Introduction
MicroRNAs (miRNAs or miRs) are small non-coding RNAs
with 17–25 nucleotides that are present in eukaryotic organisms
(1). miRNAs are important in diverse
biological processes by post-transcriptionally regulating gene
expression through base pairing to complementary sites in the
3′-untranslated region (UTR) of selected transcripts (1). Since their initial identification,
~1,000 miRNA sequences have been reported thus far (1), and their biological functions have
attracted great attention, although further investigation into
their mechanism of action is still required. Recently, miRNAs have
been observed to participate in the regulation of diverse
physiological and pathological processes, particularly in cancer
development (2). Aberrant miRNA
expression is associated with the pathogenesis of various types of
cancer, and they are associated with clinical outcomes of cancer
patients (3). However, identifying
the deregulated miRNAs and their roles in carcinogenesis and cancer
progression, particularly in osteosarcoma development, is an
ongoing process (4).
Osteosarcoma is the most common type of human
primary malignant bone tumor, and usually develops in children and
young adults, with an incidence rate of 3 per million per year
(5). It is characterized by an
aggressive clinical course, with a 5-year survival rate of 60–70%
in patients with no metastatic disease (5–7). The
clinical outcomes are considerably worse in patients with
metastatic disease (5–7). Currently, the mechanisms responsible for
the oncogenic insults in the initiation and progression of
osteosarcoma remain to be fully elucidated. To date, a number of
miRNAs have been determined to be deregulated in osteosarcoma and
to participate in osteosarcoma development, including miR-212
(8), miR-33b (9), miR-503 (10) and miR-451 (11). In addition, the present authors also
reported that miR-143 and miR-133a are downregulated in
osteosarcoma, and they function as anti-oncomiRs in osteosarcoma
development (12,13). However, the miRNA expression profile
and the roles of deregulated miRNAs in osteosarcoma carcinogenesis
and progression require further investigation.
In order to identify the deregulated miRNAs involved
in osteosarcoma, paired clinically resected osteosarcoma tissues
and adjacent normal tissues were obtained from 92 primary
osteosarcoma patients, and their miRNA expression profile was
elucidated using miRNA microarray analysis in the present study. A
set of deregulated miRNAs in osteosarcoma tissues were identified,
including upregulated miR-31, miR-210 and miR-148a, and
downregulated miR-206, miR-1 and miR-133a. Since the roles of
miR-31, miR-210, miR-206, miR-1 and miR-133a in osteosarcoma
progression have been previously reported (13–17), and
the roles of miR-148a in osteosarcoma development are elusive, the
present study focused on the expression and biological function of
miR-148a in osteosarcoma carcinogenesis and progression.
Materials and methods
Patients samples
Human osteosarcoma tumor tissues and paired adjacent
normal tissues were obtained from 92 primary osteosarcoma patients
during surgery. The surgeries of these patients were performed
between April 2006 and December 2009 at Changhai Hospital
(Shanghai, China). The clinical information of these patients was
previously published (13). The
tissues were rapidly frozen in liquid nitrogen following surgical
resection. All specimens were collected upon obtaining informed
consent from all patients. The experiments in the present study
were approved by the ethics committee of The Second Military
Medical University (Shanghai, China).
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA, including miRNA, was extracted as
previously reported (12,13). For evaluating the expression of
miR-148a, RT-qPCR analysis was performed using stem-loop RT
primers, 5′-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACACAAAG-3′,
and qPCR primers 5′-TCAGTGCACTACAGAA-3′ (forward) and
5′-GTGCAGGGTCCGAGGT-3′ (reverse). The primers were synthesized by
Invitrogen (Thermo Fisher Scientific, Inc., Waltham, MA, USA). The
RT-qPCR assay was performed and analyzed as previously reported
(12,13).
Cell culture and transfection
The human osteoblastic cell line hFOB 1.19 and the
osteosarcoma cell lines MG63 and U2OS (American Type Culture
Collection, Manassas, VA, USA) were cultured, seeded and
transfected as previously described (12,13).
miR-148a mimics, miR-148a inhibitor and their respective controls
were synthesized by Shanghai GenePharma Co., Ltd. (Shanghai,
China). miRNA mimics or inhibitor were transfected at a final
concentration of 20 or 50 nM, respectively, using
INTERFERin® transfection reagent (Polyplus-transfection
SA, Illkirch, France), following the manufacturer's protocol.
Cell proliferation analysis
MG63 and U2OS cells were subjected to in
vitro cell proliferation assay as previously reported (12,13). In
brief, cells were seeded into 96-well plates, transfected and
analyzed using the
(3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay at the indicated time points. Used cell medium was replaced
with 200 µl fresh medium containing MTT 0.5 mg/ml. Cell cultural
plates were incubated at 37°C for 4 h, then 100 µl dimethyl
sulfoxide (Sigma-Aldrich, St. Louis, MO, USA) was added and the
plates were shaken for 10 min. The absorbance was measured at 570
nm.
Tumor growth analysis in vivo
All animal experiments were performed according to
the National Institutes of Health Guide for the Care and Use of
Laboratory Animals and approved by the ethics committee of Second
Military Medical University, Shanghai, China. The nude mice used in
the present study were obtained from Shanghai Laboratory Animal
Research Center (Shanghai, China), and kept in pathogen-free
conditions with normal chow, access to water and regular light/dark
cycles. MG63 cells (1×106) suspended in 0.1 ml
phosphate-buffered saline were injected subcutaneously into the
posterior flank of BALB/c athymic nude mice. Two weeks later, 10
nmol cholesterol-conjugated small RNA agonist (ago)-miR or
antagonist (antago)-miR of miR-148a (Guangzhou RiboBio Co., Ltd.,
Guangzhou, China) in 0.1 ml saline buffer was locally injected into
the tumor mass (once every 3 days for 2 weeks). Tumor growth was
measured as previously described (13).
