Introduction
The global prevalence of diabetes has reached ~828
million cases, with an estimated 14% of the adult population
affected worldwide (1). Among
these, type 2 diabetes mellitus (T2DM) accounts for ~98% of cases.
Due to hyperinsulinemia and hyperglycemia, individuals with T2DM
are predisposed to the development of various types of cancer,
including colonic carcinoma (2,3). The
majority of colonic carcinoma cases originate from aberrant crypt
foci with abnormal cell proliferation (4). Given this clinical context, it is
noteworthy that a previous study has demonstrated that abnormal
epithelial proliferation is already present in the colonic mucosa
of a T2DM mouse model, even in the absence of colonic carcinoma
(5). Elucidating the mechanisms
underlying this abnormal proliferation provides insight into the
development of colonic carcinoma in the context of T2DM.
N6-methyladenosine (m6A) is
one of the most common RNA modifications (6). The regulation of m6A is
mediated by three functionally distinct classes of categories of
enzymes: ‘writers’ (methyltransferases) that install the m6A mark,
‘erasers’ (demethylases) that remove it and ‘readers’
(m6A-binding proteins) that specifically recognize the
mark and dictate downstream functional outcomes (7). m6A modifications influence
multiple aspects of RNA metabolism, including splicing, export,
translation efficiency and RNA stability, thereby modulating gene
expression and contributing to diverse pathological processes
(8). In the context of colonic
carcinoma, the dynamic regulation of m6A modifications
plays an important role in tumorigenesis (9). Given the implication of altered
m6A regulation in various metabolic and immune disorders
(10,11), it is noteworthy that T2DM, a
complex metabolic disease characterized by chronic low-grade
inflammation, is closely linked to dysregulated m6A
modification (12). However, the
potential involvement of m6A modification and the
specific regulatory role it may play in driving the
hyperproliferation of colonic epithelium under T2DM conditions
remains to be fully elucidated.
The present study demonstrated abnormal
proliferation of human colonic epithelium under T2DM conditions.
This was linked to the upregulation of insulin-like growth factor 2
mRNA-binding protein 2 (IGF2BP2). Furthermore, the present study
established that IGF2BP2, functioning as an m6A reader,
promoted cell proliferation by binding to and stabilizing midkine
(MDK) mRNA, thereby enhancing its expression.
Materials and methods
Analysis of m6A-related
enzyme expression in colonic epithelium of patients with T2DM
RNA sequencing data of colonic epithelium were
obtained from the Gene Expression Omnibus database (accession no.
GSE115313; http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE115313),
which included samples from 23 patients with T2DM and 19 patients
without T2DM or prediabetes (13).
Probe identifiers were mapped to gene symbols according to platform
annotation data. Probes corresponding to multiple genes were
excluded, while for genes represented by multiple probes, the
average expression value was calculated. Differential expression
analysis of genes encoding m6A-related enzymes between
the two groups was performed to identify key candidates. These,
together with other differentially expressed genes (P<0.05; fold
change >1.2) from sequencing, were subjected to further
bioinformatic analysis. Correlation analysis of their expression in
colonic epithelium was performed using the Gene Expression
Profiling Interactive Analysis (GEPIA) database (http://gepia.cancer–pku.cn/) (14). The potential for the differentially
expressed genes to be RNA-binding targets was predicted using
StarBase database (https://rnasysu.com/encori/index.php) (15). The potential m6A
modification sites were predicted using the sequence-based RNA
adenosine methylation site predictor (SRAMP; http://www.cuilab.cn/sramp) (16).
Tissue samples
Patients with colonic carcinoma who underwent
surgical resection at Sun Yat-sen Memorial Hospital (Guangdong,
China) between January 2015 and December 2020 were retrospectively
reviewed. The diagnosis criteria of the American Diabetes
Association were applied: i) Glycated hemoglobin ≥6.5%; ii) fasting
plasma glucose ≥7.0 mmol/l (126 mg/dl); iii) 2-h plasma glucose
≥11.1 mmol/l (200 mg/dl) during an oral glucose tolerance test; or
iv) random plasma glucose ≥11.1 mmol/l (200 mg/dl) in the presence
of classic hyperglycemic symptoms, such as polydipsia, polyuria or
unexplained weight loss or hyperglycemic crisis. In patients
without obvious clinical manifestations, at least two independent
test results were required to confirm the diagnosis of diabetes
(17). Exclusion criteria: i) Age
<18 years; ii) History of any preoperative anticancer therapy
(for example chemotherapy or immunotherapy); iii) Concurrent
intestinal infections, inflammatory bowel diseases or other
malignancies. A total of 50 patients with T2DM and 50 patients
without T2DM or prediabetes were recruited into the diabetes
mellitus (DM; mean age 62 range (42–74); men/women: 34/16) and the non-DM
(NDM; mean age 60; range (40–76);
men/women: 38/12) groups, respectively. Paraffin-embedded samples
of adjacent normal colonic epithelium tissue and related
clinicopathological data were collected. Ethical approval for the
present study was obtained from the Ethics Committee of Sun Yat-sen
Memorial Hospital (approval no. SYSKY-2024-1040-01).
Immunohistochemistry (IHC) and
evaluation
Tissue samples were fixed in 10% neutral buffered
formalin at room temperature for 24 h, dehydrated in graded
ethanol, embedded in paraffin. The paraffin-embedded tissue samples
were retrieved from storage in December 2024 and cut into
4-µm-thick sections. For staining, the paraffin sections were first
deparaffinized in xylene and then rehydrated through a graded
ethanol series (100, 95, 85 and 75%). Finally, they were rinsed in
PBS. Antigen retrieval was performed by immersing the sections in
citrate buffer (pH 6.0) at 95°C for 15 min. After cooling and
washing with PBS, endogenous peroxidase activity was quenched by
incubation in 3% hydrogen peroxide for 10 min at room temperature,
followed by blocking with 5% normal goat serum (cat no. G1208;
Wuhan Servicebio Technology Co., Ltd.) for 15 min at room
temperature. The sections were incubated overnight at 4°C with the
respective primary antibodies against proliferating cell nuclear
antigen (PCNA; 1:1,000; cat. no. ZM-0213; Beijing Zhongshan Jinqiao
Biotechnology Co., Ltd.), IGF2BP2 (1:200; cat. no. RM6660; Suzhou
Biodragon Immunology Technology Co., Ltd.) and MDK (1:200; cat. no.
BD-PT5177; Suzhou Biodragon Immunology Technology Co., Ltd.). After
washing with PBS, the sections were incubated at room temperature
for 1 h with the appropriate HRP-conjugated secondary antibody:
goat anti-rabbit (1:500; cat. no. BF03008; Suzhou Biodragon
Immunology Technology Co., Ltd.) or goat anti-mouse (1:500; cat.
no. BF03001; Suzhou Biodragon Immunology Technology Co., Ltd.).
Immunostaining signals were developed using the DAB chromogen kit
(cat. no. ZLI-9017; Beijing Zhongshan Jinqiao Biotechnology Co.,
Ltd.) followed by subsequent counterstaining with hematoxylin. The
stained sections were observed using a light microscope equipped
with a digital camera (Nikon TE2000-U; Nikon Corporation).
The IHC evaluation was independently assessed by two
blinded researchers (JL and QX). Any disagreement was resolved by
consulting a third researcher (JX). PCNA staining was scored
binarily as either positive or negative, with the percentage of
PCNA-positive cells calculated as the proliferative index. IGF2BP2
and MDK staining were assessed using the immunoreactive score (IRS)
according to a previous study (18). Staining intensity was classified
into three levels (1, slight; 2, moderate; 3, intense), while the
proportion of stained cells was scored as: 0, 0%; 1≤25%; 2, 26–50%;
3, 51–75%; and 4, 76–100%. The IRS was defined as the product of
intensity and proportion scores.
