Introduction
Cervical cancer (CC) is the fourth most common type
of cancer in women worldwide and the leading cause of death, with
~530,000 new cases and 275,000 deaths occurring each year (1,2).
Although CC treatments and screening methods continue to evolve,
with early diagnosis and treatment reducing its mortality rate, the
5-year cancer-specific survival rate remains poor (3). Therefore, it is of great importance to
recognize the lack of effective therapies and to identify novel
methods for improving the accuracy of diagnosis and the prognosis
of patients with CC. Discovering effective cancer treatments is
highly challenging due to tumor invasion and metastasis (4). Therefore, investigation of biomarkers
that can influence the pathogenesis and progression of cancer is of
great interest for the diagnosis and treatment of CC.
NUAK family kinase 2 (NUAK2) is a member of the
AMP-activated protein kinase family, which is located at 1q32 and
can be suppressed by the tumor suppressor hepatic kinase B1, as
well as by NF-κB, which inhibits death receptor signaling
activation (5–7). NUAK2 has been reported to serve a key
role in cancer development and tumor progression. It has been
reported that NUAK2 is upregulated in gastric cancer tissues, and
that it promotes the proliferation of gastric cancer cells and
regulates the cell cycle, resulting in upregulation of
proliferation and cancer stem cell marker expression (8). A previous study revealed that NUAK2
expression is increased in glioma tissues and is associated with
advanced disease stage, whereas in vitro experiments
demonstrated that overexpression of NUAK2 promoted the
proliferation, migration and invasion of A172 glioblastoma cells
(9).
The cytoplasmic FMR1-interacting protein (CYFIP)
gene family has two highly conserved members, CYFIP1 and CYFIP2,
which are ~145-kDa proteins with high homology (88% identity and
95% similarity) in their amino acid sequences (10). It has been reported that CYFIP2 mRNA
expression is significantly reduced in gastric cancer tissues
compared with that in non-cancerous tissues, and knockdown of
CYFIP2 has been shown to promote cell proliferation and colony
formation, and inhibit apoptosis (11). Additionally, overexpression of
CYFIP2 has been demonstrated to promote apoptosis-like death of
colorectal cancer cells (12).
The present study was undertaken to investigate the
biological functions of NUAK2 in CC. The expression of NUAK2 in CC
tissues and cells was examined, and further experiments were
conducted to determine whether interference with NUAK2 expression
could suppress the proliferation, migration, invasion and
epithelial-to-mesenchymal transition (EMT) of CC cells. The role of
CYFIP2 in this process was also investigated.
Materials and methods
Bioinformatics analysis
The interaction between NUAK2 and CYFIP2 was
predicted using the Search Tool for the Retrieval of Interacting
Genes/Proteins (STRING) database (version 11.0, http://string-db.org/) (13). The UALCAN cancer database
(http://ualcan.path.uab.edu) was used to
retrieve data regarding the expression levels of NUAK2 and CYFIP2
in the primary tumor tissues (n=305) of patients with cervical
squamous cell carcinoma (CESC) and normal tissues (n=3) from
healthy controls (14); these
databases were from The Cancer Genome Atlas.
Tissues and cell lines
CC tissues and pair-matched non-cancerous cervical
tissues (2 cm from the lesions) were obtained from 36 patients
(age, 30–65 years) diagnosed with CC at Wuhan Children's Hospital
(Wuhan, China) from April 2018 to September 2019. Patients were
excluded from the study if they received chemotherapy or
radiotherapy prior to surgery. Written informed consent was
obtained from all patients.
The HeLa cervical adenocarcinoma cell line, C-33A,
SiHa, CaSKi and HCC-94 cervical squamous cell carcinoma cell lines,
and Ect1 human normal cervical cell line were obtained from The
Cell Bank of Type Culture Collection of The Chinese Academy of
Sciences. All cell lines were cultured in Dulbecco's modified
Eagle's medium (DMEM) supplemented with 10% bovine calf serum, 100
µg/ml penicillin and 100 µg/ml streptomycin (all from Gibco; Thermo
Fisher Scientific, Inc.). Cells were incubated at 37°C in an
incubator containing 5% CO2. The medium was changed
every 2 days and cells were grown to logarithmic phase for use in
subsequent experiments.
Cell transfection
Two small interfering RNA (siRNA) sequences against
CYFIP2 (si-CYFIP2-1/si-CYFIP2-2) and a nontargeting siRNA used as a
negative control (si-NC) were designed by Shanghai GenePharma Co.,
Ltd. Targeting and nontargeting sequences were as follows:
si-CYFIP2-1, 5′-AGGCTAACTTTGACACAAACT-3′; si-CYFIP2-2,
5′-GGCTAACTTTGACACAAACTT-3′; si-NC, 5′-GCGTTCTTAACTTTGAACC-3′.
siRNAs were transfected into cells using Lipofectamine®
3000 (Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol. Briefly, HeLa cells were seeded at a
density of 2×105 cells/ml in 24-well plates and cultured
for 24 h, after which they were transfected with siRNAs targeting
CYFIP2 and si-NC; the knockdown efficiency was verified 48 h after
transfection by western blotting and reverse
transcription-quantitative PCR (RT-qPCR). All the siRNAs were used
at a final concentration of 5 nM. In addition, two NUAK2-specific
short hairpin RNAs (shRNAs) (shRNA-NUAK2-1/shRNA-NUAK2-2) and a
scrambled shRNA used as a negative control (shRNA-NC) were obtained
from Shanghai GenePharma Co., Ltd. The sequences were as follows:
shRNA-NUAK2-1, sense 5′-CCATAAGATCCTAGTGAAA-3′, antisense
5′-TTTCACTAGGATCTTATGG-3′; shRNA-NUAK2-2, sense
5′-GCATGACCATAAGATCCTA-3′, antisense 5′-TAGGATCTTATGGTCATGC-3′;
shRNA-NC, sense 5′-GATCCCCTTCTCCGAACG-3′, antisense
5′-AGCTAAAAATTCTCCGAAC-3′. The corresponding shRNA oligonucleotides
(synthesized by Guangzhou RiboBio Co., Ltd.) were cloned into the
pLentiLox 3.7 lentiviral plasmid [American Type Culture Collection
(ATCC)]. Lipofectamine 3000 was used to transfect 293T cells (ATCC)
at a ratio of 3 µg lentiviral construct and 6 µg package mix [pLP1
(3): pLP2 (2): pLP/VSVG (3); (Invitrogen; Thermo Fisher Scientific,
Inc.)] cultured at 37°C, according to the manufacturer's
recommendations. The supernatants were collected at 48 and 72 h
then mixed for ultracentrifugation (4°C, 72,000 × g, 2 h) to obtain
lentiviral particles. The lentivirus (multiplicity of infection,
10) and HeLa cells were incubated in DMEM supplemented with 10%
fetal bovine serum containing 8 µg/ml polybrene (MineBio Life
Sciences Ltd.) at 37°C for 12 h and then washed with PBS. The
medium was then replaced with fresh medium and cells were cultured
for a total of 2 days prior to subsequent experiments.
