Introduction
Cervical cancer is one of the most common
malignancies that occur in the cervical canal (1). It is estimated that ~528,000 cases of
cervical cancer, with 266,000 mortalities, occur every year
(2). Although not all of the
causes of cervical cancer are known, human papillomavirus
infections is considered to be the main risk factor (3). Currently, the standard tumor
treatments for cancer include surgery, radiation and chemotherapy,
alone or in combination (4).
Although treatment strategies are improving, the 5-year survival
rate for cervical cancer patients is <40% (5). Therefore, to further study cervical
cancer development at the molecular level is necessary.
Circular RNAs (circRNAs) are single-stranded closed
ring-like non-coding RNAs, with no 5′-terminal cap or a 3′-terminal
poly-A tail (6). Due to their
special stable structure, circRNAs cannot be degraded by RNA
exonucleases (7). circRNAs have
been shown to function as competitive endogenous RNAs and microRNA
(miRNA) sponges (8,9). Recent studies have shown that altered
circRNA levels play crucial roles in carcinogenesis (10,11).
Circular RNA SMARCA5 (cSMARCA5; circBase ID: hsa_circ_0001445) is a
novel circRNA derived from exons 15 and 16 of the SMARCA5 gene
(12). A previous study reported
that overexpression of cSMARCA5 could inhibit the proliferation and
metastasis of hepatocellular carcinoma cells (13). However, to the best of our
knowledge, the role of cSMARCA5 in cervical cancer has not been
previously investigated.
miRNAs are a class of evolutionarily conserved,
short non-coding RNA molecules of 18–25 nucleotides in length
(14). miRNAs can modulate gene
expression by binding to the 3′-untranslated regions (3′-UTRs) of
target mRNAs, resulting in repression of protein translation or
mRNA degradation (15,16). Previous studies have shown that
miRNAs are involved in multiple cellular processes, including cell
proliferation, cell cycle, apoptosis and cell differentiation
(17). Abnormal expression of
miRNAs has been identified in various types of malignancies
(18–20). Several miRNAs have been
demonstrated to be involved in cervical cancer initiation and
progression (21–23). miRNA-432 (miR-432) dysregulation
has been shown to be involved in the carcinogenesis of many cancers
(24–26). However, a limited number of studies
have reported the functional role of miR-432 in cervical cancer
development.
The ERK pathway is involved in the regulation of a
variety of growth and differentiation pathways through several
phosphorylation cascades (27).
Uncontrolled growth is a necessary step for the development of all
cancers (28). In many cancer
types, a defect in the mitogen-activated protein/ERK pathway is
believed to contribute to uncontrolled proliferation (29). Upregulation of epidermal growth
factor receptor (EGFR) is frequently detected in cervical cancer
and is considered to be an independent predictor for the prognosis
of cervical cancer (30).
The aim of the present study was to investigate the
level of cSMARCA5 expression in human cervical cancer tissues and
cell lines. In addition, the present study examined the function of
cSMARCA and its underlying mechanism.
Materials and methods
Tissue samples
The present study was approved by the Medical Ethics
Committee of Cangzhou Central Hospital. All patients (18–65 years)
were informed of the study and signed written informed consent.
Human cervical cancer tissues and adjacent normal tissues (n=56)
were collected from patients who visited the Cangzhou Central
Hospital from January 2016 to Decembr 2017. All the specimens were
immediately snap-frozen and preserved in liquid nitrogen at −80°C
until further use.
Cell culture and transfection
The human normal cervical epithelial cell line
(Ect1/E6E7) and human cervical cancer cell lines HeLa, Ca-Ski,
C-33A, and SiHa were purchased from VCANBIO Cell & Gene
Engineering Corporation, Ltd. and cultured in DMEM with 10% FBS
(both from Invitrogen; Thermo Fisher Scientific, Inc.) at 37°C with
5% CO2. Small interfering RNA (siRNA) targeting cSMARCA5
(si-cSMARCA5; 5′-AUUGGCGACUCAAUGGAUCAG-3′), miR-432 mimic
(5′-CCUCGCGUUAUAACGUUAC-3′) and their corresponding negative
controls (NCs; si-NC, 5′-UUCUCCGAACGUGUCA-3′; miR-NC,
5′-UUCUCCGAACGUGUCACGUAA-3′) were synthesized by Shanghai
GenePharma, Co., Ltd. After culturing overnight at 37°C, HeLa and
Ca-Ski cells (4×105 cells/well) were transfected with
si-cSMARCA5, miR-432 mimic or their parental negative controls
using Lipofectamine 2000 reagent (cat. no. 11668-019; Invitrogen;
Thermo Fisher Scientific, Inc.) according to the manufacturer's
protocol. The of siRNAs and miRNAs was 50 nM. Subsequent
experiments were performed 24 h after transfection.