3′-UTR luciferase reporter
analysis
Human phosphatase and tensin homolog (PTEN) 3′-UTR
luciferase reporter constructs and constructs with miR-148a target
site deletion were constructed as previously reported (12,13), and
verified by sequencing. Sequencing was performed by service
provider Jie Li Biology (Shanghai, China). Cells were transfected
and luciferase activity was analyzed as previously described
(12,13).
Western blotting
Cells or grinded human tissues were lysed, and their
total protein content was extracted and subjected to
electrophoresis and western blot analysis as previously reported
(12,13). Antibodies specific for PTEN (rabbit
monoclonal; catalog no., 9188; 1:1,000) and β-actin (mouse
monoclonal; catalog no., 3700; 1:2,000), and horseradish peroxidase
(HRP)-coupled secondary antibodies against rabbit [anti-rabbit
immunoglobulin G (IgG), HRP-linked; catalog no., 7074; 1:2,000] and
mouse (anti-mouse IgG, HRP-linked; catalog no., 7076; 1:2,000) were
obtained from Cell Signaling Technology, Inc. (Danvers, MA, USA)
and used following the standard protocols. Densitometric analysis
was performed using LabWorks Image Acquisition and Analysis
Software (UVP, Inc., Upland, CA, USA) as previously described
(13).
Statistical analysis
Data are presented as the mean ± standard deviation.
Student's t-test was used to analyze statistical comparisons
between groups, and two-tailed P<0.05 was considered to indicate
a statistically significant difference. Spearman's rank correlation
coefficient in SPSS version 17.0 software (SPSS, Inc., Chicago, IL,
USA) was used to analyze the correlation between miR-148a
expression and clinical osteosarcoma stages, while Kaplan-Meier
survival analysis with log-rank test in SPSS version 17.0 software
was used to analyze the overall survival in osteosarcoma patients.
Pearson's correlation coefficient in SPSS version 17.0 software was
used to analyze the correlation between the expression of miR-148a
and the protein levels of PTEN.
Results
Increased miR-148a expression in
osteosarcoma
To investigate the miRNA expression profile in
osteosarcoma, miRNA microarray analysis was applied to human
osteosarcoma tissue and adjacent normal tissue. The microarray
analysis of miRNA expression was performed by a service provider,
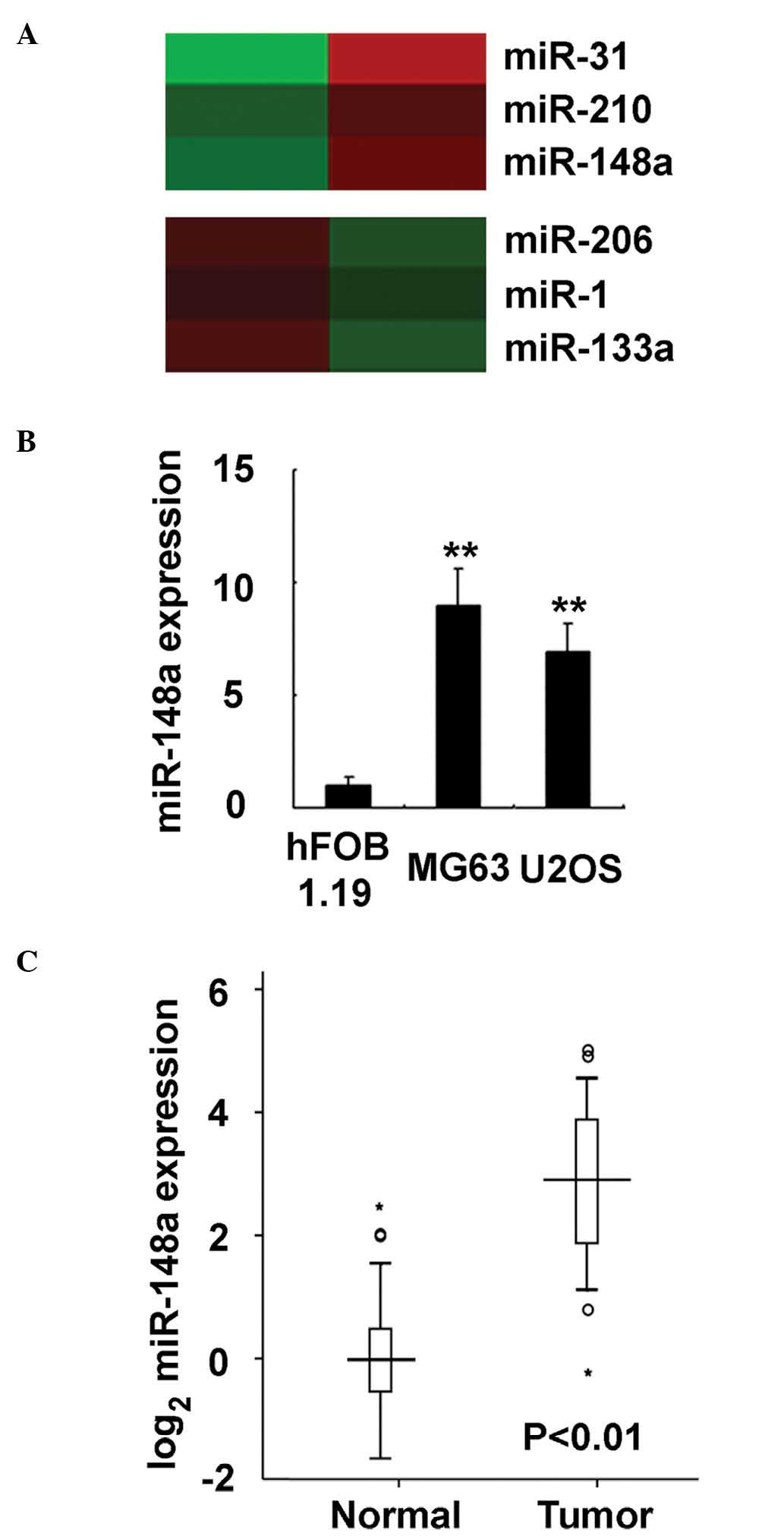
Shanghai Biotechnology Corporation (Shanghai, China) (3). A set of miRNAs were observed to be
deregulated in osteosarcoma tissue, including miR-31, miR-210 and
miR-148a (the most upregulated miRNAs identified in the analysis),
and miR-206, miR-1 and miR-133a (the most downegulated miRNAs
identified in the analysis) (Fig.
