|
1
|
Fitzgerald SP: Breast-cancer screening. N
Engl J Med. 366:191author reply 191. –192. 2012.PubMed/NCBI
|
|
2
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Colzani E, Liljegren A, Johansson AL,
Adolfsson J, Hellborg H, Hall PF and Czene K: Prognosis of patients
with breast cancer: Causes of death and effects of time since
diagnosis, age, and tumor characteristics. J Clin Oncol.
29:4014–4021. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Staiger C, Cadot S, Györffy B, Wessels LF
and Klau GW: Current composite-feature classification methods do
not outperform simple single-genes classifiers in breast cancer
prognosis. Front Genet. 4:2892013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Goswami CP and Nakshatri H: PROGgene: Gene
expression based survival analysis web application for multiple
cancers. J Clin Bioinforma. 3:222013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
van de Vijver MJ, He YD, van't Veer LJ,
Dai H, Hart AA, Voskuil DW, Schreiber GJ, Peterse JL, Roberts C,
Marton MJ, et al: A gene-expression signature as a predictor of
survival in breast cancer. N Engl J Med. 347:1999–2009. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Pawitan Y, Bjöhle J, Amler L, Borg AL,
Egyhazi S, Hall P, Han X, Holmberg L, Huang F, Klaar S, et al: Gene
expression profiling spares early breast cancer patients from
adjuvant therapy: Derived and validated in two population-based
cohorts. Breast Cancer Res. 7:R953–R964. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Paik S, Shak S, Tang G, Kim C, Baker J,
Cronin M, Baehner FL, Walker MG, Watson D, Park T, et al: A
multigene assay to predict recurrence of tamoxifen-treated,
node-negative breast cancer. N Engl J Med. 351:2817–2826. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang J, Webb-Robertson BJ, Matzke MM,
Varnum SM, Brown JN, Riensche RM, Adkins JN, Jacobs JM, Hoidal JR,
Scholand MB, et al: A semiautomated framework for integrating
expert knowledge into disease marker identification. Dis Markers.
35:513–523. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lee G, Singanamalli A, Wang H, Feldman MD,
Master SR, Shih NN, Spangler E, Rebbeck T, Tomaszewski JE and
Madabhushi A: Supervised multi-view canonical correlation analysis
(sMVCCA): Integrating histologic and proteomic features for
predicting recurrent prostate cancer. IEEE Trans Med Imaging.
34:284–297. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Davis S and Meltzer PS: GEOquery: A bridge
between the gene expression omnibus (GEO) and BioConductor.
Bioinformatics. 23:1846–1847. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang Y, Klijn JG, Zhang Y, Sieuwerts AM,
Look MP, Yang F, Talantov D, Timmermans M, Meijer-van Gelder ME, Yu
J, et al: Gene-expression profiles to predict distant metastasis of
lymph-node-negative primary breast cancer. Lancet. 365:671–679.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Miller LD, Smeds J, George J, Vega VB,
Vergara L, Ploner A, Pawitan Y, Hall P, Klaar S, Liu ET and Bergh
J: An expression signature for p53 status in human breast cancer
predicts mutation status, transcriptional effects and patient
survival. Proc Natl Acad Sci USA. 102:13550–13555. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ivshina AV, George J, Senko O, Mow B,
Putti TC, Smeds J, Lindahl T, Pawitan Y, Hall P, Nordgren H, et al:
Genetic reclassification of histologic grade delineates new
clinical subtypes of breast cancer. Cancer Res. 66:10292–10301.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chanrion M, Negre V, Fontaine H, Salvetat
N, Bibeau F, MacGrogan G, Mauriac L, Katsaros D, Molina F, Theillet
C and Darbon JM: A gene expression signature that can predict the
recurrence of tamoxifen-treated primary breast cancer. Clin Cancer
Res. 14:1744–1752. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Keshava Prasad TS, Goel R, Kandasamy K,
Keerthikumar S, Kumar S, Mathivanan S, Telikicherla D, Raju R,
Shafreen B, Venugopal A, et al: Human protein reference
database-2009 update. Nucleic Acids Res. 37:(Database issue).
D767–D772. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Nepusz T, Yu H and Paccanaro A: Detecting
overlapping protein complexes in protein-protein interaction
networks. Nat Methods. 9:471–472. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Friston KJ, Frith CD, Liddle PF and
Frackowiak RS: Functional connectivity: The principal-component
analysis of large (PET) data sets. J Cereb Blood Flow Metab.
