Introduction
A study of nasopharyngeal carcinoma (NPC) recently
demonstrated that the mortality rate of this disease was increasing
in Guangxi, China (1). The current
prevention and therapy for the disease does not indicate an
optimistic outcome. Since CNE3 was established from a liver
metastatic carcinoma tissue of primary NPC (2), it has been used in basic studies of
NPC (3–8). Certain studies revealed that the
molecular biological characteristics were different between primary
NPC and metastatic NPC, including expression of EBV-encoded small
RNA 1 (EBER1) (9), zinc levels
(10), karyotype and
differentiation (11). Therefore,
CNE3 may be useful for studies of metastatic NPC. However, the
molecular pathology of CNE3 is altered due to long-term culture
in vitro. The knowledge obtained from the continuing
progress in molecular biological technology combined with the
present study of the molecular pathology of CNE3 may provide useful
data for subsequent studies.
Materials and methods
Cell culture
The human NPC epithelial cell lines, CNE1, CNE2,
CNE3 and C666-1, were preserved in the Research Center of Medical
Sciences, The People’s Hospital of Guangxi Zhuang Autonomous Region
(Nanning, China). As a control, CNE3 was obtained from the National
Institute for Viral Disease Control and Prevention, Chinese Center
of Disease Control and Prevention (Beijing, China).
Animal experiments
In order to establish the nude mouse tumor model of
CNE3 through subcutaneous transplantation, Balb/c pure line mice
were obtained from the Guangxi Medical University Laboratory Animal
Centre (certification no. SCXK Gui 2009–0002). This study was
approved by the ethics committee of The People’s Hospital of
Guangxi Zhuang Autonomous Region.
CNE3 short tandem repeat (STR) loci
analyses
The CNE3 STR loci were authenticated using an ABI
3100 Genetic Analyzer (Microread Gene Technology, Beijing,
China).
Histomorphology experiments
The tissues were obtained from a patient’s primary
nasopharynx foci in 1982, the same patient’s metastatic liver
carcinoma of primary NPC in 1988 and nude mice transplanted tumor
in 2012. The tissues were fixed using 10% formalin and paraffin
embedding, then sliced and stained with hematoxylin and eosin (HE).
An optical analysis was then performed. Subsequent to being double
stained with uranyl acetate-lead citrate, the transplanted tumor
was observed using a H-7650 transmission electron microscope (TEM;
Hitachi, Tokyo, Japan).
Immunohistochemistry (IHC)
A non-biotin horseradish peroxidase (HRP)
ready-to-use two-step detection system (ZSGB-BIO, Beijing, China)
and BX51 fluorescence microscopy (Olympus, Tokyo, Japan) were used
in the IHC analysis. The positive brown granules, which were more
abundant than the unspecific staining background, were mainly
distributed in the cell nucleus (p63) or cytoplasm [cytokeratin
(CK)5/6, CK7]. The positive cell rates and staining intensities
were comprehensively analyzed in the intact slices using high power
fields (x200 or ×400). The results of the positive cell rates
(<10%) and weak coloring were negative. The results of the
positive cell rates (>10%) and dark brown granules were
positive.
EBER in situ hybridization
(EBER-ISH)
An EBER-ISH kit (ZSGB-BIO) identified that the
positive brown granules were mainly distributed in cell nuclei.
DNA extraction and polymerase chain
reaction (PCR)
DNA was extracted using the Genomic DNA Purification
kit (Promega, Madison, WI, USA). The following primer sequences
were used for the amplification of BamH1-A right frame 1
(BARF1; NC_007605.1): BARF1 forward, 5′-CCAGGCTGTCACCGCTTTC-3′ and
reverse, 5′-CGCCAT TTGCCGCAGTT-3′. The sequence length was 469 bp.
The reaction conditions consisted of 12 μl 2XTaq PCR Mix
(Tiangen, Beijing, China), 0.5 μl template, 0.5 μl forward primer,
0.5 μl reverse primer and 11.5 μl ddH2O. The reaction
program consisted of an initial denaturation step at 95°C for 10
min, denaturation at 94°C for 35 sec, annealing at 57°C for 35 sec,
extension at 72°C for 35 sec for 40 cycles and a final extension at
72°C for 10 min. The sequence was amplified using S1000 Thermal
Cycler PCR (Bio-Rad, Hercules, CA, USA).
