Introduction
Lung cancer represents one of the most prevalent
malignancies worldwide, with an annual incidence of ~1.2 million
new diagnoses (1). Histologically,
it is divided into two principal categories: Small cell lung cancer
(SCLC) and non-small cell lung cancer (NSCLC). NSCLC constitutes
~80% of all lung cancer cases and can be further subdivided into
several subtypes: Lung adenocarcinoma (LUAD), squamous cell
carcinoma and large cell carcinoma. In the early stages, NSCLC is
often asymptomatic, leading to delayed detection, allowing the
cancer to progress and thus high recurrence rates when treated.
Although therapeutic strategies, including radiotherapy,
chemotherapy and epidermal growth factor receptor (EGFR)-targeted
agents, have improved, the 5-year survival rate for patients with
NSCLC remains poor, ranging from 4–17% (2). While several studies have focused on
protein-coding genes, the molecular pathways driving lung cancer
pathogenesis and metastasis remain incompletely elucidated. Thus,
continued exploration of molecular alterations is essential to
uncover novel biomarkers and advance therapeutic interventions for
this disease.
Long non-coding RNAs (lncRNAs) are RNA molecules of
>200 nucleotides in length (3).
Due to the absence of a complete open reading frame, lncRNAs have
traditionally been considered incapable of encoding proteins
(4). Accumulating evidence now
indicates that a number of transcripts previously annotated as
lncRNAs contain small open reading frames and encode functional
micropeptides (5). During
transcription, lncRNAs are by-products generated by RNA polymerase
II, initially considered as ‘junk’ fragments (6). In 1991, Borsani et al
(7) confirmed that the lncRNA XIST
is involved in X-chromosome inactivation. Since then, increasing
research has demonstrated the critical role of lncRNAs in
biological functions, with abnormal expression often leading to
various diseases, particularly malignancies (8). Differential expression of lncRNAs has
been observed in cancerous and adjacent non-cancerous tissues,
suggesting their involvement in cancer initiation and progression.
For example, Li et al (9)
found that the expression of lncRNA HOXA11-AS was markedly higher
in oral squamous cell carcinoma tissue compared to adjacent
non-cancerous tissues. Liu et al (10) reported that lncRNA H19 is markedly
downregulated in nephroblastoma, highlighting its potential role in
the development of this cancer.
The multi-faceted roles of lncRNAs in tumorigenesis
are increasingly being uncovered. Current evidence indicates their
involvement in regulating gene expression through several
mechanisms, from epigenetic remodeling to post-transcriptional
processing (8,11). Such regulation directly impacts
pivotal cellular processes, including cell invasion, metastasis,
proliferation, apoptosis and angiogenesis, which define cancer
pathogenesis (12). Illustrating
their significance, lncRNAs such as HOTAIR, MALAT1, NEAT1 and MEG3
have been characterized as active contributors, either promoting or
restraining tumor progression in various contexts (13). A key translational advantage of
lncRNAs lies in their detectability in bodily fluids such as plasma
and urine, where their expression profiles dynamically reflect
disease activity (14). This
property positions them as potentially valuable biomarkers for the
early detection and prognostic monitoring of lung cancer.
While the roles of lncRNAs in various cancers have
been reviewed, a focused, critical evaluation in the context of
lung cancer is warranted. This necessity stems from the need to
decipher the specific molecular pathogenesis of the world's leading
cause of cancer death. Lung cancer's distinct etiology, most
notably its strong association with tobacco smoke, drives a unique
spectrum of molecular alterations and shapes a correspondingly
unique and therapeutically relevant landscape of lncRNA
dysregulation (15). This is
exemplified by the direct induction of smoke-responsive oncogenic
lncRNAs such as smoke and cancer-associated lncRNA 1, which is
upregulated by cigarette smoke to mediate reactive oxygen species
detoxification and promote cell survival (16). Understanding these etiology-driven
lncRNA mechanisms is therefore not merely an academic exercise, but
crucial for developing targeted strategies against this uniquely
prevalent malignancy. Additionally, the profound clinical
challenges in lung cancer, including the high frequency of
late-stage diagnosis and the rapid development of resistance to
therapy, underscore an urgent and specific need for novel
biomarkers and therapeutic targets that lncRNAs may meet (17).
Positioning of this review
While existing reviews have markedly advanced our
understanding, a synthesis organized around a dedicated
translational framework is still needed. Such a framework should
explicitly link mechanistic insights, including therapy resistance,
to clinical applications by emphasizing the role of enabling
technologies, while also critically evaluating implementation
challenges. To precisely define our contribution, this work is
positioned through a detailed comparison with five key reviews
published between 2019 and 2024.
Compared to the narrative review by Yu et al
(18), which begins by
highlighting the limitations of traditional lung cancer screening
methods and then focuses on summarizing dysregulated lncRNAs as
promising biomarkers, the present review adopted a fundamentally
different translational structure. Rather than organizing content
around biomarker discovery per se, the present review structured
its narrative as a continuous translational pipeline. This
framework begins with molecular mechanisms, extends to clinical
applications including therapy response prediction and culminates
in a discussion of the technological advances and practical
challenges that determine real world clinical implementation. Thus,
while both reviews recognize the clinical potential of lncRNAs, the
present review provided a more comprehensive roadmap for their
translation.
Whereas Hu et al (19) focus on lncRNA functions in NSCLC
tumorigenesis using a translational ‘from bench to bedside’
structure, the present review encompassed both NSCLC and SCLC.
Beyond broadening the scope, the present review provided a deeper,
dedicated analysis of predictive biomarkers for therapy response.
Moreover, it included a standalone section on advances in lncRNA
detection, a critical dimension absent from the discussion of Hu
et al (19).
In contrast to the broad overview by Lv et al
(20), which encompasses all major
non-coding RNA types, including miRNAs, lncRNAs and circular RNAs
and dedicates significant attention to exosomal ncRNAs in lung
cancer, the present review delivered a focused, in-depth synthesis
exclusively on lncRNAs across lung cancer. Its core theme is the
integrated translational pathway for lncRNAs, rather than the
exosome as a functional unit.
While the comprehensive review by Ao et al
(21) details the roles of lncRNAs
as oncogenes or tumor suppressors within NSCLC, exploring
mechanistic networks such as the competing endogenous RNA (ceRNA)
and tumor microenvironment modulation, the present review extended
the discussion by introducing a key translational pillar. A primary
distinction is its dedicated section evaluating advances in lncRNA
detection. The present review focused specifically on how
cutting-edge single-cell and spatial transcriptomics revolutionized
discovery and bridge biology with clinical application. This
represents a distinct technological dimension that complements
their mechanistic and clinical analysis.
Finally, in relation to the foundational overview by
Jiang et al (22) on
regulation patterns and biologic functions, the present review
integrated significant subsequent advances, particularly in therapy
response and detection methodologies. It employed a more modern
translational medicine framework with distinct sections on clinical
potential and detection technology and a more comprehensive
critical analysis of challenges and future solutions.
In conclusion, the present review offered a distinct
integrative and translational perspective. It connected fundamental
molecular mechanisms with the complete clinical picture, placing
particular emphasis on the role of lncRNAs in predicting therapy
response. Furthermore, the present review integrated critical
discussions on emerging detection technologies and the principal
challenges facing clinical translation, weaving them into a unified
narrative. This structured approach addresses a recognized need in
the literature and aimed to provide a clear roadmap for advancing
lncRNA research toward clinical applications across all major forms
of lung cancer, including both SCLC and NSCLC.
lncRNA mechanisms of action
lncRNAs share key biogenetic features with mRNAs,
including synthesis by RNA polymerase II from chromatin-template
regions (23). Their
classification is primarily grounded in a genomic context and
includes long intergenic, intronic, antisense, bidirectional and
enhancer-associated lncRNAs (24).
Functionally, lncRNAs exhibit a dichotomy in cancer, acting as
either oncogenes or tumor suppressors; the former are frequently
overexpressed to drive tumorigenesis, while the latter oppose
malignant progression and metastasis (25). This dualistic paradigm is clearly
illustrated in lung cancer, where disease progression is driven by
a dynamic interplay between upregulated oncogenic genes (such as
HOTAIR and MALAT1) and downregulated tumor suppressors (such as
GAS5 and MEG3). A further functional distinction lies in their
range of action: Cis-acting lncRNAs modulate local gene activity,
whereas trans-acting types influence targets distantly, reflecting
the broad regulatory versatility of these molecules across
physiological and pathological states (26).
The functions of lncRNA are largely defined by its
subcellular destination (Fig. 1).
In the nucleus, these transcripts predominantly govern
transcriptional regulation and chromatin architecture. Their
strategy often involves recruiting key modifiers, such as
transcription factors and epigenetic regulators, to specific
genomic sites to alter gene expression (27). For example, the oncogenic lncRNA
HOTAIR operates as a modular epigenetic scaffold in the nucleus,
recruiting complexes, such as PRC2, to repress histone markers
[histone H3 lysine 27 trimethylation (H3K27me3)] and silence tumor
suppressor genes (28). This
epigenetic reprogramming is associated with the hyperactivation of
oncogenic pathways, such as the Wnt/β-catenin pathway (29). A distinct nuclear role entails
functioning as molecular scaffolds that provide a structural
foundation for nuclear bodies, thereby coordinating extensive
regulatory networks (30).
Similarly, MALAT1 localizes to nuclear speckles and regulates
alternative splicing by interacting with serine/arginine-rich
proteins, while also participating in transcriptional activation
(31). Shifting to the cytoplasm,
the functional focus of lncRNAs transitions to post-transcriptional
control. Here, they exert influence over a broad spectrum of
events, including mRNA stability, translational efficiency and the
dynamics of intracellular signaling pathways, in part through
mechanisms such as molecular sponging (32). MALAT1, for example, functions as a
classic molecular sponge by sequestering tumor-suppressive miR-145,
thereby dysregulating the SOX9 oncogene and enhancing cancer
progression, a process that can be targeted by natural compounds
such as Glehnia littoralis polysaccharides (33). Conversely, the tumor-suppressive
lncRNA GAS5 functions as a decoy in the cytoplasm by binding to the
glucocorticoid receptor, preventing its transcriptional activation
of pro-survival genes and also acts as a ceRNA to modulate
pathways, such as mTOR and AKT (34). These mechanisms and associated
signaling pathways are summarized in Table I (28,29,31,33–36).
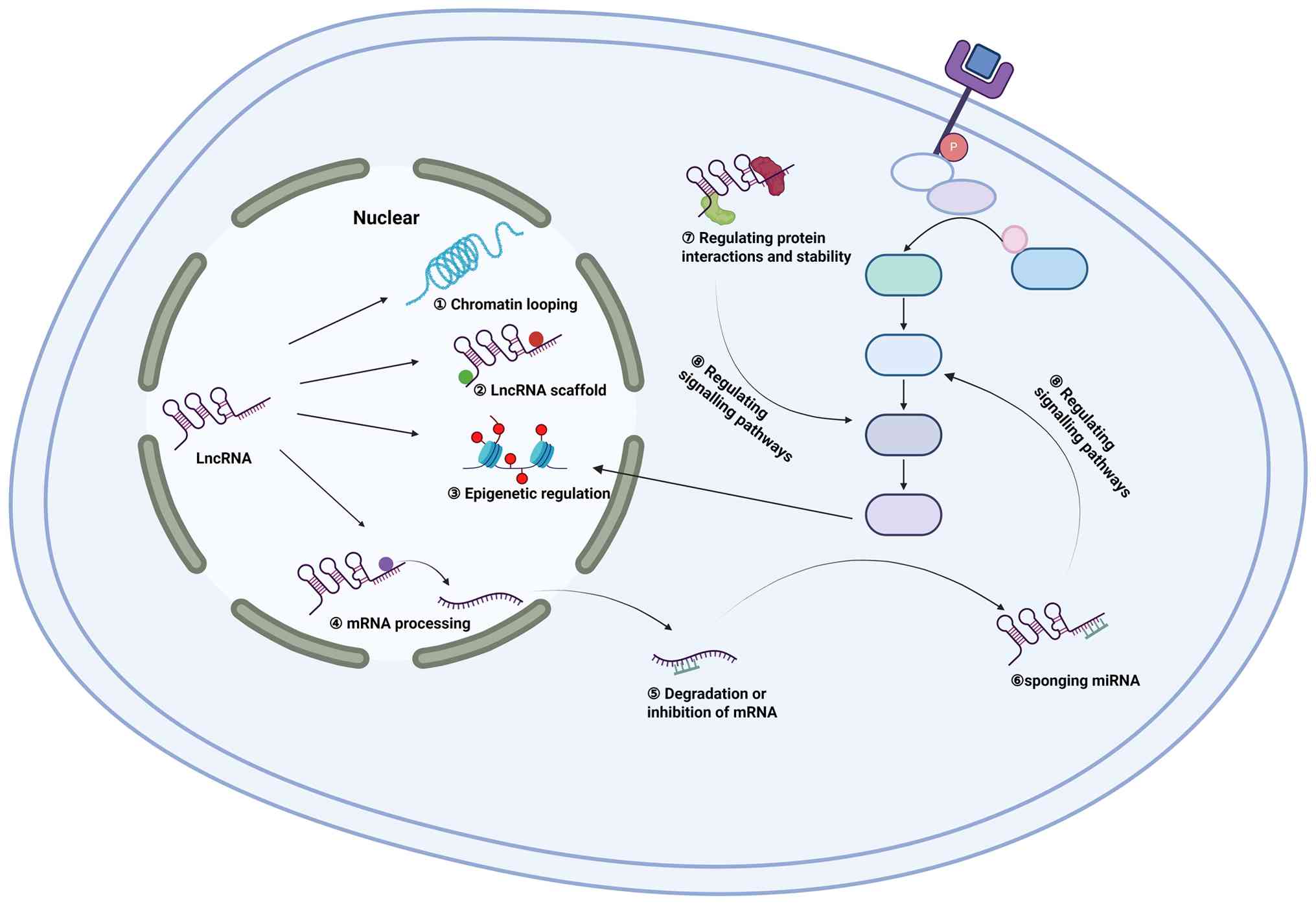 | Figure 1.Mechanisms of lncRNA-mediated
regulation in the nucleus and cytoplasm. Within the nucleus,
lncRNAs facilitate gene regulation through multiple mechanisms.
