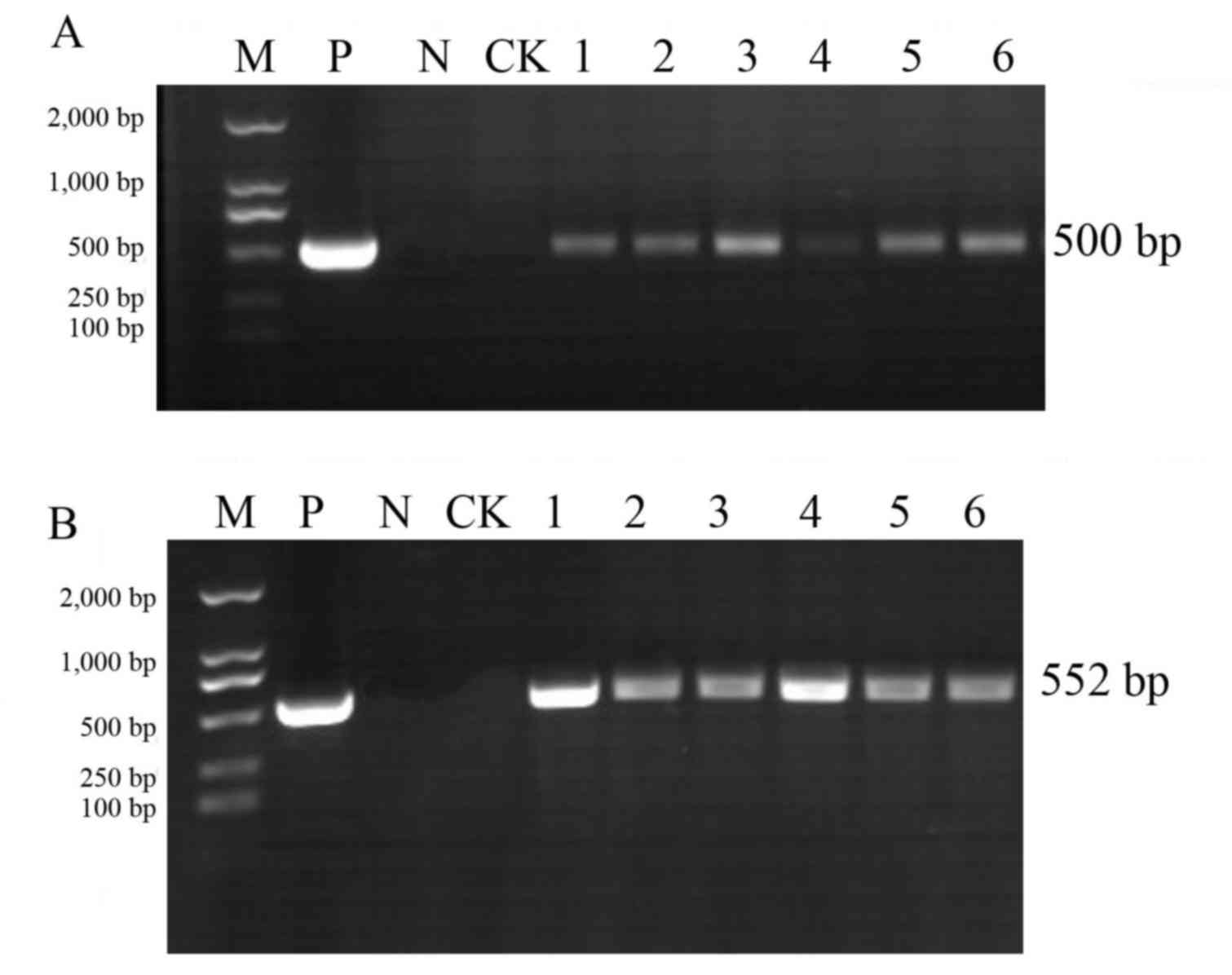
As presented in Figs.