Sequencing
Purified PCR products were analyzed by the 3730
automatic DNA sequencer (ABI, USA).
Results
Contamination status of CNE3
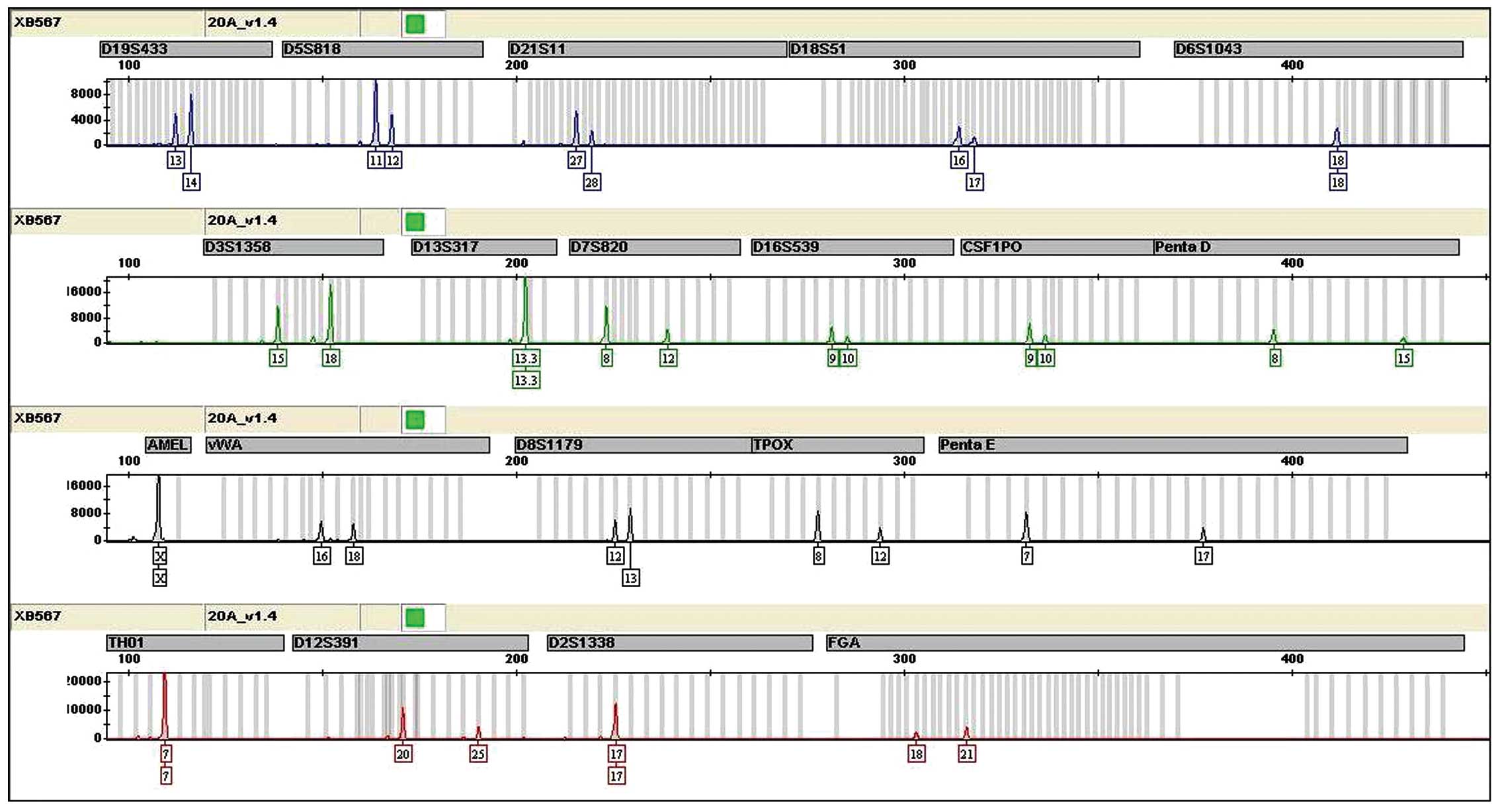
A total of 20 STR loci were not triallelic and the
results revealed that CNE3 was not cross-contaminated by other
human cells (Fig. 1).