RT-qPCR analysis
The mRNA expression levels of NUAK2 and CYFIP2 were
determined via RT-qPCR. Total RNA was extracted from tissues and
cells using TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.). cDNA was synthesized using the PrimeScript™ RT
kit (Takara Bio, Inc.) under the following conditions: 15 min at
37°C and 5 sec at 85°C. The synthesized cDNA was used as a template
for PCR amplification. The reactions were performed using SYBR
Green Taq Mix (Takara Bio, Inc.) in a Real-Time PCR Detection
System (Bio-Rad Laboratories, Inc.) under the following conditions:
45°C for 3 min, 95°C for 10 sec, 40 cycles at 95°C for 15 sec then
58°C for 1 min. Relative mRNA expression levels were quantified
using the 2−ΔΔCq method (15) and normalized to GAPDH. All RT-qPCR
experiments were performed in triplicate. The primer sequences used
were as follows: NUAK2, forward, 5′-TGAGAAACGACGGAGACAAGCTGCT-3′
and reverse, 5′-GTCTGGAGGTTTTGCTGCAGGTCTG-3′; CYFIP2 forward,
5′-TGGCGTCATCATTCCGTATCC-3′ and reverse, 5′-GTCAGGTCCTCACTCAAGC-3′;
and GAPDH forward, 5′-TGTGTCCGTCGTGGATCTGA-3′ and reverse,
5′-CCTGCTTCACCACCTTCTTGA-3′.
Western blotting
The protein expression levels of NUAK2, CYFIP2,
E-cadherin, N-cadherin, Snail and ZEB1 were determined via western
blotting. Total protein was extracted from harvested cells using
RIPA lysis buffer (Beyotime Institute of Biotechnology). The
protein concentration was measured using a bicinchoninic acid
protein assay kit (Beyotime Institute of Biotechnology). A total of
30 µg protein per lane was separated via SDS-PAGE on 15% gels and
separated proteins were subsequently transferred to polyvinylidene
fluoride membranes (EMD Millipore). Membranes were then blocked
with 10% non-fat milk for 2 h at room temperature, and incubated
with the following primary antibodies overnight at 4°C: Anti-NUAK2
(1:500; cat. no. ab107287; Abcam), anti-CYFIP2 (1:500; cat. no.
sc-134308; Santa Cruz Biotechnology, Inc.), anti-E-cadherin (1:500;
cat. no. sc-8426; Santa Cruz Biotechnology, Inc.), anti-N-cadherin
(1:500; cat. no. sc-8424; Santa Cruz Biotechnology, Inc.),
anti-Snail (1:500; cat. no. sc-271977; Santa Cruz Biotechnology,
Inc.), anti-ZEB1 (1:500; cat. no. sc-10572; Santa Cruz
Biotechnology, Inc.), anti-GAPDH (1:2,000; cat. no. sc-47724; Santa
Cruz Biotechnology, Inc.) and anti-β-actin (1:1,000; cat. no.
sc-47778; Santa Cruz Biotechnology, Inc.). Subsequently, the
membranes were washed with TBS-0.1% Tween-20 (TBST) for 5 min at
room temperature. After three washes, the membranes were incubated
at room temperature with horseradish peroxidase (HRP)-conjugated
goat anti-rabbit IgG (1:2,000; cat. no. sc-2004; Santa Cruz
Biotechnology, Inc.), HRP-conjugated goat anti-mouse IgG (1:2,000;
cat. no. sc-2005; Santa Cruz Biotechnology, Inc.) or horseradish
peroxidase (HRP)-conjugated mouse anti-goat IgG (1:2,000; cat. no.
sc-2354; Santa Cruz Biotechnology, Inc.) for 2 h. After washing
with TBST three times (10 min/wash), the protein bands were
visualized using Western Blotting Luminescent Reagent (Santa Cruz
Biotechnology, Inc.) and analyzed using ImageJ software (version
1.43; National Institutes of Health).
Cell Counting Kit-8 (CCK-8) assay
Cell proliferation was determined using a Cell
Counting Kit-8 (CCK-8) assay (cat. no. C0037; Beyotime Institute of
Biotechnology) according to the manufacturer's instructions.
Briefly, HeLa cells were seeded at a quantity of 1×104
cells/well in 96-well plates and cultured for 6 h, after which they
were incubated with 15 µl/well CCK-8 solution at 37°C for 24, 48 or
72 h. Subsequently, the optical density (OD) was calculated at a
wavelength of 450 nm to determine cell proliferation.
Immunofluorescence (IF) staining
assay
To observe the expression of Ki67, IF staining was
performed. HeLa cells were seeded onto glass coverslips in 6-well
plates. Cells were incubated with lentiviruses containing
shRNA-NUAK2 at 37°C for 2 days to silence NUAK2 gene expression.
The cells were then incubated with si-CYFIP2 for an additional 36 h
to obtain cells with low expression of NUAK2 and CYFIP2. After
that, the cells were fixed with 4% paraformaldehyde (PFA; Millipore
Sigma) at room temperature for 10 min, washed in PBS and blocked
for 15 min in QuickBlock™ Blocking Buffer for Immunol Staining
(Beyotime Biotechnology) at room temperature. After incubation with
a primary antibody against anti-Ki67 (1:200; cat. no. sc-23900;
Santa Cruz Biotechnology, Inc.) at 4°C overnight, cells were washed
in PBS and incubated with a secondary antibody conjugated with
Alexa Fluor 488 (1:1,000; cat. no. A32723; Invitrogen; Thermo
Fisher Scientific, Inc.) for 1 h at room temperature. Cells were
then washed in PBS, incubated for 5 min with DAPI (1 µg/ml; cat.
no. D1306; Invitrogen; Thermo Fisher Scientific, Inc.), and
observations were performed using a fluorescence microscope
(magnification, ×100; Olympus Corporation) and were analyzed with
ImageJ version 1.43 software.
Wound healing assay
HeLa cells were seeded at 6×104
cells/well in 6-well plates, cultured until they reached 80–90%
confluence and were subjected to serum starvation for 4 h.
Subsequently, a sterile 10-µl pipette tip was used to create a
linear scratch in the cell monolayer, the cells were washed with
PBS to remove debris, and the medium was replaced with serum-free
DMEM/F12 1:1. Images were captured under a light microscope
(Olympus Corporation) prior to and at 24 h after incubation.
Transwell invasion assay
HeLa cells (2×104 cells/well) were
cultured in the upper chamber of Transwell plates (8-µm pore size;
Corning, Inc.). The surface of the upper chamber was pre-coated
with Matrigel™ (BD Biosciences) at 37°C for 1 h. Complete medium
(500 µl) supplemented with 10% bovine calf serum (Gibco; Thermo
Fisher Scientific, Inc.) was added into the lower chamber. After
24-h incubation at 37°C, cells remaining on the upper membrane were
wiped away, whereas cells that had invaded across the membrane were
fixed with 4% PFA for 10 min at room temperature, stained with 0.1%
crystal violet for 15 min at room temperature and counted under a
light microscope.