Cell culture and viability
The normal human colonic mucosal epithelial cell
line NCM460 was obtained from the Cell Bank of the Chinese Academy
of Sciences (Shanghai, China). All cell culture was performed in a
37°C incubator with a 5% CO2 atmosphere. Cells were
cultured in Dulbecco's Modified Eagle medium (DMEM; Gibco; Thermo
Fisher Scientific, Inc.) supplemented with 10% fetal bovine serum
(FBS, Gibco; Thermo Fisher Scientific, Inc.) for 24 h. For serum
starvation, cells were incubated in DMEM supplemented with 1% FBS
for 12 h. Cells were then incubated in complete growth medium (DMEM
with 10% FBS) for 24 h to recover. Subsequently, the medium was
then replaced with complete growth medium containing glucose
concentrations of 100, 250, 350, 450, 550 or 650 mg/dl (equivalent
to 5.6, 14, 19.6, 25.2, 30.8 and 36.4 mmol/l), respectively. The
100 mg/dl glucose group was designated as the control group. To
account for osmotic pressure, additional groups were prepared with
medium containing 100 mg/dl glucose supplemented with 50, 150, 250,
350, 450 and 550 mg/dl L-glucose. All groups were cultured for a
further 48 h, after which cell viability was assessed using Cell
Counting Kit-8 (CCK-8; APeXBIO Technology LLC) according to the
manufacturer's protocol. Following a 2-h incubation with the CCK-8
reagent, the absorbance at 450 nm was measured using a
multifunctional microplate reader (SpectraMax® M5;
Molecular Devices, LLC). Cell viability was calculated as: Cell
viability (%)=(Value test-Value blank)/(Value control-Value blank)
×100. The glucose concentration yielding the highest cell viability
was selected and defined as the high glucose (HG) group for
subsequent experiments.
Colony formation assay
NCM460 cells were seeded at a density of 1,000 cells
per well in 6-well plates and cultured in medium containing either
100 mg/dl glucose (control group) or the HG concentration as
aforementioned. After 7 days of culture, colonies were fixed with
4% polymethyl methacrylate (Beyotime Biotechnology) at room
temperature for 30 min and stained with 0.1% crystal violet
solution (Beyotime Biotechnology) at room temperature for 15 min.
Colonies containing >50 cells were counted manually to assess
proliferative capacity.
Cell transfection and inhibition of
MDK
The full-length coding sequence of MDK was subcloned
into the pcDNA3.1 vector to generate the overexpression plasmid
(plaMDK; GeneChem), with an empty pcDNA3.1 vector (plaNC) serving
as the corresponding control. The DNA sequence encoding the
short-hairpin RNA (shRNA) was cloned into the pLKO.1 lentiviral
vector (Shanghai GenePharma Co., Ltd.). The sequence for knockdown
via shRNA targeting IGF2BP2 (shIGF2BP2) was
5′-GTTGGCCCAGGGCGTTAAATT-3′; a scrambled shRNA (shNC) with the
sequence 5′-CAACAAGATGAAGAGCACCAA-3′ was used as a negative
control. Mutant MDK plasmid constructs were generated using
QuikChange Multi Site-Directed Mutagenesis Kit (cat. no. 200513;
Agilent Technologies, Inc.). To suppress MDK expression in NCM460
cells, a specific MDK inhibitor (iMDK) was applied (200 nM; cat.
no. 5126; R&D Systems Europe, Ltd.). An equal volume of vehicle
was used as negative control. For plasmid transfection, NCM460
cells were seeded into six-well plates at 3×105 cells
per well and transfected upon reaching 70–80% confluence. For each
well, 2.5 µg of plasmid DNA and 5 µl of Lipofectamine®
2000 reagent (Invitrogen; Thermo Fisher Scientific, Inc.) were
diluted separately in Opti-MEM (cat. no. 31985070; Gibco; Thermo
Fisher Scientific, Inc.), combined and incubated at room
temperature for 15 min to form complexes. The complexes were added
to the cells and incubated for 6 h. Subsequently, cells were
cultured for 48 h (37°C; 5% CO2) before subsequent
experiments. For lentivirus production, 293T cells (iCell
Bioscience) at ~70% confluence in 10-cm dishes were co-transfected
with a plasmid mixture containing 15 µg of either pLKO.1-shNC or
pLKO.1-shIGF2BP2 plasmids along with the packaging plasmids psPAX2
(11.25 µg) and pMD2.G (3.75 µg). Transfection was performed using
Lipofectamine® 2000 reagent (DNA:reagent=1 µg:2 µl) at
37°C in 5% CO2 for 6 h. Pooled viral supernatants
(collected at 48/72 h after transfection) were filtered (0.45 µm)
and stored at −80°C. For transduction, NCM460 cells were then
infected(at ~70% confluence) with viral supernatant at an MOI of
10, supplemented with 8 µg/ml polybrene (Sigma-Aldrich; Merck KGaA)
for 24 h at 37°C. Following selection with puromycin (2 µg/ml;
P8833, MilliporeSigma) starting at 48 h post-transduction and
lasting for 7 days. The cells then were harvest for the subsequent
experiments.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA of the NCM460 cells was extracted using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.) and treated with DNA-free™ DNase Treatment & Removal I
Kit (Thermo Fisher Scientific, Inc.) to remove any remaining DNA.
cDNA was synthesized with PrimeScript RT Master Mix (Takara Bio,
Inc.) according to manufacturer's instructions. qPCR was performed
using TB Green® Premix Ex Taq II kit (Takara Bio, Inc.)
on a Bio-Rad Real-Time PCR system (Bio-Rad Laboratories, Inc.).
Relative gene expression was calculated using the 2−ΔΔCq
method (19), with GAPDH serving
as the internal control. The primer sequences were as follows:
IGF2BP2, forward 5′-TGGAAGCGCATATCAGAGTG-3′, reverse
5′-AGTGCCCGATAATTCTGACG-3′; MDK forward 5′-AAGGAGTTTGGAGCCGACTG-3′,
reverse 5′-CATTGTAGCGCGCCTTCTTC-3′; and GAPDH forward
5′-GGAGCGAGATCCCTCCAAAAT-3′, reverse 5′-GGCTGTTGTCATACTTCTCATGG-3′.
The amplification protocol consisted of an initial denaturation at
95°C for 30 sec, followed by 40 cycles comprising denaturation at
95°C for 5 sec, annealing at 60°C for 30 sec and extension at 70°C
for 30 sec.
Western blot analysis
NCM460 cells were lysed in RIPA lysis buffer (cat.
no. P0013C; Beyotime Biotechnology) supplemented with a protease
and phosphatase inhibitor cocktail (cat. no. P1045; Beyotime
Biotechnology) on ice for 30 min. The cell lysates were then
centrifuged at 14,000 × g for 15 min at 4°C and the supernatants
were collected. Protein concentration was determined using the
Pierce® BCA Protein Assay Kit (cat. no. 23225; Thermo
Fisher Scientific, Inc.) following the manufacturer's instructions.
Protein samples (30 µg per lane) were separated by 7.5–12.5%
gradient SDS-PAGE gels, transferred to PVDF membranes (cat. no.
IPVH00010; Merck KGaA) and blocked with 5% skim milk in
Tris-buffered saline with 0.1% Tween 20 (TBST) for 1 h at room
temperature. The membranes were then incubated overnight at 4°C
with the following primary antibodies: anti-IGF2BP2 (1:1,000; cat.
no. RM6660; Suzhou Biodragon Immunology Technology Co., Ltd.),
anti-MDK (1:1,000; cat. no. BD-PT5177; Suzhou Biodragon Immunology
Technology Co., Ltd.) and anti-β-actin (1:1,000; cat. no. 4967;
Cell Signaling Technology, Inc.). After three washes with TBST,
membranes were incubated with HRP-conjugated anti-rabbit secondary
antibody (1:5,000; cat. no. 7074; Cell Signaling Technology, Inc.)
for 1 h at room temperature. Protein detection was carried out
using the SuperSignal™ West Pico PLUS Chemiluminescent Substrate
(cat. no. 34580; Thermo Fisher Scientific) and visualized with the
ChemiDoc Touch (Bio-Rad).
MDK mRNA stability analysis
To evaluate MDK mRNA stability, NCM460 cells were
treated with 4 µg/ml actinomycin D (cat. no. A9415; MilliporeSigma)
for 0, 4, 8 or 12 h at 37°C. Following treatment, total RNA was
extracted using TRIzol® reagent and MDK mRNA levels were
quantified by RT-qPCR, as aforementioned. Relative stability was
determined by normalizing expression levels at each time point to
those measured at 0 h.
