Introduction
RNA polymerase II (Pol II) is a key enzyme of
eukaryotic cells. Pol II transcribes messenger RNAs (mRNAs) and
various non-coding RNAs (ncRNAs) that play key roles in cellular
growth, differentiation, and stress responses (1). Pol II is a large multi-subunit
complex of 12 distinct proteins, each contributing to the overall
structural integrity and functional specificity of the enzyme
(2). Among these proteins, RNA
polymerase II, I and III subunit L (POLR2L), a highly conserved
7.6-kDa subunit featuring an atypical zinc finger domain, plays
vital roles in terms of maintaining the stability and efficiency of
the Pol II complex (3). Despite
its small size, POLR2L is essential for both Pol II assembly and
enzymic interactions with DNA and RNA throughout transcription
(4). Although extensive structural
and functional studies of the RNA polymerase II complex have been
conducted, most previous research has primarily focused on the
overall architecture of the enzyme and its major subunits (5–7). As
a result, small Pol II subunits such as POLR2L, POLR2H, and POLR2K
have often been treated as structural components rather than as
subjects of systematic comparative analysis (8,9).
This gap highlights the need for a more integrative perspective
that places POLR2L in a direct structural and evolutionary context
with other small Pol II subunits (10,11).
AlphaFold, an artificial intelligence system, predicts the
three-dimensional structures of proteins based on their amino acid
sequences. This advanced machine learning approach uses multiple
sequence alignments to yield physical and biological insights into
how deep learning algorithms operate, significantly enhancing the
accuracy of protein structure prediction (12). This approach has yielded
predictions of the POLR2L, POLR2H, and POLR2K protein structures.
The structural role of POLR2L aside, the subunit is of broader
biological significance (13).
Emerging evidence indicates that POLR2L regulates transcriptional
programs that influence both cellular homeostasis and pathological
conditions (14). POLR2L serves as
a key regulator of various cellular processes, actively
participating in cell proliferation, differentiation, and energy
metabolism (15). POLR2L
dysregulation has been linked to various diseases, particularly
cancer. POLR2L overexpression has been associated with tumor
growth, metastasis, and resistance to chemotherapy (16). These findings suggest that POLR2L
is not merely a structural component of the transcriptional
machinery but also an active regulator of critical cellular
pathways (17). Recent research
has revealed the involvement of POLR2L in several major signaling
pathways, including the PI3K-Akt, Wnt/β-catenin, and TGF-β
(18). pathways that regulate
cellular survival, proliferation, the immune responses, and tissue
remodeling (19). Within these
signaling networks, POLR2L plays a crucial role in terms of
coordinating intricate molecular cascades, further emphasizing its
biological importance (20).
POLR2L critically regulates the immune responses and DNA damage
repair pathways, further highlighting the diverse functionality and
significance of the subunit in disease (21). The clinical relevance of POLR2L is
becoming increasingly apparent (16). Its overexpression has been linked
to poor prognosis, increased metastatic potential, and resistance
to chemotherapy of various cancers, including hepatocellular
carcinoma (HCC), gastric cancer, and prostate cancer (17). Some studies found that POLR2L
inhibition suppressed tumor growth and improved drug sensitivity in
preclinical models, emphasizing its potential as a therapeutic
target (18). The role of POLR2L
in cancer aside, the protein affects the progression of viral
infections and immune-mediated diseases, further underscoring its
broad significance in terms of human health. This review
comprehensively analyzes the structural and functional attributes
of POLR2L and the roles played by the subunit in disease
progression and the development of new therapeutic strategies. The
advanced methodologies used include RNA sequencing, CRISPR-Cas9
gene editing, and protein-protein interaction analysis. The review
highlights the potential of POLR2L as both a diagnostic biomarker
and a therapeutic target, emphasizing its relevance in both future
research and precision medicine. By synthesizing the findings of
POLR2L-related studies, the review establishes a foundation for
future advances in personalized medicine.
Structure and biochemical properties of
POLR2L
To contextualize the structural and functional role
of POLR2L within the RNA polymerase II complex, we performed a
comparative structural, sequence-based analysis of POLR2L alongside
two other small Pol II subunits, POLR2H and POLR2K (8,9,22).
Pol II plays a central role in the gene expression of eukaryotic
cells, facilitating the transcription of both protein-coding mRNAs
and various ncRNAs (2,6). Of the 12 subunits that constitute the
Pol II complex, POLR2L is a small (7.6 kDa), highly conserved
subunit, reflecting its profound evolutionary significance
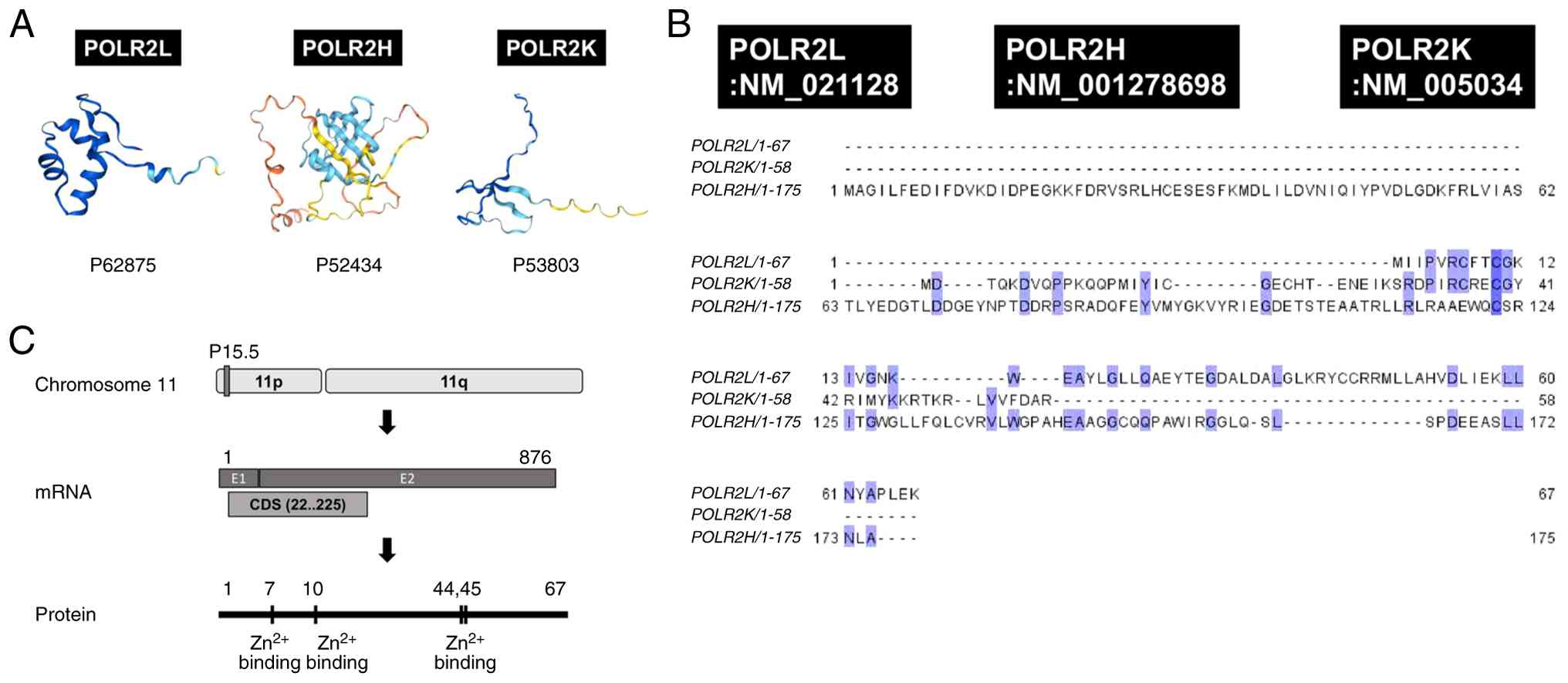
(6,23). As shown in Fig. 1A, AlphaFold-based structural models
have suggested that POLR2L adopts a compact globular fold with a
surface-exposed zinc-binding pocket (12,24),
whereas POLR2H and POLR2K display more extended conformations with
distinct surface charge distributions (2,25).
Although this in silico prediction should be interpreted as
hypothesis-generating and requires experimental validation, such
differences support the notion that POLR2L may provide a flexible
interaction module that can bridge specific subunits within the Pol
II core, in contrast to the mainly architectural roles proposed for
POLR2H and POLR2K (26,27).
Clustal Omega was used to perform multiple sequence
alignment (MSA) of POLR2L, POLR2H, and POLR2K (28). Although POLR2L, POLR2H, and POLR2K
contain 67, 175, and 58 amino acid residues, respectively, several
conserved residues were identified (Fig. 1B). Sequence alignment analysis
(Fig. 1B) revealed that POLR2L,
POLR2H, and POLR2K share conserved residues despite differences in
overall sequence length, supporting their functional relatedness as
small RNA polymerase II subunits (29,30).
