|
1
|
McCarthy N: Bladder cancer: Seemingly
similar. Nat Rev Cancer. 14:214–215. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Tomlinson D, Baldo O, Harnden P and
Knowles M: FGFR3 protein expression and its relationship to
mutation status and prognostic variables in bladder cancer. J
Pathol. 213:91–98. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Tomlinson DC, Hurst CD and Knowles MA:
Knockdown by shRNA identifies S249C mutant FGFR3 as a potential
therapeutic target in bladder cancer. Oncogene. 26:5889–5899. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Karashima T, Sweeney P, Kamat A, Huang S,
Kim SJ, Bar-Eli M, McConkey DJ and Dinney CP: Nuclear factor-kappaB
mediates angiogenesis and metastasis of human bladder cancer
through the regulation of interleukin-8. Clin Cancer Res.
9:2786–2797. 2003.PubMed/NCBI
|
|
6
|
Salazar L, Kashiwada T, Krejci P, Meyer
AN, Casale M, Hallowell M, Wilcox WR, Donoghue DJ and Thompson LM:
Fibroblast growth factor receptor 3 interacts with and activates
TGFβ-activated kinase 1 tyrosine phosphorylation and NFκB signaling
in multiple myeloma and bladder cancer. Plos One. 9:e864702014.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Reynolds A, Leake D, Boese Q, Scaringe S,
Marshall WS and Khvorova A: Rational siRNA design for RNA
interference. Nat Biotechnol. 22:326–330. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Folini M, Pennati M and Zaffaroni N: RNA
interference-mediated validation of genes involved in telomere
maintenance and evasion of apoptosis as cancer therapeutic targets.
Method Mol Biol. 487:303–330. 2009.
|
|
9
|
Barrett T and Edgar R: Gene expression
omnibus: Microarray data storage, submission, retrieval and
analysis. Methods Enzymol. 411:352–369. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Irizarry RA, Hobbs B, Collin F,
Beazer-Barclay YD, Antonellis KJ, Scherf U and Speed TP:
Exploration, normalization and summaries of high density
oligonucleotide array probe level data. Biostatistics. 4:249–264.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Gautier L, Cope L, Bolstad BM and Irizarry
RA: Affy-analysis of affymetrix genechip data at the probe level.
Bioinformatics. 20:307–315. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Smyth GK: Linear models and empirical
bayes methods for assessing differential expression in microarray
experiments. Stat Appl Genet Mol Biol. 3:Article32004.PubMed/NCBI
|
|
13
|
Cheng L, Lin H, Hu Y, Wang J and Yang Z:
Gene function prediction based on the gene ontology hierarchical
structure. Plos One. 9:e1071872014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Altermann E and Klaenhammer TR:
PathwayVoyager: Pathway mapping using the Kyoto encyclopedia of
genes and genomes (KEGG) database. BMC Genomics. 6:602005.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Dennis G Jr, Sherman BT, Hosack DA, Yang
J, Gao W, Lane HC and Lempicki RA: DAVID: Database for annotation,
visualization and integrated discovery. Genome Biol. 4:P32003.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Fogel GB, Weekes DG, Varga G, Dow ER,
Craven AM, Harlow HB, Su EW, Onyia JE and Su C: A statistical
analysis of the TRANSFAC database. Biosystems. 81:137–154. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhao M, Sun J and Zhao Z: TSGene: A web
resource for tumor suppressor genes. Nucleic Acids Res.
41:D970–D976. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chen JS, Hung WS, Chan HH, Tsai SJ and Sun
HS: In silico identification of oncogenic potential of fyn-related
kinase in hepatocellular carcinoma. Bioinformatics. 29:420–427.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C
and Jensen LJ: STRING v9.1: Protein-protein interaction networks,
with increased coverage and integration. Nucleic Acids Res.
