Introduction
Skin cutaneous melanoma (SKCM) is a malignant form
of skin cancer with an increasing incidence and mortality rate,
presenting notable challenges to patient health and survival
(1). Despite the progress made in
targeted therapies and immunotherapy against SKCM, numerous
patients develop resistance or exhibit only partial responses,
resulting in suboptimal therapeutic outcomes (2). Consequently, further investigation
into the molecular mechanisms driving melanoma progression is
required. Identifying novel therapeutic targets and reliable
biomarkers for SKCM is important for improving treatment strategies
and patient prognosis.
Signal transducer and activator of transcription
(STAT)1, a central component of the Janus kinase (JAK)-STAT
signaling pathway, regulates the expression of numerous downstream
genes involved in important cellular functions, including
proliferation, differentiation, apoptosis and immune response
modulation (3,4). In the context of cancer, STAT1 is
often described as a double-edged sword, exhibiting both
tumor-suppressing and tumor-promoting properties. For example, in
breast and colorectal cancers, STAT1 generally acts as a tumor
suppressor by inducing apoptosis and enhancing immune responses to
limit tumor progression (5).
Clinical evidence suggests that STAT1 expression is markedly
elevated in melanoma tissues compared with healthy skin, with this
upregulation associated with poor clinical outcomes (6). In line with these observations, an
in vitro study showed that silencing STAT1 reduces melanoma
cell proliferation and migration while promoting apoptosis,
underscoring its oncogenic role in melanoma (7). However, the downstream molecular
mechanisms regulated by STAT1 remain to be fully elucidated, and
the incompletely resolved interactions of STAT1 with other notable
molecular targets warrant further exploration.
Tubulin β4A (TUBB4A), a member of the tubulin
β-chain family, encodes β-tubulin, a notable protein involved in
maintaining cellular integrity and supporting important cellular
processes such as cell division and migration (8). Elevated expression of TUBB4A has been
reported in various cancers, including glioblastoma, breast cancer
and lung cancer, with its upregulation being associated with
enhanced tumor invasiveness and resistance to therapeutic
interventions (9,10). However, the precise role of TUBB4A
and its regulatory mechanisms in melanoma remain largely
unexplored. The JAK-STAT signaling pathway is important for tumor
initiation and progression, with STAT1 and STAT3 acting as central
regulators of downstream gene expression. These factors modulate
important cellular processes in cancer, including proliferation,
apoptosis and immune evasion (11–13).
In melanoma, aberrant STAT1 activation may interact with molecules
such as TUBB4A, thereby enhancing tumor invasiveness and conferring
resistance to chemotherapy (14,15).
As a well-established oncogene, STAT3 promotes tumor growth and
metastasis by modulating cytokine expression and interacting with
other transcription factors (16,17).
The interplay between STAT1 and STAT3 in melanoma is multifaceted,
characterized by both synergistic and antagonistic interactions
that necessitate further investigation (6,18,19).
Although TUBB4A upregulation has been implicated in
tumor progression and poor prognosis across various cancers, its
exact role in melanoma remains poorly defined. Evidence suggests
that TUBB4A is important for maintaining microtubule stability, a
function necessary for processes such as tumor cell mitosis
(20,21), migration (22) and invasion (10). Furthermore, TUBB4A appears to
contribute to chemotherapy resistance, with its elevated expression
linked to drug resistance in cancers, including breast cancer and
non-small cell lung cancer, particularly to agents such as
paclitaxel (23,24). Similar mechanisms may be at play in
melanoma, although tumor-specific factors are likely to influence
these processes. Understanding how STAT1 influences TUBB4A
expression is important for elucidating melanoma pathogenesis and
identifying potential therapeutic targets for SKCM.
The present study aimed to assess the roles of STAT1
and TUBB4A in melanoma cell proliferation, migration and apoptosis,
using both in vitro and in vivo models. Additionally,
the impact of these molecules on tumor growth were evaluated in a
mouse xenograft model, providing a comprehensive analysis of the
functional and mechanistic contributions of the STAT1-TUBB4A
pathway to melanoma. With the increasing use of targeted therapies
and immunotherapies (25,26), the discovery of novel biomarkers
and treatment strategies for SKCM is important for improving
patient outcomes. The present research endeavored to clarify the
STAT1-TUBB4A axis, laying the groundwork for the development of
more personalized and effective therapeutic approaches for patients
with melanoma.
Materials and methods
Bioinformatics analysis
RNA sequencing data for TUBB4A and STAT1 expression
across various cancer types were obtained from the Gene Expression
Profiling Interactive Analysis 2 (GEPIA2) database (http://gepia2.cancer-pku.cn), which processes data
from The Cancer Genome Atlas (TCGA) and the Genotype-Tissue
Expression (GTEx) project (search term, SKCM | Skin Cutaneous
Melanoma). The analysis was conducted in March 2024.
For the specific analysis of SKCM, the TCGA-SKCM
tumor dataset (n=461) was compared against the GTEx normal skin
tissue dataset (n=558). Differential expression analysis was
performed using the following parameters: A |log2fold
change (FC)| cut-off value of 1 and a P-value cut-off value of
0.05. Gene expression levels for TUBB4A and STAT1 were quantified
as transcripts per million (TPM).
To identify potential binding sites of the
transcription factor STAT1 within the regulatory region of the
TUBB4A gene promoter, bioinformatics analysis was performed using
the JASPAR database (https://jaspar.elixir.no/). First, the experimentally
validated canonical position weight matrix for STAT1 was retrieved
from JASPAR, while the promoter sequence spanning approximately
from −2,000 bp to + 500 bp relative to the transcription start site
of TUBB4A was extracted from a genomic database. Subsequently, the
‘Scan’ function in JASPAR was used to scan this promoter sequence
for STAT1 binding motifs, with a threshold set at an 85% relative
score. The analysis finally generated a prediction list containing
specific coordinates, match scores, and statistical significance of
the identified sites, thereby providing candidate targets for
subsequent experimental validation.
Patient samples and clinical data
A total of 31 SKCM tissue samples and matching
normal tissue samples were collected from patients at Pudong New
Area People's Hospital (Shanghai, China), with institutional review
board approval (approval no. CEA-2023-16) and in compliance with
ethical guidelines. The patient cohort consisted of 18 males and 13
females, with a median age of 62 years and an age range of 38–75
years. All patients provided informed consent for tissue use.
Melanoma samples were obtained from patients undergoing surgical
resection of primary tumors, and matching normal tissue was
collected from adjacent healthy skin during the same procedure.
Specimens were immediately snap-frozen in liquid nitrogen and
stored at −80°C until RNA extraction. Normal tissue was taken at
least 2 cm from the tumor to avoid potential contamination.
Cell culture and treatment
The melanoma cell lines A2058 (cat. no. CRL-3601),
SK-MEL-1 (cat. no. HTB-67), A375 (cat. no. CRL-1619) and RPMI-7951
(cat. no. HTB-66), as well as normal human epidermal melanocytes
(HEM; cat. no. PCS-200-013) and 293T cells (cat. no. CRL-3216) were
obtained from American Type Culture Collection. Cells were cultured
in DMEM (cat. no. 11995-065; Thermo Fisher Scientific, Inc.)
supplemented with 10% fetal bovine serum (FBS; cat. no. 16000-044;
Gibco; Thermo Fisher Scientific, Inc.) at 37°C in a humidified
atmosphere containing 5% CO2. Cells were seeded into
6-well plates and grown to 60–70% confluence. Transfection was
performed using Lipofectamine® 3000 (cat. no. L3000015,
Thermo Fisher Scientific, Inc.) according to the manufacturer's
protocol. Cells were transfected with small interfering RNA (siRNA)
targeting STAT1 (si-STAT1; cat. no. A09009) or a scrambled negative
control (si-NC; cat. no. A09010; GenePharma, Shanghai) at a final
concentration of 50 nM. The cells were incubated with the
transfection complexes at 37°C for 48 h prior to subsequent
experimentation. The sequences of siRNAs are listed in Table SI. Lentiviral Vector Generation
and Transduction Stable TUBB4A overexpression was achieved using a
lentiviral system. The lentiviral vector encoding the TUBB4A gene
(Ov-TUBB4A, pReceiver-Lv105) and the negative control vector
lacking the insert (Ov-NC; pReceiver-Lv105 Empty Vector) were
purchased from GeneCopoeia, Inc. Lentiviral particles were
generated using a 3rd generation packaging system in 293T cells
(ATCC). Briefly, HEK293T cells were co-transfected with 2.5 µg of
the lentiviral plasmid, along with packaging and envelope plasmids
(Lenti-Pac™ HIV Expression Packaging Kit; GeneCopoeia, Inc.) using
the EndoFectin™ Lenti transfection reagent (cat. no. EF002;
GeneCopoeia, Inc.). The ratio of the target plasmid to the
packaging mix was maintained at 1:1. Lentiviral supernatants were
collected at 48 and 72 h post-transfection and filtered through a
0.45-µm filter. Melanoma cells were infected with the viral
particles at a multiplicity of infection (MOI) of 10 in the
presence of 8 µg/ml Polybrene (cat. no. TR-1003; Sigma-Aldrich;
Merck KGaA). The transduction duration was 24 h, after which the
medium was replaced with fresh complete medium. Selection of stably
transfected cells was performed 48 h post-transduction using 2
µg/ml puromycin (cat. no. P9620; Sigma-Aldrich). At 24 h after the
transfection, the mRNA levels of STAT1 and TUBB4A were measured
using reverse transcription-quantitative (RT-q)PCR.
RT-qPCR
Total RNA was acquired from cellular and tumor
specimens through the TRIzol method (Invitrogen; Thermo Fisher
Scientific, Inc.). RT-qPCR was performed using a SuperScript™ III
Platinum™ One Step RT-qPCR Kit (cat. no. 11732088; Thermo Fisher
Scientific, Inc.). The thermocycling conditions were as follows:
cDNA synthesis at 50°C for 15 min; initial denaturation at 95°C for
2 min, followed by 40 cycles of denaturation at 95°C for 15 sec and
annealing/extension at 60°C for 30 sec. Relative transcript
abundances were determined utilizing the 2−ΔΔCq
algorithm (27). The primers were
as follows: STAT1: Forward, 5′-GACCGCACCTTCAGTCTTTTC-3′ and
reverse, 5′-TCATTCACATCTCTCAACTTCACA-3′; TUBB4A: Forward:
5′-GCTGCGGACCGAGAAACT-3′, Reverse: 5′-TCCAGTTGCAGGTCACTGTC-3′;
GAPDH: Forward: 5′-CCACTCCTCCACCTTTGAC-3′ and reverse:
5′-ACCCTGTTGCTGTAGCCA-3′.
