Introduction
Immunoglobulin A nephropathy (IgAN) is an immune
complex glomerulonephritis characterized by IgA deposition in the
mesangial area, abnormal proliferation of mesangial cells and
excessive expansion of the mesangial matrix (1,2). It is
one of the commonest causes of end-stage kidney failure and there
is at present no specific treatment targeting IgAN (3). Thus, research has focused on exploring
the pathogenesis of IgAN and developing effective treatment against
it. As IgA deposition in the mesangial region is a key step in the
development of IgAN (4) and the
etiology and pathogenesis of this disease are unclear, the
pathogenesis of IgAN can be discussed from the perspective of IgA
deposition in the mesangial region.
Dachshund family transcription factor 1 (DACH1) is a
key component of the retinal determination gene network family that
has been shown to be closely associated with organogenesis and
tumorigenesis (5,6). DACH1 participates in cell
differentiation and proliferation in renal development (7) and decreased expression of DACH1 is
associated with the progression and severity of glomerulopathy
(8). DACH1 disorder has been
reported in a variety of human malignancies such as breast cancer
(9), ovarian cancer (10), renal carcinoma (11) and gastric cancer (12). It acts as a cell-fate determination
factor that regulates cell growth and development (13,14),
but the mechanism and function of DACH1 dysregulation in IgAN has
yet to be explored.
Abnormal cell proliferation is a sign of cancer
transformation (15) and abnormal
expression of genes in cancer cells can be directly involved in the
regulation of cell growth and cell cycle progression (16,17).
Malfunctions in the cell cycle usually result in uncontrolled cell
growth characteristics, allowing cancer cells to proliferate
excessively and eventually leading to the tumorigenesis (18). DACH1 inhibits cell proliferation and
enhance cell cycle arrest. For example, Chen et al (19) demonstrated that DACH1 participates
in p53-mediated p21 induction and cell cycle arrest in
non-small-cell lung cancer. Wu et al (10) revealed that low expression of DACH1
in breast cancer cells promotes tumor growth by downregulating
Nanog and Sox2. Kalousova et al (20) confirmed that DACH1 can bind to the
promoter of the cell cycle inhibitor p27 to inhibit the
proliferation and cell cycle progression of insulin-producing
cells. The present study explored the role of DACH1 in IgAN in
terms of cell proliferation and cell cycle progression.
MicroRNAs (miRNAs) are small non-coding RNAs that
consist of ~22 nucleotides (21).
miRNAs can degrade target mRNAs or inhibit their translation by
binding to the 3′-untranslated region (3′UTR) of the target mRNAs
to negatively regulate gene expression (22). Studies have shown that miRNAs are
involved in the regulation of cell proliferation, differentiation,
cell cycle, apoptosis and inflammation (22–25).
The present study evaluated the expression and function of DACH1 in
human mesangial cells (HMCs) cultured with polymeric IgA (pIgA)
that was isolated and purified from the serum of patients with IgAN
or healthy individuals. Bioinformatics analysis was used to screen
for candidate miRNAs associated with DACH1 expression. Among the
predicted miRNAs, miRNA (miR)-140-3p was selected and its
expression and effect on cell proliferation and cell cycle
progression in IgAN were investigated. It was found that miR-140-3p
directly suppressed the expression of DACH1 and promoted the
proliferation and cell cycle progression of HMCs in IgAN. The
findings of the present study may provide a theoretical basis for
the study of the mechanism of IgAN.
Materials and methods
Human IgAN samples
The serum of 30 patients with IgAN (men=23; women=7;
age, 20–30 years n=19 and >30 years n=11) and 30 normal
individuals was collected. Inclusion criteria included cases of lgA
nephropathy and exclusion criteria included other nephropathies.
The recruitment of patients was conducted at Shanghai Tenth
People's Hospital between January 2020 and April 2020. The serum
collection was approved by the patients and all provided written
informed consent. All blood samples (each sample ~5 ml) were
anonymized. The present study was approved by the clinical research
ethics committee of Shanghai Tenth People's Hospital (approval no.
SHSY-IEC-4.1/20-117-01).
Extraction of polymeric IgA
(pIgA)
The pIgA was isolated and purified from the serum of
patients with IgAN or healthy individuals using a jacalin-agarose
column as previously described (26,27).
The extracted pIgA was stored at −80°C until use. The final
concentration of pIgA was 0.5 mg/ml.
Cell lines and culture
Human mesangial cells (HMCs) and 293T (cat. no.
KCB200744YJ) cells were purchased from the Conservation Genetics
CAS Kunming Cell Bank. HMCs were cultured in RPMI-1640 medium
(Invitrogen; Thermo Fisher Scientific, Inc.) containing 10% fetal
calf serum (Gibco; Thermo Fisher Scientific, Inc.) and 80 U/ml
penicillin and 0.1 mg/ml streptomycin (Beyotime Institute of
Biotechnology). HMCs and 293T were cultured at 37°C in an
atmosphere of 5% CO2.
Plasmids
Lentiviral vector plasmid pCDH-CMV-MCS-puro (pCDH)
and recombinant plasmid pCDH-CMV-DACH1-puro (pDACH1) were purchased
from GenScript for the transfection and production of lentiviruses.
The pGL3-Control plasmid and pGL3-DACH1 3′UTR luciferase reporter
plasmid were purchased from GenScript for use in detection of
luciferase activity following transfection.
Lentivirus transduction and
production
Lentivirus packaging vector psPAX2 (5 µg) and
envelope vector pMD2.G (10 µg) were co-transfected with lentivirus
plasmid pCDH (5 µg) or pDACH1 (5 µg) into 293T cells using
Lipofectamine 2000 (50 µl) (Thermo Fisher Scientific, Inc.) to
produce the corresponding lentivirus solution. The culture
supernatant was collected 48–72 h after transfection. The
supernatant contained the corresponding virus and was used for
subsequent cell infection experiments.
Reverse transcription-quantitative
(RT-q) PCR
Total RNA was extracted from 5×106 cells
using TRIzol® reagent kit according to the
manufacturer's protocol (Thermo Fisher Scientific, Inc.) and
reverse transcribed into single-stranded cDNA according to the
instructions of the Reverse Transcription kit (Vazyme Biotech Co.,
Ltd.). The expression of mRNA was detected by RT-qPCR using the
Rotor-Gene Q detection system (Qiagen GmbH) and the reaction volume
was 20 µl, with the following cycling conditions: 95°C for 5 min,
then 45 cycles of 95°C for 10 sec and 60°C for 30 sec. The
expression of miRNA or mRNA was calculated using the
2−ΔΔCq method (28). The
mRNA expression level was normalized to that of GAPDH and the
expression level of miRNA was normalized to U6 RNA. The primer
sequences used were: IL-6 forward, 5′-AACTCCTTCTCCACAAGCGCCTT-3′
and reverse, 5′-GTCAATTCGTTCTGAAGAGGTG-3′; IL-8 forward,
5′-GGTCTCACCTCCCAACTGC-3′ and reverse,
5′-TCAGCTCGAACACTTTGAATAT-3′; IL-13 forward,
5′-TGAGGAGCTGGTCAACATCA-3′ and reverse, 5′-CCACCTCGATTTTGGTGTCT-3′;
chemokine cc-motif ligand 1 (CXCL1) forward,
5′-CTCCTGCGAGTGGCACTGCTGCTC-3′ and reverse,
5′-GAGGCAAGCTTTCCGCCCATTCTT-3′; GAPDH forward,
5′-GAGTCAACGGATTTGGTCGT-3′ and reverse, 5′-GATCTCGCTCCTGGAAGATG-3′;
and U6 forward, 5′-CTTCGGCAGCACATATAC-3′ and reverse,
5′-TTCACGAATTTGCGTGTCAT-3′. The experiment was repeated three
times.