RNA immunoprecipitation (RIP)
assay
RIP was conducted using the Magna RIP®
Kit (cat. no. 17–700; Merck KGaA) according to the manufacturer's
protocol. Briefly, NCM460 cells were lysed using RIP lysis buffer
supplemented with protease inhibitor cocktail and RNase inhibitor,
all from the kit. For each immunoprecipitation reaction, 100 µl
cell lysate was used. A total of 5 µg of IGF2BP2 antibody (cat. no.
ab128175; Abcam) or equal amount of normal IgG (negative control)
was pre-bound to 50 µl protein A/G magnetic beads. The
antibody-bound beads were then incubated with the cell lysates at
4°C overnight. After washing, the RNA-protein complexes were
digested with Proteinase K to release the RNA. The
immunoprecipitated RNA was analyzed by RT-qPCR, as aforementioned.
Relative fold enrichment was calculated as previously described
(20) using the following
equations: ΔCqRIP=CqRIP-Cqinput;
ΔCqIgG=CqIgG-Cqinput;
ΔΔCq=ΔCqRIP-ΔCqIgG; and fold
enrichment=2−ΔΔCq.
Statistical methods
All experiments were performed independently with ≥3
biological replicates. All statistical analyses were performed
using R software (version 4.3.3; National Institutes of Health).
The normality of all datasets was assessed using the Shapiro-Wilk
test. Categorical variables were analyzed using the χ2
test, with Fisher's exact test employed for cases where >20% of
expected cell counts were <5. Normally distributed continuous
variables were expressed as mean ± standard deviation. Comparisons
between two groups were performed using unpaired Student's t-test,
while multiple comparisons were assessed by one-way analysis of
variance followed by Tukey's honest significant difference test.
Ordinal and non-normally distributed continuous data were presented
as median and range and analyzed using the Mann-Whitney U test.
Pairwise correlations in gene expression were analyzed using
Pearson's correlation coefficient. P<0.05 was considered to
indicate a statistically significant difference.
Results
Increased proliferation of colonic
epithelium under diabetic conditions
The clinicopathological characteristics of patients
with colonic carcinoma were summarized in Table I (21). No significant differences were
observed between the DM and NDM groups with respect to sex, age,
body mass index, smoking or alcohol history, tumor stage, tumor
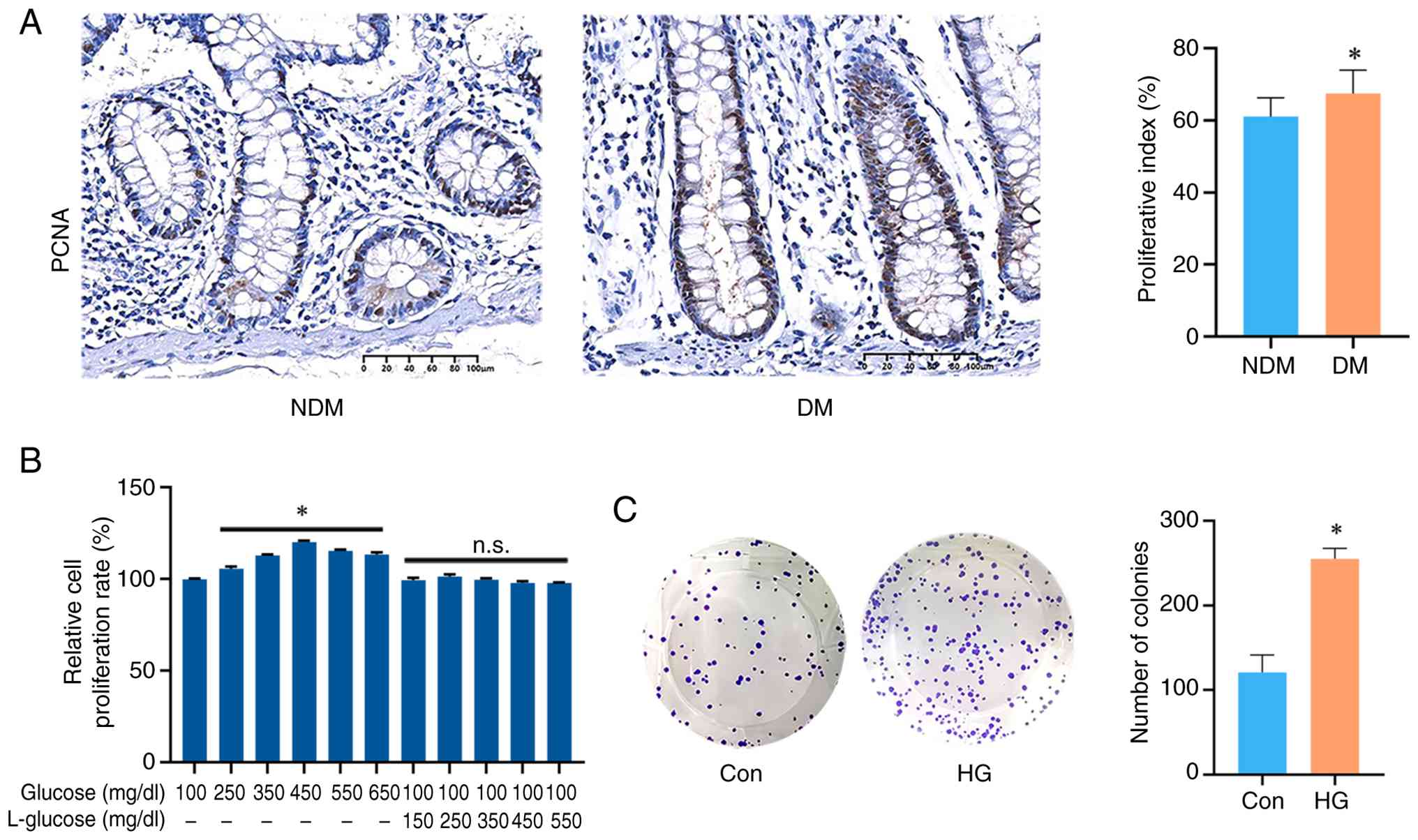
site or tumor differentiation. IHC staining of PCNA revealed that
the proliferative index was significantly higher in the DM group
compared with the NDM group (NDM, 61.05±5.22%; DM, 67.50±6.44%;
P<0.01; Fig. 1A). Previous
studies have shown that hyperglycemia induces epigenetic
reprogramming and mediates m6A mRNA modification,
thereby serving as a key pathway linking diabetes to cancer
(22,23). Consistently, hyperglycemia
contributed to the proliferative effects of T2DM on colonic
epithelium. To mimic diabetic conditions in vitro, NCM460
cells were cultured in media containing increasing concentrations
of glucose. CCK-8 assays demonstrated a significant glucose
concentration-dependent increase in cell viability. The highest
viability was observed at 450 mg/dl glucose, with a slight
reduction at 550 and 650 mg/dl. Osmotic pressure control groups
provided evidence that osmotic effects had no significant impact on
proliferation (Fig. 1B). Based on
these findings, 450 mg/dl glucose was selected as the HG condition
and 100 mg/dl glucose was used as the control condition. Colony
formation assays demonstrated a significant increase in the number
of colonies in the HG group compared with the control group
(Fig. 1C).
 | Table I.Clinicopathologic characteristics of
the DM and NDM groups. |
Table I.