2 and 3, PCR analysis of the
T1 generation transgenic plants produced the amplified 35S
and Bar bands at the correct estimated size (35S, 500
bp; Bar, 552 bp). Among them, Jinong 17 had 8 positive
strains (Fig. 2) and Jilin 30 had
6positive strains (Fig. 3).
Southern blot analysis of T1 plants
Genomic DNA of positive transgenic plants was
extracted and digested with BamHI. Southern blotting was
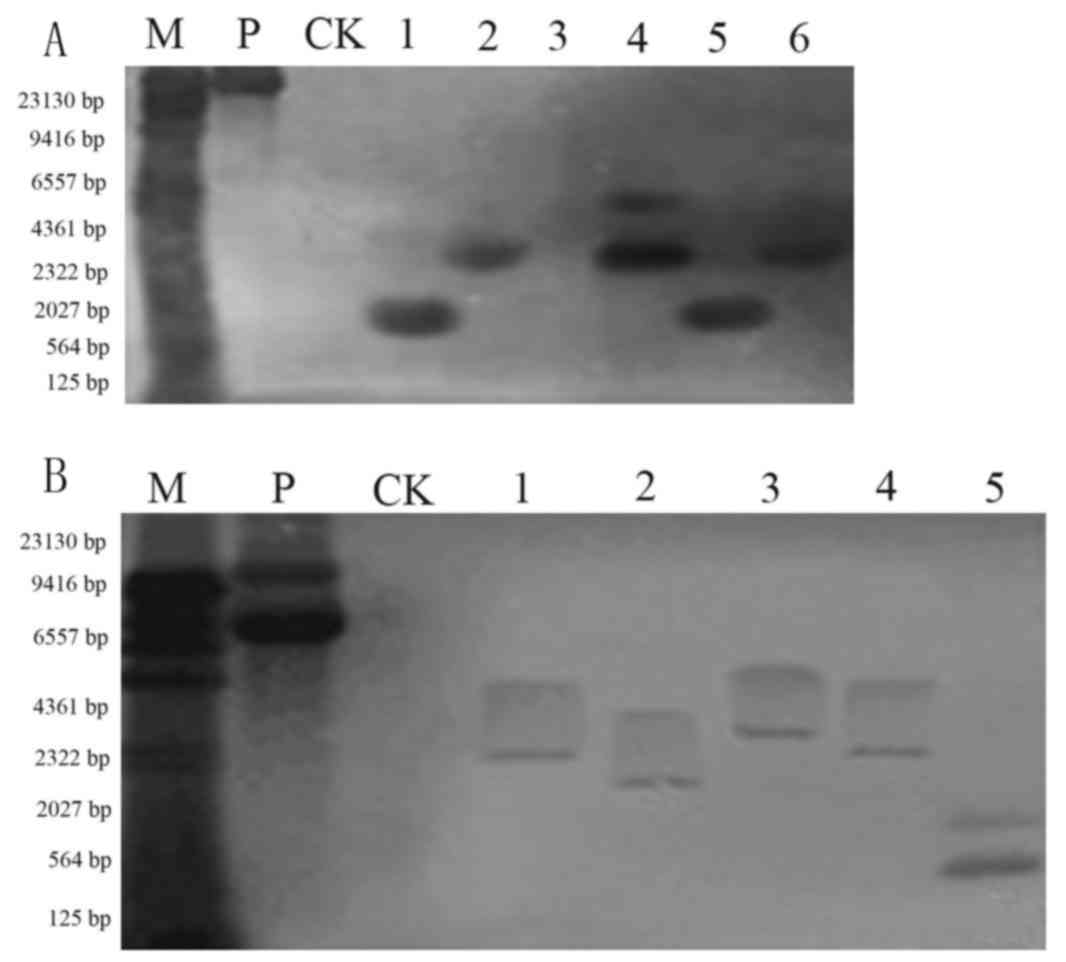
performed using the purified 35S DNA as a probe. As presented in
Fig. 4, the non-transformed plant
did not produce hybridization signals. There was observable
hybridization in Jinong 17 (Fig.