Western blot analysis
The total protein of each group was harvested and
collected using RIPA lysis buffer (Thermo Fisher Scientific, Inc.).
Protein concentration was measured with a BCA Protein Assay kit
(Beyotime Institute of Biotechnology). Equal amounts of protein (30
µg/lane) were separated by 10% sodium dodecyl
sulfate-polyacrylamide gel electrophoresis and transferred to
polyvinylidene fluoride membranes (EMD Millipore). The membrane was
blocked by incubation with 5% non-fat milk for 1 h at 24°C and then
with primary antibodies overnight at 4°C. The primary antibodies
used were: Anti-DACH1 (cat. no. ab176718; Abcam; 1:1,000),
anti-Cyclin D1 (cat. no. 55506; Cell Signaling Technology, Inc.;
1:1,000), anti-Cyclin A (cat. no. 91500; Cell Signaling Technology,
Inc.; 1:1,000) and anti-P21 (cat. no. 2947; Cell Signaling
Technology, Inc.; 1:1,000), anti-P53 (cat. no. 2527; Cell Signaling
Technology, Inc.; 1:1,000). After washing three times with Tris
buffered saline-Tween (TBS-T; 50 mM Tris, 150 mM NaCl, 0.05%
Tween-20) the membranes were incubated with goat anti-rabbit (cat.
no. ab6721; Abcam; 1:5,000) or anti-mouse (cat. no. ab205719;
Abcam; 1:5,000) IgG/horseradish peroxidase secondary antibody at
room temperature for 1 h. Protein expression was detected using a
chemiluminescence detection system (Tanon 4600SF; Tanon Science and
Technology Co., Ltd.).
Cell proliferation assay
HMCs of each group were seeded into 96 well plates
at 1×104 cells/well one day prior to cell viability
measurement at day 1, with additional measurements taken on days 3
and 5. Cell proliferation assay was determined using the CCK-8 cell
count kit (cat. no. E606335, Sangon Biotech Co., Ltd.). According
to the manufacturer's instructions, CCK-8 was mixed with DMEM at
1:10 v/v, then added to 96 well plates and incubated at 37°C for 2
h. The optical density (OD) values were measured at a wavelength of
562 nm using a multimode reader (BioTek Instruments, Inc.)
Flow cytometry
Cell cycle distribution was detected by
Annexin/propidium iodide (PI) single staining (cat. no. P4170,
Sigma-Aldrich; Merck KGaA). HMCs of each group were simultaneously
inoculated into 6-well plates at 2×105 cells/well. After
72 h, the cells were collected and detached with 0.25% trypsin
solution. The concentration of cells was adjusted to
1×106 cells/ml, with 1 ml of cells then centrifuged at
500 × g for 10 min at 4°C. The supernatant was discarded and the
cells were collected. The cell suspension (1 ml) was added to 2 ml
of PBS. This was regarded as the template for further use. The
cells were centrifuged (500 × g, 10 min, 4°C) again and the
supernatant removed. Next, the cells were reacted with precooled
70% (v/v) ethanol solution at 4°C overnight. The fixed cells were
rinsed twice with PBS and then 100 µl of the cell suspension
containing at least 1×106 cells/ml was stained with 1 ml
of 50 mg/l PI dye solution containing RNAase (20 µg/ml) for 30 min
at 25°C in darkness and then filtered. The cell cycle was detected
by red fluorescence at an excitation wavelength of 488 nm and
recorded using a flow cytometer (BD AccuriC6; BD Accuri C6
Software; BD Biosciences).
Enzyme-linked immunosorbent assay
(ELISA)
HMCs of each group were simultaneously seeded into
6-well plates at 2×105 cells/well. After 72 h, the
levels of several inflammatory cytokines including CXCL1 (cat. no.
AD10945Hu), IL-6 (cat. no. AD11099Hu), IL-8 (cat. no. AD11098Hu)
and IL-13 (cat. no. AD11110Hu) in HMC supernatants were determined
using ELISA kits (Andy Gene Biotechnology Co., Ltd.) according to
the manufacturer's instructions.
Luciferase reporter assay
miR-140-3p mimic, miR-140-3p mutant, other miRNAs
and negative control was co-transfected into 293T cells together
with DACH1 3′UTR luciferase reporter plasmid (GenScript). After 28
h of transfection, the cells were harvested using Dual-Lumi™ II
Luciferase Reporter Assay kit (Beyotime Institute of Biotechnology)
according to the manufacturer's instructions and the luciferase
activity was detected using a multimode reader (BioTek Instruments,
Inc.). The Firefly luciferase activity was normalized to the
Renilla luciferase activity.
Target prediction
miRTar (miRTar, developed by Dr. Hsien-Da Huang,
http://mirtar.mbc.nctu.edu.tw/human/)
was used to predict binding sequences of miR-140-3p in the 3′UTR of
DACH1.
miRNA mimics, mutation and
inhibition
Synthetic miR-140-3p mimic, mutation, inhibitor,
other miRNAs and negative control were obtained from GenScript. The
miR-140-3p mutant sequence was 5′-UACCACAGGGUAGAACCACGG-3′, the
miR-140-3p mutant sequence was 5′-AUGCACAGGGUAGAACCACGG-3′ and the
negative control sequence was 5′-UUCUCCGAACGUGUCACGUTT-3′. RT-qPCR
results confirmed that miR-140-3p mimic, mutation and inhibitor
were successfully transfected into HMCs or 293 (Fig. S1).
Statistical analysis
Quantitative data are presented as the mean ±
standard deviation. The Tukey multiple comparisons test was used
for statistical analysis, a one-way ANOVA was performed before
Tukey's test. P>0.05 was considered no statistical significance
(ns), P<0.05 was considered to indicate a statistically
significant difference. All graphs were generated using GraphPad
Prism 8.0 (GraphPad Software, Inc.). The experiment was repeated
three times.
Results
Exogenous DACH1 suppresses the
proliferation and altered the cell cycle of HMCs
A previous study has shown that DACH1 is expressed
at low levels in IgAN and is associated with the progression and
severity of glomerulopathy (8).