Reverse transcription-quantitative PCR
(RT-qPCR)
TRIzol® reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) was added to the tissue and cell samples
to extract total RNA. After that, total RNA was treated with RNase
R (Invitrogen; Thermo Fisher Scientific, Inc.) to remove linear
RNAs and enrich circRNAs. Total RNA was dissolved in RNase-free
water, and the concentration was measured using a NanoDrop™ 2000
spectrophotometer. cDNA was synthesized using a TaqMan MicroRNA
Reverse Transcription kit (Applied Biosystems; Thermo Fisher
Scientific, Inc.) for miR-432 and a One-Step PrimerScript cDNA kit
(Qiagen, Inc.) was used for cSMARCA5, EGFR, ERK1 and ERK2 using 50
ng total RNA with the temperature protocol of: 95°C for 30 sec and
60°C for 30 min. RT-qPCR was performed using SYBR Green PCR Master
Mix kit (Thermo Fisher Scientific, Inc.). The thermocycling
conditions were as follows: Initial denaturation at 95°C for 5 min;
followed by 30 cycles of 95°C for 10 sec and annealing at 60°C for
45 sec; then a final extension for 10 min at 72°C. All primers were
designed and synthesized by Shanghai GenePharma Co., Ltd. The
primers were as follows: cSMARCA5 forward,
5′-GCTATCAAGCTCCATCCGCAT-3′ and reverse, 5′-TAAGACGAAGCACCGGA-3′;
miR-432 forward, 5′-AACGAGACGACGACAGAC-3′ and reverse,
5′-CTTGGAGTAGGTCATTGGGT-3′; si-cSMARCA5 5′-CATGGTCCTCGAGGTTA-3′;
si-NC 5′-UGGACAACAUGGGCUCU-3′; miR-432 mimic:
5′-AUCGAGACUACGUCUGAC-3′; miR-NC 5′-AGUGCAUGCGUACGAGCUGU-3′; EGFR
forward, 5′-ATGGAATACCCTGGGTGT-3′ and reverse,
5′-GGACAAGCTGGTCAAGGT-3′; ERK1 forward, 5′-CCAGTTCCGAGAATAAGCGCA-3′
and reverse, 5′-CGTGTCGCCATGACACATGT-3′; ERK2 forward,
5′-TCATCCAACAGACAGACGTAGT-3′ and reverse, 5′-ACCAGAGCCATCAGACGA-3′;
U6 forward, 5′-GCTCGCTTCGGCAGCACA-3′ and reverse,
5′-GAGGTATTCGCACCAGAGGA-3′; GAPDH forward,
5′-ACCACAGTCCATGCCATCCAC-3′ and reverse,
5′-TCCACCACCCTGTTGCTGTA-3′. GAPDH mRNA or U6 were used as
endogenous reference genes. Gene expression was determined with the
2−ΔΔCq method (31).
MTT assay
Cells (3×103) were seeded in 96-well
plates, incubated at 37°C for 24 h and stained with 0.5 mg/ml MTT
at 37°C for 4 h. After removal of the supernatant, DMSO was added
and thoroughly mixed for 15 min. The absorbance value in each well
was measured at a wavelength of 490 nm.
Luciferase activity assay
Circinteractome (https://circinteractome.nia.nih.gov/) and TargetScan
7.2 (http://www.targetscan.org/vert_72/) were used to
predict potential targets of miR-432, and binding sites between
miR-432 and cSMARCA5/EGFR. The cSMARCA5 and EGFR 3′-UTR sequences
were amplified and cloned into the pGL3 vector (Promega
Corporation). For reporter assays, cells were cultured in 24-well
plates at 37°C and cotransfected with wild type (WT)-cSMARCA5 or
mutant (Mut)-cSMARCA5 (WT-EGFR or Mut EGFR) and 100 nM miR-432
mimics or miR-NC, using Lipofectamine® 2000 (Invitrogen;
Thermo Fisher Scientific, Inc.). After 48 h, the luciferase
activity was determined with a Dual-Luciferase Reporter System
(Promega Corporation) according to the manufacturer's protocols.
Firefly luciferase activity was normalized to Renilla
luciferase activity using the pGL3 vector.
Cell invasion assay
For cell invasion assays, 1×105 HeLa and
Ca-Ski cells were resuspended in 200 µl serum-free RPMI-1640 medium
(Gibco; Thermo Fisher Scientific, Inc.) and then plated into the
upper chambers of Transwell inserts, which were coated with
Matrigel. The lower chamber was filled with culture medium
supplemented with 20% FBS (Gibco; Thermo Fisher Scientific, Inc.).
After 24 h of incubation at 37°C, the cells on the bottom surface
were fixed with 4% polyoxymethylene at room temperature for 30 min
and stained with 0.1% crystal violet at room temperature for 20
min. Stained cells were counted and images were captured with an
Olympus BX51 light microscope (magnification, ×200; Olympus
Corporation).
Western blotting analysis
Proteins were extracted from cultured cells by RIPA
buffer (Sigma-Aldrich; Merck KGaA) containing a mixture of protease
inhibitors (100X; Beijing CoWin Biotech Co., Ltd.). Protein
concentrations were quantified using a bicinchoninic acid assay
(Beijing CoWin Biotech Co., Ltd.). Equal quantities of protein (30
µg/lane) were separated via 10% SDS-PAGE and then transferred to
PVDF membranes. The membranes were blocked with 5% skimmed milk at
room temperature for 1.5 h, followed by incubation with the
following primary antibodies overnight at 4°C: EGFR (1:1,000; cat.
no. ab32562; Abcam) and GAPDH (1:5,000; cat. no. ab185059; Abcam).
Then, a horseradish peroxidase-conjugated goat anti-rabbit IgG
secondary antibody (1:5,000; cat. no. sc-2054; Santa Cruz
Biotechnology, Inc.) was incubated with the PVDF membranes for 1 h
at room temperature. Protein signals were visualized using an
Enhanced Chemiluminescence Plus reagent (GE Healthcare Life
Sciences). Bands were quantified using Quantity One version 4.62
software (Bio-Rad Laboratories, Inc.).
Statistical analysis
Data are presented as the mean ± SEM. All
statistical analyses were performed using SPSS software (version
19.0; IBM Corp) and GraphPad Prism software (version 5.0; GraphPad
Software, Inc.). Data were analyzed using a Student's t-test for
two-group comparisons, and one-way ANOVA with a Tukey's post-hoc
test for multiple-group comparison. Spearman's correlation analysis
was used to analyze the association between cSMARCA5 and ERK1 or
ERK2 expression. P<0.05 was considered to indicate a
statistically significant difference.