Clinicopathologic characteristics of
the DM and NDM groups.
| Category | DM group No. of
patients | % | NDM group No. of
patients | % | P-value |
|---|
| Total patients | 50 |
| 50 |
|
|
| Age, median
(range), years | 64 (42–74) |
| 62 (40–76) |
| 0.366 |
| Body mass index,
median (range), kg/m2 | 23.198
(18.026–34.949) |
| 23.711
(17.715–29.936) |
| 0.620 |
| Sex |
|
|
|
| 0.373a |
|
Male | 34 | 68 | 38 | 76 |
|
|
Female | 16 | 32 | 12 | 24 |
|
| Smoker |
|
|
|
|
>0.999b |
|
Current | 13 | 26 | 12 | 24 |
|
|
Former | 3 | 6 | 3 | 6 |
|
| Alcohol use |
|
|
|
| 0.163b |
|
Current | 10 | 20 | 4 | 8 |
|
|
Former | 2 | 4 | 1 | 2 |
|
| CC stage |
|
|
|
| 0.598a |
| I | 5 | 10 | 6 | 12 |
|
| II | 16 | 32 | 21 | 42 |
|
|
III | 19 | 38 | 17 | 34 |
|
| IV | 10 | 20 | 6 | 12 |
|
| CC site |
|
|
|
| 0.420a |
|
Left | 30 | 60 | 26 | 52 |
|
|
Right | 20 | 40 | 24 | 48 |
|
| CC
differentiation |
|
|
|
| 0.824a |
|
Good | 5 | 10 | 7 | 14 |
|
|
Moderate | 39 | 78 | 37 | 74 |
|
|
Poor | 6 | 12 | 6 | 12 |
|
| Metformin use |
|
|
|
|
|
|
Yes | 23 | 46 |
|
|
|
| No | 27 | 54 |
|
|
|
IGF2BP2 knockdown decreases
proliferation of colonic epithelium under diabetic conditions
To investigate whether m6A modification
contributed to the enhanced proliferation of colonic epithelium
under T2DM conditions, sequence data from the GSE115313 dataset
detailing the colonic epithelium of 23 patients with T2DM and 19
patients without T2DM or prediabetes were re-analyzed. Among the
m6A-related enzymes analyzed, IGF2BP2 expression was
significantly upregulated in colonic epithelium from patients with
T2DM (P<0.01; Fig 2A; Table SI). This finding was validated by
IHC, which demonstrated significantly higher IGF2BP2 expression in
the DM group compared with the NDM group (NDM, 4 (range, 2–9); DM,
8 (range, 4–12); P<0.01; Fig.
2B). Consistently, IGF2BP2 expression was significantly
elevated in NCM460 cells cultured under HG conditions compared with
the control group (Fig. 2C). To
knock down IGF2BP2, a specific shRNA was employed, with its
efficiency validated in NCM460 cells under baseline conditions
(Fig. 2D). Functionally, IGF2BP2
knockdown under HG conditions significantly reduced cell viability
compared with the negative control group (Fig. 2E).
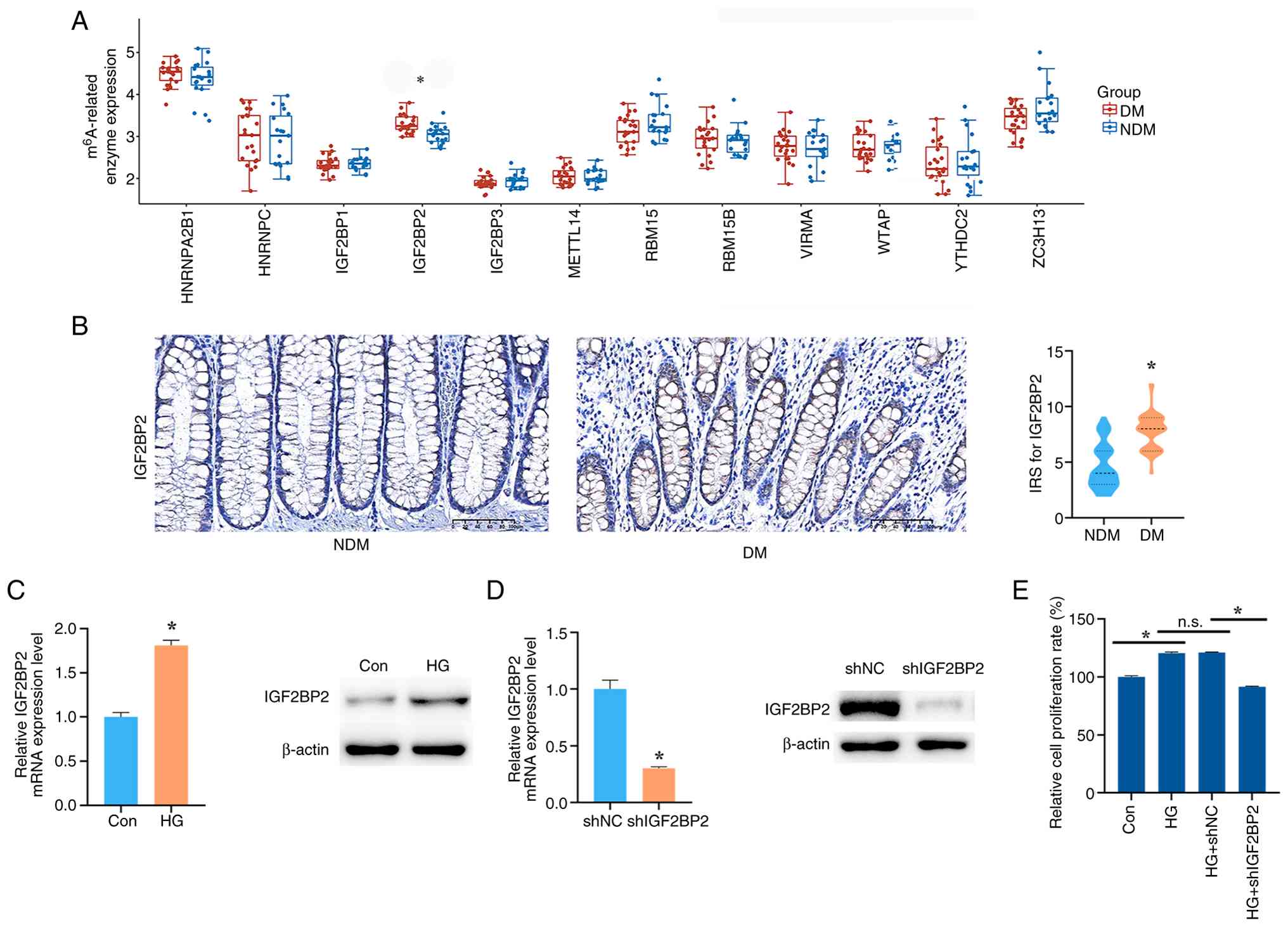 | Figure 2.IGF2BP2 knockdown decreases
proliferation of colonic epithelium under diabetic conditions. (A)
Expression of m6A-related enzymes in colonic epithelium
of DM and NDM patients based on RNA sequencing data (GSE115313;
*P<0.01). (B) Representative immunohistochemical staining and
IRS analysis of IGF2BP2 expression in colonic epithelium from NDM
and DM patients (scale bar, 100 µm; *P<0.01). (C) IGF2BP2
expression in NCM460 cells under control and HG conditions
(*P<0.01). (D) IGF2BP2 knockdown efficiency in NCM460 cells
under baseline conditions by short hairpin RNA (*P<0.01). (E)
The effect of IGF2BP2 knockdown on viability of NCM460 cells under
HG conditions measured by Cell Counting Kit-8 assay (*P<0.01).