41:D808–D815. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
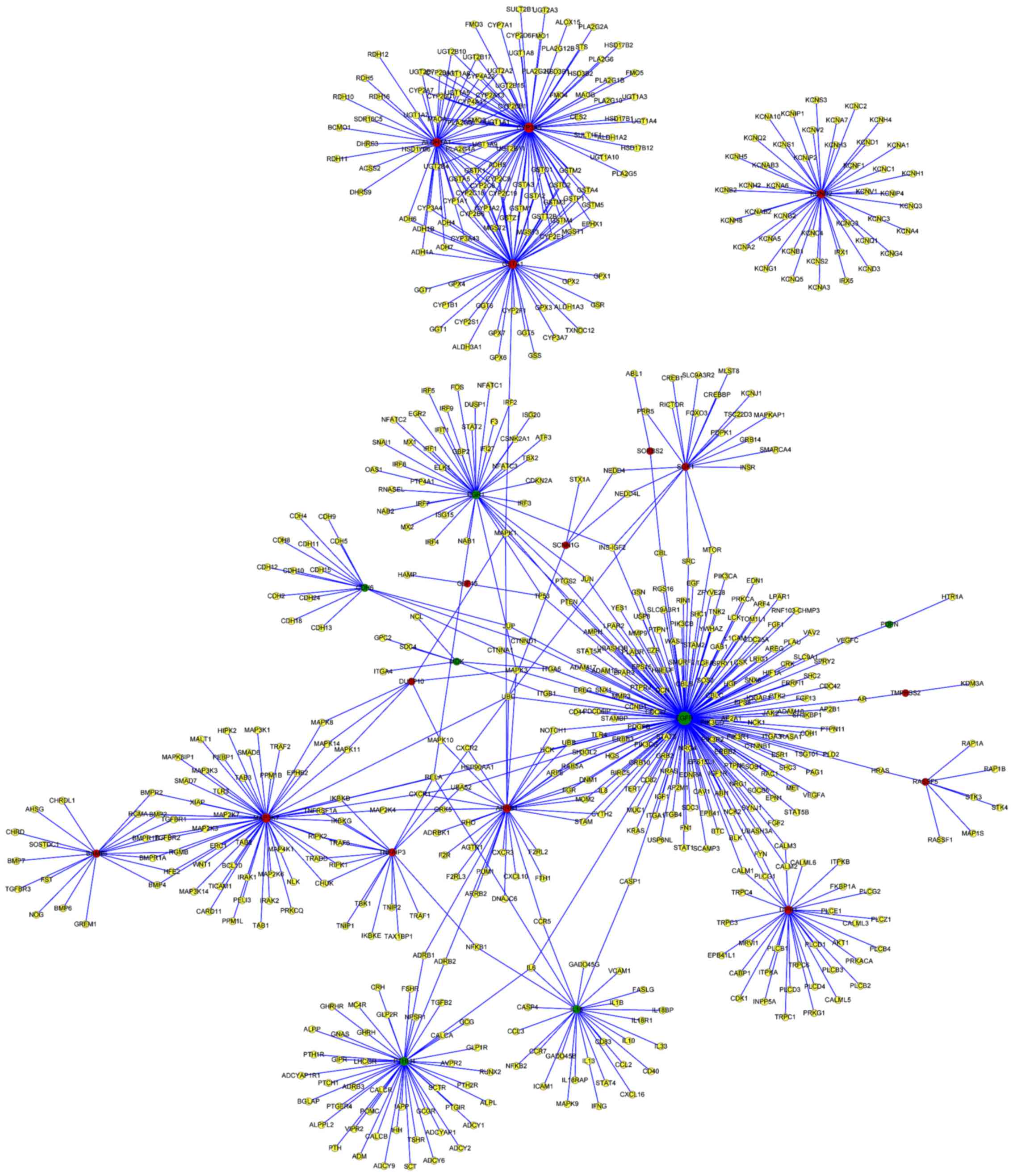
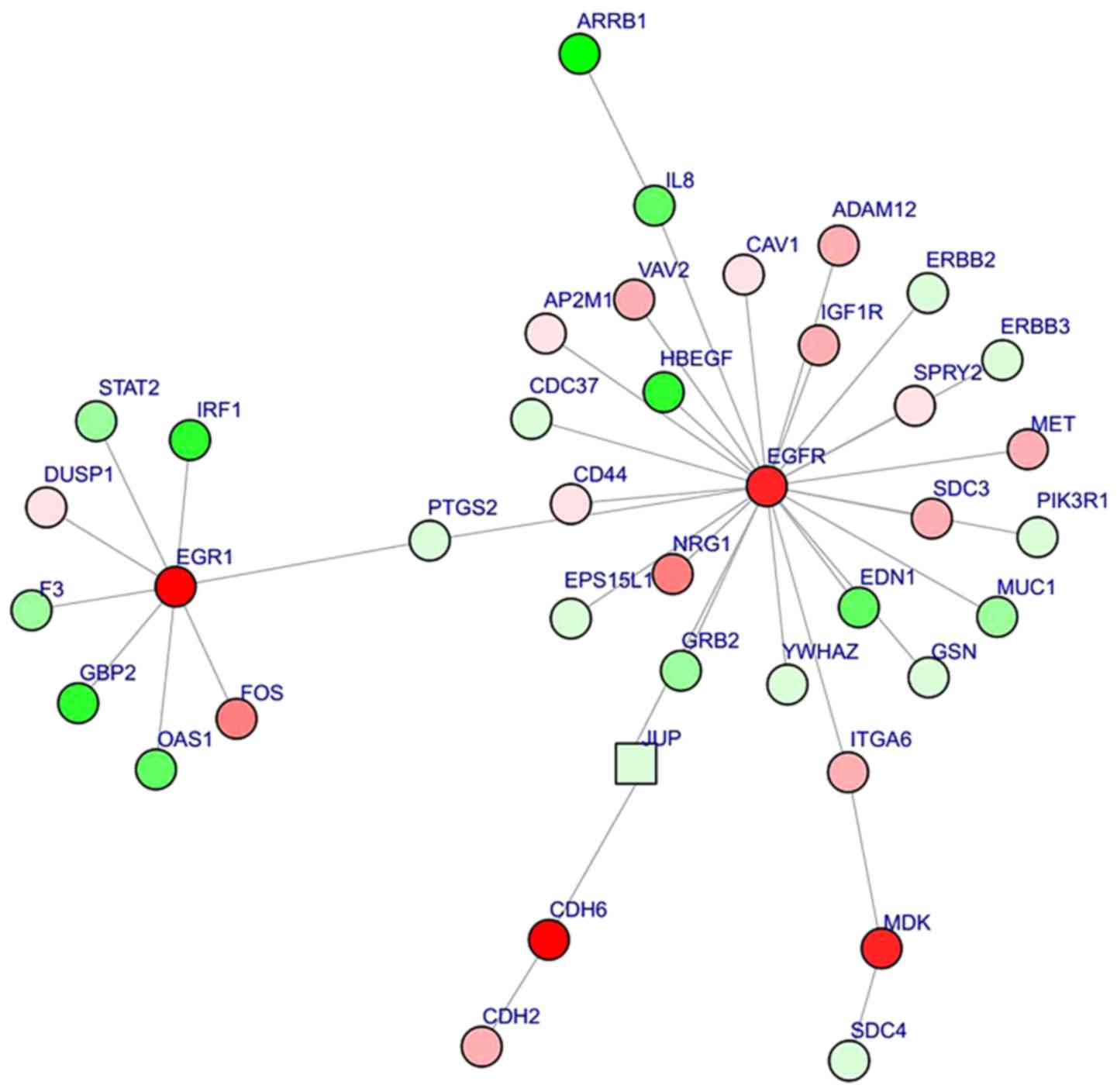
Kohl M, Wiese S and Warscheid B:
Cytoscape: Software for visualization and analysis of biological
networks. Methods Mol Biol. 696:291–303. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lah L, Podobnik B, Novak M, Korošec B,
Berne S, Vogelsang M, Kraševec N, Zupanec N, Stojan J, Bohlmann J
and Komel R: The versatility of the fungal cytochrome P450
monooxygenase system is instrumental in xenobiotic detoxification.
Mol Microbiol. 81:1374–1389. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Karpusas M, Axarli I, Chiniadis L,
Papakyriakou A, Bethanis K, Scopelitou K, Clonis YD and Labrou NE:
The interaction of the chemotherapeutic drug chlorambucil with
human glutathione transferase A1-1: Kinetic and structural
analysis. Plos One. 8:e563372013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Nebert DW and Dalton TP: The role of
cytochrome P450 enzymes in endogenous signalling pathways and
environmental carcinogenesis. Nat Rev Cancer. 6:947–960. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Fritz JH, Ferrero RL, Philpott DJ and
Girardin SE: Nod-like proteins in immunity, inflammation and
disease. Nat Immunol. 7:1250–1257. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Boorjian S, Tickoo SK, Mongan NP, Yu H,
Bok D, Rando RR, Nanus DM, Scherr DS and Gudas LJ: Reduced lecithin
retinol acyltransferase expression correlates with increased
pathologic tumor stage in bladder cancer. Clin Cancer Res.