4A) and Jilin 30 (Fig. 4B)
transgenic plants. The Southern blotting indicated that the
functional components were integrated into the soybean genome, but
that the integration site was not the same in each plant.
RT-qPCR detection of transgenic plants
Positive transgenic plants detected using Southern
blotting were analyzed by RT-qPCR with SYBR Green I. As presented
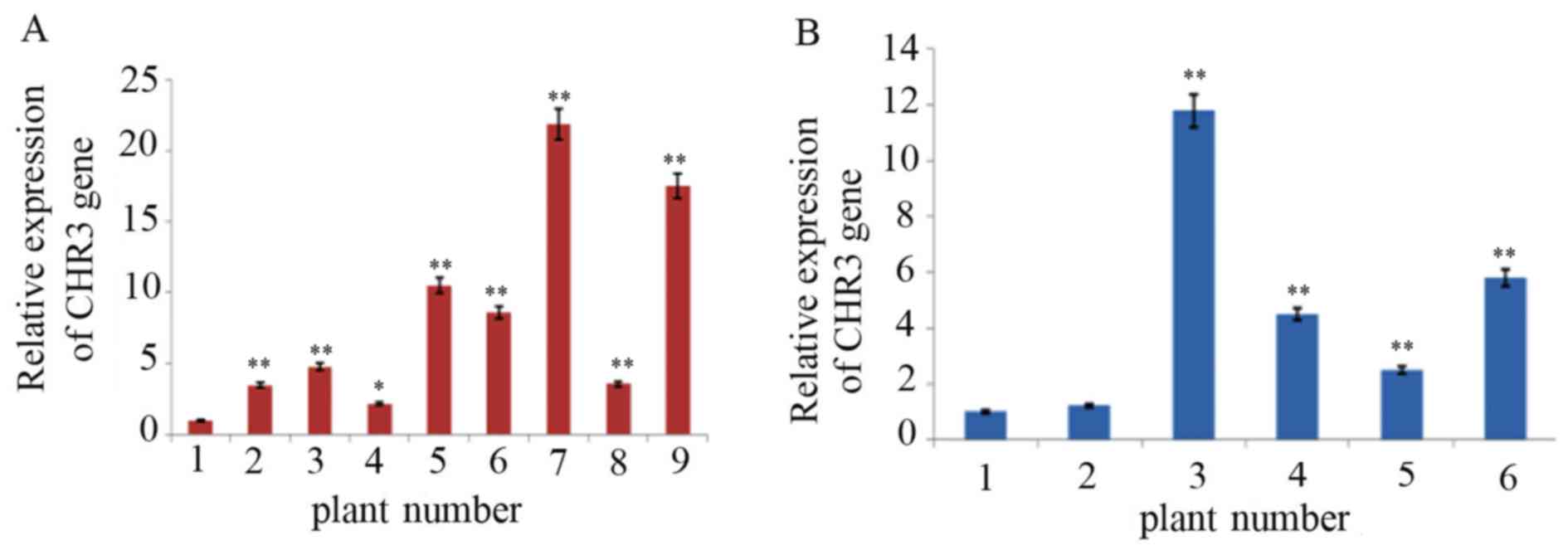
in Fig. 5, the relative expression
CHR3 mRNA in transgenic soybean plants was significantly increased
compared with control plants, and the difference ranged from 2 to
20-fold.
The average relative expression of CHR3 in
the leaf tissue of transgenic Jinong17 plants 2–9 was 3.2, 4.8,
2.0, 10.2, 8.6, 20.0, 3.3 and 17.2-fold higher than in the
non-transformed plant, respectively. The change in expression in
transgenic plant 4 compared with the non-transformed plants reached
a significance level of P<0.05; others reached P<0.01. The
average relative expression of CHR3 in the leaf tissue of
transgenic Jilin 30 plants 2–6 was 1.2, 10.5, 4.3, 2.4 and 5.5-fold
higher than in the non-transformed plants, respectively.