They can promote ① the formation of chromatin loops or ② serve as
scaffolding platforms that recruit diverse regulatory complexes to
gene promoters, leading to transcriptional activation or
repression. This is often achieved by ③ guiding epigenetic
modifiers to specific genomic loci, thereby altering DNA or histone
methylation patterns. ④ Nuclear lncRNAs can also recruit regulatory
molecules to mRNAs to regulate mRNA processing. Following their
export to the cytoplasm for translation, mRNAs become targets for
miRNAs, which typically repress gene expression by ⑤ inducing mRNA
degradation or inhibiting translation. ⑥ Cytoplasmic lncRNAs can
counteract this repression by acting as ceRNAs, sequestering miRNAs
and consequently derepressing their target mRNAs. Beyond miRNA
sponging, cytoplasmic lncRNAs also ⑦ regulate protein-protein
interactions and protein stability, thereby modulating signal
transduction pathways and the resulting gene expression profiles. ⑧
Similarly, the sequestration of miRNAs by sponging mechanisms
serves as a key regulatory layer for signaling cascades by
controlling mRNA activity. lncRNA, long non-coding RNAs; miRNAs,
microRNAs; ceRNAs, competitive endogenous RNAs. |
 | Table I.Key lncRNAs in lung cancer, their
mechanisms and associated signaling pathways. |
Table I.
Key lncRNAs in lung cancer, their
mechanisms and associated signaling pathways.
| First author/s,
year | LncRNA | Functional
role | Expression | Subcellular
localization | Primary mechanisms
of action | Associated
signaling pathways | (Refs.) |
|---|
| Herrera-Solorio
et al, 2017; Hakami et al, 2024 | HOTAIR | Oncogene | Up | Nucleus | Functions as a
modular epigenetic scaffold. Recruits PRC2 to deposit the
repressive histone mark H3K27me3, leading to silencing of tumor
suppressor genes. | Wnt/β-catenin
pathway (activated via epigenetic silencing of its
inhibitors). | (28,29) |
| Tripathi et
al, 2010; Alinejad et al, 2025 | MALAT1 | Oncogene | Up |
Nucleus/Cytoplasm | 1. Nucleus:
Localizes to nuclear speckles; regulates alternative splicing by
interacting with SR proteins and participates in transcriptional
activation. | miR-145/SOX9 axis
and other pathways promoting cancer progression. | (31,33) |
|
|
|
|
|
| 2. Cytoplasm: Acts
as a molecular sponge/ceRNA by sequestering tumor-suppressive
miR-145, leading to dysregulation of oncogenes such as SOX9. |
|
|
| Zhou and Chen,
2020 | GAS5 | Tumor
suppressor | Down | Cytoplasm | 1. Decoy mechanism:
Serves as a decoy RNA by binding to the GR, preventing its
transcriptional activation of pro-survival genes. | GR pathway, mTOR
pathway, AKT pathway. | (34) |
|
|
|
|
|
| 2. ceRNA mechanism:
Functions as a ceRNA to modulate pathways such as mTOR and
AKT. |
|
|
| Ghafouri-Fard and
Taheri, 2019; Lu et al, 2013 | MEG3 | Tumor
suppressor | Down | Nucleus | Exerts
growth-inhibitory effects primarily by activating the p53 tumor
suppressor pathway. Its frequent downregulation is often mediated
by promoter hypermethylation. | p53 tumor
suppressor pathway. | (35,36) |
lncRNAs exert their regulatory influence on gene
expression via three core mechanistic modes: Chromatin interaction,
RNA target binding and modulation of protein activity (37). A single lncRNA frequently operates
through multiple such pathways in parallel, forming intricate
regulatory circuits. This functional versatility allows them to
interface with diverse cellular components, a property of
substantial relevance to cancer (38). The frequent downregulation of
tumor-suppressive lncRNAs such asGAS5 and MEG3 in advanced lung
cancer, often through mechanisms such as promoter hypermethylation,
removes essential growth constraints (35). In the case of MEG3, this epigenetic
silencing abrogates its critical nuclear function in activating the
p53 tumor suppressor pathway, a key mechanism for its
growth-inhibitory effects (36).
Indeed, numerous studies have linked lncRNA activity to key
oncogenic processes, including uncontrolled proliferation, evasion
of apoptosis, invasion, metastasis and resistance to therapy
(36,39,40).
Therefore, a systematic exploration of lncRNA expression and
function is essential, both for advancing our fundamental
understanding of tumors and for developing novel diagnostics and
targeted therapies. Future therapeutic strategies must therefore be
dual-pronged, aiming to inhibit oncogenic lncRNAs while devising
means to reactivate silenced tumor-suppressive ones.
lncRNAs in the development of lung
cancer
The onset and progression of lung cancer are both
driven by dysregulated gene expression, typically involving the
activation of oncogenes and the inhibition of tumor suppressor
genes. lncRNAs have been identified as key regulatory factors that
influence lung cancer cell proliferation and apoptosis, as well as
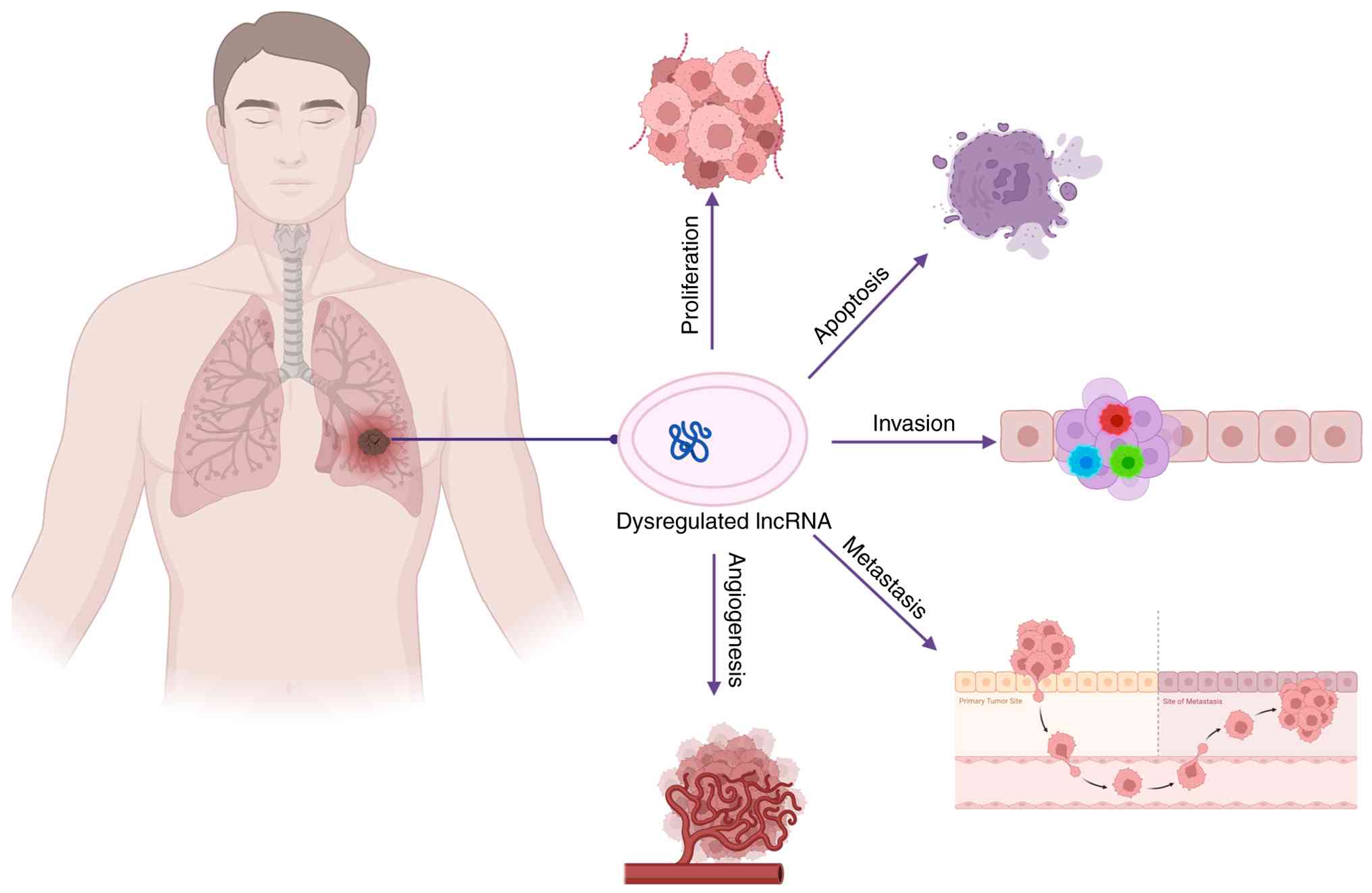
invasion, metastasis and tumor angiogenesis (Fig. 2).
lncRNAs in lung cancer cell
proliferation and apoptosis
How tumor cells achieve uncontrolled proliferation
and evade cell death is a central question in cancer research
(41). lncRNAs are important
regulators of these processes in lung cancer, primarily through
acting as ceRNAs to sponge miRNAs, modulating epigenetic states and
influencing key metabolic pathways such asglycolysis (42,43).
A prevalent mechanism is the presence of ceRNA networks, where
lncRNAs sequester specific miRNAs, thereby reducing the expression
of the miRNAs' target genes and thus driving oncogenic phenotypes.
For example, Tang et al (44) demonstrated that lncRNA UCA1
promotes lung cancer cell proliferation and inhibits apoptosis by
competitively binding miR-383 to upregulate vascular endothelial
growth factor A (VEGFA). Similarly, Liu et al (45) demonstrated that lncRNA DARS-AS1
drives tumor progression in lung cancer by enhancing cell
proliferation and suppressing apoptotic death. Mechanistically,
DARS-AS1 functions by sequestering miR-188-5p, an action that
ultimately leads to the elevated expression of the KLF12 protein.
This ceRNA mechanism also extends to the regulation of cancer cell
metabolism. lncRNA HOXA11-AS promotes proliferation and glycolysis
by binding to miR-148b-3p to enhance PKM2 expression (46), while LINC00665 facilitates aerobic
glycolysis via a let-7c-5p/HMMR axis (42).
Beyond post-transcriptional regulation, lncRNAs
directly influence the cell cycle and epigenetic programming.
lnc-TMEM132D-AS1 promotes proliferation and induces
M2/G1 cell cycle arrest by sponging miR-766-5p to
upregulate ENTPD1 (39). By
contrast, lncRNA SNHG6 employs a different strategy; it recruits
EZH2 to the p27 promoter, leading to H3K27me3-mediated epigenetic
silencing of this key cell cycle inhibitor, thereby enhancing the
G1/S transition and proliferation (47). In summary, these studies underscore
the critical role of lncRNAs in regulating lung cancer cell
proliferation, the cell cycle and apoptosis. A summary of these
mechanisms is provided in Table
II (39,42,44–60).
 | Table II.Role and mechanism of lncRNAs in
proliferation and apoptosis of lung cancer. |
Table II.