Cell Counting Kit-8 (CCK-8)
Cell viability was assessed using a CCK-8 assay
(cat. no. CK04; Dojindo Laboratories, Inc.). Following treatment
with siRNA or lentiviral vectors, cells were seeded in triplicate
into 96-well culture plates (cat. no. 3599; Corning, Inc.) at a
density of 5×103 cells/well and allowed to adhere for 24
h. The medium was subsequently replaced with 100 µl of fresh growth
DMEM supplemented with FBS as aforementioned and 10 µl of CCK-8
reagent was added to each well. The plates were incubated with
CCK-8 reagent at 37°C for 1–4 h, depending on the cell type, to
allow for the conversion of the tetrazolium salt into a formazan
product by metabolically active cells. Absorbance was measured at
450 nm using a BioTek 800 TS Absorbance Reader (cat. no. ELx800;
BioTek; Agilent Technologies, Inc.).
Colony formation assays
Melanoma cells (A375 and RPMI-7951) were seeded into
6-well plates (cat. no. 3516; Corning, Inc.) at a density of
500–1,000 cells per well and allowed to adhere for 24 h. Cells were
then transfected with si-NC (cat. no. A09010) or si-STAT1 (cat. no.
A09009; GenePharma), as well as transfected using a lentiviral
vector for TUBB4A overexpression (Ov-TUBB4A; pReceiver-Lv105;
GeneCopoeia, Inc.) or Ov-NC (pReceiver-Lv105 Empty Vector;
GeneCopoeia, Inc.). After 10–14 days incubation, colonies were
fixed with 4% paraformaldehyde (cat. no. J19943.K2; Thermo Fisher
Scientific, Inc.) for 15 min, stained with 0.5% crystal violet
solution (cat. no. C3886; MilliporeSigma; Merck KGaA) for 30 min
and washed with phosphate-buffered saline (PBS; cat. no. 10010-023;
Thermo Fisher Scientific, Inc.). Colonies, defined as clusters of
≥50 cells, were manually counted under a microscope (Nikon
Corporation) at ×20 magnification. The results were normalized to
the control group.
Flow cytometry for apoptosis
assay
Briefly, melanoma cells (A375 and RPMI-7951) were
seeded into 6-well plates and treated with siRNA or lentiviral
vectors as aforementioned. To induce apoptosis, cells were exposed
to 100 nM cisplatin (cat. no. P4394; Sigma-Aldrich) for 24 h at
37°C. Following treatment, cells were harvested, washed with cold
PBS and resuspended in Annexin V binding buffer at a concentration
of 1×106 cells/ml. Cells were then stained with 5 µl
Annexin V-FITC and 5 µl PI for 15 min at room temperature in the
dark. The cell suspension was filtered through a 40-µm nylon mesh
to remove large aggregates. Flow cytometric analysis was
immediately performed using a BD FACSCalibur™ (BD Biosciences) flow
cytometer, and data were processed using FlowJo software (version
10.8.1, BD Biosciences). A sequential gating strategy was employed:
The primary cell population was first identified on an forward
scatter area vs. side scatter area dot plot while excluding debris.
Subsequently, single cells were identified using forward scatter
height vs. forward scatter area density plots to exclude doublets
and aggregates. Apoptosis analysis was conducted on this gated
singlet population. All flow cytometry data were analyzed using a
standardized gating strategy with gating parameters maintained
consistently across all experimental groups. The proportion of
cells in early apoptosis (Annexin V-positive, PI-negative), late
apoptosis and necrosis (Annexin V-positive, PI-positive) and live
cells (Annexin V-negative, PI-negative) were determined.
Cell migration assay
Cell migration was evaluated using Transwell inserts
with 8 µm pore chambers (cat. no. 3422; Corning, Inc.). Cells (A375
and RPMI-7951) were seeded in serum-free DMEM (cat. no. 11995-065;
Thermo Fisher Scientific, Inc.) in the upper chamber at a density
of 5×104 cells/well, while DMEM supplemented with 10%
FBS (cat. no. 16000-044; Gibco; Thermo Fisher Scientific, Inc.) was
used in the lower chamber. After 24 h-incubation at 37°C, migrated
cells were fixed with 4% paraformaldehyde (cat. no. J19943.K2;
Thermo Fisher Scientific, Inc.) for 15 min at 25°C, stained with
0.1% crystal violet solution (cat. no. C3886; MilliporeSigma; Merck
KGaA) for 30 min at 25°C and washed with PBS (cat. no. 10010-023;
Thermo Fisher Scientific, Inc.). Migrated cells were visualized
under an inverted light microscope (Nikon Corporation) for
analysis. The number of migrated cells was quantified using ImageJ
software (Version 1.53e, National Institutes of Health).
Western blot analysis
A375 and RPMI-7951 cells were harvested, and
proteins were extracted using RIPA buffer (cat. no. 89900; Thermo
Fisher Scientific, Inc.) containing protease inhibitors (cat. no.
11836170001; Roche Diagnostics, Ltd.). Protein concentrations were
quantified using the Pierce™ BCA Protein Assay Kit (cat. no. 23225;
Thermo Fisher Scientific, Inc.). A total of 20 µg protein in each
lane was separated by 10% SDS-PAGE and transferred to PVDF
membranes (cat. no. IPVH00010; MilliporeSigma; Merck KGaA).
Membranes were blocked with 5% non-fat milk (cat. no. 1706404;
Bio-Rad Laboratories, Inc.) in TBST with 0.1% Tween 20 for 1 h at
25°C and incubated overnight at 4°C with primary antibodies against
STAT1 (1:1,000; cat. no. 14994; Cell Signaling Technology, Inc.)
and TUBB4A (1:1,000; cat. no. ab11315; Abcam), as well as GAPDH
(1:5,000; cat. no. 2118; Cell Signaling Technology, Inc.) serving
as a loading control. After washing three times with TBST,
membranes were incubated with HRP-conjugated goat anti-rabbit IgG
(cat. no. 7074; Cell Signaling Technology, Inc.) and horse
anti-mouse IgG (both 1:2,000, cat. no. 7076; Cell Signaling
Technology, Inc.) secondary antibodies at room temperature for 1 h.
Protein bands were visualized using SuperSignal™ West Pico PLUS
Chemiluminescent Substrate (cat. no. 34095; Thermo Fisher
Scientific, Inc.). The band intensity was semi-quantified using
ImageJ software (version 1.53e, National Institutes of Health).
Luciferase reporter assay
The promoter regions of TUBB4A were cloned into the
firefly luciferase reporter plasmid pGL3-Basic vector (cat. no.
E1741; Promega Corporation) and transfected into 293T cells. The
cells were transiently co-transfected with the reporter constructs,
the internal control Renilla luciferase vector (cat. no. E2261;
Promega Corporation), and si-STAT1 (cat. no. A09009; GenePharma,
Shanghai) as aforementioned. STAT1 knockdown was performed using
si-STAT1 (cat. no. A09009; GenePharma) to evaluate its effect on
TUBB4A promoter activity. After 48 h, both Firefly and
Renilla luciferase activities were measured using the
Dual-Luciferase Reporter Assay System (cat. no. E1910; Promega
Corporation).
Chromatin immunoprecipitation
(ChIP)
ChIP analysis was performed to investigate
endogenous STAT1 binding to the TUBB4A promoter. A375 and RPMI-7951
cells were cross-linked with 1% formaldehyde (cat. no. 28908;
Thermo Fisher Scientific, Inc.) and lysed using SDS Lysis Buffer
(cat. no. 20-163; Sigma-Aldrich; Merck KGaA) and chromatin was
fragmented by sonication using a Bioruptor® PICO (cat.
no. B01080010 Diagenode SA). For each reaction, 100 µg chromatin
lysate was diluted to a final volume of 500 µl with ChIP dilution
buffer and incubated with 5 µg of anti-STAT1 antibodies (1:100;
cat. no. 14994; Cell Signaling Technology, Inc.) or control rabbit
IgG (1:100; cat. no. 2729; Cell Signaling Technology, Inc.)
overnight at 4°C. Subsequently, ChIP-Grade Protein A/G Magnetic
Beads (cat. no. 16-663; Sigma-Aldrich; Merck KGaA) were added
according to the manufacturer's protocol. The immune complexes were
washed sequentially with low and high salt buffer. The complexes
were isolated by magnetic separation. DNA-protein complex
crosslinking was reversed using proteinase K (cat. no. EO0491;
Thermo Fisher Scientific, Inc.), and DNA was purified using the
Aquadien DNA Purification kit (cat. no. 3578121; Bio-Rad
Laboratories, Inc.). Quantitative PCR (qPCR) amplification of the
TUBB4A promoter region was performed using SYBR® Green
JumpStart™ Taq ReadyMix™ (cat. no. S4438; Sigma-Aldrich) on a CFX96
real-time PCR instrument (Bio-Rad). The thermocycling conditions
were: 50°C for 10 min, 95°C for 5 min (1×), 95°C for 10 sec/61°C
for 15 sec/72°C for 30 sec (40×). The specific primer sequences for
the TUBB4A promoter were: Forward
5′-CAGTGACCTGCAACTGGAGA-3′, reverse 5′-GATTGGCCAAACACGAAGTT-3′.
Data were quantified using the %input method, where Input DNA
(non-immunoprecipitated chromatin) served as the normalization
control.
Mouse xenograft model
A total of 25 BALB/c nude mice (male; age, 8 weeks
old, 23–25 g) were subcutaneously injected in the axillary region
with 2×106 A375 cells treated with si-STAT1 and
Ov-TUBB4A, either separately or in combination, with 5 mice in each
treatment group. BALB/c nude mice were purchased from Shanghai
Bikai Keyi Biotechnology Co., Ltd (Shanghai, China). Mice were
housed at a temperature of 22±2°C, humidity of 55±5%, with a 12 h
light/dark cycle. Food and water were provided ad libitum.
Tumor volumes were measured every 3 days using calipers, and at the
conclusion of the study, tumors were excised, weighed and
photographed. At the experimental endpoint (five weeks after A375
cell injection), mice were sacrificed by overdose with sodium
pentobarbital (150 mg/kg), with cervical dislocation . Humane
endpoints for mice were as follows: i) Tumor size >1,500
mm3; ii) ulceration, necrosis or infection were
observed, particularly where the tumor broke the skin or became
necrotic; iii) mice exhibited impaired mobility or interfered with
normal functions, as indicated by the tumor location interfering
with walking, feeding or drinking; iv) mice demonstrated signs of
pain or distress, including hunched posture, lethargy, ruffled fur
or abnormal vocalization; v) mice exhibited significant body weight
loss, with a limit of >20% from a healthy control group adhered
to; and vi) tumor growth interfered with vital organs and their
functioning. No mice reached these experimental endpoints. Death
was confirmed by the absence of a heartbeat and respiration for
>5 min. All animal experiments were approved by the
Institutional Animal Care and Use Committee of Pudong New Area
People's Hospital (approval no. AEA-2024-33) and were conducted in
accordance with ethical guidelines.
Statistical analysis
Data were presented as mean ± standard deviation
from at least three independent experiments. Comparisons between
groups were performed using paired Student's t-test or one-way
ANOVA followed by Tukey's honestly significant difference post hoc
test. Correlations were analyzed using Pearson's coefficient.
Survival analysis was conducted using Kaplan-Meier curves and the
log-rank test, with statistical significance set at P<0.05.