Co-immunoprecipitation (Co-IP)
The Pierce Co-IP Kit (cat. no. 26149; Thermo Fisher
Scientific, Inc.) was used to explore the interaction between
proteins, according to the manufacturer's protocol. Briefly, the
HeLa cells cultured in a 10-cm plate were washed twice with
pre-cooled PBS, and the PBS was discarded. Subsequently, 1 ml lysis
buffer was used to fully lyse the cells, the lysate was transferred
into 1.5-ml microcentrifuge tubes, centrifuged for 15 min (4°C,
14,000 × g), and the supernatant protein was taken for
quantification. The total volume was made up to 500 µl with lysis
buffer after 500 µg protein was taken. Subsequently, 30 µl Agarose
A + G was used to pre-clear lysate at 4°C for 2 h. The pre-cleared
lysate was centrifuged at 4°C for 5 min at 1,000 × g and the
supernatant was removed and incubated overnight with shaking at 4°C
by adding the corresponding antibody. The following antibodies were
used: NUAK2 (1:10; cat. no. sc-374348; Santa Cruz Biotechnology,
Inc.), CYFIP2 (1:50; cat. no. sc-134308; Santa Cruz Biotechnology,
Inc.) and IgG (1:50; cat. no. sc-69786; Santa Cruz Biotechnology,
Inc.), which was used as a control. After 12 h of antibody
conjugation, Agarose A + G (30 µl/tube) was added and shaken at 4°C
for 4 h. The supernatant was discarded after centrifugation at
1,500 × g for 5 min to obtain the sediment. The precipitate was
then washed three times with pre-chilled PBS to obtain the protein
sample for western blotting.
Statistical analysis
All experimental data are presented as the mean ±
standard deviation of at least three independent experiments.
Statistical analysis was performed using GraphPad Prism 8 software
(GraphPad Software, Inc.). In order to assess whether data obtained
from the tissue samples followed a normal distribution, a
Shapiro-Wilk test was used. Based on this information, data were
further analyzed using parametric tests. To compare differences
among multiple groups, one-way ANOVA followed by Tukey's post hoc
test was used. P<0.05 was considered to indicate a statistically
significant difference.
Results
Expression of NUAK2 in CC tissues and
cells
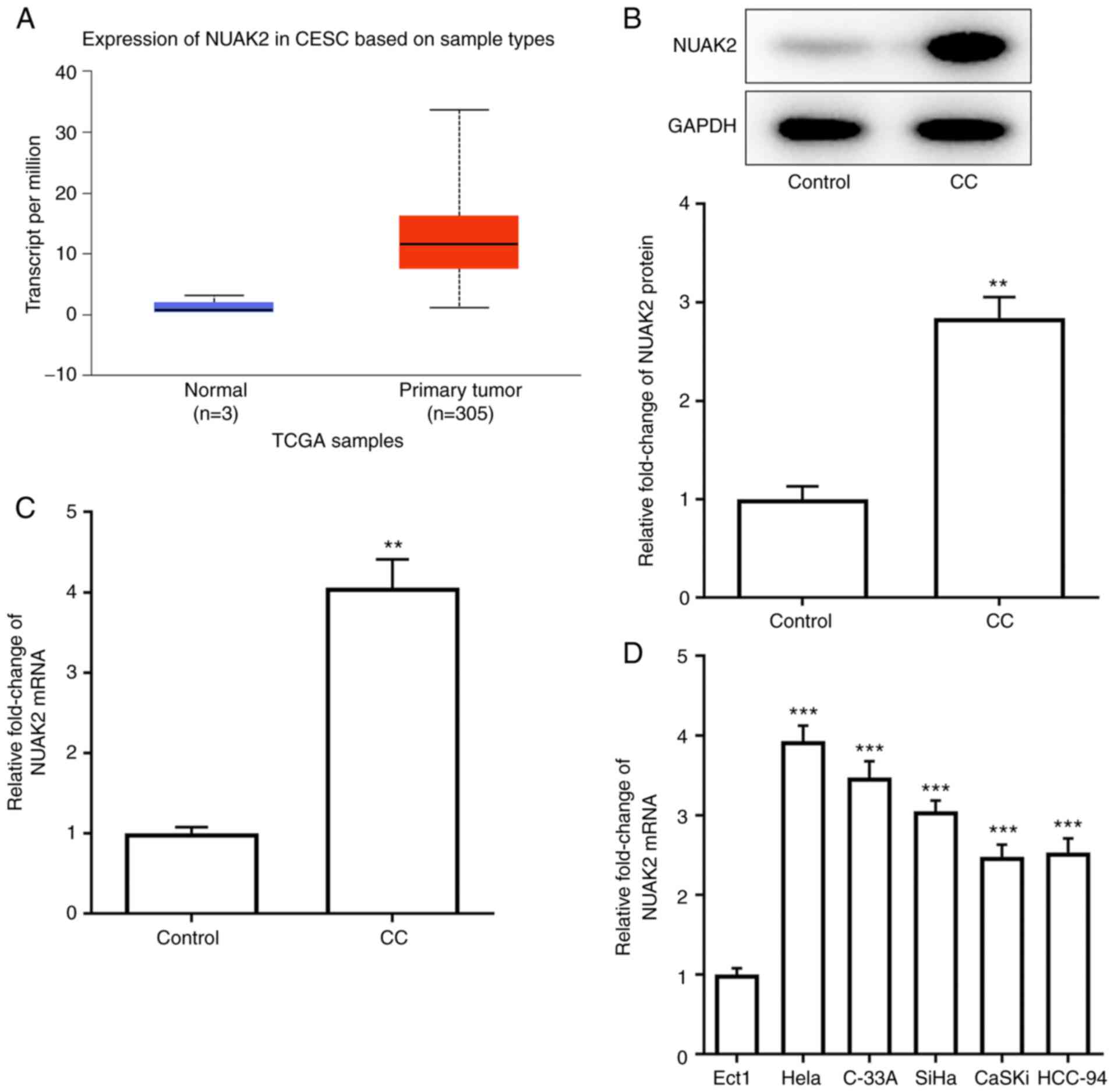
UALCAN website predicted the expression levels of
NUAK2 in CESC (Fig. 1A). The
expression of NUAK2 was detected in CC tissues by RT-qPCR and
western blotting. As shown in Fig. 1B
and C, the protein and mRNA expression levels of NUAK2 were
significantly increased in CC tissues compared with those in the
control group. Similarly, NUAK2 exhibited higher expression in CC
cell lines compared with that in Ect1 cells, with the maximum
expression observed in HeLa cells (Fig.
1D). Therefore, HeLa cells were selected for subsequent
experiments. These results indicated that NUAK2 was highly
expressed in CC tissues and cell lines.