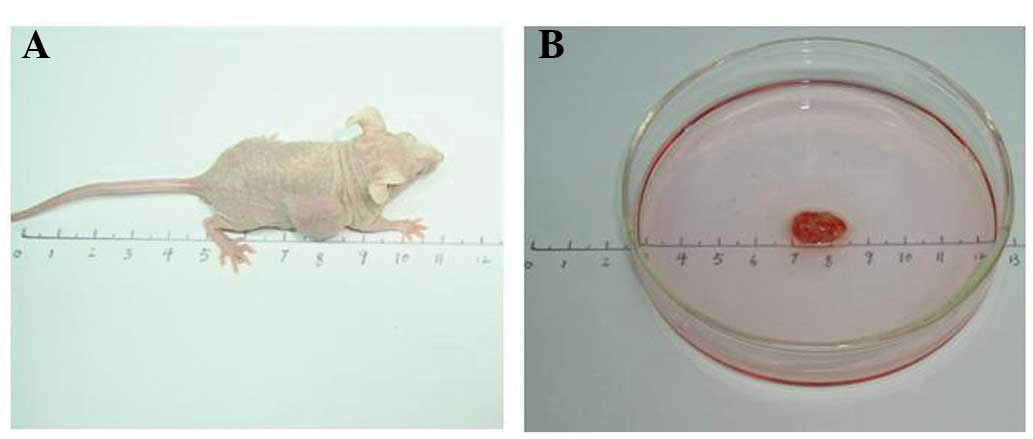
Nude mouse transplanted tumor model
The transplanted CNE3 tumor volume was 0.15
cm3 after 14 days (Fig.
2).
Adenocarcinoma morphological
characteristics
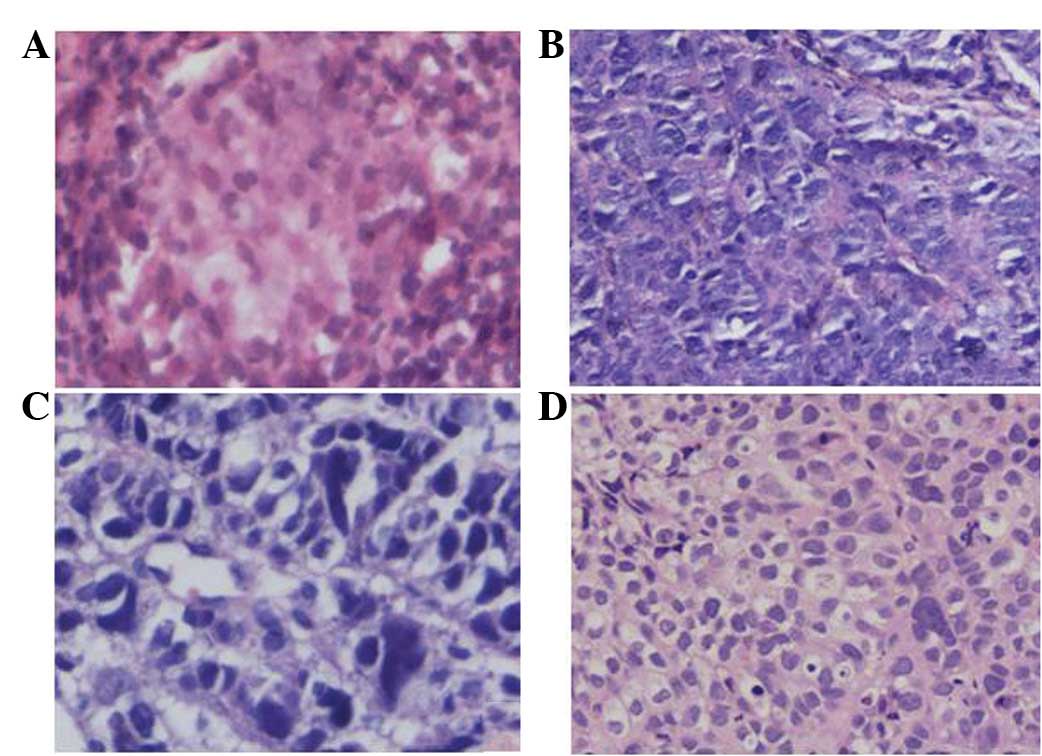
Microscopically, the cancer cells from the primary
nasopharynx foci indicated the structure of an undifferentiated
non-keratinizing carcinoma. The cells were polygonal, weakly
basophilic, contained a large nucleus with prominent nucleoli, had
little cytoplasm and an unclear cell boundary (Fig. 3A), which were arranged in sheets and