Notably, POLR2L has strongly conserved cysteine residues that are
predicted to coordinate zinc ions, indicating additional
evolutionary constraints that may underlie specialized structural
contributions within the Pol II complex (25,31).
These include regions potentially involved in both metal ion
coordination and subunit interaction, highlighting functionally
relevant similarities across these Pol II components (32). The alignment supported the
hypothesis that the three small subunits share conserved
biochemical features critical for Pol II complex assembly and
stability (33). POLR2L contains
an atypical zinc finger domain that is essential for the
maintenance of Pol II structural stability and functionality
(34). This domain appears to
support efficient complex assembly and transcriptional activity,
although direct experimental dissection of its contribution to
these processes remains limited (1,35).
The atypical zinc finger motif of POLR2L, which is
characterized by a conserved
Cys-X2-Cys-X12-Cys-X2-Cys
configuration, resembles zinc-coordinating motifs in other
multi-subunit RNA polymerases that reinforce subunit-subunit
interactions (36,37). By analogy, POLR2L may contribute to
Pol II assembly and transcriptional stability through
metal-mediated structural reinforcement, although this mechanistic
role remains to be directly validated (3,38)
(Fig. 1C). POLR2L is proposed to
contribute to Pol II stability and transcriptional precision,
primarily based on structural predictions and sequence conservation
analyses, rather than direct mutational or structural experiments
focused on POLR2L itself (25,39).
These predictions provide a useful framework for hypothesis
generation; however, dedicated biochemical and biophysical studies
will be required to establish definitively how POLR2L influences
Pol II assembly and function (8,35).
From a functional perspective, the evolutionary
conservation of POLR2L and its zinc-binding motif strongly suggests
a role in stabilizing Pol II assembly and facilitating subunit
interactions (25,35). Previous biochemical and structural
studies of RNA polymerase complexes have demonstrated that small
zinc-binding subunits frequently act as structural scaffolds that
reinforce interactions among core subunits. In yeast and archaeal
RNA polymerases, the disruption of homologous zinc-binding subunits
impairs the assembly of POLR2C-POLR2J (RPB3-RPB11) subcomplexes and
diminishes transcriptional initiation efficiency, providing a
functional precedent for the putative scaffolding role of POLR2L in
the human Pol II complex (40–42).
In this context, conserved cysteine residues within the POLR2L zinc
finger motif are likely to contribute to the stabilization of the
POLR2C-POLR2J subcomplex and maintenance of Pol II structural
integrity (3,34,43).
Although direct experimental evidence specifically
dissecting individual POLR2L residues remains limited, studies in
yeast and mammalian systems have shown that the perturbation of
small Pol II subunits disrupts polymerase assembly, reduces
transcriptional efficiency, and destabilizes elongating RNA-DNA
hybrids (44–47). Given the strong conservation of
cysteine residues within the POLR2L zinc finger motif, and the
sensitivity of small Pol II subunits to perturbation in other
systems, these residues likely constitute key anchoring points for
subunit-subunit interfaces that support Pol II assembly and
elongation competence (25,37).
Therefore, the high degree of sequence conservation observed in
POLR2L likely reflects evolutionary pressure to preserve these
structural and functional interactions within the Pol II complex
(48). Future mutational and
structural analyses will be required to validate the mechanistic
contributions of conserved residues and the zinc finger domain to
Pol II function (37). Although
the atypical zinc finger domain of POLR2L likely plays an important
role in maintaining structural integrity within the RNA polymerase
II complex, its precise functional contributions remain
incompletely defined (8). In
addition to stabilizing subunit interactions, this zinc-binding
motif may play complementary roles, such as facilitating
conformational flexibility during transcriptional initiation or
elongation, acting as a scaffold for transient protein-protein
interactions, or contributing to the dynamic regulation of Pol II
under stress or signaling conditions (49). The functional relevance of the zinc
finger domain may also be context-dependent, varying across cell
types or transcriptional states (50). Importantly, most current insights
into the POLR2L zinc finger are derived from structural
predictions, evolutionary conservation, and indirect functional
studies, underscoring the need for future biochemical and
mutational analyses (25).
A comparative analysis of POLR2L with other small
RNA polymerase II subunits, particularly POLR2H and POLR2K,
provides additional insight into the organizational principles of
the Pol II complex (1,2,6,24).
Although all three subunits are relatively small and evolutionarily
conserved, they display distinct structural and biochemical
features that suggest functional specialization (13,34,40).
POLR2H and POLR2K primarily function as architectural components
that stabilize core Pol II subcomplexes, whereas POLR2L is
distinguished by its atypical zinc-binding motif and compact fold,
which may confer greater flexibility in mediating subunit
interactions (3,14,23,34).
Sequence alignment analyses have revealed that POLR2L exhibits a
higher degree of conservation within its zinc-coordinating
residues, compared to POLR2H and POLR2K, indicating stronger
evolutionary constraints on metal binding functionality (23,28,34).
This distinction suggests that POLR2L may play a more dynamic role
in reinforcing Pol II assembly or maintaining transcriptional
stability under variable cellular conditions (4,41).
By placing POLR2L in a direct structural and evolutionary context
with POLR2H and POLR2K, this comparative framework advances our
current understanding of small Pol II subunit organization, while
highlighting features that may underlie functional specialization
within the transcriptional machinery (6).
Interaction network of POLR2L
POLR2L has been proposed to regulate transcription
with other subunits of the Pol II complex, principally POLR2C and
POLR2J. These physical connections are thought to contribute to the
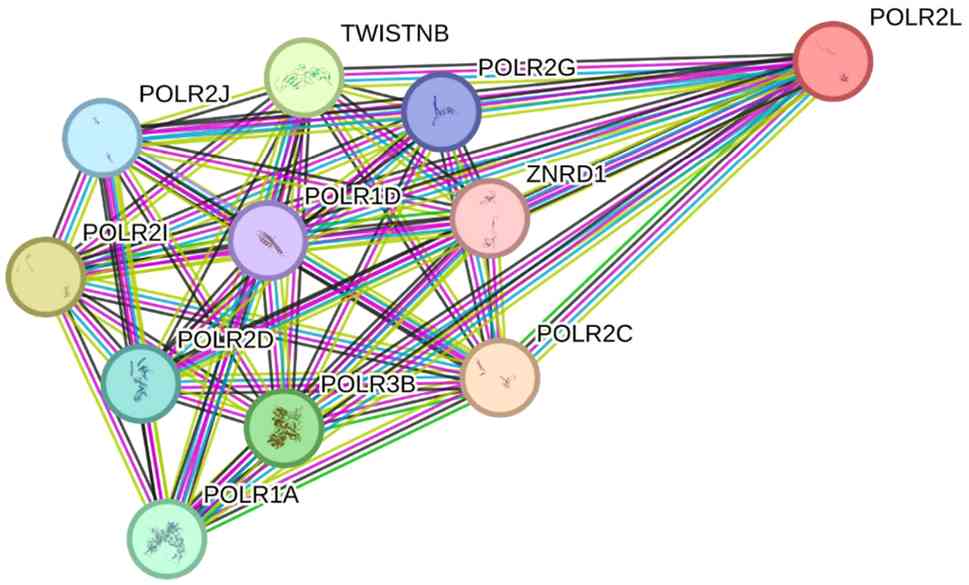
assembly and stability of the Pol II complex. POLR2L also interacts
with other proteins involved in transcription and signaling
(Fig. 2), such as POLR2C, POLR1A,
POLR3B, and POLR2E, further underscoring its importance within the
general transcription machinery (3). POLR2L has been implicated in
stabilizing RNA-DNA hybrids and may influence the elongation phase
of transcription (51). POLR2L
mutations have been reported to disrupt Pol II complex assembly and
impair transcriptional activity associated with transcriptional
repression and metabolic dysfunction (16). Such mutations have been identified
in various cancers, where they significantly reduce cell survival
and proliferation (16). Notably,
structural defects in POLR2L have been linked to chemotherapy
resistance in cancer cells, which suggests that POLR2L may affect
cancer progression and treatment responses (16).
Evolutionary conservation and biological
significance of POLR2L
POLR2L amino acid sequence and structural features
are highly conserved across eukaryotes and archaea, highlighting
its critical roles in Pol II function (40). Despite its structural simplicity,
POLR2L engages in transcriptional activation, enhancing the
adaptability of the overall enzyme complex (3). Its evolutionary conservation suggests
that POLR2L acts as a key regulatory component of the transcription
machinery, rather than merely playing a structural role (34).