Results
Expression and prognostic significance
of TUBB4A in SKCM
Following queries concerning RNA sequencing data for
TUBB4A and STAT1 expression in cancers, the GEPIA2 database
revealed notably elevated TUBB4A expression in multiple tumor
tissues compared with normal tissues, with data quantified in TPM.
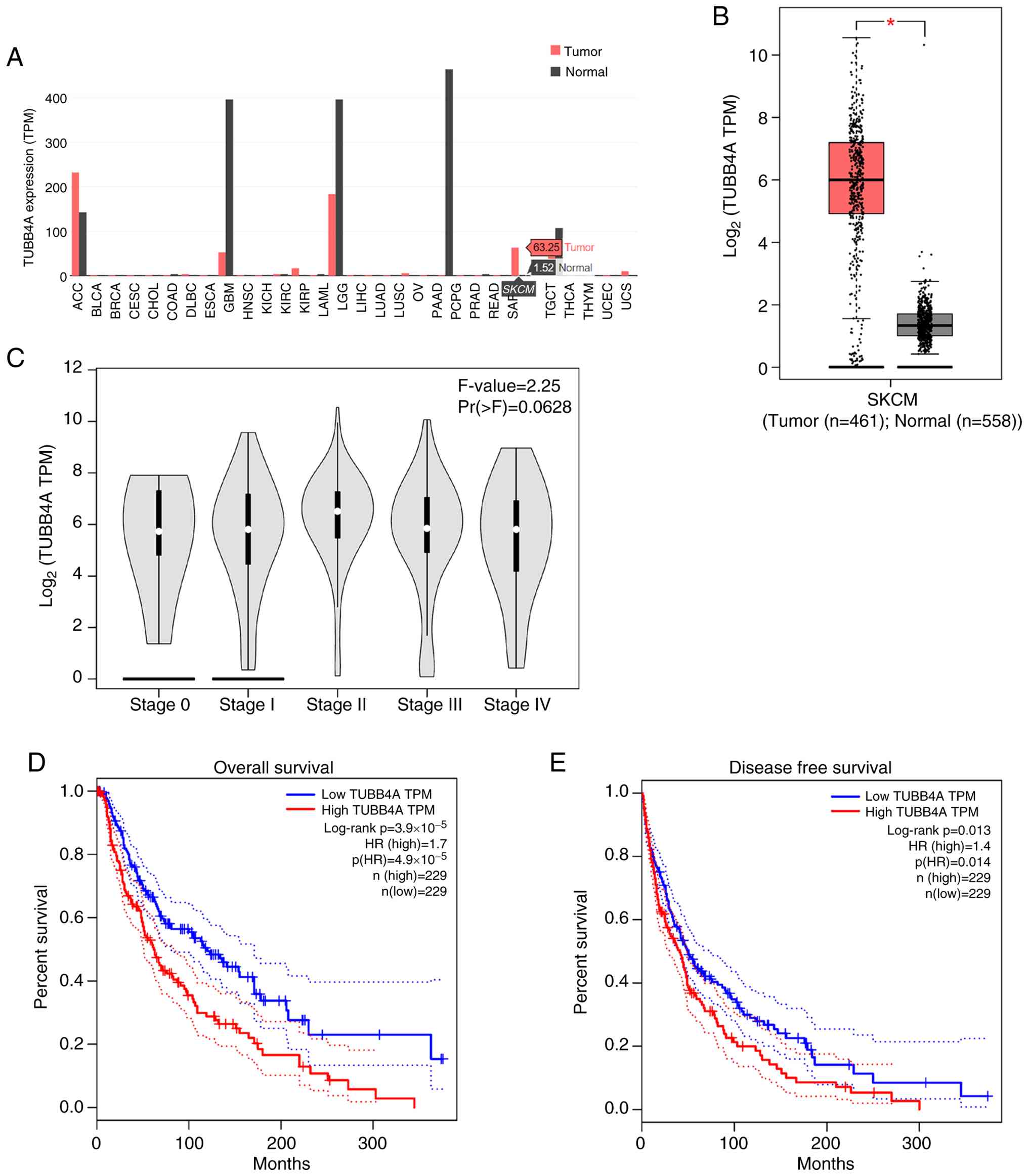
Specifically, in SKCM, TUBB4A expression was notably higher in
tumor samples (TPM, 63.25) than in normal tissues (TPM, 1.52)
(Fig. 1A). Box plot analysis
further supported that tumor samples (n=461) exhibited
significantly higher TUBB4A expression than normal samples (n=558)
(P<0.05; Fig. 1B), suggesting
that TUBB4A played a notable role in SKCM pathogenesis. Examination
of TUBB4A expression across pathological stages 0-IV of SKCM
(28) using a violin plot revealed
no statistically significant differences between stages (F=2.25;
P=0.0628; Fig. 1C). Kaplan-Meier
survival analysis demonstrated that elevated TUBB4A expression
(n=229) was strongly associated with reduced overall survival (OS)
compared with low expression (n=229), with a log-rank P-value of
3.9×10−5 and a hazard ratio (HR) of 1.7 [p(hazard ratio,
HR)=4.9×10−5; Fig. 1D].
Additionally, high TUBB4A expression associated with significantly
poorer disease-free survival (DFS), with a log-rank P-value of
0.013 and an HR of 1.4 (p(HR)=0.014; Fig. 1E). These results underscored the
negative prognostic value of increased TUBB4A expression for both
OS and DFS in patients with SKCM.
Correlation between STAT1 and TUBB4A
expression in SKCM
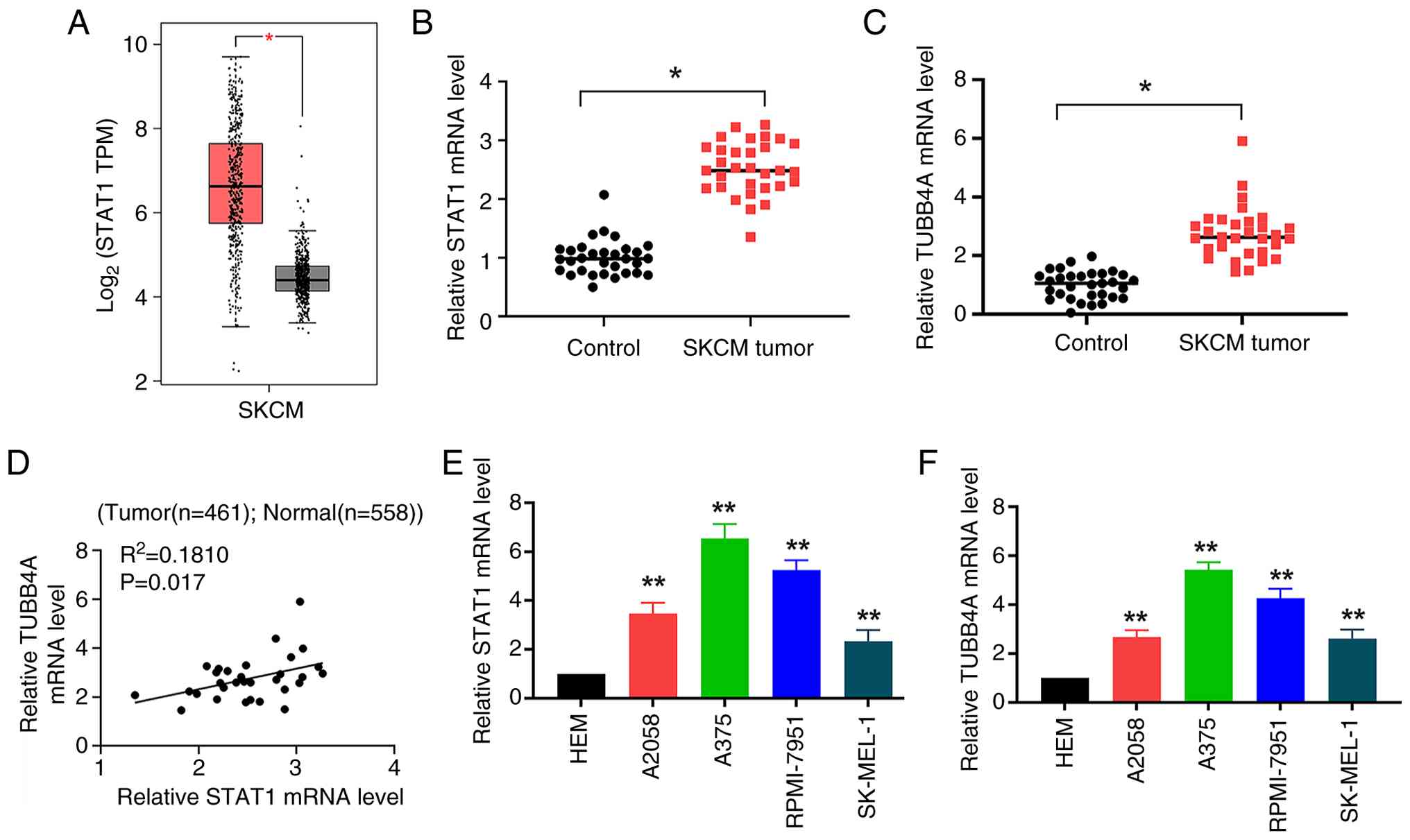
The expression levels of STAT1 and TUBB4A in SKCM
tissues were analyzed. Box plot analysis revealed significantly
elevated STAT1 expression in cancerous tissues (n=461) compared
with normal tissues (n=558) (P<0.05; Fig. 2A). STAT1 mRNA was also
significantly upregulated in SKCM tumor tissues (n=31) compared
with control tissues (n=31; P<0.05; Fig. 2B). Similarly, TUBB4A expression was
significantly higher in SKCM tumor samples than in control tissue
(P<0.05; Fig. 2C), indicating
that both STAT1 and TUBB4A were upregulated in melanoma. Pearson's
correlation analysis revealed a weak positive correlation between
the mRNA expression of STAT1 and TUBB4A in SKCM (|R=0.4254|,
R2=0.1810; P=0.017; Fig.
2D), suggesting that STAT1 may have regulated TUBB4A
expression. Further examination of STAT1 and TUBB4A levels in SKCM
cell lines demonstrated significantly elevated STAT1 mRNA levels in
the melanoma cell lines A2058, A375, RPMI-7951 and SK-MEL-1
compared with normal HEM (P<0.01; Fig. 2E), with A375 cells showing the
highest STAT1 expression. Similarly, TUBB4A expression was
significantly elevated in all melanoma cell lines relative to HEM
(P<0.01), with the highest expression once again observed in
A375 cells (Fig. 2F).