1A). Of these miRNAs, the roles of miR-133a in osteosarcoma
development were previously reported by the present authors
(13). Therefore, the present study
focused on the roles of miR-148a in human osteosarcoma progression.
miR-148a expression in osteosarcoma was further evaluated in
osteosarcoma cell lines, and it was observed to be increased in the
human osteosarcoma cell lines MG63 and U2OS, compared with the
normal osteoblastic cell line hFOB 1.19 (P<0.0001; Fig. 1B). Furthermore, in the 92 pairs of
human clinical osteosarcoma and adjacent normal tissue specimens
analyzed in the present study, miR-148a expression was determined
to be significantly increased in osteosarcoma tissues, compared
with normal tissues (Fig. 1C). Thus,
the present data indicate that miR-148a expression is increased in
osteosarcoma.
Increased miR-148a expression is
significantly correlated with progression and prognosis of
osteosarcoma
As miR-148a expression was observed to be
significantly increased in osteosarcoma, the correlation between
miR-148a upregulation and osteosarcoma progression was further
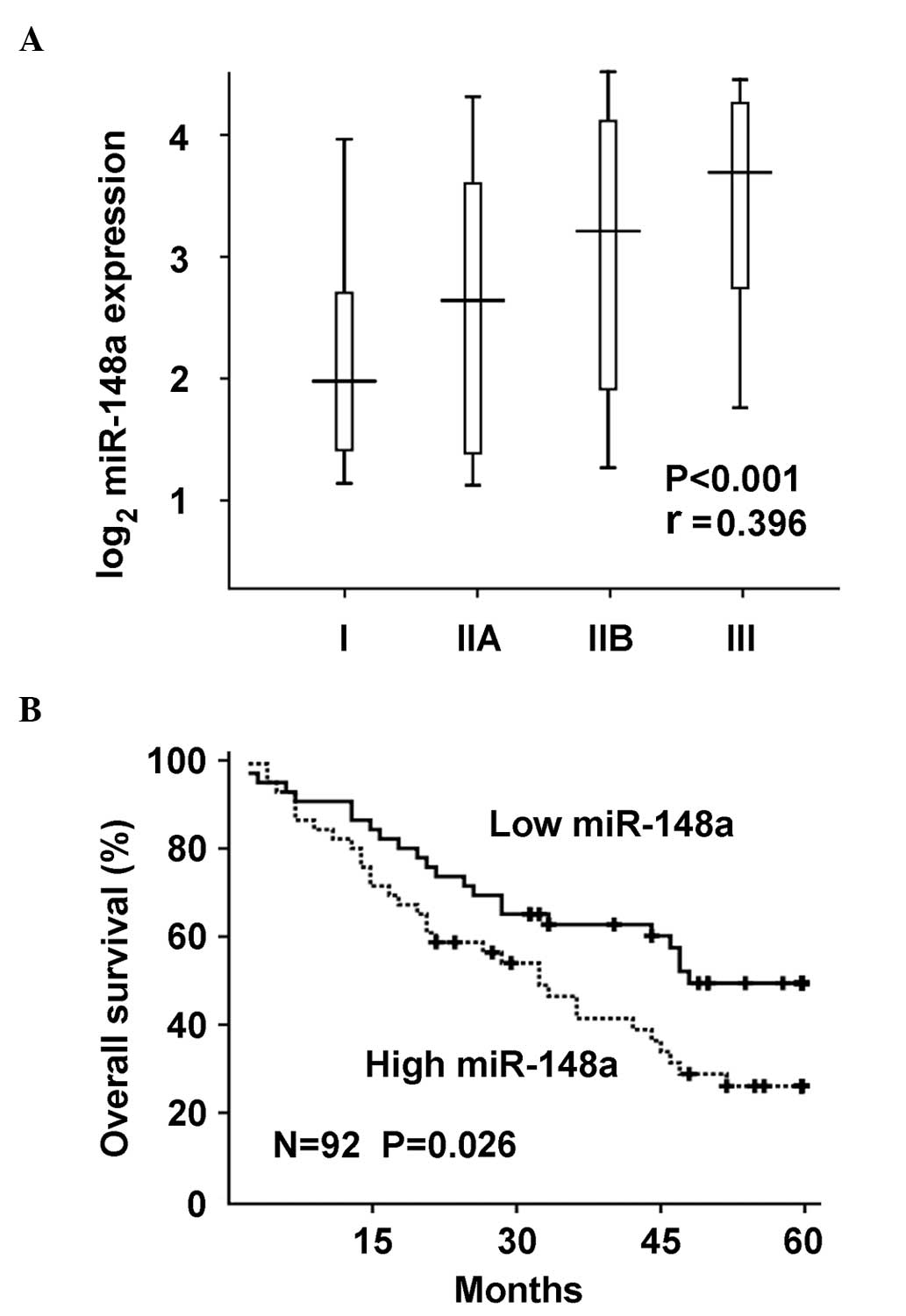
investigated. miR-148a expression was significantly
positive-correlated with tumor stages of osteosarcoma in the human
samples, according to the Spearman's rank correlation coefficient
(P<0.0001; Fig. 2A). Furthermore,
miR-148a upregulation in osteosarcoma tissues was significantly
correlated with poor overall survival of osteosarcoma patients by
Kaplan-Meier survival analysis (Fig.
2B). Taken together, these data suggest that miR-148a
upregulation may be important in osteosarcoma progression and
prognosis, and determination of miR-148a expression may be used in
the pathological identification and prediction of prognosis in
osteosarcoma patients.
miR-148a promotes osteosarcoma growth
in vitro and in vivo
Since the expression of miR-148a was increased in
osteosarcoma and correlated with poor prognosis, it was next
investigated whether miR-148a functioned as an oncogene in
osteosarcoma. When miR-148a was overexpressed in osteosarcoma MG63
and U2OS cells, it promoted cell proliferation in both cell lines
(Fig. 3A and B). In addition,
transfection of miR-148a inhibitor significantly suppressed
miR-148a expression, and miR-148a inhibition suppressed
proliferation in osteosarcoma cells (Fig.