However, its role in the development of human IgAN remains to be
elucidated. The endogenous expression of DACH1 in HMCs that were
cultured with pIgA-IgAN or pIgA-control was first detected. As
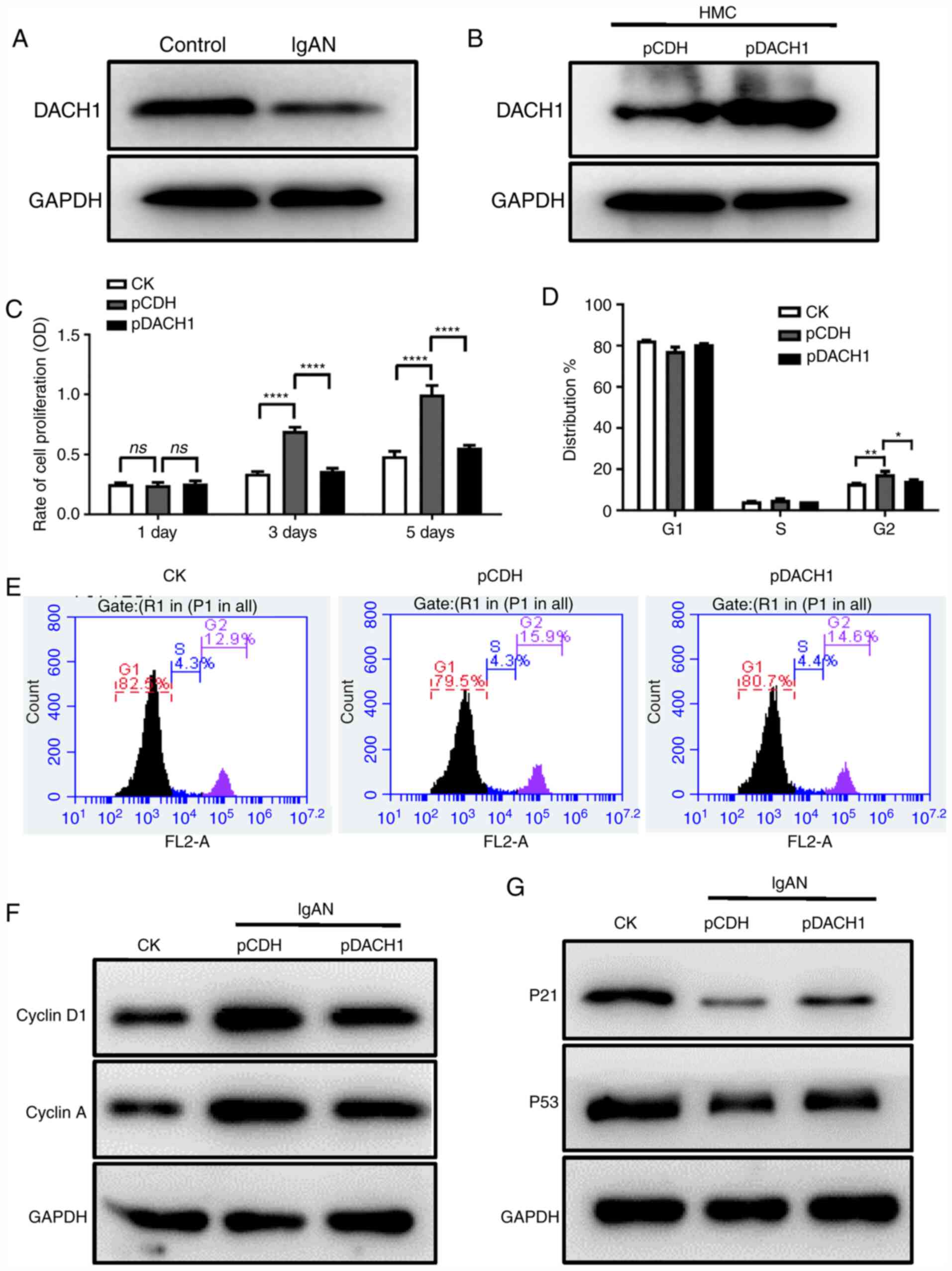
shown in Figs. 1A and S2A, the endogenous expression of DACH1
was significantly lower in HMCs cultured with pIgA-IgAN compared
with cells cultured with pIgA-control. To investigate the role of
DACH1 in IgAN, HMCs were transfected with lentiviral DACH1 vectors.
DACH1-transfected HMCs overexpressed DACH1 at the mRNA (Fig. S2B) and protein levels (Fig. 1B). The effect of DACH1 on cell
proliferation and cell cycle progression was then examined. The
cell proliferation assay demonstrated that DACH1 overexpression
reversed the ability of pIgA-IgAN to promote HMC proliferation
(Fig. 1C). The cell cycle assay
demonstrated that DACH1 enhanced pIgA-IgAN-induced G2 phase arrest
in HMCs (Fig. 1D and E). The
expression of proteins closely associated with the cell cycle were
detected by western blot. As shown in Fig. 1F and G, the expression levels of
Cyclin D1 and Cyclin A were upregulated in the presence of
pIgA-IgAN but were downregulated by DACH1 and the expression levels
of p21 and p53 were downregulated in the presence of pIgA-IgAN but
were upregulated by DACH1.
Exogenous DACH1 regulates inflammatory
response in HMCs
To further investigate the function of DACH1 in
IgAN, the release of inflammatory cytokines from HMCs that were
stably transfected with DACH1 was measured. The expression of
inflammatory factors (IL-6, IL-8, IL-13 and CXCL1) in the
supernatant of DACH1-transfected HMCs was detected by ELISA. As
shown in Fig. 2A, the
concentrations of IL-6, IL-8, IL-13 and CXCL1 in HMCs cultured with
pIgA-IgAN were higher compared with the control group, whereas
DACH1 reversed this phenomenon by reducing the pIgA-IgAN-induced
expression of IL-6, IL-8, IL-13 and CXCL1. In addition, the mRNA
expression levels of IL-6, IL-8, IL-13 and CXCL1 in HMCs incubated
with 0.5 mg/ml pIgA-IgAN were higher compared with the control
group. Similarly, DACH1 downregulated the expression of IL-6, IL-8,
IL-13 and CXCL1 (Fig. 2B).