Silencing STAT1 markedly decreases the
viability and proliferation of melanoma cells
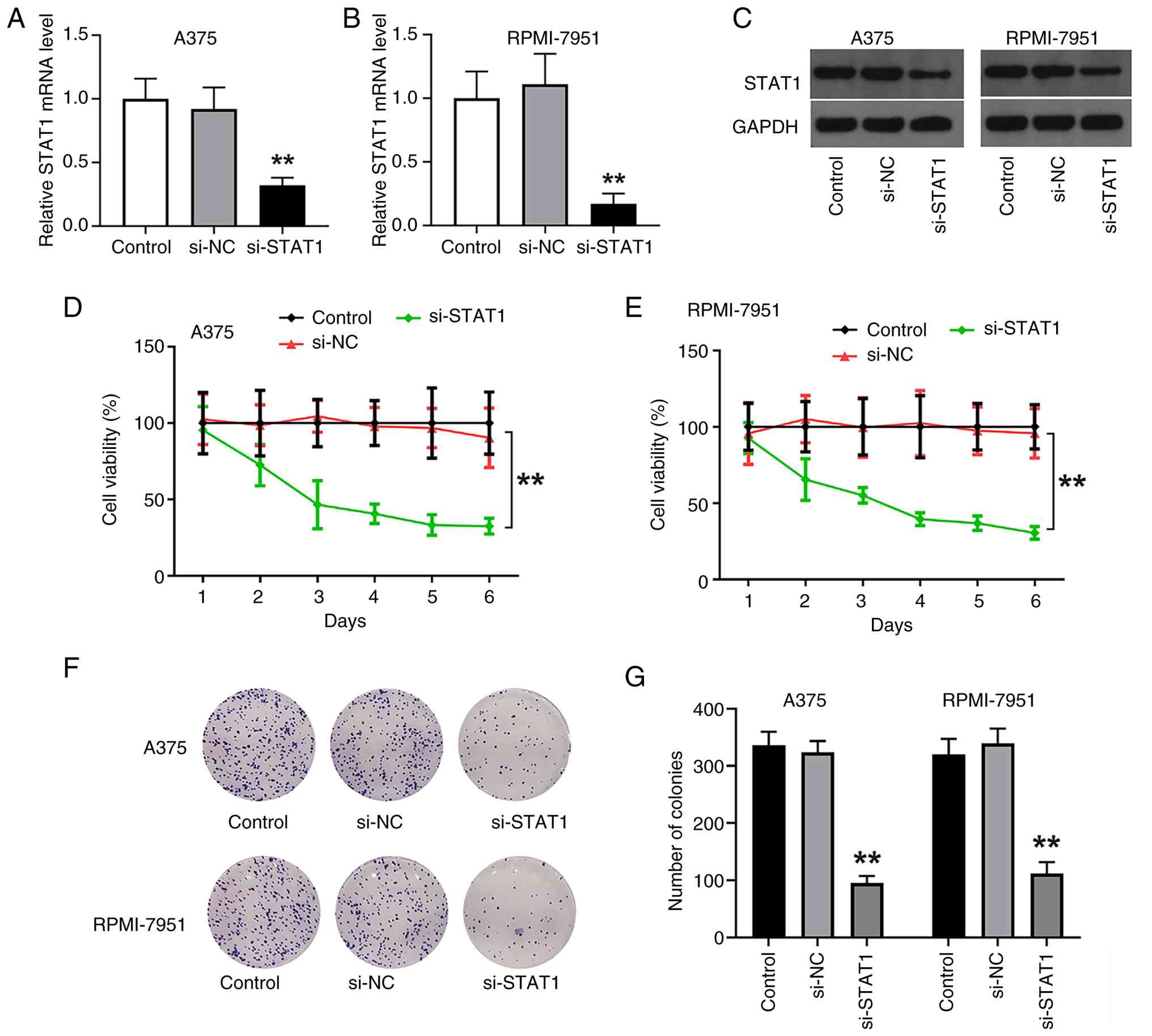
To assess the impact of STAT1 knockdown on melanoma
cells, STAT1 mRNA levels were measured in A375 and RPMI-7951 cell
lines following transfection with si-STAT1. In both cell lines,
STAT1 mRNA expression was significantly reduced in the si-STAT1
group compared with the control and si-NC groups (P<0.01),
supporting the successful knockdown of STAT1 (Fig. 3A and B). At the protein level,
western blot analysis further validated the knockdown of STAT1,
revealing a notable decrease in STAT1 protein expression in both
A375 and RPMI-7951 cells transfected with si-STAT1, while no
significant changes were observed between the control and si-NC
groups, reinforcing the effectiveness of the knockdown protocol
(Fig. 3C). To investigate the
functional consequences of STAT1 silencing, cell viability was
assessed via CCK-8 assay. A significant reduction in cell viability
over time was observed in the si-STAT1 group relative to the
control and si-NC groups (P<0.01; Fig. 3D and E), suggesting that STAT1 was
important for maintaining melanoma cell viability. Additionally,
the si-STAT1 group exhibited a notable reduction in colony
formation compared with the control and si-NC groups, indicating
that STAT1 knockdown inhibited the proliferative capacity of
melanoma cells. Quantitative analysis of colony formation assays
further revealed a significant reduction in colony counts in both
A375 and RPMI-7951 cell lines following transfection with si-STAT1
(P<0.01; Fig. 3F and G),
reinforcing the conclusion that STAT1 downregulation suppressed
melanoma cell proliferation.
Knockdown of STAT1 significantly
promotes apoptosis and inhibits cell migration in melanoma
cells
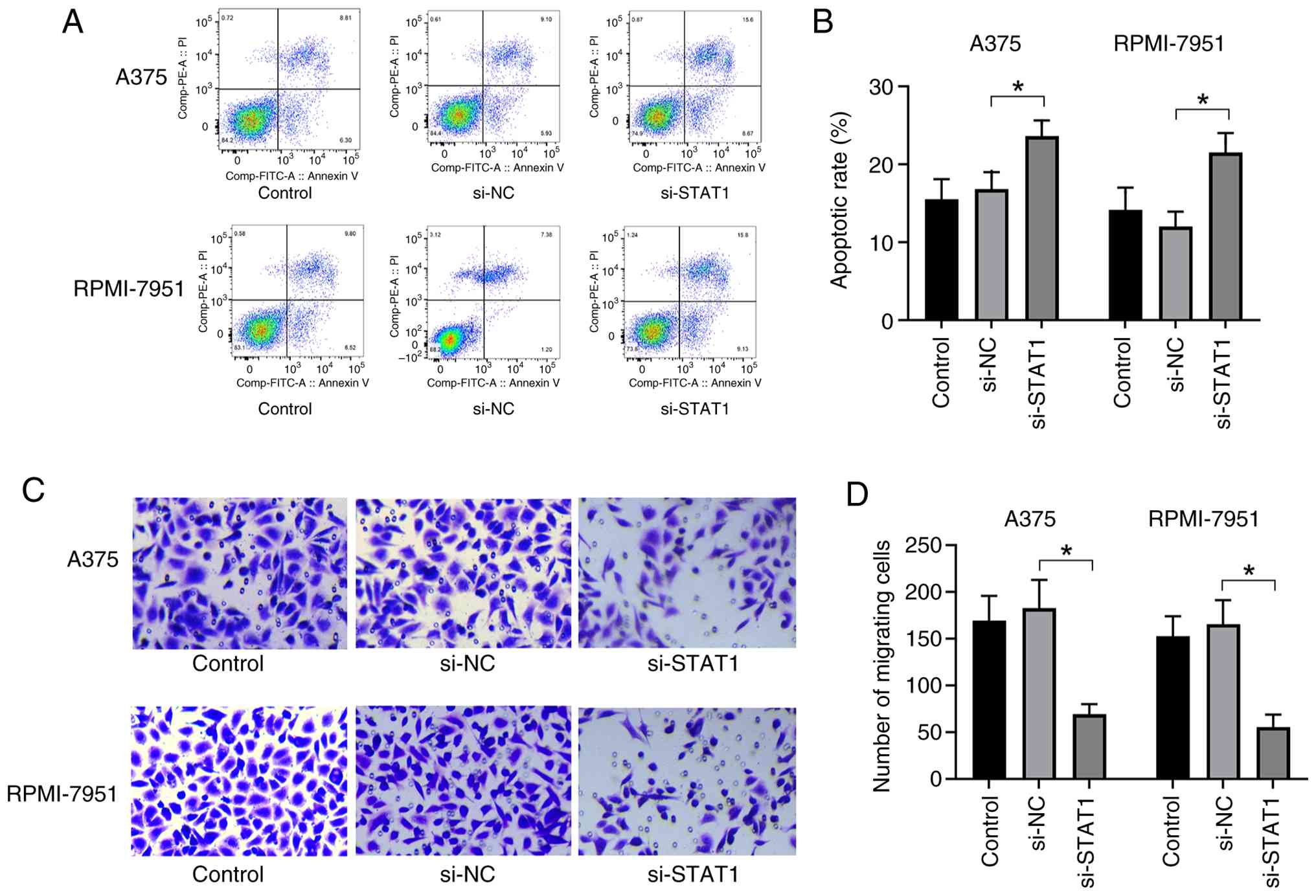
Apoptosis was assessed in A375 and RPMI-7951 cells
following STAT1 knockdown using flow cytometry. Scatter plots
revealed a significant increase in the proportion of cells
undergoing apoptosis in the si-STAT1 group compared with the
control groups, as indicated by a higher percentage of cells in the
apoptotic region (Fig. 4A).
Quantification of apoptotic rates showed that STAT1 knockdown
significantly elevated apoptosis in both A375 and RPMI-7951 cells
compared with the control and si-NC groups (P<0.05; Fig. 4B), suggesting that STAT1 knockdown
promoted melanoma cell apoptosis. Furthermore, migration assays
revealed a notable reduction in the number of migrating cells in
the si-STAT1 group compared with the control and si-NC groups
(Fig. 4C). Quantification of
migrating cell counts supported that STAT1 knockdown significantly
inhibited cell migration (P<0.05; Fig. 4D). Collectively, these results
demonstrated that STAT1 was important for the viability,
proliferation and migration of melanoma cells, with its depletion
promoting apoptosis and inhibiting cell migration. These results
underscored the notable role of STAT1 in regulating melanoma cell
survival and migratory capacity.
STAT1 regulates the expression of
TUBB4A through transcriptional mechanisms
Two independent si-STAT1 sequences, denoted
si-STAT1-1 and si-STAT1-2, were utilized for transfection. qPCR
analyses demonstrated that both siRNA sequences induced a
significant reduction in STAT1 mRNA level (P<0.01; Fig. 5A), whereas the si-NC sequence
exerted no significant effects on STAT1 expression. Following STAT1
knockdown, a significant reduction in TUBB4A mRNA expression was
observed in both A375 and RPMI-7951 melanoma cells compared with
the si-NC group (P<0.01; Fig.