DM, diabetes mellitus; NDM, non-diabetes mellitus; n.s., not
significant; IRS, immunoreactive score; Con, control group; HG,
high glucose; IGF2BP2, insulin-like growth factor 2 mRNA binding
protein 2; shNC, short hairpin RNA negative control; shIGF2BP2,
short hairpin RNA targeting IGF2BP2; HNRNPA2B1, heterogeneous
nuclear ribonucleoproteins A2/B1; HNRNPC, heterogeneous nuclear
ribonucleoproteins C1/C2; IGF2BP1, insulin-like growth factor 2
mRNA-binding protein 1; IGF2BP3, insulin-like growth factor 2
mRNA-binding protein 3; METTL14, methyltransferase-like protein 14;
RBM15, RNA-binding protein 15; RBM15B, putative RNA-binding protein
15B; VIRMA, protein virilizer homolog; WTAP, pre-mRNA-splicing
regulator WTAP; YTHDC2, 3′-5′ RNA helicase YTHDC2; ZC3H13, zinc
finger CCCH domain-containing protein 13. |
MDK knockdown decreases the
proliferation of colonic epithelial cells under diabetic
conditions
Sequence data from dataset GSE115313 were analyzed
to identify genes associated with the enhanced proliferation of
colonic epithelial cells promoted by IGF2BP2. As an m6A
reader, IGF2BP2 stabilizes target mRNAs (24), and its expression was elevated in
colonic epithelium under T2DM conditions (Fig. 2C). Genes significantly upregulated
(P<0.05; fold change >1.2) in patients with T2DM were
therefore selected as candidates for further analysis. Among these,
27 genes were identified as showing positive correlated with
IGF2BP2 expression in normal colonic epithelium based on GEPIA
database. Additionally, 30 genes were predicted to interact with
IGF2BP2 as RNA-binding targets in the StarBase database (Fig. 3A; Table SII). A total of 14 genes
overlapped between the two datasets (Fig. 3A; Table SIII). Among these, MDK, which
demonstrated a positive correlation with IGF2BP2 (R=0.41;
P=8.9×10−16; Fig. 3B and
C), was notable as it has been implicated in colonic carcinoma
progression (25). IHC performed
in the patient cohort showed significantly higher MDK expression
the DM group compared with the NDM group [NDM, 6 (range, 3–9); DM,
8.5 (range, 6–12); P<0.01; Fig.
3D]. In vitro, MDK mRNA expression was significantly
elevated in NCM460 cells under HG conditions relative to the
control group, which was reflected in MDK protein expression that
also showed a marked increase (Fig.
3E). The functional role of MDK was assessed using its
inhibitor iMDK. Once its efficacy was confirmed under baseline
settings (Fig. 3F), iMDK was
applied to NCM460 cells under HG conditions, resulting a
significant reduction in both colony formation and cell viability
(Fig. 3G and H).
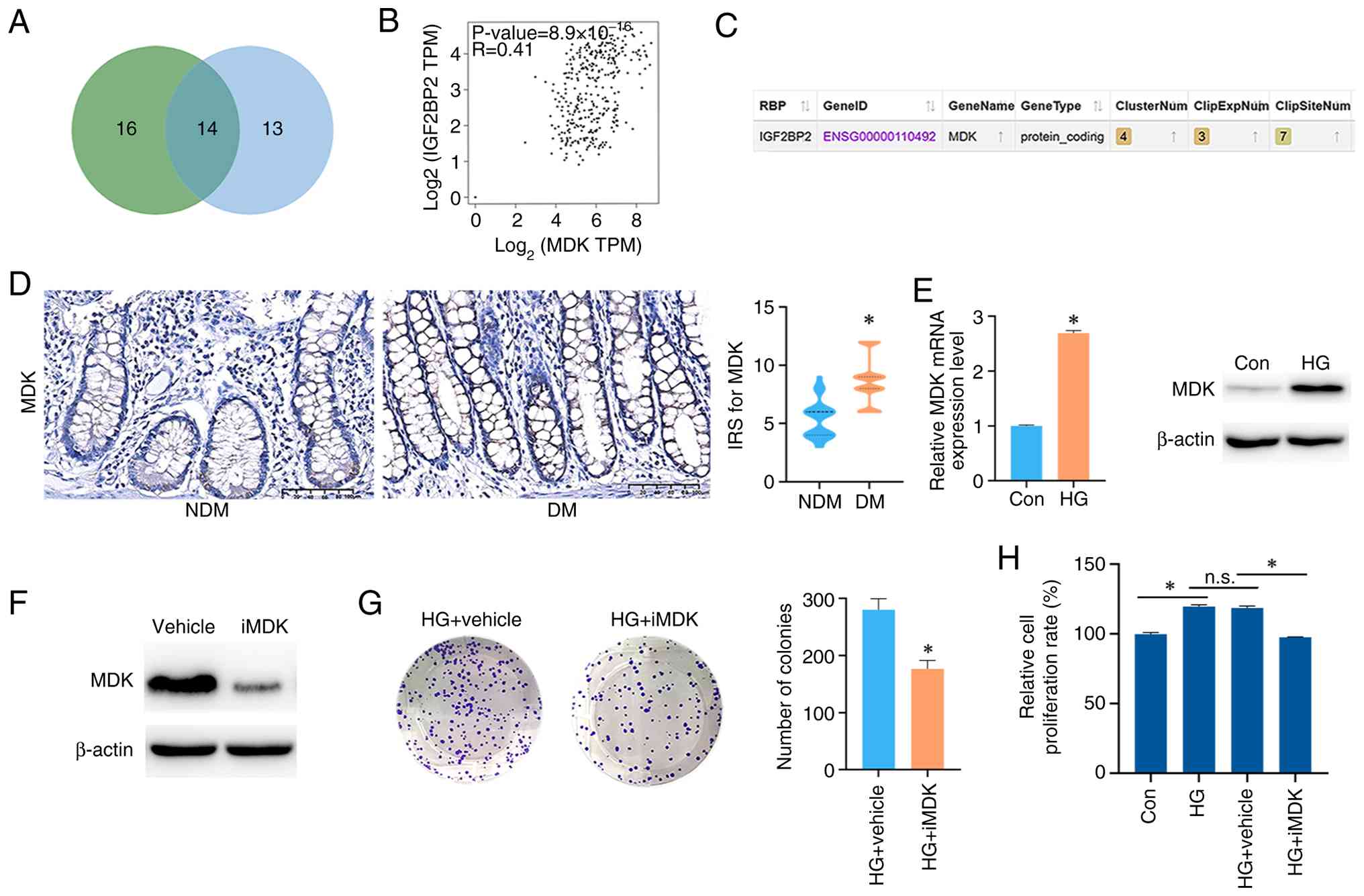 | Figure 3.MDK knockdown decreases proliferation
of colonic epithelium under diabetic conditions. (A) Venn diagram
of candidate genes, where green represents genes predicted to
interact with IGF2BP2 and blue represents genes positively
correlated with IGF2BP2 expression. (B) Correlation between MDK and
IGF2BP2 expressions in colonic epithelium analyzed by the Gene
Expression Profiling Interactive Analysis database. (C) Predicted
interaction between MDK and IGF2BP2 using the StarBase database.
(D) Representative immunohistochemical staining and IRS analysis of
MDK expression in colonic epithelium from patients in the NDM and
DM groups (scale bar, 100 µm; *P<0.01). (E) MDK expressions in
NCM460 cells under Con and HG conditions (*P<0.01). (F)
Validation of MDK inhibition by iMDK under baseline conditions. (G
and H) Effects of iMDK on proliferation and viability of NCM460
cells under HG conditions, assessed by (G) colony formation and (H)
Cell Counting Kit-8 assay (*P<0.01). MDK, midkine; iMDK,
inhibitor of MDK; n.s., not significant; DM, diabetes mellitus;
NDM, non-diabetes mellitus; Con, control group; HG, high glucose;
IRS, immunoreactive score; IGF2BP2, insulin-like growth factor 2
mRNA binding protein 2. |
IGF2BP2 promotes proliferation of
colonic epithelial cells by targeting MDK in an
m6A-dependent manner
The interaction between IGF2BP2 and MDK was
subsequently examined. IGF2BP2 knockdown under HG conditions
significantly reduced MDK expression (Fig. 4A). To test if MDK downregulation
mediated the effects of IGF2BP2 loss, a rescue experiment was
performed. MDK was overexpressed via plasmid transfection and the
efficiency was confirmed under baseline conditions in NCM460 cells
(Fig. 4B). MDK overexpression
partially rescued the suppression of proliferation induced by
IGF2BP2 knockdown under HG conditions, as demonstrated by CCK-8
(Fig. 4C) and colony formation
assays (Fig. 4D). Mechanically,
the effect of IGF2BP2 on MDK transcript stability was further
investigated and it was revealed that IGF2BP2 knockdown markedly
reduced MDK mRNA stability at 4, 8 and 12 h (Fig. 4E). To predict potential
m6A sites in MDK, the transcript sequence was analyzed
using the prediction tool, SRAMP, which identified one site with
very high confidence and five sites with high confidence (Fig. 4F). The top three predicted sites
were mutated, and RIP enrichment analysis demonstrated that
wild-type MDK mRNA was significantly enriched by IGF2BP2 compared
with the IgG control, whereas enrichment was significantly lower in
the mutant group than the wild-type group (Fig. 4G).