nests. The cells of the primary metastatic liver carcinoma revealed
a primitive adenoid structure. The cells had a circular form, rich
cytoplasm and clear cell boundaries (Fig. 3B and C). The cells of the nude mice
with the transplanted tumors indicated an adenoid structure. The
cells were a spindle or polylateral shape and there were
physaliphorous cells (Fig. 3D).
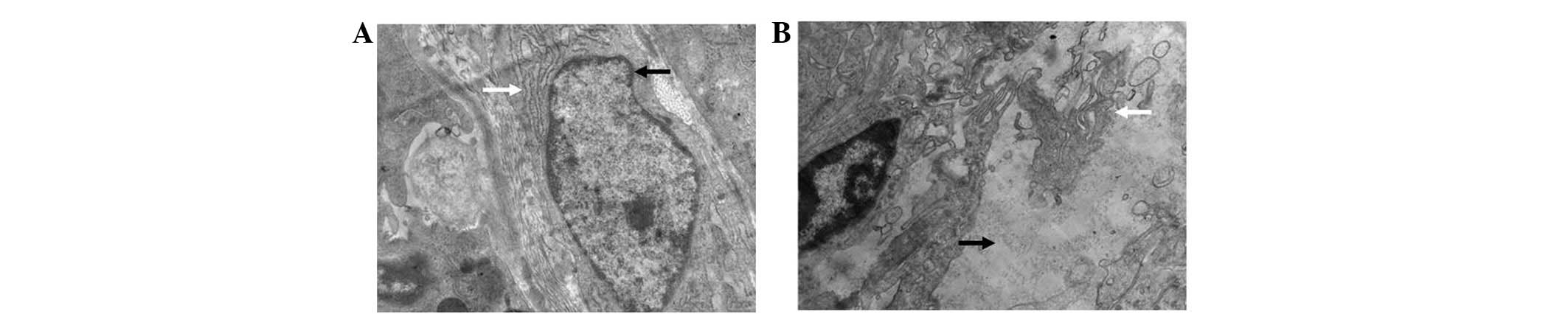
Electron microscopy observations revealed the typical
characteristics of an adenocarcinoma (Fig. 4).