Results
cSMARCA5 expression is upregulated in
cervical cancer tissues and cell lines
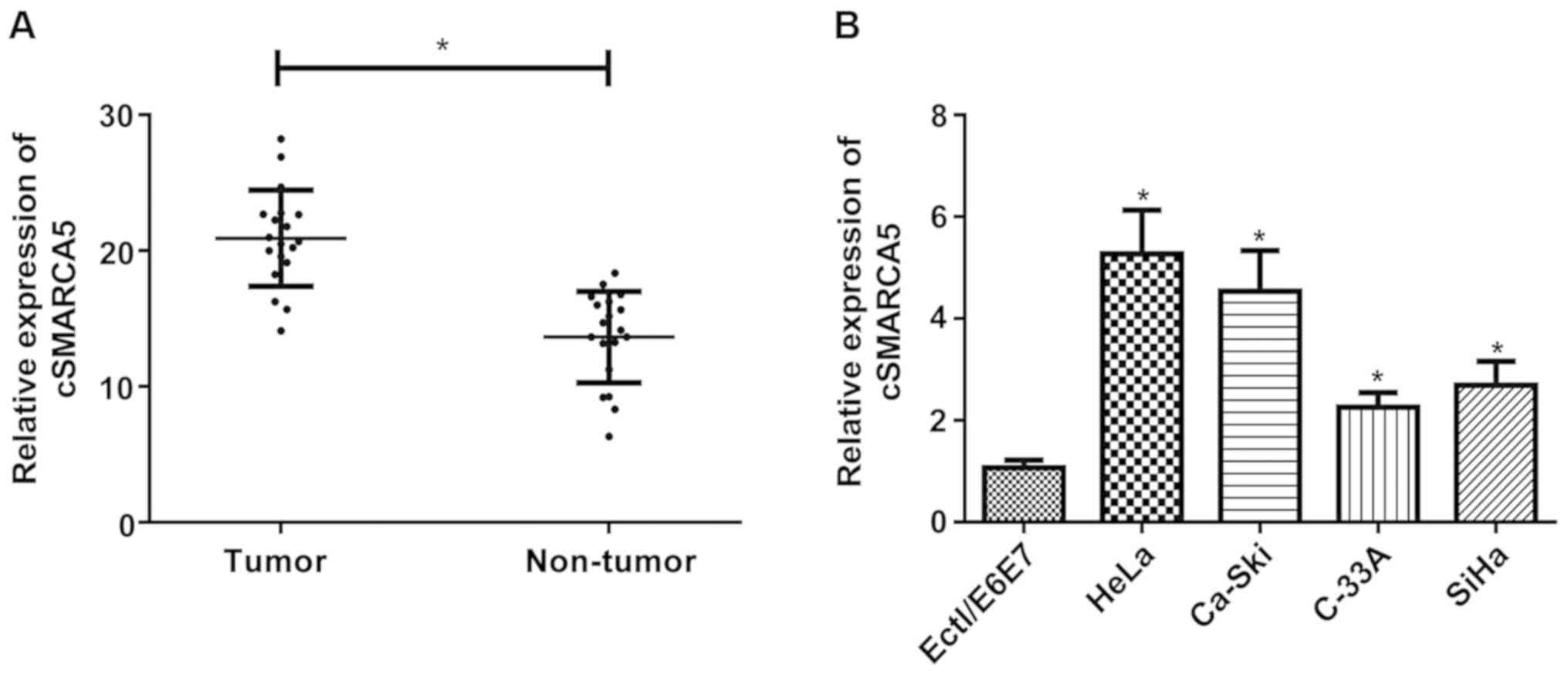
cSMARCA5 expression level in cervical cancer tissues
was investigated using RT-qPCR. The expression level of cSMARCA5
was significantly increased in cervical cancer tissues (Fig. 1A) compared with non-tumor tissues.
In addition, the present study examined the expression level of
cSMARCA5 in four cervical cancer cell lines (HeLa, Ca-Ski, C-33A,
and SiHa) and a human normal cervical epithelial cell line
(Ect1/E6E7). A significant increase in cSMARCA5 expression level
was found in the four cancer cell lines compared with Ect1/E6E7
cells (Fig. 1B).