Role and mechanism of lncRNAs in
proliferation and apoptosis of lung cancer.
| First author/s,
year | LncRNA | Expression | Molecular
mechanism | Function in lung
cancer | (Refs.) |
|---|
| Wang et al,
2023 | TMEM132D-AS1 | Up |
miR-766-5p/ENTPD1 | Cell
cycle/proliferation/apoptosis | (39) |
| Li et al,
2024 | LINC00665 | Up | Let-7c-5p/HMMR |
Proliferation/glycolysis | (42) |
| Tang et al,
2023 | UCA1 | Up | miR-383/VEGFA |
Proliferation/apoptosis | (44) |
| Liu et al,
2021 | DARS-AS1 | Up |
miR-188-5p/KLF12 |
Proliferation/apoptosis | (45) |
| Chen et al,
2023 | HOXA11-AS | Up |
miR-148b-3p/PKM2 |
Proliferation/glycolysis | (46) |
| Wang et al,
2022 | SNHG6 | Up | EZH2/p27 | Cell
cycle/proliferation | (47) |
| Zheng et al,
2020 | PIK3CD-AS2 | Up | YBX1/p53 |
Proliferation/apoptosis | (48) |
| Li et al,
2021 | MALAT1 | Up | FOXP3/GINS1 | Proliferation | (49) |
| Han et al,
2020 | UPLA1 | Up |
DSP/Wnt/β-catenin | Cell
cycle/proliferation | (50) |
| Jin et al,
2020 | FTX | Down |
miR-200a-3p/FOXA2 | Proliferation | (51) |
| Cao et al,
2020 | MBNL1-AS1 | Down |
miR-135a-5p/LOXL4 | Cell
cycle/proliferation/apoptosis | (52) |
| Yang et al,
2023 | LINC02159 | Up | ALYREF/YAP1 | Cell
cycle/proliferation/apoptosis | (53) |
| Zhu et al,
2022 | MNX1-AS1 | Up |
MNX1-AS1/IGF2BP | Cell
cycle/proliferation | (54) |
| Min et al,
2024 | MIR100HG | Up |
miR-5590-3p/DCBLD2 | Proliferation | (55) |
| Hong et al,
2024 | MACC1-AS1 | Up |
miR-579-3p/NOTCH1 |
Proliferation/apoptosis | (56) |
| Ma et al,
2024 | HAR1A | Down | MYC/HSP90β |
Proliferation/apoptosis | (57) |
| Zhai et al,
2025 | SLC7A11AR | Up |
miR-150-5p/SLC7A11 | Proliferation | (58) |
| Ke et al,
2025 | LINC00323 | Up |
LINC00323/AKAP1 | Proliferation | (59) |
| Wang et al,
2021 | GAN1 | Down |
miR-26a-5p/PTEN |
Proliferation/apoptosis | (60) |
While in vitro models have provided most of
the current insights into lncRNA function in lung cancer cell
growth and death, the true regulatory landscape within a complex
tumor microenvironment (TME) is markedly more complex. This
disparity between model systems and physiological reality
represents a key challenge that can only be addressed through
robust in vivo experimentation.
lncRNAs in lung cancer cell invasion
and metastasis
Tumor cell invasion and metastasis constitute the
primary drivers of cancer recurrence and patient mortality
(61), underscoring the critical
need to elucidate their underlying mechanisms for therapeutic
advancement. Within this paradigm, lncRNAs have been identified as
essential regulators. For example, work by Li et al
(62) illustrated how lncRNA TEX41
promoted lung cancer cell invasion and metastasis by upregulating
Runx2 and suppressing the PI3K/AKT signaling pathway. This
regulatory influence extended beyond the cancer cells themselves.
Notably, components of the TME, such as neutrophil extracellular
traps (NETs) formed during infection or inflammation, are
increasingly being recognized for their role in facilitating
metastatic spread (63).
Supporting this, Wang et al (64) demonstrated that NETs promoted NSCLC
metastasis by inhibiting lncRNA MIR503HG, an action that activated
the pro-metastatic NF-κB/NOD-like receptor protein 3 inflammasome
pathway.
Central to cancer metastasis is
epithelial-mesenchymal transition (EMT), a pivotal reprogramming
event that dismantles the epithelial phenotype and confers cells
increased migratory and invasive attributes associated with a
mesenchymal state (65). As key
regulators of this process, the dysregulation of specific lncRNAs
has been shown to drive EMT, thereby endowing lung cancer cells
with invasive and metastatic capabilities. Pan et al
(66) reported that lncRNA JPX
promoted lung cancer cell invasion and metastasis by competitively
binding miR-33a-5p to upregulate Twist1, thereby activating the
Wnt/β-catenin signaling pathway, thereby inducing EMT.
Additionally, Zhong et al (67) found that lncRNA AFAP1-AS1 induced
EMT and accelerated the migration and invasion of lung cancer cells
by interacting with SNIP1 to upregulate c-Myc. The specific
mechanism involves AFAP1-AS1 binding to SNIP1, which acts as a
molecular guide to mediate the SNIP1-c-Myc interaction. This
interaction likely masks the ubiquitination site of c-Myc, thereby
inhibiting its ubiquitination and proteasomal degradation, leading
to the accumulation of stabilized c-Myc protein. The elevated c-Myc
protein, as a transcription factor, subsequently upregulates the
transcription of key EMT master regulators, including ZEB1, ZEB2
and SNAIL. The increased expression of these factors drives the EMT
program, characterized by the loss of epithelial markers, such as
E-cadherin and the gain of mesenchymal markers. This reprogramming
confers enhanced migratory and invasive properties to lung cancer
cells, ultimately promoting metastasis (67).
In summary, invasion and metastasis are multi-step
malignant processes and lncRNAs may act as key regulatory factors
in these processes. Developing blockers targeting lncRNAs could
potentially reduce lung cancer metastasis, thus improving patient
prognosis. A summary of lncRNAs associated with lung cancer
migration, invasion, metastasis and EMT is provided in Table III (40,62,64,66–74).
 | Table III.Roles and mechanisms of lncRNAs in
migration, invasion, metastasis and EMT of lung cancer. |
Table III.
Roles and mechanisms of lncRNAs in
migration, invasion, metastasis and EMT of lung cancer.
| First author/s,
year | LncRNA | Expression | Molecular
mechanism | Function in lung
cancer | (Refs.) |
|---|
| Wang et al,
2025 | RP11-297P16.4 | Up |
miR-145-5p/MMP-2/9 |
Migration/invasion | (40) |
| Li et al,
2023 | TEX41 | Up | Runx2/PI3K/AKT |
Migration/invasion/metastasis/autophagy | (62) |
| Wang et al,
2022 | MIR503HG | Up | NF-κB/NLRP3 | Metastasis | (64) |
| Pan et al,
2020 | JPX | Up |
miR-33a-5p/Twist1 |
Metastasis/invasion/EMT | (66) |
| Zhong et al,
2021 | AFAP1-AS1 | Up | SNIP1/c-Myc |
Metastasis/invasion/EMT | (67) |
| Nie et al,
2022 | DUBR | Down |
c-Myc/DUBR/ZBTB11 |
Migration/invasion | (68) |
| Chen et al,
2025 | MSTO2P | Up | Wnt/β-catenin |
Migration/invasion | (69) |
| Ma et al,
2025 | SPAT | Down | SF1/KITLG |
Metastasis/migration | (70) |
| Chen et al,
2025 | EP300-AS1 | Down | PTBP1/PRMT5 |
Metastasis/migration/invasion | (71) |
| Xu et al,
2025 | LINC00472-ORF | Down | HDAC2/SP1 | Migration | (72) |
| Wu et al,
2025 | TMEM99 | Up |
TMEM99-FUBP3-p21/IGF2BP2 |
Migration/invasion/autophagy | (73) |
| Fang et al,
2025 | LUCAT1 | Up | ALYREF/TTYH3 |
Migration/invasion/EMT/metastasis | (74) |
lncRNA in lung cancer
angiogenesis
Angiogenesis is the formation of new blood vessels
from preexisting vessels to provide a nutrient supply that supports
tumor cell growth. Tumor-induced angiogenesis is characterized by
high permeability, irregular blood vessel formation, vascular
infiltration and immature blood vessels (75). Generally, tumor angiogenesis is a
complex process regulated by various angiogenic factors and
signaling pathways, such as VEGF and the angiopoietin (Ang)/Tie2
signaling pathway (76). VEGF is
the primary regulator of endothelial cell proliferation, directly
facilitating tumor growth and metastasis (77). Chen et al (78) found that lncRNA LINC00173.v1 acts
as a molecular sponge for miR-511-5p, thereby enhancing VEGFA
expression. VEGFA, in turn, directly acts on endothelial cells to
promote angiogenesis, accelerating the progression of lung squamous
cell carcinoma. Additionally, Hou et al (79) demonstrated that in NSCLC, lncRNA
EPIC1 stimulates endothelial cell proliferation via the Ang2/Tie2
pathway, leading to angiogenesis and the formation of vascular
channels.
The pro-angiogenic functions of lncRNAs establish
them as key facilitators of tumor progression, directly
orchestrating the shift from a localized lesion to widespread
metastatic disease.
Clinical potential of lncRNAs
As our understanding of the pathogenic roles of
lncRNAs in lung cancer increases, their translational prospects
have become increasingly compelling. Their specific expression
across different disease contexts provides a foundation for novel
approaches in early detection, treatment monitoring and outcome
prediction. This section addressed the advances in applying lncRNAs
as diagnostic biomarkers, therapeutic targets and prognostic tools,
tracing their development from bench research to clinical
utility.
lncRNAs as diagnostic biomarkers for
lung cancer
The ongoing challenge of early lung cancer
diagnosis, primarily due to a lack of reliable biomarkers,
continues to negatively impact patient survival, even as imaging
technologies advance (1). This
reality highlights the critical need for diagnostic methods that
are not only effective but also economical and minimally invasive.
Here, lncRNAs offer considerable promise. They are notable for
their high specificity, stability in body fluids such as serum and
saliva and suitability for quantitative analysis, making them
strong candidates as diagnostic biomarkers (12).
However, a critical assessment from the perspective
of non-invasive diagnosis reveals several challenges. While lncRNAs
demonstrate good stability in circulation, their typically low
abundance in body fluids poses significant technical challenges for
the reliability of detection and analytical sensitivity,
potentially limiting their clinical utility in real-world settings
(80). However, the performance of
individual lncRNAs varies. A comparative analyses of key diagnostic
metrics across different studies, such as area under the curve
(AUC), sensitivity and specificity, are summarized in the present
review, revealing the potential and limitations of various lncRNA
candidates (Table IV) (81–93).
HOTAIR, for example, is dysregulated across numerous types of
cancer yet shows a sensitivity of only 52.3% in NSCLC detection,
despite an 86.9% specificity, which is insufficient for a
standalone diagnosis (81). This
characteristically low sensitivity is particularly problematic for
non-invasive early detection, as it can lead to unacceptably high
false-negative rates in screening scenarios, potentially missing
early-stage cancer when intervention would be most beneficial.
 | Table IV.LncRNAs as diagnostic markers in lung
cancer. |
Table IV.
LncRNAs as diagnostic markers in lung
cancer.
| First author/s,
year | Biomarker | Expression | Sample | Method | AUC | Sensitivity, % | Specificity, % | (Refs.) |
|---|
| Yao et al,
2022 | HOTAIR | Up | Tissue | RT-qPCR | 0.801 | 52.3 | 86.9 | (81) |
| Li et al,
2019 | GAS5 | Down | Serum exosome | RT-qPCR | 0.857 | 85.94 | 70.00 | (82) |
| Li et al,
2019 | CEA | UP | Serum | RT-qPCR | 0.758 | 57.81 | 87.50 | (82) |
| Li et al,
2019 | GAS5+CEA | NA | NA | NA | 0.929 | 89.06 | 90.00 | (82) |
| Tantai et
al, 2015 | XIST | UP | Serum | RT-qPCR | 0.834 | NR | NR | (83) |
| Tantai et
al, 2015 | HIF1A-AS1 | UP | Serum | RT-qPCR | 0.876 | NR | NR | (83) |
| Tantai et
al, 2015 | XIST+HIF1A-AS1 | NA | NA | NA | 0.931 | NR | NR | (83) |
| Min et al,
2022 | RP5-977B1 | Up | Serum | RT-qPCR | 0.8899 | 82.86 | 84.93 | (84) |
| He et al,
2022 | HEIH | Up | Peripheral
blood | RT-qPCR | 0.860 | 72.86 | 95.71 | (85) |
| Wang et al,
2022 | LINC00313 | Up | Serum | RT-qPCR | 0.916 | 78.91 | 90.63 | (86) |
| Yang et al,
2020 | LINC00173 | Up | Serum | RT-qPCR | 0.809 | 62.96 | 89.01 | (87) |
| Zhang et al,
2019 | DLX6-AS1 | Up | Serum | RT-qPCR | 0.806 | 77.5 | 85.9 | (88) |
| Li et al,
2025 | MIR99AHG | Down | Serum | RT-qPCR | 0.908 | 85.71 | 92.38 | (89) |
| Zang et al,
2020 | UFC1 | Up | Serum exosome | RT-qPCR | 0.794 | 73.3 | 74.1 | (90) |
| Mohamed et
al, 2022 | RP11-510M2.10 | Down | Serum exosome | RT-qPCR | 0.918 | 95 | 90 | (91) |
| Wang et al,
2022 | LUCAT1 | Up | Serum exosome | RT-qPCR | 0.852 | 84.45 | 77.38 | (92) |
| Teng et al,
2019 | SOX2-OT | Up | Serum exosome | RT-qPCR | 0.815 | 76 | 73.17 | (93) |
A more productive approach involves combining
several biomarkers. One study found that the exosomal lncRNA GAS5
is downregulated in NSCLC. When GAS5 was used in conjunction with
the conventional marker CEA, the combined AUC reached 0.929,
outperforming either marker alone and demonstrating a significant
boost in positive diagnosis rates (82). Notably, combining lncRNAs
themselves can also yield improvements. Serum levels of XIST and
HIF1A-AS1 are elevated in NSCLC. Their combined use achieved an AUC
of 0.931, which was higher than that of XIST (AUC=0.834) or
HIF1A-AS1 (AUC=0.876) used independently (83). A systematic comparison of the
diagnostic performance between single and combined biomarkers is
provided in Table IV (81–93).