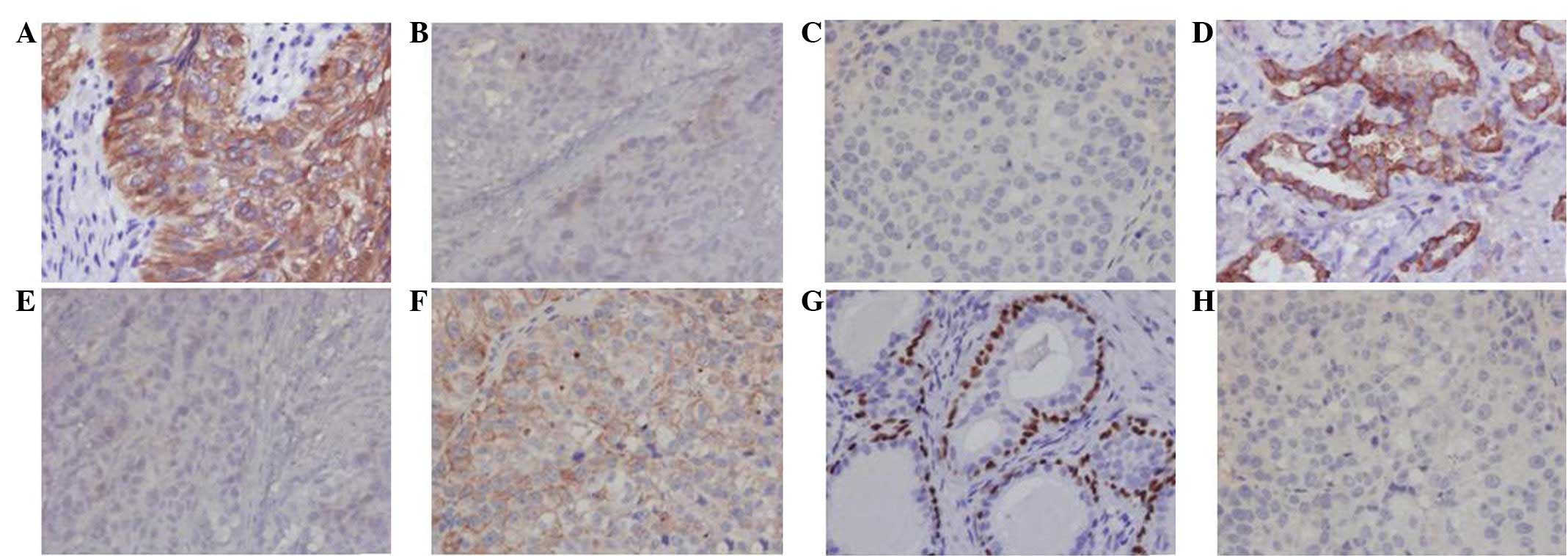
IHC results
Positive CK5/6 and CK7 results indicated that the
metastatic liver carcinoma tissues had features of adenocarcinoma
and undifferentiated non-keratinizing carcinoma. The negative
results for CK5/6 and p63 expression and the positive result for
CK7 expression indicated that CNE3 only had features that were
specific to an adenocarcinoma (Fig.
5).
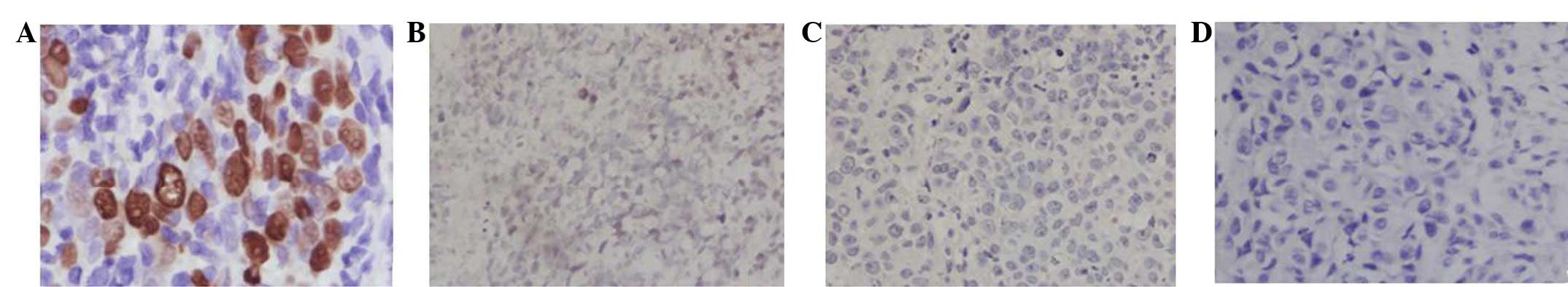
ISH results
The liver metastatic carcinoma cells were positive
for EBER; however, the nude mice transplanted tumor CNE3 cells were
negative for EBER. The results indicated that the EBV was no longer
present in the CNE3 cells (Fig.
6).