3C and D). Furthermore, osteosarcoma cell growth in vivo
was promoted by administration of miR-148a ago-miR, while it was
inhibited by its antago-miR (Fig. 3E and
F, respectively). Therefore, these data suggest that miR-148a
may be an oncogene that promotes osteosarcoma growth both in
vitro and in vivo, and inhibition of miR-148a bears
considerable therapeutic potential in osteosarcoma.
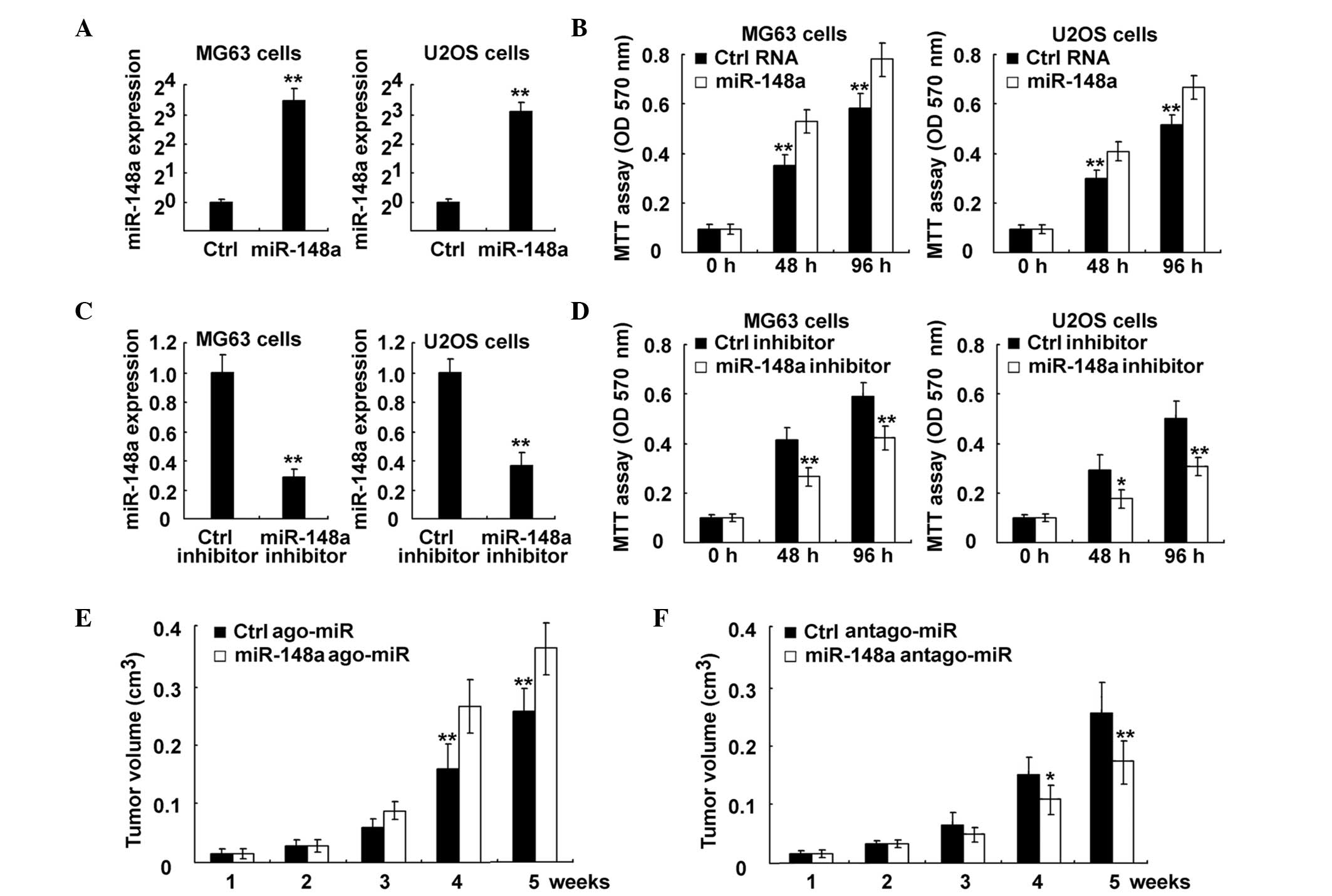 | Figure 3.Inhibition of miR-148a suppressed
osteosarcoma growth both in vitro and in vivo. (A and
B) MG63 and U2OS cells were transfected with control RNA or
miR-148a mimics, as indicated. (A) miR-148a expression was examined
using RT-qPCR. The data are the relative values of miR-148
expression normalized to the internal control of U6 expression. (B)
Cell proliferation was measured using MTT assay at the indicated
time points following transfection. (C and D) MG63 and U2OS cells
were transfected with control RNA or miR-148a inhibitor, as
indicated. (C) miR-148a expression was examined using RT-qPCR. The
data are the relative values of miR-148 expression normalized to
the internal control of U6 expression. (D) Cell proliferation was
measured using MTT assay at the indicated time points following
transfection. (E and F) MG63 cells were injected subcutaneously
into the posterior flank of nude mice. Two weeks later,
cholesterol-conjugated small RNA (E) ago-miR or (F) antago-miR of
miR-148a were locally injected into the tumor mass, and tumor
growth was measured. The growth curves obtained are indicated in
the image. Data are presented as the mean ± standard deviation
(n=4), or correspond to one representative experiment, whereby
similar results were obtained in three independent experiments.