5B). Bioinformatic analysis identified multiple high-confidence
STAT1 binding motifs within the regulatory region of the TUBB4A
promoter (Table SII). These
motifs were selected for further experimental validation. A series
of luciferase reporter constructs containing TUBB4A promoter
fragments were generated, spanning from −2,000 to −32 base pairs
relative to the transcription initiation site (Fig. 5C). Reporter assays revealed that
the −1,783 and −1,771 fragments exhibited the highest
transcriptional activity, as indicated by significantly increased
luciferase activity relative to other fragments (P<0.01;
Fig. 5D). These results suggested
that the −1,783 and −1,771 regions were important for TUBB4A
promoter activity. The mRNA results validated the efficacy of STAT1
knockdown (Fig. 5E). Upon STAT1
knockdown, luciferase activity of the promoter-gluc luciferase
(PGL)3-1783 construct significantly decreased (P<0.01; Fig. 5F), while the PGL3-1771 construct
showed no significant changes in luciferase activity. This
suggested that STAT1 specifically regulated transcription through
the −1,783 region of the TUBB4A promoter. ChIP assays further
supported that STAT1 binding to the TUBB4A promoter was
significantly enriched at the −1,783 region in both A375 and
RPMI-7951 cells compared with IgG control (P<0.01; Fig. 5G), providing direct evidence that
STAT1 interacted with the TUBB4A promoter at this site. These
results demonstrated that STAT1 directly regulated TUBB4A
expression at the transcriptional level by binding to the −1,783
region of the TUBB4A promoter.
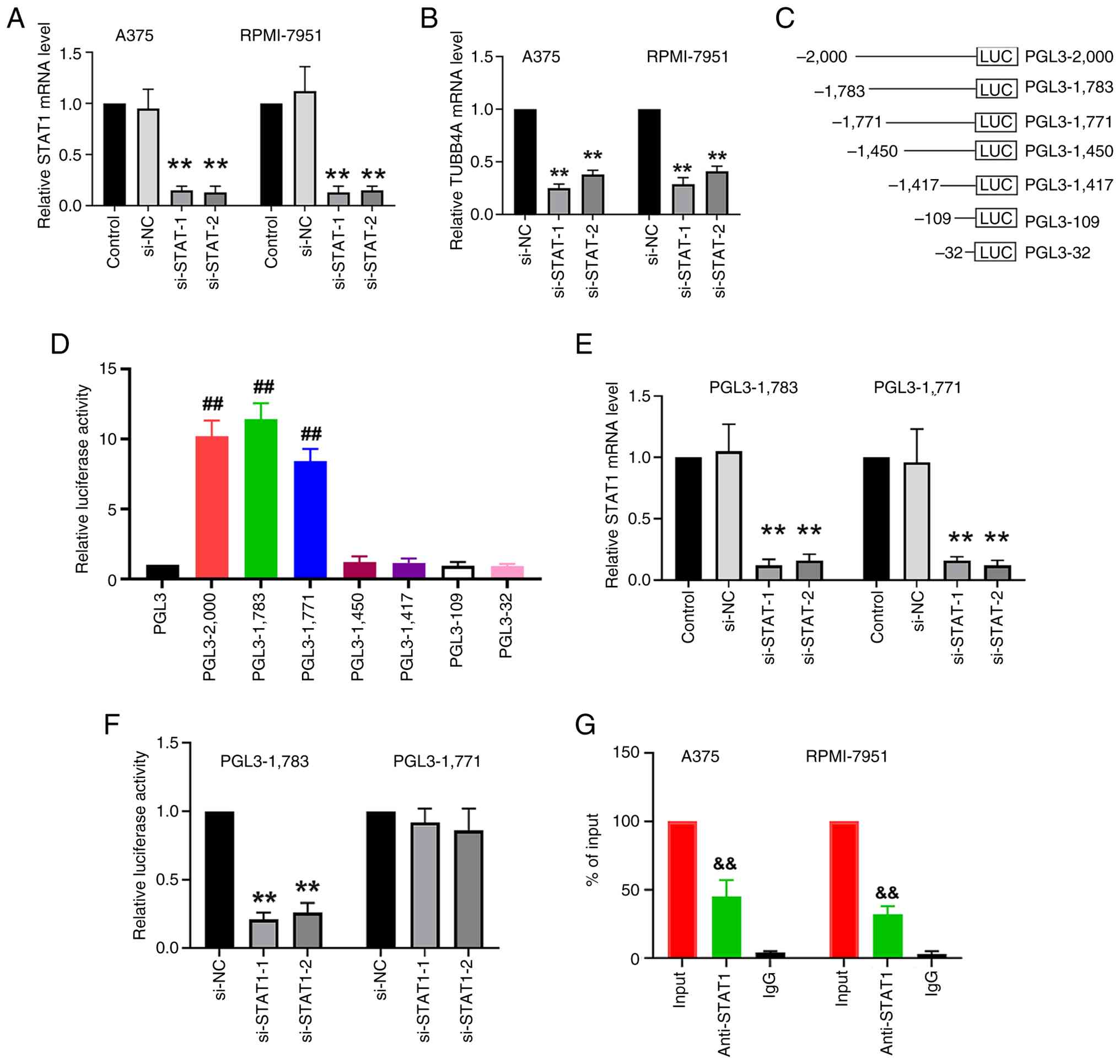 | Figure 5.STAT1 regulates TUBB4A expression at
the transcription level. (A) STAT1 mRNA levels were measured in
A375 and RPMI-7951 cells after STAT1 knockdown via transfection
with different siRNA sequences. (B) TUBB4A mRNA levels were
measured in A375 and RPMI-7951 cells after STAT1 knockdown via
transfection with different siRNA sequences. (C) Specific fragments
of the TUBB4A promoter region were cloned into the luciferase
reporter plasmids upstream of the firefly luciferase gene. (D)
Transcriptional activity of various TUBB4A promoter fragments was
analyzed by luciferase reporter assay in 293T cells, with the
−1,783 and −1,771 fragments exhibiting the highest activity. (E)
STAT1 siRNA-mediated knockdown significantly reduced STAT1 mRNA
levels in A375 cells. (F) STAT1 knockdown significantly reduced the
luciferase activity of the −1,783 fragment of the TUBB4A promoter,
but not the −1,771 fragment. (G) Chromatin immunoprecipitation
assays were performed in A375 and RPMI-7951 cells targeting the
−1,783 binding site in the TUBB4A promoter region. Quantitative PCR
provided evidence of STAT1 binding to this region. Genomic DNA
input was set to 100%. **P<0.01 vs. si-NC;
##P<0.01 vs. PGL3; &&P<0.01 vs.
IgG. STAT1, signal transducer and activator of transcription 1;
siRNA, small interfering RNA; si-NC, negative control siRNA;
si-STAT1, siRNA targeting STAT1; si-STAT1-1, siRNA targeting STAT1
sequence 1; si-STAT1-2, siRNA targeting STAT1 sequence 2; TUBB4A,
tubulin β4A; PGL3, promoter-gluc luciferase 3; LUC, firefly
luciferase gene. |
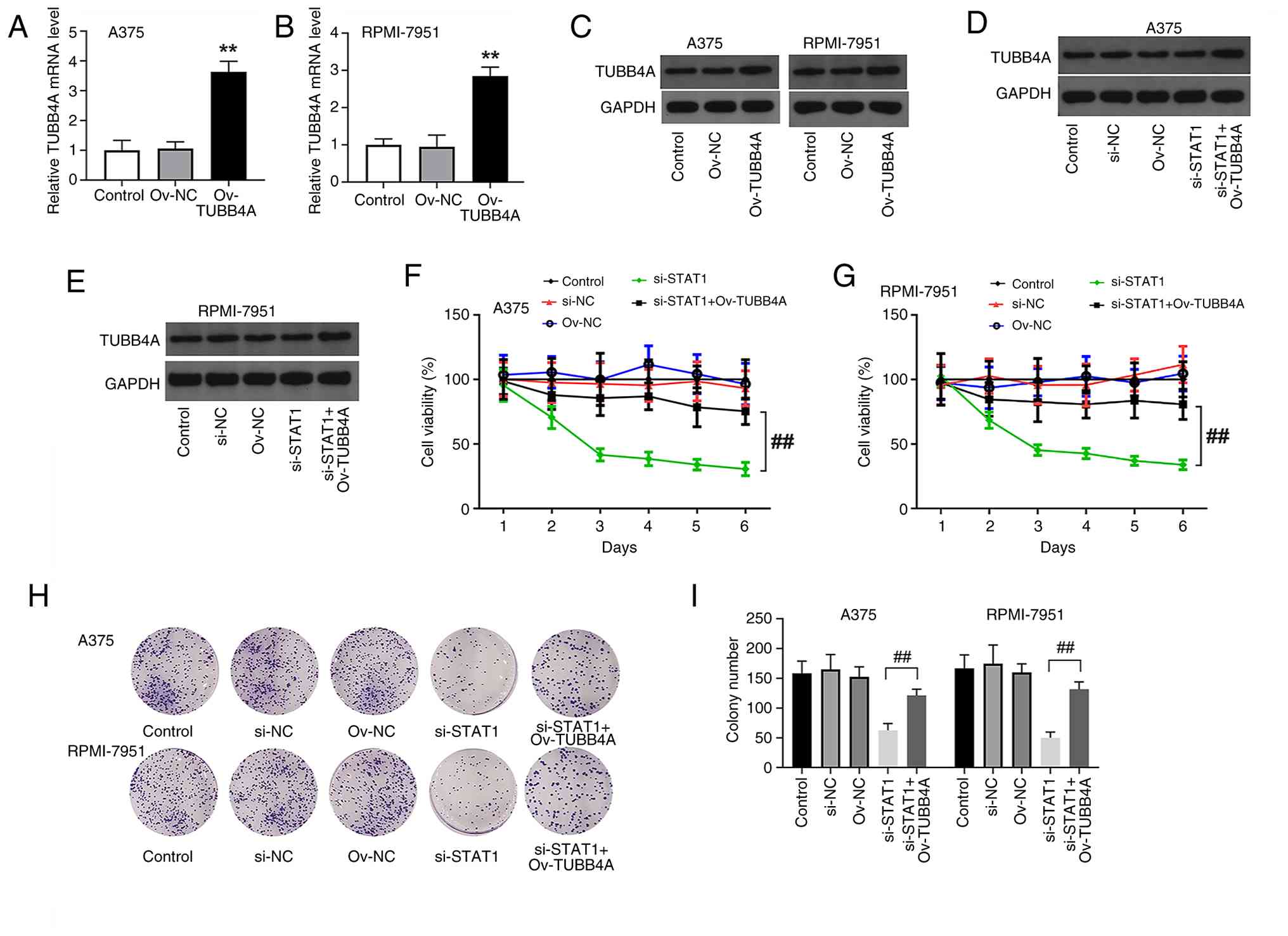
TUBB4A overexpression partially
reverses the suppressive impact of STAT1 knockdown on cell
viability
Compared with the control and Ov-NC groups, the
Ov-TUBB4A group exhibited a significant increase in TUBB4A
expression (P<0.01; Fig. 6A and
B), providing evidence of successful overexpression. Western
blot analysis further validated TUBB4A overexpression in A375 and
RPMI-7951 cells, with a marked increase in TUBB4A protein levels
observed in the Ov-TUBB4A group relative to the control and Ov-NC
groups (Fig. 6C), supporting
effective protein overexpression. TUBB4A protein levels were also
assessed under various conditions, including combined STAT1
knockdown and TUBB4A overexpression (Fig. 6D and E). To assess the impact of
STAT1 knockdown and TUBB4A overexpression on cell proliferation,
CCK-8 assays were conducted in A375 and RPMI-7951 cells. As
aforementioned, STAT1 knockdown significantly decreased cell
viability in both cell lines. Conversely, TUBB4A overexpression
significantly rescued cell proliferation (P<0.01; Fig. 6F and G). STAT1 knockdown led to a
notable reduction in colony formation, indicating impaired
proliferative capacity in STAT1-deficient melanoma cells. However,
simultaneous overexpression of TUBB4A markedly restored colony
formation compared with the si-STAT1 group (Fig. 6H). Quantitative colony formation
analysis supported a significant reduction in colony formation in
the si-STAT1 group, with TUBB4A overexpression significantly
restoring colony formation (P<0.01; Fig. 6I). These results demonstrated that
STAT1 silencing significantly inhibited melanoma cell proliferation
and colony formation in both A375 and RPMI-7951 cells, while TUBB4A
overexpression partially reversed these effects, suggesting that
TUBB4A functioned as a downstream effector of STAT1, playing a
compensatory role in regulating melanoma cell proliferation and
survival.