PCR and DNA sequencing results
The results of 3–4 unspecific amplification bands
indicated that the EBV was no longer present in the CNE1, CNE2 and
CNE3 cells. C666-1 was used as a positive control and
ddH2O was used as a negative control (Fig. 7A). The PCR products were not
sequenced, with the exception of C666-1. The PCR sequence of C666-1
was matched with the BARF1 gene, according to the NCBI blast
database (Fig. 7B).
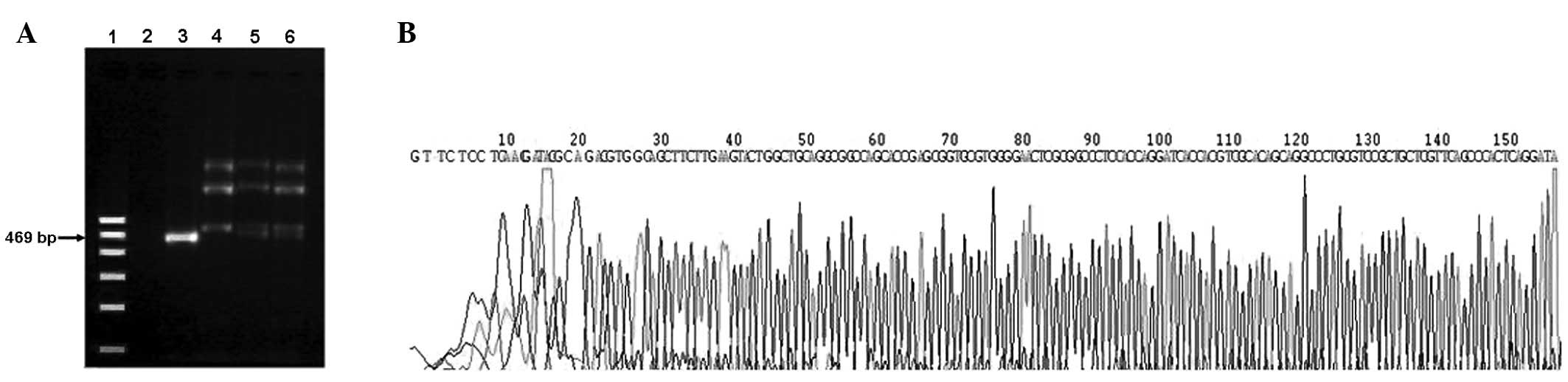 | Figure 7BARF1 PCR and sequencing in various
NPC cell lines. (A) Lanes 1–6 represent the DNA marker, negative
control, C666-1, CNE1, CNE2 and CNE3, respectively. The DNA markers
are 100, 200, 300, 400, 500 and 600 bp. The product length of the
BARF1 PCR product was 469 bp. (B) The PCR products of C666-1 were
sequenced, but other PCR products were not. BARF1, BamH1-A
right frame 1; PCR, polymerase chain reaction; NPC, nasopharyngeal
carcinoma. |
Discussion
Scanning the tissue slices of the nasopharynx
primary foci and liver metastatic carcinoma of primary NPC, the
histological type of the nasopharynx primary foci was identified as
an undifferentiated non-keratinizing carcinoma. The main area of
liver metastatic foci was the undifferentiated non-keratinizing
carcinoma structure. However, the other area indicated a primitive
adenoid structure. The change suggested that the CNE3 cells were
differentiating towards an adenocarcinoma. The CNE3 cell line has
had the features of a poorly-differentiated adenosquamous carcinoma
since it was established in 1992. EBV has been shown to express
specific proteins, including EBV nuclear antigen (EBNA) and latent
membrane protein (LMP), in the 19th passage cells of nude mouse
transplanted tumors (2). Teng et
al(4) detected EBV in the 33rd
passage cells. CNE3 has been passaged and preserved well for 20
years. The present study used the CNE3 cell line to establish a
Balb/c nude mouse transplanted tumor model and then detected the
tumor tissues by morphological and molecular pathological
experiments. As a result, certain changes were identified
subsequent to comparing the nasopharynx primary foci tissues with
the metastatic liver carcinoma tissues.
The formerly used single method of pathological
detection may cause biases of the morphological diagnosis.