Pol II is a key enzyme of eukaryotic gene
expression, transcribing both protein-coding genes and ncRNAs
(2,41). As an integral component of the Pol
II complex, POLR2L plays essential roles throughout all stages of
transcription, including initiation, elongation, and termination
(3). Although POLR2L primarily
maintains transcriptional efficiency and stability, the subunit is
also involved in various cellular processes (34), including the regulation of growth,
differentiation, the stress response, and metabolic pathways,
demonstrating its broader biological significance (15).
Role of POLR2L in transcription
POLR2L contributes to both transcription initiation
and elongation (3). POLR2L
stabilizes RNA-DNA hybrids, ensuring that Pol II remains securely
attached to the DNA template (40). POLR2L also maintains the structural
integrity of Pol II, supporting efficient RNA synthesis via
transcription and elongation (34). POLR2L function is prominent near
the promoters of actively transcribed genes, where the subunit
facilitates transcriptional initiation even in the presence of
transcriptional repressors (3),
which allows cells to adapt swiftly to environmental changes,
enhancing transcriptional flexibility under stress (15).
POLR2L function in cell growth and
differentiation
POLR2L participates in the regulation of cell growth
and differentiation (34). Its
expression is elevated in highly proliferative tissues, suggesting
that POLR2L functions as a key transcriptional regulator during the
S and G2/M phases of the cell cycle (15). This function highlights the
importance of POLR2L in facilitating the production of proteins and
RNAs essential for cell proliferation (2). Additionally, POLR2L is required for
cellular differentiation associated with both development and
tissue regeneration (3). POLR2L
inhibition disrupts the transcriptional patterns of differentiating
cells, potentially reducing the expression of proteins required for
appropriate differentiation and cell survival (34).
POLR2L and stress responses
Cells regulate gene expression when adapting to
oxidative stress or hypoxia, among other conditions (15). POLR2L then plays a crucial role
during transcription by controlling the expression of stress
response genes (1). POLR2L
stabilizes Pol II complexes at the promoters of stress-inducible
genes, enabling cells to respond rapidly to damage (3). This function is particularly
significant when cells such as cancer cells become adapted to
stress. Thus, POLR2L promotes survival by enhancing cellular
resilience under adverse conditions (16).
POLR2L in cellular metabolism and
energy regulation
POLR2L regulates cellular metabolism and the energy
balance. Many genes transcribed by Pol II are essential for energy
production and metabolism. POLR2L facilitates the transcription of
genes that regulate metabolism, thereby maintaining ATP production
and the equilibrium among metabolic intermediates. Disrupted POLR2L
expression is associated with metabolic imbalances, potentially
affecting the metabolic adaptation of cancer cells (16).
POLR2L and regulation of gene
expression
The roles played by POLR2L in transcription aside,
the subunit also contributes to epigenetic regulation (42). As part of the Pol II complex,
POLR2L interacts with histone-modifying enzymes and DNA
methyltransferases, as well as transcription factors and
co-activators that regulate gene expression (43,44).
Such epigenetic functions underscore the fact that POLR2L modifies
the transcriptional programs of specific cell types that respond to
changed environmental conditions. In addition to maintaining
transcriptional efficiency and stability, POLR2L is actively
involved in cell growth, differentiation, stress responses, and
metabolic regulation. POLR2L dysregulation has been linked to the
development of various diseases, yielding valuable research
insights and identification of potentially therapeutic targets
(16).
POLR2L and ncRNA regulation
In addition to its role in protein-coding gene
transcription, POLR2L may influence the regulation of ncRNAs
through Pol II-mediated transcriptional control. RNA polymerase II
is responsible for the transcription of numerous long ncRNAs and
microRNA precursors, suggesting that POLR2L participates indirectly
in ncRNA biogenesis (52–54). The dysregulation of POLR2L
expression may be linked with altered transcriptional programs
involving oncogenic long ncRNAs and microRNAs that regulate cancer
cell proliferation, metastasis, and drug resistance (53,55).
Although direct physical interactions between POLR2L and ncRNAs
have not yet been clearly demonstrated, POLR2L-dependent
transcriptional regulation likely shapes ncRNA expression
landscapes in disease contexts. Further studies employing RNA
immunoprecipitation and single-cell transcriptomic analyses will be
required to elucidate the mechanistic links between POLR2L and
ncRNA networks (8,56,57).
POLR2L and disease
POLR2L regulates gene expression during various
physiological processes. POLR2L dysregulation has been increasingly
associated with pathogenesis, particularly cancer, where increased
POLR2L expression may contribute to tumor progression, drug
resistance, and modulation of the immune response (16). Transcriptomic data revealed that
POLR2L expression was significantly elevated in several tumor types
compared to the corresponding normal tissues, including diffuse
large B-cell lymphoma (DLBC), glioblastoma multiforme (GBM), liver
hepatocellular carcinoma (LIHC), ovarian cancer (OV), thymoma
(THYM), prostate adenocarcinoma (PRAD), and stomach adenocarcinoma
(STAD) (Fig. 3A). POLR2L
expression levels were linked to survival rates of patients with
liver hepatocellular carcinoma (HCC), adrenocortical carcinoma,
skin cutaneous melanoma, and uveal melanoma (Fig. 3B). POLR2L can thus serve as both a
prognostic biomarker of and a possible therapeutic target for
cancer.
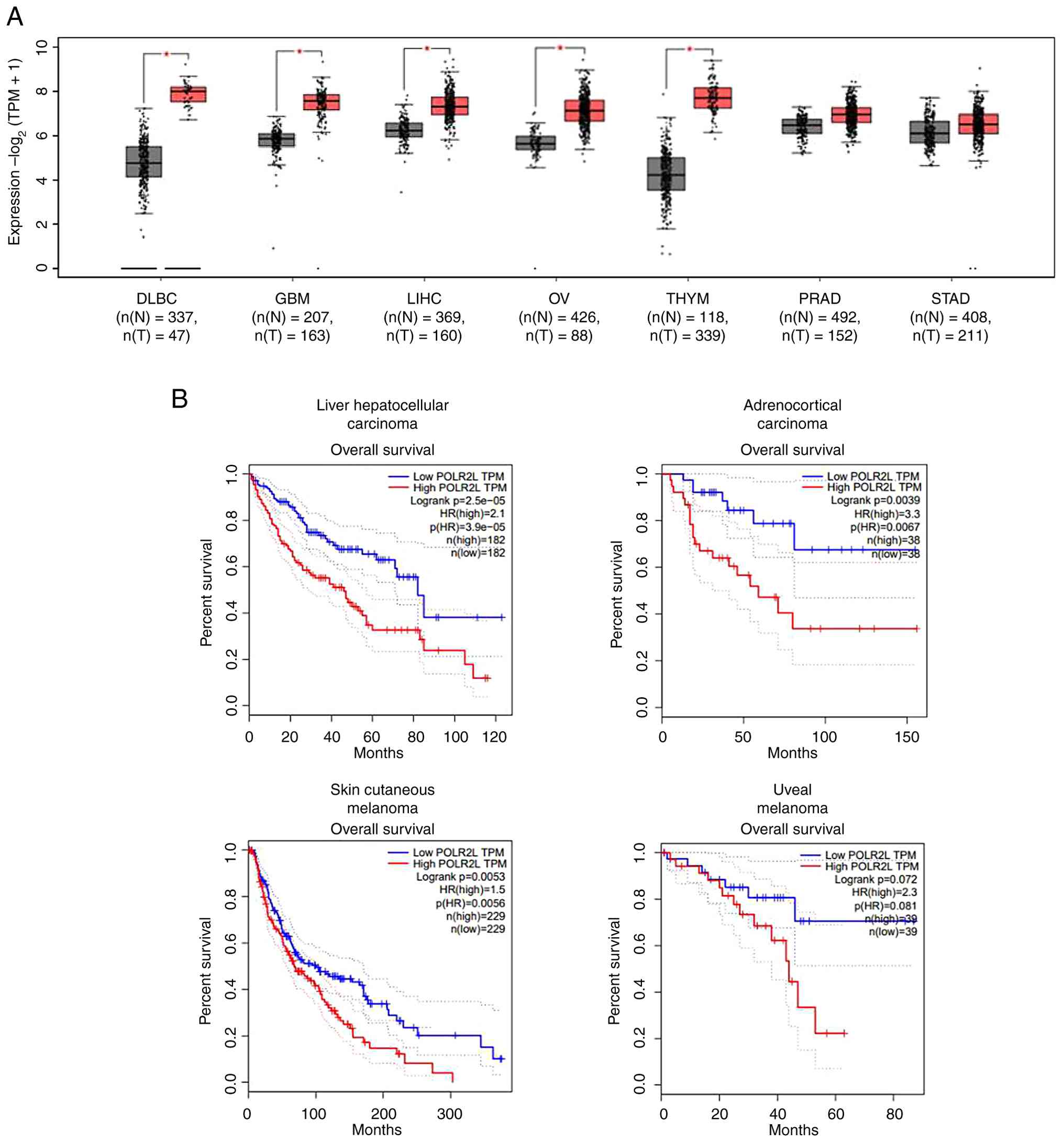 | Figure 3.POLR2L expression in various tumors
and its impact on survival rates. In cancer research, POLR2L
expression was found to be significantly elevated or showed an
increasing trend in seven types of cancer. (A) The expression
levels of POLR2L in normal tissues (gray) and tumor tissues (red)
in seven cancer types in which POLR2L expression was relatively
high (DLBC, GBM, LIHC, OV, THYM, PRAD and STAD). (B) Survival rates
are shown in relation to POLR2L expression across different tumor
types. *P<0.05. DLBC, lymphoid neoplasm diffuse large B-cell
lymphoma; GBM, glioblastoma multiforme; LIHC, liver hepatocellular
carcinoma; OV, ovarian serous cystadenocarcinoma; THYM, Thymoma;
PRAD, prostate adenocarcinoma; STAD, stomach adenocarcinoma; ACC,
adrenocortical carcinoma; SKCM, skin cutaneous melanoma; UVM, uveal
melanoma; POLR2L, RNA polymerase II, I and III subunit L. |
POLR2L is highly expressed in HCC, promoting tumor
cell proliferation and metastasis (45,46).