13:5–14. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Gene Ontology Consortium: Gene Ontology
Consortium: Going forward. Nucleic Acids Res. 43:(Database issue).
D1049–D1056. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
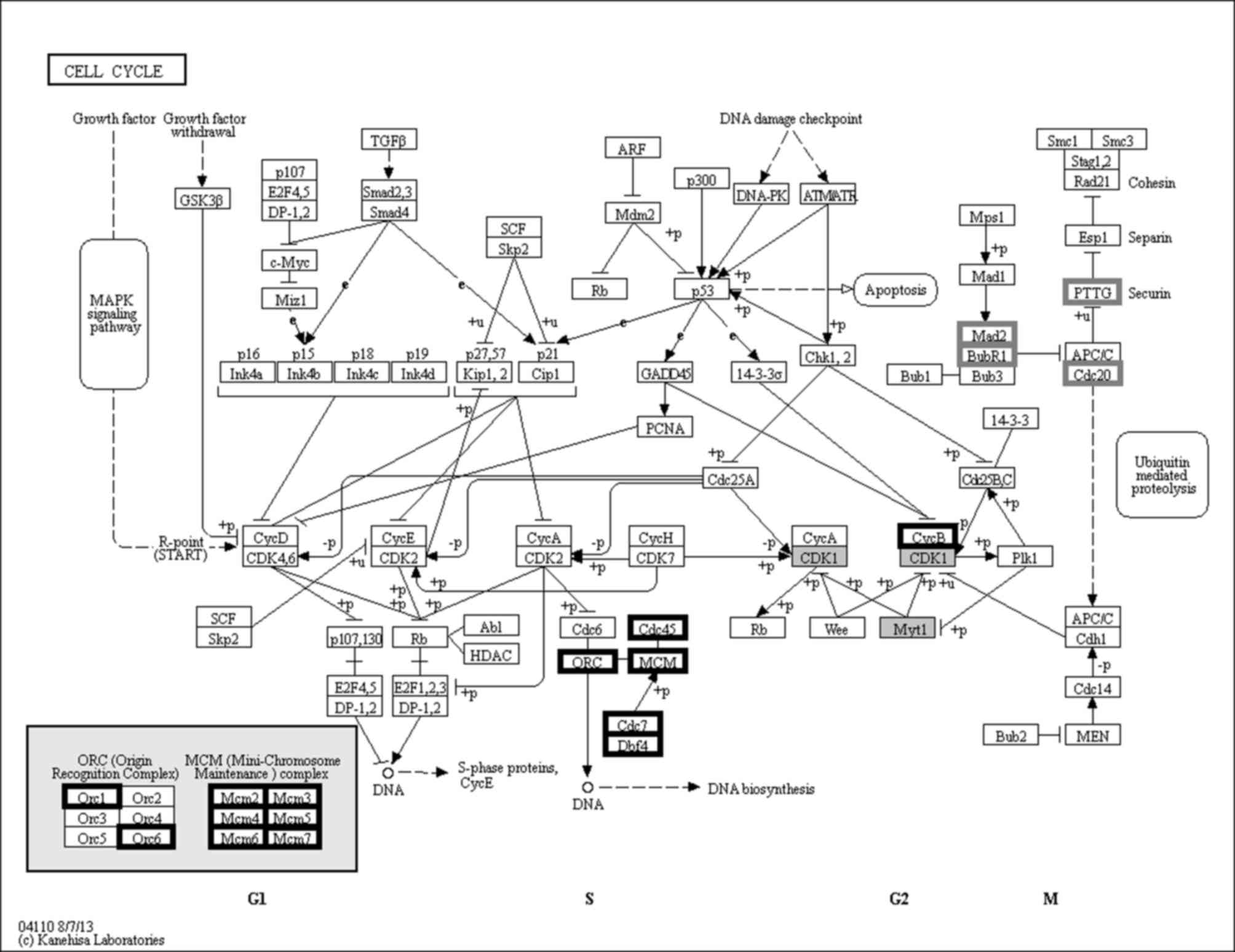
Kanehisa M, Goto S, Sato Y, Kawashima M,
Furumichi M and Tanabe M: Data, information, knowledge and
principle: Back to metabolism in KEGG. Nucleic Acids Res.