Effect of NUAK2 knockdown on CC cell
proliferation
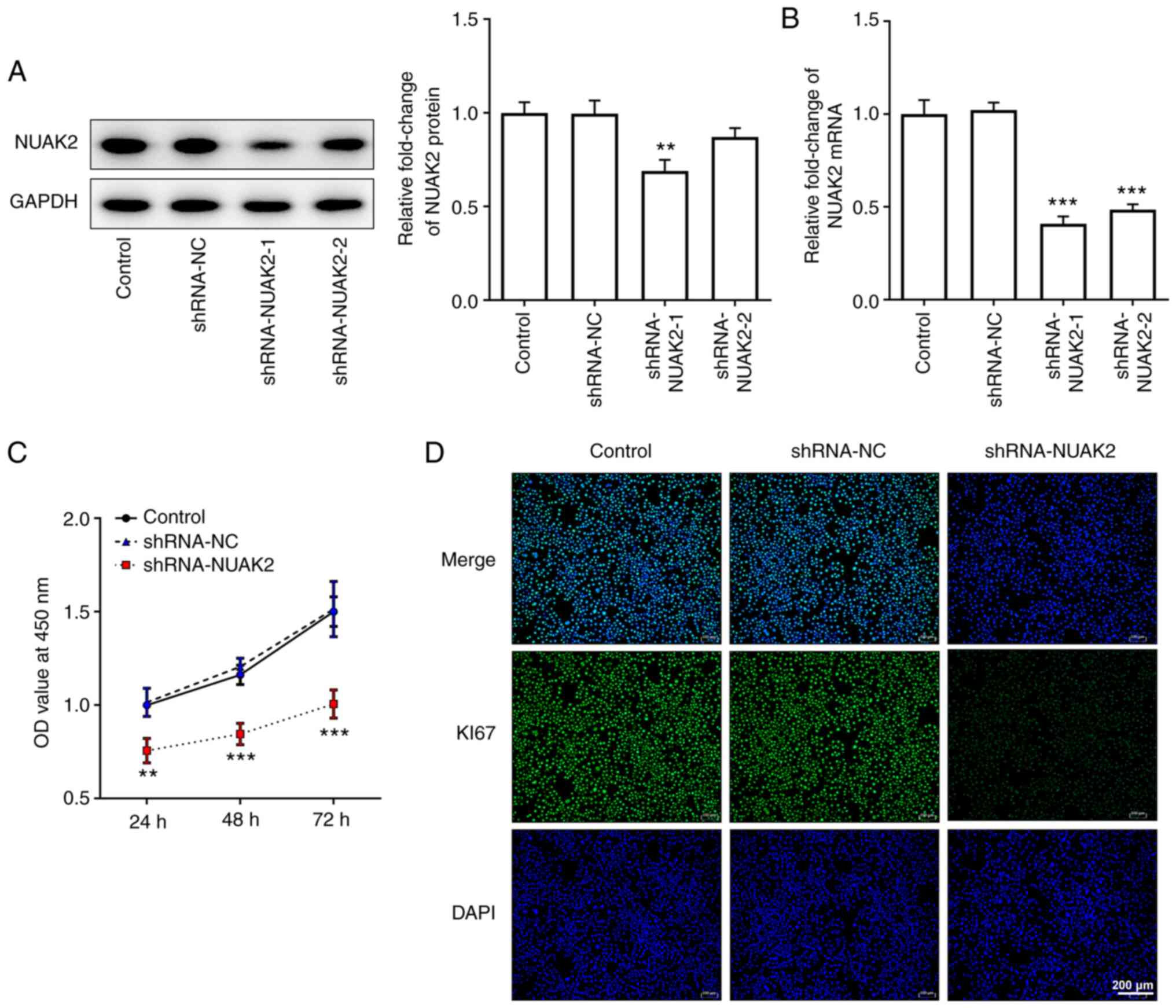
In order to determine the role of NUAK2 expression
in CC, shRNA-NUAK2 was used. As shown in Fig. 2A and B, transfection of HeLa cells
with shRNA-NUAK2-1 silenced NUAK2 expression more efficiently
compared with shRNA-NUAK2-2; therefore, shRNA-NUAK2-1 was selected
for subsequent experiments. As determined by CCK-8 assay, the OD
value at 450 nm in the shRNA-NUAK2 group was significantly
attenuated compared with that in the shRNA-NC group, whereas there
were no changes between the untransfected control group and the
shRNA-NC group (Fig. 2C).
Furthermore, the protein expression levels of Ki67 were markedly
decreased in the shRNA-NUAK2 group compared with those in the
control group and the shRNA-NC group (Fig. 2D). These results indicated that
knockdown of NUAK2 inhibited CC cell proliferation.
Effect of NUAK2 knockdown on CC cell
migration, invasion and EMT
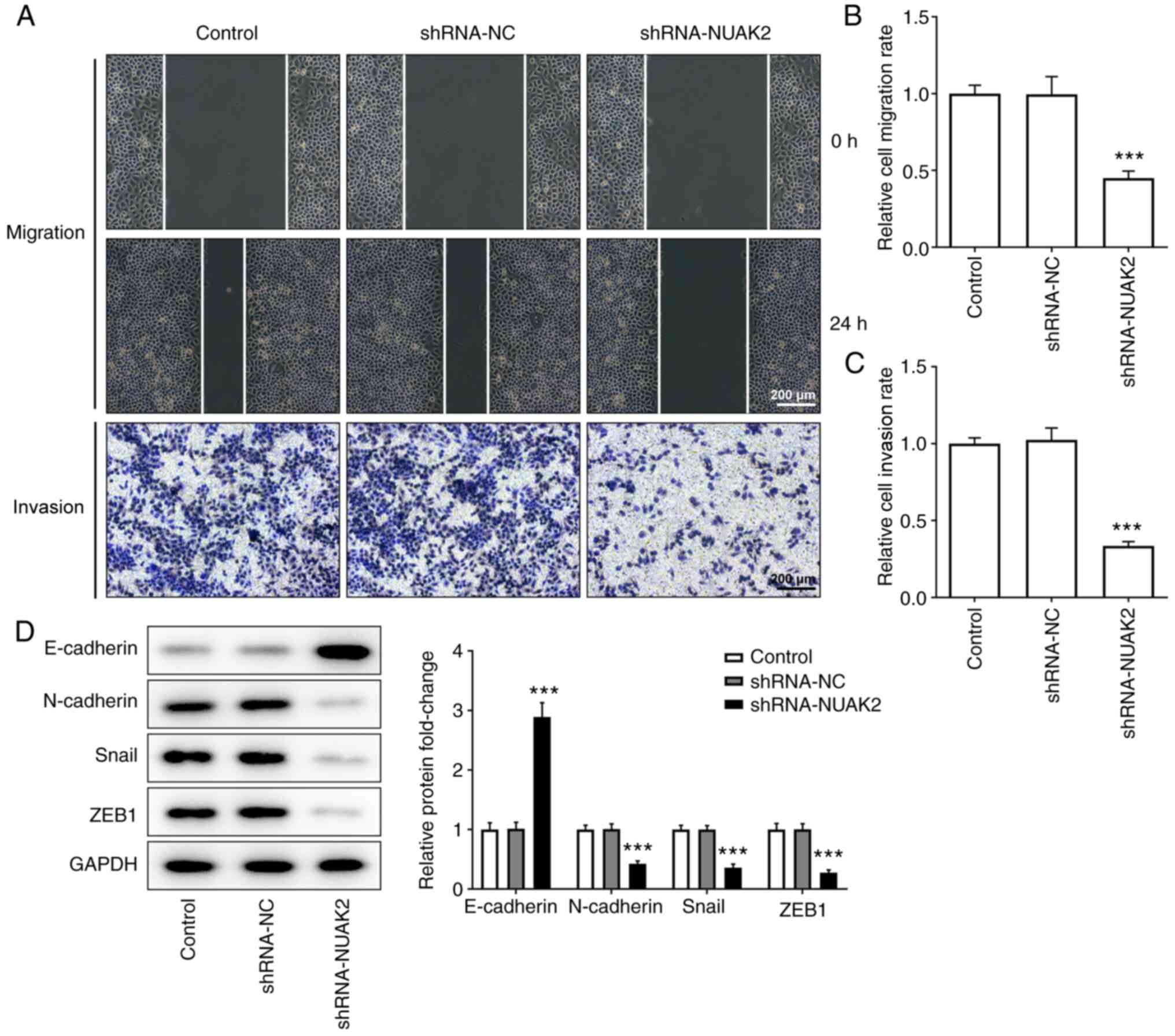