A clear pattern emerged; all combined biomarker panels demonstrated
a consistent increase in AUC values to >0.9, with synchronous
optimization of both sensitivity and specificity, markedly
outperforming any single biomarker. This comparison provides
compelling evidence of the substantial value in incorporating
lncRNAs into combined diagnostic strategies. While these
combination strategies show improved performance, they also
increase the complexity and potential cost of non-invasive tests,
which are important considerations for developing economically
viable liquid biopsy platforms suitable for widespread clinical
implementation.
Research has indicated that exosomes are selectively
packaged with distinct lncRNA profiles in malignant tissues,
facilitating their role in cell-to-cell communication and tumor
progression (94–96). Exosomal lncRNAs are crucial for
studying tumor biology as they participate in various processes
such as cell proliferation, invasion, metastasis, angiogenesis,
drug resistance and immune-suppressive microenvironments (97,98).
For example, Shi et al (97) found that lncRNA Mir100hg is
upregulated in lung cancer stem cells and can be delivered via
exosomes to target miR-15a-5p and miR-31-5p, thereby increasing
glycolytic activity and enhancing the metastatic potential of lung
cancer cells. Therefore, lncRNA Mir100hg may serve as a diagnostic
biomarker for lung cancer. Similarly, Mao et al (98) demonstrated that exosomal lncRNA
FOXD3-AS1 derived from lung cancer cells upregulated the expression
of ELAV-like RNA-binding protein 1 and activated the PI3K/Akt
pathway to promote lung cancer progression, thus making lncRNA
FOXD3-AS1 another potential diagnostic biomarker for lung cancer.
Despite this strong biological rationale, the technical challenges
in consistently isolating and characterizing tumor-derived exosomes
from blood samples remain substantial. The current lack of
standardized protocols for exosome isolation and lncRNA
quantification represents a major bottleneck in translating these
findings into clinically applicable non-invasive tests (99).
lncRNAs show promising diagnostic efficiency as
non-invasive biomarkers for lung cancer, yet their clinical
translation requires addressing key limitations. Current evidence
is constrained by insufficient sample sizes, unclear mechanisms of
dysregulation, technical challenges in reliably isolating
tumor-derived exosomes from bodily fluids and a lack of
comprehensive specificity analysis across cancer types (21). Future work should prioritize
large-scale multicenter validation using standardized liquid biopsy
protocols, mechanistic studies of lncRNA secretion and function in
biofluids and direct comparisons of leading lncRNA candidates
across a range of populations. Ultimately, transforming lncRNAs
into clinically viable non-invasive diagnostic tools will depend on
developing robust, cost-effective detection methods and rigorously
demonstrating their analytical and clinical validity in real-world
settings.
Application of lncRNAs in lung cancer
treatment
Currently, surgery, thoracic radiotherapy,
chemotherapy and targeted therapies are commonly used either alone
or in combination to treat patients with lung cancer (100). However, resistance to
chemotherapy and targeted therapies requires exploration of novel
therapeutic approaches (2). Since
lncRNAs play crucial roles in various aspects of lung cancer
development and regulate key signaling pathways, they represent
promising therapeutic targets. Moreover, several lncRNAs are
associated with enhanced chemotherapy resistance, suggesting that
targeting these lncRNAs may potentially restore cancer cell
sensitivity to chemotherapy drugs (101,102).
There are several strategies for targeting lncRNAs
in cancer treatment, including RNA interference (RNAi)-based gene
silencing, antisense oligonucleotide (ASO)-based therapies,
small-molecule regulators that modulate lncRNA-protein interactions
and the delivery of tumor-suppressive lncRNAs (36,103–105). It has been shown that
HOTAIR-siRNA, loaded into sodium alginate microspheres, can
markedly inhibit the proliferation, migration and invasion of lung
cancer cells (103). Further
study in a PC9/GR cell xenograft model in male BALB/c nude mice
confirmed that intratumoral injection of si-HOTAIR suppresses tumor
growth in vivo (103).
This anti-tumor effect is attributed to its ability to alleviate
acquired resistance to EGFR-tyrosine kinase inhibitors (TKIs) by
regulating the Hedgehog-Gli1 signaling pathway (103). Furthermore, HOTAIR siRNA-mediated
knockdown increases the sensitivity of lung cancer cells to
cisplatin treatment (106).
However, RNAi may cause off-target effects and nuclear RNA
targeting poses challenges, as several lncRNAs function within the
cell nucleus in lung cancer. By contrast, ASOs offer advantages due
to their high affinity, relatively low off-target effects and
reduced toxicity (107). A study
demonstrated that, in the mice model of experimental lung
metastasis established via intravenous injection of A549 cells,
treatment with MALAT1 ASO resulted in smaller lung tumor nodules
compared to control (105).
Additionally, Gong et al (104) developed a MALAT1-specific ASO and
nuclear-targeted TAT peptide co-functionalized Au nanoparticles,
called ASO-Au-TAT nanoparticles. These nanoparticles exhibited high
biocompatibility and markedly reduced the formation of metastatic
lung tumor nodules in an experimental lung metastasis model
established by intravenous injection of A549 cells in mice. This
suggests that MALAT1-ASO can inhibit lung cancer metastasis and may
serve as a reliable therapeutic approach for managing lung cancer.
Targeting the interaction between lncRNAs and proteins may be an
effective strategy to reduce off-target effects and enhance
targeting specificity. Several lncRNAs promote tumorigenesis in
lung cancer through interactions with the epigenetic regulator EZH2
or by modulating EZH2 activity. Researchers have developed
high-throughput screening methods to identify small molecule
inhibitors that target specific lncRNA-EZH2 interactions (108). lncRNA MEG3, a well-known tumor
suppressor, inhibits lung cancer cell migration and invasion and is
downregulated in lung cancer tissues (109). Overexpression of lncRNA MEG3
exerts potent antitumor effects in lung cancer. In vivo,
ectopic expression of MEG3 markedly suppressed the growth of SPC-A1
cell-derived xenograft tumors in female athymic BALB/c nude mice.
This was associated with reduced proliferation and induced
apoptosis of NSCLC cells in vitro (36). These findings indicate that
delivering tumor-suppressive lncRNAs, such as MEG3, represents a
promising therapeutic strategy for lung cancer. However, delivering
tumor-suppressive lncRNAs as a clinical treatment still requires
further research.
Predictive lncRNAs for therapeutic
response in lung cancer
Beyond their potential as direct therapeutic
targets, lncRNAs are increasingly recognized for their value as
predictive biomarkers. This application focuses on forecasting an
individual patient's likelihood of responding to a specific,
established therapy, such as chemotherapy, targeted agents, or
immunotherapy, thereby guiding personalized treatment decisions and
avoiding ineffective treatments and associated toxicity (110).
To date, this predictive potential has been
substantiated across all major therapeutic modalities for lung
cancer. In the realm of targeted therapy, reduced lncRNA H19
expression promotes acquired resistance to EGFR-TKIs such as
erlotinib in EGFR-mutant lung cancers by activating the PKM2/AKT
signaling axis (111). The
observation that AKT inhibition restores erlotinib sensitivity in
resistant models further supports the functional importance of this
pathway (111). In clinical
cohorts of EGFR-mutant lung cancer patients receiving EGFR-TKIs,
lower H19 levels are associated with markedly shorter
progression-free survival, indicating its utility as a predictive
biomarker for this specific patient population (111).
During chemotherapy, lncRNA UCA1 is frequently
upregulated in NSCLC and promotes resistance to platinum-based
drugs. Clinically, elevated UCA1 levels in tumor tissues or serum
are associated with poor response to platinum-doublet chemotherapy
and worse clinical outcomes, positioning it as a potential
predictive marker for chemosensitivity (112,113). Similarly, lncRNA XIST drives
cisplatin resistance through mechanisms such as modulating
glycolysis and inhibiting programmed cell death. Its high
expression is associated with poor chemotherapy response in
patients and has been validated as a predictive biomarker in
preclinical models (114,115). Furthermore, lncRNA HCG11 has been
shown to suppress gemcitabine resistance in NSCLC by acting as a
ceRNA for miR-17-5p and upregulating p21 expression. Its
tumor-suppressive role and ability to modulate chemosensitivity
highlight its potential as a predictive biomarker for responses to
gemcitabine-based chemotherapy (116).
The predictive role of lncRNAs is also prominently
exemplified in the context of immunotherapy, where they can
directly modulate the expression of immune checkpoint molecules.
Two compelling examples highlight distinct mechanistic layers of
this regulation. First, lncRNA LINC02418 functions as a
post-translational negative regulator of PD-L1. It promotes the
ubiquitination and proteasomal degradation of PD-L1 protein by
enhancing its interaction with the E3 ligase Trim21. Consequently,
higher LINC02418 expression is associated with lower PD-L1 protein
levels, increased CD8+ T cell infiltration and predicts more
favorable clinical outcomes in NSCLC patients receiving
anti-PD-1/PD-L1 therapy (117).
By contrast, lncRNA NKX2-1-AS1 operates at the transcriptional
level to suppress PD-L1. It interacts with the transcription factor
NKX2-1, interfering with its binding to the CD274 (PD-L1) promoter,
thereby repressing PD-L1 gene transcription. Loss of NKX2-1-AS1 may
thus contribute to an immune-evasive phenotype characterized by
elevated PD-L1 expression, positioning it as a potential biomarker
for identifying tumors reliant on the PD-1/PD-L1 axis (118).
These examples underscore a critical translational
avenue: the profiling of specific lncRNAs could enable pretreatment
stratification of patients into probable responders and
non-responders. By integrating such predictive lncRNA signatures
with existing clinicopathological and genetic data, a more precise
and effective personalized treatment strategy can be envisioned.
However, the clinical implementation of lncRNA-based predictive
models requires rigorous validation in large, prospective
multicenter cohorts and standardization of detection methods in
accessible biospecimens such asplasma or serum.
lncRNAs as prognostic biomarkers for
lung cancer
Beyond predicting the likelihood of response to
specific therapies, lncRNAs also hold significant value in
forecasting the long-term outcomes of lung cancer patients. The
prognosis of patients with lung cancer is closely associated with
the tumor node metastasis (TNM) staging system (119). Due to the presence of drug
resistance, the overall survival rate for patients with lung cancer
remains low. Currently, there is no accurate method to assess the
prognosis of lung cancer. Studies have shown that lncRNAs can serve
as predictive biomarkers for TNM staging, suggesting their
potential as prognostic markers for lung cancer. For example, Wang
et al (120) performed
bioinformatics analysis and experimental validation, discovering
that a novel lncRNA, AC079630.4, was markedly downregulated in lung
cancer tissues. Low expression of lncRNA AC079630.4 was associated
with later-stage disease and a worse prognosis compared to those
with high expression, indicating that it could serve as a potential
prognostic marker for lung cancer. Similarly, Chen et al
(121) found that lncRNA
AC099850.3 was markedly upregulated in LUAD. Through Cox
multivariate regression analysis, it was demonstrated that lncRNA
AC099850.3 was an independent prognostic factor associated with
overall survival (OS), disease-free survival and progression-free
survival in patients with LUAD. Liu et al (122) discovered that lncRNA KTN1-AS1
promoted the proliferation, migration, invasion and EMT of NSCLC
cells while inhibiting apoptosis. The expression of KTN1-AS1 is
associated with TNM stage, histological grade and lymph node
metastasis, with high KTN1-AS1 expression correlating with reduced
OS in patients with NSCLC. Furthermore, Song et al (123) used Cox regression and LASSO
regression analysis to identify five lncRNAs associated with LUAD
prognosis. Among them, GSEC, FAM83A-AS1, AL606489.1 and AC010980.2
were identified as potential risk factors, whereas AL034397.3 was a
potential protective factor.
These findings indicate that lncRNAs can serve as
prognostic markers and their expression levels in tumors may be
used to assess patients' clinicopathological features and OS.