Such overexpression activates the survival pathways of cancer
cells, contributing to resistance against chemotherapeutic agents
such as cisplatin (17). POLR2L
inhibition reduced HCC viability and induced apoptosis,
highlighting the fact that POLR2L may serve as a therapeutic target
for liver cancer (17).
Increased POLR2L expression has also been observed
in gastric cancer, associated with poorer patient prognosis
(38). Patients with elevated
POLR2L levels often exhibit more rapid tumor progression and
enhanced resistance to chemotherapy (16). Targeting of POLR2L structural
stability may be useful in the treatment of gastric cancer
(16).
POLR2L is a key regulator of prostate cancer
malignancy and drug resistance (38,58).
POLR2L promotes cell invasiveness by activating TGF-β and
epithelial-mesenchymal transition (EMT) pathways (17). POLR2L suppression enhanced
sensitivity to anti-androgen therapies such as bicalutamide and
chemotherapeutic agents such as docetaxel, further highlighting the
potential of POLR2L as a therapeutic target (16,59).
POLR2L in viral infections
POLR2L facilitates viral transcription and
modulation of host immune responses. POLR2L enhances hepatitis B
virus gene expression by allowing the virus to use host
transcriptional machinery for replication (60). POLR2L inhibition reduced hepatitis
B virus replication, suggesting that POLR2L targeting may exert
antiviral effects (60). POLR2L is
also essential for herpes simplex virus gene expression. POLR2L
knockdown significantly impaired replication, underscoring its
critical role in the viral life cycle and potential as a
therapeutic target.
POLR2L in immune disorders
POLR2L regulates the immune responses of autoimmune
and inflammatory diseases by controlling inflammatory signaling and
cytokine production. POLR2L overexpression was associated with
excessive immune activation (61).
POLR2L modulated the expression of various inflammatory cytokines
including interleukins (ILs) and tumor necrosis factor (TNF) that
contribute to the pathogenesis of autoimmune diseases (62). POLR2L modulated the expression of
immune checkpoint proteins including PD-1 and CTLA-4 that control
immune cell activation (63).
POLR2L therefore serves as a promising target for novel
immunotherapies, particularly for patients with cancer (64).
POLR2L and DNA damage repair
POLR2L plays a crucial role in terms of DNA damage
repair by maintaining genomic stability. POLR2L coordinates
expression of the transcriptional and DNA repair mechanisms to
ensure that damage is rapidly remedied. POLR2L deficiencies impair
DNA repair and induce genomic instability, increasing the risk of
tumorigenesis. The involvement of POLR2L in the preservation of
genomic integrity indicates that POLR2L may serve as a therapeutic
target for cancers associated with defective DNA repair (65).
Thus, POLR2L plays critical roles in various
pathological processes, including cancer, viral infections, immune
disorders, and DNA damage repair. POLR2L dysregulation contributes
to both disease onset and progression, highlighting its potential
as a therapeutic target (66). A
deeper understanding of how POLR2L affects disease could lead to
the development of novel diagnostic tools and personalized
treatment strategies, thus advancing the field of precision
medicine (67).
POLR2L as a diagnostic and prognostic
biomarker
Accumulating transcriptomic and clinical evidence
supports the potential of POLR2L as a diagnostic and prognostic
biomarker in cancer (17,68). Elevated POLR2L expression has been
consistently associated with advanced tumor stage, poor prognosis,
and reduced overall survival in multiple malignancies, including
hepatocellular carcinoma, gastric cancer, and prostate cancer
(Table I). These associations
suggest that POLR2L expression levels may reflect tumor
aggressiveness and transcriptional dependency (38,59,69,70).
 | Table I.POLR2L as a biomarker in human
cancer. |
Table I.
POLR2L as a biomarker in human
cancer.
| Cancer type | POLR2L
expression | Clinical
association | Potential
application |
|---|
| Hepatocellular
carcinoma | Upregulated | Poor prognosis,
chemoresistance | Prognostic
biomarker |
| Gastric cancer | Upregulated | Advanced stage,
reduced survival | Prognostic
marker |
| Prostate
cancer | Upregulated | EMT, drug
resistance | Predictive
biomarker |
| Glioblastoma
multiforme | Elevated | Tumor
aggressiveness | Diagnostic
support |
| Multiple
cancers | Increased in
tumors | Survival
correlation | Liquid biopsy
(potential) |
Importantly, POLR2L may also be applicable in
non-invasive clinical settings. Advances in liquid biopsy
technologies have enabled the detection of tumor-derived RNA
signatures in circulating blood, raising the possibility that
POLR2L mRNA levels could be monitored using circulating tumor RNA
or exosomal RNA (56). Although
direct clinical validation remains limited, these approaches
highlight the translational potential of POLR2L as a minimally
invasive biomarker for cancer diagnosis, prognosis, and therapeutic
response monitoring (46,71).
POLR2L and signaling pathways
POLR2L regulates multiple signaling pathways via
transcriptional control, impacting cell survival, differentiation,
stress responses, and cancer progression (16). Therefore, POLR2L involvement in key
signaling pathways has gained increasing attention, particularly in
the context of cancer and other pathological conditions (17).
PI3K-Akt signaling pathway
The PI3K-Akt pathway is a key regulator of cell
survival, growth, and transcription, and plays critical roles in
both cancer cell proliferation and therapy resistance (72). POLR2L activates the pathway by
modulating Pol II-mediated gene expression. POLR2L regulates the
transcription of genes essential for Akt activation, thereby
supporting cancer cell survival and metabolism (72) (Fig.
4). POLR2L overexpression enhances PI3K-Akt signaling,
promoting the G1/S cell cycle transition and driving cancer cell
proliferation (73). In patients
with HCC and gastric cancer, POLR2L has been implicated in the
development of chemoresistance mediated by the PI3K-Akt pathway
(74). POLR2L suppression
deactivates the pathway and triggers cancer cell apoptosis, further
highlighting the key role played by POLR2L as a regulator of
PI3K-Akt signaling (75).