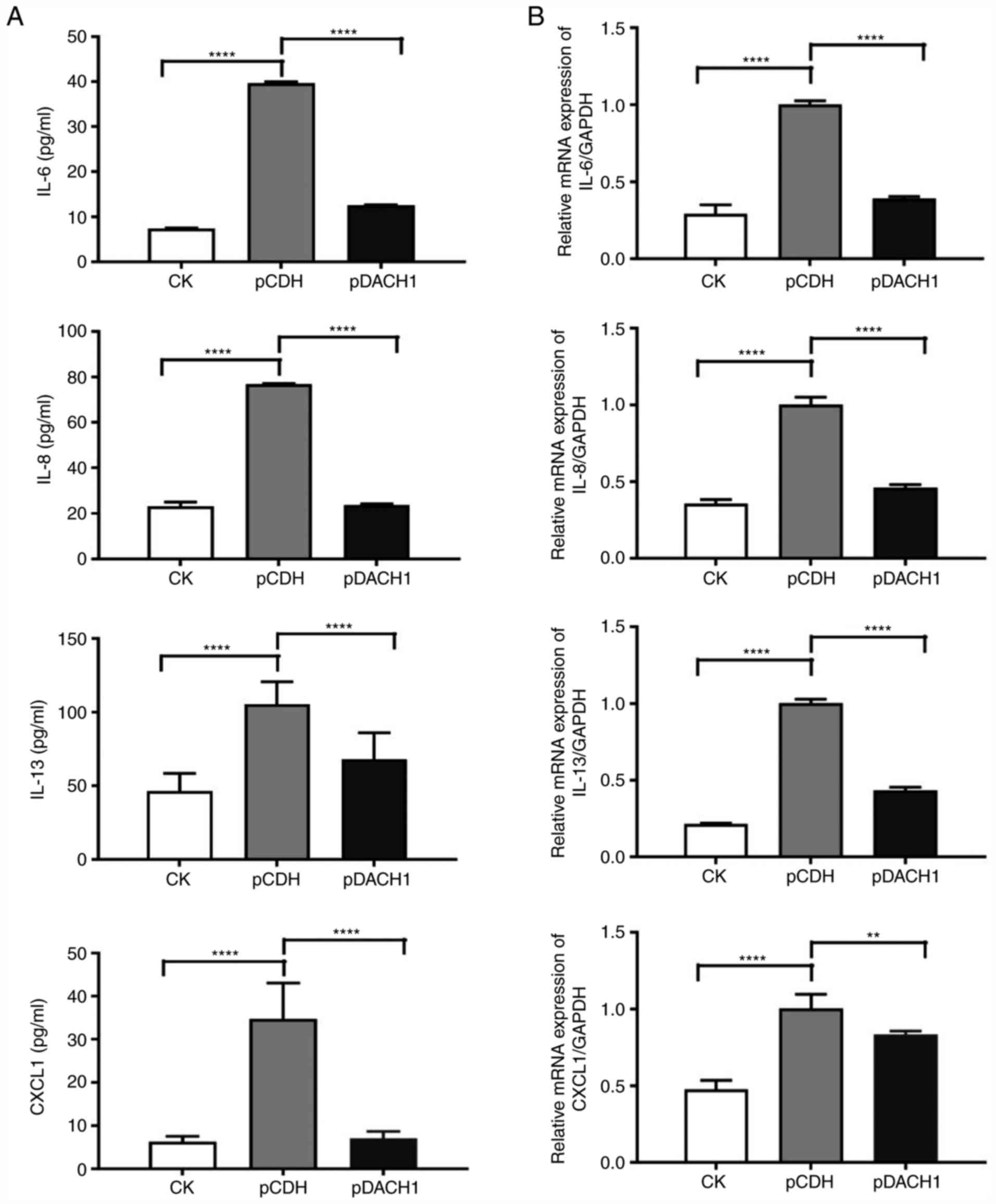 | Figure 2.Overexpression of DACH1 decreases the
release of inflammatory cytokines. (A) Secretion of IL-6, IL-8,
IL-13 and CXCL1 (****P<0.0001) in the supernatant of HMCs
transduced with the DACH1 or control (pCDH) when pIgA-IgAN was
added or not. (B) Reverse transcription-quantitative PCR analyses
of the mRNA levels of IL-6, IL-8, IL-13 and CXCL1 (**P<0.01 and
****P<0.0001) in HMCs transduced with the DACH1 or control
(pCDH) when pIgA-IgAN was added or not. Data are expressed as mean
± standard deviation. pCDH, pCDH-CMV-MCS-puro; ns, not significant;
pDACH1, pCDH-CMV-DACH1-puro; pIgA, polymeric IgA; IgAN,
immunoglobulin A nephropathy; HMCs, human mesangial cells; DACH1,
dachshund family transcription factor 1. |
Cellular miR-140-3p directly targets
DACH1 3′UTR
To assess the mechanism mediating DACH1-induced
inhibition of cell proliferation and inflammatory response, the
present study screened for miRNAs that may interact with DACH1
using bioinformatics software. A total of 10 miRNAs were selected,
synthesized and transfected into HMCs. The mRNA and protein
expression of DACH1 were detected by RT-qPCR and western blot,
respectively. As shown in Figs. 3A
and S2C, when miR-140-3p was
transfected into HMCs, the mRNA and protein expression of DACH1 was
decreased. To further verify whether DACH1 is regulated by
miR-140-3p, the luciferase reporter plasmids containing the
full-length sequence of DACH1 3′UTR region with 10 miRNAs we
co-transfected into 293 cells. In the luciferase reporter assay,
miR-140-3p reduced the luciferase activity of the 3′UTR of DACH1
(Fig. 3B). Subsequently, miR-140-3p
mimics were transfected into HMCs and the expression of DACH1
detected by RT-qPCR and western blot. As shown in Fig. 3C, the expression of DACH1 was
decreased by increasing the expression of miR-140-3p in a
dose-dependent manner. miR-140-3p mimics and mutants were then
transfected into 293 cells to examine the expression of DACH1. As
shown in Fig. 3E-G, miR-140-3p
mimics inhibited the expression of DACH1 and the luciferase
activity of the 3′UTR of DACH1, while the miR-140-3p mutant had no
effect. These data suggested that DACH1 is a direct target of, and
is regulated by, miR-140-3p.