cSMARCA5 silencing represses the
proliferation and invasion of cervical cancer cells
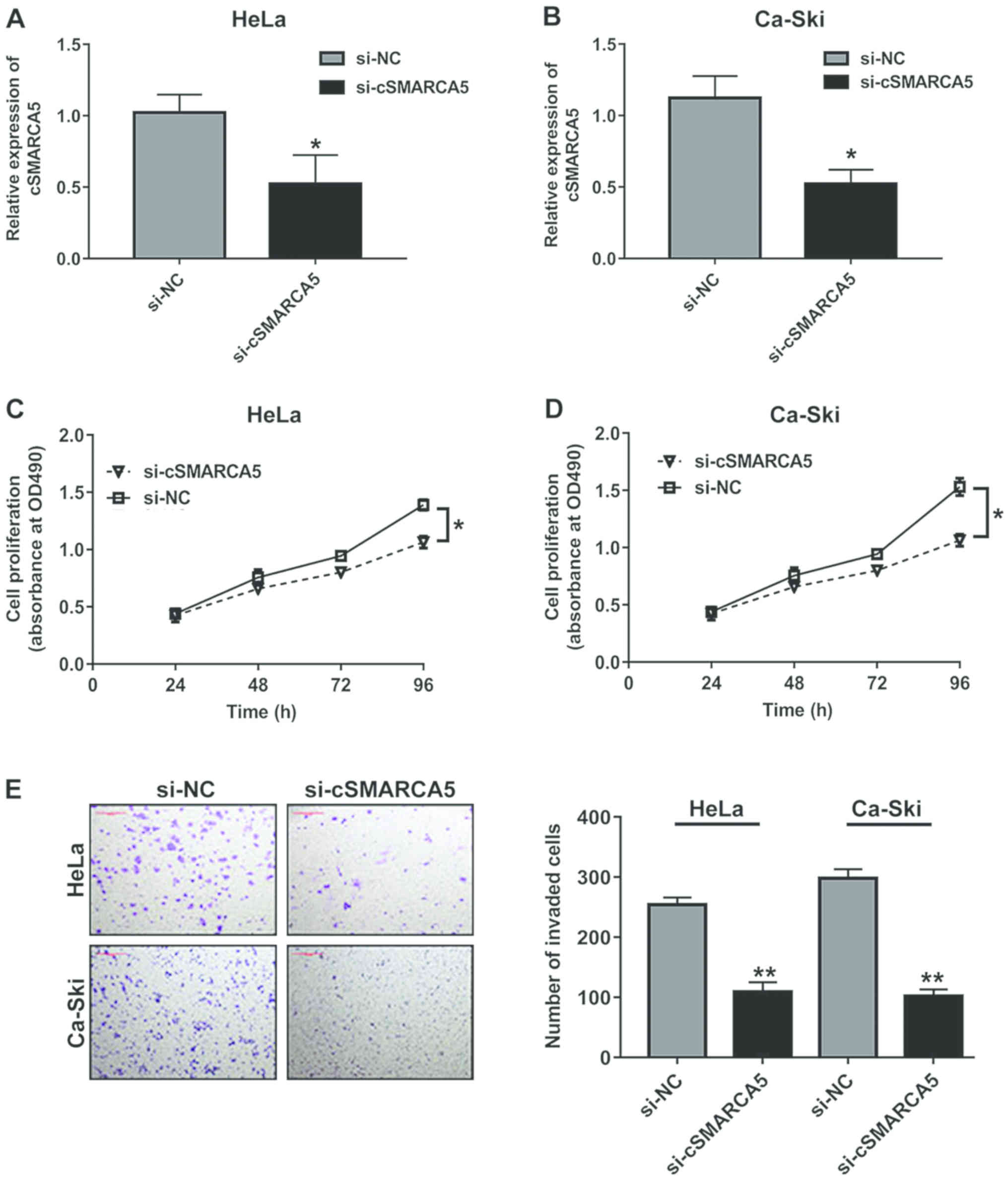
The oncogenic role of cSMARCA5 in cervical cancer
cell lines was investigated, and it was observed that cSMARCA5
expression level was remarkably decreased in two cell lines (HeLa
and Ca-Ski), which were transfected with si-cSMARCA5. The RT-qPCR
results showed that cSMARCA5 expression level was significantly
downregulated in the two cell lines transfected with si-cSMARCA5
(Fig. 2A and B). In addition, the
MTT assay revealed that the proliferation rate of cells transduced
with si-cSMARCA5 was significantly decreased at 96 h compared with
cells transduced with si-NC in the two cell lines (Fig. 2C and D). Cell invasion assays
revealed that si-cSMARCA5 suppressed the invasion of cervical
cancer lines (Fig. 2E).
miR-432 targets cSMARCA5
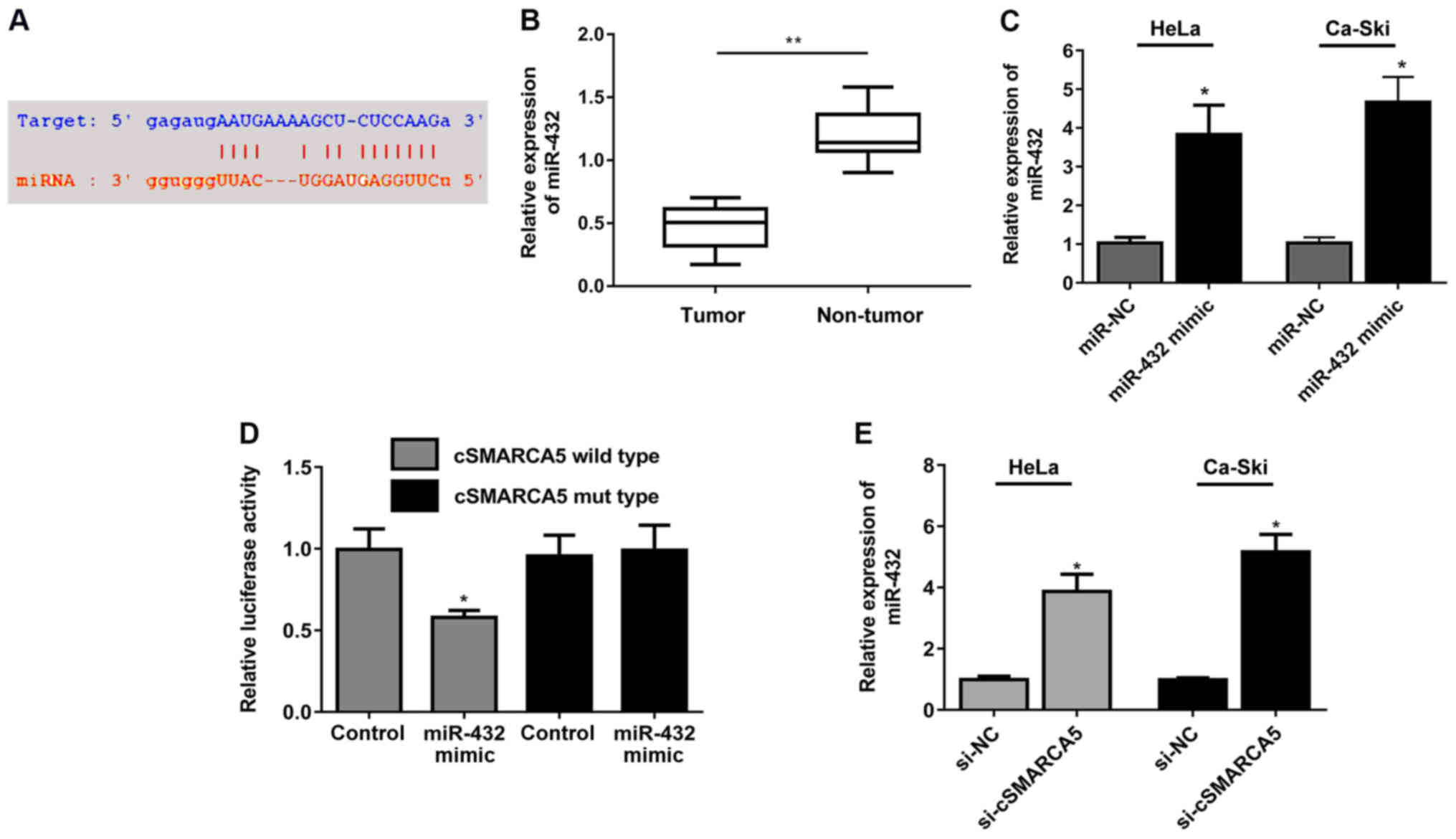
The binding sites between miR-432 and cSMARCA5 are
presented in Fig. 3A. miR-432
expression level was significantly decreased in cervical cancer
tissues (Fig. 3B). miR-432
expression was significantly increased in two cell lines (HeLa and