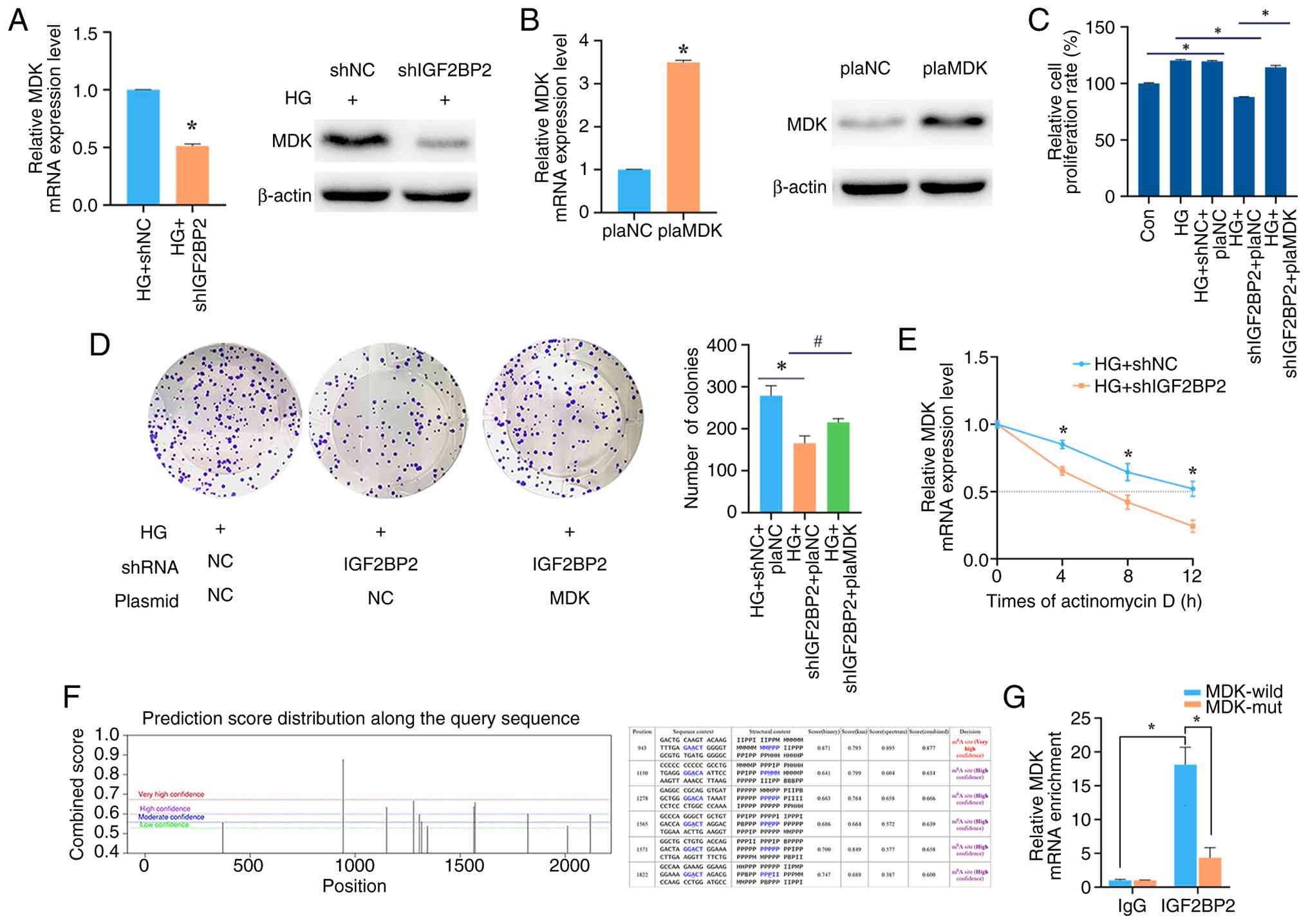 | Figure 4.IGF2BP2 promotes proliferation of
colonic epithelial cells by targeting MDK in an
m6A-dependent manner. (A) MDK mRNA and protein
expression following IGF2BP2 knockdown under HG conditions
(*P<0.01). (B) Validation of MDK overexpression by plasmid
transfection under baseline conditions (*P<0.01). (C and D)
Effects of IGF2BP2 knockdown and MDK rescue on viability and
proliferation of NCM460 cells under HG conditions, assessed by (C)
Cell Counting Kit-8 and (D) colony formation assay (*P<0.01;
#P<0.05). (E) MDK mRNA stability measured by reverse
transcription-quantitative PCR following IGF2BP2 knockdown under HG
conditions (*P<0.01). (F) Predicted m6A sites in MDK
using the sequence-based RNA adenosine methylation site predictor
prediction tool. (G) RNA immunoprecipitation assay showing
enrichment of wild-type vs. mutant MDK mRNA by anti-IGF2BP2
antibody (*P<0.01). IGF2BP2, insulin-like growth factor 2 mRNA
binding protein 2; NC, negative control; shRNA, short hairpin RNA;
shNC, short hairpin RNA negative control; shIGF2BP2, short hairpin
RNA targeting IGF2BP2; MDK, midkine; HG, high glucose; plaNC,
negative control overexpression plasmid; plaMDK, MDK overexpression
plasmid; MDK-wild, wild-type MDK mRNA; MDK-mut, mutant MDK
mRNA. |
Discussion
T2DM increases the risk of colonic carcinoma but
exerts limited effects on tumor behavior once carcinoma has
developed (26,27). This observation suggests that the
driving influence of T2DM may occur primarily at the early stage of
tumorigenesis (28). As the
majority of cases of colonic carcinoma originate from aberrant
crypt foci (29), pathological
changes in the colonic epithelium of patients with T2DM may
contribute to malignant transformation. However, few studies have
directly examined epithelial proliferation in this context. In the
present study, IHC staining of PCNA demonstrated that epithelial
proliferation in colonic tissue from patients with T2DM was
significantly higher compared with that from non-diabetic or
prediabetic patients, consistent with previous findings (30). Hyperglycemia notably links T2DM to
colonic carcinogenesis by acting as a key oncogenic driver that
disrupts epigenetic stability and dysregulates core signaling
pathways (22,31). Based on this rationale, the present
study mimicked diabetic conditions in vitro by culturing
NCM460 cells in media containing increasing concentrations of
glucose and the maximal cell viability was observed at 450 mg/dl
glucose, further supported by colony formation assay. This
concentration was therefore applied as the HG condition in
subsequent experiments to mimic the diabetic microenvironment.
m6A modification modulates RNA expression
by influencing stability, translation, selective splicing,
processing and maturation, thereby regulating cellular signaling
and contributing to the modulation of cell proliferation (32). However, whether the expression of
m6A-related enzymes is altered in the colonic epithelium
of patients with T2DM has remained to be fully elucidated.
Re-analysis of published RNA sequencing data revealed that IGF2BP2
expression was significantly elevated in patients with T2DM
(13). This finding was further
validated in a larger clinical cohort and in the human colonic
epithelial NCM460 cell line cultured under T2DM-mimicking
conditions. As an m6A reader, IGF2BP2 enhances the
expression of target genes, thereby promoting cell proliferation
and cancer progression (33–36).
Furthermore, IGF2BP2 has been identified as a T2DM-associated gene
(36). Nevertheless, to the best
of our knowledge, its role in colonic epithelial hyperproliferation
under T2DM conditions has not been previously defined. In the
present study, IGF2BP2 knockdown significantly suppressed
epithelial proliferation, indicating its functional role in
tumorigenesis.