CHR3 expression was increased significantly in transgenic
plants 3–6 compared with the non-transformed plant (P<0.01).
The isoliquiritigenin content of T1 generation
transgenic plants
The Jinong 17 plant 6 and non-transformed soybean
leaf tissue were selected and their isoliquiritigenin content was
measured by HPLC. According to the regression equation: Y=2,
52828×106X+0.223424, r=0.999 (X represents the content of
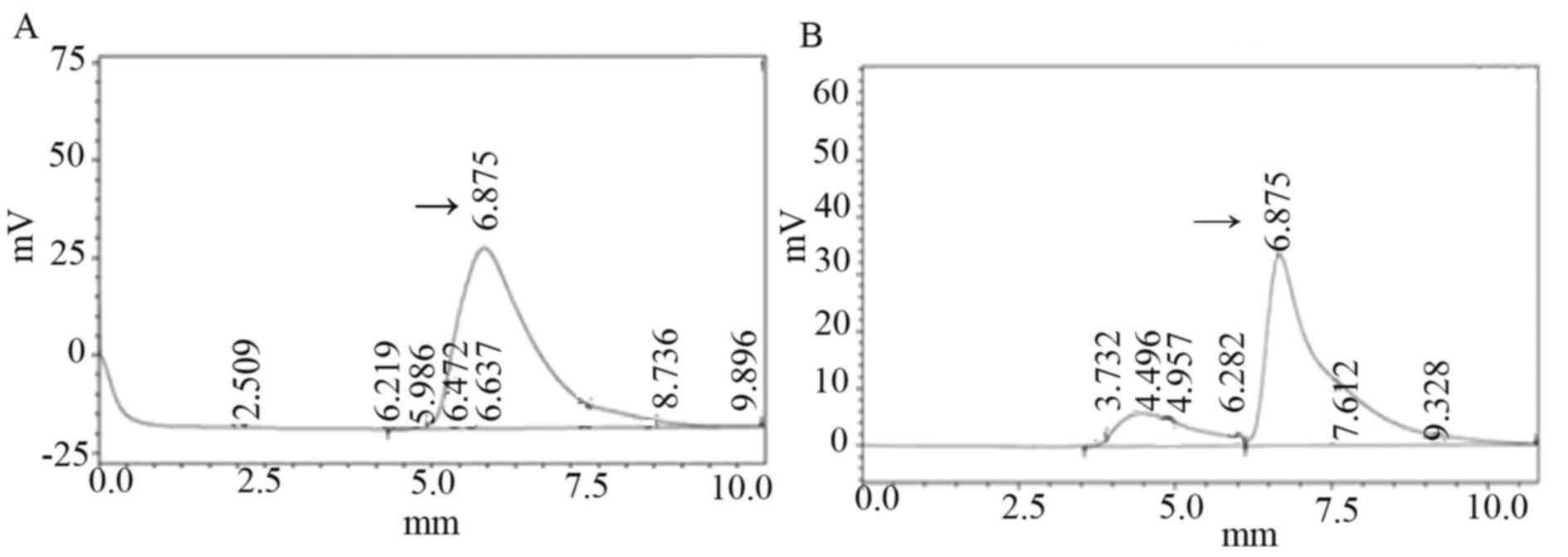
isoliquiritigenin; Y represents the peak area). As presented in
Fig. 6A, isoliquiritigenin content
of transformed plants was 1.256 µg/ml; in Fig. 6B, the isoliquiritigenin content of
untransformed plant leaf tissue was 1.157 µg/ml. The
isoliquiritigenin content was increased by 8.56% in transformed
Jinong 17 plant 6; however, no obvious increase was observed in
isoliquiritigenin content in the transgenic Jilin 30 plant compared
with non-transformed plants.