10:3429–3437. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wincent J, Schoumans J and Anderlid BM: De
novo deletion of chromosome 11q13.4-q14.3 in a boy with
microcephaly, ptosis and developmental delay. Eur J Med Genet.
53:50–53. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ferris CD, Huganir RL, Supattapone S and
Snyder SH: Purified inositol 1,4,5-trisphosphate receptor mediates
calcium flux in reconstituted lipid vesicles. Nature. 342:87–89.
1989. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Boehning D, Patterson RL, Sedaghat L,
Glebova NO, Kurosaki T and Snyder SH: Cytochrome c binds to
inositol (1,4,5) trisphosphate receptors, amplifying
calcium-dependent apoptosis. Nat Cell Biol. 5:1051–1061. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Nicholson R, Gee J and Harper M: EGFR and
cancer prognosis. Eur J Cancer. 37:(Supply 4). S9–S15. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Black PC, Brown GA, Dinney CP, Kassouf W,
Inamoto T, Arora A, Gallagher D, Munsell MF, Bar-Eli M, McConkey DJ
and Adam L: Receptor heterodimerization: A new mechanism for
platelet-derived growth factor induced resistance to anti-epidermal
growth factor receptor therapy for bladder cancer. J Urol.
185:693–700. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chadalapaka G, Jutooru I, Burghardt R and
Safe S: Drugs that target specificity proteins downregulate
epidermal growth factor receptor in bladder cancer cells. Mol
Cancer Res. 8:739–750. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Urosevic J, Garcia-Albéniz X, Planet E,
Real S, Céspedes MV, Guiu M, Fernandez E, Bellmunt A, Gawrzak S,
Pavlovic M, et al: Colon cancer cells colonize the lung from
established liver metastases through p38 MAPK signalling and PTHLH.
Nat Cell Biol. 16:685–694. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zheng L, Pu J, Jiang G, Weng M, He J, Mei
H, Hou X and Tong Q: Abnormal expression of early growth response 1
in gastric cancer: Association with tumor invasion, metastasis and
heparanase transcription. Pathol Int. 60:268–277. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Yarden Y: The EGFR family and its ligands
in human cancer. signalling mechanisms and therapeutic
opportunities. Eur J Cancer. 37:(Supply 4). S3–S8. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yonesaka K, Zejnullahu K, Okamoto I, Satoh
T, Cappuzzo F, Souglakos J, Ercan D, Rogers A, Roncalli M, Takeda
M, et al: Activation of ERBB2 signaling causes resistance to the
EGFR-directed therapeutic antibody cetuximab. Sci Transl Med.
3:99ra862011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Li LY, Li EM, Wu ZY, Cao HH, Shen JH, Xu
XE, Chen B, Wu JY and Xu LY: Overexpression of GRB2 is correlated
with lymph node metastasis and poor prognosis in esophageal
squamous cell carcinoma. Int J Clin Exp Pathol. 7:3132–3140.
2014.PubMed/NCBI
|
|
37
|
Engelman JA: Targeting PI3K signalling in
cancer: Opportunities, challenges and limitations. Nat Rev Cancer.
9:550–562. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Urick ME, Rudd ML, Godwin AK, Sgroi D,
Merino M and Bell DW: PIK3R1 (p85α) is somatically mutated at high
frequency in primary endometrial cancer. Cancer Res. 71:4061–4067.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Cheung LW, Hennessy BT, Li J, Yu S, Myers
AP, Djordjevic B, Lu Y, Stemke-Hale K, Dyer MD, Zhang F, et al:
High frequency of PIK3R1 and PIK3R2 mutations in endometrial cancer
elucidates a novel mechanism for regulation of PTEN protein
stability. Cancer Discov. 1:170–185. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Yarden Y and Sliwkowski MX: Untangling the
ErbB signalling network. Nat Rev Mol Cell Biol. 2:127–137. 2001.
View Article : Google Scholar : PubMed/NCBI
|