Subsequently, the downstream genes involved in
IGF2BP2-mediated regulation of aberrant colonic epithelial
proliferation under T2DM conditions were investigated. IGF2BP2 has
been reported to stabilize target mRNAs post-translationally
(37), and its expression was
shown to be elevated in the present study. RNA sequencing data of
colonic epithelium from a previous study were reviewed (13), and genes upregulated in colonic
epithelium from patients with T2DM were selected for further
analysis. Candidate genes were evaluated for positive correlation
with IGF2BP2 expression using GEPIA database, and potential
interactions with IGF2BP2 were predicted using the StarBase
database. A total of 14 genes met both criteria, among which MDK
was notable. MDK, a cytokine and growth factor, modulates antitumor
immunity and the cell cycle and exerts tumor-promoting effects in
several malignancies (38–40). In colonic cancer, high MDK
expression has been associated with poor overall survival and
immune tolerance, thereby facilitating carcinogenesis (25,41).
However, to the best of our knowledge, its role in T2DM-associated
epithelial proliferation has not been established. In the present
study, MDK expression was significantly increased in colonic
epithelium from patients with T2DM and in NCM460 cells cultured
under HG conditions. Inhibition of MDK using iMDK under HG
conditions significantly reduced both cell viability and colony
formation in NCM460 cells. Collectively, these findings indicated
that MDK expression was upregulated and contributed to the enhanced
proliferation of colonic epithelium under T2DM conditions.
The interaction between IGF2BP2 and MDK was further
investigated. Knockdown of IGF2BP2 led to decreased MDK expression,
consistent with the aforementioned predictions that MDK and IGF2BP2
were functionally linked. Conversely, overexpression of MDK
attenuated the proliferation-suppressive effect of IGF2BP2
knockdown under HG conditions, indicating that IGF2BP2 regulated
proliferation through MDK. As an m6A reader, IGF2BP2 has
been shown to stabilize target mRNAs (37). MDK mRNA stability analysis showed
that IGF2BP2 knockdown significantly reduced MDK transcript
stability. Bioinformatic predictions identified six putative
m6A sites in MDK with high or very high confidence. RIP
assays demonstrated the specific binding of IGF2BP2 to MDK mRNA,
and this interaction was significantly impaired by mutagenesis of a
subset of m6A sites. Collectively, these results
demonstrated that IGF2BP2 promoted colonic epithelial proliferation
by targeting MDK in an m6A-dependent manner.
Several limitations should be acknowledged in the
present study. Primarily, the sample size for IHC detection was
initially determined based on preliminary experimental results
(data not shown), yielding a final cohort of 50 patients per group.
Although this sample size was sufficient for the current findings,
it remains inadequate for more detailed investigations. Limited
research is available regarding factors that contribute to IGF2BP2
and MDK upregulation in colonic epithelium. Given that abnormal
epithelial cell proliferation is closely related to colon
carcinogenesis, together with the fact that T2DM and colonic
carcinoma share several risk factors, such as tobacco use, alcohol
consumption, excessive body weight and physical inactivity
(42,43), future studies with larger cohorts
are warranted to more effectively evaluate the impact of these
variables on T2DM and colonic carcinoma and to control for
potential confounding factors. Furthermore, in light of the
documented effects of hypoglycemic medications, such as
glucagon-like peptide-1 receptor agonists, metformin and insulin,
on colonic epithelium (44–46),
it would be valuable to perform subgroup analyses assessing the
specific influences of these drugs. Secondly, although the present
study established that MDK promoted the abnormal proliferation of
colonic epithelial cells under T2DM conditions, the specific
downstream mechanisms involved remained undefined. Previous
research indicates that in malignancies, MDK interacts with several
proliferation-related pathways, including the Wnt, MAPK and
PI3K/AKT pathways (47). To
further elucidate the mechanism of MDK action, future research
could establish MDK-deficient T2DM models and apply diverse omics
technologies, such as transcriptomics and proteomics, complemented
by experimental validation to identify the key downstream target
genes and functional processes involved.
In summary, the present study provided evidence that
colonic epithelial proliferation was abnormally increased under
T2DM conditions, both clinically and in vitro. The
upregulation of IGF2BP2 was identified as a key driver of this
effect, acting by binding to and stabilizing MDK mRNA in an
m6A-dependent manner. These findings suggested that
IGF2BP2 and MDK may represent potential therapeutic targets for
controlling abnormal epithelial proliferation in T2DM, thereby
reducing the risk of colonic carcinoma in this patient
population.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by the Natural Science
Foundation of Guangdong Province, China (grant no.
2023A1515010311).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
JL contributed to the conceptualization and design
of the study, drafted the initial manuscript, performed the in
vitro experiments, coordinated revisions with all co-authors,
provided overall supervision and guidance throughout the project
and secured funding. QX participated in performing in vitro
experiments, data collection, preliminary data analysis and
manuscript preparation. JX performed bioinformatics analyses and
collected and analyzed the clinical data. JL and QX confirm the
authenticity of all the raw data. All authors read and approved the
final version of the manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Sun Yat-sen Memorial Hospital (approval no.
SYSKY-2024-1040-01) with a waiver of informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
NCD Risk Factor Collaboration (NCD-RisC),
. Worldwide trends in diabetes prevalence and treatment from 1990
to 2022: A pooled analysis of 1108 population-representative
studies with 141 million participants. Lancet. 404:2077–2093. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Zhang YY, Li YJ, Xue CD, Li S, Gao ZN and
Qin KR: Effects of T2DM on cancer progression: pivotal
precipitating factors and underlying mechanisms. Front Endocrinol
(Lausanne). 15:13960222024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Vekic J, Zeljkovic A, Stefanovic A, Giglio
RV, Ciaccio M and Rizzo M: Diabetes and colorectal cancer risk: A
new look at molecular mechanisms and potential role of novel
antidiabetic agents. Int J Mol Sci. 22:124092021. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Gharib E and Robichaud GA: From crypts to
cancer: A holistic perspective on colorectal carcinogenesis and
therapeutic strategies. Int J Mol Sci. 25:94632024. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wang SY, Li JY, Xu JH, Xia ZS, Cheng D,
Zhong W, Lai Y, Yu T and Chen QK: Butyrate suppresses abnormal
proliferation in colonic epithelial cells under diabetic state by
targeting HMGB1. J Pharmacol Sci. 139:266–274. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Jiang X, Liu B, Nie Z, Duan L, Xiong Q,
Jin Z, Yang C and Chen Y: The role of m6A modification in the
biological functions and diseases. Signal Transduct Target Ther.
6:742021. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yang Y, Hsu PJ, Chen YS and Yang YG:
Dynamic transcriptomic m6A decoration: Writers, erasers,
readers and functions in RNA metabolism. Cell Res. 28:616–624.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Chen T, Ye W, Gao S, Li Y, Luan J, Lv X
and Wang S: Emerging importance of m6A modification in liver cancer
and its potential therapeutic role. Biochim Biophys Acta Rev
Cancer. 1880:1892992025. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Liu S, Liu M, Li Y and Song Q:
N6-methyladenosine-dependent signaling in colorectal cancer:
Functions and clinical potential. Crit Rev Oncol Hematol.
198:1043602024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Cheng C, Yu F, Yuan G and Jia J: Update on
N6-methyladenosine methylation in obesity-related diseases. Obesity
(Silver Spring). 32:240–251. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Luo J, Xu T and Sun K: N6-Methyladenosine
RNA modification in inflammation: Roles, mechanisms, and
applications. Front Cell Dev Biol. 9:6707112021. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ren Y, Li Z, Li J, Liang R, Wang Z, Bai Y,
Yang Y, Tang Q, Fu Y, Zhang X, et al: m6 A mRNA
methylation: Biological features, mechanisms, and therapeutic
potentials in type 2 diabetes mellitus. Obes Rev. 24:e136392023.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Del Puerto-Nevado L, Minguez P, Corton M,
Solanes-Casado S, Prieto I, Mas S, Sanz AB, Gonzalez-Alonso P,
Villaverde C, Portal-Nuñez S, et al: Molecular evidence of field
cancerization initiated by diabetes in colon cancer patients. Mol
Oncol. 13:857–872. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res. 45:W98–W102.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Li JH, Liu S, Zhou H, Qu LH and Yang JH:
starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA
interaction networks from large-scale CLIP-Seq data. Nucleic Acids
Res. 42:D92–D97. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Fan R, Cui C, Kang B, Chang Z, Wang G and
Cui Q: A combined deep learning framework for mammalian m6A site
prediction. Cell Genom. 4:1006972024. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
American Diabetes Association Professional
Practice Committee for Diabetes, . 2. diagnosis and classification
of diabetes: Standards of care in diabetes-2026. Diabetes Care.