Ca-Ski) following transfection of miR-432 mimic (Fig. 3C). Decreased luciferase activity
was observed in cells transfected with miR-432 mimic and cSMARCA5
wild-type reporter, but not in cells transfected with the cSMARCA5
mutant reporter plasmid and miR-432 mimic (Fig. 3D). Moreover, miR-432 expression
levels were significantly increased in cells transduced with
si-cSMARCA5 compared with the control group (Fig. 3E).
miR-432 targets the 3′-UTR of
EGFR
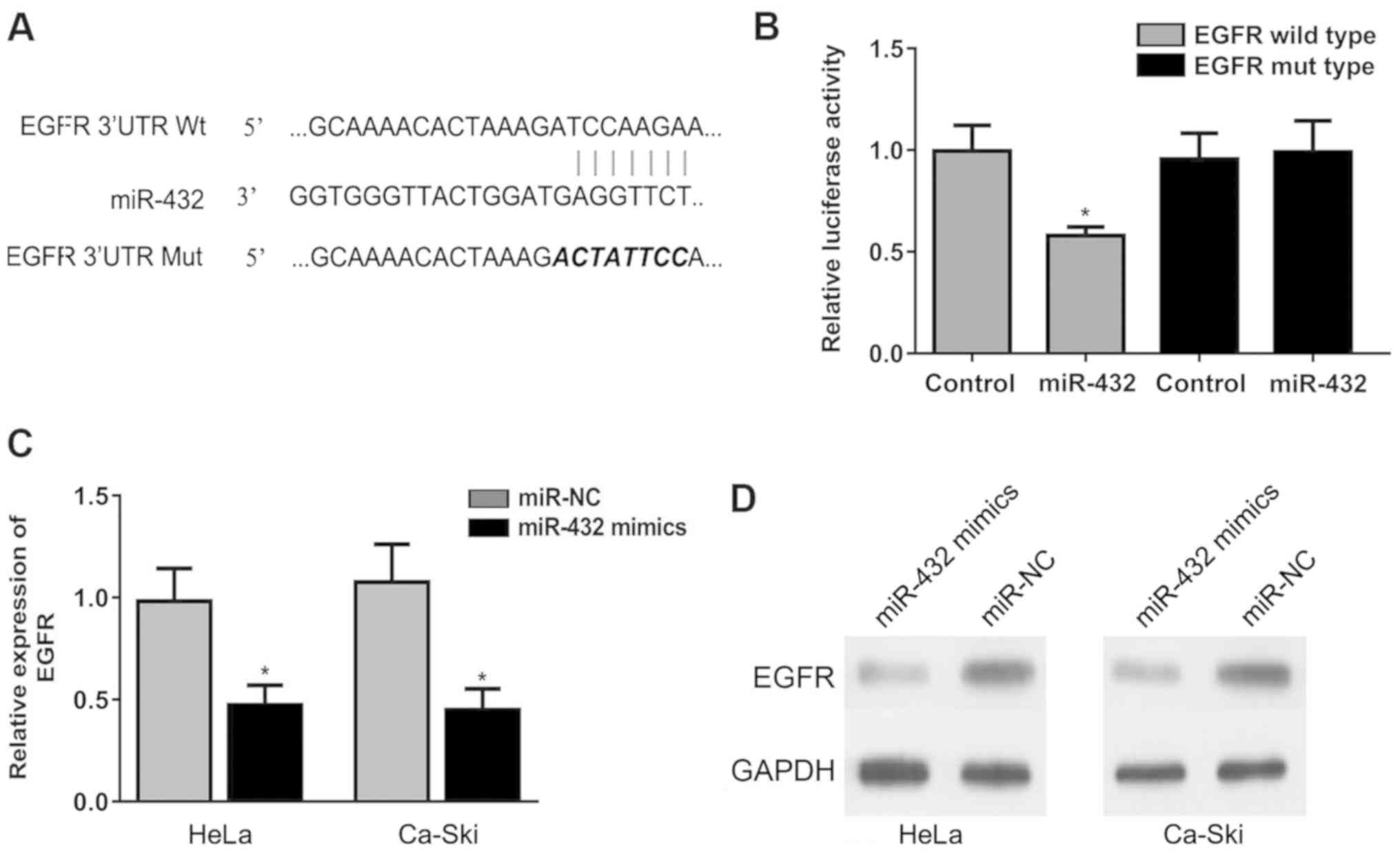
Bioinformatics analysis found that the 3′UTR of EGFR
may interact with miR-432. The binding site between miR-432 and
EGFR is shown in Fig. 4A. The
miR-432 mimic significantly reduced the relative luciferase
activity of the EGFR luciferase reporter in transfected cells
(Fig. 4B), suggesting a direct
interaction between miR-432 and EGFR. Moreover, the miR-432 mimic
significantly downregulated EGFR expression in HeLa and Ca-Ski
cells at the mRNA and protein levels (Fig. 4C and D). Collectively, the present
results suggested that miR-432 inhibited the expression of EGFR by
binding to the 3′-UTR of EGFR.