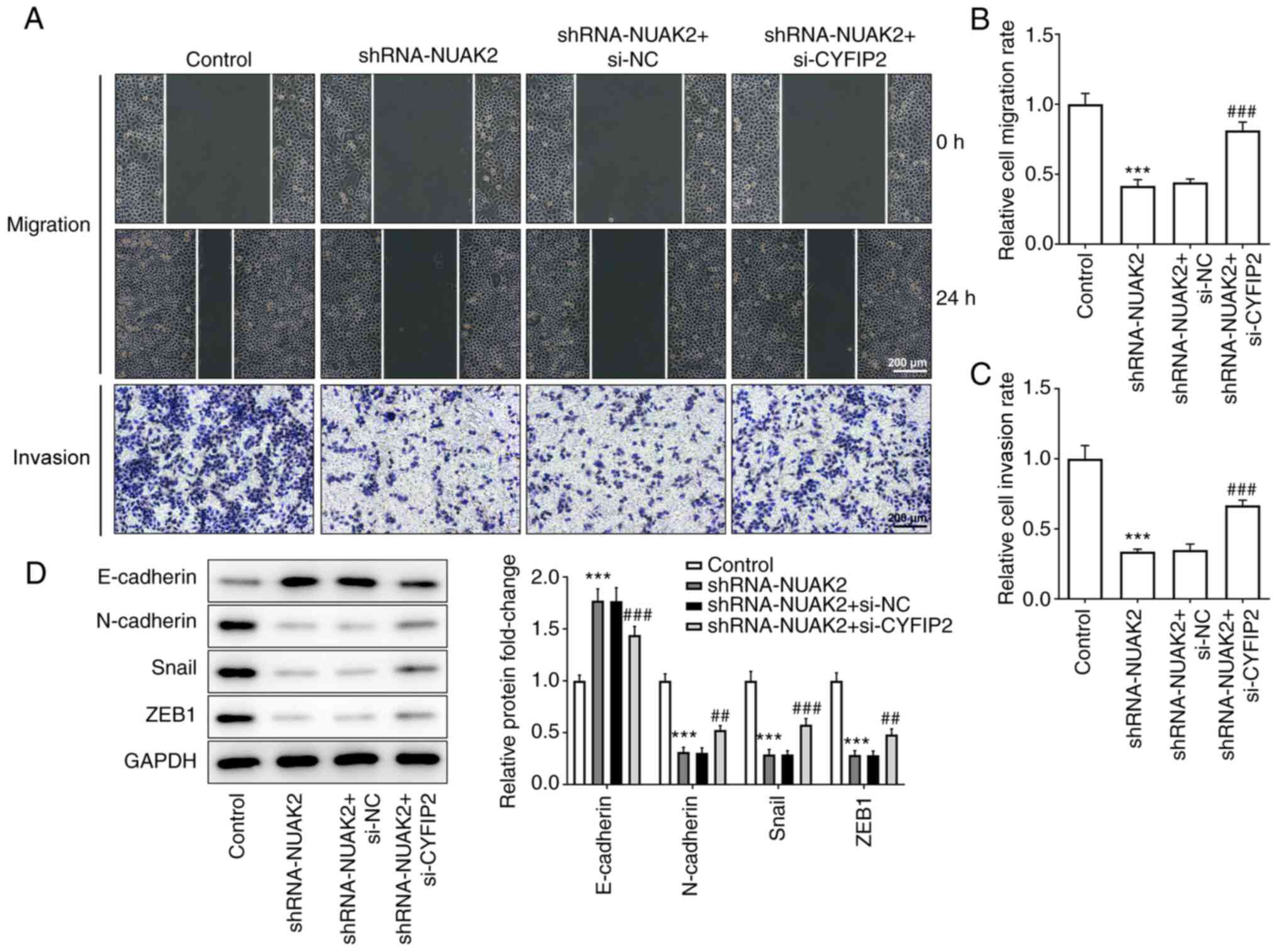
As shown in Fig.
3A-C, after 24 h of incubation, the scratches in the cell
monolayer were wider and the relative cell migration rate was lower
in the shRNA-NUAK2 group compared with that in the control group
and the shRNA-NC group, suggesting that cell migration was
inhibited when NUAK2 expression was knocked down. Similarly, the
results of the Transwell invasion assay revealed that, compared
with in the control and shRNA-NC groups, the invasive ability of
HeLa cells was suppressed in the shRNA-NUAK2 group. Furthermore,
western blotting revealed that NUAK2 knockdown elevated the
expression levels of E-cadherin, and decreased the expression
levels of N-cadherin, Snail and ZEB1 compared with those in the
shRNA-NC group (Fig. 3D),
suggesting that NUAK2 promoted EMT in CC cells. These data
confirmed that NUAK2 knockdown suppressed the EMT of CC cells.
Taken together, these data indicated that knockdown of NUAK2
expression decreased the mobility of HeLa cells after
transfection.