Advances in lncRNA detection
Since the 1970s, DNA sequencing has evolved from
first-generation technologies to third-generation sequencing,
making significant contributions to the unraveling of the human
genome sequence (124). When
applied to lncRNA research, these advanced technologies have proven
particularly powerful. For example, Qi et al (125) identified a novel lncRNA,
LNC11649, in NSCLC using PacBio third-generation sequencing
technology. This molecule was shown to promote cytoplasmic MSI1
distribution by interacting with MSI1, thereby activating the Akt
signaling pathway to regulate NSCLC cell proliferation and
migration. These findings not only reveal a novel molecular
mechanism in NSCLC progression but also highlight the potential of
LNC11649 as a diagnostic biomarker, given its specific role in
promoting oncogenic signaling pathways.
Similarly, single-cell sequencing has become
indispensable for dissecting the roles of lncRNAs within the
complex TME, as it can reveal gene expression patterns and lncRNA
regulatory networks at the cellular level, which is crucial for
understanding tumor heterogeneity (126). The power of single-cell
resolution is exemplified by several recent studies leading to the
discovery of functionally distinct lncRNAs in LUAD. For instance,
mining of the single-cell RNA-seq dataset CancerSEA identified
LINC00847, a lncRNA whose expression is associated with immune cell
infiltration and PD-L1 regulation and lncRNA PCBP1-AS1, a
metastasis suppressor whose expression is negatively associated
with metastatic states in single-cell data, these molecules may
serve as potential therapeutic targets for LUAD (127,128). Direct analysis of LUAD
single-cell RNA-seq data enabled the construction of a prognostic
gene signature, from which SFTA3 was prioritized as a key
component. Its clinical relevance was validated by prognostic
association in independent cohorts and its tumor-suppressive
function was confirmed through in functional investigations showing
that knockdown promoted proliferation and migration while
overexpression inhibited these phenotypes. Its diagnostic potential
was further indicated by markedly decreased serum levels in LUAD
patients (129). Importantly,
these single-cell-derived hypotheses often require spatial
validation. A prime example is the study of LINC01116: Initial
single-cell analysis re-localized its expression from tumor cells
to lymphatic endothelial cells, a finding that was critically
validated at spatial resolution by RNA-FISH, thereby solidifying
its role in tumor lymphangiogenesis (130). This case highlights a growing
paradigm in which high-resolution single-cell findings provide
compelling hypotheses for subsequent spatial validation.
To overcome the limitation of single-cell
sequencing, which loses spatial information, spatial
transcriptomics provides precise localization data. The integration
of these two methods is key to fully understanding cellular
heterogeneity and spatial organization (131). This powerful combination has
directly accelerated lncRNA discovery. Zhu et al (132) Combined single-cell RNA sequencing
with spatial transcriptomics in LUAD; the authors' work not only
identified the specific subpopulations of cancer cells and the TME
but also mapped their spatial topography and interactions. This
multi-faceted analysis provided a window into the molecular and
cellular dynamics driving the progression from in situ to
invasive adenocarcinoma. Such spatial and molecular
characterization of lncRNAs enhances their potential as diagnostic
biomarkers by linking their expression to specific tumor regions
and progression stages. Lin et al (96) employed single-cell RNA sequencing
and spatial transcriptomics to identify lncCRLA, a chemotherapy
resistance-associated lncRNA in LUAD. This lncRNA was shown to be
highly expressed in LUAD and demonstrated prognostic potential for
predicting disease progression and evolution. The discovery of
lncCRLA underscored the clinical relevance of lncRNAs as biomarkers
for both diagnosis and prediction of treatment response in lung
cancer.
Furthermore, to support such integrative analyses,
resources such as the updated LnCeCell 2.0 database have been
developed, which combines single-cell and spatial transcriptomics
data and provides tools for investigating lncRNA-related ceRNA
networks with high resolution, thereby improving our understanding
of regulatory mechanisms in complex ecosystems (133). These resources markedly
facilitate the identification and validation of lncRNAs as
diagnostic biomarkers by enabling comprehensive analysis of their
expression and functional networks across different cellular and
spatial contexts.
Challenges and future perspectives
Although evidence supports the potential of lncRNAs
as diagnostic, prognostic and therapeutic tools for lung cancer,
numerous limitations remain for their clinical translation,
particularly in diagnostics. For example, translating exosomal
lncRNA biomarkers into reliable clinical assays faces significant
hurdles, including their low abundance in biofluids, technical
challenges in isolation and detection and a critical lack of
standardization (134). To
overcome this, future work must focus on standardizing isolation
protocols, developing more sensitive detection technologies (such
as digital PCR and next-generation sequencing) and establishing
uniform validation criteria for exosomal lncRNAs to ensure their
reliability as clinical biomarkers (135). Additionally, while nucleic
acid-based therapies (such as ASOs and siRNAs) targeting lncRNAs
hold therapeutic promise, their development faces challenges,
including off-target effects, which could confound the functional
validation of these lncRNAs as diagnostic targets (21). Furthermore, unlike proteins, the
three-dimensional structures of several lncRNAs remain largely
unknown and the absence of conserved domains complicates the design
of small-molecule inhibitors and, more broadly, the understanding
of their precise mechanisms of action (136). The mechanisms and regulatory
networks of a number of lncRNAs in lung cancer remain poorly
understood, hindering the identification of highly specific and
reliable diagnostic or prognostic biomarkers. Although certain
lncRNAs, such as MALAT1, are conserved, most lack cross-species
conservation, hindering validation in animal models and preclinical
studies, which are crucial steps for both therapeutic and biomarker
development (137,138).
A significant obstacle for therapeutic applications
is the difficulty in the pulmonary delivery of nucleic acid-based
agents such assiRNA and ASO (139,140). Physical barriers such as mucosa
and cilia, along with immune clearance mechanisms, complicate
initial delivery (141). To
overcome these drug delivery hurdles, potential solutions include
the development of advanced nanoparticle-based delivery systems
(such as lipid nanoparticles and polymeric nanoparticles) and novel
approaches such as CRISPR-based genome editing tools that can
directly target lncRNA genes, though these strategies require
further optimization for lung-specific delivery and improved safety
profiles (142,143). Zeng et al (142) discovered that LINC02178 is a
carcinogen for LUAD and accordingly developed a nanoparticle small
interfering RNA delivery system (NPs/2178) targeting this gene. The
NPs/2178 gene delivery system markedly promoted LUAD apoptosis and
inhibited tumor growth, representing a promising gene therapy
strategy. Wang et al (143) confirmed that lncRNA FAM83H-AS1 is
a carcinogen for LUAD and targeted FAM83H-AS1 based on the CRISPR
interference method to inhibit the progression of LUAD. Therefore,
further optimization of in vivo lung drug delivery
techniques is necessary to accurately evaluate the therapeutic
efficacy of lncRNA-based therapies in lung cancer.
Looking ahead, research should focus on three key
areas. First, multi-omics integrative analyses, including studies
on lncRNA-protein interaction networks (144) and joint
epigenomics-transcriptomics analyses (145). Such integrated approaches can
help bridge the current knowledge gaps by comprehensively mapping
lncRNA regulatory circuits and identifying novel, clinically
relevant lncRNA targets. Second, high-throughput approaches can be
used to analyze in greater detail the molecular mechanisms of
cross-regulation between redox and epigenetic systems in
vitro and in vivo, elucidating regulatory mechanisms and
providing guidance for clinical treatment (146). Third, targeted drug delivery
systems for lung cancer treatment need to be developed. For
example, Tian et al (147)
found that intravenous injection of exosome-targeted doxorubicin, a
chemotherapy drug, specifically delivered it to tumor tissues in
nude mice, inhibiting tumor growth with minimal toxicity. These
platforms can be leveraged not only for the targeted delivery of
lncRNA therapeutics but also for the capture and detection of
tumor-specific exosomal lncRNAs, potentially leading to novel
liquid biopsy platforms for early diagnosis and monitoring.
In conclusion, while lncRNAs face numerous
challenges in clinical application, further research is essential
to address these issues. Specifically, future efforts should
prioritize bridging the translation gap for exosomal lncRNA
biomarkers through standardization and improved detection methods
and addressing drug-delivery challenges through innovative
engineering approaches, ultimately unlocking their full potential
in lung cancer management.
Conclusions
Lung cancer is the most common type of cancer in the
respiratory system. The pathogenesis of lung cancer is highly
complex and remains incompletely understood. Understanding the
mechanisms underlying lung cancer is crucial for developing
effective therapeutic strategies. Accumulating evidence indicates
that numerous dysregulated lncRNAs contribute to lung cancer
development and progression. These molecules are implicated in a
range of biological processes, including proliferation, apoptosis,
invasion, metastasis and angiogenesis. Moreover, aberrant lncRNA
expression can be readily detected in the body fluids of patients
and shows strong correlations with TNM stage, highlighting their
potential as non-invasive biomarkers and therapeutic targets
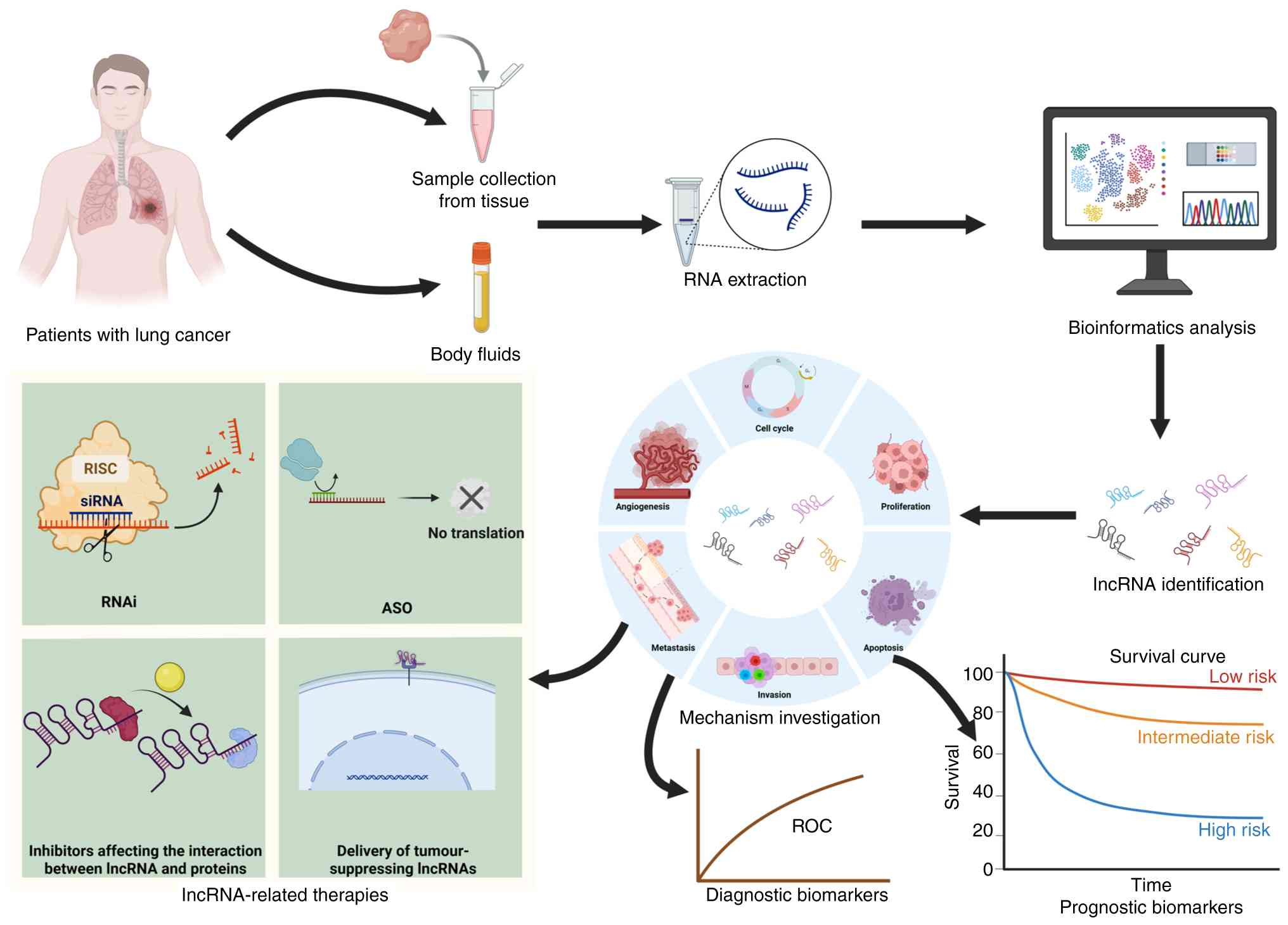
(Fig. 3).
Despite these promising findings, clinical
application of lncRNAs faces several challenges, such as the lack
of standardized detection methods, potential off-target effects and
limited cohort validation. Addressing these limitations will be
critical for their effective application in clinical practice.
Future research should focus on translating these findings into
clinical applications, such as integrating lncRNA signatures into
AI-based diagnostic tools to improve early detection and
personalized treatment strategies. Nevertheless, lncRNAs represent
valuable diagnostic and prognostic markers and may serve as
promising candidates for targeted therapies in lung cancer. Future
studies should continue to identify aberrantly expressed lncRNAs
and their precise functional roles in tumor biology and to explore
innovative lncRNA-based therapeutic approaches.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
Not applicable.