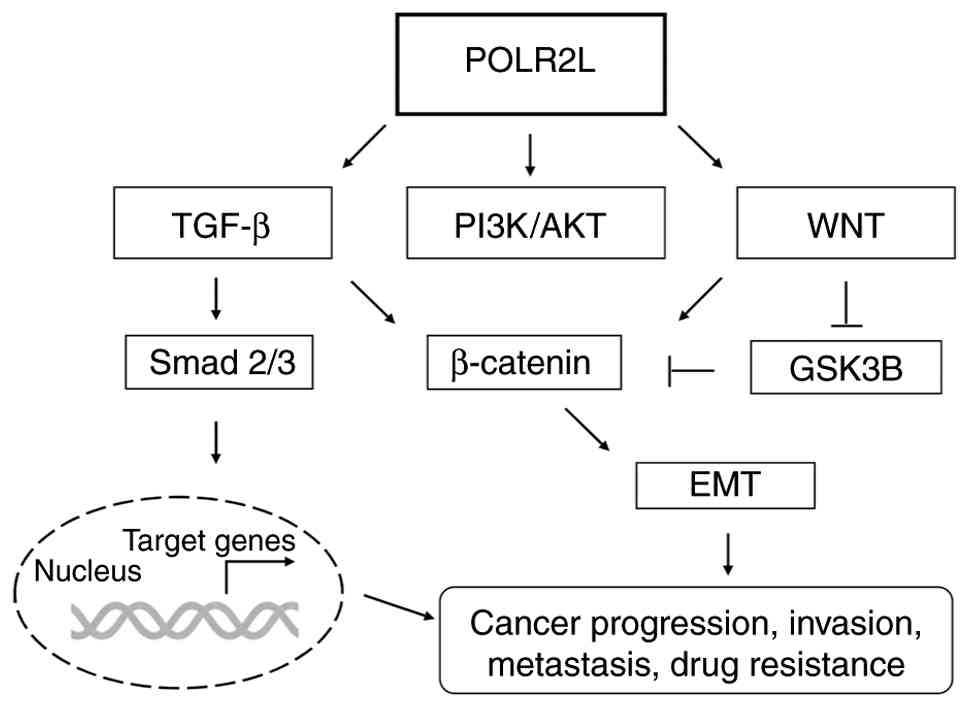 | Figure 4.POLR2L function in TGF-β, WNT and
PI3K/AKT signaling pathways. POLR2L enhances cell survival and
metabolism through the PI3K/AKT pathway and facilitates the
transcription of target genes in the nucleus by promoting Smad2/3
phosphorylation via the TGF-β pathway. Additionally, POLR2L
contributes to the activation of β-catenin through the WNT pathway,
thereby increasing EMT, a process also influenced by TGF-β
signaling. By promoting ETM through these pathways, POLR2L plays a
significant role in enhancing cancer proliferation, metastasis,
invasion and drug resistance. EMT, epithelial-mesenchymal
transition; POLR2L, RNA polymerase II, I and III subunit L. |
Wnt/β-catenin signaling pathway
The Wnt/β-catenin pathway is essential for cell
differentiation and tissue homeostasis, and plays major roles in
cancer development and metastasis (76). POLR2L modulates this pathway by
regulating β-catenin transcription and activating downstream target
genes. POLR2L enhanced the transcription of Wnt target genes,
including CCND1 and MYC (76).
Activation of β-catenin via POLR2L-dependent transcription promoted
cancer cell proliferation and motility (77) (Fig.
4). POLR2L regulated the Wnt/β-catenin pathways of prostate and
liver cancers (78). POLR2L
inhibition triggered pathway deactivation, reducing cancer cell
invasiveness and metastasis (77).
These findings suggest that POLR2L could serve as a promising
therapeutic target when seeking to block Wnt/β-catenin signaling in
cancer patients (77).
TGF-β signaling pathway
The TGF-β signaling pathway plays key roles in
various biological processes, including growth inhibition, immune
regulation, and epithelial-to-mesenchymal transition (EMT). POLR2L
regulates the expression of TGF-β-induced transcription factors,
promoting the EMT and enhancing cancer invasiveness (17). Via TGF-β signaling, POLR2L
facilitates the EMT, enabling cancer cells to acquire metastatic
properties and develop drug resistance (17) (Fig.
4). POLR2L inhibition downregulated the expression of
EMT-related genes such as ZEB1 and SNAI1, thereby reducing prostate
and gastric cancer cell migration and invasiveness (79). POLR2L increased the activation of
immunosuppressive cells, including regulatory T cells and M2
macrophages, in the tumor microenvironment, in turn promoting
immune evasion (80). These
findings highlight the potential of POLR2L as a cancer
immunotherapy target (80).
NF-κB signaling pathway
POLR2L also regulates the NF-κB signaling pathway,
which plays essential roles in both inflammation and cancer
progression (81). NF-κB controls
the expression of cytokines, inflammatory proteins, and survival
genes (36), and POLR2L
facilitates transcription of these genes (82). Elevated POLR2L expression has been
linked to NF-κB activation, which has intensified inflammatory
responses and contributed to the development of tumor-promoting
inflammatory microenvironments (83). POLR2L inhibition reduced the
expression of inflammation-related genes, such as those encoding
TNF and IL6, suggesting that POLR2L knockdown may exert an
anti-inflammatory effect (82).
Interactions between POLR2L expression and NF-κB signaling have
been observed in patients with gastric cancer and HCC (17,84).
POLR2L suppression inhibited the signaling pathway, reducing tumor
growth and metastasis, again indicating that POLR2L may be a
valuable therapeutic target (83).
POLR2L interacts with various signaling pathways via
transcriptional regulation. Such pathways include the PI3K-Akt,
Wnt/β-catenin, TGF-β, and NF-κB pathways, which play crucial roles
in pathological conditions such as cancer (16). POLR2L dysregulation distorts these
signaling networks, promoting tumor survival and metastasis
(17). These findings position
POLR2L as both a potential therapeutic target and a diagnostic
biomarker, opening up new possibilities for understanding and
treating disease (16).
Therapeutic potential of POLR2L
POLR2L plays a vital role in transcriptional
regulation and significantly impacts the progression of cancer and
other diseases (16). POLR2L
dysregulation is strongly associated with tumor cell survival,
metastasis, and drug resistance, highlighting the potential of
POLR2L as a promising candidate for both disease diagnosis and
therapeutic intervention (17).
POLR2L inhibition may exert anticancer effects.
POLR2L is overexpressed in cancer cells; POLR2L suppression reduced
cell viability, inhibited metastasis, and induced apoptosis
(16). Advanced technologies such
as RNA interference (siRNA, shRNA) and CRISPR-Cas9 have been used
to downregulate POLR2L expression (85), in turn impairing transcriptional
efficiency followed by cell cycle arrest and apoptosis (16). Such effects have been particularly
pronounced in models of HCC, gastric cancer, and prostate cancer
(16). In the HCC context, POLR2L
inhibition suppressed cell proliferation and enhanced the
effectiveness of anti-POLR2L therapies in combination with
chemotherapeutic agents (17).
Similarly, POLR2L suppression reduced metastatic potential in
gastric and prostate cancers by inhibiting TGF-β and Wnt/β-catenin
signaling. These findings highlight the therapeutic promise of
POLR2L targeting in various cancer cells (16).
Another therapeutic strategy involves POLR2L
targeting to overcome cancer-associated drug resistance. POLR2L is
a key contributor to chemoresistance, and POLR2L inhibition may
therefore be valuable. POLR2L enhances resistance to
chemotherapeutic agents such as cisplatin and bicalutamide by
activating PI3K-Akt and NF-κB signaling. POLR2L suppression
restored the drug sensitivity of drug-resistant cancer cells,
suggesting a novel approach toward improvement of therapeutic
efficacy (16). POLR2L also
upregulates immune checkpoint proteins, including PD-1 and PD-L1,
in turn allowing cancer cells to evade immune detection (86). POLR2L targeting enhanced the
effectiveness of immune checkpoint inhibitors, suggesting that
anti-POLR2L agents might be useful components of combination
therapies (86).
POLR2L exhibits significant potential as both a
diagnostic and prognostic biomarker of cancers and other diseases
(16). POLR2L expression levels
are strongly correlated with tumor progression, rendering POLR2L a
valuable disease severity indicator (17). For example, POLR2L expression has
predicted the tumor size and metastatic status of HCC and gastric
cancer (17). Analyzing POLR2L
expression levels may aid non-invasive diagnosis. Its
overexpression has been linked to lower survival rates and
increased risk of metastasis, both of which are critical to patient
outcomes (17). Thus, POLR2L
expression levels may guide the development of personalized
therapeutic strategies, in turn enhancing precision medicine
(16).
Direct POLR2L targeting is emerging as a promising
cancer treatment (16). The
structural characteristics of POLR2L, particularly the atypical
zinc finger domain, render it an ideal target of small-molecule
inhibitors (2) that selectively
block POLR2L function to suppress tumor growth and metastasis
(16). Combining POLR2L inhibitors
with existing chemotherapies or immune checkpoint inhibitors has
enhanced therapeutic efficacy. For example, an anti-POLR2L agent
and cisplatin exhibited synergistic effects against HCC and gastric
cancer (17). POLR2L is a key
regulator of transcription in cancer and other diseases, and is
therefore promising as both a therapeutic target and a biomarker
(16). POLR2L inhibition reduced
tumor cell survival, overcame drug resistance, and enhanced the
efficacy of immunotherapy (87).
Drugs that specifically target POLR2L as part of new combination
therapies may become a new cancer treatment paradigm, paving the
way for more effective personalized therapeutic strategies
(88).
Small-molecule-targeting strategies
for POLR2L
Although direct small-molecule inhibitors
specifically targeting POLR2L are not yet clinically available,
emerging evidence suggests that POLR2L represents a
pharmacologically tractable target (25,89).
Structural analyses have indicated that the atypical zinc finger
domain of POLR2L plays a critical role in stabilizing Pol II
assembly, making this motif a potential site for structure-guided
inhibitor design. Small molecules that disrupt zinc coordination or
POLR2L-mediated subunit interactions may impair transcriptional
stability in POLR2L-dependent cancer cells (25,90).