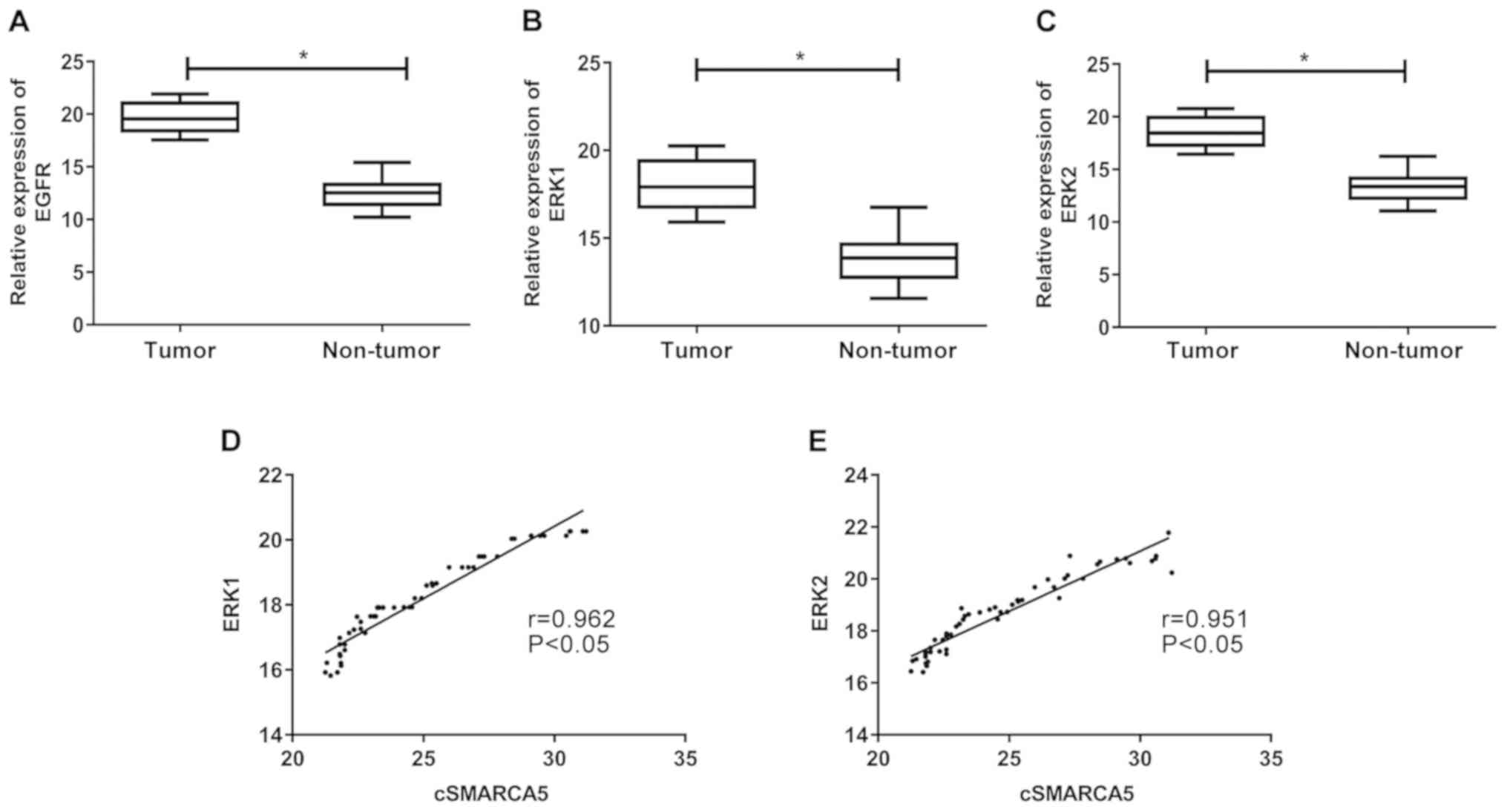
Expression levels of EGFR, ERK1 and
ERK2 in cervical cancer tissues
EGFR expression in cervical cancer tissue samples
was subsequently analyzed. EGFR levels were significantly increased
in cervical cancer tissues compared with non-tumor tissues
(Fig. 5A). The levels of ERK1 and
ERK2 expression were significantly increased in cervical cancer
tissues (Fig. 5B and C).
Furthermore, correlation analysis revealed that cSMARCA5 levels
were positively correlated with ERK1 (r=0.962, P<0.05; Fig. 5D) and ERK2 levels (r=0.951,
P<0.05; Fig. 5E).