Authors' contributions
RP prepared the original manuscript draft. CW and YT
participated in conceptualization. FZ and YZ participated in
guiding the preparation and design of this manuscript. QZ reviewed
and edited the paper. Data authentication is not applicable. All
authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Nooreldeen R and Bach H: Current and
future development in lung cancer diagnosis. Int J Mol Sci.
22:86612021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hirsch FR, Scagliotti GV, Mulshine JL,
Kwon R, Curran WJ Jr, Wu YL and Paz-Ares L: Lung cancer: Current
therapies and new targeted treatments. Lancet. 389:299–311. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bridges MC, Daulagala AC and Kourtidis A:
LNCcation: lncRNA localization and function. J Cell Biol.
220:e2020090452021. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ponting CP, Oliver PL and Reik W:
Evolution and functions of long noncoding RNAs. Cell. 136:629–641.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhang Y, Wang X, Hu C and Yi H: Shiny
transcriptional junk: LncRNA-derived peptides in cancers and immune
responses. Life Sci. 316:1214342023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Entezari M, Ghanbarirad M, Taheriazam A,
Sadrkhanloo M, Zabolian A, Goharrizi MASB, Hushmandi K, Aref AR,
Ashrafizadeh M and Zarrabi A: Long non-coding RNAs and exosomal
lncRNAs: Potential functions in lung cancer progression, drug
resistance and tumor microenvironment remodeling. Biomed
Pharmacother. 150:1129632022. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Borsani G, Tonlorenzi R, Simmler MC,
Dandolo L, Arnaud D, Capra V, Grompe M, Pizzuti A, Muzny D,
Lawrence C, et al: Characterization of a murine gene expressed from
the inactive X chromosome. Nature. 351:325–329. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Tan YT, Lin JF, Li T, Li JJ, Xu RH and Ju
HQ: LncRNA-mediated posttranslational modifications and
reprogramming of energy metabolism in cancer. Cancer Commun (Lond).
41:109–120. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li B, Wang W, Miao S, Li G, Lv Y, Xiang C
and Pei R: HOXA11-AS promotes the progression of oral squamous cell
carcinoma by targeting the miR-518a-3p/PDK1 axis. Cancer Cell Int.
19:1402019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Liu HC, Zhu WY and Ren LY: LncRNA H19
inhibits proliferation and enhances apoptosis of nephroblastoma
cells by regulating the miR-675/TGFBI axis. Eur Rev Med Pharmacol
Sci. 26:3800–3806. 2022.PubMed/NCBI
|
|
11
|
Zhang C, Wang L, Jin C, Zhou J, Peng C,
Wang Y, Xu Z, Zhang D, Huang Y, Zhang Y, et al: Long non-coding RNA
Lnc-LALC facilitates colorectal cancer liver metastasis via
epigenetically silencing LZTS1. Cell Death Dis. 12:2242021.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Fu J, Yu L, Yan H, Tang S, Wang Z, Dai T,
Chen H, Zhang S, Hu H, Liu T, et al: LncRNAs in non-small cell lung
cancer: Novel diagnostic and prognostic biomarkers. Front Mol
Biosci. 10:12971982023. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Esposito R, Bosch N, Lanzós A, Polidori T,
Pulido-Quetglas C and Johnson R: Hacking the cancer genome:
Profiling therapeutically actionable long Non-coding RNAs using
CRISPR-Cas9 screening. Cancer Cell. 35:545–557. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Palanca-Ballester C, Rodriguez-Casanova A,
Torres S, Calabuig-Fariñas S, Exposito F, Serrano D, Redin E,
Valencia K, Jantus-Lewintre E, Diaz-Lagares A, et al: Cancer
epigenetic biomarkers in liquid biopsy for high incidence
malignancies. Cancers (Basel). 13:30162021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ge L, Zhao S, Sun J, Cheng J and Wang S:
Role of long noncoding RNAs in Smoking-induced lung cancer: An in
silico study. Comput Math Methods Med. 2022:71693532022. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Cruz CRV, Ferrer JLM and Garcia RL:
Concomitant and decoupled effects of cigarette smoke and SCAL1
upregulation on oncogenic phenotypes and ROS detoxification in lung
adenocarcinoma cells. Sci Rep. 11:183452021. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chen J, Wang R, Zhang K and Chen LB: Long
non-coding RNAs in non-small cell lung cancer as biomarkers and
therapeutic targets. J Cell Mol Med. 18:2425–2436. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yu P, He X, Lu F, Li L, Song H and Bian X:
Research progress regarding long-chain non-coding RNA in lung
cancer: A narrative review. J Thorac Dis. 14:3016–3029. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hu Q, Ma H, Chen H, Zhang Z and Xue Q:
LncRNA in tumorigenesis of non-small-cell lung cancer: From bench
to bedside. Cell Death Discov. 8:3592022. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lv X, Yang L, Xie Y and Momeni MR:
Non-coding RNAs and exosomal non-coding RNAs in lung cancer:
Insights into their functions. Front Cell Dev Biol. 12:13977882024.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ao YQ, Gao J, Jiang JH, Wang HK, Wang S
and Ding JY: Comprehensive landscape and future perspective of long
noncoding RNAs in non-small cell lung cancer: It takes a village.
Mol Ther. 31:3389–3413. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Jiang L, Li Z and Wang R: Long non-coding
RNAs in lung cancer: Regulation patterns, biologic function and
diagnosis implications (Review). Int J Oncol. 55:585–596.
2019.PubMed/NCBI
|
|
23
|
Eidem TM, Kugel JF and Goodrich JA:
Noncoding RNAs: Regulators of the mammalian transcription
machinery. J Mol Biol. 428:2652–2659. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kopp F and Mendell JT: Functional
classification and experimental dissection of long noncoding RNAs.
Cell. 172:393–407. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Huang Y, Ling A, Pareek S and Huang RS:
Oncogene or tumor suppressor? Long noncoding RNAs role in patient's
prognosis varies depending on disease type. Transl Res. 230:98–110.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Lee JT: Epigenetic regulation by long
noncoding RNAs. Science. 338:1435–1439. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sun Q, Hao Q and Prasanth KV: Nuclear long
noncoding RNAs: Key regulators of gene expression. Trends Genet.
34:142–157. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Herrera-Solorio AM, Armas-López L, Arrieta
O, Zúñiga J, Piña-Sánchez P and Ávila-Moreno F: Histone code and
long non-coding RNAs (lncRNAs) aberrations in lung cancer:
Implications in the therapy response. Clin Epigenetics. 9:982017.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Hakami MA, Hazazi A, Abdulaziz O,
Almasoudi HH, Alhazmi AYM, Alkhalil SS, Alharthi NS, Alhuthali HM,
Almalki WH, Gupta G and Khan FR: HOTAIR: A key regulator of the
Wnt/β-catenin signaling cascade in cancer progression and
treatment. Pathol Res Pract. 253:1549572024. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Tsai MC, Manor O, Wan Y, Mosammaparast N,
Wang JK, Lan F, Shi Y, Segal E and Chang HY: Long noncoding RNA as
modular scaffold of histone modification complexes. Science.
329:689–693. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tripathi V, Ellis JD, Shen Z, Song DY, Pan
Q, Watt AT, Freier SM, Bennett CF, Sharma A, Bubulya PA, et al: The
nuclear-retained noncoding RNA MALAT1 regulates alternative
splicing by modulating SR splicing factor phosphorylation. Mol
Cell. 39:925–938. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Nojima T and Proudfoot NJ: Author
Correction: Mechanisms of lncRNA biogenesis as revealed by nascent
transcriptomics. Nat Rev Mol Cell Biol. 23:8532022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Alinejad T, Hao Z, Zhou W, Zareh D,
Farajtabrizi E, Mossahebi-Mohammadi M and Chen CS: Knockdown of
long noncoding RNA MALAT1 enhances the anti-cancer effects of
polysaccharides Glehnia littoralis in lung cancer cells
possibly via the regulation of miR-145/SOX9 axis. Int
Immunopharmacol. 164:1153232025. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhou Y and Chen B: GAS5-mediated
regulation of cell signaling (Review). Mol Med Rep. 22:3049–3056.
2020.PubMed/NCBI
|
|
35
|
Ghafouri-Fard S and Taheri M: Maternally
expressed gene 3 (MEG3): A tumor suppressor long non coding RNA.
Biomed Pharmacother. 118:1091292019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Lu KH, Li W, Liu XH, Sun M, Zhang ML, Wu
WQ, Xie WP and Hou YY: Long non-coding RNA MEG3 inhibits NSCLC
cells proliferation and induces apoptosis by affecting p53
expression. BMC Cancer. 13:4612013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Bergmann JH and Spector DL: Long
non-coding RNAs: Modulators of nuclear structure and function. Curr
Opin Cell Biol. 26:10–18. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Huang Y, Qiao Y, Zhao Y, Li Y, Yuan J,
Zhou J, Sun H and Wang H: Large scale RNA-binding proteins/LncRNAs
interaction analysis to uncover lncRNA nuclear localization
mechanisms. Brief Bioinform. 22:bbab1952021. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Wang N, Zhao Q, Huang Y, Wen C, Li Y, Bao
M and Wu L: Lnc-TMEM132D-AS1 as a potential therapeutic target for
acquired resistance to osimertinib in non-small-cell lung cancer.
Mol Omics. 19:238–251. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wang W, Lu Y, Qin GM, Ni LF, Xu BX, Liu
CF, Yu BF, Wang HL and Pang M: LncRNA RP11-297P16.4 promotes the
invasion and metastasis of non-small-cell lung carcinoma by
targeting the miR-145-5p/MMP-2/9 axis. Biomedicines. 13:6172025.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
de Visser KE and Joyce JA: The evolving
tumor microenvironment: From cancer initiation to metastatic
outgrowth. Cancer Cell. 41:374–403. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Li Z, Zhu T, Yao F, Shen X, Xu M, Fu L, Wu
Y, Ding J, Zhang J and Zhao J: LINC00665 promotes glycolysis in
lung adenocarcinoma cells via the let-7c-5p/HMMR axis. J Bioenerg
Biomembr. 56:181–191. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Hua Q, Jin M, Mi B, Xu F, Li T, Zhao L,
Liu J and Huang G: LINC01123, a c-Myc-activated long non-coding
RNA, promotes proliferation and aerobic glycolysis of non-small
cell lung cancer through miR-199a-5p/c-Myc axis. J Hematol Oncol.
12:912019. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Tang L, Wang S, Wang Y, Li K and Li Q:
LncRNA-UCA1 regulates lung adenocarcinoma progression through
competitive binding to miR-383. Cell Cycle. 22:213–228. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Liu Y, Liang L, Ji L, Zhang F, Chen D,
Duan S, Shen H, Liang Y and Chen Y: Potentiated lung adenocarcinoma
(LUAD) cell growth, migration and invasion by lncRNA DARS-AS1 via
miR-188-5p/KLF12 axis. Aging (Albany NY). 13:23376–23392. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Chen W, Li X, Du B, Cui Y, Ma Y and Li Y:
The long noncoding RNA HOXA11-AS promotes lung adenocarcinoma
proliferation and glycolysis via the microRNA-148b-3p/PKM2 axis.
Cancer Med. 12:4421–4433. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Wang Q, Zhang W, Yin D, Tang Z, Zhang E
and Wu W: Gene amplification-driven lncRNA SNHG6 promotes
tumorigenesis via epigenetically suppressing p27 expression and
regulating cell cycle in non-small cell lung cancer. Cell Death
Discov. 8:4852022. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zheng X, Zhang J, Fang T, Wang X, Wang S,
Ma Z, Xu Y, Han C, Sun M, Xu L, et al: The long non-coding RNA
PIK3CD-AS2 promotes lung adenocarcinoma progression via
YBX1-mediated suppression of p53 pathway. Oncogenesis. 9:342020.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Li M, Shi M, Hu C, Chen B and Li S: MALAT1
modulated FOXP3 ubiquitination then affected GINS1 transcription
and drived NSCLC proliferation. Oncogene. 40:3870–3884. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Han X, Jiang H, Qi J, Li J, Yang J, Tian
Y, Li W, Jing Q and Wang C: Novel lncRNA UPLA1 mediates
tumorigenesis and prognosis in lung adenocarcinoma. Cell Death Dis.
11:9992020. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Jin S, He J, Zhou Y, Wu D, Li J and Gao W:
LncRNA FTX activates FOXA2 expression to inhibit non-small-cell
lung cancer proliferation and metastasis. J Cell Mol Med.
24:4839–4849. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Cao G, Tan B, Wei S, Shen W, Wang X, Chu
Y, Rong T and Gao C: Down-regulation of MBNL1-AS1 contributes to
tumorigenesis of NSCLC via sponging miR-135a-5p. Biomed
Pharmacother. 125:1098562020. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Yang Q, Wang M, Xu J, Yu D, Li Y, Chen Y
and Zhang X, Zhang J, Gu J and Zhang X: LINC02159 promotes
non-small cell lung cancer progression via ALYREF/YAP1 signaling.