Indirect pharmacological approaches have also been
proposed. Cancer cells exhibiting POLR2L overexpression appear to
display heightened dependency on Pol II transcriptional stress.
Thus, transcriptional inhibitors or compounds that destabilize Pol
II assembly may exert selective anticancer effects in tumors with
high POLR2L expression (91).
These findings support the rationale for developing POLR2L-centered
therapeutic strategies, either through direct inhibition or via
synthetic lethality-based approaches (37,91).
Discussion and future perspectives
POLR2L, a crucial Pol II component, plays
fundamental roles in cell growth, differentiation, stress
responses, metabolic regulation, and disease progression (1). This review comprehensively analyzed
POLR2L in terms of its structural characteristics, physiological
functions, dysregulated expression in disease, interactions with
key signaling pathways, and potential as a therapeutic target. By
explicitly placing POLR2L in a structural and evolutionary context
with POLR2H and POLR2K, this review refines current models of Pol
II subunit organization and suggests testable hypotheses regarding
the differential contribution of individual small subunits to
complex assembly, transcriptional plasticity, and oncogenic
transcriptional dependency (35,92–94).
AlphaFold POLR2L structural prediction offers valuable insights
into the functional mechanisms in play and potential POLR2L
interaction partners, aiding further exploration of their
biological significance. The multifunctional nature of POLR2L
underscores its critical roles in the development and progression
of various diseases. POLR2L has become a key focus of researchers
who seek innovative diagnostic and therapeutic strategies (17). POLR2L is thought to support Pol II
assembly and transcriptional competence, thereby connecting its
structural role to disease associated transcriptional programs
(34). POLR2L contains an atypical
zinc finger domain that is thought to support efficient
transcription and stabilization in RNA-DNA hybrids, although the
precise mechanistic contribution of this motif remains incompletely
defined (2). POLR2L is highly
conserved across all eukaryotic species because POLR2L plays a
fundamental role in transcriptional regulation (3). POLR2L is overexpressed in several
cancers, including HCC, gastric cancer, and prostate cancer
(16). Increased POLR2L expression
has been strongly linked to tumor cell proliferation, metastasis,
and drug resistance (17).
Notably, POLR2L inhibition reduced cancer cell viability and
enhanced drug sensitivity, highlighting the potential of POLR2L as
a promising therapeutic target (16).
POLR2L contributes to cancer progression and immune
regulation by modulating transcription via key signaling pathways,
including the PI3K-Akt, Wnt/β-catenin, TGF-β, and NF-κB pathways
(16). POLR2L interacts with both
upstream and downstream components of these pathways to regulate
key disease-related processes including cell survival, metastasis,
and immune suppression (17).
Thus, POLR2L may represent a valuable diagnostic and prognostic
biomarker. POLR2L levels are strongly correlated with cancer
progression and predict both patient survival and metastatic
potential. Additionally, POLR2L may be useful a non-invasive
clinical diagnostic tool, and POLR2L inhibitors may offer novel
therapeutics for cancer and other diseases (95). POLR2L targeting may overcome the
drug resistance of cancer cells and improve the effectiveness of
chemotherapy and immunotherapy. Personalized therapeutic strategies
centered on POLR2L are required (88).
Despite these significant advances, the details of
how POLR2L interacts with transcriptional networks remain
incompletely understood. Further studies, including RNA sequencing
and protein-protein interaction network analyses, are essential.
Although the role played by POLR2L in cancer is well established,
POLR2L involvement in viral infections, immune disorders, and
metabolic diseases remains relatively unexplored. More research is
essential. Additionally, anti-POLR2L drug development is still at
an early stage (16). Further
studies must optimize the stability and efficacy of POLR2L
inhibitors in both preclinical and clinical settings to ensure that
such materials will be therapeutically active (96).
Future research will further elucidate the roles
played by POLR2L in cancers, immune disorders, and infectious
diseases, and explore whether POLR2L targeting might advance
personalized treatment. The details of how POLR2L interacts with
key signaling pathways will allow the development of combination
therapies that target both POLR2L and other critical components.
Preclinical and clinical studies will assess the stability and
efficacy of POLR2L inhibitors. POLR2L inhibitors should be combined
with existing chemotherapies and immunotherapies. The roles played
by POLR2L in viral infections and autoimmune diseases require
further study prior to the development of novel POLR2L-targeting
therapies (88). CRISPR-Cas9 gene
editing, single-cell RNA sequencing, and bioinformatics studies are
required (85). to better
understand the transcriptional networks of POLR2L and the role
played by POLR2L in disease progression (97).
In summary, POLR2L, a crucial component of the Pol
II complex, plays central roles in the progression of cancer and
other diseases. Research on POLR2L has evolved from study of the
structural and functional characteristics of the protein to
exploration of whether POLR2L might serve as a valuable therapeutic
target and diagnostic biomarker. Future studies will create new
therapeutic strategies for cancers and other complex diseases,
ultimately transforming treatment and fostering precision
medicine.
Acknowledgements
Not applicable.
Funding
This work was financially supported by research fund of Chungnam
National University (SH Kim, 2025) and by the National Research
Foundation of Korea grant funded by the Korea government (MSIT)
(grant no. RS-2025-00514760).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
BL, CS, SE, YL, SK and JS contributed to the study
conception and design. The first draft of the manuscript was
written by BL and JP, and all authors commented on previous
versions of the manuscript. All authors read and approved the final
manuscript. Data authentication is not applicable.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Cramer P, Armache KJ, Baumli S, Benkert S,
Brueckner F, Buchen C, Damsma GE, Dengl S, Geiger SR, Jasiak AJ, et
al: Structure of eukaryotic RNA polymerases. Annu Rev Biophys.
37:337–352. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cramer P, Bushnell DA and Kornberg RD:
Structural basis of transcription: RNA polymerase II at 2.8
angstrom resolution. Science. 292:1863–1876. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Acker J, de Graaff M, Cheynel I, Khazak V,
Kedinger C and Vigneron M: Interactions between the human RNA
polymerase II subunits. J Biol Chem. 272:16815–16821. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kolodziej PA and Young RA: Mutations in
the three largest subunits of yeast RNA polymerase II that affect
enzyme assembly. Mol Cell Biol. 11:4669–4678. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Cramer P: RNA polymerase II structure:
From core to functional complexes. Curr Opin Genet Dev. 14:218–226.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Schier AC and Taatjes DJ: Structure and
mechanism of the RNA polymerase II transcription machinery. Genes
Dev. 34:465–488. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Gillis A and Berry S: Global control of
RNA polymerase II. Biochim Biophys Acta Gene Regul Mech.
1867:1950242024. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ciesla M, Turowski TW, Nowotny M,
Tollervey D and Boguta M: The expression of Rpb10, a small subunit
common to RNA polymerases, is modulated by the R3H
domain-containing Rbs1 protein and the Upf1 helicase. Nucleic Acids
Res. 48:12252–12268. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ryu J and Lee C: RNA polymerase subunits
and ribosomal proteins: An overview and their genetic impact on
complex human traits. Front Biosci (Landmark Ed). 29:1852024.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Guo Z and Stiller JW: Comparative genomics
and evolution of proteins associated with RNA polymerase II
C-terminal domain. Mol Biol Evol. 22:2166–2178. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Purkayastha D and Karmodiya K: RNA
polymerase II evolution and adaptations: Insights from Plasmodium
and other parasitic protists. Infect Genet Evol. 115:1055052023.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jumper J, Evans R, Pritzel A, Green T,
Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A,
Potapenko A, et al: Highly accurate protein structure prediction
with AlphaFold. Nature. 596:583–589. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Sawadogo M and Sentenac A: RNA polymerase
B (II) and general transcription factors. Annu Rev Biochem.
59:711–754. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Myer VE and Young RA: RNA polymerase II
holoenzymes and subcomplexes. J Biol Chem. 273:27757–27760. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Dieci G, Fiorino G, Castelnuovo M,
Teichmann M and Pagano A: The expanding RNA polymerase III
transcriptome. Trends Genet. 23:614–622. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sadurni MM and Saponaro M: Deregulations
of RNA Pol II subunits in cancer. Appl Biosci. 2:459–476. 2023.
View Article : Google Scholar
|
|
17
|
Chen J, Xu J, Li L, Yuan Y, Jiang J and
Sun Y: Propofol regulates the progression of hepatocellular
carcinoma via the POLR2L/TGF-beta signaling pathway. Transl Cancer
Res. 13:2266–2281. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lv Z, Wu X, Lu P, Xu X, Wang J, Zhang C,
Liu W, Gao Y, Lu C, Zhang Y and Kou H: POLE2 knockdown suppresses
lymphoma progression via downregulating Wnt/beta-catenin signaling
pathway. Mol Cell Biochem. 479:487–497. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Roszkowska M: Multilevel mechanisms of
cancer drug resistance. Int J Mol Sci. 25:124022024. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Khan SU, Fatima K, Aisha S and Malik F:
Unveiling the mechanisms and challenges of cancer drug resistance.