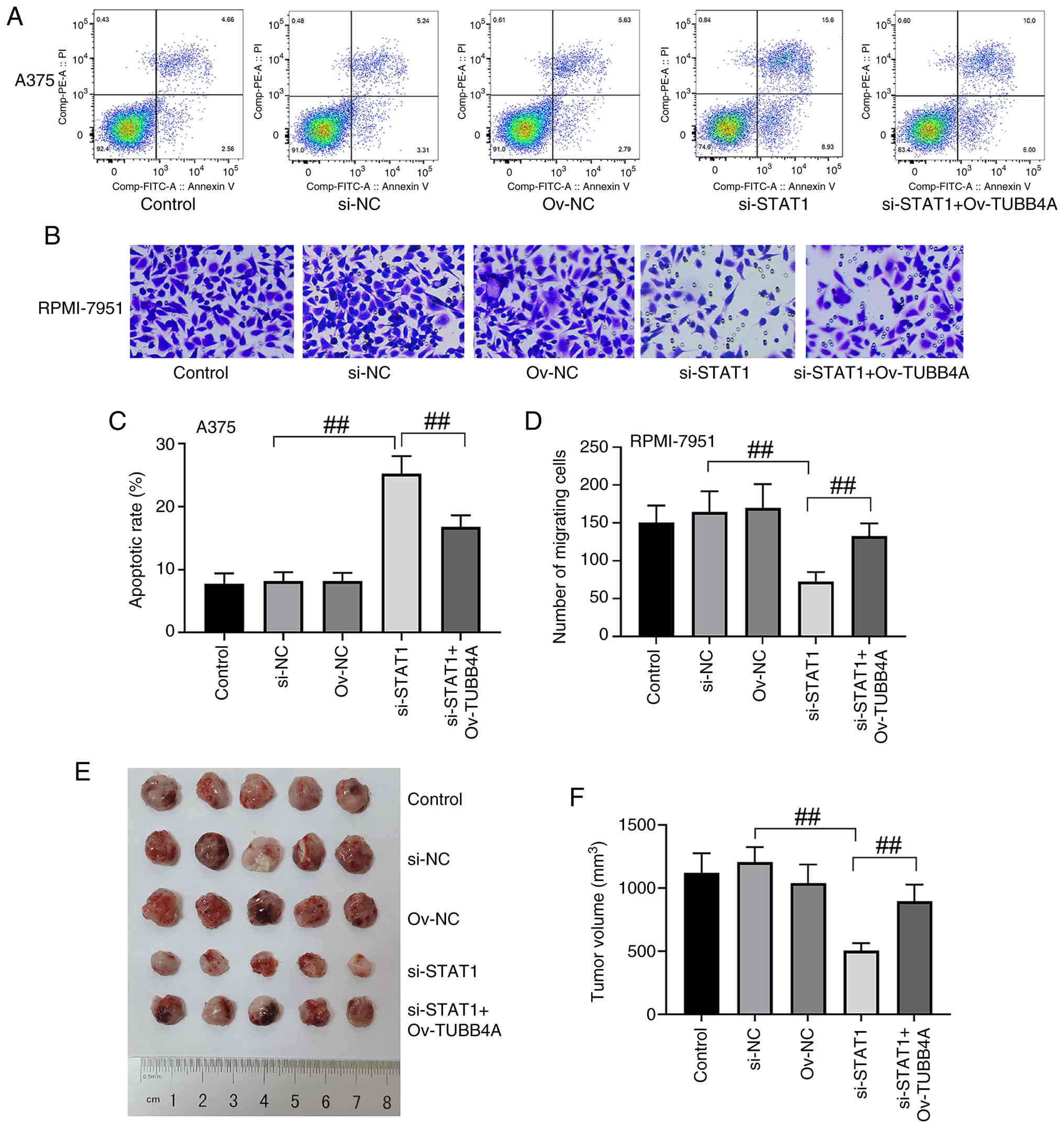
Overexpression of TUBB4A partially
reverses the effect of STAT1 knockdown on cell apoptosis, migration
and tumor growth in vivo
Flow cytometric analysis was conducted to evaluate
apoptosis in A375 cells following transfection treatments. STAT1
knockdown markedly increased the apoptotic rate compared with the
control and si-NC groups. However, co-transfection with TUBB4A
overexpression vectors partially mitigated this apoptotic effect,
as evidenced by a notable reduction in apoptotic cells (Fig. 7A). Quantification of apoptotic
rates supported a significant increase in apoptosis in the si-STAT1
group compared with the si-NC group, while the si-STAT1 + Ov-TUBB4A
group showed a significant reduction in apoptosis compared with
STAT1 knockdown alone (P<0.01; Fig.
7C). These results suggested that TUBB4A overexpression
counterbalanced the elevation in apoptosis induced by STAT1
knockdown. To assess the effects of STAT1 knockdown and TUBB4A
overexpression on cell migration, Transwell assays were performed
in RPMI-7951 cells (Fig. 7B).
STAT1 knockdown markedly inhibited cell migration, as demonstrated
by the reduced number of migrating cells. Notably, TUBB4A
overexpression following STAT1 knockdown partially restored
migration, indicating a potential rescue effect. Quantitative
migration analysis revealed a significant reduction in the number
of migrating cells in the si-STAT1 group compared with the si-NC
group, while this migration ability was partially yet significantly
restored in the si-STAT1 + Ov-TUBB4A group (P<0.01; Fig. 7D), suggesting that TUBB4A
overexpression partially counteracted the inhibitory effect of
STAT1 knockdown on melanoma cell migration.
In vivo tumorigenicity assays were performed
by injecting A375 cells into mice. Tumors derived from the si-STAT1
group were notably smaller than those from the control and si-NC
groups, indicating that STAT1 knockdown markedly suppressed tumor
growth. However, tumor volume in the si-STAT1 + Ov-TUBB4A group was
notably larger compared with the si-STAT1 group (Fig. 7E), suggesting that TUBB4A
overexpression partially rescued the tumor growth inhibition caused
by STAT1 knockdown. Quantitative analysis supported a significant
reduction in tumor volume in the si-STAT1 group compared with the
si-NC group, while the si-STAT1 + Ov-TUBB4A group exhibited a
significantly larger tumor volume than the si-STAT1 group
(P<0.01; Fig. 7F). The maximum
tumor volume and diameter observed at the endpoint of the study
were 1,320 mm3 and 13.6 mm, respectively, which were
observed in the si-NC group. These results further supported the
conclusion that TUBB4A overexpression mitigated the suppressive
effects of STAT1 knockdown on tumor growth. Collectively, these
results suggested that TUBB4A may have functionally cooperated with
STAT1 to regulate apoptosis, migration and tumor growth in melanoma
cells.
Discussion
Melanoma, an aggressive form of skin cancer,
presents ongoing challenges in clinical management. The present
study explored the complex roles of STAT1 and TUBB4A in SKCM,
focusing on their involvement in regulating cell proliferation,
migration, apoptosis and tumor progression. The findings of the
present study underscored the important role of STAT1 in promoting
melanoma progression through its regulation of cell survival and
motility, while TUBB4A served as a downstream effector that
mitigated the adverse effects of STAT1 silencing in melanoma
cells.
The present results aligned with previous studies
highlighting STAT1 as an important regulator of tumor progression.
STAT1 regulates numerous cellular processes, including immune
responses, proliferation and apoptosis (29–31).
In melanoma, STAT1 upregulation is associated with favorable or
poor prognosis, reflecting its dual role as either a tumor
suppressor or promoter, depending on the tumor context (15,32).
In the present study, silencing STAT1 significantly impaired
melanoma cell proliferation, migration and colony formation, while
simultaneously promoting apoptosis. These findings demonstrated
that STAT1 supported melanoma cell survival and motility,
consistent with observations in other cancer types where STAT1
facilitates tumor invasion and metastasis (33,34).
An apoptotic rate of ~20% following STAT1 knockdown, while modest
in absolute terms, reflected a strong 3–4 fold increase over
control levels. Similar ranges of apoptotic induction following
specific gene knockdown, with rates ranging from 15–30%, have been
frequently reported in cancer research literature and are accepted
as evidence of the notable role of a gene in cell survival
(35–38). In the context of single-gene
manipulation without additional cytotoxic agents, the discussed
magnitude of change is biologically notable and consistent with the
profound reductions observed in cell viability and long-term
proliferation in the present study, robustly supporting the
conclusion that STAT1 was important for melanoma cell survival.
Notably, the role of STAT1 in melanoma contrasted
with its generally tumor-suppressive function in cancers such as
breast and colorectal cancer (39–42).
This divergence suggested that the effects of STAT1 may have been
context-dependent, with its contribution to melanoma progression
likely modulated by interactions with other oncogenic pathways.
Although STAT1 is recognized for its role in orchestrating
anti-tumor immune responses and inflammation (43), the findings of the present study,
consistent with emerging evidence in melanoma (6,15,32),
underscored its context-dependent oncogenic capacity. The
pro-tumorigenic role of STAT1 may have extended beyond the
regulation of cell proliferation and apoptosis. Notable evidence
indicates that STAT1 serves as a core transcription factor in the
IFNγ signaling pathway and directly participates in regulating the
expression of the immune checkpoint molecule programmed cell death
1 ligand 1 (PD-L1) (44–46). A previous study in aggressive
soft-tissue sarcomas also revealed co-expression of PD-L1 with
cancer-testis antigens such as cancer-testis antigen 1, suggesting
a potential role for the STAT1 pathway in shaping the tumor immune
microenvironment (47). Therefore,
the present study speculated that in melanoma, STAT1 activation may
have concurrently driven malignant progression and immune escape.
Targeting STAT1 could potentially kill tumor cells directly by
inhibiting the expression of downstream targets such as TUBB4A; on
the other hand, targeting STAT1 may enhance T-cell-mediated
cytotoxicity by downregulating molecules such as PD-L1. Therefore,
the results of the present study provided a theoretical foundation
for combining STAT1 inhibitors with immune checkpoint blockers.