*P<0.05; **P<0.01. miR, microRNA; Ctrl, control; RT-qPCR,
reverse transcription-quantitative polymerase chain reaction; MTT,
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide; OD,
optical density; ago, agonist; antago, antagonist. |
miR-148a promotes phosphoinositide
3-kinase (PI3K) signaling pathway activation by targeting PTEN
expression
The mechanism responsible for the growth-promoting
effect of miR-148a on osteosarcoma was next investigated. The
intracellular cell signaling pathways in MG63 and U2OS cells
overexpressing miR-148a was examined, including PI3K,
mitogen-activated protein kinase, nuclear factor-κB and signal
transducer and activator of transcription signaling pathways, and
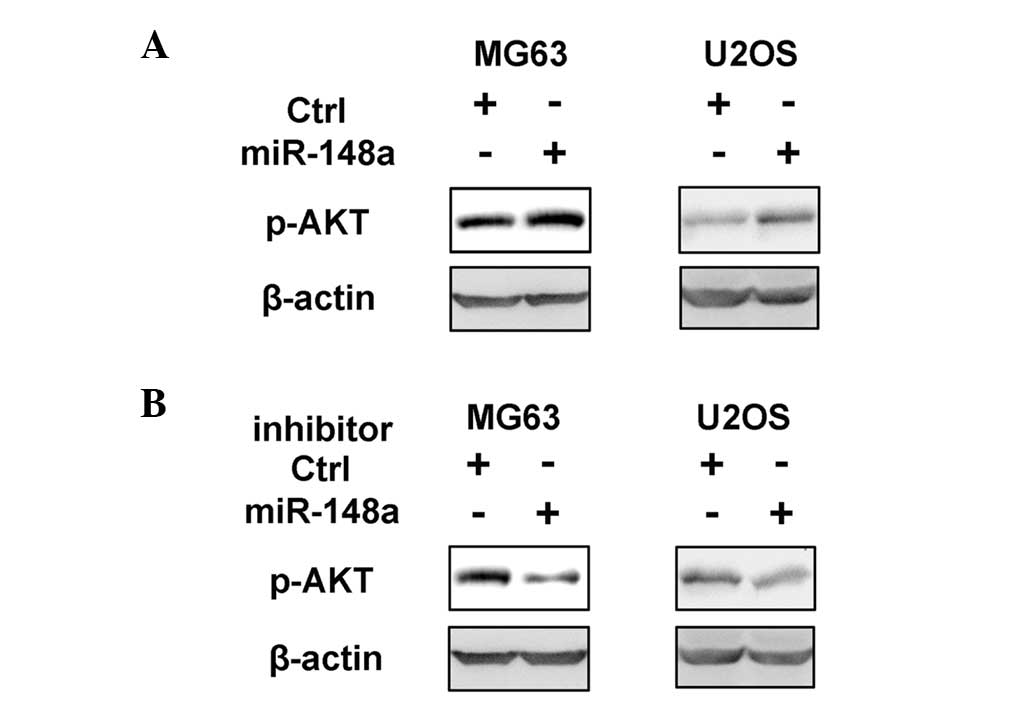
it was observed that only the activation of the PI3K signaling
pathway was significantly promoted by miR-148a overexpression
(Fig. 4A and data not shown). In
addition, the inhibition of miR-148a suppressed PI3K activation in
osteosarcoma cells (Fig. 4B). As PI3K
activation and downstream AKT phosphorylation are critical in
osteosarcoma growth and progression (18,19), it
was concluded that miR-148a may promote osteosarcoma growth by
enhancing the activation of the PI3K signaling pathway.
Since miRNAs mainly function through
post-transcriptional inhibition of their target mRNAs, the target
genes of miR-148a, which may be responsible for the activated PI3K
signaling pathway in osteosarcoma cells, were next investigated.
The predicted target genes of miR-148a were retrieved from
TargetScan (http://www.targetscan.org), and
subjected to the PI3K signaling pathway analysis of the Kyoto
Encyclopedia of Genes and Genomes pathway database (http://www.genome.jp/kegg/). Notably, PTEN, an
important PI3K inhibitor, was observed to contain two putative
target sites in the 3′-UTR sequence of its messenger (m)RNA
(Fig. 5A). To verify the potential
interaction between miR-148a and PTEN, a dual-luciferase reporter
system was performed by co-transfection of miR-148a and luciferase
reporter plasmids containing the 3′-UTR sequence of human PTEN, or
bearing deletions of the putative miR-148a target sites. As
represented in Fig. 5B, miR-148a
co-transfection with the above plasmids inhibited the luciferase
activity of wild-type PTEN 3′-UTR reporter, but failed to inhibit
that of the reporter containing deletion of both target sites.
These results indicate that miR-148a is able to target directly
PTEN mRNA 3′-UTR sequences. Furthermore, PTEN expression was
suppressed by miR-148a overexpression, while it was enhanced by
miR-148a inhibition in MG63 and U2OS cells (Fig. 5C). Thus, these data further
demonstrate that PTEN expression is directly targeted by miR-148a,
which may in turn promote the activation of the PI3K signaling
pathway in osteosarcoma.