Cell Commun Signal. 22:1092024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wei L, Levine AS and Lan L:
Transcription-coupled homologous recombination after oxidative
damage. DNA Repair (Amst). 44:76–80. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dumay-Odelot H, Durrieu-Gaillard S, Da
Silva D, Roeder RG and Teichmann M: Cell growth- and
differentiation-dependent regulation of RNA polymerase III
transcription. Cell Cycle. 9:3687–3699. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Acker J, Murroni O, Mattei MG, Kedinger C
and Vigneron M: The gene (POLR2L) encoding the hRPB7.6 subunit of
human RNA polymerase. Genomics. 32:86–90. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Cramer P: Multisubunit RNA polymerases.
Curr Opin Struct Biol. 12:89–97. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Mackereth CD, Arrowsmith CH, Edwards AM
and McIntosh LP: Zinc-bundle structure of the essential RNA
polymerase subunit RPB10 from Methanobacterium thermoautotrophicum.
Proc Natl Acad Sci USA. 97:6316–6321. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Sainsbury S, Bernecky C and Cramer P:
Structural basis of transcription initiation by RNA polymerase II.
Nat Rev Mol Cell Biol. 16:129–143. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Donaldson IM and Friesen JD: Zinc
stoichiometry of yeast RNA polymerase II and characterization of
mutations in the zinc-binding domain of the largest subunit. J Biol
Chem. 275:13780–13788. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sievers F, Wilm A, Dineen D, Gibson TJ,
Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, et al:
Fast, scalable generation of high-quality protein multiple sequence
alignments using Clustal Omega. Mol Syst Biol. 7:5392011.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kontermann R and Bautz EK: Similarity
between subunit 8 of yeast RNA polymerase II (RPB8) and the
second-largest subunits of eukaryotic RNA polymerases. Nucleic
Acids Res. 20:52311992. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Sakurai H and Ishihama A: Transcription
organization and mRNA levels of the genes for all 12 subunits of
the fission yeast RNA polymerase II. Genes Cells. 6:25–36. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Rubbi L, Labarre-Mariotte S, Chedin S and
Thuriaux P: Functional characterization of ABC10alpha, an essential
polypeptide shared by all three forms of eukaryotic DNA-dependent
RNA polymerases. J Biol Chem. 274:31485–31492. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Cramer P, Bushnell DA, Fu J, Gnatt AL,
Maier-Davis B, Thompson NE, Burgess RR, Edwards AM, David PR and
Kornberg RD: Architecture of RNA polymerase II and implications for
the transcription mechanism. Science. 288:640–649. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Buratowski S: Progression through the RNA
polymerase II CTD cycle. Mol Cell. 36:541–546. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Shpakovski GV, Acker J, Wintzerith M,
Lacroix JF, Thuriaux P and Vigneron M: Four subunits that are
shared by the three classes of RNA polymerase are functionally
interchangeable between Homo sapiens and Saccharomyces cerevisiae.
Mol Cell Biol. 15:4702–4710. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Turowski TW and Boguta M: Specific
features of RNA polymerases I and III: Structure and assembly.
Front Mol Biosci. 8:6800902021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Juang U, Lee S, Gwon S, Jung W, Nguyen H,
Huang Q, Lee B, Kwon SH, Kim SH, Kim IS, et al: Enhancement of
renal fibrosis in PHF20 transgenic mice. Toxicol Res. 41:71–80.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Trinh V, Langelier MF, Archambault J and
Coulombe B: Structural perspective on mutations affecting the
function of multisubunit RNA polymerases. Microbiol Mol Biol Rev.
70:12–36. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Zhou D, Li X, Zhao H, Sun B, Liu A, Han X,
Cui Z and Yuan L: Combining multi-dimensional data to identify a
key signature (gene and miRNA) of cisplatin-resistant gastric
cancer. J Cell Biochem. 119:6997–7008. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Woychik NA and Young RA: RNA polymerase II
subunit RPB10 is essential for yeast cell viability. J Biol Chem.
268:122301993. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Korkhin Y, Unligil UM, Littlefield O,
Nelson PJ, Stuart DI, Sigler PB, Bell SD and Abrescia NG: Evolution
of complex RNA polymerases: The complete archaeal RNA polymerase
structure. PLoS Biol. 7:e10001022009. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Garrido-Godino AI, Gupta I, Pelechano V
and Navarro F: RNA Pol II assembly affects ncRNA expression. Int J
Mol Sci. 25:5072023. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Gagnidze K and Pfaff DW: Epigenetic
mechanisms: DNA methylation and histone protein modification. In:
Neuroscience in the 21st Century: From Basic to Clinical. Pfaff DW,
Volkow ND and Rubenstein J: Springer; New York, New York, NY: pp.
1–40. 2020
|
|
43
|
Robertson KD: Epigenetic mechanisms of
gene regulation. DNA methylation and cancer therapy. Springer US;
Boston, MA: pp. 13–30. 2005, View Article : Google Scholar
|
|
44
|
Juang U, Gwon S, Jung W, Nguyen H, Huang
Q, Lee S, Lee B, Kwon SH, Kim SH and Park J: Exploring the various
functions of PHD finger protein 20: Beyond the unknown. Toxicol
Res. 41:1–11. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Yao F, Zhan Y, Li C, Lu Y, Chen J, Deng J,
Wu Z, Li Q, Song Y, Chen B, et al: Single-Cell RNA sequencing
reveals the role of phosphorylation-related genes in hepatocellular
carcinoma stem cells. Front Cell Dev Biol. 9:7342872021. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Lu Y, Wang S, Chi T, Zhao Y, Guo H, Wang H
and Feng L: DNA damage repair-related gene signature for
identifying the immune status and predicting the prognosis of
hepatocellular carcinoma. Sci Rep. 13:189782023. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Kettenberger H, Armache KJ and Cramer P:
Complete RNA polymerase II elongation complex structure and its
interactions with NTP and TFIIS. Mol Cell. 16:955–965. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Woychik NA and Young RA: RNA polymerase II
subunit RPB10 is essential for yeast cell viability. J Biol Chem.
265:17816–17819. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Hu Y and Liu B: Roles of zinc-binding
domain of bacterial RNA polymerase in transcription. Trends Biochem
Sci. 47:710–724. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Li Y, Huang J, Zhu J, Bao L, Wang H, Jiang
Y, Tian K, Wang R, Zheng H, Duan W, et al: Targeted protein
degradation reveals RNA Pol II heterogeneity and functional
diversity. Mol Cell. 82:3943–3959.e3911. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Ehara H, Yokoyama T, Shigematsu H,
Yokoyama S, Shirouzu M and Sekine SI: Structure of the complete
elongation complex of RNA polymerase II with basal factors.
Science. 357:921–924. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek
SH and Kim VN: MicroRNA genes are transcribed by RNA polymerase II.
EMBO J. 23:4051–4060. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Jia W, Chen W and Kang J: The functions of
microRNAs and long non-coding RNAs in embryonic and induced
pluripotent stem cells. Genomics Proteomics Bioinformatics.
11:275–283. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Mattick JS, Amaral PP, Carninci P,
Capenter S, Chang HY, Chen LL, Chen R, Dean C, Dinger ME,
Fitzgerald KA, et al: Long non-coding RNAs: Definitions, functions,
challenges and recommendations. Nat Rev Mol Cell Biol. 24:430–447.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Wang J, Ye C, Xiong H, Shen Y, Lu Y, Zhou
J and Wang L: Dysregulation of long non-coding RNA in breast
cancer: An overview of mechanism and clinical implication.
Oncotarget. 8:5508–5522. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Wei DM, Jiang MT, Lin P, Yang H, Dang YW,
Yu Q, Liao DY, Luo DZ and Chen G: Potential ceRNA networks involved
in autophagy suppression of pancreatic cancer caused by chloroquine
diphosphate: A study based on differentially-expressed circRNAs,
lncRNAs, miRNAs and mRNAs. Int J Oncol. 54:600–626. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zou RC, Shi ZT, Xiao SF, Ke Y, Tang HR, Wu
TG, Guo ZT, Ni F, An S and Wang L: Co-expression analysis and ceRNA
network reveal eight novel potential lncRNA biomarkers in
hepatocellular carcinoma. PeerJ. 7:e81012019. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Cui X, Li C, Ding J, Yao Z, Zhao T, Guo J,
Wang Y and Li J: Establishing a proteomics-based signature of
AKR1C3-related genes for predicting the prognosis of prostate
cancer. Int J Mol Sci. 24:45132023. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Zhang D, Xu X, Wei Y, Chen X, Li G, Lu Z,
Zhang X, Ren X, Wang S and Qin C: Prognostic role of DNA damage
response genes mutations and their association with the sensitivity
of olaparib in prostate cancer patients. Cancer Control.