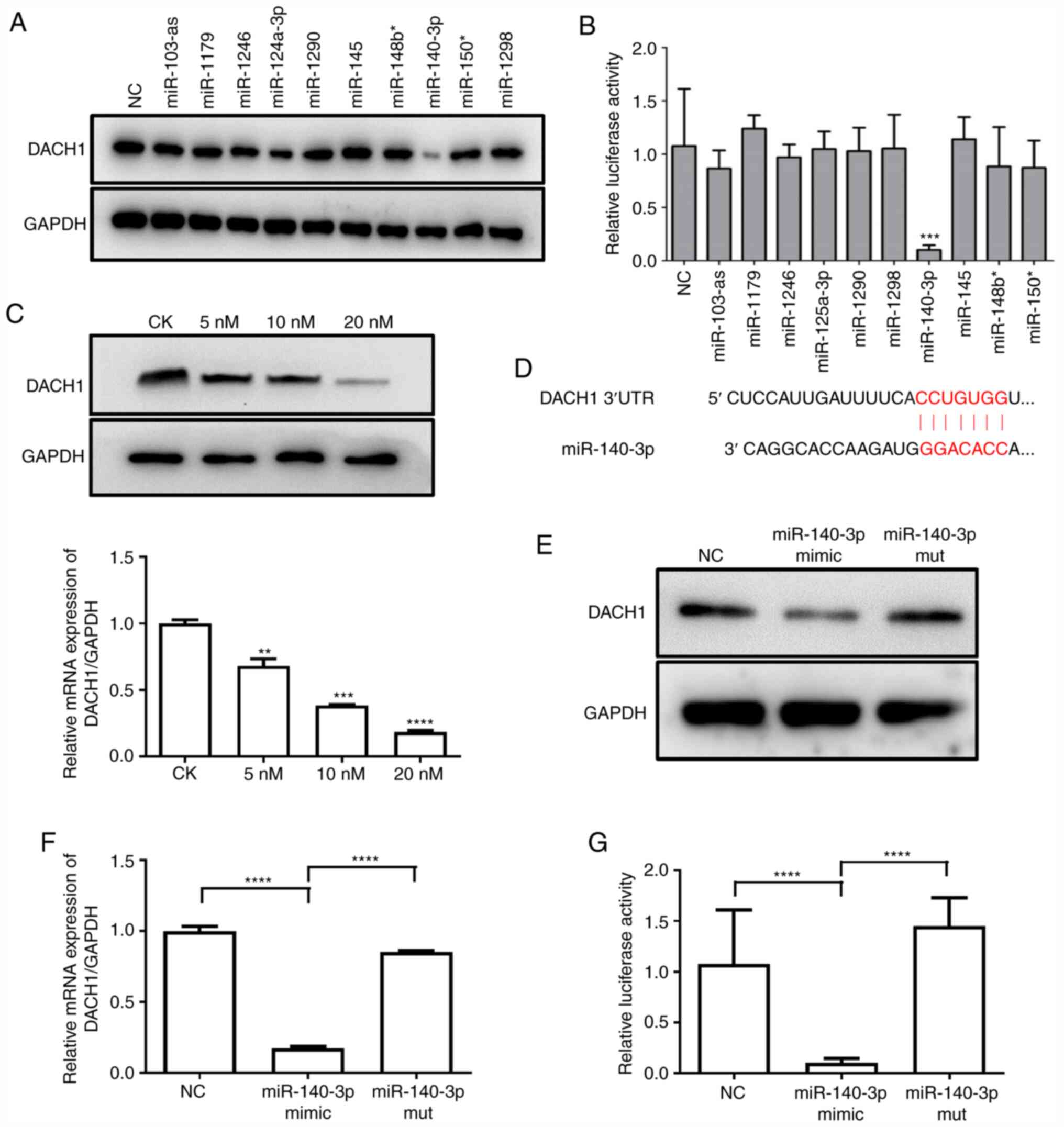 | Figure 3.miR-140-3p directly targets DACH1.
(A) The effect of miRNAs on endogenous DACH1 expression. HMCs were
transfected with 10 miRNA mimics (20 nM) and NC, western blotting
was performed at 48 h after transfection. (B) Inhibition of DACH1
3′UTR reporter activity by miRNAs. Luciferase activity was detected
in 293T cells co-transfected with 10 miRNA mimics (20 nM) or NC
together with the pGL3-DACH1 3′UTR luciferase reporter
(***P<0.001 NC vs. miR-140-3p). (C) miR-140-3p inhibited
expression of DACH1 in a dose-dependent manner. HMCs transfected
with an increasing amount (5–20 nM) of miR-140-3p mimic or NC for
48 h. The transfected cells were collected for examined by western
blotting and RT-qPCR (**P<0.01 NC vs. 5 nM miR-140-3p,
***P<0.001 NC vs. 10 nM miR-140-3p, ****P<0.0001 NC vs. 20 nM
miR-140-3p). (D) Predicted binding sequences of miR-140-3p in the
3′UTR of DACH1 (miRTar, http://mirtar.mbc.nctu.edu.tw/human/). (E and F)
Effect of mutation of miR-140-3p on endogenous DACH1 expression.
miR-140-3p mimics and mutant were transfected into HMCs for 48 h,
while the expression of DACH1 was detected by western blotting and
RT-qPCR (****P<0.0001). (G) miR-140-3p mimics/mutant or NC were
co-transfected with pGL3-DACH1 3′UTR luciferase reporter into 293T
cells for 48 h and the luciferase activity detected
(****P<0.0001). Data are expressed as mean ± standard deviation.
CK, control check; miR, microRNA; DACH1, dachshund family
transcription factor 1; HMCs, human mesangial cells; NC, negative
control; 3′UTR, 3′-untranslated region; RT-qPCR, reverse
transcription-quantitative PCR; mut, mutant. |
miR-140-3p inhibition reverses the
effect of pIgA-IgAN induced cell proliferation and cell cycle
changes
Next, cell proliferation and cell cycle assays were
performed to determine the functional role of miR-140-3p in IgAN.
First, the expression of miR-140-3p was detected in HMCs that were
cultured with pIgA-IgAN or pIgA-control. As shown in Fig. 4A, the expression of miR-140-3p was
significantly higher in HMCs cultured with pIgA-IgAN compared with
cells cultured with pIgA-control. Cell proliferation and cell cycle
assays were performed in HMCs transfected with miR-140-3p
inhibitors or negative controls in the presence of pIgA-IgAN. The
cell cycle assay demonstrated that miR-140-3p enhanced
pIgA-IgAN-induced G2 phase arrest and inhibited
G1 phase activation in HMCs (Fig. 4B and C). The cell proliferation
assay revealed that inhibition of miR-140-3p reversed the ability
of pIgA-IgAN to promote HMC proliferation (Fig. 4D). Upon miR-140-3p inhibition,
cyclin D1 and cyclin A were downregulated whereas p21 and p53 were
upregulated (Fig. 4E).