Accompanied by a widespread application of IHC, numerous studies
have demonstrated that the combined expression of p63 and CK5/6 may
improve the diagnostic accuracy of undifferentiated
non-keratinizing carcinoma (12).
CK7 was highly expressed in the nasopharyngeal glandular epithelium
(13). Therefore, a combined
application of p63, CK5/6 and CK7 was able to distinguish between
undifferentiated non-keratinizing carcinoma and adenocarcinoma.
Immunophenotyping analyses of nasopharyngeal undifferentiated
non-keratinizing carcinoma were shown to be p63+,
CK5/6+ and CK7−. Therefore, the three markers
were used to detect the tissues of the nude mouse transplanted
tumors. The CK5/6 and CK7 results from the metastatic liver
carcinoma tissues of primary NPC revealed that the CNE3 xenograft
transformed from an undifferentiated non-keratinizing carcinoma
into a poorly-differentiated adenocarcinoma. Electron microscopy
further confirmed that the transplanted tumor tissues had classical
characteristics of a poorly-differentiated adenocarcinoma,
consisting of abundant rough endoplasmic reticulum, a ranged
lamellar structure and microvilli on the surface of the
micrograndular cavities. The fast growth and predominant quantities
of the adenocarcinoma cells may have gradually hampered the growth
space of the undifferentiated non-keratinizing carcinoma in the
continuing passage.
EBER-ISH is considered to be the gold standard for
detecting EBV in cancer cells (14). The metastatic liver carcinoma cells
of primary NPC were positive for EBER. However, the nude mice
transplanted tumor CNE3 cells were negative for EBER. In 1996, EBV
markers of CNE1, CNE2 and CNE3 were detected using ISH, western
blotting, southern blotting and PCR. The techniques gave positive
results, particularly when using PCR for BARF1. The expression of
EBV was strongest in the CNE2 cell line (4). The undifferentiated C666-1 cancer cell
line was used as a positive control (15). The conservative carcinogen, BARF1
(16), was identified in order to
confirm whether EBV was present in the tissues. The PCR results
indicated that the CNE1, CNE2 and CNE3 cells were negative for
BARF1, with the exception of C666-1. Therefore, the EBV was no
longer present in the CNE3 cells.
EBNAl is a unique viral protein that is found in the
four forms of latent infection by EBV. It provides a distinct
episome maintenance function by binding to oriP, which is the
latent origin of DNA replication (17,18).
Therefore, the expression level of EBNA1 is a key factor that
episomes maintain in a steady state in vitro. If all
episomes are lost, continuously mutated or partially missed, EBV
will be lost.
The characteristics of CNE3 were studied and the
pathological type was confirmed to be a poorly-differentiated
adenocarcinoma with a low incidence rate. In conclusion, this
knowledge on the molecular pathological changes of CNE3 may aid in
the development of new research approaches for NPC.
Acknowledgements
This study was supported by grants from the Natural
Science Foundation of Guangxi Province (no. 2011 GXNSFA018308) and
the Self Foundation of the Health Department of Guangxi (Z2013389).
The authors would like to thank Dr Hai-Jun Du (National Institute
For Viral Disease Control and Prevention of Chinese Center) for
providing the CNE3 cell line.
References
|
1
|
Huang TR, Zhang SW, Chen WQ, Deng W, Zhang
CY, Zhou XJ and Zhai RH: Trends in nasopharyngeal carcinoma
mortality in China, 1973–2005. Asian Pac J Cancer Prev.
13:2495–2502. 2012.PubMed/NCBI
|
|
2
|
Jiao W: Establishment of a human
epithelial cell line of nasopharyngeal carcinoma - CNE3 and its
biological characteristics. Journal of Guangxi Medical University.
12:187–190. 1995.(In Chinese).
|
|
3
|
Chen W, Lee Y, Wang H, Yu GG, Jiao W, Zhou
W and Zeng Y: Suppression of human nasopharyngeal carcinoma cell
growth in nude mice by the wild-type p53 gene. J Cancer Res Clin
Oncol. 119:46–48. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Teng ZP, Ooka T, Huang DP and Zeng Y:
Detection of Epstein-Barr virus DNA in well and poorly
differentiated nasopharyngeal carcinoma cell lines. Virus Genes.