This combinatorial strategy represents an important direction for
future research.
The present study elucidated a novel mechanism
through which STAT1 drove melanoma progression by upregulating
TUBB4A expression. The significant association between high STAT1
expression and poor patient survival, coupled with mechanistic data
identifying the pro-survival and pro-migratory factor TUBB4A as a
direct transcriptional target of STAT1, suggested that in melanoma,
the tumor-promoting functions of STAT1 overrode its classical
tumor-suppressive roles. Kaplan-Meier survival analysis
demonstrated that elevated TUBB4A expression was strongly
associated with reduced OS and DFS, underscoring its negative
prognostic value. Notably, while TUBB4A expression was
significantly elevated in melanoma tissues overall, the cross-stage
analysis performed in the present study did not detect
statistically significant variations in its expression levels. This
suggested that TUBB4A upregulation was an early event in
melanomagenesis, remaining relatively stable throughout disease
progression. However, the present study could not rule out the
possibility that subtle changes in TUBB4A expression existed but
were not captured due to the sample size distribution across
stages. The consistent high expression of TUBB4A, irrespective of
pathological stage, further supported its potential role as a
fundamental driver of melanoma progression.
The analysis of STAT1-TUBB4A interactions conducted
within the present study revealed that the overexpression of TUBB4A
partially reversed the inhibitory effects of STAT1 knockdown on
cell proliferation, migration and colony formation. These findings
suggested that TUBB4A acted as a compensatory factor in melanoma
cells under conditions of STAT1 silencing. As a notable component
of the microtubule network, TUBB4A is important for maintaining
cell integrity and supports cellular processes such as division,
migration and survival (8,48). The present results suggested that
TUBB4A played a notable role in melanoma cell migration and
apoptosis, likely by stabilizing microtubules and influencing
cytoskeletal dynamics. The interplay between STAT1 and TUBB4A in
melanoma pointed to a potential mechanism in which STAT1 drove
tumor progression by upregulating TUBB4A. In turn, TUBB4A appeared
to support tumor cell migration and survival by stabilizing
microtubules, remodeling the cytoskeleton and enhancing motility
(49). This was consistent with
findings in other types of cancer, where tubulin family proteins,
including TUBB4A, and microtubule dynamics contribute to tumor
growth and therapy resistance (50).
In vivo experiments further demonstrated that
TUBB4A overexpression partially reversed the reduction in tumor
size induced by STAT1 knockdown. While tumor growth was not fully
restored to baseline, these results suggested that TUBB4A supported
tumor maintenance by promoting cell proliferation and survival
within the tumor microenvironment. The incomplete reversal of tumor
growth highlighted that additional molecular mechanisms were
potentially involved in melanoma progression and that TUBB4A alone
could not fully compensate for STAT1 silencing. This underscores
the complexity of tumor biology, where multiple pathways and
factors collaboratively influence cancer development (51). Future research should explore the
interactions of TUBB4A with other key molecules, such as STAT3 and
additional tubulin family proteins, to further elucidate its role
in melanoma progression.
The present study elucidated a novel mechanism
whereby STAT1 drove melanoma progression through the direct
transcriptional upregulation of TUBB4A. Placing this newly
identified axis in context alongside the well-established oncogene
STAT3 in melanoma helps to further define the findings of the
present study. STAT3 plays a central role in promoting cell
survival, proliferation and immune evasion by upregulating target
genes such as Bcl-xL, Myeloid cell leukemia-1, c-Myc
and cyclin D1 (52–54). Although STAT1 and STAT3 are
structurally related and can be activated by overlapping upstream
signals, for example through shared cytokine receptors such as
gp130, they often exhibit antagonistic or divergent functions in
melanoma (55,56). The present finding that STAT1
exerted its pro-tumorigenic effect by regulating TUBB4A, a key gene
involved in microtubule dynamics and cell motility, suggested a
distinct mechanism through which STAT1 drove malignancy,
potentially independent of the canonical STAT3 signaling network.
Nonetheless, potential synergy cannot be ruled out; STAT1 and STAT3
may cooperate to promote aggressiveness and therapy resistance in
melanoma by regulating separate yet complementary sets of
downstream targets. A key source of ambiguity in the present study
was whether TUBB4A was a shared target or was specifically
regulated by STAT1, which will be a focus of our future
investigations. In summary, the present study uncovered a
previously underappreciated, non-canonical oncogenic role for STAT1
in melanoma, independent of its classic immune-related functions,
and added a new layer of complexity to the STAT signaling paradigm
by identifying TUBB4A as a key downstream effector.
Given the role of TUBB4A in melanoma, further
studies are necessary to elucidate its underlying molecular
mechanisms, particularly in relation to tumor-associated signaling
pathways. Investigating the interaction between TUBB4A and STAT3, a
notable regulator in melanoma progression, could offer valuable
insights into how these molecules collaborate in tumor development.
Additionally, exploring how TUBB4A influences the tumor
microenvironment, including immune responses and metastatic
potential, may reveal novel therapeutic targets. The present study
also emphasized the therapeutic potential of targeting the
STAT1-TUBB4A axis. Since TUBB4A is implicated in drug resistance
across various cancers, its contribution to the resistance of
melanoma to chemotherapy and immunotherapy warrants in-depth
exploration. A combined approach targeting both STAT1 and TUBB4A
may present an innovative strategy to overcome therapeutic
resistance and enhance treatment outcomes for patients with
melanoma.
Summarily, STAT1 drove melanoma progression by
upregulating TUBB4A expression, with TUBB4A acting as a downstream
effector. The capacity of TUBB4A to counteract the effects of STAT1
knockdown underscored the potential of the STAT1-TUBB4A axis as a
therapeutic target in melanoma. These findings provided valuable
insights into refining treatment strategies for melanoma,
highlighting the importance of targeting this pathway to enhance
therapeutic efficacy.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
RZ and KF designed the study, performed in
vitro experiments and analyzed data. XZ contributed to patient
sample collection and clinical data interpretation. HL supervised
the project, designed in vivo xenograft experiments and
interpreted results. RZ and HL drafted the manuscript. KF and XZ
prepared figures and revised the manuscript. RZ and HL confirm the
authenticity of all the raw data. All authors read and approved the
final version of the manuscript.
Ethics approval and consent to
participate
Approval for clinical and animal experimental
protocols was granted by the Ethics Committee of Pudong New Area
People's Hospital (animal study, approval no. AEA-2024-33; clinical
sample study, approval no. CEA-2023-16). The requirement for
ethical approval regarding the use of primary human epidermal
melanocytes in the present study was waived by the Ethics Committee
of Pudong New Area People's Hospital. All subjects involved in the
study provided informed consent. Patients provided written consent
for participation in the present study.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
CCK-8
|
Cell Counting Kit-8
|
|
ChIP
|
chromatin immunoprecipitation
|
|
DFS
|
disease-free survival
|
|
FBS
|
fetal bovine serum
|
|
GEPIA
|
Gene Expression Profiling Interactive
Analysis
|
|
HEM
|
human epidermal melanocytes
|
|
HR
|
hazard ratio
|
|
JAK
|
Janus kinase
|
|
OS
|
overall survival
|
|
Ov-NC
|
negative control lentiviral
overexpression vector
|
|
Ov-TUBB4A
|
lentiviral vector for tubulin β4A
overexpression
|
|
PBS
|
phosphate-buffered saline
|
|
si-NC
|
negative control small interfering
RNA
|
|
si-STAT1
|
small interfering RNA targeting signal
transducer and activator of transcription 1
|
|
SKCM
|
skin cutaneous melanoma
|
|
STAT1
|
signal transducer and activator of
transcription 1
|
|
STAT3
|
signal transducer and activator of
transcription 3
|
|
TPM
|
transcripts per million
|
|
TUBB4A
|
tubulin β4A
|
References
|
1
|
Liu M, Wu M, Liu X, Zhou J, Lan Y, Zhang
H, Zhang X, Leng L, Zheng H and Li J: Assessing the quality of care
for skin malignant melanoma on a global, regional, and national
scale: A systematic analysis of the global burden of disease study
from 1990 to 2019. Arch Dermatol Res. 315:2893–2904. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Arnold M, Singh D, Laversanne M, Vignat J,
Vaccarella S, Meheus F, Cust AE, de Vries E, Whiteman DC and Bray
F: Global burden of cutaneous melanoma in 2020 and projections to
2040. JAMA Dermatol. 158:495–503. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Sarapultsev A, Gusev E, Komelkova M,
Utepova I, Luo S and Hu D: JAK-STAT signaling in inflammation and
stress-related diseases: Implications for therapeutic
interventions. Mol Biomed. 4:402023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Xue C, Yao Q, Gu X, Shi Q, Yuan X, Chu Q,
Bao Z, Lu J and Li L: Evolving cognition of the JAK-STAT signaling
pathway: autoimmune disorders and cancer. Signal Transduct Target
Ther. 8:2042023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wang W, Lopez McDonald MC, Kim C, Ma M,
Pan ZT, Kaufmann C and Frank DA: The complementary roles of STAT3
and STAT1 in cancer biology: Insights into tumor pathogenesis and
therapeutic strategies. Front Immunol. 14:12658182023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Huang L, Chen J, Zhao Y, Gu L, Shao X, Li
J, Xu Y, Liu Z and Xu Q: Key candidate genes of STAT1 and CXCL10 in
melanoma identified by integrated bioinformatical analysis. IUBMB
Life. 71:1634–1644. 2019. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Isacescu E, Chiroi P, Zanoaga O, Nutu A,
Budisan L, Pirlog R, Atanasov AG and Berindan-Neagoe I: Melanoma
cellular signaling transduction pathways targeted by polyphenols
action mechanisms. Antioxidants (Basel). 12:4072023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gao S, Wang S, Zhao Z, Zhang C, Liu Z, Ye
P, Xu Z, Yi B, Jiao K, Naik GA, et al: TUBB4A interacts with MYH9
to protect the nucleus during cell migration and promotes prostate
cancer via GSK3β/β-catenin signalling. Nat Commun. 13:27922022.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Kanakkanthara A and Miller JH: βIII
tubulin overexpression in cancer: Causes, consequences, and
potential therapies. Biochim Biophys Acta Rev Cancer.