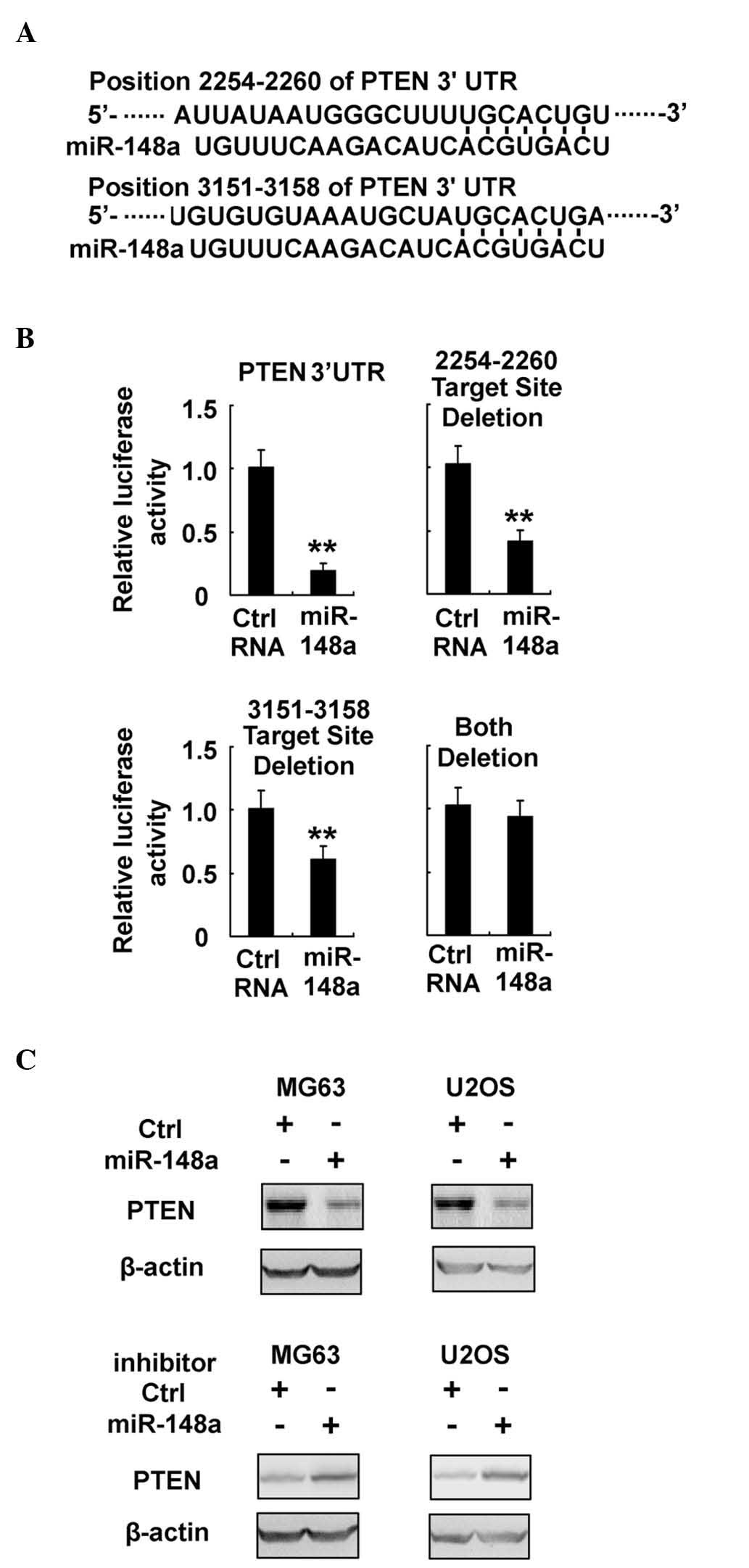 | Figure 5.miR-148a directly targets PTEN
expression in osteosarcoma. (A) Sequence alignment of miR-148a and
its predictive target sites in the 3′-UTR region of PTEN. (B) MG63
cells were co-transfected with PTEN 3′-UTR firefly luciferase
reporter plasmid, or with a reporter construct whereby the miR-148a
target sites had been deleted, together with Renilla
luciferase plasmids, control RNA or miR-148a mimics, as indicated.
Following 48 h, luciferase activity was measured and normalized.
(C) MG63 and U2OS cells were transfected with miR-148a mimics or
miR-148a inhibitor, as indicated. Protein expression of PTEN and
β-actin (which served as an internal control) were detected by
western blotting. Data are presented as the mean ± standard
deviation (n=4) of one representative experiment. Similar results
were obtained in three independent experiments. **P<0.01. miR,
microRNA; Ctrl, control; UTR, untranslated region; PTEN,
phosphatase and tensin homolog. |
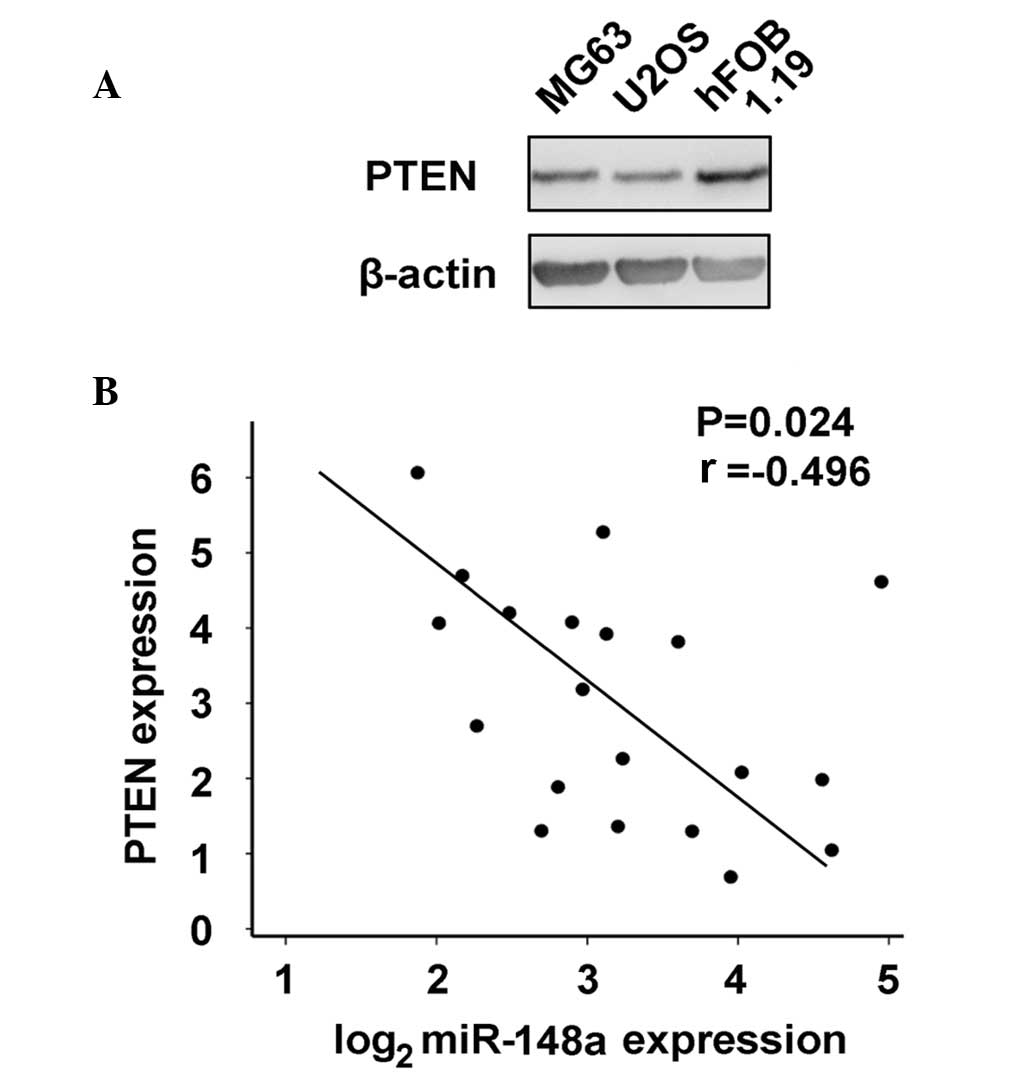
PTEN expression was further compared in osteosarcoma
MG63 and U2OS cells vs. normal osteoblastic hFOB 1.19 cells, as
increased miR-148a expression was identified in osteosarcoma. As
represented in Fig. 6A, PTEN
expression was inhibited in osteosarcoma cells. Additionally, the
correlation between miR-148a and PTEN expression in human
osteosarcoma tissues was examined by RT-qPCR and western blot
analysis. PTEN expression inversely correlated with miR-148a
expression in osteosarcoma tissues by Pearson's correlation
coefficient (Fig. 6B). Thus, these
results further suggest that increased miR-148a expression may
inhibit PTEN expression, which in turn promotes PI3K signaling
pathway activation in osteosarcoma.
Discussion
Osteosarcoma is characterized by an aggressive
clinical course, and is the most common primary malignant bone
tumor (5–7). Recently, molecular mechanisms
responsible for osteosarcoma carcinogenesis and progression have
attracted much attention in osteosarcoma development research, and
deregulated miRNAs have been identified to be important in
osteosarcoma pathogenesis (8–13). In the present study, miR-148a
expression was observed to be significantly upregulated in
osteosarcoma, and miR-148a could promote osteosarcoma growth by
inhibiting PTEN expression, thus promoting PI3K signaling pathway
activation. Furthermore, miR-148a upregulation in osteosarcoma
appeared to be correlated with osteosarcoma progression and
prognosis, and inhibiting miR-148a using antago-miR in vivo
could suppress osteosarcoma growth. Thus, the present results
suggest that miR-148a may be a novel prognosis predictor and
therapeutic target in osteosarcoma.
Previously, miR-148a expression was suggested to be
deregulated in other types of cancer, including downregulated in
gastric cancer, hepatocellular carcinoma and pancreatic cancer, and
upregulated in glioblastoma and chordomas (20–24). The
present study has determined that miR-148a is an onco-miR, and its
expression is upregulated in osteosarcoma. In combination with
previous reports of miR-148a in cancer biology, it is possible to