Discussion
Previous studies reported that there are five
CHR genotypes in alfalfa (25,26).
Young et al (27)
identified six CHR genotypes in tribulus alfalfa. Shimada
et al (28) cloned the
polyketoreductase gene, that is homologous with the chalcone
ketoreductase gene in Lotus japonicus, which was
overexpressed in morning glory [Ipomoea nil (L.) Roth]
suggesting it promotes isoliquiritigenin production. Li et
al (12) cloned the
CHR1 gene, constructed an overexpression vector and
transformed soybean plants. Tissues in the transgenic plants
exhibited increased CHR1 gene expression in soybean leaves.
In the present study, RT-qPCR analysis demonstrated that the
expression of CHR3 was increased by 2–20-fold in transgenic
plants compared with non-transformed plants.
pCAMBIA3300-CHR3 was introduced into the soybean
genome by Agrobacterium-mediated transformation, which was
confirmed using Southern blotting. Zhang et al (16) cloned a CHR3 gene from
soybean and transformed into E. coli BL21, which expressed a
protein that catalyzed the production of isoliquiritigenin in
soybean powder. The expression of CHR3 in the transgenic
Jinong 17 plant 6 was increased by 21.1-fold compared with control
plants; however, isoliquiritigenin content was only increased by
8.56% in the present study, as detected by HPLC; the reason for
this requires further investigation. Isoliquiritigenin is a
precursor of daidzein, which in turn can improve the ability of
soybean to resist phytophthora root rot.
References
|
1
|
Sun JM, Ding AL and Dong HR: High
performance liquid chromatographic determination of isoflavone
content in soybean test samples. Soybean Sci. 19:15–20. 2000.(In
Chinese).
|
|
2
|
Wang Y, Wu L, Sun MY, Zhao X, Han YP, Teng
WL and Li WB: Analysis of gene expression underlying soybean
isoflavone synthesis relative enzymes at different growth stages.
Soybean Sci. 31:887–893. 2012.(In Chinese).
|
|
3
|
Huang JX, Qu LJ, Yang J, Yin H and Gu HY:
A preliminary study on the origin and evolution of chalcone
synthase (CHS)gene in angiosperms. Acta Botanic Sinica. 46:10–19.
2004.
|
|
4
|
Choi EJ and Kim GH: Daidzein causes cell
cycle arrest at the G1 and G2/M phases in human breast cancer MCF-7
and MDA-MB-453 cells. Phytomedicine. 15:683–690. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Gao Y, Gu W, Chen L, Xu Z and Li Y: The
role of daidzein-loaded sterically stabilized solid lipid
nanoparticles in therapy forcardio-cerebrovascular diseases.
Biomaterials. 29:4129–4136. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Graham MY: The diphenylether herbicide
lactofen induces cell death and expression of defense-related genes
in soybean. Plant Physiol. 139:1784–1794. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Choi EJ and Kim GH: Daidzein causes cell
cycle arrest at the G1 and G2/M phases in human breast cancer MCF-7
and MDA-MB-453 cells. Phytomedicine. 15:683–690. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhang Z, Lui ZK, Huang Z, Ma J, Yao D, Qu
J and Wang PW: Isolation and functional characterization of soybean
chalcone reductase gene Gmchr4. Chin J Oil Crop Sci. 36:720–727.
2014.(In Chinese).
|
|
9
|
Ren HL, Ma QB, Yang CY, Song EL, Wang RP,
Ma TX, Tang YJ and Nian H: Identification of Phytophthora root rot
disease of soybean breeding materials in South China. Soybean Sci.
31:453–456. 2012.(In Chinese).
|
|
10
|
Liu J, Todd TC and Trick HN: Rapid in
plant evaluation of the root expressed transgenes in chimeric
soybean plants. Plant Cell Rep. 29:113–123. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Oliver Yu and McGonigle Brian: Metabolic
engineering of isoflavone biosynthesis. Advance Agronomy.