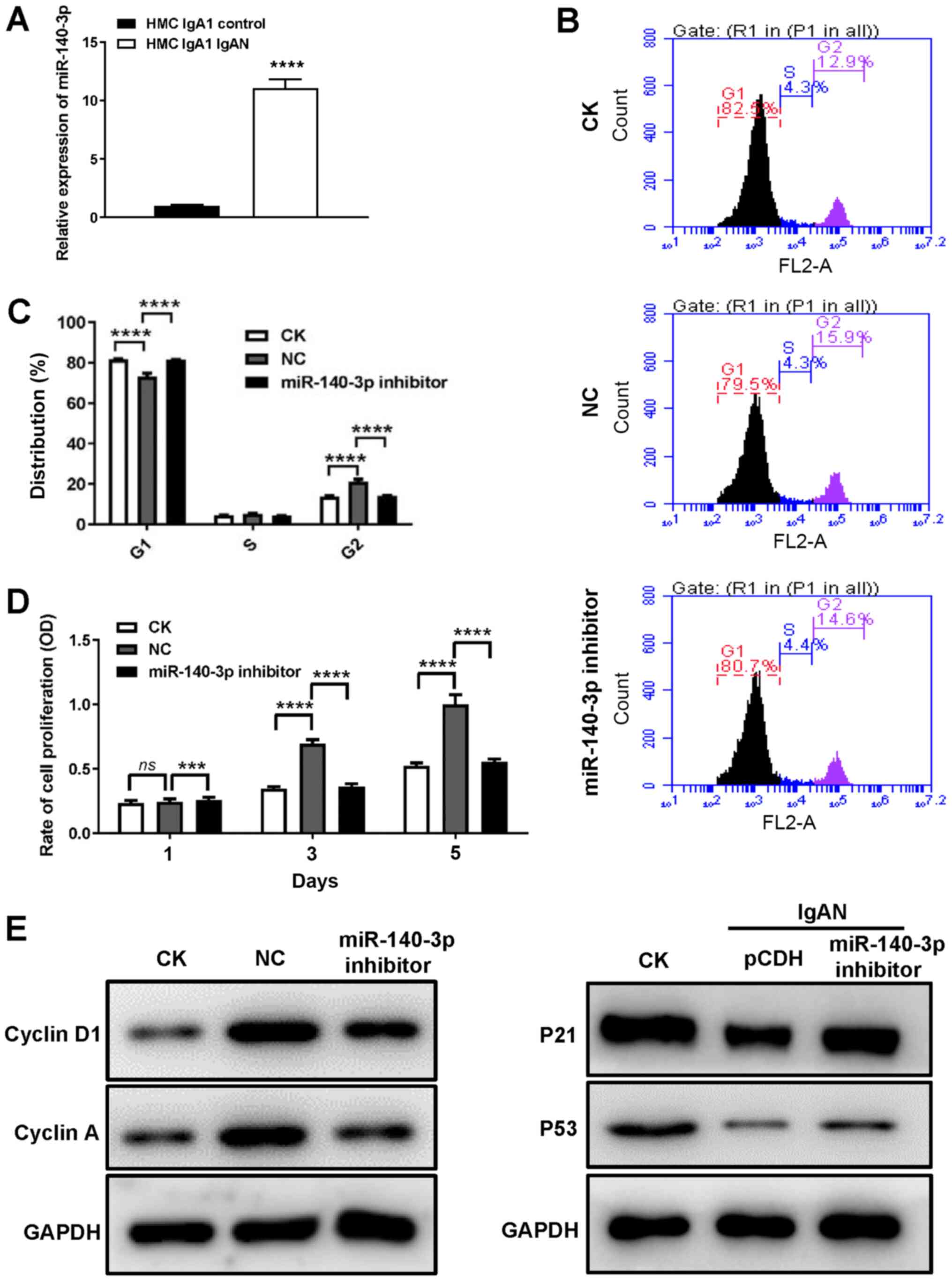 | Figure 4.DACH1 decrease IgAN cell
proliferation and cell cycle progression by regulating miR-140-3p
expression. (A) Expression of miR-140-3p in HMCs incubated with
pIgA or control was detected by reverse transcription-quantitative
PCR (****P<0.0001 control vs. IgAN). Representative (B) flow
cytometric graphs and (C) statistical analysis showing the
distribution of cell cycle in HMCs transfected with miR-140-3p
inhibitor or NC. (D) HMCs transfected with miR-140-3p inhibitor or
NC were seeded and examined for cell proliferation with CCK-8 test
on days 1, 3 and 5. (E) Protein expression of Cyclin D1, Cyclin A,
p21 and p53 in HMCs transfected with miR-140-3p inhibitor or NC was
analyzed by western blotting. Data are expressed as mean ± standard
deviation. ***P<0.001 and ****P<0.0001 by multiple
comparisons test. CK, control check; DACH1, dachshund family
transcription factor 1; IgAN, immunoglobulin A nephropathy; miR,
microRNA; HMCs, human mesangial cells; NC, negative control; ns,
not significant. |
miR-140-3p also regulates inflammatory
response in HMCs
To further evaluate the function of miR-140-3p in
IgAN, the release of inflammatory cytokines from HMCs transfected
with miR-140-3p inhibitors was detected. The supernatant was
collected after transfection and the expression of IL-6, IL-8,
IL-13 and CXCL1 was detected by ELISA. As shown in Fig. 5A and B, miR-140-3p inhibition
reversed the pIgA-IgAN-induced expression of IL-6, IL-8, IL-13 and
CXCL1 in HMCs at the supernatant and at the mRNA level.
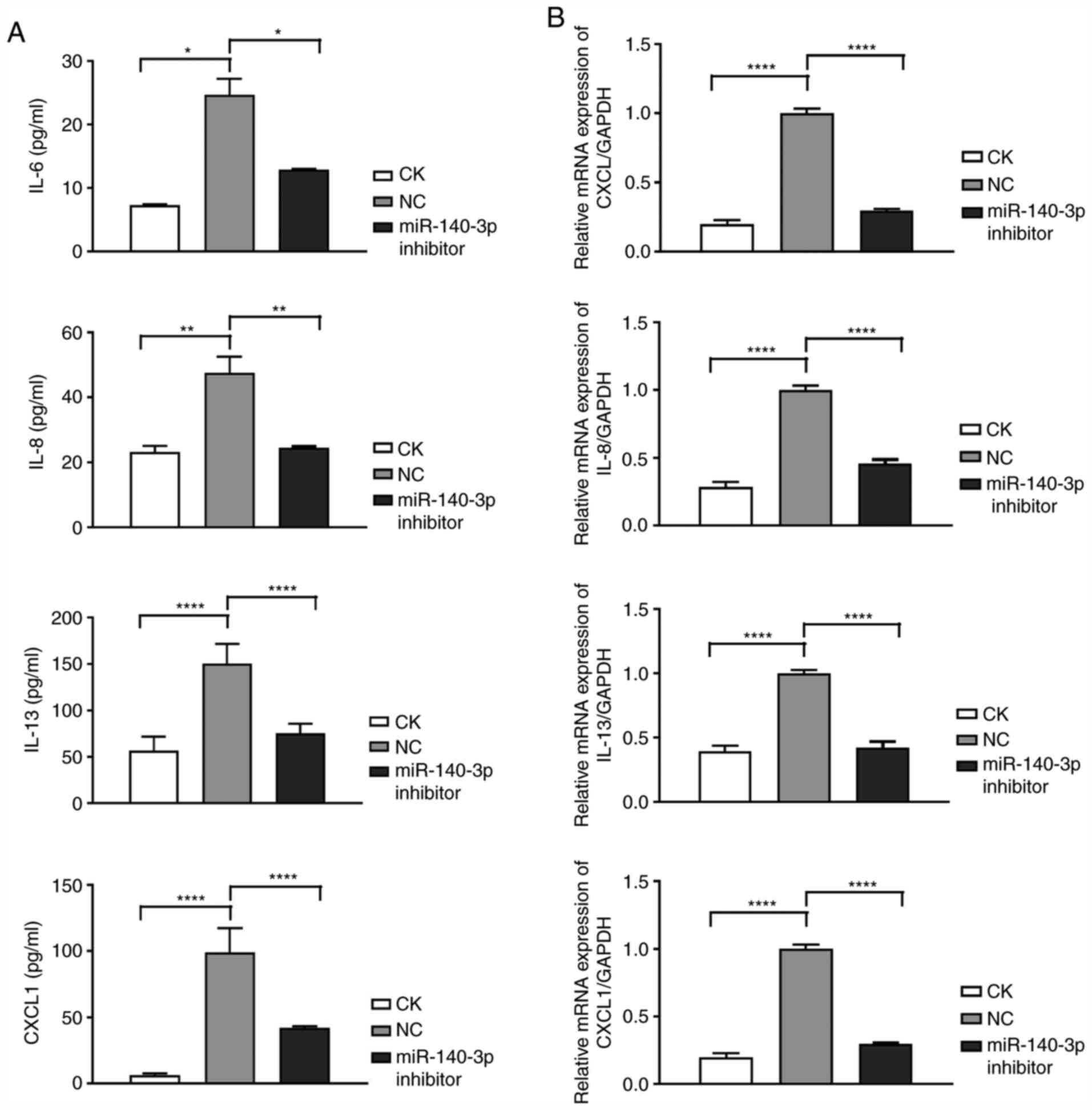 | Figure 5.DACH1 decreases the release of
inflammatory cytokines by positively regulating miRNA-140-3p
expression. (A) Secretion of IL-6, IL-8, IL-13 and CXCL1 in the
supernatant of HMCs transfected with the miR-140-3p inhibitor or NC
when pIgA-IgAN was added or not. (B) Reverse
transcription-quantitative PCR analyses of the mRNA levels of IL-6,
IL-8, IL-13 and CXCL1 in HMCs transfected with the miR-140-3p
inhibitor or NC when pIgA-IgAN was added or not. Data are expressed
as mean ± standard deviation. *P<0.05, **P<0.01 and
****P<0.0001. CK, control check; DACH1, dachshund family
transcription factor 1; miR, microRNA; NC, negative control; HMCs,
human mesangial cells; pIgA, polymeric IgA. |
Discussion
IgAN is one of the common type of glomerulonephritis
worldwide (29). According to
statistics, ~40% of patients with IgAN progress to end-stage
nephropathy within 20 years of disease onset, severely affecting
the physical and mental health of patients (30). A typical feature of IgAN is IgA
immune complex-mediated mesangial cell proliferation and the
detection of IgA deposition in mesangial cells by renal biopsy and
immunohistochemistry is a method of IgAN diagnosis (31,32).