Expression of CYFIP2 in CC tissues and
cells
The UALCAN website predicted the expression levels
of CYEIP2 in CESC (Fig. 4A). To
reveal the molecular mechanisms underlying the role of NUAK2 in CC,
the STRING database was used to identify whether NUAK2 directly
binds to CYFIP2. The results of western blotting (Fig. 4B) and RT-qPCR (Fig. 4C) revealed that CYFIP2 expression
was decreased in tissues from patients with CC compared with that
in the control group. Subsequently, the mRNA expression levels of
CYFIP2 were detected in several CC cell lines (HeLa, C-33A, SiHa,
CaSKi and HCC-94), which demonstrated that CYFIP2 was downregulated
at different levels in these CC cell lines compared with that in
Ect1 cells (Fig. 4D). To verify the
targeted binding of NUAK2 and CYFIP2, Co-IP was performed. The
results of Co-IP demonstrated that NUAK2 could interact with CYFIP2
(Fig. 4E). The western blotting
results confirmed that NUAK2 knockdown increased the expression
levels of CYFIP2 (Fig. 4F). These
results indicated that CYFIP2 was expressed at low levels in CC and
was negatively associated with NUAK2.
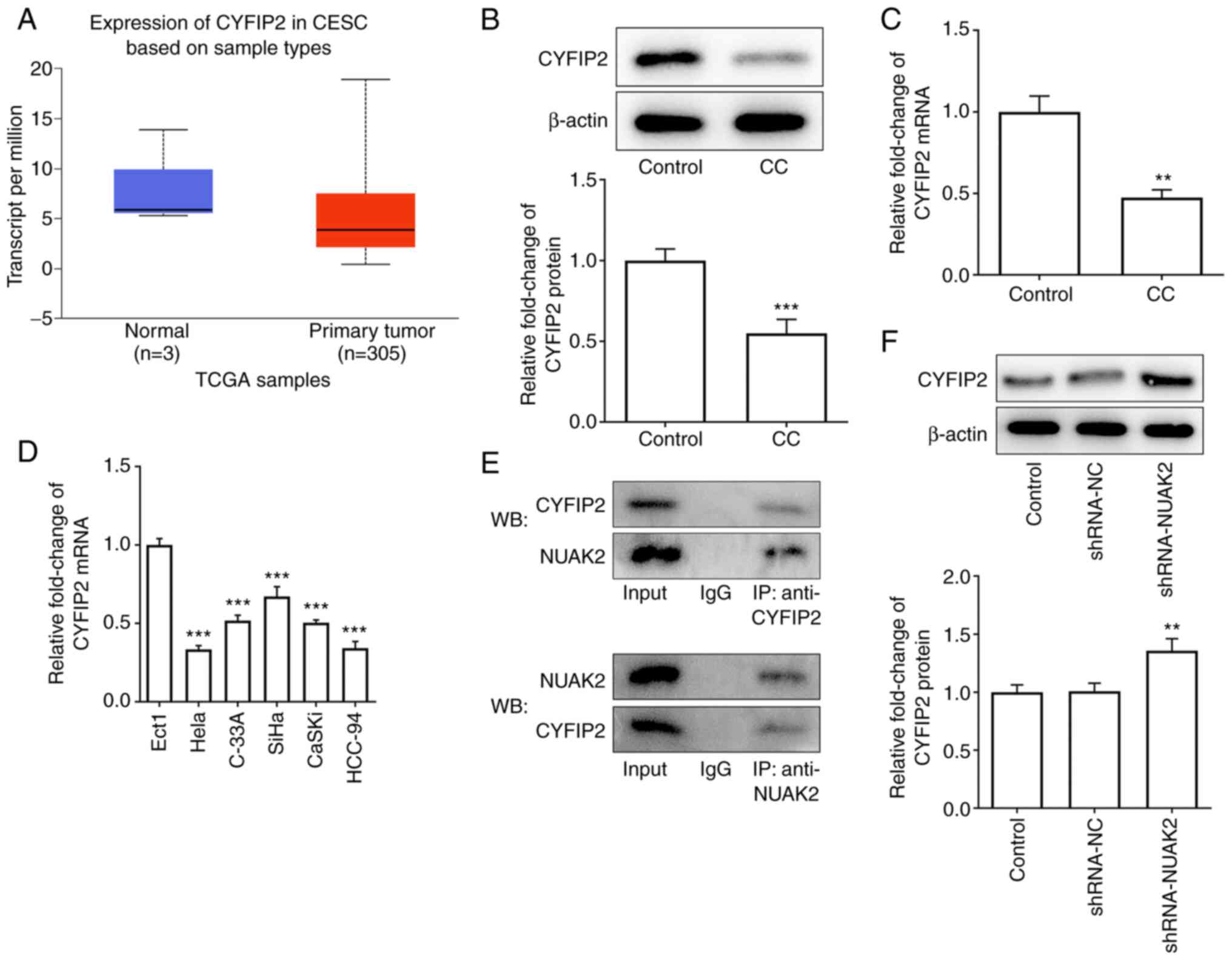 | Figure 4.CYFIP2 is downregulated in CC tissues
and cell lines, and is negatively associated with NUAK2. (A) UALCAN
website predicted the expression levels of CYFIP2 in CESC. (B)
CYFIP2 protein expression in CC samples was assessed via WB. (C)
CYFIP2 mRNA expression in CC samples was assessed via RT-qPCR.