Mol Cancer. 22:1222023. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Zhu Q, Zhang C, Qu T, Lu X, He X, Li W,
Yin D, Han L, Guo R and Zhang E: MNX1-AS1 promotes phase separation
of IGF2BP1 to Drive c-Myc-mediated Cell-cycle progression and
proliferation in lung cancer. Cancer Res. 82:4340–4358. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Min S, Zhang L, Zhang L, Liu F and Liu M:
LncRNA MIR100HG affects the proliferation and metastasis of lung
cancer cells through mediating the microRNA-5590-3p/DCBLD2 axis.
Immun Inflamm Dis. 12:e12232024. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Hong J, Gu R, Cheng W, Lu C and Wang X:
LncRNA MACC1-AS1 facilitates the cell growth of small cell lung
cancer by sequestering miR-579-3p and mediating NOTCH1-pathway. Int
J Biol Macromol. 281:1365792024. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Ma J, Zhang P, Wang Y, Lu M, Cao K, Wei S,
Qi C, Ling X and Zhu J: LncRNA HAR1A inhibits non-small cell lung
cancer growth by downregulating c-MYC transcripts and facilitating
its proteasomal degradation. Int Immunopharmacol. 142:1132642024.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Zhai H, Xiang X, Pu J, Niu X, Gao J, Mu D,
Du J, Li Y, Qu L, Liu B, et al: LncRNA SLC7A11AR promotes lung
adenocarcinoma progression by inhibiting ferroptosis via promoting
SLC7A11 expression. Int J Biol Sci. 21:4549–4566. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Ke B, Zhong H, Gong Y, Chen X, Yan C and
Shi L: LINC00323 knockdown suppresses the proliferation, migration,
and vascular mimicry of non-small cell lung cancer cells by
promoting ubiquitinated degradation of AKAP1. Noncoding RNA Res.
11:131–140. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Wang RQ, Long XR, Zhou NN, Chen DN, Zhang
MY, Wen ZS, Zhang LJ, He FZ, Zhou ZL, Mai SJ, et al: Lnc-GAN1
expression is associated with good survival and suppresses tumor
progression by sponging mir-26a-5p to activate PTEN signaling in
non-small cell lung cancer. J Exp Clin Cancer Res. 40:92021.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Gerstberger S, Jiang Q and Ganesh K:
Metastasis. Cell. 186:1564–1579. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Li R, Lin Y, Hu F, Liao Y, Tang J, Shen Y,
Li H, Guo J and Xie L: LncRNA TEX41 regulates autophagy by
increasing Runx2 expression in lung adenocarcinoma bone metastasis.
Am J Transl Res. 15:949–966. 2023.PubMed/NCBI
|
|
63
|
Yang L, Liu Q, Zhang X, Liu X, Zhou B,
Chen J, Huang D, Li J, Li H, Chen F, et al: DNA of neutrophil
extracellular traps promotes cancer metastasis via CCDC25. Nature.
583:133–138. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Wang Y, Liu F, Chen L, Fang C, Li S, Yuan
S, Qian X, Yin Y, Yu B, Fu B, et al: Neutrophil Extracellular Traps
(NETs) promote non-small cell lung cancer metastasis by suppressing
lncRNA MIR503HG to activate the NF-κB/NLRP3 inflammasome pathway.
Front Immunol. 13:8675162022. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Huang Y, Hong W and Wei X: The molecular
mechanisms and therapeutic strategies of EMT in tumor progression
and metastasis. J Hematol Oncol. 15:1292022. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Pan J, Fang S, Tian H, Zhou C, Zhao X,
Tian H, He J, Shen W, Meng X, Jin X, et al: lncRNA
JPX/miR-33a-5p/Twist1 axis regulates tumorigenesis and metastasis
of lung cancer by activating Wnt/β-catenin signaling. Mol Cancer.
19:92020. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Zhong Y, Yang L, Xiong F, He Y, Tang Y,
Shi L, Fan S, Li Z, Zhang S, Gong Z, et al: Long non-coding RNA
AFAP1-AS1 accelerates lung cancer cells migration and invasion by
interacting with SNIP1 to upregulate c-Myc. Signal Transduct Target
Ther. 6:2402021. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Nie W, Hu MJ, Zhang Q, Lu J, Qian FF,
Zhang LL, Hu F, Li CH, Cao SH, Li JW, et al: DUBR suppresses
migration and invasion of human lung adenocarcinoma cells via
ZBTB11-mediated inhibition of oxidative phosphorylation. Acta
Pharmacol Sin. 43:157–166. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Chen Y, Shi H, Dong Y and Cui W: LncRNA
MSTO2P affects the proliferation, invasion and migration of
non-small cell lung cancer by regulating the Wnt/β-catenin pathway.
Discov Oncol. 16:1502025. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Ma Y, Zhou X, Yu M, Cheng X, Yang J, Ren
J, Zheng C, Li J, Qian X, Yi J, et al: SPAT inhibits LUAD
metastasis by targeting SF1-mediated splicing. Cell Death Dis.
16:5982025. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Chen J, Fan C, Song S and Zhang D: LncRNA
EP300-AS1 interacts with PTBP1 to destabilize PRMT5 mRNA and
suppresses NSCLC growth and metastasis. Cell Death Dis. 16:6072025.
View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Xu L, Kuang H, Peng H, Wu S, Bai Y, Jia X
and Yao W: A LINC00472-encoded polypeptide impedes migration and
proliferation through modulation of the HDAC2/SP1 axis in non-small
cell lung cancer cells. Cancer Cell Int. 25:2632025. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Wu Z, Zhao Y, Peng Y, Liu P, Huang Q, Wo
Y, Pan Y, Zheng D, Yuan C, Shang Y, et al: LncRNA TMEM99 Complexes
with IGF2BP2 to inhibit autophagy in lung adenocarcinoma. Adv Sci
(Weinh). 12:e078712025. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Fang F, Zhao M, Meng J, He J, Yang C, Wang
C, Wang J, Xie S, Jin X and Shi W: Upregulation of TTYH3 by lncRNA
LUCAT1 through interacting with ALYREF facilitates the metastasis
in non-small cell lung cancer. Cancer Biol Ther. 26:24649662025.
View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Liao Y, Wu X, Wu M, Fang Y, Li J and Tang
W: Non-coding RNAs in lung cancer: Emerging regulators of
angiogenesis. J Transl Med. 20:3492022. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Zhang Y, Liu J, Zou T, Qi Y, Yi B,
Dissanayaka WL and Zhang C: DPSCs treated by TGF-β1 regulate
angiogenic sprouting of three-dimensionally co-cultured HUVECs and
DPSCs through VEGF-Ang-Tie2 signaling. Stem Cell Res Ther.
12:2812021. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Cui Y, Liu H, Wang Z, Zhang H, Tian J,
Wang Z, Song W, Guo H, Liu L, Tian R, et al: Fructose promotes
angiogenesis by improving vascular endothelial cell function and
upregulating VEGF expression in cancer cells. J Exp Clin Cancer
Res. 42:1842023. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Chen J, Liu A, Wang Z, Wang B, Chai X, Lu
W, Cao T, Li R, Wu M, Lu Z, et al: LINC00173.v1 promotes
angiogenesis and progression of lung squamous cell carcinoma by
sponging miR-511-5p to regulate VEGFA expression. Mol Cancer.
19:982020. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Hou Y, Jia H, Cao Y, Zhang S, Zhang X, Wei
P, Xie J, Dong W and Wang B: LncRNA EPIC1 promotes tumor
angiogenesis via activating the Ang2/Tie2 axis in non-small cell
lung cancer. Life Sci. 267:1189332021. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Badowski C, He B and Garmire LX:
Blood-derived lncRNAs as biomarkers for cancer diagnosis: The good,
the bad and the beauty. NPJ Precis Oncol. 6:402022. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Yao X, Wang T, Sun MY, Yuming Y, Guixin D
and Liu J: Diagnostic value of lncRNA HOTAIR as a biomarker for
detecting and staging of non-small cell lung cancer. Biomarkers.
27:526–533. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Li C, Lv Y, Shao C, Chen C, Zhang T, Wei
Y, Fan H, Lv T, Liu H and Song Y: Tumor-derived exosomal lncRNA
GAS5 as a biomarker for early-stage non-small-cell lung cancer
diagnosis. J Cell Physiol. 234:20721–20727. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Tantai J, Hu D, Yang Y and Geng J:
Combined identification of long non-coding RNA XIST and HIF1A-AS1
in serum as an effective screening for non-small cell lung cancer.
Int J Clin Exp Pathol. 8:7887–7895. 2015.PubMed/NCBI
|
|
84
|
Min L, Zhu T, Lv B, An T, Zhang Q, Shang
Y, Yu Z, Zheng L and Wang Q: Exosomal LncRNA RP5-977B1 as a novel
minimally invasive biomarker for diagnosis and prognosis in
non-small cell lung cancer. Int J Clin Oncol. 27:1013–1024. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
He C, Huang D, Yang F, Huang D, Cao Y,
Peng J and Luo X: High expression of lncRNA HEIH is helpful in the
diagnosis of non-small cell lung cancer and predicts poor
prognosis. Cancer Manag Res. 14:503–514. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Wang J, Zhang L, Wang C, Chen Y and Sui X:
LINC00313/miR-4429 axis provides novel biomarkers for the diagnosis
and prognosis of non-small cell lung cancer. Acta Biochim Pol.
69:343–348. 2022.PubMed/NCBI
|
|
87
|
Yang Q, Kong S, Zheng M, Hong Y, Sun J,
Ming X, Gu Y, Shen X and Ju S: Long intergenic noncoding RNA
LINC00173 as a potential serum biomarker for diagnosis of
non-small-cell lung cancer. Cancer Biomark. 29:441–451. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Zhang X, Guo H, Bao Y, Yu H, Xie D and
Wang X: Exosomal long non-coding RNA DLX6-AS1 as a potential
diagnostic biomarker for non-small cell lung cancer. Oncol Lett.
18:5197–5204. 2019.PubMed/NCBI
|
|
89
|
Li W, Li W and Xu Z: LncRNA MIR99AHG
serves as a diagnostic marker for early lung cancer and regulates
cancer cell growth and metastasis via miR-194-5p. Discov Oncol.
16:10502025. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Zang X, Gu J, Zhang J, Shi H, Hou S, Xu X,
Chen Y, Zhang Y, Mao F, Qian H, et al: Exosome-transmitted lncRNA
UFC1 promotes non-small-cell lung cancer progression by
EZH2-mediated epigenetic silencing of PTEN expression. Cell Death
Dis. 11:2152020. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Mohamed Gamal El-Din G, Ibrahim FK,
Shehata HH, Osman NM, Abdel-Rahman OM and Ali M: Exosomal
expression of RAB27A and its related lncRNA Lnc-RNA-RP11-510M2 in
lung cancer. Arch Physiol Biochem. 128:1479–1485. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Wang L, Xie Y, Wang J, Zhang Y, Liu S,
Zhan Y, Zhao Y, Li J, Li P and Wang C: Characterization of a novel
LUCAT1/miR-4316/VEGF-A axis in metastasis and glycolysis of lung
adenocarcinoma. Front Cell Dev Biol. 10:8335792022. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Teng Y, Kang H and Chu Y: Identification
of an exosomal long noncoding RNA SOX2-OT in plasma as a promising
biomarker for lung squamous cell carcinoma. Genet Test Mol
Biomarkers. 23:235–240. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Li X, Wu Y and Jin Y: Exosomal LncRNAs and
CircRNAs in lung cancer: Emerging regulators and potential
therapeutic targets. Noncoding RNA Res. 9:1069–1079. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Cao Y, Liu X, Liu J, Su Z, Liu W, Yang L
and Zhang L: Diagnostic value of exosomal noncoding RNA in lung
cancer: A meta-analysis. Front Oncol. 14:13572482024. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Lin S, He C, Song L, Sun L, Zhao R, Min W
and Zhao Y: Exosomal lncCRLA is predictive for the evolvement and
development of lung adenocarcinoma. Cancer Lett. 582:2165882024.
View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Shi L, Li B, Zhang Y, Chen Y, Tan J, Chen
Y, Li J, Xiang M, Xing HR and Wang J: Exosomal lncRNA Mir100hg
derived from cancer stem cells enhance glycolysis and promote
metastasis of lung adenocarcinoma through mircroRNA-15a-5p/31-5p.
Cell Commun Signal. 21:2482023. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Mao G, Mu Z and Wu DA: Exosomal lncRNA
FOXD3-AS1 upregulates ELAVL1 expression and activates PI3K/Akt
pathway to enhance lung cancer cell proliferation, invasion, and
5-luorouracil resistance. Acta Biochim Biophys Sin (Shanghai).