The pathogenesis of IgAN is unclear and it is important to explore
the mechanism of IgAN.
DACH1 has been shown to act as a tumor suppressor
gene in various malignancies (9–12) and
participates in tumor cell proliferation and metabolism by
regulating cell cycle-related proteins (20). Liu et al (8) suggested that DACH1 expression is
decreased in IgAN and that DACH1 is involved in disease progression
and severity by regulating cell cycle-related proteins. The present
study hypothesized that DACH1 may be involved in regulating the
cell cycle and proliferation of mesangial cells. In the present
study, pIgA was isolated and purified from 30 patients with IgAN
and 30 healthy individuals. HMCs were incubated with pIgA to
simulate the cell model of IgA deposition in mesangial cells in
IgAN (27,33). It was found that DACH1 was
downregulated in HMCs incubated with pIgA isolated from patients
with IgAN compared with that in cells cultured with pIgA isolated
from healthy individuals. In addition, exogenous DACH1 inhibited
the proliferation and enhanced cell cycle arrest in HMCs induced by
pIgA from patients with IgAN.
Each phase of the cell cycle is controlled by
specific regulatory proteins. Cyclin D1 mainly promotes
G1-S phase transition and regulates the G0
phase reentry of stationary cells into the cell cycle in
G1 (34,35). Cyclin A is essential for DNA
synthesis and its expression peaks at the late G1 phase
and S phase (36,37). p53 activation initiates cell cycle
arrest at the G1/S phase (38) and p21 is involved in the regulation
of G2/M transition (39). In the present study, DACH1
overexpression decreased the expression of cyclin D1 and cyclin A
in HMCs that were upregulated by pIgA from patients with IgAN.
According to previous study, abnormal miRNA
expression is present in most patients with IgAN and is closely
associated with the extent of the disease (40). miRNAs participate in almost every
process involved in cancer occurrence, development and progression
(22,41–43).
The present study confirmed that miR-140-3p expression was
upregulated in HMCs treated with pIgA-IgAN and that the
upregulation of miR-140-3p meditated the expression of DACH1. The
present study also demonstrated that miR-140-3p regulated cell
growth, cell cycle progression and the release of inflammatory
cytokines in HMCs. Although the findings of the present study
indicated that DACH1 was probably involved in the regulation of
IgAN and was regulated by miR-140-3p, a variety of other factors
are involved in regulating the occurrence and development of IgAN
and further studies on the pathogenesis of IgAN are warranted.
In summary, the present study demonstrated that
DACH1 was downregulated in HMCs in the presence of pIgA-IgAN.
Moreover, DACH1 regulated cell proliferation, cell cycle
progression and the release of inflammatory cytokines in HMCs and
these phenomena may be targeted by miR-140-3p. This novel pathway
is therefore expected to provide a weak theoretical basis for IgAN
research.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors' contributions
XZ and YL designed the study and performed the
experiments. XZ drafted the manuscript. PG and CZ were major
contributors in the conception, design and reviewing of the
manuscript. Based on their contributions, XZ was listed as the
first author, while CZ was the author for correspondence. All
authors read and approved the final manuscript.
Ethics approval and consent to
participate
All patients provided written informed consent and
all serum samples were anonymized. The present study was approved
by the clinical research ethics committee of Shanghai Tenth
People's Hospital (approval no. SHSY-IEC-4.1/20-117-01).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
IgAN
|
immunoglobulin A nephropathy
|
|
DACH1
|
dachshund family transcription factor
1
|
|
HMCs
|
human mesangial cells
|
|
pIgA
|
polymeric IgA
|
|
3′UTR
|
3′-untranslated region
|
|
CXCL1
|
chemokine cc-motif ligand 1
|
|
NC
|
negative control.
|
References
|
1
|
Lai KN: Pathogenesis of IgA nephropathy.
Nat Rev Nephrol. 8:275–283. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
McGrogan A, Franssen CF and de Vries CS:
The incidence of primary glomerulonephritis worldwide: A systematic
review of the literature. Nephrol Dial Transplant. 26:414–430.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Mestecky J, Novak J, Moldoveanu Z and
Raska M: IgA nephropathy enigma. Clin Immunol. 172:72–77. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Boyd JK, Cheung CK, Molyneux K, Feehally J
and Barratt J: An update on the pathogenesis and treatment of IgA
nephropathy. Kidney Int. 81:833–843. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhu H, Wu K, Yan W, Hu L, Yuan J, Dong Y,
Li Y, Jing K, Yang Y and Guo M: Epigenetic silencing of DACH1
induces loss of transforming growth factor-β1 antiproliferative
response in human hepatocellular carcinoma. Hepatology.
58:2012–2022. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Popov VM, Wu K, Zhou J, Powell MJ, Mardon
G, Wang C and Pestell RG: The Dachshund gene in development and
hormone-responsive tumorigenesis. Trends Endocrinol Metab.
21:41–49. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ikeda K, Watanabe Y, Ohto H and Kawakami
K: Molecular interaction and synergistic activation of a promoter
by Six, Eya, and Dach proteins mediated through CREB binding
protein. Mol Cell Biol. 22:6759–6766. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liu QQ, Zhou YQ, Liu HQ, Qiu WH, Liu H, Hu
TY, Xu Q, Lv YM and Wu KM: Decreased DACH1 expression in
glomerulopathy is associated with disease progression and severity.