**P<0.01, ***P<0.001 vs. Control. (D) mRNA expression levels
of CYFIP2 in CC cell lines were detected by RT-qPCR. ***P<0.001
vs. Ect1. (E) Co-IP was performed to detect the levels of CYFIP2 in
response to NUAK2. (F) CYFIP2 protein expression in HeLa cells
transfected with shRNA-NUAK2. **P<0.01 vs. shRNA-NC. CC,
cervical cancer; RT-qPCR, reverse transcription-quantitative PCR;
shRNA, short hairpin RNA; NC, negative control; IP,
immunoprecipitation; WB, western blotting; NUAK2, NUAK family
kinase 2; CYFIP2, cytoplasmic FMRP-interacting protein 2; TCGA, The
Cancer Genome Atlas. |
Effects of CYFIP2 knockdown on CC cell
proliferation
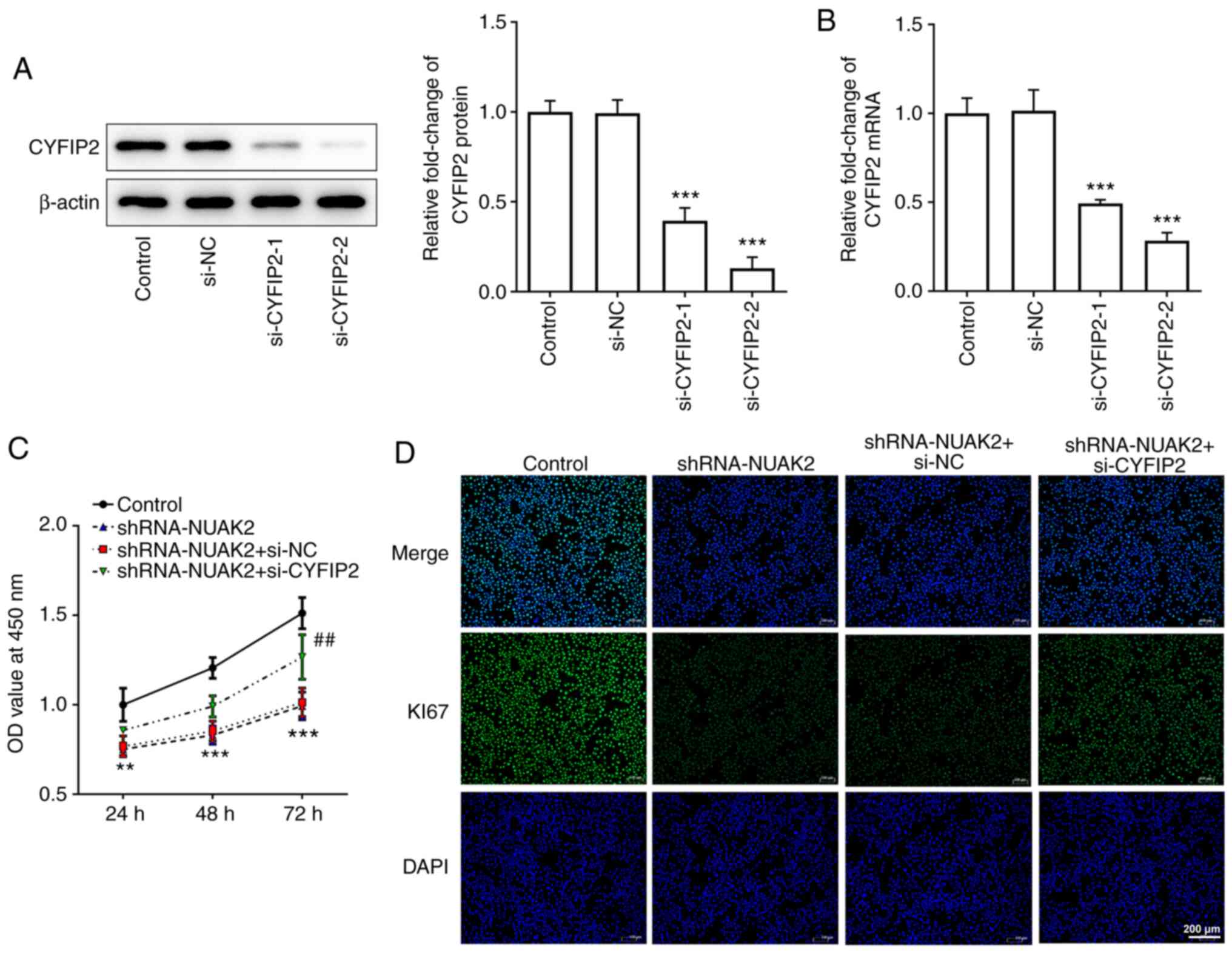
In order to further explore the role of CYFIP2 in
CC, si-CYFIP2 was used in the following experiments. The results
presented in Fig. 5A and B show
that both si-CYFIP2-1 and si-CYFIP2-2 could reduce the protein and
mRNA expression levels of CYFIP2 in HeLa cells, but si-CYFIP2-2 was
more efficient. Thus, si-CYFIP2-2 was selected to knock down CYFIP2
expression. The CCK-8 assay and IF revealed that si-CYFIP2 could
enhance cell proliferation compared with that in the shRNA-NUAK2+
si-NC group (Fig. 5C and D). These
results suggested that the effect of NUAK2 knockdown on CC cell
proliferation may be reversed by inhibition of CYFIP2
expression.
Effect of CYFIP2 knockdown on CC cell
migration, invasion and EMT
As presented in Fig.
6A-C, in HeLa cells transfected with shRNA-NUAK2, transfection
with siRNA to interfere with CYFIP2 expression could accelerate gap
closure in the cell monolayer and enhanced the invasive ability of
HeLa cells compared with in the shRNA-NUAK2 group and the
shRNA-NUAK2+ siRNA-NC group. In addition, inhibition of CYFIP2
expression reversed the effect of NUAK2 knockdown on the EMT
phenotype of HeLa cells (Fig. 6D).
These results demonstrated that interference with NUAK2 inhibited
CC cell migration and the EMT process through upregulation of
CYFIP2.
Discussion
CC has become a major health concern among women
worldwide, and the survival rate of patients with advanced CC is
poor (16). Therefore, the search
for novel biomarkers is of great importance in the diagnosis and
treatment of CC. The present study focused on NUAK2.
Previous studies have identified an important role
for NUAK2 in human tumors. Amplification of NUAK2 has been reported
to facilitate the development of melanoma (17). Furthermore, it was experimentally
demonstrated that knockdown of NUAK2 inhibited the proliferation
and migration of melanoma cells (18). NUAK2 upregulation was previously
observed in the surface epithelium of ovaries in a large cohort of
patients with ovarian plasmacytoma, and was predicted to have a
role in driving mutations in ovarian cancer; furthermore, patients
with lower NUAK2 expression levels were found to have longer
overall survival rates (19). In
addition, upregulation of NUAK2 in the maternal kidney has been
demonstrated to accelerate cellular senescence in short-lived
neonatal mice (20). These previous
studies indicated that NUAK2 may play a key a role in cancer.
However, to the best of our knowledge, there are no reports on the
role of NUAK2 in CC; therefore, the present study was undertaken to
investigate the role of NUAK2 expression in CC. Through the search
of the UALCAN cancer database, NUAK2 expression was found to be
elevated in the tissues of patients with CESC, which is consistent
with the literature reporting that NUAK2 is highly expressed in
gastric cancer and glioblastoma (8,9). The
results were also validated in tissues collected from patients with
CC and in CC cell lines.
It is commonly known that the developmental process
of tumor cells, including cell proliferation, migration and
invasion, is important in the study of cancer. EMT is the important
biological process by which epithelial-derived malignant cells are
transformed into cells with a mesenchymal phenotype with the
ability to migrate and invade; EMT has an important role in cancer
metastasis (21). Therefore, to
investigate the role of NUAK2 in CC, the present study primarily
observed its role in cell progression and EMT. The results
demonstrated that NUAK2 knockdown inhibited CC cell proliferation,
migration, invasion and EMT.
In order to further investigate the mechanism of
action of NUAK2, the online STRING database was used to search for
genes that bind to NUAK2, which determined that CYFIP2 could
directly bind to NUAK2. CYFIP2 is often studied in neurological
disorders and is associated with neuronal functions (10). CYFIP2 has also been proposed as a
candidate gene for intellectual disability and autism (22). Furthermore, CYFIP2 has been
suggested as a potential target in the treatment of Alzheimer's
disease (23). In a previous study,
CYFIP2 expression was reduced in gastric cancer, and inhibition of
CYFIP2 promoted gastric cancer cell proliferation and chemotherapy
resistance to 5-fluorouracil (11).