Discussion
In recent years, with the rapid development of
bioinformatics technology, an increasing number of circRNAs have
been reported to be involved in the development and progression of
various malignant tumors (32). A
previous study showed that hsa_circ_0005075 is involved in cell
adhesion during hepatocellular carcinoma development (33). High levels of circular RNA CCDC66
promote colorectal cancer growth and metastasis by sponging miRNAs
(34). In the present study,
cSMARCA5 expression level was found to be upregulated in cervical
cancer tissues. In addition, a significant increase in the cSMARCA5
level in the four cell lines was observed in the present study. The
proliferation and invasion of tumor cells are important
characteristics that affect the progression of tumors (35). The present results suggested that
cSMARCA5 could promote cervical cancer cell proliferation and
invasion. In addition, results suggested that cSMARCA5 may present
tumor-promoting abilities in cervical cancer cells.
It is hypothesized that circRNAs modulate diverse
biological processes, acting as miRNA ‘sponges’, as well as
regulating transcription, protein binding and translation (36). In the present study, the miRNA
‘sponge’ theory was explored in relation to cSMARCA5 in cervical
cancer. Using bioinformatics analysis, it was predicted that
miR-432 was a target of cSMARCA5. The miR-432 expression level was
increased following when cSMARCA5 knockdown. The present results
suggested that cSMARCA5 may induce the progression of cervical
cancer by targeting miR-432.
The human EGFR gene is localized on the 7th
chromosome and encodes a glycoprotein composed of ~53 amino acids
that is activated by binding to specific ligands, including EGF and
transforming growth factor α (37). Biesterfeld et al (38) found that EGFR expression was
upregulated in cervical carcinomas, suggesting that the expression
of EGFR may correlate with the aggressive and proliferative
phenotype of cervical carcinoma. In line with previous research, in
the present study, upregulation of EGFR was detected in cervical
cancer tissues. Using bioinformatics analysis, the present study
showed that miR-432 directly targeted the 3′-UTR of EGFR,
suggesting that the miR-432 mimic could directly downregulate EGFR
expression.
As a highly conserved mitogen-activated protein
kinase family member, ERK plays important roles in biological
processes such as proliferation, differentiation and apoptosis
(39). A previous study revealed
that inhibition of the ERK pathway could exert antitumor effects
(40). The present results showed
that ERK1 and ERK2 expression levels increased significantly in
cervical cancer tissues. Furthermore, the present study found a
positive correlation between cSMARCA5 and ERK1/2, suggesting that
cSMARCA5 could affect the progression of cervical cancer by
upregulating the ERK1/2 signaling pathway. The present study
suggested that cSMARCA5 promoted the progression of cervical cancer
by modulating miR-432, and induced the proliferation and invasion
of cervical cancer by upregulating the ERK1/2 signaling
pathway.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
BQ, HY and LZ participated in data analysis and
manuscript preparation. YL and QL performed the experiments and PH
interpreted the data and drafted the manuscript.
Ethics approval and consent to
participate
The present study was approved by The Medical Ethics
Committee of Cangzhou Central Hospital. All patients were informed
of the study and signed written informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Pecorelli S: Revised FIGO staging for
carcinoma of the vulva, cervix, and endometrium. Int J Gynaecol
Obstet. 105:103–104. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ferlay J, Soerjomataram I, Dikshit R, Eser
S, Mathers C, Rebelo M, Parkin DM, Forman D and Bray F: Cancer
incidence and mortality worldwide: Sources, methods and major
patterns in GLOBOCAN 2012. Int J Cancer. 136:E359–E386. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Walboomers JM, Jacobs MV, Manos MM, Bosch
FX, Kummer JA, Shah KV, Snijders PJ, Peto J, Meijer CJ and Muñoz N:
Human papillomavirus is a necessary cause of invasive cervical
cancer worldwide. J Pathol. 189:12–19. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yee GP, de Souza P and Khachigian LM:
Current and potential treatments for cervical cancer. Curr Cancer
Drug Targets. 13:205–220. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Smith RA, Brooks D, Cokkinides V, Saslow D
and Brawley OW: Cancer screening in the United States, 2013: A
review of current American Cancer Society guidelines, current
issues in cancer screening, and new guidance on cervical cancer
screening and lung cancer screening. CA Cancer J Clin. 63:88–105.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chen LL and Yang L: Regulation of circRNA
biogenesis. RNA Biol. 12:381–388. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Chen LL: The biogenesis and emerging roles
of circular RNAs. Nat Rev Mol Cell Biol. 17:205–211. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Salzman J: Circular RNA expression: Its
potential regulation and function. Trends Genet. 32:309–316. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hansen TB, Jensen TI, Clausen BH, Bramsen
JB, Finsen B, Damgaard CK and Kjems J: Natural RNA circles function
as efficient microRNA sponges. Nature. 495:384–388. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zheng Q, Bao C, Guo W, Li S, Chen J, Chen
B, Luo Y, Lyu D, Li Y, Shi G, et al: Circular RNA profiling reveals
an abundant circHIPK3 that regulates cell growth by sponging
multiple miRNAs. Nat Commun. 7:112152016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Pan H, Li T, Jiang Y, Pan C, Ding Y, Huang
Z, Yu H and Kong D: Overexpression of circular RNA ciRS-7 abrogates
the tumor suppressive effect of miR-7 on gastric cancer via
PTEN/PI3K/AKT signaling pathway. J Cell Biochem. 119:440–446. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Glazar P, Papavasileiou P and Rajewsky N:
circBase: A database for circular RNAs. RNA. 20:1666–1670. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yu J, Xu QG, Wang ZG, Yang Y, Zhang L, Ma
JZ, Sun SH, Yang F and Zhou WP: Circular RNA cSMARCA5 inhibits
growth and metastasis in hepatocellular carcinoma. J Hepatol.
68:1214–1227. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Valencia-Sanchez MA, Liu J, Hannon GJ and
Parker R: Control of translation and mRNA degradation by miRNAs and
siRNAs. Genes Dev. 20:515–524. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ebert MS and Sharp PA: Roles for microRNAs
in conferring robustness to biological processes. Cell.