assume that miR-148a may play different roles in different types of
cancer, and the target genes and corresponding mechanisms are also
likely to be different depending on the type of cancer. However,
the roles and underlying mechanism of miR-148a deregulation in
carcinogenesis and tumor progression require to be further
investigated in different types of cancer.
Recently, the identification of molecular biomarkers
correlating with progression and/or predicting prognosis of cancer
patients has attracted considerable attention (3). Previously, the present authors reported
that downregulation of miR-133a expression in osteosarcoma is
correlated with cancer stages and prognosis of osteosarcoma
patients (13). In the present study,
increased miR-148a expression in osteosarcoma was also demonstrated
to be correlated with cancer stages and able to predict prognosis.
Therefore, the combined detection of deregulated miRNAs, including
miR-148a, miR-133a or other miRNAs and coding genes, may be
valuable to identify pathological stages and predict prognosis of
osteosarcoma more accurately than the detection of single
miRNA.
In conclusion, the present study has validated both
in vitro and in vivo that miR-148a promotes
osteosarcoma growth, while inhibition of miR-148a could suppress
tumor progression. In particular, cholesterol-conjugated RNA
antago-miR of miR-148a could significantly inhibit osteosarcoma
growth in vivo. These data strongly suggest that inhibition
of miR-148a bears considerable therapeutic potential for the
treatment of osteosarcoma, as the enforced expression or inhibition
of miRNAs in vivo has been demonstrated to be promising in
cancer therapy (25). Therefore,
future studies on in vivo administration of miRNA modulators
in cancer therapy may lead to important advances on cancer
treatment, particularly for those patients who respond poorly to
radiotherapy or chemotherapy.
Acknowledgements
The authors would like to thank Professor Zhiwei
Wang and Dr Yue Wang for their helpful discussion, and Ms. Jianfang
Chen and Liqing Fu for their excellent technical assistance. The
present study was supported by grants from the National Natural
Science Foundation of China (Beijing, China; grant nos. 30973019,
81272942 and 81202122), the Key Biomedicine Research Program of the
Science and Technology Commission of Shanghai (Shanghai, China;
grant no. 10411956000) and the Shanghai Natural Science Foundation
(Shanghai, China; grant no. 064119605).
References
|
1
|
Griffiths-Jones S, Grocock RJ, van Dongen
S, Bateman A and Enright AJ: miRBase: microRNA sequences, targets
and gene nomenclature. Nucleic Acids Res. 34:(Database Issue).
D140–D144. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Bushati N and Cohen SM: microRNA
functions. Annu Rev Cell Dev Biol. 23:175–205. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hou J, Lin L, Zhou W, Wang Z, Ding G, Dong
Q, Qin L, Wu X, Zheng Y, Yang Y, et al: Identification of miRNomes
in human liver and hepatocellular carcinoma reveals miR-199a/b-3p
as therapeutic target for hepatocellular carcinoma. Cancer Cell.