1876:1886072021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Novikov NM, Zolotaryova SY, Gautreau AM
and Denisov EV: Mutational drivers of cancer cell migration and
invasion. Br J Cancer. 124:102–114. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Thomas SJ, Snowden JA, Zeidler MP and
Danson SJ: The role of JAK/STAT signalling in the pathogenesis,
prognosis and treatment of solid tumours. Br J Cancer. 113:365–371.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Hu X, Li J, Fu M, Zhao X and Wang W: The
JAK/STAT signaling pathway: From bench to clinic. Signal Transduct
Target Ther. 6:4022021. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hu Q, Bian Q, Rong D, Wang L, Song J,
Huang HS, Zeng J, Mei J and Wang PY: JAK/STAT pathway:
Extracellular signals, diseases, immunity, and therapeutic
regimens. Front Bioeng Biotechnol. 11:11107652023. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gandalovicova A, Šůchová AM, Čermák V,
Merta L, Rösel D and Brábek J: Sustained inflammatory signalling
through Stat1/Stat2/IRF9 is associated with amoeboid phenotype of
melanoma cells. Cancers (Basel). 12:24502020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Osborn JL and Greer SF: Metastatic
melanoma cells evade immune detection by silencing STAT1. Int J Mol
Sci. 16:4343–4361. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang W, Lopez McDonald MC, Hariprasad R,
Hamilton T and Frank DA: Oncogenic STAT transcription factors as
targets for cancer therapy: Innovative strategies and clinical
translation. Cancers (Basel). 16:13872024. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Prutsch N, He S, Berezovskaya A, Durbin
AD, Dharia NV, Maher KA, Matthews JD, Hare L, Turner SD, Stegmaier
K, et al: STAT3 couples activated tyrosine kinase signaling to the
oncogenic core transcriptional regulatory circuitry of anaplastic
large cell lymphoma. Cell Rep Med. 5:1014722024. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Fu XQ, Liu B, Wang YP, Li JK, Zhu PL, Li
T, Tse KW, Chou JY, Yin CL, Bai JX, et al: Activation of STAT3 is a
key event in TLR4 signaling-mediated melanoma progression. Cell
Death Dis. 11:2462020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kortylewski M, Jove R and Yu H: Targeting
STAT3 affects melanoma on multiple fronts. Cancer Metastasis Rev.
24:315–327. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen J, Kholina E, Szyk A, Fedorov VA,
Kovalenko I, Gudimchuk N and Roll-Mecak A: α-tubulin tail
modifications regulate microtubule stability through selective
effector recruitment, not changes in intrinsic polymer dynamics.
Dev Cell. 56:2016–2028.ed. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Fu G, Yan S, Khoo CJ, Chao VC, Liu Z,
Mukhi M, Hervas R, Li XD and Ti SC: Integrated regulation of
tubulin tyrosination and microtubule stability by human
alpha-tubulin isotypes. Cell Rep. 42:1126532023. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Garcin C and Straube A: Microtubules in
cell migration. Essays Biochem. 63:509–520. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Nami B and Wang Z: Genetics and expression
profile of the tubulin gene superfamily in breast cancer subtypes
and its relation to taxane resistance. Cancers (Basel). 10:2742018.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Song JX, Wang Y, Hua ZP, Huang Y, Hu LF,
Tian MR, Qiu L, Liu H and Zhang J: FATS inhibits the Wnt pathway
and induces apoptosis through degradation of MYH9 and enhances
sensitivity to paclitaxel in breast cancer. Cell Death Dis.
15:8352024. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wagstaff W, Mwamba RN, Grullon K,
Armstrong M, Zhao P, Hendren-Santiago B, Qin KH, Li AJ, Hu DA,
Youssef A, et al: Melanoma: Molecular genetics, metastasis,
targeted therapies, immunotherapies, and therapeutic resistance.
Genes Dis. 9:1608–1623. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Marconcini R, Pezzicoli G, Stucci LS,
Sergi MC, Lospalluti L, Porta C and Tucci M: Combination of
immunotherapy and other targeted therapies in advanced cutaneous
melanoma. Hum Vaccin Immunother. 18:19803152022. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Papageorgiou C, Apalla Z, Manoli SM,
Lallas K, Vakirlis E and Lallas A: Melanoma: Staging and follow-up.
Dermatol Pract Concept. 11 (Suppl):e2021162S2021. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Tolomeo M, Cavalli A and Cascio A: STAT1
and its crucial role in the control of viral infections. Int J Mol
Sci. 23:40952022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Niu M, Yi M, Dong B, Luo S and Wu K:
Upregulation of STAT1-CCL5 axis is a biomarker of colon cancer and
promotes the proliferation of colon cancer cells. Ann Transl Med.
8:9512020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Li X, Wang F, Xu X, Zhang J and Xu G: The
dual role of STAT1 in ovarian cancer: Insight into molecular
mechanisms and application potentials. Front Cell Dev Biol.
9:6365952021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Cerezo M, Guemiri R, Druillennec S,
Girault I, Malka-Mahieu H, Shen S, Allard D, Martineau S, Welsch C,
Agoussi S, et al: Translational control of tumor immune escape via
the eIF4F-STAT1-PD-L1 axis in melanoma. Nat Med. 24:1877–1886.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang J, Tan GL, Jiang M, Wang TS, Liu GH,
Xiong SS and Qing X: Effects of SENP1-induced deSUMOylation of
STAT1 on proliferation and invasion in nasopharyngeal carcinoma.
Cell Signal. 101:1105302023. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wong GS, Lee JS, Park YY, Klein-Szanto AJ,
Waldron TJ, Cukierman E, Herlyn M, Gimotty P, Nakagawa H and Rustgi
AK: Periostin cooperates with mutant p53 to mediate invasion
through the induction of STAT1 signaling in the esophageal tumor
microenvironment. Oncogenesis. 2:e592013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Huang J, Ding Z, Luo Q and Xu W: Cancer
cell-derived exosomes promote cell proliferation and inhibit cell
apoptosis of both normal lung fibroblasts and non-small cell lung
cancer cell through delivering alpha-smooth muscle actin. Am J
Transl Res. 11:1711–1723. 2019.PubMed/NCBI
|
|
36
|
Xu J, Li Y and Hu H: Effects of lycopene
on ovarian cancer cell line SKOV3 in vitro: Suppressed
proliferation and enhanced apoptosis. Mol Cell Probes.
46:1014192019. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Ji Y, Yu M, Qi Z, Cui D, Xin G, Wang B,
Jia W and Chang L: Study on apoptosis effect of human breast cancer
cell MCF-7 induced by lycorine hydrochloride via death receptor
pathway. Saudi Pharm J. 25:633–637. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Xu G, Zhang Y, Li N, Zhang JB and Xu R:
LncRNA CCHE1 in the proliferation and apoptosis of gastric cancer
cells. Eur Rev Med Pharmacol Sci. 22:2631–2637. 2018.PubMed/NCBI
|
|
39
|
Tanaka A, Zhou Y, Ogawa M, Shia J,
Klimstra DS, Wang JY and Roehrl MH: STAT1 as a potential prognosis
marker for poor outcomes of early stage colorectal cancer with
microsatellite instability. PLoS One. 15:e02292522020. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Nivarthi H, Gordziel C, Themanns M, Kramer
N, Eberl M, Rabe B, Schlederer M, Rose-John S, Knosel T, Kenner L,
et al: The ratio of STAT1 to STAT3 expression is a determinant of
colorectal cancer growth. Oncotarget. 7:51096–51106. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Legrier ME, Bièche I, Gaston J, Beurdeley
A, Yvonnet V, Déas O, Thuleau A, Château-Joubert S, Servely JL,
Vacher S, et al: Activation of IFN/STAT1 signalling predicts
response to chemotherapy in oestrogen receptor-negative breast
cancer. Br J Cancer. 114:177–187. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Wong GL, Manore SG, Doheny DL and Lo HW:
STAT family of transcription factors in breast cancer: Pathogenesis
and therapeutic opportunities and challenges. Semin Cancer Biol.
86:84–106. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wudtiwai B, Makeudom A, Krisanaprakornkit
S, Pothacharoen P and Kongtawelert P: Anticancer activities of
hesperidin via suppression of up-regulated programmed death-ligand
1 expression in oral cancer cells. Molecules. 6:53452021.
View Article : Google Scholar
|
|
44
|
Okita R, Shimizu K, Nojima Y, Saisho S and
Nakata M: Tofacitinib overcomes an IFNγ-induced decrease in NK
cell-mediated cytotoxicity via the regulation of immune-related
molecules in LC-2/ad. Thorac Cancer. 12:775–782. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Breitenecker K, Homolya M, Luca AC, Lang
V, Trenk C, Petroczi G, Mohrherr J, Horvath J, Moritsch S, Haas L,
et al: Down-regulation of A20 promotes immune escape of lung
adenocarcinomas. Sci Transl Med. 13:eabc39112021. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Yang J, Wang X, Huang B, Liu R, Xiong H,
Ye F, Zeng C, Fu X and Li L: An IFNγ/STAT1/JMJD3 axis 2021induces
ZEB1 expression and promotes aggressiveness in lung adenocarcinoma.
Mol Cancer Res. 19:1234–1246. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Hashimoto K, Nishimura S, Ito T, Kakinoki
R and Akagi M: Immunohistochemical expression and
clinicopathological assessment of PD-1, PD-L1, NY-ESO-1, and
MAGE-A4 expression in highly aggressive soft tissue sarcomas. Eur J
Histochem. 66:33932022. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Sase S, Almad AA, Boecker CA, Guedes-Dias
P, Li JJ, Takanohashi A, Patel A, McCaffrey T, Patel H,
Sirdeshpande D, et al: TUBB4A mutations result in both glial and
neuronal degeneration in an H-ABC leukodystrophy mouse model.
Elife. 9:e529862020. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Sobierajska K, Ciszewski WM, Wawro ME,
Wieczorek-Szukala K, Boncela J, Papiewska-Pajak I, Niewiarowska J
and Kowalska MA: TUBB4B downregulation is critical for increasing
migration of metastatic colon cancer cells. Cells. 8:8102019.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Vemu A, Atherton J, Spector JO, Moores CA
and Roll-Mecak A: Tubulin isoform composition tunes microtubule
dynamics. Mol Biol Cell. 28:3564–3572. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Du W and Elemento O: Cancer systems
biology: Embracing complexity to develop better anticancer
therapeutic strategies. Oncogene. 34:3215–3225. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Afrang N and Honardoost M: Cell cycle
regulatory markers in melanoma: New strategies in diagnosis and
treatment. Med J Islam Repub Iran. 33:962019.PubMed/NCBI
|
|
53
|
Becker TM, Boyd SC, Mijatov B,
Gowrishankar K, Snoyman S, Pupo GM, Scolyer RA, Mann GJ, Kefford
RF, Zhang XD and Rizos H: Mutant B-RAF-Mcl-1 survival signaling
depends on the STAT3 transcription factor. Oncogene. 33:1158–1166.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Picco ME, Castro MV, Quezada MJ, Barbero
G, Villanueva MB, Fernández NB, Kim H and Lopez-Bergami P: STAT3
enhances the constitutive activity of AGC kinases in melanoma by
transactivating PDK1. Cell Biosci. 9:32019. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Regis G, Pensa S, Boselli D, Novelli F and
Poli V: Ups and downs: The STAT1:STAT3 seesaw of Interferon and
gp130 receptor signalling. Semin Cell Dev Biol. 19:351–359. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Kortylewski M, Komyod W, Kauffmann ME,
Bosserhoff A, Heinrich PC and Behrmann I: Interferon-gamma-mediated
growth regulation of melanoma cells: Involvement of STAT1-dependent
and STAT1-independent signals. J Invest Dermatol. 122:414–422.
2004. View Article : Google Scholar : PubMed/NCBI
|