49:S27–S49. 2026. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Koo CL, Kok LF, Lee MY, Wu TS, Cheng YW,
Hsu JD, Ruan A, Chao KC and Han CP: Scoring mechanisms of p16INK4a
immunohistochemistry based on either independent nucleic stain or
mixed cytoplasmic with nucleic expression can significantly signal
to distinguish between endocervical and endometrial adenocarcinomas
in a tissue microarray study. J Transl Med. 7:252009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhang Z, Xie Z, Lin J, Sun Z, Li Z, Yu W,
Zeng Y, Ye G, Li J, Ye F, et al: The m6A methyltransferase METTL16
negatively regulates MCP1 expression in mesenchymal stem cells
during monocyte recruitment. JCI Insight. 8:e1624362023. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Weiser MR: AJCC 8th edition: Colorectal
cancer. Ann Surg Oncol. 25:1454–1455. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Li H, Li C, Zhang B and Jiang H:
Lactoferrin suppresses the progression of colon cancer under
hyperglycemia by targeting WTAP/m6A/NT5DC3/HKDC1 axis. J
Transl Med. 21:1562023. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wu D, Hu D, Chen H, Shi G, Fetahu IS, Wu
F, Rabidou K, Fang R, Tan L, Xu S, et al: Glucose-regulated
phosphorylation of TET2 by AMPK reveals a pathway linking diabetes
to cancer. Nature. 559:637–641. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu S, Liao S, He J, Zhou Y and He Q:
IGF2BP2: An m6A reader that affects cellular function
and disease progression. Cell Mol Biol Lett. 30:432025. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kemper M, Hentschel W, Grass JK, Stüben
BO, Konczalla L, Rawnaq T, Ghadban T, Izbicki JR and Reeh M: Serum
Midkine is a clinical significant biomarker for colorectal cancer
and associated with poor survival. Cancer Med. 9:2010–2018. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Albai O, Frandes M, Timar B, Paun DL,
Roman D and Timar R: Long-term risk of malignant neoplastic
disorders in type 2 diabetes mellitus patients with metabolic
syndrome. Diabetes Metab Syndr Obes. 13:1317–1326. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Prieto I, Del Puerto-Nevado L, Gonzalez N,
Portal-Nuñez S, Zazo S, Corton M, Minguez P, Gomez-Guerrero C, Arce
JM, Sanz AB, et al: Colon cancer modulation by a diabetic
environment: A single institutional experience. PLoS One.
12:e01723002017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Del Puerto-Nevado L, Santiago-Hernandez A,
Solanes-Casado S, Gonzalez N, Ricote M, Corton M, Prieto I, Mas S,
Sanz AB, Aguilera O, et al: Diabetes-mediated promotion of colon
mucosa carcinogenesis is associated with mitochondrial dysfunction.
Mol Oncol. 13:1887–1897. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chen C, Fu Q, Wang L, Tanaka S and Imajo
M: Establishment of a novel mouse model of colorectal cancer by
orthotopic transplantation. BMC Cancer. 25:4052025. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li JY, Yu T, Xia ZS, Chen GC, Yuan YH,
Zhong W, Zhao LN and Chen QK: Enhanced proliferation in colorectal
epithelium of patients with type 2 diabetes correlates with
beta-catenin accumulation. J Diabetes Complications. 28:689–697.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Vasconcelos-Dos-Santos A, Loponte HF,
Mantuano NR, Oliveira IA, de Paula IF, Teixeira LK,
de-Freitas-Junior JC, Gondim KC, Heise N, Mohana-Borges R, et al:
Hyperglycemia exacerbates colon cancer malignancy through
hexosamine biosynthetic pathway. Oncogenesis. 6:e3062017.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wei B, Zeng M, Yang J, Li S, Zhang J, Ding
N and Jiang Z: N6-Methyladenosine RNA modification: A
potential regulator of stem cell proliferation and differentiation.
Front Cell Dev Biol. 10:8352052022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Shi R, Zhao R, Shen Y, Wei S, Zhang T,
Zhang J, Shu W, Cheng S, Teng H and Wang H: IGF2BP2-modified
circular RNA circCHD7 promotes endometrial cancer progression via
stabilizing PDGFRB and activating JAK/STAT signaling pathway.
Cancer Gene Ther. 31:1221–1236. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Yi J, Peng F, Zhao J and Gong X:
METTL3/IGF2BP2 axis affects the progression of colorectal cancer by
regulating m6A modification of STAG3. Sci Rep. 13:172922023.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Kessler SM, Laggai S, Barghash A,
Schultheiss CS, Lederer E, Artl M, Helms V, Haybaeck J and Kiemer
AK: IMP2/p62 induces genomic instability and an aggressive
hepatocellular carcinoma phenotype. Cell Death Dis. 6:e18942015.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wang J, Chen L and Qiang P: The role of
IGF2BP2, an m6A reader gene, in human metabolic diseases and
cancers. Cancer Cell Int. 21:992021. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Liu S, Liao S, He J, Zhou Y and He Q:
IGF2BP2: An m6A reader that affects cellular function
and disease progression. Cell Mol Biol Lett. 30:432025. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Aller EJ, Nair HB, Vadlamudi RK and
Viswanadhapalli S: Significance of midkine signaling in women's
cancers: Novel biomarker and therapeutic target. Int J Mol Sci.
26:48092025. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Yuan F, Wang Y, Yuan L, Tang T, Ye L, Li
Y, Dai X and Cheng H: EGFRvIII-positive glioblastoma contributes to
immune escape and malignant progression via the c-Fos-MDK-LRP1
axis. Cell Death Dis. 16:4532025. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Munter D, de Faria FW, Richter M,
Aranda-Pardos I, Hotfilder M, Walter C, Paga E, Inserte C, Albert
TK, Roy R, et al: Multiomic analysis uncovers a continuous spectrum
of differentiation and Wnt-MDK-driven immune evasion in
hepatoblastoma. J Hepatol. 83:367–382. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Hashimoto M, Kojima Y, Sakamoto T, Ozato
Y, Nakano Y, Abe T, Hosoda K, Saito H, Higuchi S, Hisamatsu Y, et
al: Spatial and single-cell colocalisation analysis reveals
MDK-mediated immunosuppressive environment with regulatory T cells
in colorectal carcinogenesis. EBioMedicine. 103:1051022024.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zou D, Xin X, Xu Y, Xu H and Xu T: A
cross-sectional study on the association between physical activity
and the risk of colon cancer based on NHANES 2007–2018. Sci Rep.
15:32972025. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Cao J, Yan W, Ma X, Huang H and Yan H:
Insulin-like growth factor 2 mRNA-binding protein 2-a potential
link between type 2 diabetes mellitus and cancer. J Clin Endocrinol
Metab. 106:2807–2818. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Sun H, Shu J, Tang J, Li Y, Qiu J, Ding Z,
Xuan B, Chen M, Gan C, Lin J, et al: GLP-1 receptor agonists
alleviate colonic inflammation by modulating intestinal microbiota
and the function of group 3 innate lymphoid cells. Immunology.
172:451–468. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Wang R, Xie L, Jiang P, Hou Y, Li D and
Wang W: Metformin may improve intestinal mucosal barrier function
and help prevent and reverse colorectal cancer in mice. J Cancer.
16:3703–3711. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Juarez-Vazquez CI, Gurrola-Diaz CM,
Vargas-Guerrero B, Domínguez-Rosales JA, Rodriguez-Ortiz JF,
Barros-Núñez P, Flores-Martínez SE, Sánchez-Corona J and
Rosales-Reynoso MA: Insulin glargine affects the expression of
Igf-1r, Insr, and Igf-1 genes in colon and liver of diabetic rats.
Iran J Basic Med Sci. 21:489–494. 2018.PubMed/NCBI
|
|
47
|
Yildirim B, Kulak K and Bilir A: Midkine:
A cancer biomarker candidate and innovative therapeutic approaches.
Eur J Breast Health. 20:167–177. 2024. View Article : Google Scholar : PubMed/NCBI
|