19:232–243. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kobayashi E, Hornicek FJ and Duan Z:
MicroRNA involvement in osteosarcoma. Sarcoma. 2012:3597392012.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Longhi A, Errani C, De Paolis M, Mercuri M
and Bacci G: Primary bone osteosarcoma in the pediatric age: State
of the art. Cancer Treat Rev. 32:423–436. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Provisor AJ, Ettinger LJ, Nachman JB,
Krailo MD, Makley JT, Yunis EJ, Huvos AG, Betcher DL, Baum ES,
Kisker CT and Miser JS: Treatment of nonmetastatic osteosarcoma of
the extremity with preoperative and postoperative chemotherapy: A
report from the Children's Cancer Group. J Clin Oncol. 15:76–84.
1997.PubMed/NCBI
|
|
7
|
Ferguson WS and Goorin AM: Current
treatment of osteosarcoma. Cancer Invest. 19:292–315. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Luo XJ, Tang DG, Gao TL, Zhang YL, Wang M,
Quan ZX and Chen J: MicroRNA-212 inhibits osteosarcoma cells
proliferation and invasion by down-regulation of Sox4. Cell Physiol
Biochem. 34:2180–2188. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Xu N, Li Z, Yu Z, Yan F, Liu Y, Lu X and
Yang W: MicroRNA-33b suppresses migration and invasion by targeting
c-Myc in osteosarcoma cells. PLoS One. 9:e1153002014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Chong Y, Zhang J, Guo X, Li G, Zhang S, Li
C, Jiao Z and Shao M: MicroRNA-503 acts as a tumor suppressor in
osteosarcoma by targeting L1CAM. PLoS One. 9:e1145852014.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yuan J, Lang J, Liu C, Zhou K, Chen L and
Liu Y: The expression and function of miRNA-451 in osteosarcoma.
Med Oncol. 32:3242015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhang H, Cai X, Wang Y, Tang H, Tong D and
Ji F: microRNA-143, down-regulated in osteosarcoma, promotes
apoptosis and suppresses tumorigenicity by targeting Bcl-2. Oncol
Rep. 24:1363–1369. 2010.PubMed/NCBI
|
|
13
|
Ji F, Zhang H, Wang Y, Li M, Xu W, Kang Y,
Wang Z, Wang Z, Cheng P, Tong D, et al: MicroRNA-133a,
downregulated in osteosarcoma, suppresses proliferation and
promotes apoptosis by targeting Bcl-xL and Mcl-1. Bone. 56:220–226.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Creighton CJ, Fountain MD, Yu Z, Nagaraja
AK, Zhu H, Khan M, Olokpa E, Zariff A, Gunaratne PH, Matzuk MM and
Anderson ML: Molecular profiling uncovers a p53-associated role for
microRNA-31 in inhibiting the proliferation of serous ovarian
carcinomas and other cancers. Cancer Res. 70:1906–1915. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Cai H, Lin L, Cai H, Tang M and Wang Z:
Prognostic evaluation of microRNA-210 expression in pediatric
osteosarcoma. Med Oncol. 30:4992013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bao YP, Yi Y, Peng LL, Fang J, Liu KB, Li
WZ and Luo HS: Roles of microRNA-206 in osteosarcoma pathogenesis
and progression. Asian Pac J Cancer Prev. 14:3751–3755. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Novello C, Pazzaglia L, Cingolani C, Conti
A, Quattrini I, Manara MC, Tognon M, Picci P and Benassi MS: miRNA
expression profile in human osteosarcoma: Role of miR-1 and
miR-133b in proliferation and cell cycle control. Int J Oncol.
42:667–675. 2013.PubMed/NCBI
|
|
18
|
Zhou Y, Zhu LB, Peng AF, Wang TF, Long XH,
Gao S, Zhou RP and Liu ZL: LY294002 inhibits the malignant
phenotype of osteosarcoma cells by modulating the
phosphatidylinositol 3 kinase/Akt/fatty acid synthase signaling
pathway in vitro. Mol Med Rep. 11:1352–1357. 2015.PubMed/NCBI
|
|
19
|
Jin S, Pang RP, Shen JN, Huang G, Wang J
and Zhou JG: Grifolin induces apoptosis via inhibition of PI3K/AKT
signalling pathway in human osteosarcoma cells. Apoptosis.
12:1317–1326. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen Y, Song Y, Wang Z, Yue Z, Xu H, Xing
C and Liu Z: Altered expression of MiR-148a and MiR-152 in
gastrointestinal cancers and its clinical significance. J
Gastrointest Surg. 14:1170–1179. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Liffers ST, Munding JB, Vogt M, Kuhlmann
JD, Verdoodt B, Nambiar S, Maghnouj A, Mirmohammadsadegh A, Hahn SA
and Tannapfel A: MicroRNA-148a is down-regulated in human
pancreatic ductal adenocarcinomas and regulates cell survival by
targeting CDC25B. Lab Invest. 91:1472–1479. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Xu X, Fan Z, Kang L, Han J, Jiang C, Zheng
X, Zhu Z, Jiao H, Lin J, Jiang K, et al: Hepatitis B virus X
protein represses miRNA-148a to enhance tumorigenesis. J Clin
Invest. 123:630–645. 2013.PubMed/NCBI
|
|
23
|
Kim J, Zhang Y, Skalski M, Hayes J, Kefas
B, Schiff D, Purow B, Parsons S, Lawler S and Abounader R:
microRNA-148a is a prognostic oncomiR that targets MIG6 and BIM to
regulate EGFR and apoptosis in glioblastoma. Cancer Res.
74:1541–1553. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bayrak OF, Gulluoglu S, Aydemir E, Ture U,
Acar H, Atalay B, Demir Z, Sevli S, Creighton CJ, Ittmann M, et al:
MicroRNA expression profiling reveals the potential function of
microRNA-31 in chordomas. J Neurooncol. 115:143–151. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Mishra PJ and Merlino G: MicroRNA
reexpression as differentiation therapy in cancer. J Clin Invest.
119:2119–2123. 2009.PubMed/NCBI
|