13:53–60. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Xia Y, Wong NS, Fona WF and Tideman H:
Upregulation of GADD153 expression in the apoptotic signaling of
N-(4-hydroxyphenyl)retinamide (4HPR). Int J Cancer. 102:7–14. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Peng LX, Chen JX, Cheng JJ, Liu F, et al:
Establishment of radioresistant subline from human nasopharyngeal
carcinoma cell line by repeating irradiation. Chin J New Clin Med.
5:1107–1109. 2012.(In Chinese).
|
|
7
|
Yang XL, Liu XC, Huang L, Lan J, Zhang HY,
Qin MB, Zhong YY and Mo ZN: Effect of TSA on nasopharyngeal
carcinoma CNE3 cells and its mechanism. Chin J Public Health.
26:1029–1030. 2010.(In Chinese).
|
|
8
|
Liu XC, Lan J, Nong CZ, Pan LL and Jiao W:
Effects of mangiferin on induction of apoptosis and intracellular
Ca2+ concentration in nasopharyngeal carcinoma CNE3 cells. Chin J
New Clin Med. 3:805–807. 2010.(In Chinese).
|
|
9
|
Tsai ST, Jin YT and Su IJ: Expression of
EBER1 in primary and metastatic nasopharyngeal carcinoma tissues
using in situ hybridization. A correlation with WHO histologic
subtypes. Cancer. 77:231–236. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bay B, Chan Y, Fong C and Leong H:
Differential cellular zinc levels in metastatic and primary
nasopharyngeal carcinoma. Int J Oncol. 11:745–748. 1997.PubMed/NCBI
|
|
11
|
Lin JJ, He SY, Hao XP and Zong YS: Study
on the primary and metastatic tumours of nasopharyngeal carcinoma
using microspectrometry and immunohistochemistry. Chin J Cancer.
17:90–92. 1998.(In Chinese).
|
|
12
|
Kaufmann O, Fietze E, Mengs J and Dietel
M: Value of p63 and cytokeratin 5/6 as immunohistochemical markers
for the differential diagnosis of poorly differentiated and
undifferentiated carcinomas. Am J Clin Pathol. 116:823–830. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mitroi M, Cčpitčnescu A, Georgescu CV,
Mogoantă CA, Popescu C, Georgescu M, Mitroi G and Ioniţă E:
Expression pattern of CK7 and CK20 in nasal polyps, at patients
with chronic rhinosinusitis with nasal polyposis. Rom J Morphol
Embryol. 52(3 Suppl): S1051–S1057. 2011.PubMed/NCBI
|
|
14
|
Kuo TT, Shih LY and Tsang NM: Nasal NK/T
cell lymphoma in Taiwan: a clinicopathologic study of 22 cases,
with analysis of histologic subtypes, Epstein-Barr virus LMP-1 gene
association, and treatment modalities. Int J Surg Pathol.
12:375–387. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Cheung ST, Huang DP, Hui AB, Lo KW, Tsang
YS, Wong N, Whitney BM and Lee JC: Nasopharyngeal carcinoma cell
line (C666-1) consistently harbouring Esptein-Barr virus. Int J
Cancer. 83:121–126. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
de Turenne-Tessier M and Ooka T:
Post-translational modifications of Epstein-Barr virus BARF1
oncogene-encoded polypeptide. J Gen Virol. 88:2656–2661.
2007.PubMed/NCBI
|
|
17
|
Dittmer DP, Hilscher CJ, Gulley ML, Yang
EV, Chen M and Glaser R: Multiple pathways for Epstein-Barr virus
episome loss from nasopharyngeal carcinoma. Int J Cancer.
123:2105–2112. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Sivachandran N, Thawe NN and Frappier L:
Epstein-Barr virus nuclear antigen 1 replication and segregation
functions in nasopharyngeal carcinoma cell lines. J Virol.
85:10425–10430. 2011. View Article : Google Scholar : PubMed/NCBI
|