29:107327482211294512022. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Wang H and Ryu WS: Hepatitis B virus
polymerase blocks pattern recognition receptor signaling via
interaction with DDX3: Implications for immune evasion. PLoS
Pathog. 6:e10009862010. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
de Gruijter NM, Jebson B and Rosser EC:
Cytokine production by human B cells: Role in health and autoimmune
disease. Clin Exp Immunol. 210:253–262. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Yasmeen F, Pirzada RH, Ahmad B, Choi B and
Choi S: Understanding autoimmunity: Mechanisms, predisposing
factors, and cytokine therapies. Int J Mol Sci. 25:76662024.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Zhang H, Dai Z, Wu W, Wang Z, Zhang N,
Zhang L, Zeng WJ, Liu Z and Cheng Q: Regulatory mechanisms of
immune checkpoints PD-L1 and CTLA-4 in cancer. J Exp Clin Cancer
Res. 40:1842021. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Wojtukiewicz MZ, Rek MM, Karpowicz K,
Górska M, Polityńska B, Wojtukiewicz AM, Moniuszko M, Radziwon P,
Tucker SC and Honn KV: Inhibitors of immune checkpoints-PD-1,
PD-L1, CTLA-4-new opportunities for cancer patients and a new
challenge for internists and general practitioners. Cancer
Metastasis Rev. 40:949–982. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Bai Y, Wang W and Wang J: Targeting DNA
repair pathways: Mechanisms and potential applications in cancer
therapy. Genome Instability & Disease. 1:318–338. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Talukdar PD and Chatterji U:
Transcriptional co-activators: Emerging roles in signaling pathways
and potential therapeutic targets for diseases. Signal Transduct
Target Ther. 8:4272023. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Hartman J, Khanna V, Habib A, Farrokhyar
F, Memon M and Adili A: Perioperative systemic glucocorticoids in
total hip and knee arthroplasty: A systematic review of outcomes. J
Orthop. 14:294–301. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Li M, Liu Z, Wang J, Liu H, Gong H, Li S,
Jia M and Mao Q: Systematic analysis identifies a specific
RNA-binding protein-related gene model for prognostication and
risk-adjustment in HBV-related hepatocellular carcinoma. Front
Genet. 12:7073052021. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Sun J, Zhao J, Yang Z, Zhou Z and Lu P:
Identification of gene signatures and potential therapeutic targets
for acquired chemotherapy resistance in gastric cancer patients. J
Gastrointest Oncol. 12:407–422. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Cai J, Chen Z, Chen X, Huang H, Lin X and
Miao B: Coexpression network analysis identifies a novel Nine-RNA
signature to improve prognostic prediction for prostate cancer
patients. Biomed Res Int. 2020:42642912020. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Zhang S, Qin O, Wu S, Xu H, Huang W and
Hailiang S: A pyrimidine metabolism-related signature for
prognostic and immunotherapeutic response prediction in
hepatocellular carcinoma by integrating analyses. Aging (Albany
NY). 16:5545–5566. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Osaki M, Oshimura M and Ito H: PI3K-Akt
pathway: Its functions and alterations in human cancer. Apoptosis.
9:667–676. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Glaviano A, Foo ASC, Lam HY, Yap KCH,
Jacot W, Jones RH, Eng H, Nair MG, Makvandi P, Geoerger B, et al:
PI3K/AKT/mTOR signaling transduction pathway and targeted therapies
in cancer. Mol Cancer. 22:1382023. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Sun EJ, Wankell M, Palamuthusingam P,
McFarlane C and Hebbard L: Targeting the PI3K/Akt/mTOR pathway in
hepatocellular carcinoma. Biomedicines. 9:16392021. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Yang J, Nie J, Ma X, Wei Y, Peng Y and Wei
X: Targeting PI3K in cancer: Mechanisms and advances in clinical
trials. Mol Cancer. 18:262019. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Song P, Gao Z, Bao Y, Chen L, Huang Y, Liu
Y, Dong Q and Wei X: Wnt/β-catenin signaling pathway in
carcinogenesis and cancer therapy. J Hematol Oncol. 17:462024.
View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Zhang Y and Wang X: Targeting the
Wnt/β-catenin signaling pathway in cancer. J Hematol Oncol.
13:1652020. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Wang C, Chen Q and Xu H: Wnt/β-catenin
signal transduction pathway in prostate cancer and associated drug
resistance. Discov Oncol. 12:402021. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Parsana P, Amend SR, Hernandez J, Pienta
KJ and Battle A: Identifying global expression patterns and key
regulators in epithelial to mesenchymal transition through
multi-study integration. BMC Cancer. 17:4472017. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Imodoye SO and Adedokun KA: EMT-induced
immune evasion: Connecting the dots from mechanisms to therapy.
Clin Exp Med. 23:4265–4287. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Ma Q, Hao S, Hong W, Tergaonkar V, Sethi
G, Tian Y and Duan C: Versatile function of NF-kB in inflammation
and cancer. Exp Hematol Oncol. 13:682024. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Hoesel B and Schmid JA: The complexity of
NF-κB signaling in inflammation and cancer. Mol Cancer. 12:862013.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Park MH and Hong JT: Roles of NF-κB in
cancer and inflammatory diseases and their therapeutic approaches.
Cells. 5:152016. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Chen B, Cai T, Huang C, Zang X, Sun L, Guo
S, Wang Q, Chen Z, Zhao Y, Han Z, et al: G6PD-NF-κB-HGF signal in
gastric cancer-associated mesenchymal stem cells promotes the
proliferation and metastasis of gastric cancer cells by
upregulating the expression of HK2. Front Oncol. 11:6487062021.
View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Di Carlo E and Sorrentino C: State of the
art CRISPR-based strategies for cancer diagnostics and treatment.
Biomark Res. 12:1562024. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Ortega MA, Boaru DL, De Leon-Oliva D,
Fraile-Martinez O, García-Montero C, Rios L, Garrido-Gil MJ,
Barrena-Blázquez S, Minaya-Bravo AM, Rios-Parra A, et al:
PD-1/PD-L1 axis: Implications in immune regulation, cancer
progression, and translational applications. J Mol Med (Berl).
102:987–1000. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Mpiima DP, Salongo GW, Lugobe H, Ssemujju
A, Mulisya OM, Masinda A, Twizerimana H and Ngonzi J: Association
between prior chlamydia trachomatis infection and ectopic pregnancy
at a tertiary care hospital in South Western Uganda. Obstet Gynecol
Int. 2018:48273532018. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Seow H, O'Leary E, Perez R and Tanuseputro
P: Access to palliative care by disease trajectory: A
population-based cohort of Ontario decedents. BMJ Open.
8:e0211472018. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Krishna SS, Majumdar I and Grishin NV:
Structural classification of zinc fingers: Survey and summary.
Nucleic Acids Res. 31:532–550. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Tubon TC, Tansey WP and Herr W: A
nonconserved surface of the TFIIB zinc ribbon domain plays a direct
role in RNA polymerase II recruitment. Mol Cell Biol. 24:2863–2874.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Masud T, Soong C, Xu H, Biele J, Bjornson
S, McKinney S and Aparicio S: Ubiquitin-mediated DNA damage
response is synthetic lethal with G-quadruplex stabilizer CX-5461.
Sci Rep. 11:98122021. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Devaux S, Kelly S, Lecordier L, Wickstead
B, Perez-Morga D, Pays E, Vanhamme L and Gull K: Diversification of
function by different isoforms of conventionally shared RNA
polymerase subunits. Mol Biol Cell. 18:1293–1301. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Coulombe B and Burton ZF: DNA bending and
wrapping around RNA polymerase: A ‘revolutionary’ model describing
transcriptional mechanisms. Microbiol Mol Biol Rev. 63:457–478.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Garrido-Godino AI, Gutierrez-Santiago F
and Navarro F: Biogenesis of RNA polymerases in yeast. Front Mol
Biosci. 8:6693002021. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Singh H: Targeting RNA polymerase II
mediator subunits in cancer therapy. Proc Natl Acad Sci USA.
118:e21001151182021. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Rodriguez-Berriguete G, Ranzani M, Prevo
R, Puliyadi R, Machado N, Bolland HR, Millar V, Ebner D, Boursier
M, Cerutti A, et al: Small-molecule poltheta inhibitors provide
safe and effective tumor radiosensitization in preclinical models.
Clin Cancer Res. 29:1631–1642. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Zhang Y, Wang D, Peng M, Tang L, Ouyang J,
Xiong F, Guo C, Tang Y, Zhou Y, Liao Q, et al: Single-cell RNA
sequencing in cancer research. J Exp Clin Cancer Res. 40:812021.
View Article : Google Scholar : PubMed/NCBI
|