42:(Database issue). D199–D205. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Parmar MB, Aliabadi HM, Mahdipoor P,
Kucharski C, Maranchuk R, Hugh JC and Uludağ H: Targeting cell
cycle proteins in breast cancer cells with siRNA by using
lipid-substituted polyethylenimines. Front Bioeng Biotechnol.
3:142015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Weinberg RA: The retinoblastoma protein
and cell cycle control. Cell. 81:323–330. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Yu Z, Baserga R, Chen L, Wang C, Lisanti
MP and Pestell RG: microRNA, cell cycle, and human breast cancer.
Am J Pathol. 176:1058–1064. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Tenga MJ and Lazar IM: Proteomic snapshot
of breast cancer cell cycle: G1/S transition point. Proteomics.
13:48–60. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Achari C, Winslow S, Ceder Y and Larsson
C: Expression of miR-34c induces G2/M cell cycle arrest in breast
cancer cells. BMC Cancer. 14:5382014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kato T, Kameoka S, Kimura T, Tanaka S,
Nishikawa T and Kobayashi M: p53, mitosis, apoptosis and necrosis
as prognostic indicators of long-term survival in breast cancer.
Anticancer Res. 22:1105–1112. 2002.PubMed/NCBI
|
|
28
|
van Diest PJ, van der Wall E and Baak JP:
Prognostic value of proliferation in invasive breast cancer: A
review. J Clin Pathol. 57:675–681. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Königsberg R, Rögelsperger O, Jäger W,
Thalhammer T, Klimpfinger M, De Santis M, Hudec M and Dittrich C:
Cell cycle dysregulation influences survival in high risk breast
cancer patients. Cancer Invest. 26:734–740. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Louie MC, McClellan A, Siewit C and
Kawabata L: Estrogen receptor regulates E2F1 expression to mediate
tamoxifen resistance. Mol Cancer Res. 8:343–352. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kamel A, Mokhtar N, Elshakankiry N, Yassin
D, Elnahass Y, Zakarya O, Elbasmy A and Elmetenawy W: The
prognostic impact of some cell cycle regulatory proteins in
Egyptian breast cancer patients. J Egypt Natl Canc Inst. 18:93–102.
2006.PubMed/NCBI
|
|
32
|
Simon NE and Schwacha A: The Mcm2-7
replicative helicase: A promising chemotherapeutic target. Biomed
Res Int. 2014:5497192014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Coster G, Frigola J, Beuron F, Morris EP
and Diffley JF: Origin licensing requires ATP binding and
hydrolysis by the MCM replicative helicase. Mol Cell. 55:666–677.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Labib K, Tercero JA and Diffley JF:
Uninterrupted MCM2-7 function required for DNA replication fork
progression. Science. 288:1643–1647. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wan G, Hu X, Liu Y, Han C, Sood AK, Calin
GA, Zhang X and Lu X: A novel non-coding RNA lncRNA-JADE connects
DNA damage signalling to histone H4 acetylation. EMBO J.
32:2833–2847. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
de Munnik SA, Bicknell LS, Aftimos S,
Al-Aama JY, van Bever Y, Bober MB, Clayton-Smith J, Edrees AY,
Feingold M, Fryer A, et al: Meier-Gorlin syndrome
genotype-phenotype studies: 35 individuals with pre-replication
complex gene mutations and 10 without molecular diagnosis. Eur J
Hum Genet. 20:598–606. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
MacDermed DM, Khodarev NN, Pitroda SP,
Edwards DC, Pelizzari CA, Huang L, Kufe DW and Weichselbaum RR:
MUC1-associated proliferation signature predicts outcomes in lung
adenocarcinoma patients. BMC Med Genomics. 3:162010. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Wu G and Stein L: A network module-based
method for identifying cancer prognostic signatures. Genome Biol.
13:R1122012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Cronin M, Sangli C, Liu ML, Pho M, Dutta
D, Nguyen A, Jeong J, Wu J, Langone KC and Watson D: Analytical
validation of the Oncotype DX genomic diagnostic test for
recurrence prognosis and therapeutic response prediction in
node-negative, estrogen receptor-positive breast cancer. Clin Chem.
53:1084–1091. 2007. View Article : Google Scholar : PubMed/NCBI
|