149:515–524. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lv D, Zhen Z and Huang D: MicroRNA-432 is
downregulated in osteosarcoma and inhibits cell proliferation and
invasion by directly targeting metastasis-associated in colon
cancer-1. Exp Ther Med. 17:919–926. 2019.PubMed/NCBI
|
|
19
|
Wu K, Ma L and Zhu J: miR4835p promotes
growth, invasion and selfrenewal of gastric cancer stem cells by
Wnt/betacatenin signaling. Mol Med Rep. 14:3421–3428. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Liu F, Cai Y, Rong X, Chen J, Zheng D,
Chen L, Zhang J, Luo R, Zhao P and Ruan J: MiR-661 promotes tumor
invasion and metastasis by directly inhibiting RB1 in non small
cell lung cancer. Mol Cancer. 16:1222017. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang X, Li F and Zhu L: Clinical
significance and functions of microRNA-93/CDKN1A axis in human
cervical cancer. Life Sci. 209:242–248. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Pedroza-Torres A, Campos-Parra AD,
Millan-Catalan O, Loissell-Baltazar YA, Zamudio-Meza H, Cantú de
León D, Montalvo-Esquivel G, Isla-Ortiz D, Herrera LA,
Ángeles-Zaragoza Ó, et al: MicroRNA-125 modulates radioresistance
through targeting p21 in cervical cancer. Oncol Rep. 39:1532–1540.
2018.PubMed/NCBI
|
|
23
|
Tan D, Zhou C, Han S, Hou X, Kang S and
Zhang Y: MicroRNA-378 enhances migration and invasion in cervical
cancer by directly targeting autophagy-related protein 12. Mol Med
Rep. 17:6319–6326. 2018.PubMed/NCBI
|
|
24
|
Jiang N, Chen WJ, Zhang JW, Xu C, Zeng XC,
Zhang T, Li Y and Wang GY: Downregulation of miR-432 activates
Wnt/β-catenin signaling and promotes human hepatocellular carcinoma
proliferation. Oncotarget. 6:7866–7879. 2015.PubMed/NCBI
|
|
25
|
Das E and Bhattacharyya NP: MicroRNA-432
contributes to dopamine cocktail and retinoic acid induced
differentiation of human neuroblastoma cells by targeting NESTIN
and RCOR1 genes. FEBS Lett. 588:1706–1714. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chen L, Kong G, Zhang C, Dong H, Yang C,
Song G, Guo C, Wang L and Yu H: MicroRNA-432 functions as a tumor
suppressor gene through targeting E2F3 and AXL in lung
adenocarcinoma. Oncotarget. 7:20041–20053. 2016.PubMed/NCBI
|
|
27
|
Gollob JA, Wilhelm S, Carter C and Kelley
SL: Role of Raf kinase in cancer: Therapeutic potential of
targeting the Raf/MEK/ERK signal transduction pathway. Semin Oncol.
33:392–406. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Downward J: Targeting RAS signalling
pathways in cancer therapy. Nat Rev Cancer. 3:11–22. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Leicht DT, Balan V, Kaplun A, Singh-Gupta
V, Kaplun L, Dobson M and Tzivion G: Raf kinases: Function,
regulation and role in human cancer. Biochim Biophys Acta.
1773:1196–1212. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kersemaekers AM, Fleuren GJ, Kenter GG,
Van den Broek LJ, Uljee SM, Hermans J and Van de Vijver MJ:
Oncogene alterations in carcinomas of the uterine cervix:
Overexpression of the epidermal growth factor receptor is
associated with poor prognosis. Clin Cancer Res. 5:577–586.
1999.PubMed/NCBI
|
|
31
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao
L, Zhang Y, Wu YM, Dhanasekaran SM, Engelke CG, Cao X, et al: The
landscape of circular RNA in cancer. Cell. 176:869–881.e13. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Shang X, Li G, Liu H, Li T, Liu J, Zhao Q
and Wang C: Comprehensive circular RNA profiling reveals that
hsa_circ_0005075, a new circular RNA biomarker, is involved in
hepatocellular crcinoma development. Medicine (Baltimore).
95:e38112016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hsiao KY, Lin YC, Gupta SK, Chang N, Yen
L, Sun HS and Tsai SJ: Noncoding effects of circular RNA CCDC66
promote colon cancer growth and metastasis. Cancer Res.
77:2339–2350. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wang PL, Liu B, Xia Y, Pan CF, Ma T and
Chen YJ: Long non-coding RNA-low expression in tumor inhibits the
invasion and metastasis of esophageal squamous cell carcinoma by
regulating p53 expression. Mol Med Rep. 13:3074–3082. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Lasda E and Parker R: Circular RNAs:
Diversity of form and function. RNA. 20:1829–1842. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gullick WJ, Marsden JJ, Whittle N, Ward B,
Bobrow L and Waterfield MD: Expression of epidermal growth factor
receptors on human cervical, ovarian, and vulval carcinomas. Cancer
Res. 46:285–292. 1986.PubMed/NCBI
|
|
38
|
Biesterfeld S, Schuh S, Muys L, Rath W,
Mittermayer C and Schroder W: Absence of epidermal growth factor
receptor expression in squamous cell carcinoma of the uterine
cervix is an indicator of limited tumor disease. Oncol Rep.
6:205–209. 1999.PubMed/NCBI
|
|
39
|
Yoshioka K: Scaffold proteins in mammalian
MAP kinase cascades. J Biochem. 135:657–661. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Kohno M and Pouyssegur J: Targeting the
ERK signaling pathway in cancer therapy. Ann Med. 38:200–211. 2006.
View Article : Google Scholar : PubMed/NCBI
|