53:1484–1494. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Saadh MJ, Abdulsahib WK, Daminov B,
Sanghvi G, Ballal S, Sharma RSK, Pathak PK, Shankhyan A, Kumar A,
Sead FF and Chaitanya MVNL: Unveiling the power of exosomal
non-coding RNAs: Key drivers of lung cancer progression in the
tumor microenvironment. Med Oncol. 42:3082025. View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Albain KS, Swann RS, Rusch VW, Turrisi AT
III, Shepherd FA, Smith C, Chen Y, Livingston RB, Feins RH, Gandara
DR, et al: Radiotherapy plus chemotherapy with or without surgical
resection for stage III non-small-cell lung cancer: A phase III
randomised controlled trial. Lancet. 374:379–386. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Ju ZS, Sun B, Bao D and Zhang XF: Effect
of lncRNA-BLACAT1 on drug resistance of non-small cell lung cancer
cells in DDP chemotherapy by regulating cyclin D1 expression. Eur
Rev Med Pharmacol Sci. 24:9465–9472. 2020.PubMed/NCBI
|
|
102
|
Huang N, Guo W, Ren K, Li W, Jiang Y, Sun
J, Dai W and Zhao W: LncRNA AFAP1-AS1 Supresses miR-139-5p and
promotes cell proliferation and chemotherapy resistance of
Non-small cell lung cancer by competitively upregulating RRM2.
Front Oncol. 9:11032019. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Lu G, Zhong H, Gao J and Zhang Y: Alginate
microspheres encapsulating hox transcript antisense RNA siRNA
regulate the Hedgehog-Gli1 pathway to alleviate epidermal growth
factor receptor tyrosine kinase inhibitors resistance. J Biomater
Appl. 38:877–889. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Gong N, Teng X, Li J and Liang XJ:
Antisense Oligonucleotide-conjugated nanostructure-targeting lncRNA
MALAT1 inhibits cancer metastasis. ACS Appl Mater Interfaces.
11:37–42. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Gutschner T, Hämmerle M, Eissmann M, Hsu
J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, et al:
The noncoding RNA MALAT1 is a critical regulator of the metastasis
phenotype of lung cancer cells. Cancer Res. 73:1180–1189. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Guo F, Cao Z, Guo H and Li S: The action
mechanism of lncRNA-HOTAIR on the drug resistance of non-small cell
lung cancer by regulating Wnt signaling pathway. Exp Ther Med.
15:4885–4889. 2018.PubMed/NCBI
|
|
107
|
Chery J: RNA therapeutics: RNAi and
antisense mechanisms and clinical applications. Postdoc J. 4:35–50.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Pedram Fatemi R, Salah-Uddin S, Modarresi
F, Khoury N, Wahlestedt C and Faghihi MA: Screening for
small-molecule modulators of long noncoding RNA-protein
interactions using AlphaScreen. J Biomol Screen. 20:1132–1141.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Tu H, Zhang Z, Li J, Shi S and Costa M:
Loss of MEG3 contributes to the enhanced migration and invasion in
arsenic-induced carcinogenesis through NQO1/FSCN1 pathway. Am J
Cancer Res. 13:2307–2322. 2023.PubMed/NCBI
|
|
110
|
You H, Fan M, Yin L, Na F and You L:
Enhanced multi-omics analysis reveals a lncRNA signature with 12
RNA modifications to predict tumor heterogeneity and potential
therapy in non-small cell lung cancer. Discov Oncol. 16:18732025.
View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Chen C, Liu WR, Zhang B, Zhang LM, Li CG,
Liu C, Zhang H, Huo YS, Ma YC, Tian PF, et al: LncRNA H19
downregulation confers erlotinib resistance through upregulation of
PKM2 and phosphorylation of AKT in EGFR-mutant lung cancers. Cancer
Lett. 486:58–70. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Liu X, Huang Z, Qian W, Zhang Q and Sun J:
Silence of lncRNA UCA1 rescues drug resistance of cisplatin to
non-small-cell lung cancer cells. J Cell Biochem. 120:9243–9249.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Li C, Fan K, Qu Y, Zhai W, Huang A, Sun X
and Xing S: Deregulation of UCA1 expression may be involved in the
development of chemoresistance to cisplatin in the treatment of
non-small-cell lung cancer via regulating the signaling pathway of
microRNA-495/NRF2. J Cell Physiol. 235:3721–3730. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Hua G, Zeng ZL, Shi YT, Chen W, He LF and
Zhao GF: LncRNA XIST contributes to cisplatin resistance of lung
cancer cells by promoting cellular glycolysis through Sponging
miR-101-3p. Pharmacology. 106:498–508. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Xu X, Zhou X, Chen Z, Gao C, Zhao L and
Cui Y: Silencing of lncRNA XIST inhibits non-small cell lung cancer
growth and promotes chemosensitivity to cisplatin. Aging (Albany
NY). 12:4711–4726. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Xu Y, Tan X, Yang Q, Fang Z and Chen W:
LncRNA HCG11 enhances the chemosensitivity of non-small cell lung
cancer cells to Gemcitabine via miR-17-5p/p21 axis. Expert Rev
Anticancer Ther. 24:81–93. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Sun Z, Mai H, Xue C, Fan Z, Li J, Chen H,
Huo N, Kang X, Tang C, Fang L, et al:
Hsa-LINC02418/mmu-4930573I07Rik regulated by METTL3 dictates
anti-PD-L1 immunotherapeutic efficacy via enhancement of
Trim21-mediated PD-L1 ubiquitination. J Immunother Cancer.
11:e0074152023. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Kathuria H, Millien G, McNally L, Gower
AC, Tagne JB, Cao Y and Ramirez MI: NKX2-1-AS1 negatively regulates
CD274/PD-L1, cell-cell interaction genes, and limits human lung
carcinoma cell migration. Sci Rep. 8:144182018. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Wu J, Li L, Zhang H, Zhao Y, Zhang H, Wu S
and Xu B: A risk model developed based on tumor microenvironment
predicts overall survival and associates with tumor immunity of
patients with lung adenocarcinoma. Oncogene. 40:4413–4424. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Wang LF, Wu LP and Wen JD: LncRNA
AC079630.4 expression associated with the progression and prognosis
in lung cancer. Aging (Albany NY). 13:18658–18668. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Chen X, Guo J, Zhou F, Ren W, Pu J, Mutti
L, Niu X and Jiang X: Over-Expression of Long Non-Coding
RNA-AC099850.3 correlates with tumor progression and poor prognosis
in lung adenocarcinoma. Front Oncol. 12:8957082022. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Liu C, Li X, Hao Y, Wang F, Cheng Z, Geng
H and Geng D: STAT1-induced upregulation of lncRNA KTN1-AS1
predicts poor prognosis and facilitates non-small cell lung cancer
progression via miR-23b/DEPDC1 axis. Aging (Albany NY).
12:8680–8701. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Song J, Sun Y, Cao H, Liu Z, Xi L, Dong C,
Yang R and Shi Y: A novel pyroptosis-related lncRNA signature for
prognostic prediction in patients with lung adenocarcinoma.
Bioengineered. 12:5932–5949. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Kumar KR, Cowley MJ and Davis RL:
Next-Generation sequencing and emerging technologies. Semin Thromb
Hemost. 50:1026–1038. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Qi JZ, Zhao TT and Qi SS: Functions of
long non-coding RNA LNC11649 in non-small cell lung cancer cells as
a reprocessed form of MALAT1. Neoplasma. 69:1322–1337. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Chen L, Wang L, Xiong Z, Zhu X and Chen L:
Exploring the prognostic significance and immunotherapeutic
potential of Single-Cell Sequencing-Identified long noncoding RNA
(LncRNA) in patients With Non-small cell lung cancer. Cureus.
15:e484362023.PubMed/NCBI
|
|
127
|
Chen X and Zhang L: Integrative analysis
revealed LINC00847 as a potential target of tumor immunotherapy.
Appl Biochem Biotechnol. 195:6345–6358. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Li Z, Pan C, Wang Z, Deng X, Zhu Q, Wu W
and Chen L: LncRNA PCBP1-AS1 correlated with the functional states
of cancer cells and inhibited lung adenocarcinoma metastasis by
suppressing the EMT progression. Carcinogenesis. 42:931–939. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Zhao Y, Zhou C, Zuo L, Yan H, Gu Y, Liu H,
Yu G and Zhou X: Identification of cancer cell-intrinsic biomarkers
associated with tumor progression and characterization of SFTA3 as
a tumor suppressor in lung adenocarcinomas. BMC Cancer. 25:362025.
View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Gautier-Isola M, Lopes Goncalves R, Truchi
M, Lacoux C, Scribe C, Cadis H, Guardini L, Bekisz S, Ilié M,
Hofman P, et al: LINC01116, a hypoxia-lncRNA marker of pathological
lymphangiogenesis and poor prognosis in lung adenocarcinoma. Mol
Oncol. Dec 9–2025.doi: 10.1002/1878-0261.70175 (Epub ahead of
print). View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Huang Y, Zhu S, Yao S, Zhai H, Liu C and
Han JJ: Unraveling aging from transcriptomics. Trends Genet.
41:218–235. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Zhu J, Fan Y, Xiong Y, Wang W, Chen J, Xia
Y, Lei J, Gong L, Sun S and Jiang T: Delineating the dynamic
evolution from preneoplasia to invasive lung adenocarcinoma by
integrating single-cell RNA sequencing and spatial transcriptomics.
Exp Mol Med. 54:2060–2076. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Guo Q, Liu Q, He D, Xin M, Dai Y, Sun R,
Li H, Zhang Y, Li J, Kong C, et al: LnCeCell 2.0: An updated
resource for lncRNA-associated ceRNA networks and web tools based
on single-cell and spatial transcriptomics sequencing data. Nucleic
Acids Res. 53:D107–D115. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Wu Y, Wang Y, Wei M, Han X, Xu T and Cui
M: Advances in the study of exosomal lncRNAs in tumors and the
selection of research methods. Biomed Pharmacother. 123:1097162020.
View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Tang P, Sun D, Xu W, Li H and Chen L: Long
non-coding RNAs as potential therapeutic targets in non-small cell
lung cancer (Review). Int J Mol Med. 52:682023. View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Mishra A and Mishra S:
Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1)
lncRNA conformational dynamics in complex with RNA-Binding Protein
with Serine-Rich domain 1 (RNPS1) in the Pan-cancer splicing and
gene expression. ACS Omega. 9:42212–42226. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Monroy-Eklund A, Taylor C, Weidmann CA,
Burch C and Laederach A: Structural analysis of MALAT1 long
noncoding RNA in cells and in evolution. Rna. 29:691–704. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Vannini I, Ferracin M, Fabbri F and Fabbri
M: Overexpression of ultraconserved region 83-induces lung cancer
tumorigenesis. PLoS One. 17:e02614642022. View Article : Google Scholar : PubMed/NCBI
|
|
139
|
Statello L, Ali MM and Kanduri C: In vivo
administration of therapeutic antisense oligonucleotides. Methods
Mol Biol. 2254:273–282. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
140
|
Chen Y, Li Z, Chen X and Zhang S: Long
non-coding RNAs: From disease code to drug role. Acta Pharm Sin B.
11:340–354. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
141
|
Li Y, Lin M, Wang S, Cao B, Li C and Li G:
Novel angiogenic regulators and Anti-Angiogenesis drugs targeting
angiogenesis signaling pathways: Perspectives for targeting
angiogenesis in lung cancer. Front Oncol. 12:8429602022. View Article : Google Scholar : PubMed/NCBI
|
|
142
|
Zeng L, Wen H, Jiao T, Zhang Q, Zhao X,
Zhang L and Yang S: LINC02178 drives lung adenocarcinoma
progression and serves as a therapeutic target for
nanodelivery-based intervention. Sci Rep. 15:393442025. View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Wang S, Han C, Liu T, Ma Z, Qiu M, Wang J,
You Q, Zheng X, Xu W, Xia W, et al: FAM83H-AS1 is a noncoding
oncogenic driver and therapeutic target of lung adenocarcinoma.
Clin Transl Med. 11:e3162021. View Article : Google Scholar : PubMed/NCBI
|
|
144
|
Alhajlah S: The molecular mechanisms of
various long non-coding RNA (lncRNA) in human lung tumors: Shedding
light on the molecular mechanisms. Pathol Res Pract.
256:1552532024. View Article : Google Scholar : PubMed/NCBI
|
|
145
|
Liu Z, Jiang H, Zhao R, Quan Q and Huang
X: Long Noncoding RNA IPW is a novel diagnostic and predictive
biomarker in lung adenocarcinoma. Genet Test Mol Biomarkers.
27:18–26. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
146
|
Zhang X, Ma L and Wang J: Cross-regulation
between redox and epigenetic systems in tumorigenesis: Molecular
mechanisms and clinical applications. Antioxid Redox Signal.
39:445–471. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
147
|
Tian Y, Li S, Song J, Ji T, Zhu M,
Anderson GJ, Wei J and Nie G: A doxorubicin delivery platform using
engineered natural membrane vesicle exosomes for targeted tumor
therapy. Biomaterials. 35:2383–2390. 2014. View Article : Google Scholar : PubMed/NCBI
|