Oncotarget. 7:86547–86560. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sunde JS, Donninger H, Wu K, Johnson ME,
Pestell RG, Rose GS, Mok SC, Brady J, Bonome T and Birrer MJ:
Expression profiling identifies altered expression of genes that
contribute to the inhibition of transforming growth factor-beta
signaling in ovarian cancer. Cancer Res. 66:8404–8412. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wu K, Li A, Rao M, Liu M, Dailey V, Yang
Y, Di Vizio D, Wang C, Lisanti MP, Sauter G, et al: DACH1 is a cell
fate determination factor that inhibits cyclin D1 and breast tumor
growth. Mol Cell Biol. 26:7116–7129. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Dalgin GS, Drever M, Williams T, King T,
DeLisi C and Liou LS: Identification of novel epigenetic markers
for clear cell renal cell carcinoma. J Urol. 180:1126–1130. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yamada Y, Arao T, Gotoda T, Taniguchi H,
Oda I, Shirao K, Shimada Y, Hamaguchi T, Kato K, Hamano T, et al:
Identification of prognostic biomarkers in gastric cancer using
endoscopic biopsy samples. Cancer Sci. 99:2193–2199. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chen R, Amoui M, Zhang Z and Mardon G:
Dachshund and eyes absent proteins form a complex and function
synergistically to induce ectopic eye development in Drosophila.
Cell. 91:893–903. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Shen W and Mardon G: Ectopic eye
development in Drosophila induced by directed dachshund expression.
Development. 124:45–52. 1997.PubMed/NCBI
|
|
15
|
Fan GK, Imanaka M, Yang B and Takenaka H:
Characteristics of nasal inverted papilloma and its malignant
transformation: A study of cell proliferation and programmed cell
death. Am J Rhinol. 20:360–363. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Liu M, Zhang H, Li Y, Wang R, Li Y, Zhang
H, Ren D, Liu H, Kang C and Chen J: HOTAIR, a long noncoding RNA,
is a marker of abnormal cell cycle regulation in lung cancer.
Cancer Sci. 109:2717–2733. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang W, Dong M, Cui J, Xu F, Yan C, Ma C,
Yi L, Tang W, Dong J and Wei Y: NME4 may enhance nonsmall cell lung
cancer progression by overcoming cell cycle arrest and promoting
cellular proliferation. Mol Med Rep. 20:1629–1636. 2019.PubMed/NCBI
|
|
18
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Chen K, Wu K, Cai S, Zhang W, Zhou J, Wang
J, Ertel A, Li Z, Rui H, Quong A, et al: Dachshund binds p53 to
block the growth of lung adenocarcinoma cells. Cancer Res.
73:3262–3274. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kalousova A, Mavropoulos A, Adams BA,
Nekrep N, Li Z, Krauss S, Stainier DY and German MS: Dachshund
homologues play a conserved role in islet cell development. Dev
Biol. 348:143–152. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
He L and Hannon GJ: MicroRNAs: Small RNAs
with a big role in gene regulation. Nat Rev Genet. 5:522–531. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kloosterman WP and Plasterk RH: The
diverse functions of microRNAs in animal development and disease.
Dev Cell. 11:441–450. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Li K, Du Y, Jiang BL and He JF: Increased
microRNA-155 and decreased microRNA-146a may promote ocular
inflammation and proliferation in Graves' ophthalmopathy. Med Sci
Monit. 20:639–643. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang W, Zhao LJ, Tan YX, Ren H and Qi ZT:
miR-138 induces cell cycle arrest by targeting cyclin D3 in
hepatocellular carcinoma. Carcinogenesis. 33:1113–1120. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Tamouza H, Vende F, Tiwari M,
Arcos-Fajardo M, Vrtovsnik F, Benhamou M, Monteiro RC and Moura IC:
Transferrin receptor engagement by polymeric IgA1 induces receptor
expression and mesangial cell proliferation: Role in IgA
nephropathy. Contrib Nephrol. 157:144–147. 2007.PubMed/NCBI
|
|
27
|
Lai KN, To WY, Li PK and Leung JC:
Increased binding of polymeric lambda-IgA to cultured human
mesangial cells in IgA nephropathy. Kidney Int. 49:839–845. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Schena FP and Nistor I: Epidemiology of
IgA nephropathy: A global perspective. Semin Nephrol. 38:435–442.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Tian J, Wang Y, Liu X, Zhou X and Li R:
Rapamycin ameliorates IgA nephropathy via cell cycle-dependent
mechanisms. Exp Biol Med (Maywood). 240:936–945. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kurogi Y: Mesangial cell proliferation
inhibitors for the treatment of proliferative glomerular disease.
Med Res Rev. 23:15–31. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Rodrigues JC, Haas M and Reich HN: IgA
nephropathy. Clin J Am Soc Nephrol. 12:677–686. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Leung JC, Chan LY, Tang SC, Lam MF, Chow
CW, Lim AI and Lai KN: Oxidative damages in tubular epithelial
cells in IgA nephropathy: Role of crosstalk between angiotensin II
and aldosterone. J Transl Med. 9:1692011. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Morgan DO: Principles of CDK regulation.
Nature. 374:131–134. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Kim JK and Diehl JA: Nuclear cyclin D1: An
oncogenic driver in human cancer. J Cell Physiol. 220:292–296.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wolf G and Shankland SJ: Cell cycle
control in glomerular disease. Prog Cell Cycle Res. 5:71–79.
2003.PubMed/NCBI
|
|
37
|
Shankland SJ and Wolf G: Cell cycle
regulatory proteins in renal disease: Role in hypertrophy,
proliferation, and apoptosis. Am J Physiol Renal Physiol.
278:F515–F529. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li Y, Liu Z, Guo X, Shu J, Chen Z and Li
L: Aristolochic acid I-induced DNA damage and cell cycle arrest in
renal tubular epithelial cells in vitro. Arch Toxicol. 80:524–532.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Marshall CB and Shankland SJ: Cell cycle
and glomerular disease: A minireview. Nephron Exp Nephrol.
102:e39–48. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Dai Y, Sui W, Lan H, Yan Q, Huang H and
Huang Y: Microarray analysis of micro-ribonucleic acid expression
in primary immunoglobulin A nephropathy. Saudi Med J. 29:1388–1393.
2008.PubMed/NCBI
|
|
41
|
Silveri L, Tilly G, Vilotte JL and Le
Provost F: MicroRNA involvement in mammary gland development and
breast cancer. Reprod Nutr Dev. 46:549–556. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zhang B, Pan X, Cobb GP and Anderson TA:
microRNAs as oncogenes and tumor suppressors. Dev Biol. 302:1–12.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Khoshnaw SM, Green AR, Powe DG and Ellis
IO: MicroRNA involvement in the pathogenesis and management of
breast cancer. J Clin Pathol. 62:422–428. 2009. View Article : Google Scholar : PubMed/NCBI
|