86:147–190. 2005. View Article : Google Scholar
|
|
12
|
Li D, et al: Clone and transformation of
chalcone reductase gene CHR1 in soybean. J Northwest Sci Tech Univ
Agriculture Forestry (Social Science). 43:2015.
|
|
13
|
Wu N, Wang P, Li D, Dai L, Zheng C, Lu S,
Cai Y, Zhang Z, Qu J and Xia H: Function of chalcone reductase gene
CHR1 in soybean. Hereditas. 36:707–712. 2014.PubMed/NCBI
|
|
14
|
Graham TL, Graham MY, Subramanian S and Yu
O: RNAi silencing of genes for elicitation or biosynthesis of
5-deoxyisoflavonoids suppresses race-specific resistance and
hypersensitive cell death in phytophthora sojae infected tissues.
Plant Physiol. 144:728–740. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Liu GY: Isolation, sequence identification
and tissue expression profile of two novel soybean (glycine max)
genes-vestitone reductase and chalcone reductase. Mol Biol Rep.
36:1991–1994. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhang C, Wang PW, Zhang Z, et al: Clone of
CHR3 and its activity analysis in vitro. J Northwest Sci Tech Univ
Agriculture Forestry (Social Science). 43:2015.
|
|
17
|
Zhu C, Wu J and He C: Induction of
chromosomal inversion by integration of T-DNA in the rice genome. J
Genet Genomics. 37:189–196. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhu QH, Ramm K, Eamens AL, Dennis ES and
Upadhyaya NM: Transgene structures suggest that multiple mechanisms
are involved in T-DNA integration in plants. Plant Sci.
171:308–322. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yu L, Liu JP, Zhuang JX, Yang LQ, Zhang
RL, Ye XM and Cheng JQ: Quantitative analysis of real-time PCR
expression production by REST (C) and 2(-Delta Delta C(T)). J
Tropical Med. 7:956–968. 2007.(In Chinese).
|
|
20
|
Liu BF, Zhang YM, Yan YQ and Xu XL:
Screening transgenic tobacco Byreal-Time fluorescent quantification
Pcr technology. J Natural Sci J Harbin Normal Univ. 21:69–72.
2012.
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−ΔΔCT) method. Methods. 25:402–408. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Li YY: High performance liquid
chromatography applied to the chief ingredients of
eucommiaulmoidesoliv and isoflavones in food analysis. Chongqing
Southwest Univ. 1–53. 2012.
|
|
24
|
Gu Y: Study of detection of soybean
isoflavone using HPLC. Haerbin Northeastern Univ. 1–43. 2011.
|
|
25
|
Ballance GM and Dixon RA: Medicago sativa
cDNAs encoding chalcone reductase. Plant Physiol. 107:1027–1028.
2005. View Article : Google Scholar
|
|
26
|
Sallaud C, EL-Turk J, Breda C, Buffard D,
da I kozak, Wsnault R and Kondorosi A: Differential expression of
cDNA coding for chalcone reductase, a key enzyme of the
5-deoxyflavonoid pathway, under various stress conditions in
Medicago sativa. Plant Science. 109:179–190. 2005. View Article : Google Scholar
|
|
27
|
Young ND, Debellé F, Oldroyd GE, Geurts R,
Cannon SB, Udvardi MK, Benedito VA, Mayer KF, Gouzy J, Schoof H, et
al: The Medicago genome provides insight into the evolution of
rhizobial symbioses. Nature. 480:520–524. 2011.PubMed/NCBI
|
|
28
|
Shimada N, Aoki T, Sato S, Nakamura Y,
Tabata S and Ayabe S: A cluster of genes encodes the two types of
chalcone isomerase involved in the biosynthesis of general
flavonoids and legume-specific 5-deoxy(iso)flavonoids in Lotus
japonicus. Plant Physiol. 131:941–951. 2003. View Article : Google Scholar : PubMed/NCBI
|