A whole-exome sequencing study previously revealed that mutational
dynamics and genetic variation, as well as aberrant DNA repair,
tumor cell cycle control and apoptotic pathways were associated
with CYFIP2 in endometrial cancer in the Taiwanese population
(24). In the present study, it was
predicted that the expression levels of CYFIP2 would be decreased
in CC tissues, which was confirmed through analysis of CC tumor
samples collected from patients and CC cell lines. A Co-IP assay
was conducted to detect the direct binding between NUAK2 and
CYFIP2.
Once the present study determined that CYFIP2 was
negatively associated with NUAK2, si-CYFIP2 was selected to explore
the mechanism of the inhibitory effects of NUAK2 knockdown on CC
cell processes. The results revealed that inhibition of CYFIP2
expression could partially counteract the effects of NUAK2
knockdown on CC cell proliferation, migration and EMT.
In conclusion, NUAK2 was revealed to serve a crucial
role in CC cell proliferation, migration, invasion and EMT by
regulating CYFIP2. These findings suggested that NUAK2 may be a
potential therapeutic target for CC detection and clinical
treatment. However, there are limitations to the present study.
Firstly, the lack of validation of the results in additional cell
lines and in vivo experiments. Secondly, the effects of
NUAK2 overexpression on the progression/prognosis of CC were not
determined. Thirdly, the mechanism underlying the negative
regulation of CYFIP2 by NUAK2 was not deeply investigated. These
issues require further in-depth investigations and will be
addressed in future studies.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
LY and YL designed the study and wrote the
manuscript. YL and XS performed the experiments. LL collected,
analyzed and interpreted the data. YL and LY confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
The present study was approved by Tongji Medical
College (approval no. WHCH 2018047; Wuhan, China). Written informed
consent was obtained from all patients.
Patient consent for publication
Written informed consent was obtained from all
patients.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Olusola P, Banerjee HN, Philley JV and
Dasgupta S: Human papilloma virus-associated cervical cancer and
health disparities. Cells. 8:6222019. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Vu M, Yu J, Awolude OA and Chuang L:
Cervical cancer worldwide. Curr Probl Cancer. 42:457–465. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Cohen PA, Jhingran A, Oaknin A and Denny
L: Cervical cancer. Lancet. 393:169–182. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Mei D, Zhu Y, Zhang L and Wei W: The Role
of CTHRC1 in regulation of multiple signaling and tumor progression
and metastasis. Mediators Inflamm. 2020:95787012020. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zagorska A, Deak M, Campbell DG, Banerjee
S, Hirano M, Aizawa S, Prescott AR and Alessi DR: New roles for the
LKB1-NUAK pathway in controlling myosin phosphatase complexes and
cell adhesion. Sci Signal. 3:ra252010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bright NJ, Thornton C and Carling D: The
regulation and function of mammalian AMPK-related kinases. Acta
Physiol (Oxf). 196:15–26. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Sun X, Gao L, Chien HY, Li WC and Zhao J:
The regulation and function of the NUAK family. J Mol Endocrinol.
51:R15–R22. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Tang L, Tong SJ, Zhan Z, Wang Q, Tian Y
and Chen F: Expression of NUAK2 in gastric cancer tissue and its
effects on the proliferation of gastric cancer cells. Exp Ther Med.
13:676–680. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Fu TG, Wang L, Li W, Li JZ and Li J:
miR-143 inhibits oncogenic traits by degrading NUAK2 in
glioblastoma. Int J Mol Med. 37:1627–1635. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhang Y, Lee Y and Han K: Neuronal
function and dysfunction of CYFIP2: From actin dynamics to early
infantile epileptic encephalopathy. BMB Rep. 52:304–311. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Jiao S, Li N, Cai S, Guo H and Wen Y:
Inhibition of CYFIP2 promotes gastric cancer cell proliferation and
chemoresistance to 5-fluorouracil through activation of the Akt
signaling pathway. Oncol Lett. 13:2133–2140. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jackson RS II, Cho YJ, Stein S and Liang
P: CYFIP2, a direct p53 target, is leptomycin-B sensitive. Cell
Cycle. 6:95–103. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
von Mering C, Huynen M, Jaeggi D, Schmidt
S, Bork P and Sne B: STRING: A database of predicted functional
associations between proteins. Nucleic Acids Res. 31:258–261. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chandrashekar DS, Bashel B, Balasubramanya
SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BVSK and
Varambally S: UALCAN: A portal for facilitating tumor subgroup gene
expression and survival analyses. Neoplasia. 19:649–658. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Hung P, Zahnd WE, Brandt HM, Adams SA,
Wang S and Eberth JM: Cervical cancer treatment initiation and
survival: The role of residential proximity to cancer care. Gynecol
Oncol. 160:219–226. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Namiki T, Yaguchi T, Nakamura K, Valencia
JC, Coelho SG, Yin L, Kawaguchi M, Vieira WD, Kaneko Y, Tanemura A,
et al: NUAK2 amplification coupled with PTEN deficiency promotes
melanoma development via CDK activation. Cancer Res. 75:2708–2715.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Namiki T, Tanemura A, Valencia JC, Coelho
SG, Passeron T, Kawaguchi M, Vieira WD, Ishikawa M, Nishijima W,
Izumo T, et al: AMP kinase-related kinase NUAK2 affects tumor
growth, migration, and clinical outcome of human melanoma. Proc
Natl Acad Sci USA. 108:6597–6602. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Emmanuel C, Gava N, Kennedy C, Balleine
RL, Sharma R, Wain G, Brand A, Hogg R, Etemadmoghadam D, George J,
et al: Comparison of expression profiles in ovarian epithelium in
vivo and ovarian cancer identifies novel candidate genes involved
in disease pathogenesis. PLoS One. 6:e176172011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen JH, Tarry-Adkins JL, Matharu K, Yeo
GS and Ozanne SE: Maternal protein restriction affects gene
expression profiles in the kidney at weaning with implications for
the regulation of renal function and lifespan. Clin Sci (Lond).
119:373–384. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Aiello NM and Kang Y: Context-dependent
EMT programs in cancer metastasis. J Exp Med. 216:1016–1026. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zweier M, Begemann A, McWalter K, Cho MT,
Abela L, Banka S, Behring B, Berger A, Brown CW, Carneiro M, et al:
Spatially clustering de novo variants in CYFIP2, encoding the
cytoplasmic FMRP interacting protein 2, cause intellectual
disability and seizures. Eur J Hum Genet. 27:747–759. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Tiwari SS, Mizuno K, Ghosh A, Aziz W,
Troakes C, Daoud J, Golash V, Noble W, Hortobágyi T and Giese KP:
Alzheimer-related decrease in CYFIP2 links amyloid production to
tau hyperphosphorylation and memory loss. Brain. 139:2751–2765.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Chang YS, Huang HD, Yeh KT and Chang JG:
Identification of novel mutations in endometrial cancer patients by
whole-exome sequencing. Int J Oncol. 50:1778–1784. 2017. View Article : Google Scholar : PubMed/NCBI
|